Abstract
The growth of skin appendages, such as hair, feathers and scales, depends on terminal differentiation of epidermal keratinocytes. Here, we investigated keratinocyte differentiation in avian scutate scales. Cells were isolated from the skin on the legs of 1-day old chicks and subjected to single-cell transcriptomics. We identified two distinct populations of differentiated keratinocytes. The first population was characterized by mRNAs encoding cysteine-rich keratins and corneous beta-proteins (CBPs), also known as beta-keratins, of the scale type, indicating that these cells form hard scales. The second population of differentiated keratinocytes contained mRNAs encoding cysteine-poor keratins and keratinocyte-type CBPs, suggesting that these cells form the soft interscale epidermis. We raised an antibody against keratin 9-like cysteine-rich 2 (KRT9LC2), which is encoded by an mRNA enriched in the first keratinocyte population. Immunostaining confirmed expression of KRT9LC2 in the suprabasal epidermal layers of scutate scales but not in interscale epidermis. Keratinocyte differentiation in chicken leg skin resembled that in human skin with regard to the transcriptional upregulation of epidermal differentiation complex genes and genes involved in lipid metabolism and transport. In conclusion, this study defines gene expression programs that build scutate scales and interscale epidermis of birds and reveals evolutionarily conserved keratinocyte differentiation genes.
Similar content being viewed by others
Introduction
Keratinocytes of the epidermis form a cornified cell layer at the body surface which protects against water loss and insults from the environments. In coordination with mesenchymal cells, epidermal keratinocytes also form skin appendages, such as claws, hair, feathers and scales, which have important functions in defense, capture of prey, thermoregulation and locomotion1,2,3,4. In birds most parts of the body surface are covered by a soft epidermis which suppresses water loss whereas hard skin appendages, such as the beak, feathers, and claws are used for interactions with the environment that require mechanical resilience5. The lower legs and the toes of birds are covered by scales which can be distinguished into scutate and reticulate scales6,7.
Scutate scales are located on the tarsometatarsus and on the dorsal side of the toes. They consist of overlapping hard scales that are separated by interscale or hinge regions. The structure of scutate scales resembles that of overlapping scales of reptiles. However, the hypothesis that avian scutate scales are homologous to reptilian scales, meaning that they have been inherited from a common ancestor of archosaurs (birds and crocodilians)8,9,10, has been challenged. The alternative hypothesis holds that avian scutate scales are secondarily derived from feathers6,11,12. Like feathers and scales of squamates, avian scutate scales develop from an anatomical placode13,14,15. The epidermal compartment of scutate scale placodes is characterized by the expression of beta-catenin (CTNNB1)13. A patterning mechanism distinct from that of avian scutate scales leads to the development of non-overlapping reticulate scales7.
The protective function of scutate scales depends on their structure and molecular composition. Corneous beta-proteins (CBPs), also known as beta-keratins16, keratins and other proteins were shown to be expressed in mature scutate scales17,18,19,20,21,22,23,24,25,26. A transcriptome analysis of embryonic scutate scales provided information on genome-wide gene expression, but only a subset of selected keratin intermediate filament and CBP genes were localized by mRNA in situ hybridization in hard scales and soft interscale regions27. To the best of our knowledge, a comprehensive gene expression catalog resolving the alternating pattern of soft and hard cornification has not been reported yet. In a recent study, we isolated keratinocytes from chicken leg skin, cultured them in an in vitro model of avian skin and determined their transcriptome28. Differentiation of keratinocytes in this culture system induced the expression of many genes, including members of the keratin family29, that were not expressed in monolayer cultures. However, it remained elusive which genes are expressed in the hard and soft segments of scutate scales in vivo.
Here, we report single-cell RNA-sequencing (scRNA-seq) of chicken leg skin and the characterization of two distinct types of differentiated keratinocytes of scutate scales.
Results
Keratin KRT9LC2 is a marker of differentiated keratinocytes in scutate scales of chickens
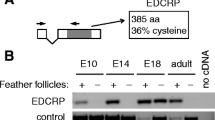
The chicken has a diversified set of keratins29 which are hypothesized to mark specific states of epithelial cell differentiation. Keratin 9-like cysteine-rich 2 (KRT9LC2), also referred to as Hard Acid Sauropsid-specific 2 (HAS2)29, was detected by RT-PCR in scutate scales and analysis of transcriptome data28 suggested that it is transcriptionally upregulated during in vitro differentiation of keratinocytes isolated from chicken leg skin (Fig. 1A). An antibody raised against a carboxy-terminal peptide of the KRT9LC2 protein (Supplementary Fig. S1) detected a prominent band at the predicted size of 51 kD in protein extracts from the stratified epidermis of an in vitro skin model but not in extracts from monolayer cultures of undifferentiated chicken keratinocytes (Fig. 1B). The KRT9LC2 protein was also detected in extracts from scutate scales but not in back skin or reticulate scales of chickens (Fig. 1C).
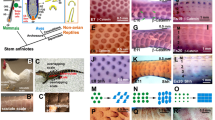
KRT9LC2 is a marker of hard scutate scales of chicken leg skin. (A) Quantitative RT-PCR analysis of KRT9LC2 expression during differentiation of chicken leg keratinocytes (KC) in vitro. RNA was isolated from chicken KCs (n = 3, biological replicates) growing in monolayer culture and in an epidermal model. KRT9LC2 mRNA was quantified by RT-PCR and normalized to the housekeeping gene HMBS. a.u., arbitrary units. (B,C) Western blot analysis of KRT9LC2 and GAPDH in cultured cells (B) and tissues of chicks (C). Bands at predicted sizes are indicated by arrows. The positions of molecular weight markers (kD, kilo-Dalton) are shown of the right. (D,E,G) Immunohistochemical detection of KRT9LC2 (red) in scutate scales (D,E) and reticulate scales (G). A negative control (neg. con.) staining in which the anti-KRT9LC2 antiserum was replaced by non-immune serum is shown in panel (F). Scale bars, 50 µm.
Immunohistochemical staining yielded a strong KRT9LC2 signal in the suprabasal keratinocytes of scutate scales whereas interscale epidermis was not immunostained (Fig. 1D,E). When the primary antibody was replaced by preimmune serum, there was no immunostaining, confirming the absence of unspecific staining (Fig. 1F). The reticulate scales were immunonegative (Fig. 1G). These results demonstrated that KRT9LC2 is a marker of differentiated keratinocytes that form the hard outer surface of scutate scales.
scRNA-seq analysis reveals two distinct populations of differentiated keratinocytes in chicken scutate scales
To characterize gene expression during keratinocyte differentiation in chicken scutate scales, we isolated cells from the legs of 1-day old chicks (n = 3) and subjected them to single-cell RNA-sequencing (scRNA-seq). The protocol was designed to enrich for epidermal keratinocytes but smaller populations of fibroblasts, smooth muscle cells, endothelial cells, Schwann cells and erythrocytes were also detected (Fig. 2A, Supplementary Fig. S2).
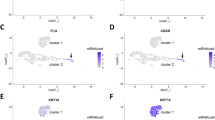
Single-cell RNA-sequencing (scRNA-seq) analysis of scutate scales of chicks. (A) Uniform manifold approximation and projection (UMAP) of cells from chicken scutate scales according to similarity of their transcriptome. Unsupervised clustering resulted in 8 clusters that are indicated by different colors. Cell types were identified by the expression of specific markers, so that 5 unsupervised clusters (KC1-KC5) could be defined as keratinocytes. (B–D) Feature plots showing the expression level of KRT14L1 (B), KRT9L3 (C) and KRT9LC2 (D) in each cell depicted in UMAP plots. The expression levels are color-coded from grey (no expression) to purple (high expression level).
According to nearest neighbor clustering as implemented in Seurat30, keratinocytes were divided into 5 clusters (Supplementary Table S1). Clusters KC1, KC2 and KC3 (Fig. 2A) represented non-differentiated cells, characterized by high expression levels of KRT14L1 (Fig. 2B) 29, which is the avian homolog of KRT14, a marker of the basal epidermal layer in mammals. KRT9L3 (Fig. 2C), a cysteine-poor keratin upregulated during differentiation of chicken keratinocytes in in vitro skin models28, was expressed at high levels in cluster KC4 (Fig. 2A), for which CBP63 was defined as another marker gene (Supplementary Table S1). Expression of CBP63 (GenBank Gene ID: 101751614), previously referred to as “β-keratin, Chr25, Ktn13” 27, was demonstrated by mRNA in situ hybridization in the interscale epidermis of scutate scales 27. Therefore, cluster KC4 contained keratinocytes of the soft interscale epidermis. Cluster KC5 (Fig. 2A) was defined by marker genes such as KRT9LC2 (Supplementary Table S1; Fig. 2D). From the immunolocalization of KRT9LC2 (Fig. 1D) we inferred that KRT9LC2-positive cells represented the hard scales. The clustering of cells was reproduced in the 3 biological replicates investigated in this study (Supplementary Fig. S3).
Keratinocyte differentiation is associated with the expression of distinct genes in scale and interscale segments of chicken scutate scales
To determine genes that are upregulated during keratinocyte differentiation in scutate scales, we compared gene expression in cells containing KRT14L1 transcripts, marking the non-differentiated state of keratinocytes, versus gene expression in cells positive for one or both of the two differentiation markers defined above, i.e. KRT9L3 and KRT9LC2. In KRT9L3-positive cells 219 genes were expressed at higher levels than in KRT14L1-positive cells (Fig. 3A, Supplementary Table S2), and in KRT9LC2-positive cells 213 genes were upregulated (> 0.25 Log2-fold average upregulation, P < 0.001) (Fig. 3B, Supplementary Table S3). The majority of these genes (n = 133), including the type II keratin, KRT78L2, the epidermal differentiation complex gene EDQL (Supplementary Fig. S4), homologs of mammalian keratinocyte differentiation-associated genes, such as DSP, FABP5, POF1B, and others (Supplementary Tables S2 and S3) were upregulated both in KRT9L3-positive and in KRT9LC2-positive cells relative to KRT14L1-positive cells.
Volcano plots of genes differentially expressed in KRT9L3-high versus KRT14L1-high and KRT9LC2-high versus KRT14L1-high cells. Significance (-log10 of adjusted P value) was plotted against Log2 of Fold-change of gene expression levels in KRT9L3-positive versus KRT14L1-positive keratinocytes (A) and KRT9LC2-positive versus KRT14L1-positive keratinocytes (B).
To identify specific markers of hard and soft epidermal differentiation in scutate scales, we compared gene expression levels in KRT9LC2-positive versus KRT9L3-positive cells and determined the genes that differed most strongly with regard to their expression in these cells (Tables 1 and 2, Supplementary Fig. S4). KRT9LC2-positive cells accumulated, amongst others, the cysteine-rich keratin KRT9LC1 (Supplementary Fig. S4B), scale-associated CBPs such as CBP53 (Supplementary Fig. S4F), the lectin LGASL1, and HOPX, whose mammalian ortholog regulates keratinocyte differentiation31 (Table 1). Likewise, CTNNB1, previously reported as a regulator of scutate scale development, was found enriched in differentiated keratinocytes of hard scales (Table 1). KRT9L3-positive cells accumulated cysteine-poor keratin KRT9L4 (Supplementary Fig. S4A), keratinocyte-associated CBPs such as CBP62 and CBP63 (Supplementary Fig. S4E), the lectin LGAS1, and PRDX1, an antioxidant enzyme32 (Table 2). Thus, the results of this study suggest catalogues of genes associated with keratinocyte differentiation in hard epidermal segments (Table 1) and genes associated with keratinocyte differentiation in soft interscale segments (Table 2) of chicken scutate scales.
Discussion
Differentiation of keratinocytes underlies the growth of epithelial skin structures, such as claws, hair, feathers and scales of mammals, reptiles and birds. The molecular control of keratinocyte differentiation is well characterized for mammalian interfollicular epidermis and skin appendages33,34,35, whereas little is known about the genetic regulation of keratinocyte differentiation in sauropsids. The results of the present study shed light into keratinocyte differentiation in scutate scales and provide a basis for the comparative analysis of further epithelial cell differentiation processes in avian claws, beak and feathers.
We have used scRNA-seq to characterize two types of keratinocyte differentiation, leading to the hard outer surface and the soft interscale epidermis of scutate scales. The methodology was adapted from successful scRNA-seq analyses of human and mouse skin36,37,38,39. scRNA-seq of mouse tail skin revealed two paths of keratinocyte differentiation into scale and interscale epidermis40. Of note, hard scales of the mouse tail were found to contain transcripts of cysteine-rich keratins such as KRT31 whereas the soft interscale regions contained epidermal keratins such as KRT10 and epidermal differentiation complex (EDC) genes such as involucrin40,41. In contrast to the availability of many antibodies against mouse keratinocyte proteins, we had only one antibody, anti-KRT9LC2, specific for a keratin expressed in chicken scutate scales. This was a significant limitation of the present study. We were able to ascertain the expression of KRT9LC2 in differentiated keratinocytes of hard scales, and mRNA in situ hybridization data published by Wu et al. 2015 supported the expression of CBP63 in interscale epidermis27. However, other putative differentiation markers, that are suggested by our results, remain to be localized in situ in future studies.
Gene expression in interscale epidermis of chicken leg skin showed several similarities to gene expression in two models of chicken epidermis with a soft cornified layer28. scRNA-seq analysis of chicken back skin and bulk RNA-seq analysis of an organotypic model of chicken skin revealed expression of EDC genes and cysteine-poor but not cysteine-rich keratins28. Many of the genes upregulated during differentiation of back skin keratinocytes28, such as KRT9L4, LOR1, KRT9L3, BDH1L, EDQM2, SPTSSB, EDQM1, AADACL4B, LIPML2, and ELOVL4 (Table 2) were enriched in interscale versus scale epidermis. Conversely, genes enriched in hard scale epidermis, such as KRT9LC2, KRT9LC1, CBP53-S, CBP54-S, CBP55-S, EDMTF1, and MT4 (Table 1) were not upregulated during differentiation of back skin keratinocytes28. Therefore, we conclude that the genetic program of keratinocyte differentiation in the soft interscale epidermis of scutate scales is similar to the keratinocyte differentiation program in the soft back epidermis.
Keratinocyte differentiation in the hard outer surface of scutate scales differs substantially from that in the interscale regions. The results of the present study establish KRT9LC2, also referred to as HAS2 keratin29, as a protein marker of the hard scutate scales and identify other genes that are co-regulated with KRT9LC2. Among these scale-associated genes was KRT9LC1 (GenBank Gene ID:772,080) (Supplementary Fig. S3B), also referred to as Hard Acidic Sauropsid-specific 1 (HAS1) keratin29. In situ hybridization of transcripts corresponding to this gene (then named KRT13A) demonstrated predominant expression in the outer surface epithelium of scutate scales27, thus validating our scRNA-seq data. Another gene co-expressed with KRT9LC2 was CBP55-S, a scale CBP (beta-keratin). In the aforementioned study of Wu and colleagues27, expression of this gene (then named β-keratin, Chr25, Scale18) was detected by in situ hybridization specifically in the outer surface epithelium of scutate scales, further validating our scRNA-seq data.
An important result of this study is the genome-wide gene expression catalog of scutate scale epidermis resolved at the single-cell level. In addition to the genes discussed above, many more genes with differentiation-dependent expression were identified both in soft interscale epidermis (Supplementary Table S2) and hard scales (Supplementary Table S3). With regard to ongoing efforts to characterize evolutionarily ancient and derived patterns of gene expression during epidermal keratinocyte differentiation28,29,42,43,44,45,46,47,48, the results of the present study support the hypothesis that EDC genes, anti-inflammatory interleukin 1 family cytokines (IL-36RN and IL-1RN) (Supplementary Tables S2 and S3; Supplementary Fig. S5) and lipid metabolism and lipid transport-related genes, such as FABP5 and GLTP28 belong to the common keratinocyte differentiation program of amniotes. The transcriptome data generated in this study will be particularly useful for characterizing the process of hard cornification in a non-mammalian model species. The single-cell transcriptomes of chicken leg skin are now accessible for data searches according to criteria not limited to keratinocyte differentiation, so that new research questions pertaining to avian skin biology can be addressed in future studies.
Methods
Tissue preparation and scRNA-seq analysis
One day old chicks (strain Lohmann) were obtained from Schropper GmbH, Gloggnitz, Austria. Skin was excised from the leg of sacrificed animals and incubated in thermolysin (0.5 mg/ml) (Sigma-Aldrich) at 37 °C for 45 min. The lower dermis was removed using forceps and the remaining tissue, including the epidermis and parts of the upper dermis, was processed further according to protocols established for human skin39,49. For the isolation of cells, the tissue was split into two fractions that were incubated either in buffer-enzyme mix of the Whole Skin Dissociation Kit human (MACS Milteny Biotech) for 2.5 h at 37 °C or with 0.05% trypsin/EDTA (Thermo Fisher Scientific) and DNase 1 (10 µg/ml) (Roche Diagnostics) at 37 °C for 15 min. Afterwards the samples were combined and processed according to the manufacturer’s protocol (Whole Skin Dissociation Kit human, MACS Milteny Biotech). In brief, the epidermis-enzyme mix was diluted in 0.5 ml medium and dissociated with the gentleMACS Dissociator. The ground tissue was filtered through 100-micron (Falcon) and 40-micron (Falcon) meshes. Subsequently, cells were stained with DAPI dye for 10s and viable cells were sorted via an AriaFusion high-speed cell sorting device (BD Biosciences, San Jose, CA, USA). Single cell RNA sequencing was performed according to a published protocol49. In brief, a 10× Genomics Chromium instrument (10× Genomics, Pleasanton, CA) was used for single cell partitioning and barcoding and Illumina HiSeq 3000/4000 (Illumina, San Diego, CA) was used for sequencing (Center for Molecular Medicine, Vienna, Austria). Using the CellRanger Fastq pipeline (10X Genomics) the demultiplexed raw sequencing data were aligned to the chicken reference genome Gallus_gallus-5.0.
Analysis of scRNA-sequencing data
We distinguished between background noise and droplets containing cells using emptyDrops50. Briefly, this method models ambient RNA background in the data set and tests for deviations from the background RNA. We used a false discovery rate of 0.05 to call cells to be included into further analysis. On the other end of the spectrum we used scran package to remove droplets containing more than one cell51. The applied approach simulates thousands of doublets by adding together two randomly chosen single cell profiles. For the doublet score calculation, cell clustering including the set randomly generated doublets is performed. Then for each cell of the original dataset, the number of simulated doublets in their neighbourhood is recoded and used as input for score calculation. We used 200 nearest neighbours for each cell and applied a threshold of doublet score > 4 to identify doublets in each dataset separately. Doublet score was log10 of the ratio between simulated doublet cells and total number of neighbours taken into consideration for each cell. The data obtained from 3 biological replicates were submitted to the NCBI Gene Expression Omnibus (GEO) database under accession numbers GSE179690 (BioProject PRJNA744554). The individual samples were referred to as “leg skin 1” (BioSample: SAMN20109848, SRA: SRX11375855), “leg skin 2” (BioSample: SAMN20109847, SRA: SRX11375856) and “leg skin 3” (BioSample: SAMN20109846, SRA: SRX11375857).
Subsequent to individual quality control of samples, raw read counts of across datasets were combined to one count matrix. Reads from 11,779 cells were included in the final analysis. We used Pearson residuals derived from a generalized negative binomial model of UMI counts as input for principal component analysis and differential gene expression analysis. This approach is implemented in the R package Seurat as sctransform52. Apart from cellular sequencing depth, which is added by default to the regression model, we also adjusted for mitochondrial RNA content, batch, and cell cycle score. Calculation of cell cycle scores was performed as implemented in Seurat package where gene expression of cell cycle marker genes are combined to score. The score consisted of 43 genes primarily expressed in G1/S and 55 primarily expressed in G2/M53. We removed cells with a mitochondrial RNA content above 15%. Batch correction between individual datasets was done as part of the principal component analysis using the Harmony algorithm54. Harmony starts off with user supplied information about batch, then uses fuzzy clustering to assign each cell to multiple clusters in a way that batch diversity in each cluster is maximized. To get a correction factor for each cell, global and batch-specific centroids are calculated for each cluster. Data are corrected for batch and the procedure is repeated until convergence of global and batch-specific centroids54. We used PC 1–15 for subsequent UMAP analysis and nearest neighbor based clustering at a resolution of 0.2.
For further analysis of differentially expressed genes (DEGs), three different clusters, characterized by a certain expression threshold of marker genes, e.g. KRT9L3 > 2, KRT9LC2 > 1.5 and KRT14L1 > 1.5, were created. Gene expression levels in different clusters were compared and adjusted P values were calculated according to the standard algorithm implemented in Seurat30.
Quantitative reverse-transcription polymerase chain reaction
RNA was isolated from chicken tissues and skin models28 and purified with TriFast according to a published protocol46 and reverse-transcribed with the iScriptTM cDNA synthesis kit (Biorad, Hercules, CA). Polymerase chain reactions (PCRs) were performed with primer pairs specific for KRT9LC2 (KRT9LC2-s, 5′-GAATGCCGCTACAACAACCAC-3′ and KRT9LC2-a, 5′-TGCTTCAGGGATCTCTCATTG-3′), IL1RN (IL1RN-s, 5′-GAGAAGGTGTTTTGGGTGCC-3′ and IL1RN-a, 5′-TAGGTGCGGAAGAAGGTGAA-3′), IL36RN (IL36RN-s, 5′-GAGCTCAGCCGTACCACTAC-3′ and IL36RN-a, 5′-AACAGCTTCACCTCCTCCAG-3′), and the housekeeping gene Hydroxymethylbilane synthase (HMBS) (HMBS-s, 5′-AACTGTGGGAAAACGCTCAG-3′ and HMBS-a, 5′-TTCTCTTCAGTCCAGCAGCA-3′) on a Roche LightCycler with LC480 SYBR Green I Master Kit according to the manufacturer’s protocol. Quantitative analysis of IL1RN and IL36RN expression in chicken tissues was performed according to a published method46. The expression levels of these genes were compared between scutate scales and other tissues, considering differences with a P value of < 0.05 significant (two-sided t-test).
Western blot analysis
Proteins were prepared from chicken skin and scales by treatment with the Precellys system (VWR, International, Radnor, PA) and from chicken keratinocytes cultured in vitro28 by sonication in Laemmli buffer containing 2% SDS. Thirty µg of protein per lane were electrophoresed through an ExcelGel SDS 8–18% polyacrylamide gel (GE Healthcare Life Sciences) and afterwards blotted onto a nitrocellulose membrane (GVS Life Sciences). Subsequently, the membrane was blocked with phosphate-buffered saline containing 5% milk powder (Sigma-Aldrich), 2% bovine serum albumin (Sigma-Aldrich) and 0.1% Tween (Sigma-Aldrich) at room temperature for one hour, and incubated with mouse anti-KRT9LC2 antibody (1:500) that was raised in mice by immunization with a synthetic peptide CAAAEIQVPCRRICD, corresponding to the carboxy-terminus of the protein (GenBank accession number XP_418162.6, GenBank definition: keratin, type I cytoskeletal 19) (Supplementary Fig. S1) conjugated to keyhole limpet hemocyanin, according to a published protocol 55. After overnight incubation at 4 °C, the membrane was washed and sheep anti-mouse immunoglobulin G (1:10,000, GE Healthcare UK Limited) coupled with horseradish peroxidase used as secondary antibody at room temperature for one hour. The chemiluminescence system (Clarity Western ECL Substrate, BioRad) served for the protein detection. For loading control, the membrane was reincubated with anti-mouse GAPDH (1:5000, HyTest) and sheep anti-mouse immunoglobulin G (1:10,000, GE Healthcare UK Limited), coupled with horseradish peroxidase. The recordings of the chemiluminescence signal over the entire blots are shown in Supplementary Fig. S6 and the relevant portions thereof are depicted in Fig. 1C,D.
Immunohistochemistry
Immunohistochemistry was performed according to published protocols with modifications46. In brief, chicken tissue samples were fixed in 7.5% formaldehyde and embedded in paraffin. Citrate buffer pH6 (DAKO) was used to retrieve the antigens and mouse anti-KRT9LC2 antibody (1:500) as primary antibody. To block unspecific binding, 10% sheep serum was added to secondary sheep anti-mouse immunoglobulin G (GE Healthcare), and further the nuclei were counterstained with haematoxylin. For control experiments, the primary antibody was replaced by the pre-immune serum.
Ethics statement
All animal procedures were approved by the Animal Care and Use Committee of the Medical University of Vienna. All procedures were performed in accordance with the guidelines established by this committee and in adherence to the ARRIVE guidelines 56.
Data availability
Single-cell transcriptomes generated in this study are available at GEO under accession number GSE179690. All other data generated or analysed during this study are included in this published article and its Supplementary Information files.
References
Alibardi, L. Adaptation to the land: The skin of reptiles in comparison to that of amphibians and endotherm amniotes. J. Exp. Zool. B Mol. Dev. Evol. 298, 12–41 (2003).
Wu, P. et al. Evo-Devo of amniote integuments and appendages. Int. J. Dev. Biol. 48, 249–270 (2004).
Dhouailly, D. A new scenario for the evolutionary origin of hair, feather, and avian scales. J. Anat. 214, 587–606 (2009).
Chang, C. et al. Reptile scale paradigm: Evo-Devo, pattern formation and regeneration. Int. J. Dev. Biol. 53, 813–826 (2009).
Sawyer, R.H., Knapp, L.W. & O’Guin, W.M. The skin of birds. Epidermis, dermis and appendages. In: Biology of the Integument (ed. Bereiter-Hahn, J., Matoltsy, A.G. & Richards, K.S.) 194–238 (Springer, 1986).
Prin, F. & Dhouailly, D. How and when the regional competence of chick epidermis is established: feathers vs. scutate and reticulate scales, a problem en route to a solution. Int. J. Dev. Biol. 48, 137–148 (2004).
Cooper, R. L. et al. Conserved gene signalling and a derived patterning mechanism underlie the development of avian footpad scales. EvoDevo 10, 19 (2019).
Alibardi, L. & Thompson, M. B. Fine structure of the developing epidermis in the embryo of the American alligator (Alligator mississippiensis, Crocodilia, Reptilia). J. Anat. 198, 265–282 (2001).
Sawyer, R. H., Rogers, L., Washington, L., Glenn, T. C. & Knapp, L. W. Evolutionary origin of the feather epidermis. Dev. Dyn. 232, 256–267 (2005).
Alibardi, L., Knapp, L. W. & Sawyer, R. H. Beta-keratin localization in developing alligator scales and feathers in relation to the development and evolution of feathers. J. Submicrosc. Cytol. Pathol. 38, 175–192 (2006).
Wu, P., Lai, Y. C., Widelitz, R. & Chuong, C. M. Comprehensive molecular and cellular studies suggest avian scutate scales are secondarily derived from feathers, and more distant from reptilian scales. Sci. Rep. 8, 16766 (2018).
Wu, P. et al. Multiple regulatory modules are required for scale-to-feather conversion. Mol. Biol. Evol. 35, 417–430 (2018).
Musser, J. M., Wagner, G. P. & Prum, R. O. Nuclear β-catenin localization supports homology of feathers, avian scutate scales, and alligator scales in early development. Evol. Dev. 17, 185–194 (2015).
Di-Poï, N. & Milinkovitch, M. C. The anatomical placode in reptile scale morphogenesis indicates shared ancestry among skin appendages in amniotes. Sci. Adv. 2, e1600708 (2016).
Lai, Y. C. et al. Transcriptome analyses of reprogrammed feather / scale chimeric explants revealed co-expressed epithelial gene networks during organ specification. BMC Genomics 19, 780 (2018).
Holthaus, K. B., Eckhart, L., Dalla Valle, L. & Alibardi, L. Review: Evolution and diversification of corneous beta-proteins, the characteristic epidermal proteins of reptiles and birds. J. Exp. Zool. B Mol. Dev. Evol. 330, 438–453 (2018).
Shames, R. B. & Sawyer, R. H. Expression of beta keratin genes during skin development in normal and sc/sc chick embryos. Dev. Biol. 116, 15–22 (1986).
Shames, R. B., Jennings, A. G. & Sawyer, R. H. The initial expression and patterned appearance of tenascin in scutate scales is absent from the dermis of the scaleless (sc/sc) chicken. Dev. Biol. 147, 174–186 (1991).
Shames, R. B., Knapp, L. W., Carver, W. E., Washington, L. D. & Sawyer, R. H. Keratinization of the outer surface of the avian scutate scale: interrelationship of alpha and beta keratin filaments in a cornifying tissue. Cell Tissue Res. 257, 85–92 (1989).
Carver, W. E. & Sawyer, R. H. Avian scale development: XI. Immunoelectron microscopic localization of alpha and beta keratins in the scutate scale. J Morphol. 195, 31–43 (1988).
Knapp, L. W., Linser, P. J., Carver, W. E. & Sawyer, R. H. Biochemical identification and immunological localization of two non-keratin polypeptides associated with the terminal differentiation of avian scale epidermis. Cell Tissue Res. 265, 535–545 (1991).
Knapp, L. W., Shames, R. B., Barnes, G. L. & Sawyer, R. H. Region-specific patterns of beta keratin expression during avian skin development. Dev. Dyn. 196, 283–290 (1993).
Sawyer, R. H. et al. The expression of beta (β) keratins in the epidermal appendages of reptiles and birds. Am. Zool. 40, 530–539 (2000).
Sawyer, R. H. et al. Origin of feathers: Feather beta (beta) keratins are expressed in discrete epidermal cell populations of embryonic scutate scales. J. Exp. Zool. B Mol. Dev. Evol. 295, 12–24 (2003).
Alibardi, L. Immunocytochemical and autoradiographic studies on the process of keratinization in avian epidermis suggests absence of keratohyalin. J. Morphol. 259, 238–253 (2004).
Zeltinger, J. & Sawyer, R. H. Avian scale development. XVI. Epidermal commitment to terminal differentiation is prior to definitive scale ridge formation. Dev. Biol. 149, 55–65 (1992).
Wu, P. et al. Topographical mapping of α- and β-keratins on developing chicken skin integuments: Functional interaction and evolutionary perspectives. Proc. Natl. Acad. Sci. USA 112, E6770–E6779 (2015).
Lachner, J. et al. An in vitro model of avian skin reveals evolutionarily conserved transcriptional regulation of epidermal barrier formation. J. Invest. Dermatol. 141, 2829–2837 (2021).
Ehrlich, F., Lachner, J., Hermann, M., Tschachler, E. & Eckhart, L. Convergent evolution of cysteine-rich keratins in hard skin appendages of terrestrial vertebrates. Mol. Biol. Evol. 37, 982–993 (2020).
Satija, R., Farrell, J. A., Gennert, D., Schier, A. F. & Regev, A. Spatial reconstruction of single-cell gene expression data. Nat. Biotechnol. 33, 495–502 (2015).
Mariotto, A., Pavlova, O., Park, H. S., Huber, M. & Hohl, D. HOPX: The unusual homeodomain-containing protein. J. Invest. Dermatol. 136, 905–911 (2016).
Ledgerwood, E. C., Marshall, J. W. & Weijman, J. F. The role of peroxiredoxin 1 in redox sensing and transducing. Arch. Biochem. Biophys. 617, 60–67 (2017).
Fuchs, E. Scratching the surface of skin development. Nature 445, 834–842 (2007).
Matsui, T. & Amagai, M. Dissecting the formation, structure and barrier function of the stratum corneum. Int. Immunol. 27, 269–280 (2015).
Rice, G. & Rompolas, P. Advances in resolving the heterogeneity and dynamics of keratinocyte differentiation. Curr. Opin. Cell Biol. 67, 92–98 (2020).
Joost, S. et al. Single-cell transcriptomics reveals that differentiation and spatial signatures shape epidermal and hair follicle heterogeneity. Cell Syst. 3, 221–237 (2016).
Cheng, J. B. et al. Transcriptional programming of normal and inflamed human epidermis at single-cell resolution. Cell Rep. 25, 871–883 (2018).
Dubois, A., Gopee, N., Olabi, B. & Haniffa, M. Defining the skin cellular community using single-cell genomics to advance precision medicine. J. Invest. Dermatol. 41, 255–264 (2021).
Kalinina, P. et al. The whey acidic protein WFDC12 is specifically expressed in terminally differentiated keratinocytes and regulates epidermal serine protease activity. J. Invest. Dermatol. 141, 1198–1206 (2021).
Dekoninck, S. et al. Defining the design principles of skin epidermis postnatal growth. Cell 181, 604–620 (2020).
Gomez, C. et al. The interfollicular epidermis of adult mouse tail comprises two distinct cell lineages that are differentially regulated by Wnt, Edaradd, and Lrig1. Stem Cell Rep. 1, 19–27 (2013).
Strasser, B. et al. Evolutionary origin and diversification of epidermal barrier proteins in amniotes. Mol. Biol. Evol. 31, 3194–3205 (2014).
Mlitz, V. et al. Trichohyalin-like proteins have evolutionarily conserved roles in the morphogenesis of skin appendages. J. Invest. Dermatol. 134, 2685–2692 (2014).
Lachner, J. et al. Immunolocalization and phylogenetic profiling of the feather protein with the highest cysteine content. Protoplasma 256, 1257–1265 (2019).
Holthaus, K. B. et al. Comparative genomics identifies epidermal proteins associated with the evolution of the turtle shell. Mol. Biol. Evol. 33, 726–737 (2016).
Lachner, J., Mlitz, V., Tschachler, E. & Eckhart, L. Epidermal cornification is preceded by the expression of a keratinocyte-specific set of pyroptosis-related genes. Sci. Rep. 7, 17446 (2017).
Davis, A. & Greenwold, M. J. Evolution of an Epidermal Differentiation Complex (EDC) gene family in birds. Genes (Basel) 12, 767 (2021).
Lin, G. W. et al. Regional specific differentiation of integumentary organs: regulation of gene clusters within the avian epidermal differentiation complex and impacts of SATB2 overexpression. Genes (Basel) 12, 1291 (2021).
Vorstandlechner, V. et al. Deciphering the functional heterogeneity of skin fibroblasts using single-cell RNA sequencing. FASEB J. 34, 3677–3692 (2020).
Lun, A. T. L. et al. EmptyDrops: distinguishing cells from empty droplets in droplet-based single-cell RNA sequencing data. Genome Biol. 20, 63 (2019).
Lun, A.T., McCarthy, D.J. & Marioni, J.C. A step-by-step workflow for low-level analysis of single-cell RNA-seq data with Bioconductor. F1000Res. 5, 2122 (2016).
Hafemeister, C. & Satija, R. Normalization and variance stabilization of single-cell RNA-seq data using regularized negative binomial regression. Genome Biol. 20, 296 (2019).
Tirosh, I. et al. Dissecting the multicellular ecosystem of metastatic melanoma by single-cell RNA-seq. Science 352, 189–196 (2016).
Korsunsky, I. et al. Fast, sensitive and accurate integration of single-cell data with Harmony. Nat Methods 16, 1289–1296 (2019).
Eckhart, L. et al. Identification of reptilian genes encoding hair keratin-like proteins suggests a new scenario for the evolutionary origin of hair. Proc Natl Acad Sci USA 105, 18419–18423 (2008).
Kilkenny, C., Browne, W. J., Cuthill, I. C., Emerson, M. & Altman, D. G. Improving bioscience research reporting: the ARRIVE guidelines for reporting animal research. PLoS Biol. 8, e1000412 (2010).
Acknowledgements
We thank Polina Kalinina, Vera Vorstandlechner, Dragan Copic and Michael Mildner for helpful discussions and Bahar Golabi for excellent technical support. This work was funded by the Austrian Science Fund (FWF): P28004 and P32777.
Author information
Authors and Affiliations
Contributions
J.L., E.T. and L.E. conceived the study, J.L., F.E. and M.H. performed experiments, J.L., F.E., M.W., M.F. and L.E. analyzed the results, J.L. and L.E. wrote the manuscript. All authors reviewed the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Lachner, J., Ehrlich, F., Wielscher, M. et al. Single-cell transcriptomics defines keratinocyte differentiation in avian scutate scales. Sci Rep 12, 126 (2022). https://doi.org/10.1038/s41598-021-04082-1
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-021-04082-1
- Springer Nature Limited
This article is cited by
-
Interactions between avian viruses and skin in farm birds
Veterinary Research (2024)
-
Comparative genomics of monotremes provides insights into the early evolution of mammalian epidermal differentiation genes
Scientific Reports (2024)