Abstract
Due to the recurring loss of antimalarial drugs to resistance, there is a need for novel targets, drugs, and combination therapies to ensure the availability of current and future countermeasures. Pyrazoleamides belong to a novel class of antimalarial drugs that disrupt sodium ion homeostasis, although the exact consequences of this disruption in Plasmodium falciparum remain under investigation. In vitro experiments demonstrated that parasites carrying mutations in the metabolic enzyme PfATP4 develop resistance to pyrazoleamide compounds. However, the underlying mechanisms that allow mutant parasites to evade pyrazoleamide treatment are unclear. Here, we first performed experiments to identify the sublethal dose of a pyrazoleamide compound (PA21A092) that caused a significant reduction in growth over one intraerythrocytic developmental cycle (IDC). At this drug concentration, we collected transcriptomic and metabolomic data at multiple time points during the IDC to quantify gene- and metabolite-level alterations in the treated parasites. To probe the effects of pyrazoleamide treatment on parasite metabolism, we coupled the time-resolved omics data with a metabolic network model of P. falciparum. We found that the drug-treated parasites adjusted carbohydrate metabolism to enhance synthesis of myoinositol—a precursor for phosphatidylinositol biosynthesis. This metabolic adaptation caused a decrease in metabolite flux through the pentose phosphate pathway, causing a decreased rate of RNA synthesis and an increase in oxidative stress. Our model analyses suggest that downstream consequences of enhanced myoinositol synthesis may underlie adjustments that could lead to resistance emergence in P. falciparum exposed to a sublethal dose of a pyrazoleamide drug.
Similar content being viewed by others
Introduction
The malaria parasite Plasmodium falciparum is the causative agent of the most severe form of malaria and is responsible for 97% of malaria-related deaths globally1. Due to worldwide efforts, malaria-related deaths have decreased from 736,000 in 2000 to 409,000 in 20192, however, this decrease in malaria mortality rate has plateaued in the last five years2. Treatment to decrease malaria-related deaths depends on the use of drugs that have been losing efficacy due to drug resistance3. In fact, P. falciparum has developed some degree of resistance to all approved antimalarial drugs to date3, forcing development of drug-combination therapies involving two (or more) drugs to treat malaria and, hopefully, also suppress drug-resistance development. As emergence of resistance to both single and combination therapies continues4, there is an unrelenting need to identify novel antimalarial drug targets.
One such global effort led to the identification of a P-type ATPase, PfATP4, which appears to be the target of multiple drugs of different chemical classes. The PfATP4-associated compounds, such as pyrazoleamides and spiroindolones, disrupt sodium ion homeostasis and cause multiple morphological and physiological changes5,6. P. falciparum carrying point mutations in PfATP4 have differential sensitivity to spiroindolone and pyrazoleamide compounds, e.g., the A211T mutation in PfATP4 increases the 50% inhibitory concentration (IC50) to GNF-Pf4492, a pyrazoleamide compound, but decreases the IC50 to KAF246, a spiroindolone compound7. How PfATP4 mutations are associated with differential sensitivity of drugs against P. falciparum is unclear. Prolonged exposure to sublethal concentrations of PfATP4-targeting drugs induce drug-resistant mutations in the gene and loss of efficacy against P. falciparum8,9,10. Similarly, pyrazoleamides induce drug-resistant mutations in the PfATP4 gene7,10. Here, we aim to understand the immediate metabolic adjustments of previously unexposed P. falciparum in response to a 48-h sublethal exposure to PA21A092. This adjustment constitutes the immediate survival phenotype and highlights the ability of the parasite to survive stress induced by the disruption of sodium ion homeostasis.
Towards this end, we first identified a dose of PA21A092 that caused a significant reduction in growth of P. falciparum Dd2 strain over a single intraerythrocytic developmental cycle (IDC). At this drug concentration, we did not observe any changes in morphologies of the treated parasites, suggesting that the identified sublethal dose did not cause major physiological alterations in P. falciparum. To characterize pyrazoleamide-induced perturbations in P. falciparum metabolism, we collected transcriptomic and metabolomic data at multiple time points during the IDC and coupled these data with a genome-scale metabolic model of P. falciparum11,12. Our model-based analyses revealed metabolic adaptations in P. falciparum that can counter the effects of sodium ion homeostasis disruptors, such as PA21A092. The identification of Plasmodium enzymes and pathways associated with drug-resistance development would allow development of rationally motivated drug-combination therapies that can mitigate evolution of drug-resistant progenies of P. falciparum.
Results
Pyrazoleamide-treated P. falciparum cultures
Prolonged culturing of P. falciparum in the presence of a sublethal pyrazoleamide dose selects for resistance-conferring mutations in its metabolic enzyme PfATP47,10. To understand the initial metabolic responses of P. falciparum in the presence of a pyrazoleamide, we first performed independent experiments in quadruplicate using various doses of PA21A092 to determine the optimal concentration of the drug that will affect parasite growth without killing the parasites prematurely (between 30 and 70% inhibition would be optimal). We treated highly synchronized cultures of ring-stage parasites with different concentrations of PA21A092 for 40 h. We then assessed the ability of these parasites to replicate by cutting the cultures by a factor of 10 with fresh erythrocytes and drug-free medium. Figure 1A shows that 12 nM PA21A092 achieves the desired level of inhibition, therefore, we used this concentration for data-collection experiments.
Experiments to determine a sublethal dose of PA21A092, a pyrazoleamide drug. (A) Parasitemia at ~ 70 h of experiment after treatment with the notated doses of PA21A092. We treated highly synchronized parasite cultures with 0, 8, 12, 16, or 24 nM PA21A092 in quadruplicate-independent experiments for 40 h. We then assessed the ability of these parasites to replicate by cutting the cultures by a factor of 10 with fresh erythrocytes and drug-free medium. The error bars denote one standard deviation from mean parasitemia. (B) Inhibition observed in the four pyrazoleamide-treated cultures used for transcriptomic and metabolomic sample collection. The ordinate shows the percentage of inhibition observed in each replicate after treatment with 12 nM PA21A092. For each replicate, the error bars denote variability in triplicate flow-cytometry measurements. The horizontal dotted line denotes the average of percentage inhibition observed across all measurements. (C) Representative Giemsa-stained images of untreated (CTL) and drug-treated (PYZ) parasites at the 0-, 8-, 16-, 24-, 32-, and 40-h time points of the experiment. Note that the 0-h time point represents the appearance of > 95% rings in the parasite culture and corresponds to ~ 0–2 h of the experiment. At this time (0 h), we added fresh medium containing the drug (PYZ) or no drug (CTL). (D) Number of merozoites per schizont in the untreated and drug-treated parasite cultures at the 40-h time point of the experiment. We found that a 40-h treatment by 12 nM PA21A092 (PYZ) resulted in a significant decrease in daughter merozoites per schizont. Magenta dotted lines represent the median of 50 observations under untreated (CTL) and drug-treated conditions. The black dotted lines denote the 25% and 75% quartiles. The asterisks denote that the difference between median number of merozoites/schizont under the two conditions is statistically significant (p < 0.0001; Wilcoxon rank-sum test).
Prior to sample collection for transcriptomic and metabolomic data profiling, we again quantified the amount of parasite growth inhibition and also monitored the morphologies of untreated and drug-treated parasites during the IDC. Figure 1B quantifies the inhibition of parasite proliferation, as a percentage of parasitemia observed in untreated parasite cultures, confirming that the identified PA21A092 dosage resulted in an inhibition within the desired range (30–70%).
To monitor morphological changes in parasites due to the drug treatment, we performed Giemsa staining every 8 h during the IDC. Figure 1C shows representative images of parasite-infected erythrocytes in the absence and presence of 12 nM PA21A092 at 0, 8, 16, 24, 32, and 40 h of the experiment. These images suggest that the identified dose of PA21A092 did not perturb parasite morphologies substantially until the 40-h time point of the experiment, however, it still caused ~ 35% of the drug-treated parasites to fail to proliferate in the drug-free medium after 40 h (Fig. 1B). We also quantified the number of merozoites per schizont at the 40-h time point under untreated (CTL) and drug-treated (PYZ) conditions. Figure 1D shows that 12 nM PA21A092 caused a significant decrease in the number of daughter merozoites per schizont (CTL: 12; PYZ: 10), suggesting that the drug negatively affected parasite development.
Transcriptomic changes due to a sublethal dose of a pyrazoleamide
To identify major coherent alterations in gene-expression data from the treated parasites, we performed hierarchical clustering analysis (HCA). Figure 2A illustrates the average expression of 5,851 parasite genes during the IDC in the absence and presence of PA21A092. We did not find any major alteration in the transcription program of the drug-treated parasites. To quantify the effect of PA21A092 treatment on P. falciparum IDC progression, we compared gene-expression profiles from untreated and drug-treated parasites with those from hourly sampled gene-expression data spanning the entire IDC of P. falciparum Dd2 strain13.
Global analysis of transcriptomic data from untreated and pyrazoleamide-treated P. falciparum. (A) Hierarchical clustering analysis of transcriptomic data from the untreated and treated parasites. Each row corresponds to a parasite gene, and each column represents a sampled time point of the intraerythrocytic development cycle (IDC). The gene-expression level, under a given perturbation (untreated or drug treated), at a particular time point represents the average of four technical replicates. To facilitate inter-study comparisons, we normalized temporal expression of each gene by its 0-h expression value (vertical gray bars). The untreated and drug-treated parasites appear to have similar expressions during the IDC, indicating a lack of alteration in the parasite’s gene-expression program due to drug treatment. (B) IDC progression of untreated and drug-treated parasites. To quantify IDC progression, we computed the Pearson correlation between gene-expression profiles of each replicate from this study and those from the hourly sampled data of Llinas et al.13. The closed circles denote the mean of maximum correlation between the two datasets (untreated or treated) at the time points indicated on the x-axis. (C) Average fold change in gene expression relative to its expression at 0 h (FC0h) in untreated and drug-treated parasites. Plasmodium genes shown with red circles have fourfold (log2 scale) different FC0h after treatment as compared to untreated conditions, while genes shown with gray circles have a less than twofold (log2 scale) difference; all other genes are shown using green circles. The open and closed circles denote genes with unknown and known (or putative) functions, respectively. h.p.i. hour post infection, PA21A092 a pyrazoleamide drug, PfEMP1 Plasmodium falciparum erythrocyte membrane protein 1.
Figure 2B demonstrates that IDC progression of the drug-treated parasites (red curve) is mostly comparable with that of the untreated parasites (black curve) for the first 24 h, but differing after 32 h into the IDC. Because the expression profile of specific genes may have been affected due to the treatment, we next quantified the average change in expression, relative to 0 h (FC0h), of each gene under untreated and drug-treated conditions. Figure 2C shows substantially altered genes in response to drug treatment using red circles and green circles. We found that only eight genes (out of 5,851) had FC0h values that were fourfold (log2 scale) different in magnitude after treatment as compared to untreated conditions (Fig. 2C, red circles). Of these eight genes, five encode paralogs of 18S ribosomal RNA, two encode the erythrocyte membrane protein (PfEMP1), and one encodes a conserved Plasmodium protein of unknown function (PF3D7_1142900). The PfEMP1 proteins are encoded by a family of approximately 60 var genes14. An individual parasite only expresses a particular var gene at a time, while keeping all other genes transcriptionally silent14. Therefore, the dramatically higher expression of one gene and lower expression (PfEMP1, Fig. 2C) of the other is likely due to var gene switching and not a result of pyrazoleamide treatment.
To further identify treatment-specific perturbations in the expression of Plasmodium genes, we next examined the functional annotation of the genes with a ≥ twofold difference in FC0h values (depicted by green circles in Fig. 2C), and identified 66 (out of a total of 102) genes with known or putative functions (Table S1: Sheet 1). Among these were carbonic anhydrase (PF3D7_1140000) and aquaglyceroporin (PF3D7_1132800). Carbonic anhydrase converts water and carbon dioxide into carbonic acid and protons15, while aquaglyceroporin is a water and ammonia transporter of the parasite16. The activities of carbonic anhydrase17 and aquaglyceroporin16 are pH dependent. PfATP4-associated drugs at lethal concentrations are known to alter pH homeostasis in malaria parasites, and these results suggest that a sublethal dose of a pyrazoleamide drug may also cause alterations in the pH of the treated parasites.
To characterize the effects of PA21A092 treatment on Plasmodium physiology, we performed an enrichment analysis using gene sets available in the Malaria Parasite Metabolic Pathway (MPMP) database18. We included previously published transcriptomic data obtained from P. falciparum maintained under three different perturbations11,12,19 to isolate treatment-specific effects. Table 1 lists gene sets altered only in pyrazoleamide-treated parasites. Our analysis revealed a substantial downregulation of genes encoding protein kinases (Cell cycle regulation in Table 1) and phosphatases (Phosphatase-related proteins in Table 1). Both protein kinases and phosphatases play critical roles in phosphorylation and dephosphorylation of regulatory proteins during the IDC20.
Although not directly related to our findings, an absence of dephosphorylation processes in P. falciparum results in blockage of parasite egress during IDC21, suggesting that a sublethal dose of PA21A092 might also be affecting parasite egress during the IDC. Interestingly, a previous study showed premature schizogony-like events in parasites exposed to lethal doses of PfATP4 inhibitors6. With a dose of 12 nM PA21A092, we did not observe any premature release of merozoites at any of the time points in our study. Table S2 presents the percentage of altered genes in all the gene sets obtained from the MPMP database18.
Metabolomic data from untreated and pyrazoleamide-treated cultures
We performed HCA on the metabolomic data from untreated and drug-treated cultures of uninfected erythrocytes (uRBC) and parasite-infected erythrocytes (iRBC) to identify drug-induced alterations in P. falciparum metabolism. Figure 3A illustrates the average abundance of metabolites detected in quadruplicate samples of uRBC and iRBC cultures. Surprisingly, HCA revealed uRBC metabolites with substantially different abundances under untreated and drug-treated conditions (annotated 1 and 2, Fig. 3A).
Global analysis of metabolomic data from untreated and pyrazoleamide-treated cultures of uninfected and parasite-infected erythrocytes. (A) Hierarchical clustering analysis of metabolomic data from untreated and drug-treated cultures of uninfected (uRBC) and parasite-infected (iRBC) erythrocytes. Each row denotes a metabolite and each column represents a sampled time point during the experiment. The abundance of a metabolite at each time point, under a particular condition (uRBC or iRBC) and perturbation (untreated or drug treated), represents the average of four technical replicates. To facilitate comparisons between different conditions and perturbations, we normalized the abundance of each metabolite by its abundance at 0 h. The vertical gray bars denote the 0-h time point under each condition and perturbation. Clades annotated “1” and “2” suggest alteration in abundance of uRBC metabolites due to the drug treatment. (B) Fold change, relative to 0 h (FC0h), in abundance of uRBC metabolites under untreated and drug-treated conditions. The uRBC metabolites with |Δ|≥ 4 are shown using red circles, where Δ = \({\text{F}}{\text{C}}_{\text{0h}}^{\text{PYZ}}/{\text{F}}{\text{C}}_{\text{0h}}^{\text{CTL}}\). \({\text{F}}{\text{C}}_{\text{0h}}^{\text{PYZ}}\) and \({\text{F}}{\text{C}}_{\text{0h}}^{\text{CTL}}\) denote FC0h of an uRBC metabolite in the presence and absence of the drug, respectively. The gray circles denote metabolites with |Δ|< 2, while green circles represent metabolites with |Δ|≥ 2 but less than 4. The open and closed circles, respectively, denote metabolites that vary substantially and minimally over a 2-day period22. (C) Volcano plot showing significantly altered metabolites (p < 0.001) due to parasite infection in untreated cultures. The red markers denote significantly altered metabolites with FCIDC ≤ 0.5, while green markers represent significantly altered metabolites with FCIDC ≥ 2; here, FCIDC denotes fold change in average abundance of a metabolite in iRBC cultures relative to uRBC cultures. (D) Volcano plot showing significantly altered metabolites (p < 0.001) due to parasite infection in drug-treated cultures. The red and green markers, respectively, denote significantly altered metabolites with FCIDC ≤ 0.5 and FCIDC ≥ 2.
To quantify the effect of pyrazoleamide treatment on uRBC metabolite abundances, we computed average fold change in abundance of each metabolite relative to 0 h (FC0h) for the untreated and drug-treated datasets. Figure 3B shows metabolites with substantially different FC0h between untreated and drug-treated conditions using red and green circles. In a recent effort, we quantified the expected variance in abundance of uRBC metabolites detected concurrently in distinct donors22. Using these expected variances in uRBC metabolite abundances22, we found that most of the substantially different metabolites, denoted by open red circles in Fig. 3B, tended to vary in abundance irrespective of a perturbation. Deoxythreonate, shown using closed red circles in Fig. 3B, was primarily detected in urine samples of humans23,24 and was not detected robustly (raw count > 1,000; see “Materials and methods”) in our previously analyzed metabolomic datasets22. Furthermore, Chen et al.25 demonstrated that serum deoxythreonate levels correlate with age of blood donors. We are not aware of the ages of the donors for the independent experiments performed under untreated and drug-treated conditions, but historically the age of the donors for our blood-collection protocol has been between 18 and 60 years.
Because uRBC metabolite abundances can vary between different donors, we normalized the metabolomic data from untreated and drug-treated cultures by an internal standard (see “Materials and methods”) prior to computing fold change in average abundance of metabolites due to parasite infection (FCIDC). Figure 3C,D, respectively, show metabolites in untreated and drug-treated cultures that are perturbed significantly (p < 0.001) due to parasite infection. Deoxythreonate appears to be depressed (Fig. 3C) or elevated (Fig. 3D) in untreated and drug-treated conditions, respectively, but this is largely due to the variation in its abundance (Fig. 3B) in the uRBC cultures of the two independent experiments.
Table 2 lists FCIDC values of significantly altered metabolites due to the pyrazoleamide treatment. The presence of substantial abundance variations of a specific metabolite in uninfected cultures can either hide or enhance any apparent effect on this metabolite due to the drug treatment22. Therefore, in addition to performing two-way analysis of variance (ANOVA), we also computed z-scores of variation in abundances of uRBC metabolites, in the absence and presence of the drug, using their expected variations reported in Tewari et al.22. This allowed us to identify uRBC metabolites having significantly different (p < 0.001) variation in abundance as compared to their expected variation22. In Table 2, we annotated such uRBC metabolites using asterisks. For example, creatinine and hippurate have FCIDC values greater than three in the presence of the pyrazoleamide drug, however, the temporal abundance of both these metabolites in the drug-treated uRBC cultures was significantly different than their expected variation in uRBC cultures22. Interestingly, hippurate is a blood metabolic biomarker of gut microbiome diversity in humans26.
The FCIDC of sedoheptulose-7-phosphate, a pentose phosphate pathway (PPP) metabolite, was smaller in the drug-treated cultures than in the untreated cultures, suggesting a lower production of reduced nicotinamide adenine dinucleotide phosphate (NADPH) due to the treatment. The FCIDC of lactate, a product of the ATP-generating phase of glycolysis, was higher in the drug-treated cultures as compared to the untreated cultures. To further investigate the effect of drug on glycolysis, we compared the temporal profile of glucose and five glycolysis intermediates at the sampled time points between the two conditions. We found that the abundance of these metabolites showed similar alterations up to the 24-h time point, but the values differed considerably afterwards (Table S3). For the untreated cultures, glycolytic intermediates are only 3% of their initial abundances at the 40-h time point, while for the drug-treated cultures, they are 32% of their initial abundances, indicating decreased consumption of these intermediate metabolites (Table S3). Similarly, glycolytic products (pyruvate and lactate) reached 748% of their initial abundance in the untreated cultures, but only 274% of their initial abundance in the drug-treated parasite cultures. Taken together, these alterations suggest a modulation of carbohydrate metabolism in pyrazoleamide-treated parasites, which is consistent with partial inhibition of parasite proliferation during the first IDC.
Although the tricarboxylic acid (TCA) cycle is not essential during the IDC27, we found significant alterations in abundance of several TCA cycle metabolites (α-ketoglutarate, citrate, fumarate, and malate). We also found that FCIDC of sphinganine-1-phosphate (Sa1P), an analogue of sphingosine-1-phosphate (S1P), in the treated cultures was almost twice as large as that in the untreated cultures. An increased abundance of S1P is associated with an increase in glycolysis in iRBCs28 and, hence, rationalizes the approximately twofold increase in FCIDC of lactate in the treated cultures as compared to the untreated cultures. However, these results do not explain how an increased lactate abundance is associated with a reduced PPP flux in the treated parasites.
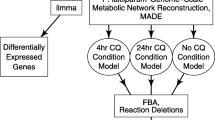
Therefore, to identify mechanistic consequences of pyrazoleamide treatment in P. falciparum, we integrated the time-resolved transcriptomic and metabolomic data from the treated parasites with a genome-scale metabolic model of P. falciparum11,12. In parallel, we also coupled transcriptomic and metabolomic data from untreated parasites with the metabolic model. The model-predicted metabolism of untreated and drug-treated parasites allowed us to gain additional insights into pyrazoleamide-induced metabolic adjustments in P. falciparum.
Metabolism of pyrazoleamide-treated P. falciparum
The integration of condition-specific and time-resolved data (transcriptomic and metabolomic) with the P. falciparum metabolic network model provides a functional description of the changes taking place in untreated and drug-treated parasites during the IDC. Figure 4A–F show rates of macromolecules synthesized during the IDC of untreated (black markers and curves) and drug-treated (red markers and curves) parasites. Our model predicted an inhibition of RNA, protein, and polyamine synthesis in the treated parasites (Fig. 4A–C). By contrast, our model predicted an increase in rates of lipid synthesis, cofactor synthesis, and inorganic ion generation in the treated parasites (Fig. 4D–F).
Model-predicted rates of macromolecule syntheses in untreated and pyrazoleamide-treated P. falciparum. (A) Rate of RNA synthesis. (B) Rate of protein synthesis. (C) Rate of polyamine synthesis. (D) Rate of lipid synthesis. (E) Rate of cofactor synthesis. (F) Rate of inorganic ion generation. The error bars denote one standard error of the mean for 10 simulations performed after adding Gaussian noise, with zero mean and 10% standard deviation in the transcriptomic data under untreated and drug-treated conditions. CTL parasites maintained in pure RPMI medium, gDW gram dry weight of the parasite, PYZ parasites maintained in RPMI medium with a sublethal dose of a pyrazoleamide, RPMI, Roswell Park Memorial Institute.
To quantify net accumulation or depletion of essential macromolecules and metabolites under both conditions, we numerically integrated their synthesis rates in the absence and presence of the drug, and then computed a ratio for each metabolite (and macromolecule) using the values obtained for untreated and drug-treated parasites (denoted by Δm). Table 3 lists Δm of select essential macromolecules and metabolites. The model predicted a net accumulation of approximately 30% and 50% in DNA and phospholipids, respectively, in the treated parasites (Table 3). In contrast to DNA and phospholipid accumulation, the model predicted a net depletion of approximately 40% and 15% in RNA and protein macromolecules, respectively, in the treated parasites (Table 3). Table S4 tabulates Δm of all essential macromolecules and metabolites.
Vaidya et al.10 found that treatment of malaria parasites by lethal doses of pyrazoleamide compounds disrupts its sodium ion homeostasis, which is essential for phosphate uptake via PfPiT29. Our model predicted inhibition of PfATP4 and PfPiT activities in the pyrazoleamide-treated parasites (Fig. S1). As compared to untreated conditions, our integrated data and model analysis predicted substantially less efflux of cytosolic protons from the pyrazoleamide-treated parasites under steady-state conditions (Fig. S1). Interestingly, several experiments5,7,10 performed using lethal doses of sodium ion homeostasis disruptors have observed an increase in cytosolic pH in the treated parasites. Note that our model predictions do not suggest that cytosolic pH decreases in parasites exposed to sublethal doses of PA21A092, but rather indicate that transport of protons is minimal across the parasitic plasma membrane under steady-state conditions. Together, these results suggest an alteration in the homeostasis of sodium ions and protons in parasites treated with a sublethal dose of PA21A092.
Pyrazoleamide treatment caused temporal upregulation in expression of the gene encoding aquaglyceroporin (Table S1: Sheet 1), which transports ammonia (or water) across the parasite’s plasma membrane16, suggesting an accumulation of ammonia (or water) in the treated parasites. In line with these observations, our model predicted an accumulation of ammonium, the ionized form of ammonia, in the treated parasites (Δm = 3.22, Table 3). The PfPdx1 enzyme of P. falciparum synthesizes pyridoxal 5-phosphate (PLP) from ammonia, glyceraldehyde 3-phosphate, and ribose 5-phosphate30, which is also captured in our model simulations (Δm = 1.76, Table 3). Specifically, in parasites treated with a sublethal dose of PA21A092, the synthesis of PLP increased 30 h post infection, prior to which the PfPdx1 activity was minimal as compared to the untreated parasites.
Metabolic consequences of a sublethal pyrazoleamide dose in P. falciparum
Figure 5 shows metabolic adjustments and adaptations in pyrazoleamide-treated P. falciparum as predicted by the integrated metabolic network model. We found that a sublethal dose of PA21A092 substantially inhibited PfATP4 activity during the late ring and early trophozoite stages of the IDC (Fig. S1). Because PfATP4 activity maintains the sodium ion gradient across the P. falciparum plasma membrane5, these results suggest that sublethal doses of pyrazoleamides also disrupt sodium ion homeostasis in P. falciparum. The model simulations predicted an inhibition of amino acid uptake in the treated parasites. Although P. falciparum obtains most amino acids from hemoglobin degradation, it relies on isoleucine uptake from its host to maintain growth during the IDC31. Accordingly, the reduction of isoleucine uptake in the treated parasites is compatible with the model-predicted 15% reduction in protein synthesis of the treated parasites (Table 3).
Effects of pyrazoleamide treatment in P. falciparum. Our integrated model analysis suggests that a sublethal dose of pyrazoleamide (PYZ) caused a reduction in PfATP4 flux, disrupting the homeostasis of sodium ions (Na+) and protons (H+) and, ultimately, reducing the osmotic pressure. The model also indicated reduced uptake of amino acids (AA), such as isoleucine, and a decreased rate of protein synthesis in drug-treated parasites. We found that the parasites adjust carbohydrate metabolism to produce an osmolyte (MI) that can counter the effects of drug treatment. However, this metabolic adjustment results in decreased flux via PPP, consequently decreasing NADPH and PRPP, and exacerbating the effects of the drug by increasing oxidative stress. Hence, our results suggest that downstream consequences of metabolic adjustments contribute to the efficacy of pyrazoleamides in P. falciparum. AA amino acid, ADP adenosine diphosphate, ATP adenosine triphosphate, G6P glucose 6-phosphate, GLC D-glucose, Hb hemoglobin, MI myoinositol, NADPH nicotinamide adenine dinucleotide phosphate, Pi inorganic phosphate, PPP pentose phosphate pathway, PRPP 5-phosphoribosyl 1-pyrophosphate, tRNA transfer RNA.
We found 49% more accumulation of phospholipids in pyrazoleamide-treated parasites as compared to untreated parasites (Table 3). Upon investigation of the metabolic network, we found that the treated parasites redirected flux via glucose 6-phosphate (G6P, Fig. 5) to enhance flux via myoinositol-3-phosphate synthase (MIPS, Fig. S2) and myoinositol-phosphate phosphatase (MIPP, Fig. S2). Together, these enzymes form the first and second steps of de novo synthesis of myoinositol (MI, Fig. 5). These results suggest that in drug-treated parasites the synthesis of MI—a precursor for phosphatidylinositol (PtdIns) synthesis—is increased causing an enhanced phospholipid synthesis in the treated parasites (Fig. 4D). The model simulations predict a downregulation in PPP flux as a direct consequence of this metabolic adaptation. They also showed that the downstream consequences of this metabolic adaptation, e.g., downregulated redox metabolism, increased oxidative stress, downregulated hemoglobin degradation, etc., are detrimental to the parasite and, hence, could contribute to effectiveness of pyrazoleamides in P. falciparum.
Discussion
In this work, we exposed Dd2 parasites to a predetermined sublethal concentration (12 nM) of a pyrazoleamide drug (PA21A092), causing an approximately 35% reduction in growth over a single IDC. The only observable morphological change was a slight decrease in the number of developing merozoites per schizont in the drug-treated parasites at the 40-h time point (Fig. 1D). We hypothesized that when exposed to a sublethal dose, P. falciparum would invoke metabolic pathways to maintain growth during the IDC to counter the effects of the administered drug. To characterize P. falciparum responses against a sublethal dose of pyrazoleamide, we collected transcriptomic and metabolomic data during the IDC from untreated and drug-treated parasites. We classified P. falciparum responses into two categories: i) immediate or proximal response to the drug and ii) a downstream overall metabolic adaptation in response to the drug perturbation.
Immediate consequences of pyrazoleamide treatment in P. falciparum
Pyrazoleamides belong to a new class of drugs that disrupt sodium ion homeostasis in P. falciparum10,32 by inhibiting the activity of the cation ATPase PfATP4. In addition to disruption of sodium ion homeostasis, pyrazoleamide treatment also causes swelling of the parasite33, changes in lipid composition of the parasite plasma membrane6, and inner membrane complexes6. Our analyses of the transcriptomic data revealed an approximately twofold (log2 scale) higher FC0h of the genes encoding carbonic anhydrase and aquaglyceroporin in drug-treated parasites as compared to the untreated parasites. Carbonic anhydrase catalyzes the conversion of water and carbon dioxide into bicarbonate ions and protons, and aquaglyceroporin transports water across the parasite plasma membrane. Because the activity of carbonic anhydrase increases with pH17, these results suggest that expression of carbonic anhydrase is upregulated due to an increase in pH that is observed in parasites treated with a lethal dose of pyrazoleamides10.
Flannery et al.7 found that parasites treated with 10 times the IC50 of a pyrazoleamide drug (GNF-Pf4492) did not progress beyond the early trophozoite stages. Vaidya et al. (unpublished data) found that conditional knockdown of PfATP4 also results in arrest at the trophozoite stage. Although the pyrazoleamide dose used in our experiments is sublethal, we found that PA21A092 inhibited PfATP4 activity (Fig. S1) and caused downregulation of genes (FCgene ≤ 0.5) related to parasite egress/invasion signaling in the treated parasites (Table S2).
Das et al.6 reported an increased accumulation of cholesterol in the parasite plasma membrane and appearance of rhoptry-like organelles in parasites treated with a lethal dose of PA21A050. Interestingly, we found substantial alterations in abundance of genes related to lipid raft and rhoptry proteins (Table S2), however, these gene-set alterations were not exclusively in the treated parasites as we also observed them in the hypoxanthine-limited parasites. In fact, we found that both hypoxanthine-limited parasites and pyrazoleamide-treated parasites shared a similar degree of gene-set alteration (Table S2). In the next Section, we discuss the mechanistic reason for this similarity between the two perturbations.
Metabolic adaptations in pyrazoleamide-treated P. falciparum
To gain a mechanistic understanding of pyrazoleamide effects, we coupled the time-resolved data with a genome-scale model of P. falciparum. Our model predicted an inhibition of PfATP4 enzyme activity in the pyrazoleamide-treated parasites. We found that pyrazoleamide treatment caused approximately 15% inhibition in the rate of protein synthesis. In our model, reduced protein synthesis was primarily due to an inhibition of isoleucine uptake, which occurs via symport of protons. We found that in the treated parasites, proton exchange was minimal (Fig. S1) and was associated with the suppressed activity of membrane-bound NADP transhydrogenase (THDpp, Fig. S2), causing an inhibition in uptake of isoleucine – an amino acid essential for parasite growth31.
In addition to protein synthesis inhibition, we found substantially suppressed RNA synthesis activity in the treated parasites (Δm = 0.61, Table 3). Upon investigation of the metabolic network, we found that the activity of orotate phosphoribosyltransferase (ORPT)—an enzyme involved in synthesis of uridine 5′-triphosphate—was lower in the treated parasites as compared to the untreated parasites. The activity of ORPT requires orotate and 5-phosphoribosyl 1-pyrophosphate (PRPP), of which PRPP was limited in treated parasites due to the suppression of the PPP (Fig. 5 and Table 2). The PRPP metabolite is also a substrate of hypoxanthine phosphoribosyltransferase, producing inositol monophosphate from hypoxanthine12. Because our gene-set analyses revealed similarities in hypoxanthine-limited and pyrazoleamide-treated parasites (Table S2), we next compared macromolecular synthesis rates of hypoxanthine-limited parasites12 with pyrazoleamide-treated parasites, identifying reduced rates of polyamine and RNA synthesis under both perturbations. A closer examination of pyrazoleamide-treated parasite metabolism revealed that the limited availability of PRPP through the PPP also caused partial inhibition of hypoxanthine phosphoribosyltransferase activity in the treated parasites; inhibition of hypoxanthine phosphoribosyltransferase activity would have mimicked hypoxanthine limitation in the treated parasites, and caused activation of gene sets that are similar to the hypoxanthine-limited parasites (Table S2).
Conclusions
To date, P. falciparum has developed some degree of resistance to all antimalarial drugs. Hence, to overcome resistance and maintain treatment efficacy, drug-combination therapies involving two (or more) drugs have become the frontline malaria treatment34,35. However, maintaining the long-term efficacies of these combination therapies is heavily influenced by the development and selection of mutations in the targeted parasite populations that allow the parasite to grow in the presences of drugs, a consideration that is difficult to a priori assess when developing or selecting combination therapies. In this study, our goal was to identify P. falciparum processes that could be relevant to the development of drug resistance. Towards this end, we cultured P. falciparum in the presence of a 12 nM pyrazoleamide dose, a drug whose prolonged exposure leads to resistance-conferring mutations in the PfATP4 enzyme7,10.
To characterize metabolic responses of P. falciparum during the IDC, we employed a genome-scale metabolic model integrating time-resolved metabolomic and transcriptomic data obtained in the absence and presence of the pyrazoleamide drug. We found that the treated parasites redirected flux via G6P to enhance synthesis of myoinositol and subsequently PtdIns, which regulates numerous cellular functions36 and plays a role in artemisinin resistance via phosphatidylinositol 3-kinase37. Moreover, because myoinositol has an osmoregulatory function in different species38,39,40, it can directly counter the effects of pyrazoleamide treatment, i.e., dysregulation of sodium ion homeostasis. Together, these results suggest that the mechanism underlying drug-resistance development in P. falciparum is specific to the administered drug. Therefore, rationally motivated drug-combination therapies that are designed to treat malaria and suppress drug-specific resistance development would be more efficacious in the long-term as compared to drug-combination therapies determined using their treatment efficacy alone.
Materials and methods
Synchronous culture of P. falciparum
We cultured the Dd2 strain of P. falciparum in human O-positive erythrocytes using methods and conditions similar to those described previously11,12,41. Briefly, we maintained parasite-infected erythrocytes in Roswell Park Memorial Institute (RPMI) 1640 medium (Gibco, Gaithersburg, MD) supplemented with 20 mM HEPES, 12.5 µg/mL hypoxanthine, 25 µg/mL gentamicin, 0.5 µM R-lipoic acid, 0.3% sodium bicarbonate, and 0.5% AlbuMAX II (Life Technologies, Inc., Carlsbad, CA). Addition of lipoic acid in the medium allows us to control the availability of this essential nutrient rather than relying on adventitious uptake from host erythrocytes. We obtained erythrocytes as part of an Institutional Review Board (IRB) approved phlebotomy protocol (NA_00019050), and used them within two days after collection. As per the Johns Hopkins Medicine IRB X, the use of human erythrocytes under the IRB protocol (NA_00019050) is not considered human-subject research. We obtained blood samples for untreated and drug-treated experiments on two different days. Prior to the experiments, we depleted the procured blood of white blood cells (WBCs). Towards this end, we first removed the buffy coat following two rounds of density gradient centrifugation and then used a NEO High Efficiency Leukocyte Reduction Filter (Haemonetics) to remove any remaining WBCs. Polymerase chain reaction using primers specific for the gene encoding 16S ribosomal RNA (5′-GGAGCAAACAGGATTAGATACCC and 5′-CACCATCTGTCACTCTGTTAACC) confirmed the absence of contaminating mycoplasma.
Pyrazoleamide dose determination
To determine the optimal concentration of PA21A092 that will allow us to assess the effect of drug on parasite processes without prematurely killing the parasites, we seeded highly synchronized ring-stage parasites (2–6 h) at 5% parasitemia and 2% hematocrit (HCT) in a 96-well flat bottom plate and grew the cultures in the presence of different concentrations of PA21A092 (8–24 nM) at 37 °C for 40 h. We used 0.1% DMSO as the no-drug control and 1 µM chloroquine as a positive control. After 40 h, we diluted the treated cultures 1:10 in drug-free media and cultured for another ~ 30 h. At this point, we quantified the parasite growth using SYBR-Green I (Invitrogen) and NxT flow cytometer as described previously19. Based on the forward and side scatter, intact erythrocytes were gated initially. A single-cell population was selected based on forward scatter. Finally, parasitemia was obtained by gating for SYBR-Green-positive erythrocytes.
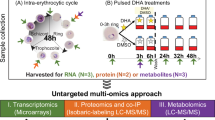
Sample collection for metabolomic and transcriptomic profiling
We collected samples of parasites treated with the pyrazoleamide (PYZ) drug for comparison to samples collected in the absence of the drug (CTL). For each sample set, we passed 300 mL of synchronized parasite culture through a magnetically activated cell sorting (MACS) XS column (Miltenyi Biotec, Auburn, CA) to seed four flasks, serving as the source for quadruplicate measurements of transcriptomic and metabolomic data. Each flask had 75 mL of culture at 2% HCT. We monitored the parasite culture via blood smears until 0–2 h post-merozoite invasion, at which point (time 0 of the experiment) we replaced the culture medium with fresh medium containing 12 nM PA21A092 (for the PYZ sample set) or drug-free medium (for the CTL sample set). Beginning at time 0, we collected 10.5 mL of the re-suspended parasite culture from each flask every 8 h until the 48-h time point and pelleted the erythrocytes by centrifugation at 400 g for 5 min. Following aspiration of the media, we transferred 100 µL and 50 µL of the cell pellets to 1.5-mL tubes for metabolomic and transcriptomic profiling, respectively. We flash-froze the tubes in liquid nitrogen and stored them at –80 °C. For each sample set (PYZ and CTL), we sent quadruplicate samples of the cell pellets collected at six time points (0, 8, 16, 24, 32, and 40 h) to Metabolon, Inc. (Durham, NC) for metabolite analysis, and quadruplicate samples of the cell pellets collected at seven time points (0, 8, 16, 24, 32, 40, and 48 h) to the Johns Hopkins Genomic Analysis and Sequencing Core Facility for microarray analysis using an Agilent microarray chip (AMADID 037237; Santa Clara, CA). We deposited the transcriptomic data in the National Center for Biotechnology Information Gene Expression Omnibus repository (GEO accession number: GSE176469).
In parallel, we maintained four flasks with 50 mL of uRBCs at 2% HCT in media containing PA21A092 to serve as controls for the PYZ-iRBC samples, and four additional flasks with no drug to serve as controls for the CTL-iRBC samples. Following the above-mentioned procedure, we collected, centrifuged, and aspirated 7 mL of uninfected culture from each flask. We transferred 100 µL of the cell pellets to 1.5-mL tubes, and immediately flash-froze the tubes and stored them at –80 °C. We sent quadruplicate samples of these control cultures collected at six time points to Metabolon, Inc. for each of the two conditions (PYZ and CTL) for metabolomic analysis. We did not send the 48 h samples for metabolomic analysis. We show the metabolomic data as provided by Metabolon, Inc. in Table S5.
To evaluate the impact of PA21A092 treatment on parasite growth, we assessed parasitemia by a blood-smear test at every time point. At 40 h, we transferred 20 µL of the pelleted iRBCs from the four drug-treated flasks to 25-cm2 flasks and diluted them to 2% HCT with fresh blood and 5 mL drug-free medium. We determined parasitemia in these diluted cultures by NxT flow cytometry at 64 h and compared them to untreated controls.
Global analyses of data
We used the built-in MATLAB function clustergram to perform HCA and identify substantial alterations in metabolite and gene abundances due to pyrazoleamide treatment. Within clustergram, we implemented Ward’s algorithm using a Euclidean distance metric to group metabolites and genes with similar temporal profiles between the two conditions, i.e., untreated and drug treated. For all analyses pertaining to metabolomic data, we only included metabolites having a robust signal (raw count > 1,000) across all the replicates and time points under both conditions.
To compute FC0h of genes under a given condition, we used the built-in MATLAB function bootstrp to generate 1,000 bootstrap samples from quadruplicate samples of the dataset for each gene and time point. We then computed a fold change in expression of a gene at each time point with respect to 0 h for all the bootstrap samples, i.e., \({{F}}{{C}}_{{i}}^{{t}} \, {=} \, \left\{{{a}}_{{i}}^{{t}}/{{a}}_{{i}}^{0}{:} \, \forall{ i} \, \epsilon {{N}}\right\}\). Here, N denotes the total number of bootstrap samples, \({\text{a}}_{0}\) and \({\text{a}}_{\text{t}}\), respectively, represent abundance levels of the gene at 0 and t hours of the experiment from any of the four replicates. We then computed FC0h of a gene by:
where Ntp denotes the number of sampled time points during the IDC and \({\text{F}}{\text{C}}_{\text{j}}\) represents the median value of \({\text{F}}{\text{C}}_{\text{i}}^{\text{t}}\) across all bootstrap samples. To compute FCgene of each gene due to a perturbation, e.g., pyrazoleamide treatment or hypoxanthine limitation, we generated 1,000 bootstrap samples, based on gene-expression data obtained from quadruplicates samples of untreated and drug-treated parasites for all time points, using the built-in MATLAB bootstrp function. Afterwards, we computed FCgene using the following:
where gcontrol and gtreatment denote the median expression value of a gene in the untreated and drug-treated parasites, respectively. To compute FCIDC of each metabolite, we followed the bootstrap procedure described in Tewari et al.41, and to estimate FC0h and identify an internal standard (a sphingomyelin metabolite), we used the procedure described in Tewari et al.22. Table S6 lists the criterion (ζ value) for selecting the identified internal metabolite.
To perform gene-set analysis, we obtained a mapping of Plasmodium genes to the 509 gene sets available in the MPMP database18. For each gene set under each perturbation, we counted genes with |FCIDC|≥ twofold and denoted the result as a fraction of the total number of genes (Ngenes). For the results shown in Table 1, we only considered gene sets with more than 10 genes.
Predicting effects of pyrazoleamide treatment on P. falciparum metabolism
To capture PfATP4-associated effects, we modified the P. falciparum network model11 to include the metabolic reaction for PfATP4 as described by Spillman et al.32. We used the built-in COBRA toolbox42 function addReaction to add the metabolic reaction of PfATP4 and assigned the Plasmodium gene PF3D7_1211900 to the reaction. The activity of PfATP4 is essential to maintain a sodium ion gradient, which in turn is required for phosphate uptake via PfPiT in P. falciparum29. We also included the metabolic reaction of PfPiT, as described in Saliba et al.29, to capture the effects of the sodium ion gradient on phosphate uptake in P. falciparum. We did not include the sodium-proton exchanger in the model because the gene (PF3D7_1303500) encoding this exchanger is not essential in P. falciparum43. The PfATP4-associated drugs cause pH changes in the treated parasites, presumably by the activity of V-type proton ATPases32. However, P. falciparum can extrude protons via potassium-dependent (PF3D7_1456800) and -independent transporters (PF3D7_1235200) as well. In our model, we encompassed all these reactions by including a proton transporter to extrude protons from the parasite (Fig. S1).
To simulate the effects of pyrazoleamide on P. falciparum metabolism, we used previously described methods11,12,44. Briefly, we integrated the time-resolved transcriptomic and metabolomic data obtained from untreated and pyrazoleamide-treated Dd2 parasites with the P. falciparum metabolic network model. To capture the effects of the drug, we assumed that pyrazoleamide treatment caused a 50% reduction in the biomass of the treated parasites as compared to untreated parasites. We used the built-in MATLAB function trapz to numerically integrate the simulated rates of essential macromolecules and metabolites (Table 3).
Data availability
The experimental datasets generated and analyzed during the current study are included with this manuscript. The MATLAB© scripts used to generate the computational results can be obtained from corresponding authors on reasonable request.
References
WHO. Report on Antimalarial Drug Efficacy, Resistance and Response: 10 Years of Surveillance (2010–2019) (World Health Organization, 2020).
WHO. World Malaria Report 2019 (World Health Organization, 2019).
Thu, A. M., Phyo, A. P., Landier, J., Parker, D. M. & Nosten, F. H. Combating multidrug-resistant Plasmodium falciparum malaria. FEBS J. 284, 2569–2578 (2017).
Fairhurst, R. M. Understanding artemisinin-resistant malaria: What a difference a year makes. Curr. Opin. Infect. Dis. 28, 417–425 (2015).
Spillman, N. J. & Kirk, K. The malaria parasite cation ATPase PfATP4 and its role in the mechanism of action of a new arsenal of antimalarial drugs. Int. J. Parasitol. Drugs Drug Resist. 5, 149–162 (2015).
Das, S. et al. Na+ influx induced by new antimalarials causes rapid alterations in the cholesterol content and morphology of Plasmodium falciparum. PLOS Pathog. 12, e1005647 (2016).
Flannery, E. L. et al. Mutations in the P-type cation-transporter ATPase 4, PfATP4, mediate resistance to both aminopyrazole and spiroindolone antimalarials. ACS Chem. Biol. 10, 413–420 (2015).
Rottmann, M. et al. Spiroindolones, a potent compound class for the treatment of malaria. Science 329, 1175–1180 (2010).
Jimenez-Diaz, M. B. et al. (+)-SJ733, a clinical candidate for malaria that acts through ATP4 to induce rapid host-mediated clearance of Plasmodium. Proc. Natl. Acad. Sci. U.S.A. 111, E5455–E5462 (2014).
Vaidya, A. B. et al. Pyrazoleamide compounds are potent antimalarials that target Na+ homeostasis in intraerythrocytic Plasmodium falciparum. Nat. Commun. 5, 5521 (2014).
Tewari, S. G. et al. Metabolic survival adaptations of Plasmodium falciparum exposed to sublethal doses of fosmidomycin. Antimicrob. Agents Chemother. 65, e02392-e2420 (2021).
Tewari, S. G. et al. Short-term metabolic adjustments in Plasmodium falciparum counter hypoxanthine deprivation at the expense of long-term viability. Malar. J. 18, 86 (2019).
Llinas, M., Bozdech, Z., Wong, E. D., Adai, A. T. & DeRisi, J. L. Comparative whole genome transcriptome analysis of three Plasmodium falciparum strains. Nucleic Acids Res. 34, 1166–1173 (2006).
Pasternak, N. D. & Dzikowski, R. PfEMP1: An antigen that plays a key role in the pathogenicity and immune evasion of the malaria parasite Plasmodium falciparum. Int. J. Biochem. Cell Biol. 41, 1463–1466 (2009).
Krungkrai, S. R. & Krungkrai, J. Malaria parasite carbonic anhydrase: Inhibition of aromatic/heterocyclic sulfonamides and its therapeutic potential. Asian Pac. J. Trop. Biomed. 1, 233–242 (2011).
Zeuthen, T. et al. Ammonia permeability of the aquaglyceroporins from Plasmodium falciparum, Toxoplasma gondii and Trypansoma brucei. Mol. Microbiol. 61, 1598–1608 (2006).
Berg, J. M., Tymoczko, J. L., Gatto, G. J. Jr. & Stryer, L. Biochemistry 8th edn. (W. H. Freeman, 2015).
Ginsburg, H. & Abdel-Haleem, A. M. Malaria parasite metabolic pathways (MPMP) upgraded with targeted chemical compounds. Trends Parasitol. 32, 7–9 (2016).
Swift, R. P. et al. A mevalonate bypass system facilitates elucidation of plastid biology in malaria parasites. PLOS Pathog 16, e1008316 (2020).
Khalife, J., Freville, A., Gnangnon, B. & Pierrot, C. The multifaceted role of protein phosphatase 1 in Plasmodium. Trends Parasitol. 37, 154–164 (2021).
Paul, A. S. et al. Co-option of Plasmodium falciparum PP1 for egress from host erythrocytes. Nat. Commun. 11, 3532 (2020).
Tewari, S. G. et al. Inter-study and time-dependent variability of metabolite abundance in cultured red blood cells. Malar. J. 20, 299 (2021).
Kassel, D. B., Martin, M., Schall, W. & Sweeley, C. C. Urinary metabolites of L-threonine in type 1 diabetes determined by combined gas chromatography/chemical ionization mass spectrometry. Biomed. Environ. Mass Spectrom. 13, 535–540 (1986).
Appiah-Amponsah, E. et al. Identification of 4-deoxythreonic acid present in human urine using HPLC and NMR techniques. J. Pharm. Biomed. Anal. 50, 878–885 (2009).
Chen, L. et al. Comparative blood and urine metabolomics analysis of healthy elderly and young male singaporeans. J. Proteome Res. 19, 3264–3275 (2020).
Pallister, T. et al. Hippurate as a metabolomic marker of gut microbiome diversity: Modulation by diet and relationship to metabolic syndrome. Sci. Rep. 7, 13670 (2017).
Ke, H. et al. Genetic investigation of tricarboxylic acid metabolism during the Plasmodium falciparum life cycle. Cell Rep. 11, 164–174 (2015).
Sah, R. K., Pati, S., Saini, M. & Singh, S. Erythrocyte sphingosine kinase regulates intraerythrocytic development of Plasmodium falciparum. Sci. Rep. 11, 1257 (2021).
Saliba, K. J. et al. Sodium-dependent uptake of inorganic phosphate by the intracellular malaria parasite. Nature 443, 582–585 (2006).
Muller, I. B. et al. The assembly of the plasmodial PLP synthase complex follows a defined course. PLoS ONE 3, e1815 (2008).
Babbitt, S. E. et al. Plasmodium falciparum responds to amino acid starvation by entering into a hibernatory state. Proc. Natl. Acad. Sci. U.S.A. 109, E3278–E3287 (2012).
Spillman, N. J. et al. Na+ regulation in the malaria parasite Plasmodium falciparum involves the cation ATPase PfATP4 and is a target of the spiroindolone antimalarials. Cell Host Microbe 13, 227–237 (2013).
Dennis, A. S. M., Lehane, A. M., Ridgway, M. C., Holleran, J. P. & Kirk, K. Cell swelling induced by the antimalarial KAE609 (cipargamin) and other PfATP4-associated antimalarials. Antimicrob. Agents Chemother. 62, e00087-e118 (2018).
Alven, S. & Aderibigbe, B. Combination therapy strategies for the treatment of malaria. Molecules 24, 3601 (2019).
van der Pluijm, R. W. et al. Triple artemisinin-based combination therapies versus artemisinin-based combination therapies for uncomplicated Plasmodium falciparum malaria: A multicentre, open-label, randomised clinical trial. Lancet 395, 1345–1360 (2020).
Arendse, L. B., Wyllie, S., Chibale, K. & Gilbert, I. H. Plasmodium kinases as potential drug targets for malaria: Challenges and opportunities. ACS Infect. Dis. 7, 518–534 (2021).
Mbengue, A. et al. A molecular mechanism of artemisinin resistance in Plasmodium falciparum malaria. Nature 520, 683–687 (2015).
Ma, A. et al. Osmoregulation by the myo-inositol biosynthesis pathway in turbot Scophthalmus maximus and its regulation by anabolite and c-Myc. Comp. Biochem. Physiol. A Mol. Integr. Physiol. 242, 110636 (2020).
Dai, G., Yu, H., Kruse, M., Traynor-Kaplan, A. & Hille, B. Osmoregulatory inositol transporter SMIT1 modulates electrical activity by adjusting PI(4,5)P2 levels. Proc. Natl. Acad. Sci. U.S.A. 113, E3290–E3299 (2016).
Thurston, J. H., Sherman, W. R., Hauhart, R. E. & Kloepper, R. F. Myo-inositol: A newly identified nonnitrogenous osmoregulatory molecule in mammalian brain. Pediatr. Res. 26, 482–485 (1989).
Tewari, S. G., Swift, R. P., Reifman, J., Prigge, S. T. & Wallqvist, A. Metabolic alterations in the erythrocyte during blood-stage development of the malaria parasite. Malar. J. 19, 94 (2020).
Heirendt, L. et al. Creation and analysis of biochemical constraint-based models using the COBRA Toolbox vol 3.0. Nat. Protoc. 14, 639–702 (2019).
Sanderson, T. & Rayner, J. C. PhenoPlasm: A database of disruption phenotypes for malaria parasite genes. Wellcome Open Res. 2, 45 (2017).
Tewari, S. G., Prigge, S. T., Reifman, J. & Wallqvist, A. Using a genome-scale metabolic network model to elucidate the mechanism of chloroquine action in Plasmodium falciparum. Int. J. Parasitol. Drugs Drug Resist. 7, 138–146 (2017).
Acknowledgements
The authors thank Ms. Maria E. Kuhrmann for her comments on a previous version of this manuscript. The authors also thank Ms. Anne E. Jedlicka and Ms. Amanda Dziedzic at Johns Hopkins University’s Genomic Analysis and Sequencing Core Facility for transcript profiling. The opinions and assertions contained herein are the private views of the authors and are not to be construed as official or as reflecting the views of the U.S. Army, the U.S. Department of Defense, or the Henry M. Jackson Foundation for the Advancement of Military Medicine, Inc. This paper has been approved for public release with unlimited distribution.
Funding
This research was primarily funded by the U.S. Army Medical Research and Development Command Network Science Initiative, Fort Detrick, MD, under Awards W81XWH-14–2-0134 and W81XWH-20-C-0031. This work was also supported by the U.S. Army Medical Research and Development Command under Contract No. W81XWH-15-C-0061 (STP), National Institutes of Health Grant R01 AI125534 (STP), training grant T32 AI007417 (KR), the Johns Hopkins Malaria Research Institute, and the Bloomberg Family Foundation.
Author information
Authors and Affiliations
Contributions
S.G.T. carried out the omics-data analysis, computational analysis, and wrote the initial draft of the manuscript. B.K., R.E., and K.R. performed all the experiments. S.T.P. conceived the study, supervised the experiments, and helped write the manuscript. J.R. conceived the study, supervised the computational analysis, and helped edit the manuscript. A.B.V. supervised the data analysis and helped edit the manuscript. A.W. conceived and supervised the study, and helped edit the manuscript. All authors have read and approved the final version of the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Tewari, S.G., Kwan, B., Elahi, R. et al. Metabolic adjustments of blood-stage Plasmodium falciparum in response to sublethal pyrazoleamide exposure. Sci Rep 12, 1167 (2022). https://doi.org/10.1038/s41598-022-04985-7
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-022-04985-7
- Springer Nature Limited
This article is cited by
-
Metabolic responses in blood-stage malaria parasites associated with increased and decreased sensitivity to PfATP4 inhibitors
Malaria Journal (2023)
-
Recent metabolomic developments for antimalarial drug discovery
Parasitology Research (2022)