Abstract
Huntington’s disease (HD) is a neurodegenerative disorder caused by a CAG repeat expansion in exon 1 of huntingtin (HTT). While there are currently no disease-modifying treatments for HD, recent efforts have focused on the development of nucleotide-based therapeutics to lower HTT expression. As an alternative to siRNA or oligonucleotide methods, we hypothesized that suppression of HTT expression might be accomplished by small molecules that either (1) directly decrease HTT expression by suppressing HTT promoter activity or (2) indirectly decrease HTT expression by increasing the promoter activity of HTT-AS, the gene antisense to HTT that appears to inhibit expression of HTT. We developed and employed a high-throughput screen for modifiers of HTT and HTT-AS promoter activity using luminescent reporter HEK293 cells; of the 52,041 compounds tested, we identified 898 replicable hits. We used a rigorous stepwise approach to assess compound toxicity and the capacity of the compounds to specifically lower huntingtin protein in 5 different cell lines, including HEK293 cells, HD lymphoblastoid cells, mouse primary neurons, HD iPSCs differentiated into cortical-like neurons, and HD hESCs. We found no compounds which were able to lower huntingtin without lowering cell viability in all assays, though the potential efficacy of a few compounds at non-toxic doses could not be excluded. Our results suggest that more specific targets may facilitate a small molecule approach to HTT suppression.
Similar content being viewed by others
Introduction
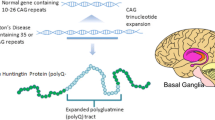
Huntington’s disease (HD) is an autosomal dominant neurodegenerative disorder characterized by abnormalities of movement, cognition, and emotion, with relentless progression to death1. HD is caused by expansion of a CAG repeat in exon 1 of the ubiquitously expressed Huntingtin (HTT) gene; the repeat is in-frame to encode polyglutamine2. In individuals with HD, the mutant huntingtin protein (mHTT) containing a long polyglutamine expansion is found throughout the CNS and also in non-CNS tissues3. HD pathogenesis is thought to derive primarily from a toxic gain-of-function conferred on mHTT by the polyglutamine expansion4, although some evidence exists for other forms of toxicity, including loss-of-function5,6 and toxicity from the mutant mRNA transcript7,8. There are currently no disease-modifying treatments for HD. Multiple attempts to target downstream pathways related to excitotoxicity, mitochondrial dysfunction, and inflammation have thus far met with little success9. More recent studies of pridopidine10, coenzyme Q1011, and dimebon12 show some benefit, although results have not always been consistent.
As an alternative to therapeutic approaches that target downstream pathways, suppression of expression of the mutant HTT allele has the considerable theoretical advantage of simultaneously preventing dysfunction of most or all of these pathways. Preclinical studies of HTT-lowering via RNA interference (RNAi), antisense oligonucleotides (ASOs), and synthetic zinc finger repressors have demonstrated the potential of this approach, with evidence of amelioration of HD pathology and increased survival in animal models13. These RNA-targeting methods are undergoing rapid development. Among the most salient developments, a recent phase 1/2a trial of a nonallele-specific modified ASO targeting the HD transcript demonstrated tolerability and decreased CSF levels of mHTT, a biomarker of disease activity14; a phase 3 trial is now underway (ClinicalTrials.gov NCT03342053). An allele-specific ASO approach is in the midst of two 1a/2b clinical trials (ClinicalTrials.gov NCT03342053 and NCT03225833). A phase 1/2 trial using adeno-associated virus (AAV)-expressed RNAi at an early stage of disease is also underway (ClinicalTrials.gov NCT04120493). Gene editing approaches are another intriguing alternative, still in the early stages of development15. While very exciting, these approaches face numerous challenges: off-target inhibition of other genes, immunostimulation, lack of specificity for the mutant allele, the need for broad brain distribution, and the development of clinically feasible delivery mechanisms.
A recently discovered gene antisense to HTT (huntingtin antisense, HTT-AS)16 represents another potential drug target. HTT-AS falls within the category of natural antisense transcripts, a subset of long non-coding RNAs (lncRNAs) which at least partially overlap with a gene on the opposite strand. As many as 72% of genes in mice and 40% in humans may have corresponding antisense genes17. These are typically expressed at much lower levels than the sense transcript, but often have critical regulatory roles. For instance, in endothelial cells, the expression of sONE, the antisense transcript to eNOS, is about 500 × lower than eNOS. Nonetheless, stimulation of sONE expression with the small molecule TSA (a histone deacytylase inhibitor) leads to an 80% decrease in eNOS protein expression, apparently via post-translational mechanisms18,19. Similar examples exist in genes relevant to neurological diseases. The expression level of a splice variant of ASFMR1, the gene antisense to fragile X mental retardation gene (FMR1), may influence the risk for development of the fragile X tremor ataxia syndrome (FXTAS)20. HTT-AS and HTT appear to have a similar relationship: in cell systems, overexpression of the HTT-AS transcript in cis resulted in a decrease in endogenous HTT transcript levels, while siRNA knockdown of HTT-AS increased HTT transcript levels16.
The manipulation of natural antisense transcripts (NATs) is of therapeutic interest, particularly through AntagoNATs, a term coined to describe modified oligonucleotides that interfere with sense-antisense interactions21. AntagoNATs were used to target BDNF-AS to increase BDNF transcript levels by 2–7 ×; similar effects were observed with AntagoNAT suppression of NATs of both GDNF and EPHB2. AntagoNAT-induced decrease of the transcript antisense to the SCN1A gene results in upregulation of SCNA1, with phenotypic improvement in a Dravet syndrome mouse model22. Knockdown of SMN-AS1 in fibroblasts by treatment with a chemically modified oligonucleotide increased SMN2 expression sixfold, a potential approach to the degenerative disease spinal muscular atrophy (SMA)23. As therapeutic agents AntagoNATs have the same advantages and disadvantages of other oligonucleotide-based approaches, as noted above.
Alternatively, small molecules that penetrate the blood–brain barrier do not require intraventricular or intrathecal infusions and avoid some of the safety concerns associated with the viral delivery typically required for shRNA-based therapeutics24. Screens using assays in which promoters were linked to a luciferase reporter have yielded small molecules that upregulate expression of the Notch pathway-associated transcription factor ATOH125, suppress expression of HAMP (which encodes hepcidin, a regulator of iron homeostasis)26,27, and suppress expression of PHOX2B, a gene involved in neurogenesis28. Optimism for the use of small molecules to modify gene expression has been encouraged by the success of risdiplam, a small molecule that modifies SMN2 splicing, increasing full length expression of SMN2 to correct the loss of SMN1 in patients with spinal muscular atrophy29. Risdiplam improved patient outcome in multiple clinical trials (ClinicalTrials.gov NCT03779334, NCT02908685, NCT02913482, NCT03032172), and recently received FDA approval for use in SMA patients30. A second agent, branaplam, also acts on SMN2 splicing31 and is currently entering clinical trials (ClinicalTrials.gov NCT02268552).
In HD, attempts to find small molecules that decrease HTT expression have thus far not yielded agents of sufficient interest to reach the stage of clinical trial1,32. Recognition that mHTT itself has a widespread effect, direct and indirect, on expression of other genes33,34 has led to attempts to reverse these effects, including efforts to inhibit DNA methyltransferases, histone deacetylases, and histone methyltransferases35. Clinical progress has been limited due to pleiotropic targets and evidence of toxicity.
We developed quantitative high-throughput screening (HTS) assays for small compounds that act either to suppress activity of the HTT promoter or activate the HTT-AS promoter. The HTS was followed by a rigorous multistep validation process in which HTT protein expression and cytotoxicity was examined in 5 different disease relevant cell systems. While many compounds demonstrated apparent activity at the promotor level, none of them were both efficacious and non-toxic in all the validation assays. We review the advantages and disadvantages of this rigorous approach to a drug screen for HTT expression modulators, and comment on a few compounds that remain of interest.
Materials and methods
Compound libraries
All compound libraries used in this study are listed in Table 1. Identity of compounds can be found in the NCBI PubChem repository (Assay IDs: 1508621, 1508622, 1508623, 1508624).
Cell lines and culture methods
The cell lines used in this study are listed in Table 2. Flp-In T-Rex Human Embryonic Kidney 293 (HEK293) cells were purchased from ThermoFisher Scientific, while the parental HEK293 cell line was obtained from ATCC (Manassas, VA, USA). Both HEK293 lines were grown in Dulbecco’s modified Eagle’s medium (DMEM) supplemented with 10% fetal bovine serum (FBS) and 1 × Penicillin–Streptomycin-Amphotericin B (P/S/A). GM05539 lymphoblastoid cells from a male HD patient were obtained from the Coriell Institute for Medical Research and grown in Roswell Park Memorial Institute 1640 (RPMI1640) medium supplemented with 15% FBS and 1 × P/S/A. STHdhQ7 and STHdhQ111 cells (a gift from Marcy MacDonald) were grown in DMEM supplemented with 1% FBS, 5 mM sodium pyruvate and 0.3 × P/S as previously described36. All cells lines were cultured in 37 °C incubators at 5% CO2, except for STHdh cells which were cultured at 33 °C.
Stem cell differentiation and compound treatment
Neural progenitor cells derived from induced pluripotent stem cells (iPSCs)37 were revived from day in vitro (DIV) 18 onto 6-well plates and differentiated into human cortical neurons (hCNs) until DIV25, at which point cells were seeded onto coated 24-well plates and penicillin/streptomycin was added to media. Media was topped up at DIV26 and cells underwent a 50% media change at DIV29. Cells were incubated with compound from DIV32 to DIV39, at which point lysates from each plate were split for use in two cell-based assays. For assays involving the GENEA020 human embryonic stem cells (hESCs)38, cells were plated onto collagen-coated plates and allowed to attach overnight prior to treatment with compound for 48 h. Lysate from one plate was split for use in three cell-based assays.
Dual-luciferase luminescence measurement
Baseline promoter activity of stable expression cell lines was measured in 96-well white-walled clear-bottom plates plated with 10,000 cells in 100 μL growth media per well. Doxycycline was added to CMV tet-on promoter cells in a volume of 25 μL during plating. Luminescence measurements were made following 72 h of culture at 37 °C. Validation experiments were conducted in 96-well white-walled plates containing 60,000 cells per 100 μL per well cultured for 24 h. Compounds were added in a volume of 25 μL per well and incubated for 24 h at 37 °C. Treatment media was replaced with 50 μL fresh growth media prior to luminescence measurements. High-throughput screening, measuring luminescence from a luciferase reporter, was performed in 1536-well white-walled tissue-culture treated plates. Briefly, 2500 cells were plated in 4 μL of Opti-MEM per well using a Multidrop Combi Reagent Dispenser (ThermoFisher) and cultured for 24 h. The compound libraries were transferred in a volume of 23 nL per well. Cells were then grown for 24 h at 37 °C prior to luminescence measurements. In all cases, luminescence was measured using the Dual-Glo Luciferase Reporter Assay System (Promega) following the manufacturer’s instructions.
Time-resolved fluorescence energy transfer (TR-FRET) assay for total HTT and mutant HTT protein levels
For initial TR-FRET assays, 4000 HEK293 cells or 8000 lymphoblastoid cells were plated in a 1536-well plate in 6 μL of growth media per well and cultured overnight (~ 16 h). Compounds were added to wells in 23 nL and incubated for 24 h. Cells were lysed with 2 μL of Cisbio-Lysis buffer #2 and incubated at room temperature for 2 h. Antibodies (Table 3) were added to the lysate in a volume of 2 μL and the plate was incubated at 4 °C for 23 h. Time-resolved fluorescent signal was measured at 665 and 615 nm using an Envision Multimode Plate Reader (Perkin Elmer) to determine FRET ratio for each well. For assays in HEK293 and lymphoblastoid cells, mutant and total HTT were measured. For assays in hESCs, lysate collected from compound-treated cells was used for a multiplexed mutant HTT, total HTT, and AKT. For assays in cortical neurons, lysate was used for a multiplexed mHTT and Tau assay.
Cell viability assays
For all ATP-based cell viability assays, the Cell Titer Glo kit (Promega) was used according to manufacturer’s instructions. For total nuclear stain in hESCs, cells were fixed and permeabilized followed by Hoechst 33342 staining. Cells were imaged using the InCell2200 widefield automated microscope (GE Healthcare) and data analyzed using the InCell Developer software.
The STHdhQ111 1536-well viability assay was performed as previously described39. Briefly, cells were plated in black-wall, clear-bottom 1536-well cyclic olefin polymer-type imaging plates (Edition Eight; Whitefish, MT) at 1200 cells per well in 5 μL volume using a Multidrop Combi Reagent Dispenser (ThermoFisher). Cells were incubated for 16 h in an incubator at 33 °C and 5% CO2. 46 nL of compounds were transferred using a pin-transfer tool and plates were returned to 33 °C for 2 h. Cells were then shifted to 37 °C and 5% CO2 for 24 h prior to staining and imaging. Hoechst 33342 and propidium iodide (PI) were prepared in phosphate buffer solution (PBS) and 1 μL was added to each well; final concentration was 4 μg/mL Hoechst 33342 and 5 μg/mL PI. Plates were incubated at room temperature for 30 min prior to imaging on an InCell2200 microscope using a 10 × 0.45 NA air objective and standard DAPI (390/18 ×, 432/48 m) and Cy3 (542/27 ×, 587/45 m) filter sets. One field of view per well, encompassing the entire well, was chosen for imaging and percent PI positive cells were determined using Columbus Image Data Analysis System (PerkinElmer) to identify Hoechst nuclei and PI positive nuclei on a cell-by-cell basis. A mean PI intensity of more than 3 standard deviations above the background mean was classified as PI positive. Data were normalized to controls on a per-plate basis (Q7 + vehicle = 100% protection, and Q111 + vehicle = 0% protection).
Data analysis
Concentration–response curves (CRCs) were generated using NCATS software (https://tripod.nih.gov/curvefit/) and analyzed using previously described methods40. Curves are classified based on quality of fit to the data, the magnitude of response, and the number of asymptotes to the calculated curve. A positive CRC would indicate a positive correlation of activity with dose concentration, while a negative CRC would indicate a negative correlation. Structural clustering of active compounds was performed using Leadscope Hosted Client (Leadscope Inc., Columbus, OH). The EC50 values of compounds in the confirmation and follow-up experiments were calculated from the dose–response curves by nonlinear regression analysis using Prism software (GraphPad Software, San Diego, CA).
Results
Development of stable reporter cell lines to measure HTT and HTT-AS promoter activity
In order to identify small molecules that alter HTT or HTT-AS expression, we developed a reporter-based screen to assess the effects of small molecules on their respective promoters. We transfected Flp-In T-Rex HEK293 cells with a dual-reporter gene plasmid linked with the HTT (1.7 kb) or HTT-AS (1.5 kb) promoter. The HTT promoter sequence included 1736 bp upstream of the transcription initiation site (chr4:3073089–3074829 human genome 38; hg38)41. The HTT-AS promoter sequence (1503 bp upstream of the transcription initiation site; chr4:3074930–3076433 hg38) was chosen based on previously published reporter experiments16. Additional control plasmids were also constructed, including no promoter (“Null” control), or a CMV tet-on inducible promoter (Fig. 1A). We performed stable monoclonal selection and analyzed a series of monoclones. HTT-2 and HTT-AS-8 clones were selected based on results showing uniform baseline firefly and renilla luciferase activities (Fig. 1B–E), normal cell morphology, lack of effect of DMSO (1%) on growth and reporter expression, high proliferation rates, and no loss of transgene through at least P10.
Strategy for generation of stable cell lines. (A) Constructs containing a promoter linked to two luciferase reporters were cloned into a pcDNA5/FRT backbone. Plasmids were transfected into HEK293 host cells with an integrated Flp recombination site. (B–E) Baseline luciferase activity was assayed in clones expressing firefly linked to (B) CMV promoter, with or without doxycycline (Dox) treatment (n = 3), (C) no promoter (“Null”; n = 4), (D) HTT promoter (n = 4), and (E) HTT-AS promoter (n = 4). Raw luminescence signal values are shown (luciferase activity); error bars are ± SD.
Primary screen: screening for compounds which activate the HTT-AS promoter or inhibit the HTT promoter using a luciferase reporter system
52,041 compounds from five compound libraries (Table 1) were tested for their effect on the HTT and HTT-AS promoter-expressing cell lines. Compounds were screened at four concentrations (57, 11, 2.3, and 0.46 μM). Renilla reporter results were excluded due to low signal-to-noise ratio (Supplement Figure 1). The primary screen yielded 898 compounds (hit rate = 1.73%) which had active concentration response curves (CRCs) indicating increased expression of HTT-AS (curve class 1/2/3) or decreased expression of HTT (curve class − 1/− 2/− 3). Of the 898 total actives, 809 inhibited the HTT promoter, 92 activated the HTT-AS promoter, and 3 compounds had both effects. The 898 compounds from the primary screening could also be subdivided into the following five activity profiles:
-
1.
Decrease in HTT and HTT-AS promoter activity (687); potential general transcriptional inhibitors.
-
2.
Increase in HTT and HTT-AS promoter activity (84); potential general transcriptional activators.
-
3.
Decrease in HTT promoter activity, increase in HTT-AS activity (3); desirable profile (both).
-
4.
Decrease in HTT promoter activity, no change in HTT-AS activity (119); desirable profile (HTT).
-
5.
No change in HTT promoter activity, increase in HTT-AS activity (5); desirable profile (HTT-AS).
Confirmation screen: identification of reproducible hits from dual-reporter screen
898 compounds identified in the initial promoter screen were reevaluated at 7 concentration (57 μM top concentration, 1:3 titration) to confirm their dose–response activity based on CRCs (1/2/3 for HTT-AS or − 1/− 2/− 3 for HTT) (Fig. 2). 739/809 HTT lowering compounds (91%) and 52/92 (57%) HTT-AS activating compounds confirmed the validation screen, for a composite confirmation rate of 88% (791/898). Compounds that were active in the confirmation screen were further triaged by structural clustering, elimination of compounds with promiscuous and undesirable functionalities, and removal of compounds which showed an increase in HTT promoter activity as such compounds were unlikely to prove of clinical interest. 123 compounds met these criteria and were selected for further study. Of this group, 112 decreased HTT promoter activity and 15 increased HTT-AS activity, with four compounds both decreasing HTT and increasing HTT-AS promoter activity.
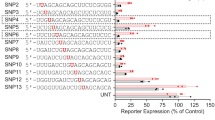
Replication of compound HTT and HTT-AS promoter activity. HTT or HTT-AS stable reporter cell lines were treated with 898 compounds identified in the HTS to replicate the initial findings. (A,B) Maximal response of luciferase activity at the highest dosage of compound treatment (57 μM) in (A) HTT- and (B) HTT-AS-promoter reporter cells. Hits were determined based on CRC values and are indicated in red (HTT downregulators; HTT CRC = − 1/− 2/− 3), blue (HTT-AS upregulators; HTT-AS CRC = 1/2/3), and green (HTT downregulators AND HTT-AS upregulators). The remaining compounds with unfavorable or inactive CRCs are indicated in gray. One outlier (449%) from (A) and 3 outliers (843%, 458%, 403%) from (B) are not shown. (C,D) Waterfall plots of luciferase activity in (C) HTT and (D) HTT-AS reporter cells in response to treatment by 123 compounds chosen for further study. Red: strongly inactivating compounds (CRC = − 1.1/− 1.2/− 2.1/− 2.2). Orange: weakly inactivating compounds (CRC = − 1.3/− 1.4/− 2.3/− 2.4/− 3). Green: inactive compounds (CRC = 4/5). Cyan: weakly activating compounds (CRC = 1.3/1.4/2.3/2.4/3). Blue: strongly activating compounds (CRC = 1.1/1.2/2.1/2.2).
Secondary screen: effect on huntingtin protein in HEK293 and HD patient lymphoblast cell
In order to ascertain huntingtin protein-lowering capacity, a FRET-based assay was used to measure the dose–response effect of the 123 selected compounds on huntingtin protein levels. Compounds were screened at 10 concentration dose response in both HEK293 cells (38 μM with 1:3 dilutions) and a lymphoblast cell line from an HD patient (long allele = CAG82; 17 μM with 1:3 dilutions). HD patient-derived lymphoblasts were selected for use in this study as they have been successfully used to identify novel aspects of HD pathophysiology42,43,44, as well as evaluate the effectiveness of potential therapeutic agents45,46. An ATP-measuring assay was performed in parallel using the same concentration range to determine compound toxicity.
A total of 17 compounds strongly lowered huntingtin protein levels as measured by the Homogeneous Time-Resolved Fluorescence (HTRF) assay, and evaluated by CRCs (− 1.1/− 1.2/− 2.1/− 2.2): 10 were active in HEK293 cells, 4 in HD lymphoblasts, and 3 in both cell lines. After eliminating toxic compounds, 14 were classified as active (Table 4; Fig. 3); 10 in HEK293 cells only, 6 in HD lymphoblasts only, and 2 compounds in both cell lines. Of the two compounds that lowered huntingtin protein in both cell types, one (NCGC00274038) was a hit derived from the HTT-AS screen, and one (NCGC00104681) from the HTT screen.
Huntingtin protein-lowering and cytotoxicity assays in HEK293 cells and HD lymphoblasts of 123 compounds that altered HTT or HTT-AS promoter activity. 14 compounds (A–N) out of 123 tested lowered huntingtin protein in either HEK 293 cells (HTRF-HEK) or HD Lymphoblasts (HTRF-Lymph) based on an HTRF assay and were non-toxic according to ATP-based cytotoxicity assays (Tox-HEK; Tox-Lymph). Each compound was screened at 10 concentrations (HEK293 screen, highest concentration = 38 μM, with 1:3 dilutions; Lymphoblast screen, highest concentration = 17 μM with 1:3 dilutions; single well per concentration). Compounds were chosen based on active CRCs for huntingtin lowering (− 1/− 2/− 3) and inactive CRCs for cytotoxicity (≥ 4). Values are normalized to measurements in untreated HEK293 and HD lymphoblast cells.
Tertiary screen: huntingtin protein lowering in stem cell and neuronal HD lines
12 out of the 14 compounds active compounds in the HTRF assay above were further tested in human embryonic stem cells (hESCs) from an HD patient (Charles River, long allele = CAG48) and cortical neurons differentiated from an HD induced pluripotent stem cell (iPSC) line (Evotec, long allele = CAG72). Two compounds were excluded due to hazardous material transportation restrictions or limited supplies.
Cortical neurons were treated with compound for seven days at 10 doses (30 μM with 1:3 dilutions). A FRET-based assay was used to assess the capacity of each compound to lower mutant huntingtin, with an ATP-based cell viability assay run in parallel. One compound (NCGC00274038) exhibited some lowering of mHTT at concentrations that did not alter ATP levels (Fig. 4A,B). However, it also lowered Tau as assayed by FRET, indicating a lack of specificity. HD hESCs were treated with compound at 10 doses (30 μM with 1:3 dilutions). Both mutant and total huntingtin were measured, as well as protein kinase B (Akt) to assess specificity of action. Cell viability was measured both by an ATP-based assay and by total cell count based on Hoechst staining. One compound (NCGC00274038) showed measurable huntingtin-lowering activity relative to ATP-based cell viability (Fig. 4C–F). However, this compound had equipotent effects on huntingtin-lowering cell viability with toxicity measured by cell count. Consistent with this effect and with results in cortical neurons, Akt suppression also paralleled HTT-lowering, indicating non-specific action by the compound.
Tertiary screening for huntingtin-lowering and toxicity in HD neuronal and stem cell lines. 12 compounds were tested in (A,B) HD cortical neurons and (C–F) HD embryonic stem cells, and 8 compounds in (G) HD mouse striatal cells. HTRF assays were used to measure mHTT, total HTT, Akt, and Tau concentrations at each dosage. Cytotoxicity was determined by total nuclear cell count (Tox-TNS) or ATP-measurement assays (Tox-ATP). (A) mHTT suppression IC50 is plotted against ATP-based cytotoxicity. One compound (NCGC00274038; N) out of 12 shows ability to lower mHTT suppression with a window over cytotoxicity. (B) Dose–response curve of NCGC00274038 Tau protein suppression, mutant HTT (mHTT) protein suppression and ATP-based cytotoxicity (Tox-ATP) in HD cortical neurons. Values are normalized to measurements in untreated cells. Error bars are ± SD; n = 2. (C) mHTT suppression IC50 is plotted against ATP cytotoxicity IC50. NCGC00274038 suppresses mHTT with a window over ATP-based cytotoxicity in HD hESCs. (D) mHTT suppression IC50 is plotted against cell count-based cytotoxicity IC50. No compounds show ability to suppress mHTT with a window over cell count-based cytotoxicity. (E) Total HTT suppression IC50 is plotted against cell count-based cytotoxicity IC50. No compounds show specific suppression of total HTT over Akt. (F) Dose–response curve of protein suppression (Tau, mutant HTT, total HTT) as well as ATP-based (Tox-ATP) and cell count-based (Tox-TNS) cytotoxicity in HD hESCs in response to NCGC00274038 treatment. Values are normalized to measurements in untreated HD hESCs. Error bars are ± SD; n = 4. (G) Dose response curves of 8 compounds tested for protective activity in STHdhQ111 model. Proportion of live cells detected by PI (negative) was measured relative to total cells detected by Hoechst 33342 (% Protection), normalized to measurements in untreated STHdhQ7 cells. Error bars are ± SD; n = 2. (H) Letters in IC50 plots indicate compounds IDs, also shown in Table 4.
8 of the 14 hits derived from HEK293 and HD lymphoblast studies were also tested for their cytoprotective capacity in STHdhQ111 mouse striatal cells. Striatal neurons were treated with compound for 24 h at 11 doses (30 μM with 1:3 dilutions) while under stress conditions (low temperature, low serum). Cells were stained with Hoechst to label all nuclei and propidium iodide (PI) to detect dead cells. Percentage of PI positive neurons in compound-treated cells relative to untreated Q7 neurons was used to measure cytoprotection. Two compounds (NCGC00100369, NCGC00113437) showed strong cytoprotective effects (Fig. 4G), three compounds showed mild cytoprotective effects (NCGC00122597, NCGC00136813, NCGC00140752), two compounds were cytotoxic (NCGC00099051, NCGC00274038), and one compound was inactive (NCGC00131485).
Discussion
We developed a high-throughput luciferase reporter assay to screen for compounds that act on the HTT or HTT-AS promoter, using HEK293 cells expressing the HTT or HTT-AS promoter linked to a luciferase reporter. This assay is 1536-well compatible, enabling us to screen a diverse collection of ~ 50,000 compounds. We identified 739 compounds that reproducibly decreased HTT promoter-driven luciferase activity, and 52 that increased HTT-AS promoter-driven luciferase activity. Validation, based on the capacity of hits to lower huntingtin protein at non-toxic concentrations, was performed in two tiers, first, using HEK293 cells and lymphoblasts from HD patients; and second, using HD patient ESCs and iPSCs, and primary neurons from an HD mouse model. None of the small molecules identified in the initial screen were efficacious at non-toxic doses in all tests of validity (Fig. 5).
Summary of high throughput screening for HTT and HTT-AS promoter modifiers. 52,041 compounds were tested in the primary screen, with a total of 123 hits (112 hits in the HTT promoter assay and 15 in the HTT-AS promoter assay, with 4 compounds active in both assays. Of these 123 compounds, 14 were identified as huntingtin-lowering and non-toxic in the secondary screen. 10 were found to lower huntingtin protein in HEK293 cells and 6 in HD-patient derived lymphoblasts, with 2 active in both cell types. 12 of these compounds were tested in HD human embryonic stem cells (hESCs) and HD cortical neurons (hCNs), and 8 in immortalized HD mouse striatal neurons (mStNs). Compound IDs (Table 4) in chart indicate which compounds moved forward to each tertiary screen. 2 compounds (NCGC00100369; B) and (NCGC00113437; D) were identified as active in the mStNs, but not in hESCs or hCNs.
The discrepancy of results between HEK cells, lymphoblasts, and neuronal and stem cell lines may in part derive from differences in the mechanisms that regulate HTT expression. While huntingtin is ubiquitously expressed, expression levels are presumably under at least some cell-type specific regulation, as indirectly demonstrated by varying levels of HTT expression in different tissues (e.g., the expression of normal HTT is 5 × higher in transformed lymphocytes than in caudate). Other small molecules detected by moderate- or high-throughput screenings as potential HD therapeutic agents have also acted differently on specific cell types, and different assays may yield different results in the same cell-type47,48. The mechanisms of HTT transcriptional regulation, and how this regulation differs by cell type, is poorly understood49. Our strategy of testing efficacy, specificity, and toxicity in non-neuronal cells first was intended to reduce false positives while increasing the efficiency and decreasing the cost of the screen. The trade-off is a likely increase in false negatives. Other high-throughput screenings for small molecule therapeutics for HD have faced similar obstacles50,51.
Our approach emphasized cultured neurons as a final step in validation and test of toxicity. However, cultured neurons are more vulnerable to toxicity than either neurons in vivo or in co-culture with astrocytes52,53,54, increasing the likelihood that potential compounds of interest will be screened-out based on toxicity55. Use of co-culture systems, previously developed for investigation of HD pathogenesis56, or 3D cultures57, will likely decrease this source of false negatives. In addition, our results reflect the methodology used for evaluation of toxicity. ATP-based toxicity screening was used to assess toxicity in HEK293 cells, HD lymphoblasts, and induced cortical neurons, while both ATP and cell count-based methods were used to assess toxicity in HD hESCs. While cell counting methods avoid the issues of cell growth that can confound assays based on ATP production, the advantage comes with the cost of scalability.
One compound, NCGC00274038, a TGBβ-activated kinase (TAK1) inhibitor, lowered HTT and was not toxic by ATP measures in induced neurons, though it was toxic when assayed by cell count in hESCs and in a mouse striatal-derived cell line. We excluded this compound from further consideration based on our a priori analytic plan and evidence that direct cell counting is superior to ATP-based assays, recognizing that this decision is quite conservative. However, this does not necessarily exclude this compound or related compounds from further investigation. Previous studies have shown TAK1 inhibitors to have neuroprotective properties58,59. TAK1 is a positive regulator of MAPK signaling, and dysregulation of this pathway has been implicated in HD60 and other neurodegenerative diseases61. Two recent studies have identified MAPK-related kinases as a positive regulators of mHTT protein levels, potentially via increased stability of HTT mRNA62,63. NCGC00274038 may therefore reduce HTT expression via suppression of MAPK signaling; its detection in our screen for compounds acting on the HTT promoter may have been artifactual, from an unrelated or indirect activity, or due to a stabilizing effect on luciferase mRNA. Other TAK1 inhibitors without the toxicity we detected in NGC00274038 may therefore merit investigation in HD, though likely independent of an effect on the HTT promoter.
This study included a screen for small molecules that upregulated HTT-AS, guided by the rationale that, as found in sense-antisense pairs at other loci, HTT-AS appears to suppress HTT. This screen had the advantage of minimizing detection of compounds on the basis of nonspecific toxicity. We focused on the HTT-AS promoter that drives transcription of HTT-AS-v1, which includes exon 1 with the CUG repeat. One challenge is that over-induction of this transcript may be toxic via the toxicity of expanded CUG repeats64,65. More generally, the regulation of expression at bidirectional loci is complex, and it is possible that small molecule manipulation of the antisense promoter is insufficient to elicit the response detected in model systems16. We have recently detected a second promoter that appears to drive expression of HTT-AS-v2; this transcript does not incorporate the HTT-AS exon 1, which contains the repeat, but does incorporate HTT-AS exon 2, which is precisely located antisense to the HTT promoter region. While the extent to which activating this second promoter can lead to suppression of HTT remains unclear, it may prove a more amenable target for manipulation than the promoter targeted in this study.
Our assay was designed to detect any potential activator of the HTT or HTT-AS promoter, in part based on limited information on more specific regulators of HTT or HTT-AS promoter function. If detected, such factors might provide a more focused therapeutic target. Work outside of HD suggests that other targets with the potential of lowering HTT expression might include promoter specific G-Quadruplexes66, the point of interaction between a transcription factor and Pol II67, promoter-regulating lncRNAs68, or translational machinery69.
We conclude that reducing mHTT expression remains an important goal in the field. Antisense and RNAi strategies look promising, but a small molecule approach may avoid some of the obstacles facing these strategies or could lead to an adjuvant therapy. Our results suggest that identifying more specific targets may facilitate this approach.
Data availability
The datasets generated during the current study are available in the NCBI PubChem repository with the following Assay IDs: 1508621, 1508622, 1508623, 1508624. Note, classification of active versus inactive compounds using the PubChem Activity Score is not identical to the criterion described herein.
References
Kieburtz, K., Reilmann, R. & Olanow, C. W. Huntington’s disease: Current and future therapeutic prospects. Mov. Disord. 33, 1033–1041. https://doi.org/10.1002/mds.27363 (2018).
The Huntington’s Disease Collaborative Research Group. A novel gene containing a trinucleotide repeat that is expanded and unstable on Huntington’s disease chromosomes. Cell 72, 971–983. https://doi.org/10.1016/0092-8674(93)90585-e (1993).
Sharp, A. H. et al. Widespread expression of Huntington’s disease gene (IT15) protein product. Neuron 14, 1065–1074. https://doi.org/10.1016/0896-6273(95)90345-3 (1995).
Bates, G. P. et al. Huntington disease. Nat. Rev. Dis. Primers 1, 15005. https://doi.org/10.1038/nrdp.2015.5 (2015).
Dragatsis, I., Levine, M. S. & Zeitlin, S. Inactivation of Hdh in the brain and testis results in progressive neurodegeneration and sterility in mice. Nat. Genet. 26, 300–306. https://doi.org/10.1038/81593 (2000).
McKinstry, S. U. et al. Huntingtin is required for normal excitatory synapse development in cortical and striatal circuits. J. Neurosci. 34, 9455–9472. https://doi.org/10.1523/JNEUROSCI.4699-13.2014 (2014).
Banez-Coronel, M. et al. A pathogenic mechanism in Huntington’s disease involves small CAG-repeated RNAs with neurotoxic activity. PLoS Genet. 8, e1002481. https://doi.org/10.1371/journal.pgen.1002481 (2012).
Sun, X. et al. Nuclear retention of full-length HTT RNA is mediated by splicing factors MBNL1 and U2AF65. Sci. Rep. 5, 12521. https://doi.org/10.1038/srep12521 (2015).
Dickey, A. S. et al. PPARdelta activation by bexarotene promotes neuroprotection by restoring bioenergetic and quality control homeostasis. Sci. Transl. Med. https://doi.org/10.1126/scitranslmed.aal2332 (2017).
Reilmann, R. et al. Safety and efficacy of pridopidine in patients with Huntington’s disease (PRIDE-HD): A phase 2, randomised, placebo-controlled, multicentre, dose-ranging study. Lancet Neurol. 18, 165–176. https://doi.org/10.1016/S1474-4422(18)30391-0 (2019).
McGarry, A. et al. A randomized, double-blind, placebo-controlled trial of coenzyme Q10 in Huntington disease. Neurology 88, 152–159. https://doi.org/10.1212/WNL.0000000000003478 (2017).
Horizon Investigators of the Huntington Study Group. European Huntington’s Disease Network. A randomized, double-blind, placebo-controlled study of latrepirdine in patients with mild to moderate Huntington disease. JAMA Neurol. 70, 25–33. https://doi.org/10.1001/2013.jamaneurol.382 (2013).
Tabrizi, S. J., Flower, M. D., Ross, C. A. & Wild, E. J. Huntington disease: New insights into molecular pathogenesis and therapeutic opportunities. Nat. Rev. Neurol. https://doi.org/10.1038/s41582-020-0389-4 (2020).
Tabrizi, S. J. et al. Targeting huntingtin expression in patients with Huntington’s disease. N. Engl. J. Med. 380, 2307–2316. https://doi.org/10.1056/NEJMoa1900907 (2019).
Karimian, A. et al. CRISPR/Cas9 novel therapeutic road for the treatment of neurodegenerative diseases. Life Sci. 259, 118165. https://doi.org/10.1016/j.lfs.2020.118165 (2020).
Chung, D. W., Rudnicki, D. D., Yu, L. & Margolis, R. L. A natural antisense transcript at the Huntington’s disease repeat locus regulates HTT expression. Hum. Mol. Genet. 20, 3467–3477. https://doi.org/10.1093/hmg/ddr263 (2011).
Katayama, S. et al. Antisense transcription in the mammalian transcriptome. Science 309, 1564–1566. https://doi.org/10.1126/science.1112009 (2005).
Fish, J. E. et al. Hypoxia-inducible expression of a natural cis-antisense transcript inhibits endothelial nitric-oxide synthase. J. Biol. Chem. 282, 15652–15666. https://doi.org/10.1074/jbc.M608318200 (2007).
Robb, G. B. et al. Post-transcriptional regulation of endothelial nitric-oxide synthase by an overlapping antisense mRNA transcript. J. Biol. Chem. 279, 37982–37996. https://doi.org/10.1074/jbc.M400271200 (2004).
Vittal, P. et al. ASFMR1 splice variant: A predictor of fragile X-associated tremor/ataxia syndrome. Neurol. Genet. 4, e246. https://doi.org/10.1212/NXG.0000000000000246 (2018).
Modarresi, F. et al. Inhibition of natural antisense transcripts in vivo results in gene-specific transcriptional upregulation. Nat. Biotechnol. 30, 453–459. https://doi.org/10.1038/nbt.2158 (2012).
Hsiao, J. et al. Upregulation of haploinsufficient gene expression in the brain by targeting a long non-coding RNA improves seizure phenotype in a model of Dravet syndrome. EBioMedicine 9, 257–277. https://doi.org/10.1016/j.ebiom.2016.05.011 (2016).
d’Ydewalle, C. et al. The antisense transcript SMN-AS1 regulates SMN expression and is a novel therapeutic target for spinal muscular atrophy. Neuron 93, 66–79. https://doi.org/10.1016/j.neuron.2016.11.033 (2017).
Aronin, N. & DiFiglia, M. Huntingtin-lowering strategies in Huntington’s disease: Antisense oligonucleotides, small RNAs, and gene editing. Mov. Disord. 29, 1455–1461. https://doi.org/10.1002/mds.26020 (2014).
Zeng, X. et al. Screen for modulators of atonal homolog 1 gene expression using notch pathway-relevant gene transcription based cellular assays. PLoS ONE 13, e0207140. https://doi.org/10.1371/journal.pone.0207140 (2018).
Gaun, V. et al. A chemical screen identifies small molecules that regulate hepcidin expression. Blood Cells Mol. Dis. 53, 231–240. https://doi.org/10.1016/j.bcmd.2014.06.002 (2014).
Kawabata, H. et al. A HAMP promoter bioassay system for identifying chemical compounds that modulate hepcidin expression. Exp. Hematol. 43, 404-413.e405. https://doi.org/10.1016/j.exphem.2015.01.005 (2015).
Di Zanni, E., Fornasari, D., Ravazzolo, R., Ceccherini, I. & Bachetti, T. Identification of novel pathways and molecules able to down-regulate PHOX2B gene expression by in vitro drug screening approaches in neuroblastoma cells. Exp. Cell Res. 336, 43–57. https://doi.org/10.1016/j.yexcr.2015.03.025 (2015).
Ratni, H. et al. Discovery of risdiplam, a selective survival of motor neuron-2 (SMN2) gene splicing modifier for the treatment of spinal muscular atrophy (SMA). J. Med. Chem. 61, 6501–6517. https://doi.org/10.1021/acs.jmedchem.8b00741 (2018).
Cheung, A. K. et al. Discovery of small molecule splicing modulators of survival motor neuron-2 (SMN2) for the treatment of spinal muscular atrophy (SMA). J. Med. Chem. 61, 11021–11036. https://doi.org/10.1021/acs.jmedchem.8b01291 (2018).
Mestre, T. A. & Guttman, M. The dawn of a new era for neurodegenerative disorders: Huntington’s disease leading the way. Mov. Disord. 34, 1301–1302. https://doi.org/10.1002/mds.27826 (2019).
Achour, M. et al. Neuronal identity genes regulated by super-enhancers are preferentially down-regulated in the striatum of Huntington’s disease mice. Hum. Mol. Genet. 24, 3481–3496. https://doi.org/10.1093/hmg/ddv099 (2015).
Nucifora, F. C. Jr. et al. Interference by huntingtin and atrophin-1 with cbp-mediated transcription leading to cellular toxicity. Science 291, 2423–2428. https://doi.org/10.1126/science.1056784 (2001).
Singh, A. K., Halder-Sinha, S., Clement, J. P. & Kundu, T. K. Epigenetic modulation by small molecule compounds for neurodegenerative disorders. Pharmacol. Res. 132, 135–148. https://doi.org/10.1016/j.phrs.2018.04.014 (2018).
Trettel, F. et al. Dominant phenotypes produced by the HD mutation in STHdh(Q111) striatal cells. Hum. Mol. Genet. 9, 2799–2809. https://doi.org/10.1093/hmg/9.19.2799 (2000).
Park, I. H. et al. Disease-specific induced pluripotent stem cells. Cell 134, 877–886. https://doi.org/10.1016/j.cell.2008.07.041 (2008).
Dumevska, B., Peura, T., McKernan, R., Goel, D. & Schmidt, U. Derivation of Huntington disease affected Genea020 human embryonic stem cell line. Stem Cell Res. 16, 430–433. https://doi.org/10.1016/j.scr.2016.02.009 (2016).
Pei, F. et al. Connecting neuronal cell protective pathways and drug combinations in a Huntington’s disease model through the application of quantitative systems pharmacology. Sci. Rep. 7, 17803. https://doi.org/10.1038/s41598-017-17378-y (2017).
Inglese, J. et al. Quantitative high-throughput screening: A titration-based approach that efficiently identifies biological activities in large chemical libraries. Proc. Natl. Acad. Sci. U.S.A. 103, 11473–11478. https://doi.org/10.1073/pnas.0604348103 (2006).
Wang, R. et al. Sp1 regulates human huntingtin gene expression. J. Mol. Neurosci. 47, 311–321. https://doi.org/10.1007/s12031-012-9739-z (2012).
Naseri, N. N. et al. Novel metabolic abnormalities in the tricarboxylic acid cycle in peripheral cells from Huntington’s disease patients. PLoS ONE 11, e0160384. https://doi.org/10.1371/journal.pone.0160384 (2016).
Panov, A. V. et al. Early mitochondrial calcium defects in Huntington’s disease are a direct effect of polyglutamines. Nat. Neurosci. 5, 731–736. https://doi.org/10.1038/nn884 (2002).
Sawa, A. et al. Increased apoptosis of Huntington disease lymphoblasts associated with repeat length-dependent mitochondrial depolarization. Nat. Med. 5, 1194–1198. https://doi.org/10.1038/13518 (1999).
Evers, M. M. et al. Targeting several CAG expansion diseases by a single antisense oligonucleotide. PLoS ONE 6, e24308. https://doi.org/10.1371/journal.pone.0024308 (2011).
Naia, L. et al. Comparative mitochondrial-based protective effects of resveratrol and nicotinamide in Huntington’s disease models. Mol. Neurobiol. 54, 5385–5399. https://doi.org/10.1007/s12035-016-0048-3 (2017).
Aiken, C. T., Tobin, A. J. & Schweitzer, E. S. A cell-based screen for drugs to treat Huntington’s disease. Neurobiol. Dis. 16, 546–555. https://doi.org/10.1016/j.nbd.2004.04.001 (2004).
Wang, W. et al. Compounds blocking mutant huntingtin toxicity identified using a Huntington’s disease neuronal cell model. Neurobiol. Dis. 20, 500–508. https://doi.org/10.1016/j.nbd.2005.03.026 (2005).
Thomson, S. B. & Leavitt, B. R. Transcriptional regulation of the huntingtin gene. J. Huntingtons Dis. 7, 289–296. https://doi.org/10.3233/JHD-180331 (2018).
Heiser, V. et al. Identification of benzothiazoles as potential polyglutamine aggregation inhibitors of Huntington’s disease by using an automated filter retardation assay. Proc. Natl. Acad. Sci. U.S.A. 99(Suppl 4), 16400–16406. https://doi.org/10.1073/pnas.182426599 (2002).
Hockly, E. et al. Evaluation of the benzothiazole aggregation inhibitors riluzole and PGL-135 as therapeutics for Huntington’s disease. Neurobiol. Dis. 21, 228–236. https://doi.org/10.1016/j.nbd.2005.07.007 (2006).
Aschner, M. & Kimelberg, H. K. The use of astrocytes in culture as model systems for evaluating neurotoxic-induced-injury. Neurotoxicology 12, 505–517 (1991).
De Simone, U., Caloni, F., Gribaldo, L. & Coccini, T. Human co-culture model of neurons and astrocytes to test acute cytotoxicity of neurotoxic compounds. Int. J. Toxicol. 36, 463–477. https://doi.org/10.1177/1091581817739428 (2017).
Limbad, C. et al. Astrocyte senescence promotes glutamate toxicity in cortical neurons. PLoS ONE 15, e0227887. https://doi.org/10.1371/journal.pone.0227887 (2020).
Malik, N. et al. Compounds with species and cell type specific toxicity identified in a 2000 compound drug screen of neural stem cells and rat mixed cortical neurons. Neurotoxicology 45, 192–200. https://doi.org/10.1016/j.neuro.2014.10.007 (2014).
Garcia, V. J. et al. Huntington’s disease patient-derived astrocytes display electrophysiological impairments and reduced neuronal support. Front. Neurosci. 13, 669. https://doi.org/10.3389/fnins.2019.00669 (2019).
Slanzi, A., Iannoto, G., Rossi, B., Zenaro, E. & Constantin, G. In vitro models of neurodegenerative diseases. Front. Cell Dev. Biol. 8, 328. https://doi.org/10.3389/fcell.2020.00328 (2020).
Neubert, M., Ridder, D. A., Bargiotas, P., Akira, S. & Schwaninger, M. Acute inhibition of TAK1 protects against neuronal death in cerebral ischemia. Cell Death Differ. 18, 1521–1530. https://doi.org/10.1038/cdd.2011.29 (2011).
White, B. J. et al. Protection from cerebral ischemia by inhibition of TGFbeta-activated kinase. Exp. Neurol. 237, 238–245. https://doi.org/10.1016/j.expneurol.2012.05.019 (2012).
Fan, J. et al. P38 MAPK is involved in enhanced NMDA receptor-dependent excitotoxicity in YAC transgenic mouse model of Huntington disease. Neurobiol. Dis. 45, 999–1009. https://doi.org/10.1016/j.nbd.2011.12.019 (2012).
Ahmed, T. et al. Map kinase signaling as therapeutic target for neurodegeneration. Pharmacol. Res. 160, 105090. https://doi.org/10.1016/j.phrs.2020.105090 (2020).
Yu, M. et al. Suppression of MAPK11 or HIPK3 reduces mutant Huntingtin levels in Huntington’s disease models. Cell Res. 27, 1441–1465. https://doi.org/10.1038/cr.2017.113 (2017).
Zhao, Q. et al. HuR stabilizes HTT mRNA via interacting with its exon 11 in a mutant HTT-dependent manner. RNA Biol. 17, 500–516. https://doi.org/10.1080/15476286.2020.1712894 (2020).
Li, P. P. et al. ATXN2-AS, a gene antisense to ATXN2, is associated with spinocerebellar ataxia type 2 and amyotrophic lateral sclerosis. Ann. Neurol. 80, 600–615. https://doi.org/10.1002/ana.24761 (2016).
Rudnicki, D. D. et al. Huntington’s disease–like 2 is associated with CUG repeat-containing RNA foci. Ann. Neurol. 61, 272–282. https://doi.org/10.1002/ana.21081 (2007).
Dhamodharan, V. & Pradeepkumar, P. I. Specific recognition of promoter G-quadruplex DNAs by small molecule ligands and light-up probes. ACS Chem. Biol. 14, 2102–2114. https://doi.org/10.1021/acschembio.9b00475 (2019).
Bahat, A., Lahav, O., Plotnikov, A., Leshkowitz, D. & Dikstein, R. Targeting Spt5-Pol II by small-molecule inhibitors uncouples distinct activities and reveals additional regulatory roles. Mol. Cell 76, 617–631614. https://doi.org/10.1016/j.molcel.2019.08.024 (2019).
Shi, Y. et al. Stabilization of lncRNA GAS5 by a small molecule and its implications in diabetic adipocytes. Cell Chem. Biol. 26, 319-330.e316. https://doi.org/10.1016/j.chembiol.2018.11.012 (2019).
Zhu, B. L. et al. MMP13 inhibition rescues cognitive decline in Alzheimer transgenic mice via BACE1 regulation. Brain 142, 176–192. https://doi.org/10.1093/brain/awy305 (2019).
Acknowledgements
We thank Dr. Samarjit Patnaik for inspiration, Dr. James Inglese for providing the luminescent reporter vector, Qi Sun for assistance with manuscript preparation, the staff of Evotec and Charles River Laboratories for performing hESC and iPSC cell studies, and CHDI, including Drs. Ramee Li, Douglas MacDonald, and Liz Doherty, for supporting this effort. This work was supported by the intramural research program of the National Center for Advancing Translational Sciences, NIH, and by NIH NS08532, NIH NS100783, the Hereditary Disease Foundation, and the ABCD Charitable Trust.
Author information
Authors and Affiliations
Contributions
H.G.K. contributed to assay design and strategy, data collection and analysis, and drafting and revision of the manuscript. H.F. contributed to compound assays, manuscript drafting. X.H. contributed to HTS development, performance, and interpretation. X.S. contributed to the generation and validation of stable cell lines. W.Z. designed and contributed to the generation of stable cell lines. P.P.L. contributed to experimental construct development, compound testing, and experimental design. D.D.R. contributed to experimental and construct design and data analysis. W.Y. implemented the primary HTS and secondary validation assays. Y.C. implemented the HTRF assays in HEK293 cells. N.S. contributed to overall strategy and experimental design. J.M. contributed to overall strategy and experimental design. C.A.R. contributed to strategic design and data interpretation. M.F. contributed to implementation, design, trouble shooting, and data analysis of HTS and HTRF assays, data analysis, and manuscript editing. M.J.H. contributed to experimental design, data analysis, and editing of the manuscript. R.L.M. provided overall supervision for strategy, data analysis, and manuscript drafting. All authors reviewed and approved the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Khaled, H.G., Feng, H., Hu, X. et al. A high-throughput screening to identify small molecules that suppress huntingtin promoter activity or activate huntingtin-antisense promoter activity. Sci Rep 11, 6157 (2021). https://doi.org/10.1038/s41598-021-85279-2
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-021-85279-2
- Springer Nature Limited