Abstract
This study aimed to develop a model to predict the 5-year risk of developing end-stage renal disease (ESRD) in patients with type 2 diabetes mellitus (T2DM) using machine learning (ML). It also aimed to implement the developed algorithms into electronic medical records (EMR) system using Health Level Seven (HL7) Fast Healthcare Interoperability Resources (FHIR). The final dataset used for modeling included 19,159 patients. The medical data were engineered to generate various types of features that were input into the various ML classifiers. The classifier with the best performance was XGBoost, with an area under the receiver operator characteristics curve (AUROC) of 0.95 and area under the precision recall curve (AUPRC) of 0.79 using three-fold cross-validation, compared to other models such as logistic regression, random forest, and support vector machine (AUROC range, 0.929–0.943; AUPRC 0.765–0.792). Serum creatinine, serum albumin, the urine albumin-to-creatinine ratio, Charlson comorbidity index, estimated GFR, and medication days of insulin were features that were ranked high for the ESRD risk prediction. The algorithm was implemented in the EMR system using HL7 FHIR through an ML-dedicated server that preprocessed unstructured data and trained updated data.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Introduction
Type 2 diabetes mellitus (T2DM) is known to be a leading cause of end-stage renal disease (ESRD) worldwide1,2. ESRD is the final and permanent stage of chronic kidney disease (CKD), where kidney function has declined to the point which the kidneys cannot function any longer on their own3. As the population of individuals with T2DM is increasing rapidly, the population of individuals with ESRD is also accelerating4,5,6. In Korea, the prevalence of T2DM increased over 17 years from 2001 (8.6%) to 2018 (13.8%) in adults ≥ 30 years of age7. This prevalence is not much different from that of adults in the United States8. Furthermore, although many anti-diabetic and anti-hypertensive medications have been developed, the prevalence of ESRD has not decreased9.
Recent studies have proven that at least 90% of patients with diabetic kidney disease (DKD) are at a higher risk of mortality because of comorbidities such as cardiovascular disease and kidney failure10. Furthermore, in contrast to other diabetic complications, mortality associated with renal complications is continuously increasing11. Therefore, there is a need to predict and prevent ESRD, which is the most severe stage of DKD, to slow or stop the progression of DKD by performing early diagnosis and treatment, thereby minimizing the medical costs associated with kidney failure treatment.
Many studies have focused on determining the predictive factors for the development of DKD, including clinical markers such as baseline values of the glomerular filtration rate (GFR), systolic blood pressure, fasting blood glucose, triglycerides12, and genetic markers, such as the angiotensin-converting enzyme genotype13. Additionally, various risk prediction models have been developed that incorporate these known risk factors14,15,16,17,18. These studies successfully presented the risk prediction for DKD. For example, one study conducted in China with 8.3 years of follow-up predicted the 3-year, 5-year, and 8-year risk of ESRD in type 2 diabetes patients with good accuracy (area under the receiver-operating characteristic [AUROC] curve of 0.90, 0.86, and 0.81, respectively) using discriminatory values such as age, sex, age at the time of diabetes onset, creatinine, albuminuria, variations in HbA1c, combined statuses of hypertension, diabetes, and hyperlipidemia in risk-scoring systems14,15,16.
However, variables other than the known risk factors can also influence the development of ESRD. Thus, machine learning using various risk factors should be adopted to improve the accuracy of the predictive model for ESRD, as in the case of previous studies where machine learning was applied to increase accuracy in diagnosing T2DM19,20. To our knowledge, no previous study has used machine learning to develop a patient-level ESRD risk prediction model. Furthermore, any attempt to deploy such a prediction model in the real-world clinical setting such that it is of service to patients and clinicians has been deficient. Integrating the prediction model into the electronic medical records (EMR) system would provide additional benefits such as allowing the identification of patients at a high risk of ESRD in busy hospital environments.
In this study, we aimed to develop a patient-level prediction model for ESRD in adults with type 2 diabetes mellitus that presents a risk score for developing ESRD within 5 years. We also aimed to distinguish the model from similar pre-existing tools, such as the Kidney Failure Risk Equation and the tools from the Chronic Kidney Disease Prognosis Consortium21,22. Furthermore, we aimed to create a model that is applicable in actual clinical practice in a multitude of hospitals and can be implemented in the EMR system.
Results
Clinical characteristics of the patients
The baseline characteristics of 19,159 individuals in the cohort are presented in Table 1. During 16 years of follow-up, 1,583 patients (8.3%) developed ESRD. These patients were older and had higher blood pressure and poor lipid profiles compared to individuals without ESRD. However, patients with ESRD were less obese than their counterparts.
Model discrimination and calibration performance
Our model had good discriminatory power, which indicates how well our model discriminates between patients with and without ESRD, with an AUROC curve of 0.947 and area under precision recall curve (AUPRC) of 0.785 (Table 2).
The calibration performance was also assessed with the calibration plot23. The plot was created by discretizing the [0, 1] interval into 10 uniform bins. For each bin, the mean predicted probability and true fraction of positive cases were plotted on the x-axis and y-axis, respectively. A perfectly calibrated model is represented by a diagonal line. If the model was above the diagonal line, it indicated that the model underestimated the risks. Similarly, if the model was below the diagonal line, it was an indication of an overestimation of risks. We observed that our model moderately overestimated higher risk score groups (Fig. 1).
Model comparison
We compared the discriminatory power of the XGBoost model against other types of models, such as linear regression, support vector machine, decision tree, and random forest models (Table 3 and Supplementary Fig. 1). Our XGBoost model had the best discrimination power (AUROC curve, 0.947; AUPRC, 0.792) compared to other models (AUROC curves range, 0.929–0.943; AUPRC range, 0.765–0.792).
Cost–benefit analysis (decision curve analysis)
To assess the recall of our model, which indicates the proportion of true-positive cases compared to all positive cases (true-positive and false-negative cases), we plotted the recall curve (Fig. 2a), which shows the recall rate as a function of the number of cases treated.
Similarly, we also assessed the precision of the model, which is the proportion of true-positive cases compared to all predicted cases, by ranking predicted cases in descending order according to their predicted probabilities (Fig. 2b).
To evaluate our model performance from the perspective of clinical value, we conducted a decision curve analysis (Fig. 2c). The main advantage of the decision curve analysis is that it incorporates clinical consequences into the model evaluation and does not require additional data, such as an explicit assessment of health outcomes or treatment-related costs. Instead, it considers a threshold probability as an informative indicator of relative harms of a false-positive and a false-negative prediction24,25. We also analyzed changes of precision and recall across a range of probability thresholds (Fig. 2d).
Model explainability
We used the Shapley Additive Explanations (SHAP) analysis to interpret the data acquired from the machine learning process of the XGBoost model and evaluate the importance of the individual features of the prediction of ESRD. SHAP is a framework that interprets predictions that a machine learning model conducted. It calculates the contribution of each feature to the prediction and assigns an importance value to each feature depending on the result of the calculation26. Figure 3 shows the magnitude and direction of the contribution of each feature compared to the average model prediction.
SHAP summary plot. If a feature is located on the upper side of this figure, then it implies a higher contribution of the feature to the prediction. More specifically, each dot represents the data of each patient, and the color of the dot indicates whether the respective feature value is low or high (as shown on the y-axis on the right). The location of the dot indicates whether the feature increases (right) or decreases (left) the risk prediction. The farther a dot is from 0, the greater the contribution to the prediction. SHAP Shapley Additive Explanations.
To assess the stability of the importance of these features, we checked the ranges and standard deviations of their rankings with regard to different index dates (Supplementary Fig. 2). According to the results of iterative experiments, the features with a high rank with the greatest importance did not show much distinction from those with other ranks. Serum creatinine, serum albumin, the urine albumin-to-creatinine ratio, Charlson comorbidity index, estimated GFR, and medication days of insulin were features that were ranked high for the ESRD risk prediction27. However, our prediction model incorporated new parameters such as blood albumin level, and non-invasive markers of hepatic fibrosis such as nonalcoholic fatty liver disease, fibrosis score, and fibrosis-4 index.
Implementation of the machine learning-based clinical decision support system
The developed model was implemented with a dedicated server for the Machine Learning-based Clinical Decision Support System (Supplementary Fig. 3). The server extracted structured data through the Fast Healthcare Interoperability Resources server from EMR, such as visit history, medication history, laboratory values, and vital signs (Supplementary Table 2). For the extraction of unstructured data, such as smoking history, reading of imaging studies, and results of electrocardiography, the server directly accessed the EMR and preprocessed the unstructured data into structured input features for the calculation of the prediction model based on the XGBoost algorithm.
Dashboard
A prototype of a comprehensive dashboard for the Machine Learning-based Clinical Decision Support System was designed to provide the results of the prediction algorithm and related test results of an individual patient. The ESRD risk determined by the prediction model was presented with the SHAP analysis to enable users to identify modifiable risk factors among the high-ranking input features according to the SHAP results (Supplementary Fig. 4).
Discussion
We successfully developed a 5-year ESRD risk prediction model for type 2 diabetes mellitus using a machine learning algorithm based on the medical data of the study cohort consisting of 19,159 patients. Among various machine learning methods, the XGBoost classifier showed the best discriminatory performance when processing medical data. Additionally, we applied the SHAP analysis to evaluate the relative importance of each feature, which could provide specific medical information to physicians.
Current treatment guidelines for chronic kidney disease in patients with diabetes have suggested stratifying the patient’s risk according to GFR and albuminuria categories28. However, this classification is too simple to correctly predict the individual risk of ESRD. Precise prediction of the prognosis of renal function is necessary when deciding whether to refer the patient to a nephrologist, preparing a long-term plan (e.g., renal transplantation), and providing appropriate medical intervention. Our model provided the 5-year risk of ESRD with good discrimination power. Additionally, the risk factors for ESRD that we identified in our model are well-known, which means our model is clinically explainable. A decreased serum albumin level was one of the strongest predictive factors in our model. The albumin level might represent the patient’s nutritional status and could be a marker for the poor prognosis of chronic kidney disease29. Systemic inflammation in a critically ill patient is known to cause altered albumin homeostasis and lead to hypoalbuminemia30. Additionally, hepatic fibrosis indices were also introduced in our model, which could be related to advanced stages of diabetes complications31,32. Nonalcoholic fatty liver disease has been proven to accelerate the decline in kidney function in chronic kidney disease patients through the activation of a pathway that enhances the transcription of pro-inflammatory genes and amplification of immunologic inflammatory responses33.
The high AUROC curve showed that our model was sufficiently able to distinguish between positive and negative cases. The high AUPRC of our model also provided significant benefits because it was much higher than the prevalence of ESRD in the training data. The recall rate of our model with a default threshold of 0.5 (i.e., treating patients when the risk score is higher than 0.5) had a 95% confidence interval between 0.62 and 0.64. If a physician provides treatment for patients with a risk score of more than 0.5, then this plan would treat 62–64% of patients who will develop ESRD. Physicians can always adjust the threshold based on their case and depending on the desired levels of recall and precision (Fig. 2d). For example, if physicians would like to increase the coverage (recall) to 80%, they may use a risk score cutoff of 0.1 while still maintaining a precision value as high as 0.6.
Similarly, the precision of our model had a 95% confidence interval between 0.82 and 0.84. Therefore, if the model predicts a patient with T2DM will have ESRD, 82–84% of them will actually have ESRD in the future.
Compared to a conventional model from cox regression by Tangri et al.34, our model has several advantages. First, our model predicts ESRD, not CKD stages 3 to 5 as in the conventional model, which means our model targets most severe case of CKD in T2DM patients. Second, new predictors such as serum albumin and makers of fatty livers, which were ranked high in the SHAP analysis. Third, our model can be used directly with EHR systems using FHIR resources.
Although our model provided significant improvements in terms of discrimination and calibration performance for ESRD risk prediction, there are several areas that can be improved. First, several features related to disease or medication were engineered as binary indicators. However, their effects can vary depending on the severity of other comorbidities or the dosage of medication, which were not fully captured in our model. Second, although the classification modeling approach using XGBoost enabled us to handle missing values, the drawback was that the model required every patient to have sufficient observation periods (i.e., 5 years) to correctly identify outcome labels. Therefore, the most recent index date that could be used in our model was at least 5 years before the last date of the available data; consequently, almost 10 years of EMR data could not be used in our model. Third, no imputations were performed to handle missing data, which could have impacted the quality of the modeling process. Future work should consider incorporating data imputation. Fourth, we did not reflect the definition of a period in which of eGFR < 15 ml/min/1.73 m2 and the need for dialysis persist for more than 3 months to accurately define ERSD. It might overestimate the incidence of patients with ESRD in the study cohort. Finally, we did not validate our model for the independent data set and prove the clinical effectiveness of our model Therefore, we cannot generalize our results to other environments. Further study is needed to evaluate accuracy and clinical effectiveness of the developed model in other environments.
Despite these limitations, our model provides several meaningful clinical and practical implications. The inclusion of additional data items from the EMR system contributed to its better performance. Previous studies could not find significant improvements in model performance by incorporating more variables beyond the traditional chronic kidney disease risk model variables14. It would be beneficial for future studies to assess the extent to which additional EMR data items contributed to the improvement in performance. Our model will be a great resource for physicians to assess the developmental and progression risk of ESRD within the next 5 years for type 2 diabetes mellitus patients, and it will help them make appropriate treatment decisions to prevent or slow the progression of ESRD.
Methods
Development of models
The development of the ESRD prediction model was accomplished through six steps. First, the study cohort was selected from the initial dataset by applying the inclusion and exclusion criteria. Second, we assigned a random index date for every patient in the cohort. Third, to identify patients with ESRD, a primary composite outcome of the study, we labeled the outcome and prediction timeline according to the clinical criteria described. Fourth, as a result of processing the time-series data of patients using the eight feature generators, 49 features were retained to compose the ESRD prediction model. Subsequently, we trained and validated the model through k-fold cross-validation. Finally, we repeated step 1 through step 4 N times; therefore, the performance of the prediction model could be based on the average of K*N iterations.
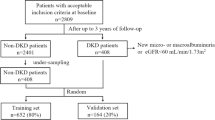
To select the study cohort, we used the medical data extracted from the EMR system of Seoul National University Bundang Hospital, a tertiary academic hospital in South Korea. Seoul National University Bundang Hospital developed and adopted an electronic health system in 2003. Of 61,936 patients from the initial dataset, 60,909 patients remained after excluding those without at least three outpatient visit records. Thereafter, we continued constructing the final dataset by applying our inclusion criteria and exclusion criterion for the ESRD prediction model.
Our inclusion criteria were as follows: age between 18 and 90 years on the index date, no history of ESRD or dialysis before the index date, more than one type 2 diabetes mellitus medication record before the index date, and a history of more than 5 years of observation after the index date or first diagnosis of ESRD within the prediction window after the index date. Our exclusion criterion was a diagnosis of ESRD within 14 days of the first visit. As a result of including only patients who satisfied these criteria, we obtained a final cohort of 19,159 patients (Fig. 4).
The primary composite outcome of the study was ESRD. To identify patients with this comorbidity, we precisely defined ESRD as a diagnosis including the ICD-10 diagnosis codes N18.5 and N18.6, a history of dialysis treatment, a history of renal transplantation with ICD-9 operation codes 55.6 and 55.69, a history of continuous ambulatory peritoneal dialysis catheter insertion with ICD-9 operation codes 38.95 and 39.43, and an estimated GFR < 15 mL/min/1.73 m235,36.
Next, for each patient, we randomly selected one index date to construct a cohort. For each patient in the cohort with a selected index date, we labeled the outcome as 1 if the patient developed the comorbidity of interest during the prediction window (5 years); in other cases, the outcome was labeled as 0. Consider the scenarios depicted in Fig. 5. The horizontal line represents the timeline of a patient’s hospital visits, which are denoted as v1, v2, and so on. Both scenarios in Fig. 5 represent the same patient, who was diagnosed with the comorbidity during v6. Depending on the selection of the index date for this patient, the outcome can be labeled as 0 or 1. If v3 was considered as the index date, then it was considered that this patient will not develop the comorbidity because the comorbidity was not observed within the prediction window for the given time at risk. However, if we considered v5 as the index date, then it was considered that this patient will develop the comorbidity because the prediction window includes the visit when the comorbidity was diagnosed.
We used the eight feature generators to process the time-series data of 19,159 patients. Although 52 features were initially generated, three (cacs_ewma, imt_max, and lipoprotein_ewma) were excluded because their values for more than 90% of patients were missing. Therefore, a total of 49 features were obtained for our ESRD prediction model. For handling time depending variables, we used most recent value, exponential weighted moving average value, max value, or length of records depending on the characteristics of the variable. Supplementary Table 1 shows the types of feature generators we used and the actual examples of predictors, as well as how to handle multiple measures.
New predictors were included for model development. Albumin was included because lower pre-ESRD serum albumin was associated with the incidence of ESRD in previous studies27,37. Markers of fatty livers disease were included because non-alcoholic fatty liver disease was related to the increased risk of chronic kidney disease in previous studies38,39.
For model training, we split the data into training and validation sets. We trained the binary XGBoost classification models using the training set. Then, we calculated the model performance using the validation set by repeating this procedure k times (i.e., k-fold cross-validation).
The model was evaluated using N-epoch K-fold cross-validation, which first used regular K-fold validation (step 4) and then repeated step 1 to step 4 N times. The final performance was the average of these K*N iterations.
Ethics
This research was approved by the Institutional Review Board of Human Research of Seoul National University Bundang Hospital. Informed consent was waived because of the retrospective nature of the research and the analysis used deidentified clinical data (B-1904-535-001). The present research was conducted in accordance with the Declaration of Helsinki.
Data availability
Data is not available to public due to the regulation of IRB in SNUBH. S.Y.J. should be contacted if someone wants to ask a question about the data from this study.
References
Benjamin, O. & Lappin, S. L. in StatPearls [Internet] (StatPearls Publishing, 2021).
Ghaderian, S. B., Hayati, F., Shayanpour, S. & Mousavi, S. S. B. Diabetes and end-stage renal disease: A review article on new concepts. J. Renal Inj. Prev. 4, 28 (2015).
Abbasi, M. A., Chertow, G. M. & Hall, Y. N. End-stage renal disease. BMJ Clin. Evid. 2010 (2010).
Nasri, H. & Rafieian-Kopaei, M. Diabetes mellitus and renal failure: Prevention and management. J. Res. Med. Sci. 20, 1112 (2015).
Lim, A. K. Diabetic nephropathy–complications and treatment. Int. J. Nephrol. Renov. Dis. 7, 361 (2014).
Narres, M. et al. The incidence of end-stage renal disease in the diabetic (compared to the non-diabetic) population: A systematic review. PLoS ONE 11, e0147329 (2016).
Jung, C.-H. et al. Diabetes fact sheets in Korea, 2020: An appraisal of current status. Diabetes Metab. J. 45, 1–10 (2021).
Lin, J. et al. Projection of the future diabetes burden in the United States through 2060. Popul. Health Metrics 16, 1–9 (2018).
Gregg, E. W., Hora, I. & Benoit, S. R. Resurgence in diabetes-related complications. JAMA 321, 1867–1868 (2019).
Foundation, N. K. CKDinform, https://www.kidney.org/CKDinform (2022).
Ling, W. et al. Global trend of diabetes mortality attributed to vascular complications, 2000–2016. Cardiovasc. Diabetol. 19, 1–12 (2020).
Rodriguez-Romero, V. et al. Prediction of nephropathy in type 2 diabetes: An analysis of the ACCORD trial applying machine learning techniques. Clin. Transl. Sci. 12, 519–528 (2019).
William, J., Hogan, D. & Batlle, D. Predicting the development of diabetic nephropathy and its progression. Adv. Chronic Kidney Dis. 12, 202–211 (2005).
Lin, C.-C. et al. Development and validation of a risk prediction model for end-stage renal disease in patients with type 2 diabetes. Sci. Rep. 7, 1–13 (2017).
Jardine, M. J. et al. Prediction of kidney-related outcomes in patients with type 2 diabetes. Am. J. Kidney Dis. 60, 770–778 (2012).
Desai, A. S. et al. Association between cardiac biomarkers and the development of ESRD in patients with type 2 diabetes mellitus, anemia, and CKD. Am. J. Kidney Dis. 58, 717–728 (2011).
Elley, C. R. et al. Derivation and validation of a renal risk score for people with type 2 diabetes. Diabetes Care 36, 3113–3120 (2013).
Wan, E. Y. F. et al. Prediction of new onset of end stage renal disease in Chinese patients with type 2 diabetes mellitus: A population-based retrospective cohort study. BMC Nephrol. 18, 1–9 (2017).
Balasubramaniyan, S., Jeyakumar, V. & Nachimuthu, D. S. Panoramic tongue imaging and deep convolutional machine learning model for diabetes diagnosis in humans. Sci. Rep. 12, 1–18 (2022).
Hasan, M. K., Alam, M. A., Das, D., Hossain, E. & Hasan, M. Diabetes prediction using ensembling of different machine learning classifiers. IEEE Access 8, 76516–76531 (2020).
Tangri, N. et al. Multinational assessment of accuracy of equations for predicting risk of kidney failure: A meta-analysis. JAMA 315, 164–174 (2016).
Grams, M. E. et al. Predicting timing of clinical outcomes in patients with chronic kidney disease and severely decreased glomerular filtration rate. Kidney Int. 93, 1442–1451 (2018).
Bella, A., Ferri, C., Hernández-Orallo, J. & Ramírez-Quintana, M. J. in Handbook of Research on Machine Learning Applications and Trends: Algorithms, Methods, and Techniques 128–146 (IGI Global, 2010).
Osawa, I., Goto, T., Yamamoto, Y. & Tsugawa, Y. Machine-learning-based prediction models for high-need high-cost patients using nationwide clinical and claims data. NPJ Digit. Med. 3, 1–9 (2020).
Vickers, A. J. & Elkin, E. B. Decision curve analysis: A novel method for evaluating prediction models. Med. Decis. Mak. 26, 565–574 (2006).
Lundberg, S. M. & Lee, S.-I. A unified approach to interpreting model predictions. Adv. Neural Inf. Process. Syst. 30 (2017).
Walther, C. P. et al. Serum albumin concentration and risk of end-stage renal disease: The REGARDS study. Nephrol. Dial. Transplant. 33, 1770–1777 (2018).
de Boer, I. H. & Rossing, P. Kidney Disease: Improving Global Outcomes (KDIGO) Diabetes Work Group. KDIGO 2020 Clinical Practice Guideline for Diabetes Management in Chronic Kidney Disease Introduction. Kidney Int. 98, S20 (2020).
Kikuchi, H. et al. Combination of low body mass index and serum albumin level is associated with chronic kidney disease progression: The chronic kidney disease-research of outcomes in treatment and epidemiology (CKD-ROUTE) study. Clin. Exp. Nephrol. 21, 55–62 (2017).
Haller, C. Hypoalbuminemia in renal failure: Pathogenesis and therapeutic considerations. Kidney Blood Press. Res. 28, 307–310 (2005).
Han, E., Kim, M. K., Jang, B. K. & Kim, H. S. Albuminuria is associated with steatosis burden in patients with type 2 diabetes mellitus and nonalcoholic fatty liver disease. Diabetes Metab. J. 45, 698–707 (2021).
Targher, G. et al. Non-alcoholic fatty liver disease is independently associated with an increased prevalence of chronic kidney disease and proliferative/laser-treated retinopathy in type 2 diabetic patients. Diabetologia 51, 444–450 (2008).
Jang, H. R. et al. Nonalcoholic fatty liver disease accelerates kidney function decline in patients with chronic kidney disease: A cohort study. Sci. Rep. 8, 1–9 (2018).
Tangri, N. et al. A predictive model for progression of chronic kidney disease to kidney failure. JAMA 305, 1553–1559 (2011).
Perkovic, V. et al. Canagliflozin and renal outcomes in type 2 diabetes and nephropathy. N. Engl. J. Med. 380, 2295–2306 (2019).
Vistisen, D. et al. A validated prediction model for end-stage kidney disease in type 1 diabetes. Diabetes Care 44, 901–907 (2021).
Kawai, Y. et al. Association between serum albumin level and incidence of end-stage renal disease in patients with Immunoglobulin A nephropathy: A possible role of albumin as an antioxidant agent. PLoS ONE 13, e0196655 (2018).
Kaps, L. et al. Non-alcoholic fatty liver disease increases the risk of incident chronic kidney disease. United Eur. Gastroenterol. J. 8, 942–948 (2020).
Targher, G., Chonchol, M., Zoppini, G., Abaterusso, C. & Bonora, E. Risk of chronic kidney disease in patients with non-alcoholic fatty liver disease: Is there a link?. J. Hepatol. 54, 1020–1029 (2011).
Acknowledgements
We thank Yong Jun Kim, Andrew Young, Vikram Pawar, Vy Tran, Gary Zakon, and Adlar Jeewook Kim for their guidance and feedback regarding model development and implementation.
Author information
Authors and Affiliations
Contributions
S.W. and J.H. drafted the entire manuscript as the first authors. S.Y, S.L., H.H., H.Y.L., and H.H. contributed to the discussion of the results. S.Y.J. and T.J.O. supervised the entire process as the corresponding authors.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Wang, S., Han, J., Jung, S.Y. et al. Development and implementation of patient-level prediction models of end-stage renal disease for type 2 diabetes patients using fast healthcare interoperability resources. Sci Rep 12, 11232 (2022). https://doi.org/10.1038/s41598-022-15036-6
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-022-15036-6
- Springer Nature Limited