Abstract
Complete ammonia oxidizer (Comammox) can complete the whole nitrification process independently, whose niche differentiation is important guarantee for its survival and ecological function. This study investigated the niche differentiation of comammox Nitrospira in the sediments of three typical tributaries of the Three Gorges Reservoir (TGR). Clade A and clade B of comammox Nitrospira coexisted in all sampling sites simultaneously. The amoA gene abundance of clade A and B was gradually increased or decreased along the flow path of the three tributaries with obvious spatial differentiation. The amoA gene abundance of comammox Nitrospira clade A (6.36 × 103 − 5.06 × 104 copies g−1 dry sediment) was higher than that of clade B (6.26 × 102 − 6.27 × 103 copies g−1 dry sediment), and the clade A amoA gene abundance was one order of magnitude higher than that of AOA (7.24 × 102 − 6.89 × 103 copies g−1 dry sediment) and AOB (1.44 × 102 − 1.46 × 103 copies g−1 dry sediment). A significant positive correlation was observed between comammox Nitrospira clade A amoA gene abundance and flow distance (P < 0.05). The number of operational taxonomic units (OTUs) in two sub-clades of clade A accounted for the majority in different tributaries, indicating that clade A also had population differentiation among different tributaries. This study revealed that comammox Nitrospira in the sediments of TGR tributaries have niche differentiation and clade A.2 played a more crucial role in comammox Nitrospira community.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Introduction
Nitrification transforms 2330 teragrams of nitrogen during the earth’s nitrogen cycle every year1. For over a century, nitrification process was considered to consist of two steps: first, ammonia was oxidized to nitrite, and then nitrite was oxidized to nitric acid2,3. However, the recent discovery of comammox has broken through traditional conception of two-step nitrification process4,5. As a bacterium associates with Nitrospira sublineage II, comammox can completely oxidize ammonia into nitrate in a single cell because it can encode all the enzymes required to complete nitrification6,7. Previous study has demonstrated that comammox Nitrospira can be classified into two clades, namely clade A and clade B, and clade A was further grouped into sub-clade A.1 and A.28.
Many studies have revealed a wide existence of comammox Nitrospira in freshwater and terrestrial ecosystems such as basin9, lake10, riparian ecosystem11, coastal wetland12, agricultural soil13, and forest soil14. Comammox Nitrospira have also been found in engineering environments including drinking water systems6,15, wastewater treatment systems16,17 and activated sludge18. These studies show that comammox Nitrospira exhibit strong distinct ecological distribution. In some environments, amoA gene of comammox Nitrospira is undetectable9, but in others, comammox Nitrospira are the main ammonia-oxidizing microorganism (AOM). In addition, different lineages of comammox Nitrospira also show niche differences. Clade A.1 is the main lineage in mudflat sediments19, while clade A.2 and clade B are abundant in agricultural soils20,21. Clade A.1 is more widely distributed than clade A.2 in natural aquatic ecosystems22. Moreover, Ca. Nitrospira inopinata, belonging to clade A.1, is the only successfully cultured pure comammox Nitrospira strain23. Comammox Nitrospira clade B has a higher affinity for NH4+-N than clade A, which may be due to the existence of an ammonium transporter in clade B7,24,25. Such a niche differentiation ensures that comammox Nitrospira can widely survive in nature and exert their ammonia oxidation function.
The Three Gorges Reservoir (TGR), situated in the middle reaches of the Yangtze River of China, is the largest reservoir in the world with a length of 663 km, a water storage of 39.3 billion m3, and a water surface area of 1084 km226. TGR has a function of a periodic anti-seasonal water storage and discharge, and it discharges water in the rainy season and stores water in the dry season every year with a water level of 145 − 175 m27. The storage capacity of the TGR tributaries accounts for about 25.0% of the total storage capacity of the reservoir area, and eutrophication has gradually appeared in TGR tributaries in recent years due to the enrichment of nutrients28,29. The main stream in the reservoir area exhibits a certain supporting effect on the tributaries in the process of integrating the tributaries into the main stream, which makes the flow velocity of the tributaries gradually slow down and the sediments gradually settle down, resulting in the spatial difference among the tributaries along the flow path.
One previous study on the Yangtze River continuum has shown that comammox Nitrospira in the sediments of Yangtze River mainstream present large-scale niche differentiation at different altitudes30. However, the distribution and diversity of comammox Nitrospira in the sediments of the TGR tributaries remain largely unknown. In this study, we hypothesized that the spatial differences among the TGR tributaries might lead to niche differentiation of comammox Nitrospira in the sediments. We investigated abundance, distribution, and diversity of comammox Nitrospira in sediments of three typical TGR tributaries, trying to test this hypothesis.
Methods
Study areas and sample collection
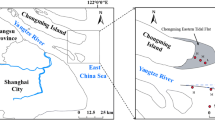
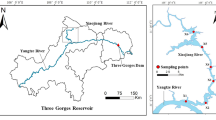
Xiaojiang River, Daning River, and Xiangxi River are the main tributaries of the Yangtze River in TGR area with a water surface area of 1000 km2. The main stream of XiaoJiang River, Daning River, and Xiangxi River is 182.4 km, 250 km, and 97.3 km, respectively. Xiaojiang River basin is mainly surrounded by agricultural land, while Daning River basin and Xiangxi River basin are mainly surrounded by forestry and urban land. According to the length of each sampling tributary, five (J1 − J5), seven (D1 − D7), and ten (X1 − X10) sampling sites were set in Xiaojiang River, Daning River, and Xiangxi River, respectively (Fig. 1). The flow distance between the sampling sites of each tributary was calculated using ArcGIS 10.7. The samples were collected in December 2019. Three parallel sediment samples were collected from a surface of 10 cm at each site, stored in sterile ziplock bags, and taken back to the laboratory on ice. Upon return to the laboratory, the sediments from different sampling sites belonging to same tributary were mixed into one sample, and stored at -80℃ for high-throughput sequencing. The sediment samples were divided into two parts. One part was stored at − 80℃ for DNA extraction, and the other was stored at − 20℃ for physicochemical properties determination.
Sampling sites in three typical tributaries of the Yangtze River in TGR area. The three tributaries were (a) Xiaojiang River, (b) Daning River, and (c) Xiangxi River. The basic geographic information was obtained from the National Geomatics Center of China (NGCC) and generated using the ArcGIS 10.7 (http://www.esri.com/).
Physicochemical analysis
Moisture content of the sediments was measured by weight loss after wet sediment was dried at 105℃ to constant weight. The ammonia nitrogen (NH4+-N), nitrite nitrogen (NO2–-N), and nitrate nitrogen (NO3–-N) were extracted from the sediment samples which were air dried and passed through a 2 mm sieve with 2 mol L−1 KCl. The pH of 1:2.5 dry soil / 2 mol L−1 KCl (wt/vol) suspensions after 30-min shaking was determined as pH of sediment samples using a pH analyzer (METTLER TOLEDO, Switzerland). Total nitrogen (TN) and total carbon (TC) in sediments were measured using an Element analyzer (Elementar Vario PYRO cube, Germany). Total phosphorus (TP) in sediments was determined by the perchloric acid sulfuric acid method. The physicochemical parameters of all sediment samples were analyzed in triplicate.
DNA extraction and PCR amplification
The total genomic DNA was extracted from sediment samples using Fast DNA® Spin for Soil Kit (MPBIO, USA) according to the manufacturer’s instructions. The purity and concentration of the extracted DNA samples were determined using a super differential spectrophotometer (NanoPhotometer-N60, IMPLEN, Germany). The specific primer pair including pmoA-189b-F (GGNGACTGGGACTTYTGG) and Com-amoA_1_R (CGAGATCATGGTGCTGTGAC) was used to amplify the sediment comammox Nitrospira amoA gene31. PCR reactions were performed in a total volume of 25μL containing 2μL of template DNA, 2μL of dNTP (2.5 mM, TransGen, China), 5μL of 5 × reaction buffer, 5μL of 5 × GC buffer, 1μL of each primer (10 μM), 8.75μL ddH2O, and 0.25μL of Q5® High-Fidelity DNA Polymerase (New England Biolabs, USA). PCRs were conducted as follows: predenaturation at 98 ℃ for 2 min; followed by 30 cycles at 98 ℃ for 15 s, 55 ℃ for 30 s, and 72 ℃ for 30 s; and a final extension at 72 ℃ for 5 min. Agarose gel electrophoresis was conducted to determine the specificity of the amplified products.
Amplicon sequencing and phylogenetic analysis
PCR products were sequenced using Illumina NovaSeq PE250 by Shanghai Personal Biotechnology Company Limited (Shanghai, China). Raw data were processed using the Vsearch (v2.13.4_linux_x86_64)32. The specific treatment process was as follows. First, cutadapt (v2.3) was used to cut the primer fragment and discard the sequences unmatched with primer. Afterwards, paired-end reads were merged and quality filtered using Vsearch. The sequences were clustered to operational taxonomic units (OTUs) according to 97% nucleic acid similarity with chimeras eliminated. Then, singletons OTUs and their representative sequences were removed from the OTU table. Finally, RDP FrameBot (v1.2) was used to correct errors of insertion or deletion in OTU sequences according to the seed protein sequence of comammox Nitrospira amoA gene33. Neighbor-joining phylogenetic tree of one representative sequence of each main OTU and its closest reference sequence retrieved from GenBank was created using MEGA X with 1000 bootstrap replicates to evaluate the reliability of the tree topologies34,35.
Quantitative real-time PCR (qRT-PCR)
The qRT-PCR was performed to determine the amoA gene abundance of comammox Nitrospira clade A, comammox Nitrospira clade B, AOA, and AOB using QuantStudio™ 6 Flex quantitative PCR instrument (Thermo Fisher Scientific, Singapore). Four primer pairs, namely, CA377f/C576r36, CB377f/C576r36, Arch-amoAF/Arch-amoAR37, and amoA–1Fmod/GenAOBR38 were used for the quantification of the aforementioned four amoA genes, respectively. Quantitative PCR was carried out in 10 μL amplification system containing 5.0 μL of T5 Fast qPCR Mix (2 ×), 0.4 μL of each Primer (10 μM), 0.2 μL of ROX Reference Dye II (50 ×), 1 μL of template DNA, and 3.0 μL of distilled deionized water (ddH2O). The qRT-PCR amplification conditions were shown in Table S1.
Statistical analysis
The differences in the sediment properties and AOMs amoA gene abundance between different groups were tested by one-way ANOVA and Duncan’ test from agricolae package (https://cran.r-project.org/web/packages/agricolae/) and dplyr package (https://dplyr.tidyverse.org) in R v3.6.139 (P < 0.05). Standard curve of qPCR process and data generated were analyzed by QuantStudio™ Real-Time PCR Software (version 1.2). Detrended correspondence analysis (DCA) was performed to preliminarily determine environmental parameters. Since the largest value of gradient length obtained from DCA was less than 3.0, redundancy discriminate analysis (RDA) was further performed using vegan package (https://github.com/vegandevs/vegan) in R v3.6.1. Spearman correlation analysis and mapping were conducted using corrplot package (https://github.com/taiyun/corrplot) in R v3.6.1 to test correlations among diversity, abundance and environmental parameters. Heat map of relative abundance of main OTUs was completed using pheatmap package (https://cran.r-project.org/web/packages/pheatmap/) in R v3.6.1. The α-diversity was calculated using QIIME40. Co-occurrence network was constructed by the OTUs with relative abundance ≥ 0.02%. Network topological parameters and sparCC coefficients were calculated by igraph package (https://igraph.org) and SpiecEasi package (https://github.com/zdk123/SpiecEasi) in R v3.6.1 and image of co-occurrence network were generated by Gephi 0.9.2 with Fruchterman-Reingold layout41,42. Other graphs were prepared using Origin 2018 software. P < 0.05 was considered as significantly different.
Nucleotide sequence accession numbers
The nucleotide sequence of comammox Nitrospira amoA gene obtained in this study was submitted to the GenBank database with the accession number of MZ669776 − MZ669807.
Ethics approval and consent to participate
Not applicable.
Consent for publication
Not applicable.
Results
Environmental parameters
Physical and chemical indexes of sediments from three typical TGR tributaries of the Yangtze River in China were shown in Table 1. The sediments at each sampling point were weakly alkaline (pH > 7). There was no significant difference in moisture content among Xiaojiang River, Daning River, and Xiangxi River. There is little difference in TN content among the three tributaries. However, the NH4+-N in Xiangxi River was lower than that in other two tributaries with an average of 5.42 mg kg−1. Daning River had the highest NO2–-N with an average of 0.43 mg kg−1. The order ranking from high to low in terms of the average NO3–-N content in three rivers were Xiaojiang River (2.10 mg kg−1), Xiangxi River (1.25 mg kg−1), and Daning River (0.80 mg kg−1). The average TC content in Xiaojiang River was 1.68 g kg−1, which was about half as much as that in Daning River (3.26 g kg−1). Xiangxi Rievr had the highest average TP content (87.22 mg kg−1) among three tributaries.
Abundance of Comammox Nitrospira amoA gene and Canonical Ammonia Oxidizers amoA gene
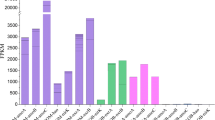
In the sediments from three tributaries in TGR area, the comammox Nitrospira clade A amoA gene showed a gradual increasing abundance along the flow path (Fig. 2). The clade A abundance reached the maximum at the estuary of the three tributaries. Similar to the spatial variation of clade A abundance, clade B amoA gene abundance was also increased gradually with the flow path, and its abundance reached the highest at the estuary of Daning River. However, the clade B amoA gene abundance in the other two rivers showed an overall decreasing trend along the flow path.
The amoA gene abundance distribution of AOA, AOB, Comammox Nitrospira clade A, and Comammox Nitrospira clade B in (a) Xiaojiang River, (b) Daning River, and (c) Xiangxi River. Different lower-case letters indicate a significant difference among different AOMs at the level of P < 0.05 by Duncan’ test.
In the total of 22 samples from three TGR tributaries, the amoA gene abundance of comammox Nitrospira clade A (6.36 × 103 − 5.06 × 104 copies g−1 dry sediment) was significantly higher than that of comammox Nitrospira clade B (6.26 × 102 − 6.27 × 103 copies g−1 dry sediment), AOA (1.03 × 102 − 2.14 × 103 copies g−1 dry sediment), and AOB (1.44 × 102 − 1.46 × 103 copies g−1 dry sediment) (Fig. 3, P < 0.05). Moreover, the amoA gene abundance of comammox Nitrospira clade B was also significantly higher than that of AOA and AOB in the three tributaries (Fig. 3, P < 0.05). In addition, AOA amoA gene abundance was significantly higher than AOB amoA gene abundance in Xiaojiang River and Xiangxi River, but there was no significant difference in amoA gene abundance between these two microorganisms in Daning River. In AOMs, AOA was the only one exhibiting significant difference in amoA gene abundance among tributaries. AOA amoA gene abundance in Daning River was significantly higher than that in Xiaojiang River (Fig. 3, P < 0.05, Duncan’ test).
Effect of physicochemical characteristics of sediments on amoA gene abundance of AOMs
The first two RDA axes in three tributaries’ RDA diagram jointly explained 83.8% of species-environment relationships (Fig. 4). Comammox Nitrospira clade A amoA gene abundance was significantly positively correlated with flow distance (Fig. 5, r = 0.564, P < 0.01, n = 22). AmoA gene abundance showed a significant positive correlation between the two lineages of comammox Nitrospira (Fig. 5, r = 0.433, P < 0.05, n = 22), and a significant positive correlation in amoA gene abundance was also observed between comammox Nitrospira clade A and AOA (Fig. 4).
Heatmap exhibits the pairwise Spearman’s correlations among environmental parameters, amoA gene abundance of AOMs, and α-diversity indices of comammox Nitrospira community. Only the significant correlations (P < 0.05) are labeled in circles. Different colors of scale bars (at the bottom) indicate different correlation coefficient values.
In addition, the amoA gene abundance of AOA and AOB was positively correlated with sediment TP and NH4+-N, respectively (Fig. 4). AOB amoA gene abundance was also significantly positively correlated with sediment NO2–-N content (r = 0.542, P < 0.01, n = 22), TC content (r = 0.517, P < 0.05, n = 22), and C:N ratio (r = 0.523, P < 0.05, n = 22) (Fig. 5).
Biodiversity and community of comammox Nitrospira
All the generated 191692 high-quality comammox Nitrospira amoA gene sequences were clustered into 5086 OTUs based on 97% nucleotide similarity. The α-diversity for each point was shown in Table S2. Spearman correlation analysis indicated that sediment pH, NH4+-N, and TP were significantly correlated with all the α-diversity indices of comammox Nitrospira community (P < 0.05, Fig. 5).
The phylogenetic analysis revealed that the 32 main OTUs with a relative abundance > 0.3% accounted for 76.9% of the total comammox Nitrospira amoA gene sequences (Fig. 6). The phylogenetic tree supported the division of the 32 main OTUs of comammox Nitrospira amoA gene into comammox Nitrospira into clade A (27 OTUs) and clade B (5 OTUs). Clade A was further subdivided into clade A.1 (11 OTUs) and A.2 (16 OTUs). Clade A.1, clade A.2, and clade B were found in the sediments from all three tributaries. Clade A.1 accounted for 40.6% and 35.2% of comammox Nitrospira community in Xiaojiang River and Xiangxi River, and it was the most dominant lineage in these two tributaries. Clade A.2 was the most dominant lineage in Daning River, accounting for 38.0% of comammox Nitrospira community (Fig. 7). Clade B was not the dominant lineage in all three tributaries with its highest percentage found in Xiangxi River (12.4%). The relationship of 214 OTUs with relative abundance > 0.02% was explored with co-occurrence network analysis (Fig. 8). The connections among the members in network were mostly positive, accounting for 54.0% (Table S3). It was noteworthy that nodes of clade A.1, cladeA.2, and clade B accounted for 33.2%, 51.4%, and 15.4% of all nodes, respectively, and 9, 17, and 2 of 28 hub nodes, respectively. In addition, the average degrees of clade A.1, clade A.2 and clade B were 10.4, 12.0 and 8.1, respectively (Table S4). Clade A.2 not only had the most OTUs and the highest relative abundance in phylogenetic analysis, but also occupied the most nodes and had the highest average degree in network analysis.
Neighbor-joining phylogenetic tree and relative abundances of Comammox Nitrospira amoA gene sequences. The reference sequences are from the GenBank. The sequence and its corresponding sequence number (consisting of the letters and numbers, behind sequence) are listed in a row. Percentages in brackets following the OTUs indicate the percentage of each OTU in the total comammox amoA gene sequences. The calculation method of neighbor-joining phylogenetic tree is Maximum Composite Likelihood, and the scale bar (on the top) represents 20% sequence divergence.
Co-occurrence network analysis of comammox Nitrospira. The nodes represented OTUs, the relationship between two OTUs in the network was shown by an edge. The inferred correlations were restricted to those having correlations > 0.3 or < − 0.3 (P < 0.05, two-sided). Nodes with different colors represent different clades of comammox Nitrospira, and the size of the nodes was proportional to their relative abundance. The bule and red edges represent the positive and negative correlations, respectively, and the thickness of each edge between two nodes is proportional to the value of correlation coefficient. Hub nodes (degree > 20) were labeled in network.
Discussion
Ecological differentiation of two lineages of comammox Nitrospira
In the TGR area of the Yangtze River in China, the amoA gene abundance of comammox Nitrospira clade A in three tributaries and clade B in one tributary was gradually increased along the flow path, but amoA gene abundance of clade B was gradually decreased in two tributaries, indicating that the spatial change of tributaries led to the niche change of comammox Nitrospira. One previous study found that comammox Nitrospira had niche differentiation along the flow path in the main stream of the Yangtze River30. Similarly, the niche differentiation of comammox Nitrospira have also been reported in the estuary ecosystem19.
Landforms and nutrient conditions have been assumed to lead to the differentiation of comammox Nitrospira in the main stream of the Yangtze River30. However, local ammonium concentration might play the main role in the niche differentiation between comammox Nitrospira and canonical AOMs in agricultural ecosystem43. In this study, the abundance of clade A showed a significant positive correlation with the flow path (P < 0.05, Fig. 5), indicating that the spatial change of hydrology and water quality of tributaries might be the main driving force for niche differentiation of this microorganism. In the TGR, the closer the tributary is to the estuary, the slower the flow velocity is since the main stream of Yangtze River has a supporting effect on the tributary. Fine sediments are easy to deposit in the area close to the estuary, resulting in spatial differences in tributary sediments along the flow. In addition, the tributary gradually mixes with the main stream in the process of flowing downstream, thus leading to spatial difference in the water bodies of tributaries along the flow path. These two spatial differences might be the main reason for the niche differentiation of comammox Nitrospira along the flow path in this study.
Among the three tributaries in the TGR area, comammox Nitrospira community also exhibited ecological differentiation. Clade A was the dominant lineage in all three tributaries, of which clade A.1 was the dominant lineage in Xiaojiang River and Xiangxi River, and clade A.2 was the dominant lineage in Daning River. Clade B was dominant in none of the three tributaries. These results showed that different comammox Nitrospira lineages were adapted to the water environment of different tributaries, resulting in niche differentiation. One study found that soil pH was mainly responsible for the differentiation of comammox Nitrospira lineage44, while another study reported that low nutrient environment was the main reason for the differentiation of comammox Nitrospira lineage45. In this study, the difference in hydrological regimes and sediment nutrient conditions might lead to the dominance of clade A.1 and clade A.2 in different tributaries. However, the more specific reasons remain to be further explored in combination with the tributary environment and the physiological characteristics of the two sub-clades.
Coexistence and niche differentiation of Comammox Nitrospira and Canonical AOMs
This study found the coexistence of comammox Nitrospira with AOA and AOB in all sampling sites. Many studies have also revealed that AOMs could coexist in forest soil, agricultural soil, lake sediment, and other environments46,47,48. In this study, amoA gene abundance of comammox Nitrospira clade A was the highest, followed by clade B. The amoA gene abundance of comammox Nitrospira at each sampling site was about one order of magnitude higher than that of canonical AOMs, indicating that comammox Nitrospira might play a major role in the ammonia oxidation process in the sediment of TGR’s tributaries. Comammox Nitrospira have been reported to be dominant in forest, grassland, and agricultural soil environments49,50,51.
The niche differentiation of AOA and AOB in soil environment was mainly determined by ammonia limitation, pH, and mixotrophy2. AOA and AOB had competitive advantages respectively in acidic and high-ammonium concentration environments52. Kinetic and genomic studies showed that comammox Nitrospira was more likely to grow in microaerobic and oligotrophic environments23,24. Moreover, comammox Nitrospira also exhibited a preference for slightly alkaline environments53. In this study, the α-diversity indices of comammox Nitrospira community (either Chao1 index and Observed Species index representing species richness, or Shannon index and 1/Simpson index representing diversity) showed a significant negative correlation with sediment NH4+-N content (P < 0.05, Fig. 5). These evidences showed that comammox Nitrospira were more likely to grow in the ammonium limitation area in the TGR. In addition, niche differentiation among these AOMs led to subsequent environmental effects. The generation of nitrite in two-step nitrification could be avoided in one-step nitrification process, comammox Nitrospira, thus reducing nitrite concentration in the environment. Comammox Nitrospira genomes contain no enzymes related to nitrogen oxide metabolism such as cytochrome c nitric oxide reductase (cNOR), thereby decreasing the risk of N2O production7. Kits et al. (2019) showed that the contribution of Ca. Nitrospira inopinata to N2O emission was similar to that of AOA, but much lower than that of AOB54. Han et al. (2021) also reported that the N2O and NOy production capacity of comammox Nitrospira was only 3 − 15% of AOA and AOB55. More attention should be paid to the subsequent environmental effects induced by these niche differentiations.
Community structure of comammox Nitrospira
The main OTUs in this study exhibited high similarity with the sequences of comammox Nitrospira existing in a variety of freshwater and terrestrial ecosystems (Fig. 6). The sequences of OTU 3, OTU 7, OTU 16, OTU 26, and OTU 58 in clade A.1 were extremely similar to the sequences of comammox Nitrospira in the sediments of river, reservoir and tidal flat56. In addition, OTU 7 and OTU 124 in the same lineage (clade A.1) had high homology with the sequences of comammox Nitrospira from wetland soil of Qinghai-Tibetan plateau and lake sediment, respectively9,48. The sequences of OTU 49, OTU 44, and OTU 11 in clade A.2 had high homology with the sequences of comammox Nitrospira from estuary tidal flat wetland, alpine glacier-forefield soil and lake sediment, respectively10. The sequences of OTU 4 in clade B had high similarity with the sequences of comammox Nitrospira from forest soils (> 95%, 1000 replicates)57.
Phylogenetic analysis also showed that comammox Nitrospira in the sediments of the TGR tributaries had a complex community structure. In all the samples, Clade A and clade B coexisted, and clade A.1 and clade A.2 also coexisted. The coexistence of clade A and clade B has also been reported in tidal flat wetland and agricultural soil12,45. However, only clade A was found in tidal sediments, plateau wetland and estuarine sediments56,58,59. Our data showed the coexistence of clade A.1, clade A.2, and clade B in the sediment from the same tributary in the TGR area, indicating that the TGR was suitable for the survival of comammox Nitrospira. Co-occurrence network analysis showed that clade A.2 was critical to comammox Nitrospira community, which was similar to the conclusion reported in plateau wetland sediments and forest soils44,59. Comammox Nitrospira may play an important role in the nitrogen cycle of the TGR. However, its ammonia oxidation effect needs to be further studied.
Conclusions
This study revealed that comammox Nitrospira widely existed in tributary sediments of the TGR, meanwhile, clade A and clade B existed simultaneously. The amoA gene abundance of clade A and clade B exhibited a spatial change trend of gradual increase or decrease along the flow path. The amoA abundance of comammox Nitrospira clade A in the sediments showed a significant positive correlation with the flow distance between sampling sites. The two sub-clades of clade A were dominant in different tributaries, indicating that clade A had niche differentiation among different tributaries. These findings confirmed that niche differentiation of comammox Nitrospira community appeared in the tributary of the TGR and clade A.2 played an important role in comammox Nitrospira community.
Data availability
The nucleotide sequence of comammox Nitrospira amoA gene obtained in this study was submitted to the GenBank database with the Accession Number of MZ669776 – MZ669807.
References
Kuypers, M. M. M., Marchant, H. K. & Kartal, B. The microbial nitrogen-cycling network. Nat. Rev. Microbiol. 16(5), 263–276 (2018).
Prosser, J. I. & Nicol, G. W. Archaeal and bacterial ammonia-oxidisers in soil: the quest for niche specialisation and differentiation. Trends Microbiol. 20(11), 523–531 (2012).
Costa, E., Pérez, J. & Kreft, J.-U. Why is metabolic labour divided in nitrification?. Trends Microbiol. 14(5), 213–219 (2006).
van Kessel, M. A. H. J. et al. Complete nitrification by a single microorganism. Nature 528(7583), 555–559 (2015).
Daims, H. et al. Complete nitrification by Nitrospira bacteria. Nature 528(7583), 504–509 (2015).
Pinto, A. J. et al. Metagenomic evidence for the presence of comammox Nitrospira -like bacteria in a drinking water system. mSphere 1(1), e0005415. https://doi.org/10.1128/mSphere.00054-15 (2016).
Palomo, A. et al. Comparative genomics sheds light on niche differentiation and the evolutionary history of comammox Nitrospira. ISME J. 12(7), 1779–1793 (2018).
Xia, F. et al. Ubiquity and diversity of complete ammonia oxidizers (Comammox). Appl. Environ. Microbiol. 84(24), e01390-e1418. https://doi.org/10.1128/AEM.01390-18 (2018).
Zhang, S. et al. Ammonia oxidizers in river sediments of the Qinghai-Tibet Plateau and their adaptations to high-elevation conditions. Water Res. 173, 115589. https://doi.org/10.1016/j.watres.2020.115589 (2020).
Xu, Y. et al. Diversity and abundance of comammox bacteria in the sediments of an urban lake. J. Appl. Microbiol. 128(6), 1647–1657 (2020).
Wang, S. et al. Abundance and functional importance of complete ammonia oxidizers and other nitrifiers in a riparian ecosystem. Environ. Sci. Technol. 55(8), 4573–4584 (2021).
Sun, D. et al. Distribution and diversity of Comammox Nitrospira in Coastal Wetlands of China. Front. Microbiol. 11, 589286. https://doi.org/10.3389/fmicb.2020.589268 (2020).
Wang, X. et al. Comammox bacterial abundance, activity, and contribution in agricultural rhizosphere soils. Sci. Total Environ. 727, 138563. https://doi.org/10.1016/j.scitotenv.2020.138563 (2020).
Liu, Z. et al. Temperature and salinity drive comammox community composition in mangrove ecosystems across southeastern China. Sci. Total Environ. 742, 140456. https://doi.org/10.1016/j.scitotenv.2020.140456 (2020).
Wang, Y. et al. Comammox in drinking water systems. Water Res. 116, 332–341. https://doi.org/10.1016/j.watres.2017.03.042 (2017).
Annavajhala, M. K., Kapoor, V., Santo-Domingo, J. & Chandran, K. Comammox functionality identified in diverse engineered biological wastewater treatment systems. Environ. Sci. Technol. Lett. 5(2), 110–116 (2018).
Shao, Y.-H. & Wu, J.-H. Comammox Nitrospira species dominate in an efficient partial nitrification-anammox bioreactor for treating ammonium at low loadings. Environ. Sci. Technol. 55(3), 2087–2098 (2021).
Wang, Y., Zhao, R., Liu, L., Li, B. & Zhang, T. Selective enrichment of comammox from activated sludge using antibiotics. Water Res. 197, 117087. https://doi.org/10.1016/j.watres.2021.117087 (2021).
Wang, X. et al. Niche differentiation of comammox Nitrospira in the mudflat and reclaimed agricultural soils along the north branch of Yangtze river estuary. Front. Microbiol. 11, 618287. https://doi.org/10.3389/fmicb.2020.618287 (2021).
Wang, Z. et al. Comammox Nitrospira clade B contributes to nitrification in soil. Soil Biol. Biochem. 135, 392–395. https://doi.org/10.1016/j.soilbio.2019.06.004 (2019).
Xu, S. et al. Ubiquity, diversity, and activity of comammox Nitrospira in agricultural soils. Sci. Total Environ. 706, 135684. https://doi.org/10.1016/j.scitotenv.2019.135684 (2020).
Zhu, G. et al. Towards a more labor-saving way in microbial ammonium oxidation: a review on complete ammonia oxidization (comammox). Sci. Total Environ. 829, 154590. https://doi.org/10.1016/j.scitotenv.2022.154590 (2022).
Kits, K. D. et al. Kinetic analysis of a complete nitrifier reveals an oligotrophic lifestyle. Nature 549(7671), 269–272 (2017).
Koch, H., van Kessel, M. A. H. J. & Lücker, S. Complete nitrification: insights into the ecophysiology of comammox Nitrospira. Appl. Microbiol. Biotechnol. 103(1), 177–189 (2019).
Palomo, A. et al. Metagenomic analysis of rapid gravity sand filter microbial communities suggests novel physiology of Nitrospira spp. ISME J. 10(11), 2569–2581 (2016).
He, Q., Peng, S., Zhai, J. & Xiao, H. Development and application of a water pollution emergency response system for the Three Gorges Reservoir in the Yangtze River China. J. Environ. Sci. 23(4), 595–600 (2011).
Xiang, R., Wang, L., Li, H., Tian, Z. & Zheng, B. Water quality variation in tributaries of the Three Gorges Reservoir from 2000 to 2015. Water Res. 195, 116993. https://doi.org/10.1016/j.watres.2021.116993 (2021).
Xiang, R., Wang, L., Li, H., Tian, Z. & Zheng, B. Temporal and spatial variation in water quality in the Three Gorges Reservoir from 1998 to 2018. Sci. Total Environ. 768, 144866. https://doi.org/10.1016/j.scitotenv.2020.144866 (2021).
Zhu, L., Xu, Q., Zhang, O., Yang, Y. & Wang, W. Sedimentation at estuary of 66 tributaries in the Three Gorges Reservoir. Sci. Sin. Technol. 49(5), 552–564 (2019).
Liu, S. et al. Comammox Nitrospira within the Yangtze River continuum: community, biogeography, and ecological drivers. ISME J. 14(10), 2488–2504 (2020).
Bartelme, R. P., McLellan, S. L. & Newton, R. J. Freshwater recirculating aquaculture system operations drive biofilter bacterial community shifts around a stable nitrifying consortium of ammonia-oxidizing archaea and comammox Nitrospira. Front. Microbiol. 8, 101. https://doi.org/10.3389/fmicb.2017.00101 (2017).
Rognes, T., Flouri, T., Nichols, B., Quince, C. & Mahé, F. VSEARCH: a versatile open source tool for metagenomics. PeerJ 4, e2584. https://doi.org/10.7717/peerj.2584 (2016).
Wang, Q. et al. Ecological patterns of nifH Genes in four terrestrial climatic zones explored with targeted metagenomics using framebot, a new informatics tool. MBio 4(5), 1–9. https://doi.org/10.1128/mBio.00592-13 (2013).
Kumar, S., Stecher, G., Li, M., Knyaz, C. & Tamura, K. MEGA X: Molecular evolutionary genetics analysis across computing platforms. Mol. Biol. Evol. 35(6), 1547–1549 (2018).
Tamura, K., Nei, M. & Kumar, S. Prospects for inferring very large phylogenies by using the neighbor-joining method. Proc. Natl. Acad. Sci. 101(30), 11030–11035 (2004).
Jiang, R. et al. Use of newly designed primers for quantification of complete ammonia-oxidizing (Comammox) bacterial clades and strict nitrite oxidizers in the genus Nitrospira. Appl. Environ. Microbiol. 86(20), e01775-e1820. https://doi.org/10.1128/AEM.01775-20 (2020).
Francis, C. A., Roberts, K. J., Beman, J. M., Santoro, A. E. & Oakley, B. B. Ubiquity and diversity of ammonia-oxidizing archaea in water columns and sediments of the ocean. Proc. Natl. Acad. Sci. 102(41), 14683–14688 (2005).
Meinhardt, K. A. et al. Evaluation of revised polymerase chain reaction primers for more inclusive quantification of ammonia-oxidizing archaea and bacteria. Environ. Microbiol. Rep. 7(2), 354–363 (2015).
R Core Team. R: A language and environment for statistical computing. R Foundation for Statistical Computing, Vienna, Austria, https://www.R-project.org/ (2020).
Caporaso, J. G. et al. QIIME allows analysis of high-throughput community sequencing data. Nat. Methods 7(5), 335–336 (2010).
Kurtz, Z. D. et al. Sparse and compositionally robust inference of microbial ecological networks. PLOS Comput. Biol. 11(5), e1004226. https://doi.org/10.1371/journal.pcbi.1004226 (2015).
Weiss, S. et al. Correlation detection strategies in microbial data sets vary widely in sensitivity and precision. ISME J. 10(7), 1669–1681 (2016).
He, S. et al. Ammonium concentration determines differential growth of comammox and canonical ammonia-oxidizing prokaryotes in soil microcosms. Appl. Soil Ecol. 157, 103776. https://doi.org/10.1016/j.apsoil.2020.103776 (2021).
Li, C., Hu, H.-W., Chen, Q.-L., Chen, D. & He, J.-Z. Niche differentiation of clade A comammox Nitrospira and canonical ammonia oxidizers in selected forest soils. Soil Biol. Biochem. 149, 107925. https://doi.org/10.1016/j.soilbio.2020.107925 (2020).
Wang, J., Wang, J., Rhodes, G., He, J.-Z. & Ge, Y. Adaptive responses of comammox Nitrospira and canonical ammonia oxidizers to long-term fertilizations: Implications for the relative contributions of different ammonia oxidizers to soil nitrogen cycling. Sci. Total Environ. 668, 224–233. https://doi.org/10.1016/j.scitotenv.2019.02.427 (2019).
Pjevac, P. et al. AmoA-targeted polymerase chain reaction primers for the specific detection and quantification of comammox Nitrospira in the environment. Front. Microbiol. 8, 1508. https://doi.org/10.3389/fmicb.2017.01508 (2017).
Orellana, L. H., Chee-Sanford, J. C., Sanford, R. A., Löffler, F. E. & Konstantinidis, K. T. Year-round shotgun metagenomes reveal stable microbial communities in agricultural soils and novel ammonia oxidizers responding to fertilization. Appl. Environ. Microbiol. 84, e01646-e1717. https://doi.org/10.1128/AEM.01646-17 (2018).
Xu, Y. et al. The diversity of comammox bacteria and the effect of sewage discharge on their abundance in eutrophic lake sediments. J. Soils Sediments 20(5), 2495–2503 (2020).
Hu, J. et al. Dominance of comammox Nitrospira in soil nitrification. Sci. Total Environ. 780, 146558. https://doi.org/10.1016/j.scitotenv.2021.146558 (2021).
Li, C., Hu, H. W., Chen, Q. L., Chen, D. & He, J. Z. Comammox Nitrospira play an active role in nitrification of agricultural soils amended with nitrogen fertilizers. Soil Biol. Biochem. 138, 107609. https://doi.org/10.1016/j.soilbio.2019.107609 (2019).
Osburn, E. D. & Barrett, J. E. Abundance and functional importance of complete ammonia-oxidizing bacteria (comammox) versus canonical nitrifiers in temperate forest soils. Soil Biol. Biochem. 145, 107801. https://doi.org/10.1016/j.soilbio.2020.107801 (2020).
Verhamme, D. T., Prosser, J. I. & Nicol, G. W. Ammonia concentration determines differential growth of ammonia-oxidising archaea and bacteria in soil microcosms. ISME J. 5(6), 1067–1071 (2011).
Wang, D.-Q., Zhou, C.-H., Nie, M., Gu, J.-D. & Quan, Z.-X. Abundance and niche specificity of different types of complete ammonia oxidizers (comammox) in salt marshes covered by different plants. Sci. Total Environ. 768, 144993. https://doi.org/10.1016/j.scitotenv.2021.144993 (2021).
Kits, K. D. et al. Low yield and abiotic origin of N2O formed by the complete nitrifier Nitrospira inopinata. Nat. Commun. 10, 1836. https://doi.org/10.1038/s41467-019-09790-x (2019).
Han, P. et al. N2O and NOy production by the comammox bacterium Nitrospira inopinata in comparison with canonical ammonia oxidizers. Water Res. 190, 116728. https://doi.org/10.1016/j.watres.2020.116728 (2021).
Yu, C. et al. Evidence for complete nitrification in enrichment culture of tidal sediments and diversity analysis of clade a comammox Nitrospira in natural environments. Appl. Microbiol. Biotechnol. 102(21), 9363–9377 (2018).
Reay, D. S., Radajewski, S., Murrell, J. C., McNamara, N. & Nedwell, D. B. Effects of land-use on the activity and diversity of methane oxidizing bacteria in forest soils. Soil Biol. Biochem. 33(12–13), 1613–1623 (2001).
Jiang, Q., Xia, F., Zhu, T., Wang, D. & Quan, Z. Distribution of comammox and canonical ammonia-oxidizing bacteria in tidal flat sediments of the Yangtze River estuary at different depths over four seasons. J. Appl. Microbiol. 127(2), 533–543 (2019).
Yuan, D. et al. Comammox activity dominates nitrification process in the sediments of plateau wetland. Water Res. 206, 117774. https://doi.org/10.1016/j.watres.2021.117774 (2021).
Acknowledgements
Great gratitude goes to linguistics Prof. Ping Liu from Huazhong Agriculture University, Wuhan, China for her work at English editing and language polishing.
Funding
This study was supported by the National Natural Science Foundation of China (No. U2040211, No. U1802241, No. 92047204, No. 92047203, and No. 42177383), the National Key Research and Development Project (2021YFC3201002), the project of China Three Gorges Corporation (No. 201903144), China Institute of Water Resources and Hydropower Research (SKL2020TS07), and Follow-up Work of the Three Gorges Project (2136902).
Author information
Authors and Affiliations
Contributions
J.H.Z.: Methodology, Investigation, Formal analysis, Data curation, Writing-original draft. M.H.: Investigation, Writing-review & editing, Funding acquisition. Y.W: Conceptualization, Visualization, Funding acquisition. J.W.Z: Conceptualization, Visualization, Writing-review & editing, Project administration, Funding acquisition. S.L.: Writing—review & editing. Y.B.: Visualization, Funding acquisition. J.W.: Methodology, Investigation. J.H.: Methodology, Writing—review & editing. M.Z.: Writing—review & editing. All authors reviewed the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Zhang, J., Hu, M., Wang, Y. et al. Niche differentiation of comammox Nitrospira in sediments of the Three Gorges Reservoir typical tributaries, China. Sci Rep 12, 6820 (2022). https://doi.org/10.1038/s41598-022-10948-9
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-022-10948-9
- Springer Nature Limited
This article is cited by
-
Soil health improvement in a karst area with geogenic Cd enrichment using biochar and clay-based amendments
Journal of Soils and Sediments (2024)
-
Comammox Nitrospira clade A recovers faster than clade B after cyanobacteria outbreak in the Three Gorges tributary, China
Journal of Soils and Sediments (2023)