Abstract
Long non-coding RNAs (LncRNAs) play vital roles in the tumorigenesis of many cancers. Single nucleotide polymorphisms (SNPs) of the lncRNA also play vital roles in tumorigenesis. We explored lncRNA rs944289 and rs7990916 polymorphisms and analyzed the relationship between these lncRNA polymorphisms with the colorectal cancer (CRC) risk in a Chinese population. We recruited 1003 CRC patients from the Affiliated People’s Hospital of Jiangsu University and the Fujian Medical University Union Hospital from October 2014 to August 2017. Genomic DNA was extracted using a DNA Kit from lymphocytes of peripheral blood and the genotyping was performed with a SNPscan method. We found that the rs944289 TT homozygote was associated with the decreased CRC risk in the overall population. LncRNA rs944289 TT decreased the CRC risk in the subgroup of female, male, age ≥ 61, without alcohol intake, smoking and BMI ≥ 24 by logistic regression. The subgroup analysis revealed that lncRNA rs7990916 was not associated with CRC risk except for age < 61. Logistic regression analysis revealed that lncRNA rs944289 TT homozygote was associated with the increased risk of rectum cancer (TT vs. CC + CT: adjusted OR = 1.29, 95% CI 1.10–1.66, P = 0.041) or colon cancer. In summary, we proved that lncRNA rs944289 might be significantly related to the decreased CRC risk in the Chinese Han populations and lncRNA rs7990916 was not associated with the CRC risk except for patients of age < 61. In the future, studies with larger samples should be conducted to validate our results.
Similar content being viewed by others
Introduction
Colorectal cancer (CRC) has ranked third in terms of incidence but second in terms of mortality in the world1. In China, CRC has ranked both fifth in terms of incidence and mortality2. The incidence of CRC has been increasing, mostly due to unhealthy lifestyle, aging and environmental factors3, but accumulating evidences have shown that an individual’s inherited factors also contribute to the development of CRC.
Long noncoding RNAs (lncRNAs) are a sort of noncoding RNA molecules with more than 200 nucleotides in length, and are disease- or tissue-specific expression patterns4,5. LncRNAs play crucial roles in chromatin dynamics, genome packaging, gene regulation, cellular pathways and biological processes6. LncRNAs also play viotal roles in the oncogenesis and metastasis of CRC7,8. Yu et al. found that linc-UFC1 participated in the progression of CRC, and overexpression of linc-UFC1 in CRC patients was positively associated with tumor grade and stage7. Zhao et al. also reported overexpression of Linc-A was correlated with poor survival in CRC patients8.
Single nucleotide polymorphisms (SNPs) occurring in the functional region of lncRNAs can influence disease risk and can also promote cancer development9. LncRNA rs944289 and rs7990916 polymorphisms increased the risk of different diseases. For example, lncRNAs rs7990916 TT genotype was found to increase Alzheimer’s disease (AD) risk10. Papillary thyroid carcinoma susceptibility candidate 3 (PTCSC3) rs944289 polymorphism increased the risk of papillary thyroid cancer (PTC)11,12. Cao et al. also reported that lncRNA rs944289 may increase the risk of esophagogastric junction adenocarcinoma (EGJA)13. However, Xu et al. found that lncRNA rs944289 have no effect on the risk of breast cancer (BC)14. These data indicate SNPs in lncRNAs play important roles in tumorigenesis. Although lncRNA rs944289 and rs7990916 polymorphisms have a risk effect on some diseases, there are no studies concerning the relationship of these two variants with CRC risk. Based on previous studies, we explored lncRNA rs944289 and rs7990916 polymorphisms in CRC patients and analyzed the relationship between these lncRNA polymorphisms and CRC risk.
Materials and methods
Patients and samples
We recruited 1003 CRC patients from the Affiliated People’s Hospital of Jiangsu University and the Fujian Medical University Union Hospital from October 2014 to August 2017. Patients with histologically confirmed CRC were enrolled and the major exclusion criteria were as follows: (1) presence of immunological diseases, (2) with other primary cancers, and (3) with other colonic diseases (e.g. Crohn’s disease), (4) exposure to antitumor treatments before surgery. One thousand three hundred and three healthy subjects were selected as healthy controls from the departments of physical examination in these two hospitals. The healthy controls were Chinese Han populations with the following exclusion criteria: (1) with history of any cancer, (2) with metabolic or autoimmune diseases, and (3) with liver/kidney dysfunction, (4) with any other systemic diseases. The demographic characteristics were recorded, including body mass index (BMI), age, gender, alcohol intake and smoking.
The study was approved by the Ethics Committee of the Affiliated People's Hospital of Jiangsu University and conducted according to the Declaration of Helsinki. All participants signed an informed consent before enrolled in this research and all methods were performed in accordance with the relevant guidelines and regulations.
SNP genotyping
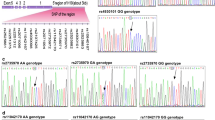
We collected blood samples from each participant after their admission to the hospital. The blood samples collected in a test tube containing EDTA were used for genotyping assay. Genomic DNA was extracted using a DNA Kit (Promega, Madison, USA) from lymphocytes of peripheral blood. The DNA was quantified by a measurement of OD260 and then was stored at -80℃. SNPs of lncRNA rs944289 and rs7990916 were analyzed by a SNPscan Kit (Genesky Biotechnologies Inc., Shanghai, China)15. 4% DNA samples were selected randomly and analyzed by PCR/Sanger sequencing. The rs944289 locus primers were: 5-TGGTTGAAAGATAGTCATTG-3 (forward) and 5-AGATTTGTAATAGCTGGGAA-3(reverse). The rs7990916 locus primers were: 5-CTTTGTATCCTTCATTCTTA-3 (forward) and 5-CAAGTTGACACTCAGAATTA-3(reverse). For quality control, replicate blinded samples were included to check for the reproducibility of the results by another technician and the results were unchanged.
Statistical analysis
All data were analyzed by using SPSS 23.0 (IBM Corporation, Armonk, NY, USA). The chi-square or Fisher’s exact test was applied to compare the categorical variables, whereas the continuous variables were evaluated by the student’s t-test or Wilcoxon signed-rank test. The relationship between the lncRNA rs944289/7990916 polymorphism and CRC risk was assessed by odds ratios (OR) and 95% confidence intervals (CIs). Logistic regression model was conducted to analyze the associations among lncRNA rs944289 and rs7990916 polymorphisms, the clinical characteristics and CRC risk. Hardy–Weinberg equilibrium (HWE) was used to analyze the genotype distributions of lncRNA rs944289/7990916. A P < 0.05 (two-tailed) was adopted as the statistically significant level.
Ethics approval and consent to participate
The Ethics Committee of the Affiliated People’s Hospital of Jiangsu University approved the protocol of the study (K-20210105-W), and all participants signed an informed consent before enrolled in this research.
Consent for publication
The authors appreciate all the patients in this work for their cooperation and permission for the publication of the article.
Results
Characteristics of CRC patients
In our study, 1003 CRC cases (431 colon cancer and 572 rectum cancer patients) and 1303 healthy controls were enrolled to investigate the correlation of the two SNPs (rs944289 and rs7990916) with CRC risk. Table 1 listed detailed demographics data. The mean age of CRC patients and controls were 61.10 ± 12.17 years and 61.40 ± 9.61 years, respectively. There were no statistically significant differences in age and gender between CRC patients and controls (both P > 0.05). However, smoking, alcohol intake and BMI increased the risk of CRC (both P < 0.05), so we adjusted these factors by multiple logistic regression analyses.
Primary information for lncRNA rs944289 and rs7990916
The genotypic frequencies of lncRNA rs944289 and rs7990916 met the HWE (P = 0.105 and P = 0.359, respectively). Minor allele frequency (MAF) of lncRNA rs944289 polymorphism was 0.28, which was similar to SNP database for Chinese populations (MAF = 0.24). MAF of lncRNA rs7990916 polymorphism was 0.23, which was similar to SNP database for Chinese populations. Table 2 summarized the corresponding information of lncRNA rs944289 and rs7990916.
Association of lncRNA polymorphisms with CRC risk in the overall population
The genotypes and allele distributions of lncRNA rs944289 and rs7990916 in CRC patients and controls were presented in Table 3. LncRNA rs944289 frequencies were 29.55% (CC), 46.73% (CT) and 23.72% (TT) in CRC patients, whereas in controls, the distributions of those genotypes were 29.69%, 51.62% and 18.69%, respectively. We found that rs944289 TT homozygote was associated with decreased CRC risk when compared with the CC or CC + CT (TT vs. CC: adjusted OR = 0.88, 95% CI 0.78–0.99, P = 0.037; TT vs. CC + CT: adjusted OR = 0.86, 95% CI 0.78–0.95, P = 0.004). LncRNA rs7990916 frequencies were 79.69% (CC), 19.49% (CT) and 0.82% (TT) in CRC patients, and 77.31% (CC), 20.92% (CT) and 1.77% (TT) in controls. There was no significant difference between lncRNA rs7990916 and the risk of CRC (all P > 0.05) in the overall population.
Stratified analyses between lncRNA rs944289 polymorphism and CRC risk
Stratified analysis was performed and revealed that rs944289 TT genotype was associated with the decreased CRC risk in the subgroup of male (adjusted OR = 0.88, 95% CI 0.77–0.99, P = 0.045), female (adjusted OR = 0.81, 95% CI 0.68–0.99, P = 0.097), age ≥ 61 (adjusted OR = 0.86, 95% CI 0.74–0.99, P = 0.035), smoking (adjusted OR = 0.66, 95% CI 0.53–0.81, P < 0.001), never alcohol intake (adjusted OR = 0.88, 95% CI 0.78–0.98, P = 0.021) and BMI ≥ 24 (adjusted OR = 0.82, 95% CI 0.696–0.94, P = 0.0016) (Table 4).
Stratified analyses between lncRNA rs7990916 polymorphism and CRC risk
We further analyzed stratified effects of lncRNA rs7990916 on CRC risk by logistic regression model. Genotype distributions of rs7990916 were evaluated with age, sex, BMI, drinking and smoking. Results show that there was no significant association between CRC patients and controls except for age < 61 (CT vs. CC: adjusted OR = 0.67, 95% CI 0.48–0.93, P = 0.018) (Table 5).
LncRNA rs944289/rs7990916 polymorphism and CRC risk by site of tumor
Logistic regression analysis revealed that lncRNA rs944289 was associated with increased risk of colon cancer (TT vs. CC + CT: adjusted OR = 1.44, 95% CI 1.11–1.88, P = 0.007) or rectum cancer (TT vs. CC + CT: adjusted OR = 1.29, 95% CI 1.10–1.66, P = 0.041) when compared with CC + CT (Table 6). We also found lncRNA rs7990916 did not alter the risk of colon cancer (CT vs. CC: adjusted P = 0.18, TT vs. CC: adjusted P = 0.21, CT + TT vs.CC: adjusted P = 0.11, and TT vs. CC + CT: adjusted P = 0.24) or rectal cancer (CT vs. CC: adjusted P = 0.58, TT vs. CC: adjusted P = 0.11, CT + TT vs. CC: adjusted P = 0.37, and TT vs. CC + CT: adjusted P = 0.12) (Table 6).
Discussion
LncRNAs play an essential role in various biological processes, including cell proliferation, apoptosis, genomic imprinting, transcriptional interference and other critical processes16,17. Furthermore, lncRNAs have been reported to participate in the process of tumorigenesis in CRC. For instance, lncRNA CCAL could activate Wnt/β-catenin signaling pathway and induce multidrug resistance in CRC18. LncRNA SPRY4-IT1 could promote invasion and proliferation as a ceRNA of miRNA-101-3p in CRC19. Recently, numerous investigations have suggested lncRNA polymorphisms as one of the contributors to CRC risk. For example, Yang et al. reported lncRNA PCAT1 rs2632159 polymorphism increased CRC risk in a Chinese population20. Wang et al. also reported H19 rs2839698 polymorphism was associated with increased CRC risk21. On the other hand, lncRNA PRNCR1 rs13252298/rs1456315 and MALAT1 rs1194338 polymorphisms decreased the CRC risk22,23. Considering this background, we performed a case–control study to determine the relationship between lncRNA rs944289/rs7990916 polymorphism and CRC risk.
In this study, we found that lncRNA rs944289 TT homozygote could decrease CRC risk. After adjustment by multiple logistic regression, the TT genotype of lncRNA rs944289 was associated with the decreased CRC risk in the subgroups of female, male, age ≥ 61, BMI ≥ 24, smoking and never alcohol intake populations. LncRNA rs944289 have been confirmed to increase the risk of PTC11,24, and rs944289 predisposes to PTC by inhibiting the expression of PTCSC311,12. PTCSC3 is a large intergenic noncoding RNA gene that is involved in the regulation of tumorigenesis. The SNP rs944289 is located 3.2 kb upstream of PTCSC3 and suppresses PTCSC3 by destroying a transcription factor-binding site in the promoter of PTCSC325,26. Cao et al. also found that lncRNA rs944289 may increase the risk of EGJA in the smoking and age < 60 years populations13. However, in a Chinese population, Xu et al. discovered that lncRNA rs944289 have no significant effect on the risk of BC, which might be due to the specific tumorigenesis of BC14. However, up to now, no study has reported the association between lncRNA rs944289 polymorphism and CRC risk, and in this study, we observed significant relationships between the lncRNA rs944289 polymorphism and CRC risk.
Our results also revealed that there was no significant difference between lncRNA rs7990916 polymorphism and the CRC risk in an overall comparison. But when stratified by age < 61 years, lncRNA rs7990916 polymorphism decreased the risk of CRC. Similar to our results, Cao et al. observed no close relationship between lncRNA rs7990916 polymorphism and EGJA patients. In contrast, Jendrzejewski et al. found significant relationship between lncRNA rs7990916 and the risk of AD in the Europe populations. But the etiology of CRC is primarily different from that of AD21. Thus, these distinct conclusions on the lncRNA rs7990916 polymorphism may be mainly attributed to the heterogeneity of ethnics and disease types.
Nevertheless, the mechanism of lncRNA polymorphisms on the tumorigenesis of CRC is unclear. Previous studies reported that lncRNA GAS5 rs55829688 polymorphism increased CRC risk by modulating the binding affinity of the transcription factors YY1 to GAS5 promotor27. Moreover, MALAT1 rs664589 polymorphism increased CRC risk by binding to miRNA-194-5p, which resulted in an overexpression of MALAT128. Taken together, lncRNA polymorphisms could induce the occurrence of tumor by binding to transcription factors, binding to miRNA and so on.
To the best of our knowledge, our research focused on the possible association of lncRNA rs944289 and rs7990916 polymorphisms with CRC risk for the first time. There still existed several limitations in the study. Firstly, this case–control study only focused on Chinese Han populations. Secondly, the two lncRNAs selected in our study may not be comprehensive because there may be other genetic polymorphisms that affected the risk of CRC. Thirdly, we did not perform the association of lncRNA rs944289 and rs7990916 polymorphisms with tumor stages, patients’ prognosis, or other associated risk factors, which might affect the precision of estimating the effect of these two polymorphisms on the CRC risk. Finally, we did not investigate the mechanism of the two polymorphisms during the tumorigenesis of CRC. Nevertheless, our study provides clews for a further mechanism study and provides possible biomarkers for CRC diagnosis.
Conclusions
In summary, we proved that the lncRNA rs944289 might be significantly related to the decreased risk of CRC in the Chinese Han populations. However, there was no significant difference between lncRNA rs7990916 polymorphism and CRC risk except for the subgroup of age < 61. In the future studies, larger samples including detailed clinical information and different ethnic populations should be conducted to validate our findings.
Data availability
All data analyzed or generated during this study are included in the article.
References
Bray, F. et al. Cancer statistics 2018: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J. Clin. 68(6), 394–424 (2018).
Chen, W. et al. Cancer statistics in China, 2015. CA Cancer J. Clin. 66(2), 115–132 (2016).
Mathers, J. C. Obesity and bowel cancer: From molecular mechanisms to interventions. Nutr. Res. 70, 26–31 (2019).
Beckedorff, F. C. et al. Long non-coding RNAs and their implications in cancer epigenetics. Biosci. Rep. 33(4), e00061 (2013).
Enfield, K. S. et al. Mechanistic roles of noncoding RNAs in lung cancer biology and their clinical implications. Genet. Res. Int. 2012, 737416 (2012).
Bhan, A., Soleimani, M. & Mandal, S. S. Long noncoding RNA and cancer: A new paradigm. Cancer Res. 77(15), 3965–3981 (2017).
Yu, T. et al. Knockdown of linc-UFC1 suppresses proliferation and induces apoptosis of colorectal cancer. Cell Death Dis. 7(5), e2228 (2016).
Zhao, C. et al. High expression of long non-coding RNA Linc-A associates with poor survival in patients with colorectal cancer. Mol. Biol. Rep. 47(10), 7497–7504 (2020).
Benjelloun, B. et al. An evaluation of sequencing coverage and genotyping strategies to assess neutral and adaptive diversity. Mol. Ecol. Resour. 19(6), 1497–1515 (2019).
Chen, G. et al. Genome-wide analysis of human SNPs at long intergenic noncoding RNAs. Hum. Mutat. 34(2), 338–344 (2013).
Rogounovitch, T. I. et al. The common genetic variant rs944289 on chromosome 14q13.3 associates with risk of both malignant and benign thyroid tumors in the Japanese population. Thyroid 25(3), 333–340 (2015).
Ai, L. et al. Associations between rs965513/rs944289 and papillary thyroid carcinoma risk: A meta-analysis. Endocrine 47(2), 428–434 (2014).
Cao, R. et al. Association of long noncoding RNAs polymorphisms with the risk of esophagogastric junction adenocarcinoma: A three-center study of 1063 cases and 1677 controls. DNA Cell Biol. 39(5), 828–835 (2020).
Xu, T. et al. Association between SNPs in long non-coding RNAs and the risk of female breast cancer in a Chinese population. J. Cancer 8(7), 1162–1169 (2017).
Tang, W. et al. Association of miRNA-499 rs3746444 A>G variants with adenocarcinoma of esophagogastric junction (AEG) risk and lymph node status. Onco Targets Ther. 12, 6245–6252 (2019).
Hu, X. et al. The role of long noncoding RNAs in cancer: The dark matter matters. Curr. Opin. Genet. Dev. 48, 8–15 (2018).
Schmitt, A. M. & Chang, H. Y. Long noncoding RNAs in cancer pathways. Cancer Cell 29(4), 452–463 (2016).
Ma, Y. et al. Long non-coding RNA CCAL regulates colorectal cancer progression by activating Wnt/β-catenin signalling pathway via suppression of activator protein 2α. Gut 65(9), 1494–1504 (2016).
Jin, J. et al. Long non-coding RNA SPRY4-IT1 promotes proliferation and invasion by acting as a ceRNA of miR-101-3p in colorectal cancer cells. Tumour Biol. 39(7), 1010428317716250 (2017).
Yang, M. et al. LncRNA-PCAT1 rs2632159 polymorphismcould be a biomarker for colorectal cancer susceptibility. Biosci. Rep. 39(7), BSR20190708 (2019).
Wang, R. et al. Identification of long noncoding RNAs as potential novel diagnosis and prognosis biomarkers in colorectal cancer. J. Cancer Res. Clin. Oncol. 142(11), 2291–2301 (2016).
Fardet, A. et al. Do alcoholic beverages, obesity and other nutritional factorsmodify the risk of familial colorectal cancer? A systematic review. Crit. Rev. Oncol. Hematol. 119, 94–112 (2017).
Li, L. et al. Association between polymorphisms in long non-coding RNA PRNCR1 in 8q24 and risk of colorectal cancer. J. Exp. Clin. Cancer Res. 32(1), 104 (2013).
Wang, Y. L. et al. Confirmation of papillary thyroid cancer susceptibility loci identified by genome-wide association studies of chromosomes 14q13, 9q22, 2q35 and 8p12 in a Chinese population. J. Med. Genet. 50(10), 689–695 (2013).
Jendrzejewski, J. et al. The polymorphism rs944289 predisposes to papillary thyroid carcinoma through a large intergenic noncoding RNA gene of tumor suppressor type. Proc. Natl. Acad. Sci. USA 109(22), 8646–8651 (2012).
Jendrzejewski, J. et al. PTCSC3 is involved in papillary thyroid carcinoma development by modulating S100A4 gene expression. J. Clin. Endocrinol. Metab. 100(10), E1370–E1377 (2015).
Wang, Y., Wu, S., Yang, X., Li, X. & Chen, R. Association between polymorphismin the promoter region of lncRNA GAS5 and the risk of colorectal cancer. Biosci. Rep. 39(4), BSR20190091 (2019).
Wu, S. et al. MALAT1 rs664589 polymorphism inhibits binding to miR-194-5p, contributing to colorectal cancer risk, growth, and metastasis. Cancer Res. 79(20), 5432–5441 (2019).
Funding
This research was funded by “Liu Ge Yi Gong Cheng”of Jiangsu Province (Grant No. LGY2017022), Youth Foundation of Jiangsu Province (Grant No. QNRC2016448), Social Development Foundation of Zhenjiang (SH2019069) and Science Foundation of the Affiliated People's Hospital of Jiangsu University (Y2019021-S).
Author information
Authors and Affiliations
Contributions
All authors contributed to the study conception and design. Clinical data collection, genetic counseling and follow-up were performed by Y.W., Z.Q. and G.T. The experiment was performed by R.Q. and Y.P. SNP analysis was performed by Q.Z. and Z.Z. Formal analysis was performed by W.T. The manuscript was written by S.Z. and Y.X., and all authors read and approved the final manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Wang, Y., Qiu, Z., Tian, G. et al. Association between long noncoding RNA rs944289 and rs7990916 polymorphisms and the risk of colorectal cancer in a Chinese population. Sci Rep 12, 2495 (2022). https://doi.org/10.1038/s41598-022-06474-3
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-022-06474-3
- Springer Nature Limited
This article is cited by
-
A review on the biological roles of LncRNA PTCSC3 in cancerous and non-cancerous disorders
Cancer Cell International (2023)