Abstract
Ubiquitylation is critical for preventing aberrant DNA repair and for efficient maintenance of genome stability. As deubiquitylases (DUBs) counteract ubiquitylation, they must have a great influence on many biological processes, including DNA damage response. To elucidate the role of DUBs in DNA repair in Drosophila melanogaster, systematic siRNA screening was applied to identify DUBs with a reduced survival rate following exposure to ultraviolet and X-ray radiations. As a secondary validation, we applied the direct repeat (DR)-white reporter system with which we induced site-specific DSBs and affirmed the importance of the DUBs Ovarian tumor domain-containing deubiquitinating enzyme 1 (Otu1), Ubiquitin carboxyl-terminal hydrolase 5 (Usp5), and Ubiquitin carboxyl-terminal hydrolase 34 (Usp34) in DSB repair pathways using Drosophila. Our results indicate that the loss of Otu1 and Usp5 induces strong position effect variegation in Drosophila eye following I-SceI-induced DSB deployment. Otu1 and Usp5 are essential in DNA damage-induced cellular response, and both DUBs are required for the fine-tuned regulation of the non-homologous end joining pathway. Furthermore, the Drosophila DR-white assay demonstrated that homologous recombination does not occur in the absence of Usp34, indicating an indispensable role of Usp34 in this process.
Similar content being viewed by others
Introduction
DNA damage-induced cellular response is a primary mechanism to protect cells against cancerous malformation. To maintain genomic integrity, this process is critical for the recruitment and activation of specific proteins, including writers, readers, and erasers of the epigenetic marks, implicated in the DNA repair-related rearrangement of the chromatin structure. These posttranslational modifications (PTMs) contribute to the fine-tuned regulation of the proteins involved in the DNA damage response (DDR) and the subsequent repair processes1,2,3. These modifications are also critical for the appropriate recruitment of downstream repair proteins to modulate the effectiveness and dynamics of DNA repair4.
Among these PTMs, ubiquitylation is one of the well described mechanisms, involving a cascade of activating, conjugating, and ligating enzymes. The ubiquitin enzyme cascade catalyzes the formation of an isopeptide bond between the C-terminal glycine (G) residue of the ubiquitin molecule and lysine (K) residue(s) of the target proteins. Removal of ubiquitin groups is catalyzed by deubiquitylases (DUBs), and the mechanism has received increasing attention for understanding of their importance in DNA repair5,6. In recent years, several DUBs have already been described to participate in DNA repair. In U2OS cells, human ubiquitin-specific peptidase 1 (USP1) was demonstrated to be essential to promote homologous recombination (HR)7,8. Furthermore, human ubiquitin-specific peptidase 3 (USP3) is implicated in the deubiquitylation of histone H2A/H2B and is required for the efficient resolution of replication-coupled stress conditions9. In addition, in human cells, USP4, USP5, USP7, and USP28 were shown to be involved in controlling the stability of p53 during DNA repair10,11,12,13. USP20 regulates the Ataxia telangiectasia and Rad3-related protein (ATR)-related DDR, whereas USP7 and USP47 facilitate Ataxia telangiectasia mutated (ATM)-related DDR13,14,15. USP7 together with USP34 stabilizes Ring Finger Protein 168 (RNF168) implicated in H2A monoubiquitylation16,17. Moreover, the human ovarian tumor domain-containing protein 4 (OTUD4) is implicated in the regulation of DNA repair through an ATM/p53-mediated pathway18, whereas OTU Deubiquitinase, OTU domain-containing ubiquitin aldehyde-binding protein 1 (OTUB1) and OTU Deubiquitinase, OTU domain-containing ubiquitin aldehyde-binding protein 2 (OTUB2) are essential for the DNA damage-induced foci formation19,20,21. On the basis of the expanding knowledge on the DNA damage-induced ubiquitylation process, different ubiquitin linkages have major impacts on the choice of the appropriate repair mechanism. Interconnection between DUBs and other PTMs has been also revealed. For instance, studies have elucidated the crosstalk between poly(ADP-ribosylation) polymerases and DUBs, but its exact mechanism still remains unclear 22,23. These data emphasize the importance of the characterization of DUBs which have been reported to be involved in DNA repair.
For elucidating the function of these enzymes and identifying new DUB players implicated in DNA repair, ultraviolet (UV) and X-ray irradiation-based siRNA screenings were applied to identify novel DUBs that may be recruited to DNA lesions. We identified five DUBs (Usp14, Usp34, Usp45, Usp47, and Otu1), which exhibit synthetic lethality following UV irradiation, of those we tested. In addition, after X-ray irradiation, the absence of Usp5 and Otu1 caused more defects in the imaginal disc development of Drosophila than those found in the wild-type strain, suggesting the regulatory roles of Otu1 and Usp5 in the ionizing irradiation-induced repair pathways. In a secondary approach, we applied a DSB repair reporter assay which generates I-SceI endonuclease-induced DSBs in euchromatic and heterochromatic regions. During these analyses, we affirmed the importance of Usp34 DUB in DSB repair pathways in Drosophila and we further determined the effect of these DUBs. These findings provide novel insights into the functional link between DUBs and DNA repair.
Results
DUBs are implicated in the regulation of UV-induced DNA repair
For identifying DUBs in Drosophila melanogaster, we applied bioinformatics homology search analysis using the sequences of known yeast and human DUBs24,25. On the basis of our search results, we identified 44 DUBs that were classified into the following DUB subgroups: 23 Ubiquitin specific proteases (USPs), 4 ubiquitin C-terminal hydrolases (UCHs), 1 Machado-Joseph Disease (MJD), 7 ovarian tumour deubiquitinases (OTUs), and 9 JAB1/MPN/Mov34 metalloenzymes (JAMMs) (Table 1). To identify the putative DUBs that might be implicated in DNA repair, we conducted a DUB siRNA screening and, in some cases, such as Otu1, Usp5 and not, we also used Drosophila null mutant stocks. We used individual D. melanogaster transgenic lines, which enabled the siRNA silencing of the 44 different DUBs by the daughterless-galactose-responsive transcription factor (da-GAL4) driver (Table 1). As the readout of the screening, we evaluated the viability of L3 larvae and their pupation rate after exposure to UV and X-ray irradiation. We first investigated the sensitivity of DUB siRNA and null mutant Drosophila L3 larvae following exposure to 15 mJ/cm2 UV. Then, the irradiated larvae were collected and maintained for 24 h on a standard medium, and the survived larvae were counted (Supplementary Fig. 1). Of the DUBs we tested, usp47 (CG5486), usp34 (CG5794), usp14 (CG5384), Otu1 (CG4603), and usp16–45 (CG4165) DUB mutant Drosophila stocks demonstrated a significant difference in viability compared with the control (w1118). The most significant difference was measured in the case of Usp16–45, Usp14 siRNA-silencing D. melanogaster strains and Otu11 (w+;+;CG4603Δ101/2/CG4603Δ101/2) Drosophila null mutants, whereas the suppression of Usp34 and Usp47 resulted in milder, but still significant difference compared with those of the control (Fig. 1a). This experiment indicated that several DUBs play an indispensable role in the UV-induced DNA repair pathways.
Survival rate of DUB mutant L3 larvae in response to UV irradiation. (a) The mean values of the survival rate of the UV-irradiated DUB mutant L3 Drosophila larvae are represented. L3 larvae were exposed to 15 mJ/cm2 UV. (b) Heterozygous Usp51/TM6b GFP and not[P]/TM6b GFP DUB mutant, L3 larvae were irradiated with 15 mJ/cm2 UV. For the statistical test, one-way ANOVA in combination with Tukey's post-hoc test was used. Significant values are indicated as follows: **P < 0.01, ***P < 0.001, ****P < 0.0001. Data were obtained from three independent experiments.
Previous studies demonstrated that the loss of Usp5 causes lethality in the L3 larval stage26,27. Furthermore, the da-Gal4-driven Usp5RNAi (17657) exhibited subsequent lethality, but primarily in the pupa stages27. In addition, the da-Gal4-driven notRNAi 45775 and the not[5-HA-1189] (not[P]) mutant alleles resulted in L2–L3 lethality in homozygous form (Supplementary Fig. 2 and Supplementary Table 1). Therefore, we investigated the Usp51/TM6b GFP (green fluorescent protein) and not[P]/TM6b GFP heterozygous mutant Drosophila stocks and monitored their sensitivity to UV light. Unfortunately, in this case, we did not determine any significant phenotypic changes compared with those in the control (Fig. 1b). These results indicate that several DUBs are required in the UV-induced DSB repair pathway in D. melanogaster. Previous research also demonstrated that USP45 plays a role in the UV-induced DNA repair pathway and USP45-knockout exhibits hypersensitivity upon UV irradiation28. According to our UV screening results, the Usp16–45 DUB mutant demonstrated the highest sensitivity to UV exposure, which is consistent with the results of previous human cell culture-based research28. Interestingly, the Usp47RNAi (103743) Drosophila strain also exhibited decreased viability, which confirmed a previous study that implicated it in the base excision repair (BER) pathway29, whereas Otu1 and Usp14 were found to be involved in DNA double-strand break repair21,30.
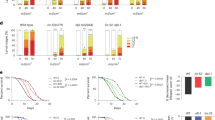
Otu1 and Usp5 are involved in the X-ray-induced DNA double-strand break repair
In the first screening, we identified several DUBs whose functions are essential in the UV-induced DNA repair pathway. In the next step, we evaluated whether these DUBs are also essential for the repair of other types of DNA damage. For this purpose, we applied X-ray irradiation that causes DSBs, and we checked the pupation and eclosion rates of Drosophila harboring DUB mutations. We used Usp51/TM6b GFP and Otu11 (w+;+;CG4603Δ101/2/CG4603Δ101/2) Drosophila mutants which exhibited different sensitivities in response to UV irradiation. The DUB mutant third instar larvae were irradiated with 40 Gray (Gy) X-ray, and after the treatment we evaluated the survival rate of the larvae. We observed that the 40 Gy dose had no significant effect on either the pupation or eclosion rate of the DUB mutant compared with that of controls (Fig. 2a and b and Supplementary Fig. 3). However, following hatching, we detected abnormal eye and wing phenotypes in Otu11 and Usp51 mutant flies. The wing and eye abnormalities were also detected in w1118 flies, but the occurrence of these defects was significantly higher in DUB mutants (Fig. 2c,d). Interestingly, although the X-ray irradiation directly caused DSBs, we found that the Otu1 mutant flies were more sensitive to UV irradiation than to X-ray exposure compared with the control (Figs. 1a and 2a–c). These results indicated that the loss of Otu1 resulted in a more severe phenotype in the UV-induced DNA repair pathways than in the X-ray irradiation-induced DSB repair pathway, suggesting a diverse function of Otu1. However, the Usp51 mutant flies showed no significant differences subsequently to either UV or X-ray irradiation, although the occurrence of eye and wing abnormalities was significantly higher after the ionizing irradiation, indicating the putative regulatory but dispensable role of Usp5 in DSB repair pathways.
Effect of X-ray irradiation on the homozygous Otu11 (w+; + ;CG4603Δ101/2/CG4603Δ101/2) mutant and the heterozygous Usp51 mutant (w+; + ;Usp51/TM6b GFP) Drosophila. (a) and (b) L3 DUB mutant larvae were irradiated with 40 Gy X-ray, and the rates of pupation and eclosion were measured (a and b, respectively). (c) The percentage of wing and eye abnormalities was represented. In both the mutants and the control lines, the number of Drosophila with abnormale eye and wing phenotype was compared to the number of the hatched Drosophila which survived the X-ray treatment. (d) The detected wing and eye defects in adult DUB mutants are depicted. For the statistical analysis, one-way ANOVA in combination with Tukey’s post-hoc test was used. Significant values are indicated as follows: *P < 0.05, ***P < 0.001, ****P < 0.0001. Data were obtained from three independent experiments.
Usp34 is essential for euchromatic site-specific DSB repair in DR-white system
Our results demonstrated that Otu1 and Usp5 play an indispensable role in the damage-induced DNA repair pathways. DSB formation may also be induced by inaccurately repaired T-T dimers caused by UV exposure. Regarding the cell cycle phase, DSBs can induce either non-homologous end joining (NHEJ) or HR, which are the two predominant DNA double-strand breaks repair (DSBR) pathways. In addition, DUBs have already been shown to be involved in DNA repair pathways through the deubiquitylation of regulatory proteins involved in DDR, NHEJ, or HR. To examine whether Otu1 is in fact required for HR, we used the previously mentioned DR-white assay to investigate the homology-directed repair frequency in the absence of this DUB. The Drosophila DR-white assay was developed by Anthony T. Do and Joseph T. Brooks et al., which allows determining the pathway choice among HR, NHEJ, and SSA during DSBR31. This in vivo system contains two nonfunctional repeats of the white gene, which enclose the yellow (y+) transgene: (I) the upstream part, referred to as Sce.white, and contains I-SceI recognition sequences, and (II) the downstream non-functional white repeat, which due to the truncation does not possess either the 5′ and 3′ UTRs or the promoter and transcription start sequences of the white gene. The heat-shock-induced expression of I-SceI generates site-specific DSBs at I-SceI recognition sites and activates DSBR. In the progenies, we could monitor the phenotypic changes caused by the activation of the appropriate DNA repair pathway. In the yw genetic background, the brown bodies and white eyes indicated no DSB induction, HR, or NHEJ repair. If the more accurate HR repair occurs, the I-SceI recognition site will be repaired from the adjacent iwhite gene, resulting in red-eye progenies. The DR-white transgenes were integrated in both euchromatic and heterochromatic regions, and therefore, the site-specific DSB repair events can be measured in both regions (Supplementary Fig. 4). In our experiment, we investigated whether the HR-related recombination frequency was affected in the absence of DUBs. For this purpose, we used the Otu1RNAi (21894) (yw; DR-white.y+ (DR-white_2eu_1)/hsp70.HA.I-Sce, Sco; da-GAL4/Otu1RNAi (21894)) and Usp34RNAi (27517) RNAi stocks (yw; DR-white.y+ (DR-white_2eu_1)/hsp70.HA.I-Sce, Sco; da-GAL4/Usp34RNAi (27517)) that also contained the heat-shock-inducible I-SceI-coding gene and the DR-white transgene on the 2nd chromosome. We also applied the da-GAL4 driver to induce the depletion of Otu1 and Usp34. As a control, we used the yw; DR-white.y+ (DR-white_2eu_1)/hsp70.HA.I-Sce,Sco; da-GAL4/+ Drosophila strain that contains the same DR-white and I-SceI transgenes on the 2nd chromosome and the da-GAL4 driver on the 3rd chromosome. The euchromatic DR-white assay showed that the ratio of red-eye offsprings was much lower in the Usp34RNAi (27517) than in the control strain. However, in the absence of Otu1, we found no significant differences in HR efficiency compared with the control (Fig. 3). These results suggested that Usp34 is an important regulator of HR repair in D. melanogaster. Previous research demonstrated that OTU protein family members in human cells can regulate RNF168 activity which together with RNF8 are important regulators of DDR, although the loss of Drosophila Otu1 did not result in any difference in HR efficiency compared with the control. We also did not detect any Drosophila adults with mosaic red-eye color in the heterochromatic DR-white assay in either the control or the RNAi DUB Drosophila strain (data not shown). This result indicated that either I-SceI is not effective in the heterochromatic region with regard to site-specific DSB induction or these DUBs are not essential for repairing heterochromatic DSBs.
Phenotypic changes were obtained using I-SceI-induced site-specific DSB system in Otu1RNAi (21894) and Usp34RNAi (27517) DUB mutants (DR-white assay). The DR-white assay allows determining the choice of the repair pathway following I-Sce1-induced site-specific DSBs. Otu1 and Usp34 RNAi transgenes were driven by da-GAL4, the L3 larvae were heat-shocked to induce site-specific DSBs in the DR-white transgenes, and the mosaic red-eye phenotype was detected. Representation of the ratio of red-eye phenotype applying the euchromatic DR-white system. In case of Otu1RNAi (21894) we used heat-shock-induced yw; DR-white.y + (DR-white_2eu_1)/hsp70.HA.I-Sce, Sco; da-GAL4/Otu1RNAi (21894) adult flies and compared the number of flies with red eye to the total number of the flies with the above-described genotype. In case of Usp34RNAi (27517) we used the same quantification by measuring the red-eye mosaic phenotype. The heat-shock-induced yw; DR-white.y + (DR-white_2eu_1)/hsp70.HA.I-Sce, Sco; da-GAL4/Usp34RNAi (27517), flies were counted and the mosaic red-eye adult flies were compared to the total number of the adult flies with the above-described genotype. For the control, we used yw; DR-white.y + (DR-white_2eu_1)/hsp70.HA.I-Sce, Sco; da-GAL4/+ Drosophila lines with the same quantification method as was used for Usp34RNAi (27517) and Otu1RNAi (21894). For the statistical analysis, Welch's ANOVA in combination with Dunnett's T3 post-hoc test was used. Significant values are indicated as follows: *P < 0.05, **P < 0.01, ****P < 0.0001. Data were obtained from nine independent experiments. More details about the DR-white system are described in a previous study31.
Discussion
Reversible PTMs play pivotal roles during the regulation of DNA repair pathways. Among these PTMs, ubiquitylation is not only involved in controlling the stability of DNA repair proteins, but it also mediates the recruitment of additional repair proteins to the damaged locus. Although several E3 ubiquitin ligases have been shown to be involved in K48- or K63-linked polyubiquitylation of specific DNA repair proteins, the DUBs that remove ubiquitin moieties from DNA repair proteins are poorly characterized32. For identifying novel DUBs implicated in DNA repair, we conducted an RNAi screening using specific DUB siRNA-expressing Drosophila strains to explore their function in UV-, X-ray-, and site-specific I-SceI-induced DNA damage. Among the DUB strains we tested, we found that Usp47, Usp34, Usp14, Otu1, Usp16–45, Usp51, and not[P] DUB mutant Drosophila strains exhibited severe UV light-induced phenotype compared with the control. We identified a milder effect in Usp51 and not[P] DUB mutants, indicating that these DUBs are required, but probably dispensable for the appropriate repair of UV light-induced DNA damage in D. melanogaster. Furthermore, the silencing of Usp47, Usp34, Usp4, and Otu1 was fatal in these larvae, and Usp16–45-silencing resulted in the most robust phenotype alteration compared with the control.
Consistently, earlier studies also revealed that USP45 is important in the UV-induced DNA repair pathways, especially in the deubiquitylation of Excision repair cross-complementation group 1 (ERCC1), which is the subunit of Xeroderma pigmentosum, complementation group F-Excision repair cross-complementation group 1 (XPF-ERCC1) implicated in Transcription-Coupled Nucleotide Excision Repair (TC-NER) regulation28,33. These data also supported the sensitivity of our screening in which we identified additional DUBs, including Usp47, Usp34, Usp14, and Otu1, required for the UV-induced DNA damage-related repair processes.
Although the Usp47 mutant larvae showed less sensitivity to UV irradiation compared with larvae with the loss of other DUBs, Usp47 may affect the survival rate upon UV irradiation through a different pathway. As demonstrated by Parsons et al. that USP47 regulates the level of Pol β in BER, the lower sensitivity of USP47 can be explained by the diverse role of the desired DUBs in the DNA repair pathways29. This assumption is further supported by the fact that USP34 and USP14 are also implicated in the regulation of the HR pathway, whereas OTU1/OTUB1 mediates the RNF168-dependent K63-linked polyubiquitylation17,20,21,30,34.
All these results are consistent with our findings showing that several DUBs, which we identified in the UV screening, have already been demonstrated to be essential in other DNA repair pathways than NER, emphasizing the specificity of the screening, and that these proteins are in fact essential players in the UV-induced DNA repair pathways.
As a next step, we induced DSBs using ionizing irradiation to explore the role of Otu1 and Usp5 in DSB repair. We found that X-ray irradiation did not affect the survival rate of Drosophila larvae in either Otu11 or Usp51 DUB mutant Drosophila, but eye and wing abnormalities were detected at a significantly higher rate than that in the control. These results suggest that the absence of Otu1 and Usp5 does not play an essential role in the X-ray-induced DNA repair pathway, but it probably decreases the efficiency of the repair itself. Our hypothesis is that it delays the repair in fast-dividing diploid tissues, which resulted in milder phenotypic abnormalities, but does not affect the overall viability of the flies. The Drosophila Otu1 and its human homology (YOD1) are poorly characterized in DNA repair pathways, but previous studies have demonstrated that both the OTU family members of DUBs such as OTUB1 and USP5 play a role in DSB repair, especially in HR21,35. It was also demonstrated that in the absence of USP5, γH2AX removal is delayed from the DSB sites in HR and the improper HR increases the sensitivity against DSB-induced agents35,36. This finding also supports our results that in the absence of Usp5 and Otu1, DNA repair is still functional, but less effective in DSB repair. In our system, the inappropriate HR repair might result in the appearance of phenotypic alterations detected in Drosophila tissues. To verify this phenomenon, we used the DR-white D. melanogaster assay which is an I-SceI meganuclease-based site-specific DSB-inducer system. This experimental setup depends on the white transgene that causes mosaic red–white eye color in Drosophila when DSBR occurs. Using this system, we detected significantly lower red-eye-color-containing progenies in the case of Usp34 silencing, whereas the loss of Otu1 did not cause any significant difference compared with the control. Earlier, it was also demonstrated that OTU family members of proteins (OTUB1 and OTUB2) can differently influence the pathway choice. The depletion of OTUB2 suppresses the HR pathway through the RNF8-mediated ubiquitylation; in contrast, the depletion of OTUB1 caused increased HR through the inhibition of ATM21. Furthermore, USP34 can stabilize RNF168 through deubiquitylation, and in the absence of USP34, the level of RNF168 is decreased17. This effect can influence the pathway choice through the accumulation of 53BP1 at DSB sites37. We also observed that the absence of DUBs can influence the pathway choice in Drosophila differently. We showed that HR was not suppressed with the decreased level of Otu1, but the silencing of Usp34 can inhibit HR, indicating a diverse role of the two DUBs in the HR pathways as well.
In our experimental setup, we used various types of DNA-damaging agents, such as UV irradiation, ionizing radiation, and oxidative damage, which can cause different types of DNA lesions. For repairing these DNA lesions, cells have evolved various well regulated DNA repair pathways. Since the activation of an inappropriately chosen DNA repair circuit can be harmful for the cells, these pathways are tightly regulated by PTMs, such as ubiquitylation, and therefore, DUBs can be key regulators of these processes. Our screening demonstrated the diverse function and importance of several DUBs in the UV- and X-ray irradiation-induced DSB repair pathways. We observed that the absence of Usp47, Usp34, Usp14, Otu1, and Usp45 can affect the survival rate of Drosophila after UV irradiation, indicating an important role of these DUBs in DNA repair. We also demonstrated that Otu1, Usp5, and Usp34 play a regulatory, but dispensable role in DSBR. These results emphasize the importance of using multicellular in vivo model system to study the function of DUBs implicated in DNA repair, and the phenotypic changes observed in our system can provide a more sensitive method for identifying novel DUBs implicated in NER and DSBR.
Materials and methods
Identification of DUBs in D. melanogaster
In the first step, the sequences of already known yeast and human DUBs were used for the identification of Drosophila DUBs, which have been reviewed by Amerik et al. and Nijman et al.24,25. Moreover, a preliminary bioinformatics analysis was performed to identify the DUB-sequences in Drosophila based on the yeast sequence homology. Sequences were downloaded from NCBI, UniProt, and FlyBase, and for identifying the homology sequences, BLAST program was used. EMBOSS Needle Pairwise Sequence Alignment program of EMBL's European Bioinformatics Institute (EMBL-EBI's) allowed comparing the protein sequences, and EMBL-EBI's Interpro Scan program ensured to identify domains in the proteins. This complex identification system enabled the accurate identification of the various DUBs in Drosophila.
UV irradiation screening
The da-GAL4 driver stock was used to diminish the levels of Usp47, Usp34, Usp14, Otu1, and Usp16–45 using transgenic RNAi Drosophila stocks. We applied the da-GAL4 driver in the homozygous form on the 2nd chromosome, and the RNAi transgene was inserted into the 3rd chromosome. All of the RNAi stocks used in this study were kindly provided by Péter Deák (Table 1). The early L3 larvae were UV-irradiated with a dose of 15 mJ/cm2. For measuring the survival rate of the UV-treated larvae, the animals were collected and maintained on standard Drosophila medium. Data were obtained from three independent biological replicates, and in each experiment, 50 larvae were exposed to UV irradiation. The w1118 stock was used as a control (Supplementary Fig. 1).
Drosophila stocks and mutation generation
The fly stocks were maintained on standard medium at 25 °C. The transgenic RNA interference and the P element insertion-containing lines were obtained from Bloomington Drosophila Stock Center, Vienna Drosophila Resource Center, and NIG-Fly National Institute of Genetics. The RNAi lines with Stock IDs are depicted in Table 1. We used the following three null mutant Drosophila lines:
usp5 1 mutant allele carrying Drosophila line
Using P element remobilization technique, the imprecise excision of P{EPgy2}EY20760 generated 275-bp deletion, eliminating the catalytic histidine box and UBA2 domain coding sequence. The deletion causes L3 lethality in homozygous form. More information about the stock is available in Kovács et al.27. In our UV screening, the mutant allele was balanced with TM6b GFP.
PCR not [P] (not [5-HA-1189]) mutation carrying Drosophila line
not[5-HA-1189] mutation causes L2 and L3 lethality in the homozygous form (Supplementary Table 1). We measured the expression of not genes using semi-quantitative reverse transcription–polymerase chain reaction (RT-PCR). We did not detect any expression of not genes in the not[5-HA-1189] homozygous mutant Drosophila larvae compared with the control (Supplementary Fig. 2). The not[5-HA-1189]-containing stock is available in the Drosophila Genetic Resource Center Kyoto with the Stock ID number 125130.
Otu1 1 (CG4603 Δ101/2) mutant allele carrying Drosophila line
For CG4603Δ101/2 deletion, the P element remobilization technique was used. The imprecise excision of P{EPgy2}CG4603(EY07831) generated the CG4603Δ101/2 deletion allele containing 1348-bp deletion which eliminates two-third part of the gene (Fig. 4). We did not detect significant lethality in the homozygous form compared with the control (Supplementary Table 1).
Schematic representation of CG4603 P element insertion and CG4603Δ101/2 deletion allele. The blue triangle represents the P element insertion, and the broken line shows the deleted segment of CG4603 gene. CG4603-RC, CG4603-RB, and CG4603-RA are the transcripts of CG4603. Figure was prepared on the basis of the FlyBase Genome Browser.
Semi-quantitative RT-PCR
Total RNA was extracted from 10 larvae using the Tri Reagent extraction kit (Sigma-Aldrich, USA). RNA samples were treated with RQ1 RNase-free DNase (Promega Corporation, USA). The Thermo Fisher Scientific cDNA synthesis kit was used for reverse transcription, according to the manufacturer’s instruction. Samples were normalized to the rpL17A level using the following primers: RpL17A forward 5′-GAGCCAAGAACCTGTACG-3′ and reverse 5′-CAGGCATGACCTTCTTCC-3′. Gene expressions were determined using 20-25-cycle PCRs with exon-specific primers. The RT-PCR was quantified by using Fiji Image J (Supplementary Fig. 2).
X-ray irradiation
The null mutant Otu11 and Usp51 Drosophila stocks were kindly provided by Péter Deák. Because of the strong L3 lethality, in the case of Usp51, we used w;+; Usp51/TM6b GFP Drosophila larvae for X-ray irradiation. The Otu11 (w;+; CG4603Δ101/2/CG4603Δ101/2) null mutant Drosophila was used as the homozygous mutant form during X-ray irradiation. w1118 stock was used as a control. In case of control (w1118) and DUB mutant Drosophila [Otu11 (w;+; CG4603Δ101/2/CG4603Δ101/2) and Usp51, we used w;+; Usp51/TM6b GFP], the early L3 stage larvae were collected and inserted into the sample holder of the X-ray machine and then exposed to an X-ray dose of 40 Gy (Supplementary Fig. 3). Subsequently to X-ray irradiation, 100 irradiated larvae were collected in each experiment (three independent experiments were performed) and maintained on standard Drosophila medium.
Drosophila DR-white assay
For the euchromatic DR-white assay, yw; DR-white.y+ (DR-white_2eu_2)/CyO; da-GAL4 females were crossed either with yw; hsp70.HA.I-Sce,Sco/CyO; UAS-RNAi CG4603 or with yw; hsp70.HA.I-Sce,Sco/CyO; UAS-RNAi Usp34 males. The embryos from these crosses yw; DR-white.y+ (DR-white_2eu_1)/hsp70.HA.I-Sce,Sco; da-GAL4/Otu1RNAi (21894) and DR-white.y+ (DR-white_2eu_1)/hsp70.HA.I-Sce,Sco; da-GAL4/UAS-RNAi Usp34 were heat-shocked at 38 °C for 1 h. For the mosaic red-eye phenotype quantification, we used heat-shock-induced yw; DR-white.y+ (DR-white_2eu_1)/hsp70.HA.I-Sce, Sco; da-GAL4/Otu1RNAi (21894) adult flies and we compared the number of red-eye flies to the total number of flies with the above-described genotype. In case of Usp34RNAi (27517), we used the same quantification method by measuring the red-eye mosaic phenotype. The heat-shock-induced yw; DR-white.y+ (DR-white_2eu_1)/hsp70.HA.I-Sce, Sco; da-GAL4/Usp34RNAi (27517) were counted and the mosaic red-eye adult flies were compared to the total number of the adult flies with the above-described genotype. For the control, we used yw; DR-white.y+ (DR-white_2eu_1)/hsp70.HA.I-Sce, Sco; da-GAL4/+ Drosophila lines with the same quantification method as was used in case of Usp34RNAi (27517) and Otu1RNAi (21894).
For the heterochromatic DR-white assay, we used yw; DR-white.y+ (DR-white_2het_2)/CyO; da-GAL4 females, which were crossed with transgenic yw; hsp70.HA.I-Sce, Sco/CyO; UAS-RNAi CG4603 or yw; hsp70.HA.I-Sce, Sco/CyO; UAS-RNAi Usp34 Drosophila males. The original Drosophila DR-white stocks (yw; DR-white.y+ (DR-white_2eu_2)/CyO and yw; DR-white.y+ (DR-white_2het_2)/CyO) and the yw; hsp70.HA.I-Sce,Sco/CyO Drosophila stock were kindly provided by Jeannine R. LaRocque and Anthony T. Do. More details about the transgenic DR-white stock are described in Janssen et al.38. The RNAi Drosophila stocks were kindly provided by Péter Deák. For the DR-white assay, nine independent experiments were performed.
References
Jackson, S. P. & Bartek, J. The DNA-damage response in human biology and disease. Nature 461, 1071–1078. https://doi.org/10.1038/nature08467 (2009).
Jackson, S. P. & Durocher, D. Regulation of DNA damage responses by ubiquitin and SUMO. Mol. Cell. 49, 795–807. https://doi.org/10.1016/j.molcel.2013.01.017 (2013).
van Attikum, H. & Gasser, S. M. Crosstalk between histone modifications during the DNA damage response. Trends Cell. Biol. 19, 207–217. https://doi.org/10.1016/j.tcb.2009.03.001 (2009).
Dantuma, N. P. & van Attikum, H. Spatiotemporal regulation of posttranslational modifications in the DNA damage response. EMBO J. 35, 6–23. https://doi.org/10.15252/embj.201592595 (2016).
McClurg, U. L. & Robson, C. N. Deubiquitinating enzymes as oncotargets. Oncotarget 6, 9657–9668. https://doi.org/10.18632/oncotarget.3922 (2015).
Jacq, X., Kemp, M., Martin, N. M. & Jackson, S. P. Deubiquitylating enzymes and DNA damage response pathways. Cell. Biochem. Biophys. 67, 25–43. https://doi.org/10.1007/s12013-013-9635-3 (2013).
Nijman, S. M. et al. The deubiquitinating enzyme USP1 regulates the Fanconi anemia pathway. Mol. Cell. 17, 331–339. https://doi.org/10.1016/j.molcel.2005.01.008 (2005).
Huang, T. T. et al. Regulation of monoubiquitinated PCNA by DUB autocleavage. Nat. Cell. Biol. 8, 339–347. https://doi.org/10.1038/ncb1378 (2006).
Nicassio, F. et al. Human USP3 is a chromatin modifier required for S phase progression and genome stability. Curr. Biol. 17, 1972–1977. https://doi.org/10.1016/j.cub.2007.10.034 (2007).
Li, M. et al. Deubiquitination of p53 by HAUSP is an important pathway for p53 stabilization. Nature 416, 648–653. https://doi.org/10.1038/nature737 (2002).
Dayal, S. et al. Suppression of the deubiquitinating enzyme USP5 causes the accumulation of unanchored polyubiquitin and the activation of p53. J. Biol. Chem. 284, 5030–5041. https://doi.org/10.1074/jbc.M805871200 (2009).
Zhang, X., Berger, F. G., Yang, J. & Lu, X. USP4 inhibits p53 through deubiquitinating and stabilizing ARF-BP1. EMBO J. 30, 2177–2189. https://doi.org/10.1038/emboj.2011.125 (2011).
Khoronenkova, S. V. et al. ATM-dependent downregulation of USP7/HAUSP by PPM1G activates p53 response to DNA damage. Mol. Cell. 45, 801–813. https://doi.org/10.1016/j.molcel.2012.01.021 (2012).
Wang, C. et al. Deubiquitinating enzyme USP20 is a positive regulator of Claspin and suppresses the malignant characteristics of gastric cancer cells. Int. J. Oncol. 50, 1136–1146. https://doi.org/10.3892/ijo.2017.3904 (2017).
Georges, A., Marcon, E., Greenblatt, J. & Frappier, L. Identification and characterization of USP7 targets in cancer cells. Sci. Rep. 8, 15833. https://doi.org/10.1038/s41598-018-34197-x (2018).
Zhu, Q., Sharma, N., He, J., Wani, G. & Wani, A. A. USP7 deubiquitinase promotes ubiquitin-dependent DNA damage signaling by stabilizing RNF168. Cell Cycle 14, 1413–1425. https://doi.org/10.1080/15384101.2015.1007785 (2015).
Sy, S. M., Jiang, J. O. W. S., Deng, Y. & Huen, M. S. The ubiquitin specific protease USP34 promotes ubiquitin signaling at DNA double-strand breaks. Nucleic Acids Res. 41, 8572–8580. https://doi.org/10.1093/nar/gkt622 (2013).
Wu, Z. et al. OTU deubiquitinase 4 is silenced and radiosensitizes non-small cell lung cancer cells via inhibiting DNA repair. Cancer Cell. Int. 19, 99. https://doi.org/10.1186/s12935-019-0816-z (2019).
Altun, M. et al. The human otubain2-ubiquitin structure provides insights into the cleavage specificity of poly-ubiquitin-linkages. PLoS ONE 10, e0115344. https://doi.org/10.1371/journal.pone.0115344 (2015).
Wiener, R., Zhang, X., Wang, T. & Wolberger, C. The mechanism of OTUB1-mediated inhibition of ubiquitination. Nature 483, 618–622. https://doi.org/10.1038/nature10911 (2012).
Nakada, S. et al. Non-canonical inhibition of DNA damage-dependent ubiquitination by OTUB1. Nature 466, 941–946. https://doi.org/10.1038/nature09297 (2010).
Kee, Y. & Huang, T. T. Role of deubiquitinating enzymes in DNA repair. Mol. Cell. Biol. 36, 524–544. https://doi.org/10.1128/MCB.00847-15 (2016).
Pahi, Z. G., Borsos, B. N., Pantazi, V., Ujfaludi, Z. & Pankotai, T. PARylation during transcription: Insights into the fine-tuning mechanism and regulation. Cancers https://doi.org/10.3390/cancers12010183 (2020).
Amerik, A. Y., Li, S. J. & Hochstrasser, M. Analysis of the deubiquitinating enzymes of the yeast Saccharomyces cerevisiae. Biol. Chem. 381, 981–992. https://doi.org/10.1515/BC.2000.121 (2000).
Nijman, S. M. et al. A genomic and functional inventory of deubiquitinating enzymes. Cell 123, 773–786. https://doi.org/10.1016/j.cell.2005.11.007 (2005).
Ristic, G. et al. USP5 is dispensable for monoubiquitin maintenance in drosophila. J. Biol. Chem. 291, 9161–9172. https://doi.org/10.1074/jbc.M115.703504 (2016).
Kovacs, L. et al. Role of the deubiquitylating enzyme DmUsp5 in coupling ubiquitin equilibrium to development and apoptosis in Drosophila melanogaster. PLoS ONE 10, e0120875. https://doi.org/10.1371/journal.pone.0120875 (2015).
Perez-Oliva, A. B. et al. USP45 deubiquitylase controls ERCC1-XPF endonuclease-mediated DNA damage responses. EMBO J. 34, 326–343. https://doi.org/10.15252/embj.201489184 (2015).
Parsons, J. L. et al. USP47 is a deubiquitylating enzyme that regulates base excision repair by controlling steady-state levels of DNA polymerase beta. Mol. Cell. 41, 609–615. https://doi.org/10.1016/j.molcel.2011.02.016 (2011).
Sharma, A. et al. USP14 regulates DNA damage repair by targeting RNF168-dependent ubiquitination. Autophagy 14, 1976–1990. https://doi.org/10.1080/15548627.2018.1496877 (2018).
Do, A. T., Brooks, J. T., Le Neveu, M. K. & LaRocque, J. R. Double-strand break repair assays determine pathway choice and structure of gene conversion events in Drosophila melanogaster. G3 4, 425–432. https://doi.org/10.1534/g3.113.010074 (2014).
Borsos, B. N., Majoros, H. & Pankotai, T. Ubiquitylation-mediated fine-tuning of DNA double-strand break repair. Cancers https://doi.org/10.3390/cancers12061617 (2020).
Borsos, B. N., Majoros, H. & Pankotai, T. Emerging roles of post-translational modifications in nucleotide excision repair. Cells https://doi.org/10.3390/cells9061466 (2020).
Sato, Y. et al. Molecular basis of Lys-63-linked polyubiquitination inhibition by the interaction between human deubiquitinating enzyme OTUB1 and ubiquitin-conjugating enzyme UBC13. J. Biol. Chem. 287, 25860–25868. https://doi.org/10.1074/jbc.M112.364752 (2012).
Nakajima, S. et al. Ubiquitin-specific protease 5 is required for the efficient repair of DNA double-strand breaks. PLoS ONE 9, e84899. https://doi.org/10.1371/journal.pone.0084899 (2014).
Reyes-Turcu, F. E. et al. The ubiquitin binding domain ZnF UBP recognizes the C-terminal diglycine motif of unanchored ubiquitin. Cell 124, 1197–1208. https://doi.org/10.1016/j.cell.2006.02.038 (2006).
Nakada, S. Opposing roles of RNF8/RNF168 and deubiquitinating enzymes in ubiquitination-dependent DNA double-strand break response signaling and DNA-repair pathway choice. J. Radiat. Res. 57(Suppl 1), i33–i40. https://doi.org/10.1093/jrr/rrw027 (2016).
Janssen, A. et al. A single double-strand break system reveals repair dynamics and mechanisms in heterochromatin and euchromatin. Genes Dev. 30, 1645–1657. https://doi.org/10.1101/gad.283028.116 (2016).
Acknowledgements
We thank Hajnalka Majoros and Vasiliki Pantazi for their expert help in commenting and correcting the manuscript.
Funding
Open access funding provided by University of Szeged. This research was funded by National Research, Development and Innovation Office Grant GINOP-2.2.1-15-2017-00052, GINOP-2.3.2-15-2016-00032 and NKFI-FK 132080, the János Bolyai Research Scholarship of the Hungarian Academy of Sciences BO/27/20, ÚNKP-20-5-SZTE-265 and ÚNKP-21-5-SZTE-563.
Author information
Authors and Affiliations
Contributions
G.Z.P. and T.P. conceived the idea designed and supervised the experiments. G.Z.P., L.K., D.S. conducted all laboratory work. G.Z.P., B.N.B., P.D. and T.P. facilitated the laboratory analysis. G.Z.P., B.N.B., L.K., P.D. and T.P. prepared initial draft of the manuscript. G.Z.P., B.N.B., L.K., P.D. and T.P. were responsible for the final paper following contributions from all authors.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Páhi, Z.G., Kovács, L., Szűcs, D. et al. Usp5, Usp34, and Otu1 deubiquitylases mediate DNA repair in Drosophila melanogaster. Sci Rep 12, 5870 (2022). https://doi.org/10.1038/s41598-022-09703-x
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-022-09703-x
- Springer Nature Limited