Abstract
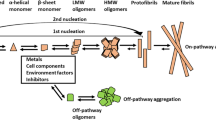
Amyloid β (Aβ) peptide aggregation plays a central role in Alzheimer’s disease (AD) etiology. AD drug candidates have included small molecules or peptides directed towards inhibition of Aβ fibrillogenesis. Although some Aβ-derived peptide fragments suppress Aβ fibril growth, comprehensive analysis of inhibitory potencies of peptide fragments along the whole Aβ sequence has not been reported. The aim of this work is (a) to identify the region(s) of Aβ with highest propensities for aggregation and (b) to use those fragments to inhibit Aβ fibrillogenesis. Structural and aggregation properties of the parent Aβ1–42 peptide and seven overlapping peptide fragments have been studied, i.e. Aβ1–10 (P1), Aβ6–15 (P2), Aβ11–20 (P3), Aβ16–25 (P4), Aβ21–30 (P5), Aβ26–36 (P6), and Aβ31–42 (P7). Structural transitions of the peptides in aqueous buffer have been monitored by circular dichroism and Fourier transform infrared spectroscopy. Aggregation and fibrillogenesis were analyzed by light scattering and thioflavin-T fluorescence. The mode of peptide-peptide interactions was characterized by fluorescence resonance energy transfer. Three peptide fragments, P3, P6, and P7, exhibited exceptionally high propensity for β-sheet formation and aggregation. Remarkably, only P3 and P6 exerted strong inhibitory effect on the aggregation of Aβ1–42, whereas P7 and P2 displayed moderate inhibitory potency. It is proposed that P3 and P6 intercalate between Aβ1–42 molecules and thereby inhibit Aβ1–42 aggregation. These findings may facilitate therapeutic strategies of inhibition of Aβ fibrillogenesis by Aβ-derived peptides.
Similar content being viewed by others
Introduction
Extracellular fibrillar deposits of the amyloid β (Aβ) peptide in cerebral parenchyma and vasculature constitute a major histopathological trait of Alzheimer’s disease (AD)1,2,3. The presence of Aβ plaques in brains of AD patients led to the amyloid cascade hypothesis, positing direct involvement of Aβ aggregates in AD etiology1,4. The cytotoxicity of Aβ has been documented in numerous in vivo and in vitro studies, lending support for the central role of Aβ in AD1,3,5,6,7,8.
Aβ is derived from the amyloid precursor protein (APP) through cleavage by β- and γ-secretases9,10. Due to poor sequence specificity of these enzymes, Aβ species of varying numbers of amino acid residues are present in human brain. Along with the most prevalent forms, i.e. 42- and 40-residue species (Aβ1–42 and Aβ1–40), many shorter peptides, generated by post-cleavage enzymatic processing, are also present. In addition, extensive somatic gene recombination in human neurons results in numerous APP variants with insertions, deletions, or missense mutations, including early termination codons and mutations linked to early onset AD11. In effect, the total Aβ pool in human brain is a heterogenous assembly of peptide species of different size and with distinct structural features and toxicities12,13,14,15,16,17.
Aβ peptides are inherently polymorphic and undergo dynamic structural transitions during fibrillogenesis18,19,20,21,22,23,24,25,26,27,28,29. Monomers and oligomers can adopt a variety of secondary/tertiary structures, including fractions of α-helix, β-sheet, various types of turn and loop conformations18,19,20,21,30. Fibrils of Aβ1–40 and Aβ1–42 acquire characteristic parallel, in-register cross-β structure, i.e. intermolecular β-sheets with H-bonding along the fibrillar axis24,31,32,33,34. The atomic resolution structures of Aβ1–40 and Aβ1–42 fibrils have been determined by solid state NMR and other techniques and revealed relatively similar secondary structures but distinct 3-dimensional folds. Aβ1–40 monomers in the fibrillar structure adopt a U-shaped conformation with two β-strands (residues 10–22 and 30–40) separated by a loop and with unordered N-terminus31 (Fig. 1). In contrast, Aβ1–42 forms S-shaped structure with a larger number (3–5) of shorter β-strands32,33,34 (Fig. 1).
Aβ1–42 amino acid sequence in single-letter format, with stretches corresponding to peptides P1 through P7 shown by horizontal bars of different colors above or below the sequence, along with respective theoretical isoelectric points (according to ExPASy ProtParam function, https://web.expasy.org/protparam). Horizontal blue arrows below the sequence show the segments corresponding to β-sheet structures in Aβ1–40 or Aβ1–42 fibrils determined experimentally, with Protein Data Bank entries shown on the right-hand side. Segments left blank are described as loop or turn regions. In the lower panel, one structure for Aβ1–40 and three structures for Aβ1–42 in fibrillar form are shown (fibril axis and the direction of interstrand H-bonding are perpendicular to the plane of the image). Backbone regions in β-sheet and turn/loop conformations are colored blue and rose, respectively.
Toxicity depends on particular Aβ species, the aggregation state, and the structural features of peptide assemblies3,20,35,36,37,38. Monomers and mature fibrils are essentially nontoxic or mildly toxic; the main cytotoxic effect is exerted by intermediate oligomeric assemblies3,20,37,38,39,40,41,42,43. In human brain, Aβ1–40 is found in higher concentrations than Aβ1–42, but the latter is more toxic, which is believed to result from its increased hydrophobicity and tendency towards β-sheet formation and aggregation26,35,36,44,45,46,47,48. Enzymatically modified forms of Aβ are produced in vivo by N-terminal trimming followed by cyclization of glutamic acid, which possess superior cytotoxicity deemed to stem from their distinct aggregation and structural properties such as faster aggregation into β-sheet fibrillar assemblies49,50 or stabilization of toxic oligomeric structures5. Apparently, the aggregation pathways and accompanying structural transitions in various Aβ species underlie the ultimate cytotoxic effects. Therefore, detailed biophysical studies, including per-residue or segmental secondary structure formation properties and their role in peptide aggregation, are important for better understanding and, potentially, regulation of the amyloidogenic behavior of Aβ. On the other hand, identification of specific structural and aggregation propensities of defined stretches of Aβ and their effects on the behavior of the full-length Aβ1–42 may help develop peptide-based drugs to inhibit Aβ aggregation or immunotherapy approaches for Aβ clearance.
Such strategies of AD drug development have generated promising results. For example, DNA plasmid vaccine targeting Aβ N-terminus (Aβ1–11) elicited strong immune response and inhibited Aβ pathology in various animal models of AD51,52,53. The first potentially disease-modifying AD drug aducanumab was recently approved by the Food and Drug Administration based on its ability to clear Aβ from brain parenchyma. Short peptides, related or unrelated to Aβ sequence, have been employed to inhibit Aβ aggregation or disperse preformed oligomers and fibrils and impede Aβ-induced cytotoxicity54,55,56,57,58,59,60,61,62,63,64,65,66,67. The inhibitory effect of peptides on Aβ aggregation and toxicity stems from their ability to intercalate into Aβ assemblies and prevent toxic oligomer formation, which may or may not correlate with their own self-aggregation propensities. Thus, a mutated peptide, KLVFWAK, mimicking the Aβ sequence 16–22 (KLVFFAE), had low self-aggregation tendency yet preferentially interacted with oligomeric and fibrillar structures of Aβ67. The native sequence of Aβ16–22, on the other hand, has been identified as a fibrillogenic segment with high antiparallel β-sheet formation propensity22,68,69. Among several Aβ segments incorporated in a macrocyclic structure, the 16–22 segment inhibited Aβ1–40 aggregation and cytotoxicity60. These effects were interpreted in terms of binding of the fragment to Aβ oligomers and inhibition of fibril nucleation. Interestingly, the same 16–22 sequence, as well as the hydrophobic C-terminal stretch from Lys28 onward, were found to be the most fibrillogenic regions in the context of several Aβ species containing 38 to 43 amino acids21. Consistent with these findings, Aβ28–42 and Aβ29–42 displayed fast aggregation and β-sheet fibril formation propensities unlike some other C-terminal Aβ fragments36. Moreover, among twelve C-terminal Aβ1–42 fragments, Aβ28–42 exerted maximum toxic effect on cultured PC-12 cells36 and suggested correlation between amyloidogenicity and toxic effects of the peptides. Furthermore, two C-terminal fragments, Aβ31–42 and Aβ39–42, were able to rescue neuronal cells from Aβ1–42 cytotoxicity by interfering with formation of toxic oligomers70.
Thus, fragments of Aβ appear to affect the aggregation and toxicity of full-length Aβ by direct interaction with the latter. Studies on the details of binding of fragments to the full-length peptide revealed diverse modes of interaction. A C-terminal Aβ fragment Aβ33–37 (GLMVG) was shown to bind to the C-terminus of Aβ1–42, possibly the homologous sequence, and suppress its aggregation, neuronal membrane damage and synaptotoxicity61. Puzzlingly, another short C-terminal fragment, Aβ39–42, preferentially interacted with the N-terminus of Aβ1–42 in oligomeric form71,72,73. The aforementioned mutated peptide, KLVFWAK, also interacted in a non-homologous manner with the C-terminus of Aβ aggregates67.
These findings indicate that Aβ aggregation can be effectively modulated by Aβ-derived peptides. However, a complete understanding of the differential effects of Aβ segments along the entire Aβ1–42 sequence has not been achieved. This work aimed at systematic studies on 10 to 12 amino acid residue long overlapping fragments of Aβ1–42 (Fig. 1) to understand their intrinsic β-sheet formation and fibrillogenesis properties and their effects on same properties of the parent peptide. Out of seven peptide fragments, three stretches, corresponding to sequence 11–20, 26–36, and 31–42, have been identified that demonstrated strong β-sheet and aggregation propensity. The former two sequences were able to inhibit Aβ1–42 fibrillogenesis by up to 80%, whereas the C-terminal dodecapeptide, as well as the fragment 6–15, displayed moderate (20–30%) inhibitory potency. Further spectroscopic studies shed light on the mode of peptide-peptide interactions. Our data thus provide significant new information on intrinsic biophysical properties of Aβ1–42 segments and their effects on the behavior of the full-length peptide, which may be useful in designing peptide-based AD therapies.
Results
Choice of peptide fragments
Seven overlapping fragments of Aβ1–42 have been studied, i.e. sequences 1–10, 6–15, 11–20, 16–25, 21–30, 26–36, and 31–42, named P1, P2, P3, P4, P5, P6, and P7, respectively. All peptides have been N-terminally acetylated and C-terminally amidated to eliminate the charge effects of terminal amine and carboxyl groups. We chose to study overlapping peptides to avoid missing the effects of sequences at boundaries of consecutive segments. The locations of these seven fragments along the Aβ1–42 sequence and their respective theoretical isoelectric point (pI) values are presented in Fig. 1.
Secondary structure from FTIR spectroscopy
The lyophilized peptides were dissolved in hexafluoroisopropanol (HFIP) to disperse possible pre-existing aggregates74. HFIP solution of the Aβ1–42 peptide was spread over a CaF2 disk, air-dried and desiccated for 15 min, followed by measurements of Fourier transform infrared (FTIR) spectra. The dry peptide acquired α-helical structure manifested by a sharp amide I peak at 1658 cm−1 (Figure S1 of Supplementary Information). Before cleavage from APP, most of the Aβ sequence corresponds to the transmembrane α-helix embedded in neuronal membranes75. Hence, this result is consistent with the intrinsic α-helical propensity of the Aβ1–42 sequence in dehydrated state. On the other hand, determining the structure of the dehydrated peptide is important for analysis of structural transitions once the peptide is exposed to an aqueous medium.
Next, HFIP solutions of Aβ1–42 and the seven fragments were separately dried in glass vials and aqueous (D2O) buffer was added to reach concentrations of 50 μM for Aβ1–42 and 100 μM for the fragments. The samples were constantly stirred with a small magnetic stir bar, and FTIR spectra of 70 μL aliquots were measured periodically over 100 h to examine structural changes of the peptides. The amide I spectrum of Aβ1–42 initially showed two prominent components at 1672 and 1647 cm−1, suggesting mostly β-turn and unordered or loop conformations76,77,78 (Fig. 2a). Upon incubation in buffer, the 1647 cm−1 peak was gradually replaced by a component at 1627 cm−1, indication formation of intermolecular β-sheet structure76,78. This is consistent with the fibrillar structure of Aβ1–42 comprising β-sheet and β-turn/loop conformations32,33,34 (Fig. 1) and indicates that formation of turn/loop structures precedes aggregation and fibrillogenesis through intermolecular H-bonding.
FTIR spectra at indicated times of incubation in D2O-based buffer (25 mM NaCl, 25 mM Na,K-phosphate, pD 7.2) for Aβ1–42 (a) and peptides P1 (b), P2 (c), P3 (d), P4 (e), P5 (f), P6 (g), and P7 (h) at 25 °C. The concentrations of the peptides were 50 μM for Aβ1–42 and 100 μM for the fragments. Times of incubation in buffer of all fragments are indicated in panel (b).
The fragments exhibited distinct structural transitions in aqueous buffer. All fragments except for P7 initially displayed significant fraction of β-turn structure in addition to irregular structure, indicated by amide I components in 1672–1670 cm−1 and 1640 cm−1 regions, respectively (Fig. 2). Among these 6 peptides, only P3 and P6 showed β-sheet component at 1628–1623 cm−1 (Fig. 2d, g). In case of P3, both β-turn and β-sheet structures grew at the expense of irregular structure (Fig. 2d), whereas in P6 the β-turn became the dominant structure upon prolonged incubation in buffer (Fig. 2g). P7 exhibited a unique behavior; it rapidly formed β-sheet structure with a distinctive peak around 1626 cm−1, which gradually shifted to higher wavenumbers possibly reflecting transition to turn and/or α-helix structures. Thus, peptides P3, P6, and P7 display β-sheet formation propensity with distinct dynamics of structural transitions whereas the other four peptides tend to adopt turn/loop conformations.
Secondary structure from CD spectroscopy
The structural transitions of the peptides in aqueous medium were further analyzed by a complementary method, circular dichroism (CD). In the same time, the question was addressed as to how the structural transitions of the peptides are related to fibrillogenesis. This was accomplished by near-simultaneous measurements of CD spectra, light scattering, and thioflavin-T (ThT) fluorescence of same peptide samples in aqueous buffer for over 100 h. CD spectra were measured for peptides dissolved in HFIP, in dry state, and in aqueous buffer. The spectrum of the full-length Aβ1–42 peptide in HFIP displayed a deep minimum at 204 nm and a shoulder around 220 nm (Figure S2a of Supplementary Information), indicating the presence of type I and II β-turns, β-strand, and possibly α-helix conformations79,80. CD spectra of seven peptide fragments exhibited two negative ellipticities as well. The stronger component occurred in the 205–208 nm region (P1 and P2) or 195–200 nm region (P3 through P7) (Figure S2b–h of Supplementary Information), assigned to β-turn and unordered structures, respectively81,82,83. The weaker component was located between 220 and 230 nm, which can be assigned to α-helix, β-turn and β-strand conformations.
The solvent was removed by desiccation and CD spectra of dry peptides were measured. These spectra showed significant structural changes upon drying. Aβ1–42 adopted mostly α-helical structure, evidenced by a negative component at 223 nm and weaker one at 208 nm (Figure S2a of Supplementary Information), which are generated by n–π* and π–π* electronic transitions80,83. Similar spectral features were displayed by P1, P3, and P6. Weaker intensity of the π–π* transition can be attributed to a more flexible or distorted α-helix80. In spectra of peptides P4 and P5, the π–π* transition was stronger, suggesting formation of more stable α-helix. Dry peptides P2 and P7 showed just one component around 220–223 nm, which is most likely to represent type-I β-turn and β-strand structures79,81.
Following structural characterization of the peptides dissolved in HFIP and in dry state, aqueous buffer was added and structural transitions were followed by recording consecutive CD spectra. Spectra of Aβ1–42 initially showed a shallow negative band around 217 nm, which upon incubation in buffer for up to 48 h became a well-defined band accompanied with a positive component of nearly equal intensity around 200 nm, clearly indicating formation of β-sheet structure (Fig. 3a). Structural transitions in peptide fragments occurred at slower rate, therefore they were examined for longer time. Peptides P1, P2, P4, and P5 retained strong negative ellipticity around 200 nm and weaker components between 225 and 235 nm (217 nm for P2), implying these peptides assume mostly unordered conformation with fractions of β-turn/β-strand structures (Fig. 3b, c, e, f). In contrast, peptides P3, P6, and P7 readily (in less than 1 h) adopted defined secondary structures (Fig. 3d, g, h). Spectra of these three peptides display a negative n-π* band around 220–225 nm and strong positive ellipticity in the 200 nm region. These spectral features can be assigned to twisted β-sheet structure, as the intense π–π* transition is diagnostic for twisted β-strands, and the n–π* band can be significantly red-shifted depending on the degree of strand twist80,81,82,84,85. Note that CD spectra become excessively noisy below 200 nm due to the relatively long optical path-length (4 mm) of the cuvette, which was used for simultaneous light scattering and fluorescence measurements. However, this does not invalidate above interpretations of CD spectra in terms of peptides’ structural features.
CD spectra of 50 μM Aβ1–42 (a) and 100 μM of peptides P1 (b), P2 (c), P3 (d), P4 (e), P5 (f), P6 (g), and P7 (h) constantly stirred by a tube rotator in buffer (25 mM NaCl, 25 mM phosphate, pH 7.2) for time periods indicated in panel a for Aβ1–42 and in panel (b) for all fragments. All measurement were conducted at 25 °C.
Some differences have been observed between peptide structures deduced from FTIR and CD measurements, such as different relative contents of β-turn and irregular structures. These differences can be attributed to non-identical sample preparation and handling protocols, as Aβ structure is exquisitely sensitive to sample preparation conditions, including the method of sample stirring24,32,86,87,88. In addition, both N-terminal acetylation and C-terminal amidation introduce additional amide bonds that generate FTIR signal in the unordered region, i.e. between 1650 and 1638 cm−1. Nonetheless, both FTIR and CD data unveil important common structural trends of these peptides, i.e., P3, P6, and P7 stand out by exhibiting strong β-sheet propensity, like the parent peptide, while peptides P1, P2, P4, and P5 tend to stay in β-turn or unordered conformations.
Aggregation and fibrillogenesis
ThT fluorescence and static light scattering at 90 degrees were used to examine the kinetics of aggregation and fibrillogenesis of the peptides in aqueous buffer. Of note, these measurements were conducted in parallel with CD on same samples, which allowed analysis of structural changes, peptide aggregation and fibril formation in real time, excluding sample-to-sample errors. Increase in ThT fluorescence around 480 nm when excited at 435 nm was interpreted as fibril formation with an understanding that non-fibrillar structures could also bind ThT and produce fluorescence in that region. Aβ1–42 underwent fibrillogenesis without delay, as indicated by increase in ThT fluorescence around 480 nm (Fig. 4a, b). Saturation of ThT fluorescence was reached by around 1 day of incubation in buffer with constant stirring on a rotary mixer. Light scattering at 550 nm increased with steeper initial rate and approached saturation by 10–20 h of incubation (Fig. 4c, d). The difference in the initial kinetics of fluorescence and light scattering may reflect rapid formation of non-fibrillar aggregates that do not bind ThT.
ThT fluorescence with excitation at 435 nm (a) and light scattering with incident light at 550 nm (c) of 35 μM Aβ1–42 in buffer (25 mM NaCl, 25 mM Na,K-phosphate, pH 7.2) stirred by a tube rotator for time periods indicated in panel a. Time dependence of ThT fluorescence averaged between 475 and 485 nm and of light scattering averaged between 550 and 555 nm are presented in panels (b) and (d), respectively, which represent average data and standard deviations from three independent experiments.
In samples with peptide fragments P1, P2, P4, and P5, very weak ThT fluorescence was detected (Fig. 5a, b, d, e). Peptides P3 and P6 generated higher level of ThT fluorescence, and peptide 7 produced maximum ThT fluorescence (Fig. 5c, f, g), unraveling a trend of fibrillogenesis propensity P7 > P6 ≈ P3 >> P4 ≈ P5 ≈ P2 ≈ P1 ≈ 0 (Fig. 5h). Statistical t-test analysis of all data of Fig. 5 between 24 and 104 h of incubation in buffer produced nearly 98% credibility for significant difference between fibrillogenesis propensities of P7 and P3 (p = 0.0214 ± 0.0084) and more than 96% credibility between P7 and P6 (p = 0.0385 ± 0.0101). P3 and P6 were not significantly different (p = 0.593 ± 0.241), and the difference between P3 and P1, P2, P4, P5 was 96.9–98.7% significant.
ThT fluorescence spectra of P1, P2, P3, P4, P5, P6, and P7 (panels a through g, respectively) with excitation at 435 nm, at 25 °C. Time of incubation of peptide samples in buffer (25 mM NaCl, 25 mM Na,K-phosphate, pH 7.2) at 70 μM concentration with constant stirring by a tube rotator is as follows: 0.1 h (dark blue), 2.0 h (light blue), 4.0 h (turquoise), 16.0 h (green), 40.0 h (yellow), 64.0 h (orange), 104.0 h (red). Panel h shows the time dependence of ThT fluorescence averaged between 475 and 485 nm for all seven peptides. Average data from three independent experiments. Error bars are shown only for P1, P3, P4, P6, P7 to maintain clarity. Error bars for P2 and P5 are of similar magnitude as for those of P1 and P4.
Concurrent light scattering measurements indicated relatively weak signal for peptides P1, P2, P4, and P5 and about threefold higher signal for peptides P3, P6, and P7 (Figure S3 of Supplementary Information). For the first group of peptides, t-test analysis produced p values between 0.172 ± 0.085 (P2/P5) and 0.747 ± 0.232 (P1/P4), and for the second group of peptides, p values varied between 0.366 ± 0.270 (P3/P7) and 0.685 ± 0.159 (P3/P6), indicating little statistical difference within each group. The difference between these two groups was significant (> 96%), as indicated by p values for P2/P3 of 0.0367 ± 0.0122. Negligible ThT fluorescence but appreciable light scattering of peptides P1, P2, P4, and P5 apparently reflects formation of non-fibrillar aggregates by these fragments. Higher level of ThT fluorescence by P7 compared to P3 and P6 but similar light scattering (cf. Fig. 5h and S3h) suggests a higher ThT binding capacity of P7, plausibly due to its exceptional hydrophobicity and/or a larger number of amino acid residues. Overall, these data provide evidence that peptides P3, P6, and P7 possess significant aggregation/fibril formation capability as compared to the other four Aβ fragments. In conjunction with structural data, it appears that the aggregation/fibrillogenesis property of these peptides correlates with their β-sheet formation ability.
Effects of Aβ fragments on Aβ fibrillogenesis
If some of peptide fragments demonstrate strong fibrillogenesis propensity, they might be able to intercalate into the assemblies of the parent peptide by binding to respective stretches and thereby interfere with its aggregation. To test this hypothesis, fibrillogenesis of Aβ1–42 was measured in the absence and presence of all seven peptide fragments at two-fold molar excess. In the presence of peptides P1, P2, P4, P5, and P7, ThT fluorescence of Aβ1–42 samples gradually increased and levelled off by 48 h of incubation with little effect of these fragments (Fig. 6a, b, d, e, g, h). In contrast, P3 and P6 strongly reduced the level of ThT fluorescence (Fig. 6c, f, h). Quantitative evaluation of inhibition of Aβ1–42 fibril formation by peptide fragments was performed by calculating the inhibition percentages (IP) for each fragment, as described in “Materials and methods”. P3 and P6 strongly inhibited Aβ1–42 fibrillization by 80.5 ± 9.0% and 77.7 ± 5.8%, respectively, with insignificant statistical difference between them (p = 0.673) (Fig. 7). Peptides P2 and P7 exhibited moderate inhibitory potencies (IP = 18.8 ± 2.1% and 28.7 ± 3.8%, respectively), with considerable statistical difference (p = 0.0163), and the other three peptides (P1, P4, and P5) were not potent inhibitors (IP < 5.0%; p values from 0.113 to 0.851). The difference between these three groups, presented by bars of distinct colors in Fig. 7, was statistically significant (p < 0.01).
ThT fluorescence spectra of 35 μM Aβ1–42 plus 70 μM P1 (a), P2 (b), P3 (c) P4 (d), P5 (e), P6 (f), and P7 (g) with excitation at 435 nm, at 25 °C. Time of incubation in buffer (25 mM NaCl, 25 mM Na,K-phosphate, pH 7.2) with constant stirring by a tube rotator is indicated in panel (a). Vertical axes are standardized for better comparison of relative fluorescence intensities. Panel (h) shows the time dependence of ThT fluorescence averaged between 475 and 485 nm. Average data from three independent experiments. Error bars are shown only for P1, P3, P6, P7 to maintain clarity. Error bars for P2, P4, and P5 are of similar magnitude as for those of P1 and P7.
Inhibition percentages for P1 through P7 calculated based on ThT fluorescence data (Figs. 4, 5, 6) averaged between 32 and 48 h of incubation of Aβ1–42 alone and with each peptide fragment at twofold molar excess in buffer (25 mM NaCl, 25 mM Na,K-phosphate, pH 7.2). Statistical difference between bars of different colors is significant (p < 0.01). Difference between bars of same color is less significant or insignificant: for blue bars, p values vary between 0.113 and 0.851, for green bars p = 0.0163, and for red bars p = 0.673.
Mode of peptide interactions from Phe-to-Tyr energy transfer
The next question we explored was whether the intrinsic fluorescence of peptide fragments can be utilized to gain insight into the mode of their interactions. Aβ1–42 is devoid of tryptophan, but has one tyrosine (Tyr) at position 10 and three phenylalanines (Phe) at positions 4, 19, 20. Phe to Tyr resonance energy transfer (RET) has been described earlier for a tryptophan-less protein, with Förster distance of R0 = 13.5 Å89,90, which is much shorter than that of many other donor–acceptor pairs, typically in the 20 Å to 60 Å range90. This provides a unique opportunity to examine the interactions between peptides containing Phe and Tyr by RET.
As both Tyr and Phe can be excited in a wide wavelength range90, an optimal wavelength needs to be identified where Phe absorbs maximally and Tyr absorbs minimally. Our measurements showed that this wavelength range was around 220 nm. Figure S4 of Supplementary Information shows how RET works for the P3/P1 donor/acceptor pair. Upon excitation at 220 nm, Phe and Tyr of peptides P3 and P1 emit around 287 and 310 nm, respectively (Figure S4a,b). In equimolar mixture of these peptides at same total peptide concentration, the intensity of Phe emission gradually decreases, indicating energy transfer to Tyr. Negative change of Phe fluorescence (DFPhe < 0) and positive change in Tyr fluorescence (DFTyr > 0) are clearly seen in difference spectra around 30 h of incubation of P1 and P3 (Figure S4c), indicating efficient RET.
As seen from Fig. 1, only peptide fragments P1, P2, P3, and P4 contain Phe and/or Tyr fluorophores, hence these four peptides have been analyzed in RET experiments. Figure 8 shows RET data for five peptide pairs that satisfy this criterion, i.e. possibility of Phe-to-Tyr RET. These data indicate slightly negative DFPhe in P1/P2 and P1/P4 systems, implying little RET in these peptide samples (Fig. 8a, c, f, h). Significant RET is seen in combinations of P3 with P1 and P2; in both cases negative DFPhe develops soon after mixing the peptides, reaches maximum values between 10 and 30 h, and returns to zero level between 50 and 100 h of incubation in buffer (Fig. 8b, d, g, i). In P2/P4 combination, there is relatively moderate RET (DFPhe < 0) for the whole time period of the experiment, i.e. ~ 200 h (Fig. 8e, j), indicating stable hetero-aggregation between peptides P2 and P4. Values of DFTyr are always positive, consistent with RET; their variations can be attributed to quenching effects of water and/or carbonyl groups of peptides during the aggregation process, as described earlier71.
Fluorescence resonance energy transfer difference spectra (a through e) and values for tyrosine and phenylalanine, averaged between 300 and 320 nm for tyrosine and 270 and 290 nm for phenylalanine (f through j) for five peptide systems, as indicated at the top. Fluorescence spectra have been smoothed by 13-point Savitzky–Golay least-squares polynomial algorithm before subtraction. See details in “Materials and methods”.
These data indicate that P3 not only readily self-aggregates, as seen from ThT fluorescence and light scattering data (Figs. 5h and S3h), but also effectively interacts with non-homologous stretches of Aβ, such as P1 and P2 segments, pointing to the “sticky” nature of P3. In the P1/P3 system, the peptides are likely to interact in parallel in-register manner, positioning Tyr10 against Phe20 (Figure S5a of the Supplementary Information). Strong RET between P2 and P3 suggests parallel out-of-register pairing, with Phe19 facing Tyr10 (Figure S5n of the Supplementary Information). These interactions apparently start falling apart at the end of first day of incubation (Fig. 8g, i), as P3 predominantly forms homologous aggregates, as described below. Data of Fig. 8e, j suggest that P2 and P4 form persistent aggregates, presumably of parallel out-of-register sense allowing juxtaposition of Phe19 and Tyr10 (Figure S5c of the Supplementary Information). It is interesting to note that in these proposed models of peptide interactions, ionic bonds between side chains of Asp7 and Lys16 play an important role by stabilizing hetero-aggregation in both P2/P3 and P2/P4 systems (Figure S5b,c of the Supplementary Information).
To gain insight into the mode of interaction of P3 with Aβ1–42, RET experiments have been conducted with this peptide system. Fluorescence spectra were measured for 35 μM Aβ1–42, 70 μM P3, and 35 μM Aβ1–42 + 70 μM P3 over a time period of 48 h, with excitation at 220 nm (Figure S6 of the Supplementary Information). First, we checked if RET effect was involved in self-aggregation of Aβ1–42 by subtracting the spectrum of Aβ1–42 measured at 1 h from spectra measured at 2 h and beyond. These difference spectra showed time-dependent changes in Tyr fluorescence, reflecting at least two effects, i.e., suspension of the dry peptide into the aqueous medium (decrease of fluorescence because of quenching by water) and sequestration from the aqueous medium through fibrillogenesis (increase of fluorescence), but a combination of negative ΔFPhe and positive ΔFTyr has not been detected (Figure S6d of the Supplementary Information). This means absence of RET between Aβ1–42 molecules, consistent with formation of parallel, in-register β-sheet aggregates. A similar procedure for the Aβ1–42 + P3 system, as well as subtraction of spectra of both Aβ1–42 and P3 from those of the combined sample at respective times, yielded RET effect at 32–48 h of co-incubation, i.e. DFPhe < 0 and DFTyr > 0 (Figure S6e,f of the Supplementary Information). This outcome implies that during interaction of P3 with Aβ1–42, Phe19 and Phe20 of at least a fraction of P3 come to close proximity to Tyr10 of Aβ1–42 at 32–48 h. However, the inhibitory effect of P3 on Aβ1–42 fibrillogenesis starts at earlier times (Fig. 6h), indicting that P3 interacts more effectively with other regions of Aβ1–42, which need to be identified in further studies.
Discussion
This work reports an analysis of structural and aggregation propensities of Aβ1–42 fragments and their effects on Aβ1–42 fibrillogenesis. The results facilitate a better understanding of segment-specific properties of Aβ1–42 and identify peptide fragments capable of inhibiting its aggregation. We chose 10- to 12-residue overlapping fragments because fragments shorter than 10 residues might be unable to exhibit robust secondary structure formation and fibrillogenesis abilities because of insufficient length. Although short Aβ fragments containing 6–8 residues have been shown to adopt parallel or antiparallel β-sheets in crystal form91,92, the hydrophobic C-terminal sequences of Aβ1–42 containing eight amino acid residues, as well as an 11-residue stretch (Aβ32–42), did not aggregate into β-sheet fibrils in aqueous buffer even at concentrations exceeding 100 mM36. On the other hand, use of overlapping sequences was important for characterization of segments at boundaries of consecutive stretches; properties of the boundary between P(i) and P(i + 2) were reflected in P(i + 1), which covers the C- and N-termini of flanking sequences.
Peptides P1 and P2 represent the N-terminus of Aβ, which has been consistently found in unordered conformation in the context of Aβ1–40 or Aβ1–4224,31,32,34,86,93. The Aβ1–8 fragment was found in extended coil confirmation in complex with a monoclonal antibody94. Despite its proposed role in stabilizing Aβ fibrils95, above data and our findings indicate that the N-terminus of Aβ is intrinsically unordered and unable to undergo fibrillogenesis. Consistent with this, RET data indicate that P1 and P2 do not form hetero-aggregates as well (Fig. 8a, f). This feature is apparently related with the amino acid composition of these segments; the paucity of nonpolar side chains and the excess negative charge (pI 4.54 and 5.72 for P1 and P2, respectively) may prevent aggregation due to electrostatic repulsion and lack of hydrophobic interactions. P1 is also a poor inhibitor of Aβ1–42 aggregation, whereas P2 demonstrates moderate inhibitory activity (Fig. 7), which probably stems from its C-terminal part, as P3 is highly aggregation prone and a potent inhibitor of Aβ1–42 fibrillogenesis.
The P3 segment corresponds to one of the two major β-strands of Aβ1–40 fibrillar structure Aβ1–4224,31,86 (Fig. 1). The whole P3 sequence or at least its central part are in β-strand conformation in fibrils formed by of Aβ1–42 as well32,33,34 (Fig. 1). P3 has one glutamic acid (Glu) and one lysine (Lys) residues and nearly zero net charge (pI = 7.01). These features would allow the peptide to form β-strand-like non-H-bonded out of register aggregates in antiparallel sense with oppositely charged side chains facing each other, as shown in Figure S7 of the Supplementary Information. This arrangement will juxtapose one Glu-Lys pair and one histidine pair between strands so the structure will be stabilized by Glu-Lys salt bridges and π-π stacking interactions between histidines96,97. As a fraction of imidazole side chains will be protonated due to the close-to-neutral pK = 6.3 ± 0.198, His-His attraction will be further strengthened by nucleophile–electrophile interactions. This structure may grow laterally beyond tetramer until decreasing internal energy is compensated by decreasing entropy at a minimum of free energy. The fibril will grow by means of parallel β-sheet formation stabilized by H-bonding perpendicular to the plane of side chains. The nonpolar overhangs may form turn or parallel β-sheet structure stabilized by hydrophobic interactions and π–π stacking interactions between C-terminal phenylalanines96. These conjectures are consistent with earlier observations of dimer formation by Aβ10–23 peptide that grow into β-sheet fibrillar structure99. FTIR data (Fig. 2d) show a low frequency (1623 cm−1) β-sheet component without a high-frequency counterpart diagnostic for antiparallel β-sheet, consistent with parallel β-sheet formation. Figures 5, 6, and 7 show that P3 forms fibrils and substantially blocks the fibrillogenesis of Aβ1–42. Inhibition of Aβ1–42 fibrillogenesis presumably occurs through intercalation of P3 aggregates between Aβ1–42 molecules and/or binding of P3 to the homologous or other segments of Aβ1–42. Heterologous binding is supported by the ability of P3 to effectively interact with P1 and P2 (Figures S5 and 8b, d, g, i). Consistent with our findings, it was reported earlier that Aβ segment 11–17 (EVHHQKL), i.e. 0.7 part of P3, but not other Aβ-derived short peptides, effectively blocked Aβ1–40-induced Ca2+ influx into cultured cells possibly by inhibiting membrane pore structure formation58.
Aβ region corresponding to P4 (Aβ16–25) contains the central hydrophobic core 17-LVFFA-21 that is deemed to be important for Aβ aggregation. Molecular modeling and experimental studies have shown that the Aβ16–22 fragment (KLVFFAE) forms antiparallel in-register β-sheet stabilized by nonpolar contacts between the central segment and ionic interactions between the side chains of N- and C-terminal Lys and Glu residues22,68,69,85,100,101,102. Monomeric Aβ16–25 was in coil or extended β-strand-like conformation and transitioned into antiparallel β-sheet through intermediate α-helical structure103. Our data show unordered plus β-strand conformation for P4 in HFIP in monomeric form (Figure S2e) but transition to β-sheet in aqueous buffer is not observed (Figs. 2e and 3e). This difference apparently results from the additional three residues in P4 compared to Aβ16–22. The presence of an additional anionic side chain of aspartic acid at position 23 results in excess negative charge and a low pI of 4.37 (Fig. 1). These considerations suggest that long-range electrostatic repulsion between short peptides may interfere with initial peptide aggregation and subsequent fibrillization. Furthermore, an analogue of Aβ16–22 with N-methyl amino acids in alternating positions was a potent inhibitor of Aβ1–40 fibrillogenesis unlike the unmodified peptide55, which is consistent with our data of negligible inhibitory effect of P4 on Aβ1–42 fibril formation (Figs. 6h and 7).
P5 encompasses the Aβ region that forms a loop between two major β-strands of Aβ1–40 fibrils characterized by two- or three-fold symmetry24,31,86 and has been viewed as a folding nucleus for Aβ104,105. This region is mostly in loop conformation in fibrils of Aβ1–42 as well32,33,34 (Fig. 1). In complex with an affibody protein, Aβ1–40 was found in a β-hairpin conformation with residues 24–29 forming a loop between two β-strands but without the Asp23-Lys28 salt bridge present in Aβ1–40 fibrils of two-fold symmetry106. The isolated P5 segment itself was mostly unordered36 or formed a loop with a central β-turn structure formed by residues 24 and 27 and stabilized by ionic interaction of Lys28 with Glu22/Asp23 and nonpolar contacts between Val24 and Lys28104. P5 did not undergo aggregation in aqueous medium36,104, in accord with sequence-based prediction of low aggregation propensity21. Transmission electron microscopic studies revealed relatively large (25–120 nm) globular particles formed by P5 with uncapped termini whereas the N-acetylated/C-amidated peptide formed smaller particles, implying low aggregation tendency105. Our FTIR data for P5 display a broad amide I band dominated by signal in the1670–1640 cm−1 range (Fig. 2f), revealing mostly unordered/turn structure of the peptide. This is confirmed by CD spectra with a deep minimum around 200 nm and a shallow negative band around 228 nm (Fig. 3f). Figure S3e,h and 5e, h demonstrate negligible light scattering and ThT fluorescence in P5 peptide samples, and Figs. 6h and 7 document the inability of P5 to affect Aβ1–42 fibrillogenesis, implying a poor potency of P5 towards homo- or hetero-aggregation.
P6 (Aβ26–36) resembles the Aβ25–35 peptide, which has been studied extensively due to its highly cytotoxic nature. In aqueous buffer at pH 4.0–5.5, the Aβ25–35 peptide underwent concentration-dependent unordered-to-β-sheet transition whereas its C-amidated counterpart was in random coil state107. At close to neutral pH, the N-acetylated/C-amidated peptide acquired β-sheet/β-turn structures108. In detergent micelles and in HFIP/water environment, Aβ25–35 was α-helical in the C-terminal region and less ordered or turn conformation in the N-terminus109,110. ThT and transmission electron microscopy assays revealed fibril formation by Aβ25–3559,61, and β-sheet breaking short peptides inhibited fibrillogenesis and cytotoxicity of the peptide59. Slight sequence differences in this region of Aβ resulted in significant differences in structural and aggregation properties. Biophysical and computational studies produced Aβ26–36 and Aβ25–35 structural models as β-hairpins containing two short β-strands separated by a turn111, in accord with our FTIR data (Fig. 2g). P6 formed β-sheet structure in aqueous buffer with red-shifted (~ 220 nm) n-π* CD band and exhibited higher fibril formation capability than similar undecapeptides Aβ25–35 and Aβ24–34111, consistent with our results (Figs. 3g, 5h).
P7 (Aβ31–42) represents the most hydrophobic C-terminal stretch of Aβ and plays an important role in its aggregation into β-sheet oligomers and fibrils35,36,70. Dynamic light scattering experiments identified two populations of Aβ1–42 aggregates with hydrodynamic radii (RH) 8–12 nm and 20–60 nm, which upon prolonged incubation in buffer exceeded 500 nm, while the P7 fragment formed large aggregates with RH around 100 nm36,70. In combination, P7 affected the aggregation of Aβ1–42 in a nontrivial way; it increased the abundance of larger particles but reduced the rate of their formation. Molecular modeling simulations showed that P7 intercalates into Aβ1–42 aggregates and thereby inhibits formation of toxic assemblies of Aβ1–4270. CD experiments showed that structural transitions of C-terminal fragments of Aβ1–42 were concentration dependent; at 62 μM, P7 gradually transitioned from unordered state to β-sheet structure during 96 h of incubation in buffer and formed fibrils36, in good agreement with our data (Figs. 3h and 5g, h).
In conclusion, this work thoroughly characterizes the intrinsic aggregation and structural properties of overlapping peptide segments throughout the Aβ1–42 sequence and the effects of these fragments on Aβ1–42 fibrillogenesis. Although previous studies have utilized a similar approach and have focused on certain parts of Aβ1–42, such as the C-terminal hydrophobic stretch, systematic dissection and analysis of the whole Aβ1–42 sequence has not been reported. Or data identify Aβ fragments that are capable of self-aggregation and β-sheet structure formation and substantially inhibit fibrillogenesis of Aβ1–42, presumably by intervention into forming aggregates or prevention of efficient intermolecular contacts between Aβ1–42 molecules. However, there is no universal correlation between self-aggregation/β-sheet propensities of the fragments and suppression of Aβ1–42 fibril formation. For example, P7, the most hydrophobic segment that undergoes efficient fibrilization (Fig. 5h) and β-sheet formation (Fig. 2h and 3h) is a relatively week inhibitor of Aβ1–42 fibrillization of (Figs. 6h and 7). This leads to a conclusion that peptide fragments with high homo-aggregation propensity prefer to self-associate instead of forming hetero-aggregates with the full-length peptide. These findings, specifically the potent inhibitory effects of P3 and P6 peptides on Aβ1–42 aggregation, may help develop new DNA-, mRNA- or peptide-based anti-AD therapeutic strategies.
Materials and methods
Materials
The synthetic Aβ1–42 peptide was from Innovagen (Lund, Sweden) and was > 98% pure. Synthetic peptide fragments, acetylated at N-terminus and amidated at C-terminus, were purchased from Peptide 2.0 Inc. (Chantilly, VA, USA) and were 98–99% pure, as verified by high performance liquid chromatography and mass-spectrometry. HFIP, ThT, salts, buffers, and most of other reagents were purchased from Sigma-Aldrich (St. Louis, MO, USA) or Fisher Scientific (Hanover Park, IL, USA). Quartz cuvettes for CD measurements were from Starna Calls, Inc. (Atascadero, CA, USA). CaF2 windows for FTIR studies were from Buck Scientific (East Norwalk, CT, USA). D2O (99.8% pure) was from Cambridge Isotope Laboratories (Andover, MA, USA).
Experimental procedures
Circular dichroism and fluorescence
Circular dichroism (CD) and fluorescence spectra were measured using a J-810 spectropolarimeter equipped with a fluorescence attachment and a temperature controller (Jasco, Tokyo, Japan) at 25 °C. Lyophilized peptides were dissolved in HFIP at a desired concentration and CD spectra were measured using a 4 mm × 4 mm rectangular quartz cuvette. Then, the solvent was removed by desiccation and CD spectra of the dry peptide stuck on the walls of the cuvette were measured. This was followed by addition of aqueous buffer (25 mM NaCl, 25 mM Na,K-phosphate, 20 μM ThT, pH 7.2) to achieve a target peptide concentration. The cuvette was capped with a Teflon stopper, sealed with parafilm, and rotated on a Fisherbrand™ Multi-Purpose Tube Rotator for several days. CD, ThT or intrinsic peptide fluorescence, and light scattering at 90 degrees were measured periodically on same sample, at pre-determined time pints, to follow the time dependence of structural changes and aggregation/fibrillogenesis of the peptides, using the multifunctional J-810 instrument. Measurements of these parameters on same samples in real time excluded sample to sample errors. For each peptide system, time-dependent spectroscopic measurements were repeated in three or four independent experiments.
ThT fluorescence was averaged between 475 and 485 nm, when excited at 435 nm, and plotted against time of incubation in buffer. For fluorescence RET experiments, incident light at 220 nm was used for predominant excitation of phenylalanine as compared to tyrosine, and spectra were measured between 200 and 400 nm. For each donor/acceptor peptide pair, the fluorescence of individual peptides was measured at 100 μM concentration, then spectra were recorded for the two peptides mixed at 50 μM each to maintain the same total peptide concentration. To determine the presence of RET, spectra of each peptide were subtracted from the spectrum of the combination multiplied a factor of 2. Reduction of the donor (Phe) but not acceptor (Tyr) fluorescence in the combination indicated RET effect. For light scattering, incident light at 550 nm was used to avoid generation of fluorescence signal and scattered light intensity at 90 degrees was measured between 530 and 570 nm. The signal intensity was averaged between 550 and 555 nm and plotted as a function of time.
The degree of inhibition of fibrillization of Aβ1–42 by peptide fragments was evaluated based on ThT fluorescence data. Total saturating ThT fluorescence between 32 and 48 h in the presence of both Aβ1–42 and a given fragment was considered as a sum of fluorescence due to each component. Then, fluorescence owing to Aβ1–42 fibrils in the presence of a fragment, F*, was determined by subtracting the signal measured for the fragment alone from total fluorescence. Fluorescence signal measured for pure fragments was corrected for concentration before subtraction, i.e. multiplied by 0.7, assuming a linear relationship between fluorescence and peptide concentration. Inhibition percentage of Aβ1–42 fibrillogenesis was calculated as IP = (1 − F*/F0) × 100%, where F0 is ThT fluorescence of Aβ1–42 in the absence of fragments.
FTIR spectroscopy
HFIP solution of the peptide was placed in small glass vials and the solvent was removed by desiccation. D2O-baseb buffer containing 25 mM NaCl and 25 mM Na,K-phosphate, adjusted to pD 7.2, was added to the dry peptide to achieve 50 μM concentration for the full-length peptide Aβ1–42 and 100 μM for the fragments. Peptide samples were continuously stirred by a small (3 mm in diameter) magnetic stir bar for > 100 h. At defined time points, a 70-μL peptide sample was placed between two CaF2 disks (32 mm in diameter, 3 mm thick), separated by a 50 μm thick Teflon spacer, and FTIR spectra were measured at 2 cm−1 nominal resolution, using a Vector-22 FTIR spectrometer (Bruker Optics, Billerica, MA, USA) equipped with a liquid nitrogen-cooled Hg-Cd-Te detector. Transmittance spectra of the blank buffer were measured and used as reference to calculate the absorbance spectra. For measurements of FTIR spectra of dry peptide sample, HFIP solution of the peptide was spread on a CaF2 disk, dried for 15 min in a desiccator, followed by measurement of the transmittance spectrum. A transmittance spectrum of the clean CaF2 disk was used as reference to calculate the absorbance spectrum. The instrument was constantly purged with dry air to avoid contribution of H2O vibrational/rotational features in the amide I region. Absorbance spectra of atmospheric H2O were measured separately and subtracted from spectra of peptides when such contribution was detected.
References
Selkoe, D. J. & Hardy, J. The amyloid hypothesis of Alzheimer’s disease at 25 years. EMBO Mol. Med. 8, 595–608 (2016).
Nortley, R. et al. Amyloid β oligomers constrict human capillaries in Alzheimer’s disease via signaling to pericytes. Science 365, 250 (2019).
Walsh, D. M. & Selkoe, D. J. Amyloid b-protein and beyond: the path forward in Alzheimer’s disease. Curr. Opin. Neurobiol. 61, 116–124 (2020).
Hardy, J. A. & Higgins, G. A. Alzheimer’s disease: The amyloid cascade hypothesis. Science 256, 184–185 (1992).
Goldblatt, G. et al. Unmodified and pyroglutamylated amyloid β peptides form hypertoxic hetero-oligomers of unique secondary structure. FEBS J. 284, 1355–1369 (2017).
Schilling, S., Rahfeld, J. U., Lues, I. & Lemere, C. A. Passive Aβ immunotherapy: Current achievements and future perspectives. Molecules 23, 1068 (2018).
Zott, B. et al. A vicious cycle of β amyloid-dependent neuronal hyperactivation. Science 365, 559–565 (2019).
Quintana, D. D. et al. Amyloid-β causes mitochondrial dysfunction via a Ca2+-driven upregulation of oxidative phosphorylation and superoxide production in cerebrovascular endothelial cells. J. Alzheimers Dis. 75, 119–138 (2020).
O’Brien, R. J. & Wong, P. C. Amyloid precursor protein processing and Alzheimer’s disease. Annu. Rev. Neurosci. 34, 185–204 (2011).
Agostinho, P., Pliássova, A., Oliveira, C. R. & Cunha, R. A. Localization and trafficking of amyloid-β protein precursor and secretases: Impact on Alzheimer’s disease. J. Alzheimers Dis. 45, 329–347 (2015).
Lee, M. H. et al. Somatic APP gene recombination in Alzheimer’s disease and normal neurons. Nature 563, 639–645 (2018).
Kubo, T., Nishimura, S., Kumagae, Y. & Kaneko, I. In vivo conversion of racemized β-amyloid ([D-Ser26]Aβ1-40) to truncated and toxic fragments ([D-Ser26]Aβ25-35/40) and fragment presence in the brains of Alzheimer’s patients. J. Neurosci. Res. 70, 474–483 (2002).
Antonios, G. et al. N-truncated Abeta starting with position four: Early intraneuronal accumulation and rescue of toxicity using NT4X-167, a novel monoclonal antibody. Acta Neuropathol. Commun. 1, 56 (2013).
Peters, C., Bascuñán, D., Opazo, C. & Aguayo, L. G. Differential membrane toxicity of amyloid-β fragments by pore forming mechanisms. J. Alzheimers Dis. 51, 689–699 (2016).
Scheidt, H. A. et al. Pyroglutamate-modified amyloid β(11–40) fibrils are more toxic than wildtype fibrils but structurally very similar. Chemistry 23, 15834–15838 (2017).
Dunys, J., Valverde, A. & Checler, F. Are N- and C-terminally truncated Aβ species key pathological triggers in Alzheimer’s disease?. J. Biol. Chem. 293, 15419–15428 (2018).
Hao, X., Zheng, J., Sun, Y. & Dong, X. Seeding and cross-seeding aggregations of Aβ40 and its N-terminal-truncated peptide Aβ11–40. Langmuir 35, 2821–2831 (2019).
Kirkitadze, M. D., Condron, M. M. & Teplow, D. B. Identification and characterization of key kinetic intermediates in amyloid beta-protein fibrillogenesis. J. Mol. Biol. 312, 1103–1119 (2001).
Xu, Y. et al. Conformational transition of amyloid beta-peptide. Proc. Natl. Acad. Sci. U.S.A. 102, 5403–5407 (2005).
Ono, K., Condron, M. M. & Teplow, D. B. Structure-neurotoxicity relationships of Amyloid β-protein oligomers. Proc. Natl. Acad. Sci. USA 106, 14745–14750 (2009).
Vandersteen A, Masman MF, De Baets G, Jonckheere W, van der Werf K, Marrink SJ, Rozenski J, Benilova I, De Strooper B, Subramaniam V. et al. Molecular plasticity regulates oligomerization and cytotoxicity of the multipeptide-length amyloid-β peptide pool. J. Biol. Chem. 287, 36732–36743 (2012).
Ma, B. Y. & Nussinov, R. Stabilities and conformations of Alzheimer’s beta-amyloid peptide oligomers (A beta(16–22′) A beta(16–35′) and A beta(10–35)): Sequence effects. Proc. Natl. Acad. Sci. USA 99, 14126–14131 (2002).
Urbanc, B. et al. Dokholyan NV molecular dynamics simulation of amyloid beta dimer formation. Biophys. J. 87, 2310–2321 (2004).
Paravastu, A. K., Leapman, R. D., Yau, W. M. & Tycko, R. Molecular structural basis for polymorphism in Alzheimer’s beta-amyloid fibrils. Proc. Natl. Acad. Sci. USA 105, 18349–18354 (2008).
Vivekanandan, S., Brender, J. R., Lee, S. Y. & Ramamoorthy, A. A. Partially folded structure of amyloid-beta(1–40) in an aqueous environment. Biochem. Biophys. Res. Commun. 411, 312–316 (2011).
Ball, K. A., Phillips, A. H., Wemmer, D. E. & Head-Gordon, T. Differences in β-strand populations of monomeric Aβ40 and Aβ42. Biophys. J. 104, 2714–2724 (2013).
Lu, J. X. et al. Molecular structure of β-amyloid fibrils in Alzheimer’s disease brain tissue. Cell 154, 1257–1268 (2013).
Tycko, R. Physical and structural basis for polymorphism in amyloid fibrils. Protein Sci. 23, 1528–1539 (2014).
Qiang, W., Yau, W. M., Lu, J. X., Collinge, J. & Tycko, R. Structural variation in amyloid-β fibrils from Alzheimer’s disease clinical subtypes. Nature 541, 217–221 (2017).
Tay, W. M., Huang, D. & Rosenberry, T. L. Paravastu AK The Alzheimer’s amyloid-b1-42 peptide forms offpathway oligomers and fibrils that are distinguished structurally by intermolecular organization. J. Mol. Biol. 425, 2494–2508 (2013).
Tycko, R. Solid state NMR studies of amyloid fibril structure. Annu. Rev. Phys. Chem. 62, 279–299 (2011).
Xiao, Y. et al. Aβ(1–42) fibril structure illuminates self-recognition and replication of amyloid in Alzheimer’s disease. Nat. Struct. Mol. Biol. 22, 499–505 (2015).
Wälti, M. A. et al. Atomic-resolution structure of a disease-relevant Aβ(1–42) amyloid fibril. Proc. Natl. Acad. Sci. USA 113, E4976–E4984 (2016).
Colvin, M. T. et al. Atomic resolution structure of monomorphic Aβ42 amyloid fibrils. J. Am. Chem. Soc. 138, 9663–9674 (2016).
Li, H. et al. Mechanistic investigation of the inhibition of Abeta42 assembly and neurotoxicity by Abeta42 C-terminal fragments. Biochemistry 49, 6358–3664 (2010).
Li, H., Monien, B. H., Fradinger, E. A., Urbanc, B. & Bitan, G. Biophysical characterization of Abeta42 C-terminal fragments: Inhibitors of Abeta42 neurotoxicity. Biochemistry 49, 1259–1267 (2010).
Wang, W. et al. Toxic amyloid-β oligomers induced self-replication in astrocytes triggering neuronal injury. EBioMedicine 42, 174–187 (2019).
Katzmarski, N. et al. Aβ oligomers trigger and accelerate Aβ seeding. Brain Pathol. 30, 36–45 (2020).
Kirkitadze, M. D., Bitan, G. & Teplow, D. B. Paradigm shifts in Alzheimer’s disease and other neurodegenerative disorders: The emerging role of oligomeric assemblies. J. Neurosci. Res. 69, 567–577 (2002).
Bernstein, S. L. et al. Amyloid-β protein oligomerization and the importance of tetramers and dodecamers in the aetiology of Alzheimer’s disease. Nat. Chem. 1, 326–331 (2009).
Benilova, I., Karran, E. & De Strooper, B. The toxic A β oligomer and Alzheimer’s disease: An emperor in need of clothes. Nat. Neurosci. 15, 349–357 (2012).
Lesné, S. E. et al. Brain amyloid-β oligomers in ageing and Alzheimer’s disease. Brain 136, 1383–1398 (2013).
Yasumoto, T. et al. High molecular weight amyloid β1–42 oligomers induce neurotoxicity via plasma membrane damage. FASEB J. 33, 9220–9234 (2019).
Bitan, G. et al. Amyloid β-protein (Aβ) assembly: Aβ40 and Aβ42 oligomerize through distinct pathways. Proc. Natl. Acad. Sci. USA 100, 330–335 (2003).
Urbanc, B., Betnel, M., Cruz, L., Bitan, G. & Teplow, D. B. Elucidation of amyloid β-protein oligomerization mechanisms: Discrete molecular dynamics study. J. Am. Chem. Soc. 132, 4266–4280 (2010).
Cukalevski, R. et al. The Aβ40 and Aβ42 peptides self-assemble into separate homomolecular fibrils in binary mixtures but cross-react during primary nucleation. Chem. Sci. 6, 4215–4233 (2015).
Wang, Y., Shi, Y. & Wei, H. Calcium dysregulation in Alzheimer’s disease: A target for new drug development. J. Alzheimers Dis. Parkinsonism. 7, 374 (2017).
Baronio, C. M., Baldassarre, M. & Barth, A. Insight into the internal structure of amyloid-β oligomers by isotope-edited Fourier transform infrared spectroscopy. Phys. Chem. Chem. Phys. 21, 8587–8597 (2019).
He, W. & Barrow, C. J. The A beta 3-pyroglutamyl and 11-pyroglutamyl peptides found in senile plaque have greater beta-sheet forming and aggregation propensities in vitro than full-length A beta. Biochemistry 38, 10871–10877 (1999).
Dammers, C. et al. Structural analysis and aggregation propensity of pyroglutamate Aβ(3–40) in aqueous trifluoroethanol. PLoS ONE 10, e0143647 (2015).
Movsesyan, N. et al. Reducing AD-like pathology in 3xTg-AD mouse model by DNA epitope vaccine—A novel immunotherapeutic strategy. PLoS ONE 3, e2124 (2008).
Ghochikyan, A. et al. Refinement of a DNA based Alzheimer’s disease epitope vaccine in rabbits. Hum. Vaccin. Immunother. 9, 1002–1010 (2013).
Davtyan, H. et al. MultiTEP platform-based AD epitope vaccine activates broad repertoire of T helper cells in non-human primates. Alzheimers Dement. 10, 271–283 (2014).
Sigurdsson, E. M., Permanne, B., Soto, C., Wisniewski, T. & Frangione, B. In vivo reversal of amyloid-beta lesions in rat brain. J. Neuropathol. Exp. Neurol. 59, 11–17 (2000).
Gordon, D. J., Sciarretta, K. L. & Meredith, S. C. Inhibition of beta-amyloid(40) fibrillogenesis and disassembly of beta-amyloid(40) fibrils by short beta-amyloid congeners containing N-methyl amino acids at alternate residues. Biochemistry 40, 8237–8245 (2001).
Gordon, D. J. & Meredith, S. C. Probing the role of backbone hydrogen bonding in beta-amyloid fibrils with inhibitor peptides containing ester bonds at alternate positions. Biochemistry 42, 475–485 (2003).
Sciarretta, K. L., Gordon, D. J. & Meredith, S. C. Peptide-based inhibitors of amyloid assembly. Methods Enzymol. 413, 273–312 (2006).
Sepulveda, F. J., Parodi, J., Peoples, R. W., Opazo, C. & Aguayo, L. G. Synaptotoxicity of Alzheimer beta amyloid can be explained by its membrane perforating property. PLoS ONE 5, e11820 (2010).
Giordano, C. et al. Inhibition Of Amyloid Peptide Fragment Aβ25–35 fibrillogenesis and toxicity by N-terminal β-amino acid-containing esapeptides: Is taurine moiety essential for in vivo effects?. Chem. Biol. Drug. Des. 79, 30–37 (2012).
Cheng, P. N., Liu, C., Zhao, M., Eisenberg, D. & Nowick, J. S. Amyloid β-sheet mimics that antagonize amyloid aggregation and reduce amyloid toxicity. Nat. Chem. 4, 927–933 (2012).
Peters, C. et al. Inhibition of amyloid beta-induced synaptotoxicity by a pentapeptide derived from the glycine zipper region of the neurotoxic peptide. Neurobiol. Aging 34, 2805–2814 (2013).
Bharadwaj, P. et al. Modulation of amyloid-beta 1–42 structure and toxicity by proline-rich whey peptides. Food Funct. 4, 92–103 (2013).
Barucker, C. et al. Aβ42-oligomer Interacting Peptide (AIP) neutralizes toxic amyloid-β42 species and protects synaptic structure and function. Sci. Rep. 5, 15410 (2015).
Rajasekhar, K., Madhu, C. & Govindaraju, T. A natural tripeptide-based inhibitor of multifaceted amyloid β toxicity. ACS Chem. Neurosci. 7, 1300–1310 (2016).
Shea, D. et al. α-Sheet secondary structure in amyloid β-peptide drives aggregation and toxicity in Alzheimer’s disease. Proc. Natl. Acad. Sci. USA 116, 8895–8900 (2019).
Ghosh, A. et al. Inhibition and degradation of amyloid beta (Aβ40) fibrillation by designed small peptide: A combined spectroscopy, microscopy, and cell toxicity study. ACS Chem. Neurosci. 8, 718–722 (2017).
Aoraha, E., Candreva, J. & Kim, J. R. Engineering of a peptide probe for β-amyloid aggregates. Mol. Biosyst. 11, 2281–2289 (2015).
Röhrig, U. F., Laio, A., Tantalo, N., Parrinello, M. & Petronzio, R. Stability and structure of oligomers of the Alzheimer peptide Abeta16-22: From the dimer to the 32-mer. Biophys. J. 91, 3217–3229 (2006).
Petty, S. A. & Decatur, S. M. Experimental evidence for the reorganization of b-strands within aggregates of the Aβ(16–22) peptide. J. Am. Chem. Soc. 127, 13488–13489 (2005).
Fradinger, E. A. et al. C-terminal peptides coassemble into Abeta42 oligomers and protect neurons against Abeta42-induced neurotoxicity. Proc. Natl. Acad. Sci. USA 105, 14175–14180 (2008).
Li, H. et al. C-terminal tetrapeptides inhibit Aβ42-induced neurotoxicity primarily through specific interaction at the N-terminus of Aβ42. J. Med. Chem. 54, 8451–8460 (2011).
Urbanc, B. et al. Structural basis for Aβ1–42 toxicity inhibition by Aβ C-terminal fragments: Discrete molecular dynamics study. J. Mol. Biol. 410, 316–328 (2011).
Gessel, M. M. et al. Aβ(39–42) modulates Aβ oligomerization but not fibril formation. Biochemistry 51, 108–117 (2012).
Stine, W. B. Jr., Dahlgren, K. N., Krafft, G. A. & LaDu, M. J. In vitro characterization of conditions for amyloid-beta peptide oligomerization and fibrillogenesis. J. Biol. Chem. 278, 11612–11622 (2003).
Barrett, P. J. et al. The amyloid precursor protein has a flexible transmembrane domain and binds cholesterol. Science 336, 1168–1171 (2012).
Tatulian, S. A. FTIR Analysis of proteins and protein-membrane interactions. Methods Mol. Biol. 2003, 281–325 (2019).
Krimm, S. & Bandekar, J. Vibrational spectroscopy and conformation of peptides, polypeptides, and proteins. Adv. Protein Chem. 38, 181–364 (1986).
Jackson, M. & Mantsch, H. H. The use and misuse of FTIR spectroscopy in the determination of protein structure. Crit. Rev. Biochem. Mol. Biol. 30, 95–120 (1995).
Perczel A., Hollósi M. Turns. In Circular Dichroism and the Conformational Analysis of Biomolecules 285–380; Fasman, G. D., Ed. (Plenum Press, 1996).
Sreerama, N., Venyaminov, S. Y. & Woody, R. W. Estimation of the number of alpha-helical and beta-strand segments in proteins using circular dichroism spectroscopy. Protein Sci. 8, 370–380 (1999).
Venyaminov SY, Yang JT. Determination of protein secondary structure. In Circular Dichroism and the Conformational Analysis of Biomolecules 69–107; Fasman, GD., Ed.; (Plemun Press, 1996).
Woody RW. Theory of circular dichroism of proteins. In Circular Dichroism and the Conformational Analysis of Biomolecules 25–67; Fasman, GD., Ed.; (Plenum Press, 1996).
Sreerama N, Woody RW. Circular dichroism of peptides and proteins. In Circular Dichroism: Principles and Applications 601−620; Berova, N., Nakanishi, K., Woody, R. W. Eds. John Wiley & Sons (2000).
Hamley, I. W. et al. Influence of the solvent on the self-assembly of a modified amyloid beta peptide fragment. II. NMR and computer simulation investigation. J. Phys. Chem. B 114, 940–951 (2010).
Wang, J. et al. Tuning self-assembled morphology of the Aβ(16–22) peptide by substitution of phenylalanine residues. Colloids Surf. B Biointerfaces 147, 116–123 (2016).
Petkova, A. T. et al. Self-propagating, molecular-level polymorphism in Alzheimer’s beta-amyloid fibrils. Science 307, 262–265 (2005).
Kodali, R., Williams, A. D., Chemuru, S. & Wetzel, R. Aβ(1–40) forms five distinct amyloid structures whose β-sheet contents and fibril stabilities are correlated. J. Mol. Biol. 40, 503–517 (2010).
Qiang, W., Kelley, K. & Tycko, R. Polymorph-specific kinetics and thermodynamics of β-amyloid fibril growth. J. Am. Chem. Soc. 135, 6860–6871 (2013).
Searcy, D. G., Montenay-Garestier, T. & Hélène, C. Phenylalanine-to-tyrosine singlet energy transfer in the archaebacterial histone-like protein HTa. Biochemistry 28, 9058–9065 (1989).
Lakowicz, J. R. Principles of Fluorescence Spectroscopy 2nd edn. (Kluwer Academic/Plenum Publishers, 1999).
Sawaya, M. R. et al. Atomic structures of amyloid cross-beta spines reveal varied steric zippers. Nature 447, 453–457 (2007).
Colletier, J. P. et al. Molecular basis for amyloid-beta polymorphism. Proc. Natl. Acad. Sci. USA 108, 16938–16943 (2011).
Vugmeyster, L. et al. Effect of post-translational modifications and mutations on amyloid-β fibrils dynamics at N terminus. Biophys. J. 117, 1524–1535 (2019).
Gardberg, A. S. et al. Molecular basis for passive immunotherapy of Alzheimer’s disease. Proc. Natl. Acad. Sci. USA 104, 15659–15664 (2007).
Söldner, C. A., Sticht, H. & Horn, A. H. C. Role of the N-terminus for the stability of an amyloid-β fibril with three-fold symmetry. PLoS ONE 12, e0186347 (2017).
An, Y., Bloom, J. W. & Wheeler, S. E. Quantifying the π-stacking interactions in nitroarene binding sites of proteins. J. Phys. Chem. B 119, 14441–14450 (2015).
Du, M. et al. Noncovalent self-assembly of protein crystals with tunable structures. Nano Lett. 21, 1749–1757 (2021).
Hansen, A. L. & Kay, L. E. Measurement of histidine pKa values and tautomer populations in invisible protein states. Proc. Natl. Acad. Sci. USA 111, E1705–E1712 (2014).
Hilbich, C., Kisters-Woike, B., Reed, J., Masters, C. L. & Beyreuther, K. Human and rodent sequence analogs of Alzheimer’s amyloid beta A4 share similar properties and can be solubilized in buffers of pH 7.4. Eur. J. Biochem. 201, 61–69 (1991).
Balbach, J. J. et al. Amyloid fibril formation by A beta 16–22, a seven-residue fragment of the Alzheimr’s beta-amyloid peptide, and structural characterization by solid state NMR. Biochemistry 39, 13748–13759 (2000).
Santini, S., Wei, G., Mousseau, N. & Derreumaux, P. Pathway complexity of Alzheimer’s beta-amyloid Abeta16–22 peptide assembly. Structure 12, 245–255 (2004).
Tao, K. et al. Self-assembly of short aβ(16–22) peptides: Effect of terminal capping and the role of electrostatic interaction. Langmuir 27, 2723–2730 (2011).
Klimov, D. K. & Thirumalai, D. Dissecting the assembly of Abeta16–22 amyloid peptides into antiparallel beta sheets. Structure 11, 295–307 (2003).
Lazo, N. D., Grant, M. A., Condron, M. C., Rigby, A. C. & Teplow, D. B. On the nucleation of amyloid beta-protein monomer folding. Protein Sci. 14, 1581–1596 (2005).
Roychaudhuri, R., Yang, M., Condron, M. M. & Teplow, D. B. Structural dynamics of the amyloid β-protein monomer folding nucleus. Biochemistry 51, 3957–3959 (2012).
Hoyer, W., Grönwall, C., Jonsson, A., Ståhl, S. & Härd, T. Stabilization of a beta-hairpin in monomeric Alzheimer’s amyloid-beta peptide inhibits amyloid formation. Proc. Natl. Acad. Sci. USA 105, 5099–5104 (2008).
Terzi, E., Hölzemann, G. & Seelig, J. Reversible random coil-beta-sheet transition of the Alzheimer beta-amyloid fragment (25–35). Biochemistry 33, 1345–1350 (1994).
Kandel, N., Zheng, T., Huo, Q. & Tatulian, S. A. membrane binding and pore formation by a cytotoxic fragment of amyloid β peptide. J. Phys. Chem. B 121, 10293–10305 (2017).
Kohno, T., Kobayashi, K., Maeda, T., Sato, K. & Takashima, A. Three-dimensional structures of the amyloid b-peptide (25–35) in membrane-mimicking environment. Biochemistry 35, 16094–16104 (1996).
D’Ursi, A. M. et al. Solution structure of amyloid beta-peptide (25–35) in different media. J. Med. Chem. 47, 4231–4238 (2004).
Do, T. D. et al. Amyloid β-protein C-terminal fragments: Formation of cylindrins and β-barrels. J. Am. Chem. Soc. 138, 549–557 (2016).
Acknowledgements
Financial support from Florida Department of Health, Ed and Ethel Moore Alzheimer’s Disease Research Program (Grants 7AZ27 and 21A06), is gratefully acknowledged.
Author information
Authors and Affiliations
Contributions
F.A. and N.K. conducted all experiments and data analysis. S.A.T. designed the research, conducted data analysis, prepared the final figures, and wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Abedin, F., Kandel, N. & Tatulian, S.A. Effects of Aβ-derived peptide fragments on fibrillogenesis of Aβ. Sci Rep 11, 19262 (2021). https://doi.org/10.1038/s41598-021-98644-y
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-021-98644-y
- Springer Nature Limited
This article is cited by
-
FoxG1 as a Potential Therapeutic Target for Alzheimer’s Disease: Modulating NLRP3 Inflammasome via AMPK/mTOR Autophagy Pathway
Cellular and Molecular Neurobiology (2024)
-
Impact of seed amplification assay and surface-enhanced Raman spectroscopy combined approach on the clinical diagnosis of Alzheimer’s disease
Translational Neurodegeneration (2023)