Abstract
Camellia is a genus of flowering plants in the family Theaceae, and several species in this genus have economic importance. Although a great deal of molecular makers has been developed for molecular assisted breeding in genus Camellia in the past decade, the number of simple sequence repeats (SSRs) publicly available for plants in this genus is insufficient. In this study, a total of 28,854 potential SSRs were identified with a frequency of 4.63 kb. A total of 172 primer pairs were synthesized and preliminarily screened in 10 C. japonica accessions, and of these primer pairs, 111 were found to be polymorphic. Fifty-one polymorphic SSR markers were randomly selected to perform further analysis of the genetic relationships of 89 accessions across the genus Camellia. Cluster analysis revealed major clusters corresponding to those based on taxonomic classification and geographic origin. Furthermore, all the genotypes of C. japonica separated and consistently grouped well in the genetic structure analysis. The results of the present study provide high-quality SSR resources for molecular genetic breeding studies in camellia plants.
Similar content being viewed by others
Introduction
Camellia is a genus of flowering plants in the family Theaceae, which is widely distributed in Southeast Asia, from the Himalayas to Japan and from southern China (Guangxi and Yunnan) to Java and Sumatra1. Several species in the genus Camellia have economic importance. Species such as C. japonica, C. reticulata, C. saluenensis and C. sasanqua, are well-known as camellias with attractive flowers. The young leaves of the important economic species C. sinensis are used to produce tea. A few species such as C. oleifera and C. semiserrata are used to produce high-quality edible and pharmaceutical seed oil1,2.
Camellia is the largest genus in the family Theaceae. This genus is believed to comprise more than 300 species3, indicating genetic instability and the high outbreeding nature of the genus. Ornamental camellias (chahua in Chinese) have been grown in China for 2,000 years4. The common camellia was introduced to Japan over 1,000 years ago4. Ornamental camellias were brought to Europe and the Americas in the late 1870s5 and are now popular flowering and landscaping shrubs in many regions worldwide1. Currently, there are more than 3,000 cultivated varieties of ornamental camellia worldwide1. Due to natural and artificial interspecific hybridization Camellia, this genus exhibits important taxonomic and systematic conflicts6.
Traditionally, the breeding of camellia plants was based on hybridization among species and cultivars and the phenotypic selection of novel or improved offspring7,8. Although traditional breeding still plays a crucial role in the quality improvement of Camellia plants, it is limited by the long selection term and tremendous resources required in the breeding of new varieties. Marker assisted selection (MAS) is potential to accelerate plant breeding and avoids the problems associated with traditional plant breeding by improving the selection criteria from phenotypes to genes, thus saving time and resources9,10. Moreover, molecular markers are not restrained to environmental regulations, and are not affected by the conditions that the plants are grown and detectable at all stages of plant growth11.
Simple sequence repeats (SSRs) are genomic fragments that composed of tandemly repeated units of 1–6 nucleotide sequence motifs flanked by unique sequences12. SSR markers are widely used in plant genetic improvement and MAS breeding. Due to their high polymorphism, codominant inheritance, and wide distribution traits throughout the genome12,13, SSRs have been extensively used in genetic linkage map construction, genetic identification, genetic diversity analyses and fingerprinting construction14,15,16,17.
A great deal of effort has been made to develop SSR markers in the genus Camellia in the past decade. Based on 454-sequencing data, 36 polymorphic expressed sequence tag (EST)-SSR markers have been developed in tea plant (C. sinensis)18. By using high-throughput Illumina RNA sequencing data, 431 polymorphic SSR markers were developed, and a consensus SSR-based linkage map was constructed that covered 1,156.9 cM with 237 SSR markers distributed in 15 linkage groups in tea plant19. Based on transcriptome data, 450 polymorphic SSR markers were developed, and 406 were successfully added to genetic linkage maps of tea plant20. Polymorphic SSR markers were developed via genome sequence analysis of tea plant and then used for genetic studies and fingerprinting construction13,21. Several efforts have also been made to develop SSR markers in ornamental camellias and oil camellias22,23,24,25,26,27.
However, the number of SSRs publicly available for plants in the genus Camellia is insufficient for some applications, such as the construction of high-resolution linkage maps, genome comparative mapping, genetic studies, and increasing marker density in specific map regions. Therefore, more efforts are still needed to develop SSR markers for further progress in camellia genetic and genomic studies.
In our previous study, we obtained approximately 1,006 million RNA-Seq reads by deep sequencing of the ornamental camellia C. japonica leaf transcriptome, using the Illumina sequencing platform28. These sequence data provide a good resource for the development of genic SSR markers. In the present study, we developed a set of novel genic SSR markers based on de novo transcriptome sequencing of C. japonica, determined transferable and polymorphic SSR markers for other Camellia species, and revealed the genetic relationships and genetic structure of Camellia germplasms. The results of this study will provide essential information for further studies in camellia plants such as taxonomic studies, diversity analysis, MAS and SSR-based genetic linkage mapping.
Results
Characteristics of genic SSRs in the C. japonica leaf transcriptome
SSRs were highly abundant in the assembled C. japonica leaf transcriptome. In total, 28,854 potential SSRs with a minimum of five repetitions for all motifs were identified from 24,368 contigs, representing 11.74% of the total 207,592 unigenes28 generated by Illumina sequencing. The frequency of occurrence of SSR loci was one in every 4.63 kb (221.17 SSRs/Mb) of the unigene sequence. The length of SSRs ranged from 12 to 279 bp, with an average of 20.54 bp.
Incidences of different repeat types and frequencies for each motif were evaluated based on the repeat unit number (Table 1). SSRs existed primarily as dinucleotide repeats and trinucleotide repeats, accounting for 97.59% of all SSRs. Dinucleotide repeats (74.56%) were the most abundant repeat unit, followed by tr- (23.08%), tetra-(2.04%), hexa-(0.19%) and pentanucleotides (0.11%), with the repeat unit number from 5 to 18. Most (99.4%) of the motifs had 5–10 repeats, while motifs with more than 10 repeat were rare (0.59%). Among the identified SSRs, AG/CT was the most common type of all dinucleotide repeat motifs (62.19%) (Supplementary Fig. 1a). The predominant trinucleotide repeat motifs were AAG/CTT and AAT/ATT, which accounted for 24.94% and 19.56% of the SSRs, respectively (Supplementary Fig. 1b). For tetranucleotide repeats, the most frequent motif was AAAT/ATTT (31.30%), followed by AAAC/GTTT (13.37%) and AAAG/CTTT (11.68%) (Supplementary Fig. 1c). These results may reflect the AG/CT-rich nature of the C. japonica transcriptome.
Functional annotation of SSR-containing unigenes
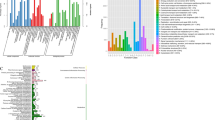
To explore the potential function of these SSR-containing unigenes we used Gene Ontology (GO) assignments to classify the predicted C. japonica genes. A total of 24,214 SSR-containing unigenes were assigned to three major functional categories (Biological Process, Molecular Function and Cellular Component) (Fig. 1a). In the Biological Processes category, ‘biosynthetic process’, ‘cellular protein modification process’, ‘nucleobase-containing compound metabolic process’, and ‘cellular process’ were the top four GO terms. In the Molecular Function category, the unigenes were predominantly assigned to the ‘nucleotide binding’, ‘hydrolase activity’ and ‘binding’ groups. In the Cellular Component category, the unigenes were frequently assigned to ‘membrane’, ‘nucleus’, ‘cytoplasm’ and ‘plastid’.
To identify the biological pathways associated with the annotated SSR-containing unigenes, we annotated the unigenes to the reference pathways in the Kyoto Encyclopedia of Genes and Genomes (KEGG) using KeggArray software, and 4,325 SSR-containing unigenes were assigned to five specific pathways, including ‘Metabolism’, ‘Genetic Information Processing’, ‘Environmental Information Processing’, ‘Cellular Processes’, and ‘Organism Systems’ (Fig. 1b). Among these pathways, the ‘signal transduction’ cluster represented the largest group, followed by ‘carbohydrate metabolism’ and ‘cell growth and death’.
A total of 1,015 putative transcription factors (TFs) were identified in the annotated SSR-containing unigenes. These TFs were classified into 51 common families based on the classification of their Arabidopsis homologues (Supplementary Table 1). Among these families, the WRKY family was the largest group (161, 16%), followed by the NAC family (149, 15%), the bHLH family (71, 7%), the C2H2 family (64, 6%) and ethylene-responsive TF (ERF) (52, 5%) (Fig. 1c).
Development and validation of genic SSR markers
A total of 24,368 SSR-containing unigenes were employed for primer design, from which 12,194 (50.04%) unigenes could be successfully used for SSR primer development. A total of 13,323 SSR primers were developed. After screening by e-PCR, 8,442 SSR primers were inferred to have unique amplification sites in the C. japonica leaf transcriptome. Of the 8,442 primer pairs, 172 were selected for primer synthesis. The 172 markers were tested for amplification using a panel of 10 C. japonica accessions (Fig. 2a), the details which are available in Supplementary Table 2. Of these SSR markers, 30 (17.44%) amplified smear or nonspecific products, and 3 (1.74%) generated no products in any of the C. japonica accessions. A total of 139 (80.81%) SSR markers yielded amplification products in the 10 accessions, of which 111 (64.53%) exhibited polymorphism. Among these polymorphic markers, trinucleotide repeats were the most abundant (66, 59.46%), followed by di- (30, 27.03%), tetra- (14, 12.61%) and hexanucleotide (1, 0.9%), whereas no polymorphic markers were identified in pentanucleotides (Table 2).
PAGE results of marker detection and population analysis. (a) Marker detection in 10 C. japonica accessions. (b) Population analysis in 89 accessions across genus Camellia. CjSSR094, CjSSR021, CjSSR128 and CjSSR166 were chosen from SSR markers with di-, tri-, tetra-, and hexanucleotide repeat, respectively.
A total of 495 alleles were identified across these 111 polymorphic genic SSR loci, and the number of alleles ranged from 1 to 12 with an average of 4.46 alleles per locus. To evaluate and characterize these polymorphisms so that they can potentially be used for assessing molecular diversity or genetic structure analysis, the polymorphism information content (PIC) values of these polymorphic primers were calculated. The PIC values of the polymorphic markers ranged from 0.15 to 0.86, with a mean value of 0.59. The PIC values of the polymorphic markers with di-, tri-, tetra-, and hexanucleotide repeat were 0.63, 0.57, 0.58 and 0.37, respectively (Table 2). The sequences of these informative newly developed genic SSR primers and other major information are shown in Supplementary Table 3.
In total, 51 genic SSR markers, which demonstrated polymorphic in these 10 C. japonica accessions, were randomly selected and further used to assess their transferability in 12 other accessions across 8 species in the genus Camellia, including C. sasanqua, C. chuongtsoensis, C. rosthorniana, C. nitidissima, C. oleifera, C. reticulata, C. sasanqua and C. sinensis. In contrast to five primer pairs, CjSSR014 and CjSSR021, that could not amplify any fragments in C. sinensis, CjSSR048 and CjSSR101, that could not amplify fragments in two C. sasanqua accessions, CjSSR098, that could not amplify fragments in C. azalea cultivar 'Nanyue Hongxia', the other forty-six primer pairs successfully amplified PCR products in all the tested accessions. Thus, the transferability rate across 8 species ranged from 94.12% to 100%, with an average transferability rate of 98.69% (Supplementary Table 4).
Cluster analyses of natural accessions in the genus Camellia
To verify the applicability of the newly developed genic SSR markers, a total of 89 accessions across the genus Camellia (Supplementary Table 5) were genotyped by the above 51 genic SSR primers (Fig. 2b). In total, 503 alleles were obtained among these natural population, with 9.9 alleles per SSR on average, which was much higher than the previous studies13,15,23,27, implying that these 51 SSRs harbored abundant variation loci of Camellia. Therefore, these 51 SSRs as core molecular markers had typical and highly representative alleles. Based on the genetic distance results, several clusters were visible in the neighbor-joining (NJ) dendrogram. To simplify the description of the results, we distinguished five major clusters, I to V, by separating the longest branch into five branches. Cluster I comprised the accessions from C. japonica and was further divided into five subclusters: Ia to Ie. The accessions from C. saluenensis hybrids were grouped in cluster III. Cluster IV contained two accessions from C. sasanqua. The accessions from C. reticulata and its hybrids were closely clustered in cluster V. Accessions collected from other species in the genus Camellia were located in cluster II (Fig. 3). This result showed groupings of genotypes that agreed with groupings based on taxonomic classification and geographic origin. Notably, several accessions known to have similar genetic backgrounds were closely clustered and clearly distinguished with each other in the dendrogram, such as ‘Coquettii’ and ‘Coquettina’, ‘Yudan’ and ‘Yuanyang Fengguan’, and ‘Grand Marshal’ and ‘Grand Marshal Variegated’ (Fig. 3). The above results implied that the newly developed genic SSR markers are powerful for determination relationship of genotypes.
It is interesting that when labeling according to flower color, some accessions that were grouped in same cluster or subcluster have similar flower colors. Cluster V (85.7%), subclusters Ib (64.7%) and Ie (83.3%) majority comprised of the accessions with pink flowers. Subclusters Ic (85.7%) and Id (92.3%) mainly contained accessions with red flowers. Cluster II contained accessions with light-colored (yellow or white) flowers. Interestingly, the accessions with orchid flowers were all grouped in cluster III (Fig. 4).
Principal component analysis (PCA) analysis of natural accessions in the genus Camellia
Principal component analysis (PCA) using the first and second eigenvectors identified two major groups (Fig. 5). The eigenvalues of first and second axes were 6.17% and 4.46%, respectively. The accessions from C. japonica were noticeably distinct from the accessions that collected from C. reticulata and its hybrids, C. sasanqua, C. saluenensis hybrids, and other species in the genus Camellia.
Genetic structure analyses of selected accessions in the C. japonica
The genetic structure of the 69 accessions from C.japonica was analyzed with the Bayesian model-based clustering algorithm implemented in STRUCTURE software. The most likely number of clusters was identified by calculating delta K (ΔK), which is based on the rate of change in the log probability of data between successive K values (K = 1 to K = 10). The peak of the ΔK graph corresponds to the most likely number of populations in the data set. The highest number of ΔK was found at K = 3 (Fig. 6a,b), where all 69 accessions were divided into three main groups: the first group contained cluster Ib and Id that determined in cluster analysis, while cluster Ia and Ic together assigned to group 2, the third group contained cluster Ie that determined in cluster analysis (Fig. 6c). The results from the population structure analysis were generally consistent with the results from the cluster analysis.
Discussion
SSRs can be divided into genomic SSRs and genic SSRs, and genomic SSRs are usually developed from genomic libraries or random genomic sequences, while genic SSRs are developed from coding regions of the EST or transcriptome sequences20,24. Although the discovery of genomic SSR loci using whole genome sequences had been successfully applied in many plant species, such as peanut29, pear30, sweet potato31 and tea plant13, genic SSR development based on transcriptome sequences is still fundamental., Genic SSR markers, which are linked to the loci of agronomic phenotypes, are considered more useful for MAS, especially when polymorphic genic SSR markers are identified in breeding lines compared with genomic SSR markers32. In previous study, we obtained the high-quality assembly transcriptome sequence of C. japonica var. ‘Jiangxue’ based on high-throughput sequencing data28, which enabled us to develop SSRs by transcriptome-wide analysis. In this study, 28,854 putative SSRs were identified for approximately 11.74% of the total unigenes. The average frequency of SSRs was 1/4.63 kb, which is similar to those estimated by Ma et al. (1/3.98 kb) and Wu et al. (1/4.99 kb) in tea plants18,20 and higher than that estimated by Wen et al. (1/6.5 kb) in oil camellia C. chekiangoleosa24.
Frequency analysis of motif repeats in C. japonica revealed that dinucleotides and trinucleotides were the most abundant types of SSRs and together accounted for 97.59% of all the SSRs identified; tetra-, penta- and hexanucleotides were less abundant and together accounted for 2.35% (Table 1). This tendency of highly abundant di- and trinucleotides was in conformity to reports of several other species in the genus Camellia13,18,20. Furthermore, the repeat numbers analyze in SSR motifs showed that the distributions of all di-, tri-, tetra-, penta- and hexanucleotides were generally skewed towards fewer repeats (Table 1). Similar tendencies were also found in other species in the genus Camellia, such as tea plant13 and oil camellia23.
Within a certain type of SSR, the frequency of specific repeat motifs may differ. Previous studies on tea plant and other plant species showed that the base composition of SSR motifs is largely biased towards As and Ts13,33. Similar results were found in this study: the AG/CT and AT/TA motifs were the most abundant, while the CG/GC motif was very rare among dinucleotide repeats (Supplementary Fig. 1a). Similarly, the motifs AAG/CTT and AAAT/ATTT were the most abundant in tr- and tetranucleotide repeats, respectively, but the percentages of GC-rich repeat motifs were extremely low (Supplementary Fig. 1b,c), indicating that genic SSRs with GC-rich repeats are rare in C. japonica. Overall, the abundant genic SSR motifs offer new perspectives for the development of SSR markers in C. japonica.
The effectiveness and success of applying SSR markers are considerably dependent on the quality of the markers. In this study, of the 172 newly designed SSR primer pairs, 111 (64.53%) exhibited polymorphism, and the polymorphism ratio was much higher than those obtained in previous studies in the genus Camellia13,18,20,23,34. The high polymorphism ratio may largely due to the rigorous e-PCR screening that performed in our study. The mean PIC value of the polymorphic markers was 0.59, which was similar to those reported in previous studies on camellia plants18,23,24. The high PIC values indicate that these newly developed genic SSR markers are suitable for phylogenetic and genetic diversity analyses and linkage map construction. Moreover, the transferability rates of SSRs in this study were extremely high (from 92.6% to 100%, with an average of 99.38%, Supplementary Table 4), which were higher than those reported in other studies on camellia plants34,35. Overall, the high polymorphism ratio and high transferability of the SSR markers developed in this study may be due to the relatively conserved nature of transcriptional sequences. It is interesting that trinucleotide repeats were the most abundant motifs among polymorphic markers (Table 2), although dinucleotide repeats were the most abundant motifs in the C. japonica leaf transcriptome (Table 1), implying that the trinucleotide repeat motifs may have a high specificity compared with other repeat motifs in C. japonica. Together with the high percentage of trinucleotide repeat motif-containing polymorphic markers among synthesized markers and the high PIC values of the trinucleotide repeat motif-containing polymorphic markers (Table 2), our results in this study indicate that SSR markers developed by trinucleotide repeat motif were the most efficient with high polymorphism and specificity for genetic improvement study in genus Camellia.
Highly polymorphic and stable SSR markers are important resources for genetic relationship analysis. Genetic distance measures applied to SSR data can yield useful estimators for phylogenetic relationships among closely related populations, as well as among species, accessions or cultivars27,30. In the present study, the NJ dendrogram demonstrated that the accessions from C. japonica, C. reticulata, C. sasanqua, C. saluenensis hybrids, and other species were clearly divided into five main clusters, and several accessions known to have similar genetic backgrounds were closely clustered and clearly distinguished with each other in the dendrogram (Fig. 3), indicate the newly developed SSR markers not only can separate the various genus, but also can distinguish the minor/individual variations between accessions. Therefore, the newly developed SSR markers as the core markers had high power for Camellia genotyping. Several previous studies showed a significant positive correlation between the genetic distance and geographic distance of ornamental camellia populations26,27,36. In the present study, we found that accessions with the similar flower color were grouped together in several clusters/subclusters in the dendrogram (Fig. 4), whereas no regular clustering based on flower type were found in this study (data not shown), implying that flower type may exhibit high probability variations compared with flower color in camellia plants.
The PCA analysis divided all accessions into two main clusters: the first cluster included all C. japonica cultivars, while the second cluster contained the accessions that were collected from other species in the genus Camellia (Fig. 5). Similar results were obtained in other studies that distinguished C. japonica cultivars from other cultivars in the genus Camellia using SSR markers6,26. A recent study in C. nitidissima grouped 96 individuals into 4 subpopulations and found some overlap between subpopulations27. In the present study, all the genotypes in C. japonica separated and consistently grouped well in the Bayesian model-based genetic structure analysis, although some overlap was also found in subpopulations 1 (clusters Ib and Id) and subpopulation 2 (clusters Ia and Ic) (Fig. 6c). In our study, the selected accessions in C. japonica cover almost all flower colors, flower types and breeding approaches (Supplementary Table 5), although the selected accessions in our study do not constitute a classical sexual population, the structure analysis of C. japonica may still provide clues for better understanding the origin and evolution of C. japonica plants and help us make use of these resources.
Conclusions
In this study, we developed 28,854 genic SSR markers with a frequency of 4.63 kb from transcriptome sequences of C. japonica. A total of 13,323 SSR primers were developed, and among them, 111 were found to be polymorphic in 9 Camellia species, which is by far the largest number of SSR markers developed in a single study in C. japonica to date. The obtained data also confirmed that the set of 51 SSR markers used in the study revealed a realistic picture of the genetic relationships between 89 camellia accessions. In conclusion, the results of this study will enable further genetic mapping, genetic diversity, and germplasm characterization studies in the genus Camellia.
Methods
Plant materials and DNA extraction
A total of 89 camellia accessions, planted in the Camellia Germplasm Resource Garden in Wuhan, Forestry and Fruit Tree Research Institute, Wuhan Academy of Agricultural Sciences, were used in the present study. The selected accessions include the original species and cultivars from C. japonica, C. sinensis, C. sasanqua, C. rosthorniana, C. reticulata, C. azalea, C.nitidissima, C. oleifera, C. chuongtsoensis, as well as their interspecific hybrids, which cover the most of the important camellia breeding species in genus Camellia. The selected accessions also cover almost all flower colors, flower types and breeding approaches (Supplementary Table 5). Qingyuan Li is responsible for a formal and more detailed description of each of these accessions. No specific permissions were required for these plant materials, since these studies did not involve endangered or protected species. All experiments including the collection of plant material in this study are in compliance with relevant institutional, national, and international guidelines and legislation.
Genomic DNA was extracted using a Plant Genomic DNA Extraction Kit (TIANGEN, China) following the manufacturer’s instructions. The DNA quality and concentration were determined by electrophoresis in 1% agarose gel and an Agilent 2100 Bioanalyzer (USA). DNA was diluted with sterilized ultrapure water, normalized to 50 ng/μl, and stored at -20 °C until use.
The identification and annotation of SSR motifs
SSR motifs were identified using MISA37. The search parameters were set for the detection of all di-, tri-, tetra-, penta-, and hexanucleotide SSR motifs with a minimum of five repeats. The SSR-containing unigenes were searched using BLASTX against the GO and KEGG38 databases (E-value ≤ 1E-5) to retrieve protein functional annotations based on sequence similarity. Plant TFs were predicted using iTAK software39.
Primer design and marker validation
The SSR loci were subjected to primer design using Primer 3 web based software40. The parameters were as follows: the length of the primers was in the range of 18–28 bp, with 20 bp as the optimum; the product size range was from 100 to 300 bp; and the melting temperature (Tm) was in the range of 55–65 °C, with 60 °C as the optimum and a maximum Tm difference of 1 °C. All the primer sequences were compared with each other, and primers with more than one copy were deleted to verify that each primer pair amplified only a single SSR. The transcriptome sequences of C. japonica were used as templates for SSR markers discovering by using electronic PCR (e-PCR) with the following parameters: 4-bp mismatch, 1-bp gap, and 80–2000 bp product size41.
A total of 172 primers were randomly selected from the newly designed primers to first validate PCR and SSR polymorphisms among 10 C. japonica accessions (Supplementary Table 2). The primers were synthesized by TSINGKE Biological Technology Co., Ltd. (Beijing, China), and the PCR reagents (buffer, MgCl2+, dNTPs and Taq) were purchased from TransGen Biotech Co., Ltd. (Beijing, China). Amplification was programmed as 5 min at 94 °C for initial denaturation; 30 cycles consisting of 30 s at 94 °C for denaturation, 30 s at the annealing temperature (Ta) for annealing, and 45 s at 72 °C for extension; and finally, a 10-min extension step at 72 °C. The PCR products were loaded for electrophoresis in an 8% polyacrylamide gel. Based on the screening results, 51 polymorphic primer pairs were further used to genotype the 89 camellia accessions using the same volume and PCR program.
Data analysis
Based on the PCR results, a binary matrix was constructed, where the presence of an amplified product was scored as 1 and the absence of the product as 0. Because the templates used in SSR analysis were pooled DNA from 89 plants and more than two alleles per sample were observed, separate bands were treated as individual alleles. The PIC of each locus was calculated using the software PowerMarker version 3.2542. The dendrogram was constructed based on Nei’s genetic distances using the NJ method and was viewed by MEGA743.
STRUCTURE software44 was used to infer population structure. To identify the number of populations (K) capturing the major structure in the data, a burn-inperiod of 100,000 Markov Chain Monte Carlo (MCMC) iterations was used, with a 100,000run length and an admixture model following Hardy–Weinberg equilibrium. The average ln likelihood value when K changed from 1 to 10 was calculated according to genetic similarity, and each run was replicated five times to ensure consistency of the results.
Abbreviations
- SSR:
-
Simple sequence repeats
- MAS:
-
Marker assisted selection
- GO:
-
Gene ontology
- KEGG:
-
Kyoto Encyclopedia of Genes and Genomes
- TFs:
-
Transcription factors
- PIC:
-
Polymorphism information content
- NJ:
-
Neighbour-joining
- PCA:
-
Principal component analysis
- EST:
-
Expressed sequence tag
References
Mondal, T. K. in Wild Crop Relatives: Genomic and Breeding Resources 15–39 (Springer, 2011).
Ming, T. & Bartholomew, B. Theaceae. Flora of China 12, 366–478 (2007).
Prince, L. & Parks, C. Estimation on relationships of Theoideae (Theaceae) infreed from DNA Data. Int Camellia J 32, 79–84 (2000).
Zhang, S. et al. Virome of Camellia japonica: discovery of and molecular characterization of new viruses of different taxa in camellias. Front. Microbiol. 11, 945 (2020).
Bartholomew, B. The Chinese species of Camellia in cultivation. Arnoldia 46, 15 (1986).
Caser, M., Torello Marinoni, D. & Scariot, V. Microsatellite-based genetic relationships in the genus Camellia: potential for improving cultivars. Genome 53, 384–399 (2010).
Xu, X. et al. Assessing the maternal origin in the polyploid complex of Camellia reticulata based on the chloroplast rpl16 intron sequences: implications for camellia cross breeding. Mol. Breed. 38, 123 (2018).
Fu-ping, L. Advance in research of camellia breeding. Guangxi Agricult. Sci. 6 (2008).
Xu, Y. & Crouch, J. H. Marker-assisted selection in plant breeding: from publications to practice. Crop Sci. 48, 391–407 (2008).
Francia, E. et al. Marker assisted selection in crop plants. Plant Cell Tissue Organ Cult. 82, 317–342 (2005).
Avise, J. C. Molecular Markers, Natural History and Evolution. (Springer, 2012).
Gupta, P., Balyan, H., Sharma, P. & Ramesh, B. Microsatellites in plants: a new class of molecular markers. Curr. Sci., 45–54 (1996).
Liu, S. et al. Genome-wide identification of simple sequence repeats and development of polymorphic SSR markers for genetic studies in tea plant (Camellia sinensis). Mol. Breed. 38, 59 (2018).
Singh, R. B., Jugran, A. K., Singh, R. K. & Srivastava, R. K. Assessing genetic diversity and population structure of sugarcane cultivars, progenitor species and genera using microsatellite (SSR) markers. Gene, 144800 (2020).
Tong, Y. & Gao, L. Z. Development and characterization of EST‐SSR markers for Camellia reticulata. Appl. Plant Sci., e11348.
Li, T. et al. Reconstruction of an SSR-based Magnaporthe oryzae physical map to locate avirulence gene AvrPi12. BMC Microbiol. 18, 47 (2018).
Zietkiewicz, E., Rafalski, A. & Labuda, D. Genome fingerprinting by simple sequence repeat (SSR)-anchored polymerase chain reaction amplification. Genomics 20, 176–183 (1994).
Wu, H. et al. De novo characterization of leaf transcriptome using 454 sequencing and development of EST-SSR markers in tea (Camellia sinensis). Plant Mol. Biol. Report. 31, 524–538 (2013).
Tan, L.-Q. et al. Floral transcriptome sequencing for SSR marker development and linkage map construction in the tea plant (Camellia sinensis). PloS One 8 (2013).
Ma, J.-Q. et al. Construction of a SSR-based genetic map and identification of QTLs for catechins content in tea plant (Camellia sinensis). PloS One 9 (2014).
Liu, S. et al. Construction of fingerprinting for tea plant (Camellia sinensis) accessions using new genomic SSR markers. Mol. Breeding 37, 93 (2017).
Tong, Y., Wu, C.-Y. & Gao, L.-Z. Characterization of chloroplast microsatellite loci from whole chloroplast genome of Camellia taliensis and their utilization for evaluating genetic diversity of Camellia reticulata (Theaceae). Biochem. Syst. Ecol. 50, 207–211 (2013).
Shi, J. et al. Discovery and experimental analysis of microsatellites in an oil woody plant Camellia chekiangoleosa. Plant Syst. Evol. 299, 1387–1393 (2013).
Wen, Q., Xu, L., Gu, Y., Huang, M. & Xu, L. Development of polymorphic microsatellite markers in Camellia chekiangoleosa (Theaceae) using 454-ESTs. Am. J. Bot. 99, e203–e205 (2012).
Chen, Z.-Y. et al. Development and characterization of microsatellite markers for Camellia nitidissima. Conserv. Genet. 11, 1163–1165 (2010).
Zhao, Y., Ruan, C., Ding, G. & Mopper, S. Genetic relationships in a germplasm collection of Camellia japonica and Camellia oleifera using SSR analysis. Genet. Mol. Res. 16, 16019526 (2017).
Li, X.-l., Wang, J., Fan, Z.-q., Li, J.-y. & Yin, H.-f. Genetic diversity in the endangered Camellia nitidissima assessed using transcriptome-based SSR markers. Trees, 1–10 (2019).
Li, Q. et al. RNA-seq based transcriptomic analysis uncovers alpha-linolenic acid and jasmonic acid biosynthesis pathways respond to cold acclimation in Camellia japonica. Sci. Rep. 6, 36463. https://doi.org/10.1038/srep36463 (2016).
Lu, Q. et al. Genome-wide identification of microsatellite markers from cultivated peanut (Arachis hypogaea L). BMC Genomics 20, 1–9 (2019).
Xue, H. et al. Genome-wide characterization of simple sequence repeats in Pyrus bretschneideri and their application in an analysis of genetic diversity in pear. BMC Genomics 19, 473 (2018).
Feng, J. et al. Genome-wide genetic diversity detection and population structure analysis in sweetpotato (Ipomoea batatas) using RAD-seq. Genomics 112, 1978–1987 (2020).
Varshney, R. K., Graner, A. & Sorrells, M. E. Genic microsatellite markers in plants: features and applications. Trends Biotechnol. 23, 48–55 (2005).
Zhu, H. et al. Genome wide characterization of simple sequence repeats in watermelon genome and their application in comparative mapping and genetic diversity analysis. BMC Genomics 17, 557 (2016).
Jia, B. et al. Development and cross-species transferability of unigene-derived microsatellite markers in an edible oil woody plant, Camellia oleifera (Theaceae). Genet. Mol. Res. 14, 6906–6916 (2015).
Sharma, H. et al. Identification and cross-species transferability of 112 novel unigene-derived microsatellite markers in tea (Camellia sinensis). Am. J. Bot. 98, e133–e138 (2011).
Lin, L., Hu, Z.-Y., Ni, S., Li, J.-Y. & Qiu, Y.-X. Genetic diversity of Camellia japonica (Theaceae), a species endangered to East Asia, detected by inter-simple sequence repeat (ISSR). Biochem. Syst. Ecol. 50, 199–206 (2013).
Beier, S., Thiel, T., Münch, T., Scholz, U. & Mascher, M. MISA-web: a web server for microsatellite prediction. Bioinformatics 33, 2583–2585 (2017).
Kanehisa, M. & Goto, S. KEGG: kyoto encyclopedia of genes and genomes. Nucleic Acids Res. 28, 27–30 (2020).
Zheng, Y. et al. iTAK: a program for genome-wide prediction and classification of plant transcription factors, transcriptional regulators, and protein kinases. Mol. Plant 9, 1667–1670 (2016).
Rozen, S. & Skaletsky, H. in Bioinformatics Methods and Protocols, 365–386 (Springer, 2000).
Schuler, G. D. Electronic PCR: bridging the gap between genome mapping and genome sequencing. Trends Biotechnol. 16, 456–459 (1998).
Liu, K. & Muse, S. V. PowerMarker: an integrated analysis environment for genetic marker analysis. Bioinformatics 21, 2128–2129 (2005).
Kumar, S., Stecher, G. & Tamura, K. MEGA7: molecular evolutionary genetics analysis version 7.0 for bigger datasets. Mol. Biol. Evolut. 33, 1870–1874 (2016).
Pritchard, J. K., Stephens, M. & Donnelly, P. Inference of population structure using multilocus genotype data. Genetics 155, 945–959 (2000).
Acknowledgements
This research was financially supported by funding from the National Natural Science Foundation of China (31701965), National Key R&D Program of China (2019YFD1000400) and Hubei Province Major Projects of Technological Innovation (2017ABA162). We would like to express our sincere thanks to Prof. Ling Min (Huazhong Agricultural University) and Prof. Guangqin Cai (Oil Crops Research Institute, Chinese Academy of Agricultural Sciences) for helping in the analysis of the data and the revision of the manuscript.
Author information
Authors and Affiliations
Contributions
QL, LX and KD conceived and designed the experiments. CX, MY and TF performed the DNA isolation experiment. XS, HM and QZ performed the PCR experiment and PCR products determination. QL, SF and BC analyzed the data and wrote the manuscript. LX and QL coordinated the study and revised the manuscript. All authors read and approved the final manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Li, Q., Su, X., Ma, H. et al. Development of genic SSR marker resources from RNA-seq data in Camellia japonica and their application in the genus Camellia. Sci Rep 11, 9919 (2021). https://doi.org/10.1038/s41598-021-89350-w
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-021-89350-w
- Springer Nature Limited
This article is cited by
-
Assembly and characterization analysis of the complete mitochondrial genome of Lithocarpus litseifolius (Hance) Chun
Genetic Resources and Crop Evolution (2024)
-
Assembly and comparative analysis of the complete mitochondrial genome of Trigonella foenum-graecum L.
BMC Genomics (2023)
-
Discrimination of Camellia cultivars using iD-NA analysis
Scientific Reports (2023)
-
Identification of Pueraria spp. through DNA barcoding and comparative transcriptomics
BMC Plant Biology (2022)
-
Assembly and comparative analysis of the complete mitochondrial genome of Bupleurum chinense DC
BMC Genomics (2022)