Abstract
KRAS mutation is responsible for 40–50% of colorectal cancers (CRCs). RNA-seq data and bioinformatics methods were used to analyze the transcriptional profiles of KRAS mutant (mtKRAS) in comparison with the wild-type (wtKRAS) cell lines, followed by in-silico and quantitative real-time PCR (qPCR) validations. Gene set enrichment analysis showed overrepresentation of KRAS signaling as an oncogenic signature in mtKRAS. Gene ontology and pathway analyses on 600 differentially-expressed genes (DEGs) indicated their major involvement in the cancer-associated signal transduction pathways. Significant hub genes were identified through analyzing PPI network, with the highest node degree for PTPRC. The evaluation of the interaction between co-expressed DEGs and lncRNAs revealed 12 differentially-expressed lncRNAs which potentially regulate the genes majorly enriched in Rap1 and RAS signaling pathways. The results of the qPCR showed the overexpression of PPARG and PTGS2, and downregulation of PTPRC in mtKRAS cells compared to the wtKRAS one, which confirming the outputs of RNA-seq analysis. Further, significant upregualtion of miR-23b was observed in wtKRAS cells. The comparison between the expression level of hub genes and TFs with expression data of CRC tissue samples deposited in TCGA databank confirmed them as distinct biomarkers for the discrimination of normal and tumor patient samples. Survival analysis revealed the significant prognostic value for some of the hub genes, TFs, and lncRNAs. The results of the present study can extend the vision on the molecular mechanisms involved in KRAS-driven CRC pathogenesis.
Similar content being viewed by others
Introduction
According to the GLOBOCAN database, colorectal cancer (CRC) is known as one of the most prevalent malignancies with 9.2% of mortality in 2018, which ranks as the second leading cause of the cancer-induced mortality worldwide1. It has been estimated that 45% of sporadic CRCs frequently arises from preneoplastic lesions through KRAS proto-oncogene mutation2. It encodes a small guanosine triphosphate (GTP)/guanosine diphosphate (GDP) binding protein involved in regulating many cellular responses to several extracellular stimuli by activating over 20 known downstream effectors3,4. In addition, the RAS-specific guanine nucleotide exchange factors (GEFs) and guanosine triphosphate hydrolase (GTPase)-activating proteins (GAPs) tightly regulate cycling between an inactive GDP-bound (“off”) and an active GTP-bound (“on”) states of wild-type KRAS (wtKRAS) protein5,6. Mutation in KRAS proto-oncogene results in conformational changes, leading to its insensitivity to GAPs and many different effects such as cell growth promotion, cell transformation, angiogenesis, migration, and apoptosis suppression7. Therefore, KRAS mutation in CRCs is associated with more reduced survival, increased tumor aggressiveness, and resistance to anti-epidermal growth factor receptor (EGFR) targeted-therapies8,9. Following the discovery of an association between the mutant forms of RAS protein with human cancers, numerous studies have been conducted on inhibiting oncogenic RAS signaling for cancer treatment10,11. So far, the specific inhibition of oncogenic RAS has not been clinically approved despite extensive efforts on understanding the mechanisms of regulation, intracellular trafficking, and RAS protein signaling activity. Due to the high prevalence and mortality of KRAS mutant CRCs, studying potential molecular biomarkers and mechanisms to improve patient outcomes is becoming increasingly important.
The hundreds of genes, long non-coding RNAs (lncRNAs), and microRNAs (miRs) involved in various signaling pathways, molecular functions, and biological processes have been identified through applying the gene microarray and next-generation RNA sequencing (RNA-seq) which are high-throughput platforms for analyzing gene expression12,13. Somatic mutations cause a series of downstream, secondary alterations in the transcriptome, leading to the large-scale disturbances of transcriptional profiles in most cancers14,15. For instance, microarray analysis of CRCs with KRAS and EGFR mutations, demonstrated that mutation status leads to the dysregulated transcriptional landscape of the cells16. Further, the assessment of differential gene expression signatures revealed an alteration in the signaling pathways such as Wnt, NF-kappa B, and the TGF-beta, and metabolic ones, along with changing the KRAS-related signaling pathways, between CRCs with and without KRAS mutations17,18. LncRNAs and miRs, as new and fundamental transcriptional and post-transcriptional regulators with a pivotal role in tumorigenesis and malignant tumor metastasis, have been studied widely19,20. Furthermore, the contribution of 282 lncRNAs to CRC heterogeneity has been recognized by monitoring the expression of 4898 lncRNA genes across 566 CRC samples14. Some researchers analyzed differential miR expression in colorectal tumors with KRAS and BRAF mutations15,21. Activating mutations in KRAS changes the transcriptional profiles of CRC cells significantly. Thus, the further characterization of CRC exploring all specific molecular and cellular aberrations, along with KRAS mutation, seems to be critical for detecting specific therapeutic opportunities17.
In the present study, RNA-seq data were downloaded from the Sequence Read Archive database (SRA) to identify the genes and lncRNAs having differential expression between mtKRAS and wtKRAS CRC cell lines. For this purpose, gene ontology (GO), pathway enrichment, and protein–protein interaction (PPI) analyses, as well as evaluating the interaction between co-expressed DEGs and differentially-expressed lncRNAs (DElncRNAs) were conducted. Additionally, the survival analyses of hub genes and transcription factors (TFs) were performed to estimate their prognostic performance as potential biomarkers in CRC patients. The results of qPCR confirmed the outputs of the bioinformatics analysis. In this regard, the expression pattern of the hub DEGs including PPARG, PTGS2, and PTPRC in mtKRAS cells compared to the wtKRAS sample, showed the overexpression of PPARG and PTGS2 and the downregulation of PTPRC both in qPCR and RNA-seq results. The pivotal hub genes (mainly PPARG, PTGS2, PTPRC, CDKN2A, and PRKACB) and lncRNAs (e.g., OGFRL1, DGCR5, and LINC01842) have been previously reported to have critical roles in the tumorigenesis of many cancers harboring KRAS mutation. Finally, the results provide novel insights into the potential biomarkers for KRAS mutant CRC, which may contribute to a better understanding of the molecular and cellular mechanisms mediating the KRAS mutation-induced malignant transformation.
Results
Identification of differentially expressed genes and non-coding RNAs
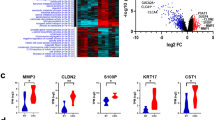
In this study, a multi-step strategy was developed to identify DEGs and differentially-expressed non-coding RNAs (DEncRNAs) (Fig. 1), allowing to distinguish the mechanisms involved in the KRAS mutation-induced malignancy by using publicly available datasets. In this regard, significant DEGs were first recognized by comparing the gene expression profiles between mtKRAS (HCT-116 and LoVo) and wtKRAS CRC cells (SW48). A total of 1979 and 1984 genes reached the criteria of 1.5-fold changes and adjusted p-value < 0.05 in HCT-116 versus SW48 and LoVo versus SW48, respectively (Supplementary Datas 1 and 2). Based on the results, 600 DEGs consisting of 492 coding and 108 non-coding (including 88 lncRNAS and miR-23b) Ensembl gene IDs were overlapped between the two comparisons. The identified DEGs with 423 up- and 149 downregulated gene IDs, in mtKRAS samples compared to wtKRAS, could be assigned to a KRAS-dependent gene expression signature (Supplementary Data 3). The cluster analysis of 600 DEGs revealed the significant differences between the wtKRAS and mtKRAS CRC cell lines (Fig. 2). Principle component analysis (PCA)-based multidimensional scaling visualization, linearly separated CRCs with and without KRAS mutation, on the basis of the gene expression data (Supplementary Data 4).
Heatmap plots based on the DEGs showing clustering of mtKRAS and wtKRAS cells lines into two distinct groups based on DEGs. (a) Clustering of HCT-116 and SW48 samples. (b) Clustering of LoVo and SW48 samples. Red shows overexpression and blue indicates underexpression with absolute log2FC > 1.5 and adjusted p-value < 0.05. Expression data are represented as normalized values (Z‑scores). Heatmap plot was created using gplots package in R environment and variance stabilizing transformation (VST) was applied to the normalized count data from DESeq2 package.
Gene set enrichment analysis (GSEA)
The results obtained from the GSEA analysis of the overrepresented KRAS signaling gene set reflected the clustering of LoVo and HCT-116 (mtKRAS) cells versus SW48 (wtKRAS) ones into two distinct groups (Supplementary Table 1). Additionally, GSEA was performed on the whole transcriptome of the CRC cell lines by using GSEA v2.07 software with 1000 gene set permutations to elucidate the discrimination of mtKRAS CRC cell line from wtKRAS one in the oncogene-associated signatures. Oncogenic signatures were derived from the Molecular Signatures Database v4.0. The results of GSEA indicated upregulated oncogenic signature gene sets like KRAS signaling in mtKRAS compared to the wtKRAS, while no significant oncogenic signature was upregulated while analyzing the use of SW48 versus mtKRAS cell lines as phenotype labels (Fig. 3).
GSEA analysis for oncogenic signature gene sets. (a) Enrichment plot of GSEA analysis for HCT-116. (b) Enrichment plot of GSEA analysis for LoVo. Enrichment plot on the whole transcriptome of the CRC cell lines showing the up-regulation of KRAS signaling oncogenic signature in HCT-116 vs SW48 and LoVo vs SW48. FDR < 25% after 1000 random permutations was set as the cut-off criterion.
GO analysis and signaling pathway enrichment of DEGs
Figure 4 displays the results related to the functional enrichment analysis of the 600 DEGs. All DEGs were computationally uploaded to the DAVID and EnrichR resources for the better revelation of the molecular and cellular mechanisms underlying the carcinogenicity of KRAS mutations. Based on the results of GO enrichment analysis, the DEGs were divided into 23 functional categories including 11 molecular function terms and 12 biological processes (Fig. 4a). Regarding the biological process group, DEGs were enriched in regulation of epithelial to mesenchymal transition (EMT), regulation of small GTPase mediated signal transduction, regulation of vascular endothelial cell proliferation, and positive regulation of GTPase activity. In the molecular function category, DEGs were mainly enriched in the GTP-dependent protein binding, GTP binding, GTPase activator activity, GTPase activity, Wnt-protein binding. The results of KEGG pathway enrichment analysis demonstrated that DEGs were mainly enriched in Rho-GTPase cycle, TOR signaling, TGF-beta regulation of extracellular matrix, proteoglycans in cancer, pathways in the cancer, PI3K-Akt signaling pathway, and RAS signaling pathway (Fig. 4b).
Functional enrichment analysis. (a) GO analysis of differentially expressed genes (DEGs) based on the molecular function (Red) and biological process (Blue). (b) KEGG pathway analysis of DEGs. (c) Functional enrichment analysis by Toppcluster. (d) Cluster of miR-23b and its target DEGs. (e) Protein–protein interaction network of DEGs. (f). One functional model identified from the PPI network by MCODE.
Further, ToppCluster was used for implementing a multi-cluster gene functional enrichment analysis to obtain a functional modular map based on the human disease and computational features. The genes in the cancer module (11 and 86) and colorectal neoplasm were significantly enriched for computational and human disease genes, respectively (Fig. 4c). Furthermore, a significant number of the DEGs, known as miR-23b targets, was obtained by considering their feature enrichment analysis by using ToppCluster (with Bonferroni correction and a 0.05 p-value cut-off) (Fig. 4d).
PPI network of DEGs
To verify the functional connectivity of the DEGs, the STRING database was interrogated to dissect the PPI networks. A total of 600 DEGs were analyzed by STRING database and following the removal of the partially-connected and isolated nodes, a network of 474 nodes with 601 edges was constructed by using the Cytoscape software (Fig. 4e). The top 10 hub genes with at least 10 degrees were PTPRC, PPARG, PRKACB, GATA3, PTGS2, CDKN2A, SCN5A, GFAP, CDH17, and EOMES, with the maximum node degree of which was observed in PTPRC as 25 (Fig. 4e). One functional module was identified from the PPI network of DEGs by using the MCODE composed of 5 nodes and 10 edges (Fig. 4f).
Interaction between co-expressed DEncRNAs and DEGs
Among DEncRNAs, 12 differentially-expressed lncRNAs exhibited possible interactions with some DEGs (Table 1). The results related to the interaction analysis of co-expressed DEG-DElncRNA predicted the presence of several mechanisms for lncRNAs targeting mRNAs. The co-expression analysis revealed the DElncRNAs regulating the expression of the genes which are majorly enriched in actin cytoskeleton regulation, gap junction, and Rap1, RAS, PI3K-AKT, and MAPK signaling pathways. Thus, these potential pathways which are all recognized as the cancer-related pathways could be described in the context of the KRAS mutation.
Transcription factor analysis
An EnrichR web-based tool was used for searching the overrepresented TFs regulating the identified DEGs. EnrichR was utilized for exploring the relevant TFs to find the possible regulatory mechanisms which may affect these target genes. The analysis was performed through detecting the binding motif sites in the gene list using the PWMs from TRANSFAC and JASPAR (Table 2). The regulatory biomolecules influencing the transcriptional regulation of DEGs were discovered by considering a statistical significance level of p-value < 0.01. Overall, 11 TFs of TFAP2A, SP1, ZBTB7A, HNRNPK, EGR1, TEAD4, HOXD9, SP3, PCBP1, MTF1, and UBTF were detected, which are mostly related to colon malignancy.
In-silico verification of hub genes and TFs
In order to test the stability of the results, the expression levels of hub genes and TFs were further explored in the same CRC cell lines using from Gene Expression Atlas database for verification (Table 3). In general, the expression levels related to 77% of these genes are in line with the results of differential expression analysis, which reflects the reliability and reproducibility of the results of the present study. In addition, the expression level of hub genes and TFs was compared with the expression data of CRC tissue samples deposited in TCGA databank. Regarding the simulations, 60% of hub proteins and 70% of reporter TFs were considered as distinct biomarker sets exhibiting their contribution to the discrimination of normal and tumor patient samples (Figs. 5 and 6).
Survival Analysis of hub genes and transcription factors
The Kaplan–Meier survival analyses of the hub genes, TFs, and lncRNAs in CRC patients was evaluated using GEPIA tools (Supplementary Figs. S1–S6). Among the hub genes, PTGS2 and CDKN2A, and GFAP were associated statistically significant with overall survival and disease-free survival of CRC patients. In addition, among the TFs, HNRNPK and EGR1 were associated statistically significant with overall survival and disease-free survival, respectively. Survival analysis for lncRNAs which were included in the interaction network with DEGs (Table 1) showed the significant connection between DGCR5 and overall survival of CRC patients (significance level at log-rank p < 0.05) (Fig. 7).
Quantitative real-time PCR verification
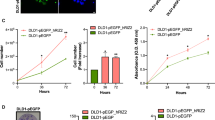
To validate the reliability of RNA-seq results, the expression level of the three selected hub genes possessing the highest degrees was measured using qRT-PCR between KRAS mutant CRC cell lines and their corresponding wtKRAS one (Fig. 8). Based on the results, PPARG and PTGS2 transcripts were significantly upregulated in KRAS mutant cell lines compared to the wtKRAS. In contrast, an elevation was obtained for the expression level of PTPRC in SW48 in comparison with the LoVo and HCT-116. The results are consistent with those of RNA sequencing, which confirms the reliability of RNA sequencing analysis.
Quantitative real-time PCR validated three dysregulated genes in association with KRAS mutation. Expressions of PPARG and PTGS2 were significantly elevated in KRAS-mutant cell lines (HCT-116 and LoVo) compared non-mutant cell line (SW48). The expressions of PTPRC in SW48 compared with other cell lines was upregulated.
Discussion
The recent advances in high-throughput technologies such as next-generation sequencing have enabled the global analyses of the key genes, lncRNAs, and miRs playing important roles in the CRC malignant transformation. In this study, the transcriptional profiles of mtKRAS and wtKRAS CRC cell lines were analyzed and compared by considering publicly available RNA-seq data to identify the critical signaling pathways involved in the tumorigenesis of the KRAS-driven CRCs.
Regarding the bioinformatics analysis of KRAS mutation effects on carcinogenesis, the RNA-seq data sets were selected from the largest publicly available repository of high-throughput sequencing data. Further, 600 DEGs were detected in comparing the transcriptome of mtKRAS cell lines with the wtKRAS one. The results of GO term analysis indicated the enrichment of DEGs in the biological processes such as the regulation of EMT, regulation of small GTPase mediated signal transduction, regulation of vascular endothelial cell proliferation, and positive regulation of GTPase activity, which are the critical causes of malignant transformation and cancer progression22,23. Epithelial plasticity or EMT is a key cellular program that can be activated by KRAS contributing to tumor progression24. Rho GTPases, the members of the Ras superfamily of small GTPases, are known as important signal transducers in the signaling pathways regulating cell proliferation, migration, survival, and death, all of which are the cellular processes crucial for cancer progression22.
The functional enrichment analysis revealed that DEGs were mainly enriched in key pathways like Rho-GTPase cycle, TOR signaling, TGF-beta regulation of extracellular matrix, proteoglycans in cancer, pathways in the cancer, PI3K-Akt signaling pathway, and RAS signaling pathway, which are all cancer-related pathways could be involved in KRAS-associated carcinogenesis25,26,27. Another functional enrichment analysis of the putative DEGs associated with KRAS was generated from ToppCluster. The results of analyzing based on the human disease and computational features were corresponded to the genes of cancer module and colorectal neoplasms, respectively. Therefore, the signaling pathways attributed to the DEGs under study are mainly involved in cancer and may provide new insights for understanding the cellular events underlying KRAS tumorigenesis.
In addition, 10 hub genes were identified from the constructed PPI network in terms of the highest degree, which included PTPRC, PPARG, PRKACB, GATA3, PTGS2, CDKN2A, SCN5A, GFAP, CDH17, and EOMES. Peroxisome proliferator activated receptors (PPARs) are the nuclear hormone receptors contributing to the genetic control of many cellular events. It, as a critical TF, regulates the important cellular functions which contribute to the regulation of metabolism and inflammation, and promotion of tumor survival and growth28. Some researchers have found the possible contribution of the crosstalk between PPARG signaling and epigenetic machinery to CRC development29. PPARG is overexpressed in different human cancers such as CRCs and pancreatic one, where its upregulation is correlated with increased pathological grade30,31,32. Further, prostaglandin E2 enhances PPARG transcriptional activity to promote CRC cell survival in vitro and intestinal tumorigenesis in mice33. JAK/STAT signal transduction pathway plays an essential role in the CRC malignancy through inactivating tumor suppressor genes and activating oncogenes[3435]. Receptor-type protein tyrosine phosphatases (PTPRs) are considered as a subgroup of protein tyrosine phosphatases with a crucial role in the cellular signaling pathways related to proliferation, survival, apoptosis, migration, and invasion. The genetic and epigenetic alterations of the phosphatases lead to aberrant cell signaling, which suggests that the PTPRs are critical components in the carcinogenesis of several cancers such as CRCs36,37. Consistent with previous studies, protein kinase CAMP-activated catalytic subunit beta (PRKACB) had a significant discrepancy of expression in normal and tumor tissues and it was also associated with poor patient-free survival and overall survival38.
Numerous studies have linked CRC to inflammation and prostaglandin-endoperoxide synthase (PTGS) 2 expression39. Other epidemiological, clinical, and animal studies have reported PTGS2 and epithelial growth factor (EGF) signaling pathways with key roles in promoting CRC growth and metastasis40. GATA3, as another hub gene in the present study, is expressed in CRC with a suppressive effect on the invasive behavior of CRC cells41. Methylation of cyclin dependent kinase inhibitor 2A (CDKN2A) promoter and subsequent gene silencing have been documented in many tumors including colon cancer and it has been associated with the CpG island methylator phenotype42. Microarray data established a definite role of sodium voltage-gated channel alpha subunit (SCN5A) as a high level regulator of CRC invasion network, including genes involved in Wnt signaling, cell migration, and cell cycle control43. The reduction in the density of glial fibrillary acidic protein (GFAP) glial cell type in the enteric nervous system was related with tumor grading of CRC and the inverse variation with the tumor proliferative activity and tumor-infiltrating leukocytes which could serve as a potential prognostic factor in this cancer44. Interaction between Cadherin-17 (CDH17) and α2β1 integrin has been revealed to regulate cell proliferation and adhesion in CRC cells causing liver metastasis45. In patients with CRC, high expression of Eomesodermin (EOMES) was associated with poor overall survival compared with individuals exhibiting low EOMES levels. However, a divergent role in cancer development, with tumor suppressor or oncogenic activities, has been reported for EOMES depending on stage and tissue context46.
Among the hub genes, the the three selected genes with the significant role in CRC tumorigenesis were selected for further verification by qPCR. The expression of PPARG, PTGS2, and PTPRC was validated in the CRC cell lines with and without KRAS mutation. The results represented the higher expression of PPARG and PTGS2 in KRAS mutant cell line than the SW48 (wtKRAS), which is in agreement with SRA data. The PTPRC expression was significantly upregulated in SW48 compared with HCT-116 and LoVo as the KRAS mutant samples.
To date, an increasing number of dysregulated lncRNAs with critical roles in human cancers such as CRC has been recognized47. In the present study, the dysregulated lncRNAs between mtKRAS and wtKRAS CRC cell lines were analyzed by using RNA-seq datasets from SRA. Among the lncRNAs, some with no annotation were excluded, and then the effects of the lncRNAs were highlighted, which could be included in the co-expressed DEG-DElncRNAs interaction network. The results of the pathway analysis of DEGs, which could potentially interact with DElncRNAs, reflected their major involvement in the actin cytoskeleton regulation, and Rap1, Ras, PI3K-AKT, and MAPK signaling pathway, and a positive correlation with cancer hallmarks. Various mechanisms are available for lncRNAs to activate, repress, or modulate the expression of target genes48. Cis-acting lncRNAs, which constitute a substantial fraction of lncRNAs, regulate the expression of their target genes in a location-dependent manner49. They can repress or activate their target genes through different mechanisms according to the possible architectures of lncRNA-target units. Antisense lncRNAs overlap their target genes in the antisense orientation50, while the sense ones overlap their target genes in the sense orientation and are typically contained within target gene introns51. Based on the competing endogenous RNA (ceRNA) hypothesis, the sequestration of the miRs from their mRNA target is another mechanism for regulating gene expression by lncRNAs52,53,54. Regarding the present study, the differential expression analysis of non-coding transcripts revealed the significant upregulation of miR-23b in wtKRAS cell line compared to the mtKRAS one. The downregulation of miR-23b in plasma is associated with poor prognosis among CRC patients55. Additionally, the PDGFR-modulated miR-23b cluster suppresses lung tumorigenesis by targeting the multiple members of KRAS and NF-kB pathways56. The results obtained from the feature enrichment analysis of DEGs with ToppCluster represented a significant number of DEGs as miR-23b targets. In the present study, a lncRNA-miRNA-mRNA cross-talk was recognized among AL589935.1-miR-23b-OGFRL1. Further, miR-23b was downregulated while upregulating AL589935.1 and OGFRL1 in mtKRAS versus wtKRAS, by indicating the ceRNA hypothesis. Considering some studies proposing the potential role of opioid signaling axis in cancer, the results can be utilized for understanding the molecular basis of the KRAS mutation-induced tumorigenesis57.
The significant TFs regulating the DEGs have been discovered, which play a role in colon malignancy. Among TFs, TFAP2A expression reduces in high-grade colorectal adenocarcinomas58, the high expression of SP1 is ascribed to CRC poor prognosis59, and ZBTB7A is known as an oncogene in CRC60. Furthermore, POU2F1 regulates colon malignancy61, the overexpression of HNRNPK is attributed to CRC poor prognosis62, EGR1 is related to CRC cell prolifration63, and TEAD4 and HOXD9 promote CRC tumorigenesis64,65. The survival analysis of the hub genes and TFs reflected their high potential as prognostic biomarkers with worse survival outcomes of the CRC patients.
In the present study, the dysregulated genes tending to be differentially expressed in the context of KRAS mutation were examined based on the SRA database, which included the hub genes derived from the PPI network such as PPARG, PTGS2, PTPRC, CDKN2A, and PRKACB. In addition, DElncRNAs were identified and the putative molecular interactions between DEGs and lncRNAs was provided. The results obtained from the GO and pathway analysis of dysregulated genes and lncRNAs proposed their role in the cancer-related pathways. Further, 11 overrepresented TFs regulating the identified DEGs were found, which were mostly related to colon malignancy. The survival analysis of hub genes and TFs revealed the contribution of almost all genes and TFs in discriminating the risk groups. The results of analyzing the differential expression of the non-coding transcripts represented that miR-23b was significantly upregulated in wtKRAS. Furthermore, a lncRNA-miRNA-mRNA crosstalk was observed among AL589935.1-miR-23b-OGFRL1. AL589935.1 and OGFRL1 were upregulated, and miR-23b was downregulated in mtKRAS versus wtKRAS, which suggests the possible ceRNA effect. The altered expression of PPARG, PTGS2, and PTPRC in mtKRAS and wtKRAS cells was confirmed by using qPCR and in-silico verification using Gene Expression Atlas and TCGA databases.
Some limitations of this study should be considered for the better interpretation of the results. The results were based on the transcriptional profile of cancer cell lines, which should be verified by patient sample data. However, the results were confirmed by the in-silico validation using TCGA database. Finally, more molecular biology experiments and computational method analysis for big data are required to clarify the relationship among the predicted regulatory lncRNAs, miR-23b, and DEGs.
Conclusion
In the present study, a comprehensive bioinformatics analysis of DEGs was performed, which may be involved in the KRAS-driven tumorigenesis. Along with affecting the KRAS-associated pathways, the possible effect of KRAS mutation on the gene expression of other signaling pathways such as those related to Rho-GTPase cycle, TOR signaling, TGF-beta regulation of extracellular matrix, proteoglycans in cancer, and pathways in the cancer is one of the important results of this study. The altered pathways may be associated with more significant cancer hallmarks induced by KRAS oncogenic effects. The identification of the differentially-expressed lncRNAs with crucial roles in regulating the potential cancer-related pathways may provide reference lncRNAs for diagnosing and treating KRAS mutant CRC. Additionally, a set of hub genes was recognized in the constructed PPI network, which may be considered as therapeutic targets. These discriminating genes such as PPARG, PTPRC, PTGS2, CDKN2A and PRKACB were reported with regard to their role in colorectal carcinogenesis although their expression was not described in the context of KRAS status in CRCs. Finally, further experiments are needed for understanding the corresponding roles and molecular mechanisms of the lncRNAs and mRNAs in detail to specify the mechanism of the KRAS-mediated tumorigenesis in CRC.
Materials and methods
Data collection
The RNA-seq data were extracted from the Sequence Read Archive database (https://www.ncbi.nlm.nih.gov/sra), which corresponded to the three human CRC cell lines of HCT-116 (SRR1030462, SRR1030463, SRR1756569, and SRR8615282), LoVo (SRR1756570, SRR8532655, and SRR8616185), and SW48 (ERR208907, SRR3228439, and SRR8615504). HCT-116, LoVo, and SW48 cell lines were obtained from The Research Institute of Biotechnology, Ferdowsi University of Mashhad.
Data preprocessing and differential expression analysis
The data were downloaded as SRA files and converted to FASTQ format using fastq-dump from the SRA toolkit (https://github.com/ncbi/sra-tools). The RNA-seq files were filtered for quality by using FLEXBAR, AfterQC, and Trimmomatic66,67,68. In addition, the human genome (http://ftp.ensembl.org/pub/release95/fasta/homo_sapiens/dna/Homo_sapiens.GRCh38.dna.toplevel.fa.gz) was indexed using Bowtie269,70. The filtered reads were mapped with Bowtie2, which the resultant SAM files were processed by using the htseq-count program for counting the aligned reads overlapped with the exons of each gene, given the GTF file (https://ftp.ensembl.org/pub/release95/gtf/homo_sapiens/Homo_sapiens.GRCh38.98.gtf.gz)71. Further, the expression profiles of HCT-116 and LoVo cell lines (mtKRAS) were compared with the transcriptome profile of SW48 one (wtKRAS) (as the control). The read counts were normalized for all samples and differentially-expressed Ensembl gene IDs were identified by using the DESeq2 package (version 1.38.0) from Bioconductor in R environment (version 3.6.1, https://www.rproject.org/)72. Furthermore, an adjusted p-value < 0.05 and a |log2FC|≥ 1.5 were defined as screening criteria. The Ensembl gene IDs with differential expression were annotated with Ensembl Biomart (https://asia.ensembl.org/biomart/martview). The Ensembl and human genes (GRCh38.p13) (as mart databases), as well as transcript type (as an attribute), were utilized for annotation to categorize the differentially-expressed Ensembl gene IDs into coding genes and non-coding RNAs. Hierarchical cluster analysis was performed to distinguish the differential expression of the overlapped DEGs in CRC samples with KRAS mutation compared to the control sample. Heatmap plot was created using gplots package in R environment and variance stabilizing transformation (VST) was applied to the normalized count data from DESeq2 package. Distance measurement and linkage analysis were based on Euclidean distance and complete linkage, respectively. Adjusted P value < 0.01 and a |log2FC|> 2 were defined as the cut-off criteria. Expression data are represented as normalized values (Z‑scores). Variations between CRC cell lines with and without KRAS mutations were also compared by PCA based on the multigene expression data.
Gene set enrichment analysis
Gene set enrichment analysis (https://www.gsea-msigdb.org/) was applied on the whole transcriptome to explore the potential altered pathways of mtKRAS cell lines in comparison with those of the wtKRAS one. To identify significantly enriched gene sets associated with cancer, between two biological states related to the KRAS mutation, gene sets were obtained from C6-Oncogenic signatures database. Two gene lists were analyzed consisting of the total number of the DEGs in HCT-116 versus SW48 and LoVo versus SW48 with 1972 and 1984 genes, respectively. The new GSEA Ensemble CHIP files provide this analysis for RNA-seq data and in this study Human-ENSEMBLE-Gene-MSigDB chip was selected as the chip platform. False discovery rate (FDR) less than 25% after 1000 random permutations was set as the cut-off criterion.
GO and pathway enrichment analysis of DEGs
To better understand the biological functions and potential mechanisms of overlapped DEGs in the mechanism of the KRAS-related tumorigenesis, we applied GO enrichment and pathway analyses. Briefly, GO analyses (www. geneontology.org) consisted of two components: biological process, and molecular function. Gene ontology analysis was performed by using the Database for Annotation, Visualization, and Integrated Discovery (DAVID) bioinformatics tool (version 6.8) (https://david.ncifcrf.gov/) and comprehensive gene set enrichment analysis web server EnrichR (https://amp.pharm.mssm.edu/Enrichr)73,74. Pathway analyses was carried to investigate the potential significant KEGG pathways using DAVID database and EnrichR resource75,76. The GO terms and KEGG pathways with p-value < 0.05 were considered as significantly enriched function annotations. Additionally, functional enrichment analysis was carried out by ToppCluster (https: //toppcluster.cchmc.org) as the other bioinformatics resource for detecting the functional enrichment of the candidate genes77. The ToppCluster output with Bonferroni correction and 0.05 p-value cut-off was visualized in Cytoscape software (version 3.7.2) (https://cytoscape.org/)78.
Protein–protein interaction network and module analysis
The potential DEG-encoded protein interactions were explored by using the Search Tool for the Retrieval of Interacting Genes (STRING, version 11.0) (https://string-db.org)79. The PPI network was visualized using Cytoscape 3.9.0 software, which the PPI confidence score lower than 0.4 and genes in the network with at least 10 degrees were respectively set as significant and hub genes. Subsequently, the Cytoscape plugin Molecular Complex Detection (MCODE) (version 2.0.0) was utilized to screen the modules of the PPI network with scores > 3, node ≥ counts 5, node score = 0.2, k-core = 2, and maximum depth = 100 as cut-off criteria.
Interaction analysis between co-expressed DEncRNAs and DEGs
According to the results of differential expression analysis, DElncRNAs were assessed for their co-expression with their mRNA targets. Finding the relevant co-expressed DEG-lncRNAs was based on the two different strategies. The first approach was on the basis of the hypothesis posing the ability of lncRNAs and mRNAs to co-regulate each other by sharing common miRNA response elements competitively, and the cis-regulation of mRNAs with DElncRNAs was considered as the second solution. lncRNAs, as the key regulators of gene expression are likely to function as competing endogenous RNAs (ceRNAs) through miRNA sponging, thereby indirectly regulating the expression of the genes targeted by these miRNAs. With this concept, miRNA targets of lncRNAs were predicted by LncRNA2Target v2.0 (http://www.lncrna2target.org) and LncTarD (http://bio-bigdata.hrbmu.edu.cn/LncTarD)80,81. Then, miRTarBase (http://mirtarbase.mbc.nctu.edu.tw/) and DIANA-TarBase v8 (http://www.microrna.gr/tarbase) were employed for predicting mRNA targets of candidate miRNAs to find our DEGs among the gene targets82,83. Regarding cis-regulation analysis, the genes transcribed within a 300-kb window upstream or downstream of lncRNAs were considered as cis target genes according to the Ensembl genome browser. These cis-regulated genes by DElncRNAs were checked to find the common genes with our DEGs which shows the interaction between co-expressed DElncRNAs and DEGs.
Transcription factor analysis
To identify overrepresented TF molecules from DEGs, TF discovery module was implemented, and processing was performed against both the TRANSFAC and JASPAR databases using EnrichR tool74,84. Based on the 35 different gene-set libraries, EnrichR computed enrichment and binding motif sites in the intended gene were detected through applying the position weight matrix (PWM) analysis from TRANSFAC and JASPAR85. The PWMs from TRANSFAC and JASPAR were utilized to scan the promoters of all human genes in the region of − 2000- + 500 from the transcription factor start site (TSS). Supposing the statistical significance of p-value < 0.05, TFs were recognized as reporter regulatory molecules, around which significant changes occur at the transcriptional level of DEGs.
Evaluation of the prognostic performance of hub genes and TFs
The prognostic power of hub genes and TF regulatory biomolecules were analyzed through utilizing interactive web-based tool GEPIA (Gene Expression Profiling Interactive Analysis), which is based on the gene expression RNA-seq datasets of TCGA (The Cancer Genome Atlas) and GTEx (Genotype-Tissue Expression)86. This resource aids in in-silico validation of target genes and for identifying tumor sub-group specific candidate biomarkers, on the basis of the expression analysis of the query genes across normal and tumor samples. The statistical significance of differences in gene expression levels between the normal and tumor groups were summarized for hub genes, TFs, and lncRNAs.
Culture of CRC cell lines
The human colon cancer cell lines HCT-116 and LoVo harboring KRAS mutation were selected as the mtKRAS ones. In addition, the SW48 cell line having wild-type KRAS, BRAF, PIK3CA, and TP53 was adopted as the control in the RNA-seq comparisons with the other mtKRAS cell lines. LoVo and SW48 cells were grown at 37 °C under 5% CO2 in the RPMI medium (Invitrogen) supplemented with 10% FBS (Invitrogen) and penicillin (100 U/mL)-streptomycin (100 μg/mL) (Betacell). However, HCT-116 cells were grown at 37 °C under 5% CO2 in the DMEM media (Gibco) complemented with 10% FBS and penicillin–streptomycin.
RNA extraction and quality monitoring
Total RNA was extracted from three independent experiments with a total RNA extraction kit (Parstous) according to the manufacturer's instructions. Further, RNA quantity and quality were measured on a Thermo Fisher Scientific NanoDrop 2000c spectrophotometer and an agarose gel electrophoresis. DNase treatment was performed by using a DNA-free DNase kit (SINACLON) and following the manufacturer's instruction.
Quantitative real-time PCR (RT-PCR) verification
The qPCRs were conducted by using SYBR green dye (Parstous) in a thermal cycler (BioRad CFX96). Briefly, cDNA was reverse-transcribed from 1 μg RNA with random hexamers, oligo d(t) primers, and H-minus MMLV reverse transcriptase by using an Easy cDNA synthesis kit (Parstous). The qPCR reaction components included 2 μL of cDNA, 10 μL of SYBR green master mix, 1 μL of each primer (100 pmol/μL), and 6 μL of nuclease-free water. Thermal cycling program was pre-denaturation at 95 °C for 4 min, followed by 40 cycles of denaturation at 95 °C for 30 s, annealing at 60 °C for 30 s, and extension at 72 °C for 30 s. All experiments were performed in technical and biological triplicate, the results of which were normalized against the geometrical mean of the expression levels of two reference genes ACTB and GAPDH. Furthermore, the first three hub genes with the highest degree (PPARG, PTGS2, and PTPRC) were selected for qPCR assay. Table 4 summarizes primer sequences. The relative gene expression levels between samples were calculated through employing 2−△△Ct method87.
In-silico validation of the expression levels of hub genes
The expression levels of hub genes were validated in the Expression Atlas database (https://www.ebi.ac.uk/gxa/) providing information on gene expression patterns from RNA-seq, microarray studies, and protein expression in order to confirm the validity of the results88. The baseline expression of each hub gene was compared among three CRC cell lines by considering cancer cell line as the biological condition and transcripts per million (TPM) as data unit to represent expression level.
Statistical analysis
All of the data were presented as the mean ± SD of three independent experiments (Prism-GraphPad Software) and analyzed through using unpaired Student's t-test (two-tailed). The p-values less than 0.05 were considered as statistically significant.
References
Bray, F. et al. Global cancer statistics 2018: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA 68(6), 394–424 (2018).
Porru, M., Pompili, L., Caruso, C., Biroccio, A. & Leonetti, C. Targeting KRAS in metastatic colorectal cancer: Current strategies and emerging opportunities. J. Exp. Clin. Cancer Res. 37(1), 57 (2018).
Stolze, B., Reinhart, S., Bulllinger, L., Fröhling, S. & Scholl, C. Comparative analysis of KRAS codon 12, 13, 18, 61, and 117 mutations using human MCF10A isogenic cell lines. Sci. Rep. 5, 8535 (2015).
Yuan, T. L. et al. Differential effector engagement by oncogenic KRAS. Cell Rep. 22(7), 1889–1902 (2018).
Simanshu, D. K., Nissley, D. V. & McCormick, F. RAS proteins and their regulators in human disease. Cell 170(1), 17–33 (2017).
Vigil, D., Cherfils, J., Rossman, K. L. & Der Channing, J. Ras superfamily GEFs and GAPs: Validated and tractable targets for cancer therapy?. Nat. Rev. Cancer 10(12), 842–857 (2010).
Liu, P., Wang, Y. & Li, X. Targeting the untargetable KRAS in cancer therapy. Acta Pharm. Sin. B 9, 871–879 (2019).
Misale, S. et al. Emergence of KRAS mutations and acquired resistance to anti-EGFR therapy in colorectal cancer. Nature 486(7404), 532–536 (2012).
Okamoto, K., Zaanan, A., Kawakami, H., Huang, S. & Sinicrope, F. A. Reversal of mutant KRAS-mediated apoptosis resistance by concurrent Noxa/Bik induction and Bcl-2/Bcl-xL antagonism in colon cancer cells. Mol. Cancer Res. 13(4), 659–669 (2015).
Saliani, M., Jalal, R. & Ahmadian, M. R. From basic researches to new achievements in therapeutic strategies of KRAS-driven cancers. Cancer Biol. Med. 16(3), 435–461 (2019).
Verma, R. K., Yu, W., Shrivastava, A., Shankar, S. & Srivastava, R. K. α-Mangostin-encapsulated PLGA nanoparticles inhibit pancreatic carcinogenesis by targeting cancer stem cells in human, and transgenic (Kras(G12D), and Kras(G12D)/tp53R270H) mice. Sci. Rep. 6, 32743–32743 (2016).
Siavoshi, A. et al. Network analysis of differential gene expression to identify hub genes in ovarian cancer. J. Cell Mol. Res. 12(1), 1–9 (2020).
Chen, J. W. & Dhahbi, J. Lung adenocarcinoma and lung squamous cell carcinoma cancer classification, biomarker identification, and gene expression analysis using overlapping feature selection methods. Sci. Rep. 11(1), 13323 (2021).
JamesdeBony, E. et al. Comprehensive identification of long noncoding RNAs in colorectal cancer. Oncotarget 9(45), 27605–27629 (2018).
Choi, Y. W. et al. MicroRNA expression signatures associated with BRAF-mutated versus KRAS-mutated colorectal cancers. Medicine 95(15), e3321 (2016).
Planck, M. et al. Genomic and transcriptional alterations in lung adenocarcinoma in relation to EGFR and KRAS mutation status. PLoS ONE 8(10), e78614 (2013).
Charitou, T. et al. Transcriptional and metabolic rewiring of colorectal cancer cells expressing the oncogenic KRAS G13D mutation. Br. J. Cancer 121(1), 37–50 (2019).
Watanabe, T. et al. Differential gene expression signatures between colorectal cancers with and without KRAS mutations: Crosstalk between the KRAS pathway and other signalling pathways. Eur. J. Cancer 47(13), 1946–1954 (2011).
Carlevaro-Fita, J. et al. Cancer LncRNA Census reveals evidence for deep functional conservation of long noncoding RNAs in tumorigenesis. Commun. Biol. 3(1), 1–16 (2020).
Goodall, G. J. & Wickramasinghe, V. O. RNA in cancer. Nat. Rev. Cancer 21(1), 22–36 (2021).
Lundberg, I. V. et al. MicroRNA expression in KRAS-and BRAF-mutated colorectal cancers. Anticancer Res. 38(2), 677–683 (2018).
Haga, R. B. & Ridley, A. J. Rho GTPases: Regulation and roles in cancer cell biology. Small GTPases 7(4), 207–221 (2016).
Sanchez-Vega, F. et al. Oncogenic signaling pathways in the cancer genome atlas. Cell 173(2), 321–337 (2018).
Arner, E. N., Du, W. & Brekken, R. A. Behind the wheel of epithelial plasticity in KRAS-driven cancers. Front. Oncol. 9, 1049 (2019).
Makrodouli, E. et al. BRAF and RAS oncogenes regulate Rho GTPase pathways to mediate migration and invasion properties in human colon cancer cells: A comparative study. Mol. Cancer 10(1), 118 (2011).
Polosukhina, D. et al. Functional KRAS mutations and a potential role for PI3K/AKT activation in Wilms tumors. Mol. Oncol. 11(4), 405–421 (2017).
Byun, J. K. et al. Oncogenic KRAS signaling activates mTORC1 through COUP-TFII-mediated lactate production. EMBO Rep. 20(6), e47451 (2019).
Liu, Y. et al. The role of PPAR-δ in metabolism, inflammation, and cancer: Many characters of a critical transcription factor. Int. J. Mol. Sci. 19(11), 3339 (2018).
Sabatino, L., Fucci, A., Pancione, M. & Colantuoni, V. PPARG epigenetic deregulation and its role in colorectal tumorigenesis. PPAR Res. 2012, 1–12 (2012).
Abdollahi, A. et al. Transcriptional network governing the angiogenic switch in human pancreatic cancer. Proc. Natl. Acad. Sci. USA 104(31), 12890–12895 (2007).
Takayama, O. et al. Expression of PPAR δ in multistage carcinogenesis of the colorectum: Implications of malignant cancer morphology. Br. J. Cancer 95(7), 889–895 (2006).
Zuo, X. et al. Metastasis regulation by PPARD expression in cancer cells. JCI Insight 2(1), e91419 (2017).
Wang, D. et al. Prostaglandin E2 promotes colorectal adenoma growth via transactivation of the nuclear peroxisome proliferator-activated receptor δ. Cancer Cell 6(3), 285–295 (2004).
Slattery, M. L., Lundgreen, A., Kadlubar, S. A., Bondurant, K. L. & Wolff, R. K. JAK/STAT/SOCS-signaling pathway and colon and rectal cancer. Mol. Carcinog. 52(2), 155–166 (2013).
Tang, S. et al. Association analyses of the JAK/STAT signaling pathway with the progression and prognosis of colon cancer. Oncol. Lett. 17(1), 159–164 (2019).
Chu, S. & Wang, HYu. M. A putative molecular network associated with colon cancer metastasis constructed from microarray data. World J. Surg. Oncol. 15(1), 115–115 (2017).
Du, Y. & Grandis, J. R. Receptor-type protein tyrosine phosphatases in cancer. Chin. J. Cancer 34(2), 61–69 (2015).
Yao, X. et al. Application of cAMP-dependent catalytic subunit β (PRKACB) low expression in predicting worse overall survival: A potential therapeutic target for colorectal carcinoma. J. Cancer 11(16), 4841–4850 (2020).
Benelli, R., Venè, R. & Ferrari, N. Prostaglandin-endoperoxide synthase 2 (cyclooxygenase-2), a complex target for colorectal cancer prevention and therapy. Transl Res. 196, 42–61 (2018).
Wang, D., Xia, D. & Dubois, R. N. The crosstalk of PTGS2 and EGF signaling pathways in colorectal cancer. Cancers 3(4), 3894–3908 (2011).
French, C. L. et al. Linking patient outcome to high throughput protein expression data identifies novel regulators of colorectal adenocarcinoma aggressiveness. F1000Research 4, 99 (2015).
Shima, K. et al. Prognostic significance of CDKN2A (p16) promoter methylation and loss of expression in 902 colorectal cancers: Cohort study and literature review. Int. J. Cancer 128(5), 1080–1094 (2011).
House, C. D. et al. Voltage-gated Na+ channel SCN5A is a key regulator of a gene transcriptional network that controls colon cancer invasion. Cancer Res. 70(17), 6957–6967 (2010).
Târtea, E. A. et al. Implications of inflammation and remodeling of the enteric glial cells in colorectal adenocarcinoma. Rom. J. Morphol. Embryol. 58(2), 473–480 (2017).
Bartolomé, R. A. et al. Cadherin-17 interacts with α2β1 integrin to regulate cell proliferation and adhesion in colorectal cancer cells causing liver metastasis. Oncogene 33(13), 1658–1669 (2014).
Wang, R. et al. Reciprocal regulation of BMF and BIRC5 (Survivin) linked to Eomes overexpression in colorectal cancer. Cancer Lett. 381(2), 341–348 (2016).
Siddiqui, H., Al-Ghafari, A., Choudhry, H. & Al Doghaither, H. Roles of long non-coding RNAs in colorectal cancer tumorigenesis: A review. Mol. Clin. Oncol. 11(2), 167–172 (2019).
Fernandes, J. C., Acuña, S. M., Aoki, J. I., Floeter-Winter, L. M. & Muxel, S. M. Long non-coding RNAs in the regulation of gene expression: Physiology and disease. Non-coding RNA 5(1), 17 (2019).
Gil, N. & Ulitsky, I. Regulation of gene expression by cis-acting long non-coding RNAs. Nat. Rev. Genet. 21, 102–117 (2019).
Latgé, G., Poulet, C., Bours, V., Josse, C. & Jerusalem, G. Natural antisense transcripts: Molecular mechanisms and implications in breast cancers. Int. J. Mol. Sci. 19(1), 123 (2018).
Tahira, A. C. et al. Long noncoding intronic RNAs are differentially expressed in primary and metastatic pancreatic cancer. Mol. Cancer 10(1), 141 (2011).
López-Urrutia, E., Montes, L. P. B., de Guevara Cervantes, D. L., Pérez-Plasencia, C. & Campos-Parra, A. D. Crosstalk between long Non-coding RNAs, Micro-RNAs and mRNAs: Deciphering molecular mechanisms of master regulators in cancer. Front. Oncol. 9, 669 (2019).
Salmena, L., Poliseno, L., Tay, Y., Kats, L. & Pandolfi, P. P. A ceRNA hypothesis: The Rosetta Stone of a hidden RNA language?. Cell 146(3), 353–358 (2011).
Shi, L. et al. A KRAS-responsive long non-coding RNA controls microRNA processing. Nat. Commun. 12(1), 1–19 (2021).
Kou, C. H., Zhou, T., Han, X. L., Zhuang, H. J. & Qian, H. X. Downregulation of mir-23b in plasma is associated with poor prognosis in patients with colorectal cancer. Oncol. Lett. 12(6), 4838–4844 (2016).
Naidu, S. et al. PDGFR-modulated miR-23b cluster and miR-125a-5p suppress lung tumorigenesis by targeting multiple components of KRAS and NF-kB pathways. Sci. Rep. 7(1), 1–14 (2017).
Yamashita, H., Shuman, L., Warrick, J. I., Raman, J. D. & Degraff, D. J. Androgen represses opioid growth factor receptor (OGFR) in human prostate cancer LNCaP cells and OGFR expression in human prostate cancer tissue. Am. J. Clin. Exp. Urol. 6(4), 164–171 (2018).
Ropponen, K. et al. Expression of transcription factor AP-2 in colorectal adenomas and adenocarcinomas; comparison of immunohistochemistry and in situ hybridisation. J. Clin. Pathol. 54(7), 533–538 (2001).
Bajpai, R. & Nagaraju, G. P. Specificity protein 1: Its role in colorectal cancer progression and metastasis. Crit. Rev. Oncol. Hematol. 113, 1–7 (2017).
Wang, L., Zhang, M.-X. & Zhang, M.-F.-W. ZBTB7A functioned as an oncogene in colorectal cancer. BMC Gastroenterol. 20(1), 1–7 (2020).
Vázquez-Arreguín, K. et al. Oct1/Pou2f1 is selectively required for colon regeneration and regulates colon malignancy. PLoS Genet. 15(5), e1007687 (2019).
Carpenter, B. et al. Heterogeneous nuclear ribonucleoprotein K is over expressed, aberrantly localised and is associated with poor prognosis in colorectal cancer. Br. J. Cancer 95(7), 921–927 (2006).
Myung, D.-S. et al. Expression of early growth response-1 in colorectal cancer and its relation to tumor cell proliferation and apoptosis. Oncol. Rep. 31(2), 788–794 (2014).
Liu, M. et al. HOXD9 promote epithelial-mesenchymal transition and metastasis in colorectal carcinoma. Cancer Med. 9, 3932–3943 (2020).
Tang, J.-Y. et al. TEAD4 promotes colorectal tumorigenesis via transcriptionally targeting YAP1. Cell Cycle 17(1), 102–109 (2018).
Dodt, M., Roehr, J. T., Ahmed, R. & Dieterich, C. FLEXBAR-flexible barcode and adapter processing for next-generation sequencing platforms. Biology 1(3), 895–905 (2012).
Chen, S. et al. AfterQC: Automatic filtering, trimming, error removing and quality control for fastq data. BMC Bioinform. 18(Suppl 3), 80–80 (2017).
Bolger, A. M., Lohse, M. & Usadel, B. Trimmomatic: A flexible trimmer for Illumina sequence data. Bioinformatics 30(15), 2114–2120 (2014).
Cunningham, F. et al. Ensembl 2019. Nucleic Acids Res. 47(D1), D745–D751 (2018).
Langmead, B. & Salzberg, S. L. Fast gapped-read alignment with Bowtie 2. Nat. Methods 9(4), 357–359 (2012).
Anders, S., Pyl, P. T. & Huber, W. HTSeq: A Python framework to work with high-throughput sequencing data. Bioinformatics 31(2), 166–169 (2015).
Anders, S. & Huber, W. Differential expression analysis for sequence count data. Genome Biol. 11(10), R106 (2010).
Dennis, G. et al. DAVID: Database for annotation, visualization, and integrated discovery. Genome Biol. 4(9), R60 (2003).
Kuleshov, M. V. et al. Enrichr: A comprehensive gene set enrichment analysis web server 2016 update. Nucleic Acids Res. 44(W1), W90–W97 (2016).
Kanehisa, M. & Goto, S. KEGG: Kyoto encyclopedia of genes and genomes. Nucleic Acids Res. 28(1), 27–30 (2000).
Kanehisa, M., Goto, S., Furumichi, M., Tanabe, M. & Hirakawa, M. KEGG for representation and analysis of molecular networks involving diseases and drugs. Nucleic Acids Res. 38, D355–D360 (2010).
Kaimal, V., Bardes, E. E., Tabar, S. C., Jegga, A. G. & Aronow, B. J. ToppCluster: A multiple gene list feature analyzer for comparative enrichment clustering and network-based dissection of biological systems. Nucleic Acids Res. 38, W96–W102 (2010).
Shannon, P. et al. Cytoscape: A software environment for integrated models of biomolecular interaction networks. Genome Res. 13(11), 2498–2504 (2003).
Szklarczyk, D. et al. STRING v11: Protein-protein association networks with increased coverage, supporting functional discovery in genome-wide experimental datasets. Nucleic Acids Res. 47(D1), D607–D613 (2019).
Cheng, L. et al. LncRNA2Target v2.0: A comprehensive database for target genes of lncRNAs in human and mouse. Nucleic Acids Res. 47(D1), D140–D144 (2019).
Zhao, H. et al. LncTarD: A manually-curated database of experimentally-supported functional lncRNA-target regulations in human diseases. Nucleic Acids Res. 48(D1), D118–D126 (2020).
Chou, C.-H. et al. miRTarBase update 2018: A resource for experimentally validated microRNA-target interactions. Nucleic Acids Res. 46(D1), D296–D302 (2018).
Karagkouni, D. et al. DIANA-TarBase v8: A decade-long collection of experimentally supported miRNA-gene interactions. Nucleic Acids Res. 46(D1), D239–D245 (2018).
Chen, E. Y. et al. Enrichr: Interactive and collaborative HTML5 gene list enrichment analysis tool. BMC Bioinformatics 14(1), 128 (2013).
Badr, M. & Häcker, G. Gene expression profiling meta-analysis reveals novel gene signatures and pathways shared between tuberculosis and rheumatoid arthritis. PLoS ONE 14(3), e0213470 (2019).
Tang, Z. et al. GEPIA: A web server for cancer and normal gene expression profiling and interactive analyses. Nucleic Acids Res. 45(W1), W98-w102 (2017).
Zhang, Z., Li, B., Xu, P. & Yang, B. Integrated whole transcriptome profiling and bioinformatics analysis for revealing regulatory pathways associated with quercetin-induced apoptosis in HCT-116 cells. Front. Pharmacol. 10, 798 (2019).
Papatheodorou, I. et al. Expression Atlas: Gene and protein expression across multiple studies and organisms. Nucleic Acids Res. 46(D1), D246–D251 (2018).
Acknowledgements
Authors are thankful to Akram Siavoshi for her insightful suggestions. We acknowledge Mehrnaz Mehrabipour for her help to prepare the figures. Authors are grateful to Iran National Science Foundation (INSF). This project was supported by Ferdowsi University of Mashhad, grant number 3/52762.
Author information
Authors and Affiliations
Contributions
NGS data analysis and preparing the manuscript draft: M.S.; Supervision, study design, and critical review of the final draft: R.J.; Supervision, study design, and critical review of the final draft: A.J.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Saliani, M., Jalal, R. & Javadmanesh, A. Differential expression analysis of genes and long non-coding RNAs associated with KRAS mutation in colorectal cancer cells. Sci Rep 12, 7965 (2022). https://doi.org/10.1038/s41598-022-11697-5
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-022-11697-5
- Springer Nature Limited
This article is cited by
-
DeepRisk network: an AI-based tool for digital pathology signature and treatment responsiveness of gastric cancer using whole-slide images
Journal of Translational Medicine (2024)
-
Long non-coding RNA signature in colorectal cancer: research progression and clinical application
Cancer Cell International (2023)