Abstract
The long-term survival of a species requires, among other things, gene flow between populations. Approaches for the evaluation of fragmentation in the frame of freshwater habitats consider only a small amount of the information that combined demography and geography are currently able to provide. This study addresses two species of Austropotamobius crayfish in the light of population genetics, spatial ecology and protected areas of the Carpathians. Advancing the classical approaches, we defined ecological distances upon the rasterised river network as a surrogate of habitat resistance to migration, quantifying the deviations from the species´ suitability range for a set of relevant geospatial variables in each cell of the network. Molecular analyses revealed the populations of the two Austropotamobius crayfish species are clearly distinct, lacking hybridisation. Comparing pairs of populations, we found, in some cases, a strong disagreement regarding genetic and ecological distances, potentially due to human-mediated translocations or the geophysical phenomena of regressive erosion, which may have led to unexpected colonisation routes. Protected areas were found to offer appropriate local habitat conditions but failed to ensure connectivity. The methodology applied in this study allowed us to quantify the contribution of each geospatial (environmental) variable to the overall effect of fragmentation, and we found that water quality was the most important variable. A multilevel approach proved to reveal a better understanding of drivers behind the distribution patterns, which can lead to more adequate conservation measures.
Similar content being viewed by others
Introduction
For species with insular distributions, conservation measures through the establishment of Protected Areas (PAs) are only partially useful in the long run, as these measures usually fail to ensure the connectivity between populations1,2. Connectivity for freshwater organisms depends on the physical distance between the populations on the river path as well as the quality of habitats this path traverses3. The success of efforts to establish corridors for migration, especially in case of rare and sensitive species, is substantially dependent on the accuracy of ecological assessment methods employed4,5. Fragmentation is one of the most challenging issues in conservation biology because its assessment is based on the evaluation of a wide array of interlocking factors4,6. Current techniques for molecular analysis can accurately pinpoint the degree of separation between populations, but the tools for measuring the ecological fragmentation within hydrological networks are still underdeveloped.
In the context of increasing global human demand7,8 and pressure on freshwater systems9,10, the future of biodiversity is marked by the question whether PAs are able to face the next challenges in ecosystems services11. Good management of PAs should focus, among others, on minimising the loss of priority species and their habitat12,13 and restoring the connectivity of populations1,14. Many crayfish species are protected as ecologically sensitive, requiring long-term habitat stability15,16. The most sensitive indigenous European crayfish species in terms of ecological and conservation requirements (see e.g. ref.17) are the Austropotamobius species: the white-clawed crayfish A. pallipes sensu lato (Lereboullet, 1858) and the stone crayfish A. torrentium (von Paula Schrank, 1803). Recently, a new Austropotamobius species, namely the idle crayfish A. bihariensis18,20, was described by phylogenetic and taxonomic evidence. Both A. torrentium and A. bihariensis reside in the Romanian Carpathians19,20 and prefer habitats with clean waters in the mountainous and submountainous regions18,21. The range of A. torrentium19 comprises Central Europe and the Balkans, and the metapopulation in Romania is representative for this species as it consists of its most common haplogroup20,22. Austropotamobius bihariensis is an endemic species restricted to the Apuseni Mountains, Romania18.
Multilevel approaches are expected to reveal the relationship between species’ biology and their habitats more accurately23,24,25. Large-scale evaluations in spatial ecology require remotely computable variables rather than pointwise field measurements26,27. In our assessment of fragmentation, we started from the idea that, in order for two populations to exchange genes, the individuals need to travel through a sector of river with characteristics that may or may not conform to the species’ ecological requirements2. Working on a rasterised river network, we introduced ecological distances, a surrogate of habitat resistance to migration widely used in landscape ecology. The ecological distance between a pair of populations is determined by the deviation from the species suitability range in each cell of the network pathway, for a relevant set of spatial variables. Coupled with population genetics data, this allowed us to obtain a better understanding of the distribution of crayfish and the efficiency of the PAs for their conservation.
Methods
Field data
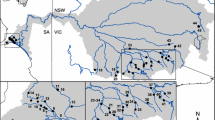
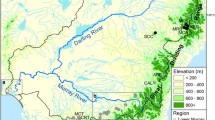
Survey datasets of presence and absence of A. torrentium and A. bihariensis in the Romanian Carpathians were obtained from recent published papers18,21,28, totalling 274 sites (126 presences and 148 absences, Fig. 1). We considered a population to be all the individuals from one stream, and therefore sampled accordingly. The sampling locations were randomly selected, considering the species’ distributions. In situ, the small tributaries were investigated by hand sampling from the riverbed by thoroughly checking galleries in the riverbanks and spaces between or beneath rocks and roots, comprising approximately 200 m of river stretch, while large rivers where inspected using baited traps (Pirate type, with double entrance), left over night. Local residents were also asked about the presence of crayfish, but this information was considered only if it was confirmed in the field. All the sites used in this study were visited at least twice over the past years, confirming the presence or absence of targeted crayfish. Out of the 274 sites, 50.7% were situated in PAs. Additionally, we sampled 411 individuals from 23 sites (at least 10 individuals from each site) for further genetic analyses (see Fig. 1).
Distribution map of sampling sites in the investigated area. The map was generated by using the ArcGIS version 10.4 software (https://www.esri.com) and designed in Inkscape 0.92.4 (https://inkscape.org).
Sample collection for the present study did not significantly affect the wild animals (according to ref.29). After one segment from the last pair of walking legs from each individual was detached and stored in 96% ethanol, the animals were released back into their habitat, in compliance with national and international ethical guidelines. This study was approved by the Romanian Academy, Commission for Monuments of Nature Protection (decision no. 3131/08.04.2009), and additionally, licences were granted from the legal administrators of Natura 2000 sites in the investigated area.
The raster of spatial variables
Four GIS-derived environmental variables (see below) were selected based on existing literature as relevant for A. torrentium ecology (e.g. refs.30,31,32). The SRTM V3 void filled 33 digital elevation model (DEM) at a 100-m resolution was processed and used for the extraction of the river network of the analysed area in ArcGIS version 10.4 (ArcMap; Environmental Systems Resource Institute, Redlands, California). Each cell (100 × 100 m) of the river network was populated with the values of the four selected geospatial variables as follows.
We used RIVERenhancer to remotely estimate water quality by using the variable RWQ as defined by ref.34, based on land cover data extracted from remote sensing images. Considering that each cell in the river raster collects environmental influences from its upstream drainage basin, RWQ is defined using the Corine Land Cover (version 18.5.1; https://land.copernicus.eu) categories found in the catchment area of each cell in the river network grouped by anthropogenic influence potential35 and weighted by their respective areas in the catchment34. Flash-flood potential (FFP) was used to quantify the riverbed stability32; it estimates the potential for flood-related disturbance to streams by taking into account the potential drainage velocity both upstream and at the site. This variable was calculated as a product of two land-surface variables, namely slope gradient and catchment slope, derived from the DEM. The multiannual mean temperature (MMT) was extracted from WorldClim global climate layers (www.worldclim.org) raster datasets of 1-km spatial resolution, resampled at 100 m. The altitude (ALT) was extracted directly from the DEM.
Statistics for the assessment of ecological fragmentation
As seen in Table 1, A. bihariensis appears to occupy a slightly narrower ecological niche than A. torrentium; its occupied range based on field observations was fully included in that of A. torrentium for all the considered variables, but no statistically significant differences were found between the species regarding the distribution of any of the environmental variables (Mann–Whitney, p > 0.1 in all cases). For this reason, and considering the fact that until very recently18, the two Austropotamobius species were regarded as being one and the same, their records were analysed in a single dataset. The presence/absence data were used to establish the most suitable range of each variable for the species distribution. The values of these characteristics in each point of the raster river network situated outside the species suitability range imposed an “ecological cost”, i.e. a stress that the travelling individuals would have to face (see e.g. ref.36). An “ecological distance” between pairs of populations was defined by summing the ecological costs along the river path connecting them.
Specifically, a Random Forest classification model was fit to the species presence/absence data, considering RWQ, FFP, ALT and MMT as predictors. We used the randomForest package37, version 4.6-12, of the R software38. In total, 5,000 trees were generated, with two predictors randomly chosen for each split. The cutoff probability for discrimination between classes was determined so that predicted species prevalence would equal observed prevalence39 and set at 0.35. Variable importance was assessed by the amount that the Gini index decreased by splits over the respective predictor, averaged over all the trees (Fig. S1). Response curves for habitat variables were obtained as partial dependence plots, illustrating the effect of the selected variable on presence probability after integrating out the other predictors. A species suitability range was then computed for each variable from the response curve, as the range where the predicted probability of presence exceeded the class discrimination threshold. This suitability range was validated against the species-occupied range (as observed in the field), and the two were found to be in high agreement (see Table 1).
Next, the full range of each variable across the analysed river network was linearly mapped to the interval [0,1]. The transformation was performed in order to assign similar importance to each variable in further cost computation. A cost per variable was computed in each cell of the river network as the distance between the scaled value of the variable in that cell and the scaled suitability range. The ecological cost for a cell x was then computed as
where the summation is made over all predictor variables i (i.e. the ecological cost is the Euclidean distance from the four-dimensional point defined by the scaled values of the habitat variables in that cell to the hyper-rectangle given by the scaled suitability ranges). The considered predictors are of comparable importance in the model (see Fig. S1) and were therefore weighed equally in the computation of costs. The cumulative ecological cost on a path connecting a pair of populations defines the “ecological distance” between those populations.
Spatial data management
We used the network analyst extension in ArcGIS version 10.4 to connect each of the 126 crayfish populations via the hydrological network to calculate the origin–destination matrix for the ecological distance (see Fig. 2A). Hierarchical clustering on the ecological distance matrix was performed using Ward's method to identify population clusters. Visual inspection of the resulting dendrogram showed the presence of 15 well-separated populations clusters.
A new polygon shapefile was created to delineate the population clusters within the river network. Each polygon contained the populations within a cluster, all the cells of the upstream branching network, as well as the downstream cells up to the point where the last tributary with a crayfish population from that cluster flows into the main river.
To assess the influence of protected areas (PAs) on the crayfish distribution pattern, the shapefiles of PAs in the region were superimposed over the river network. Sites of Community Importance in Romania (https://www.mmediu.ro/articol/date-gis/434) and Hungary (https://www.protectedplanet.net/country/HU) were considered for the EU region and PAs in the Republic of Serbia (https://www.protectedplanet.net/country/RS).
Each cell of the river network was finally assigned, based on location, as belonging to one of the 15 ecological clusters or to the river sector connecting those clusters, to PA or outside of PA.
Population genetics
Microsatellite analysis was performed on samples from 23 populations, of which 17 belonged to A. torrentium (South-Western Populations) and 6 to A. bihariensis (North-Western Populations). Sample size ranged from 10 to 66 individuals per population. The DNA extraction and PCR conditions followed the protocols described by ref.40. We used a set of five polymorphic microsatellites: aas5, aas6, aas3040, ap140 and Tor11, with the following sequence: Tor11F (5′- TGCTGCCCATGATATCCTTT-3′) and Tor11R (5′-TGGGGATGGTCATTTTGTT-3′) (unpublished data).
We ran PCR products on a LI-COR 4300 DNA analyser platform and visualised the results using IRD Dye 700 and 800 and a 50–700-bp size standard (LI-COR Biosciences, Li-COR Inc., Lincoln, NE, USA). The program SAGA v. 3.3 (LI-COR Biosciences, Li-COR Inc., Lincoln, NE, USA) was used to score the microsatellite alleles and to determine the genotypes.
Observed and expected heterozygosity (HO and HE), number of alleles and inbreeding coefficient (FIS) for each locus and each population were calculated in GenAlEx v. 6.50241. Allelic richness (AR) was calculated using FSTAT v. 2.9.342. Deviations from Hardy–Weinberg equilibrium (HWE) and linkage disequilibrium between all pairs of loci were tested in Genepop v. 4.643, and significance was determined using 100 batches and 1,000 iterations per batch.
We used MicroChecker v. 2.2.344 software to check for null alleles, genotyping error rates, allelic dropout or stuttering and to calculate null alleles frequencies. We also estimated null allele frequencies, as well as the corrected FST values using the Expectation Maximization algorithm45 and ENA method, implemented in FreeNA46.
Genetic differentiation was quantified calculating pairwise FST values47, calculated in GenAlEx v. 6.502; in addition, we compared FST values with and without the correction for null alleles calculated in FreeNA, using a t-test performed in Past v.1.5548. We also performed AMOVA to determine how the genetic variation was partitioned within and among populations grouped in the hierarchical schemes according to the clusters inferred from STRUCTURE, using GenAlEx v. 6.502. Isolation by distance was tested using a Mantel test implemented also in GenAlEx v. 6.502. A phylogenetic tree was constructed based on the microsatellite data using Neighbour-Joining algorithm and Da genetic distance49, implemented in POPULATIONS v. 1.2.3250.
We carried out a Bayesian analysis in STRUCTURE v. 2.3.451 to establish the number of genetic clusters of investigated samples, using all available individuals. For each value of K, from K = 1 to K = 25, we performed 10 independent runs of 750,000 generations, following 250,000 generations of burn-in under the admixture model and with the assumption that allele frequencies among populations are correlated. We identified the optimal number of clusters using both the Evanno method (ΔK; ref.52) and the highest probability for K following ref.51 on the CLUMPAK online platform (https://clumpak.tau.ac.il)53. Individuals were assigned as belonging to a particular cluster with an assignment probability (Q) of ≥ 0.8, with individuals with an assignment probability of 0.2 < Q < 0.8 being classified as ‘admixed’54,55). We used two other alternative methods to Bayesian clustering algorithms in order to identify the number of clusters and to characterize the population structure. The first one is discriminant analysis of principal components (DAPC), a multivariate method to identify clusters and describe genetic structure in biological populations, implemented in ADEGENET package for R56. The optimal number of clusters KDAPC is chosen as the minimum number of clusters after which the Bayesian information criterion (BIC) increases or decreases by a negligible amount56. Results were visualized as scatter plots in ADEGENET.
The second method is a multilocus maximum likelyhood algorithm57 that partitions a collection of genotypes into K groups in an iterative approach, implemented in FLOCK v. 3.158. We applied 20 re-allocations for the algorithm, with 50 runs for each K and a LLOD threshold score of 0. The optimal number of clusters, KF, was determined by ad hoc “stopping” rules58.
Results
Ecological and spatial assessments
Overall, 27.7% of the analysed river network in this study were found suitable for A. torrentium and A. bihariensis, clustered in 15 groups according to the ecological cost (Fig. 2). The range of values for each variable in the cells of the river network where the species were present can be found in Table 1. The random forest classifier out-of-bag estimate of the error rate was 28.6%, with error rates per class of 39.8% for absences and 14% for presences, respectively. An explanation for the model’s appearance to be under-predicting absences may be that some of the absences in the original data set may not have been caused by inadequate habitat conditions, but by various other factors (i.e. field detectability). A potential consequence of this behaviour could be an estimation of slightly wider suitability ranges, but the very good agreement with the observed occupied ranges suggests that this is unlikely. The ecological costs per cell ranged from 0 to 0.773 in the entire network, the distribution of values is provided in Figure S2.
Of the total analysed network, 30.4% were found inside a PA (Fig. 3). Table 2 summarises the basic statistics for the river network in this study. Analysing the ecological cost in the network, we found that for the network situated inside the 15 population clusters, the cost is significantly smaller compared to the cost associated to the network interconnecting them (Mann–Whitney, p < 2.2 × 10–16) (Fig. 3A). The ecological cost in the network situated in PAs was significantly smaller compared to that in the network outside PAs (Mann–Whitney, p < 2.2 × 10–16). Moreover, a significantly higher suitable network length was found in PAs (39.6%) than outside PAs (22.5%) (Fig. 3B).
The river network corresponding to crayfish population clusters according to the ecological cost (A) and protected areas (B). The distribution of ecological costs on the river network within and between clusters of populations, and inside and outside of protected areas, is presented below maps (A) and (B), respectively.
Genetic diversity
The dataset of 411 individuals was successfully genotyped, with only 1.02% missing data. No loci showed significant linkage disequilibrium tested over all populations after Bonferroni correction. Only three populations significantly differed from HWE expectations after Bonferroni correction: N1 for loci Aas3040 and Aas5, S9 and N6 for locus Tor11. Diversity values for the five loci ranged from 0.489 to 0.802 (HO) for A. torrentium and from 0.325 to 0.834 (HO) for A. bihariensis. The expected heterozygosity HE ranged from 0.5196 to 0.755 for A. torrentium and from 0.349 to 0.755 for A. bihariensis.
No evidence of large allele dropout and scoring errors due to stuttering was found. Also, MicroChecker found evidence of null alleles in four out of 23 populations in one or two loci (N1 at aas3040 and aas5, S5 at ap1 and tor 11, S11 at ap1 and S13 at aas6). FreeNA on the other hand, suggested the existence of null alleles in 55 out of 115 combinations of loci and populations, with values ranging from 0.0001 to 0.1655 (N2 in locus ap1).
The genetic diversity at the population level, measured by the allelic richness (AR), observed heterozygosity (HO) and expected heterozygosity (HE) varied from 3.91 to 6.98, 0.567 to 0.850 and 0.526 to 0.785, respectively, in A. torrentium and from 3.25 to 5.35, 0.485 to 0.752 and 0.503 to 0.680, respectively, in A. bihariensis. The inbreeding coefficient (FIS) varied from − 0.203 to 0.165 for A. torrentium and from − 0.113 to 0.074 for A. bihariensis populations (Table S1). One or two private alleles were identified in 13 out of 23 populations, with frequencies varying from 0.023 to 0.125. The highest FST values were registered between the populations from NW (i.e. A. bihariensis) versus populations from SW (i.e. A. torrentium). FST values and FST ENA values, uncorrected and corrected for null alleles, respectively calculated in FreeNA did not show any significant difference (t = − 0.6257, p = 0.53177). The FST index for A. torrentium populations (Table S2) indicated great population differentiation, with an overall value of 0.191 and pairwise FST values varying between 0.027 and 0.209. Pairwise FST values for A. bihariensis populations varied between 0.058 and 0.209, and overall FST had a value of 0.209, indicating high differentiation between analysed populations (Table S2). The Neighbour-Joining tree generated from the Da genetic distances visually illustrates the level of differentiation between A. torrentium and A. bihariensis as well as further differentiation between the populations of both species (Fig. 4A).
Neighbour-Joining dendrogram constructed with 1,000-bootstrap re-sampling, based on ref.49. DA calculated distance showing the relationship among A. torrentium and A. bihariensis populations from Romanian Carpathians (A); the numbers and coloured boxes in this figure represent the population clusters based on ecological cost. Clustering results obtained using STRUCTURE (B); each individual is represented by a vertical line partitioned into k segments, whose length is proportionate to their membership to each inferred cluster (subdivisions according to species and populations).
Bayesian analysis in STRUCTURE (Fig. S3) showed that two genetic groups of the investigated crayfish were present in the analysed region (ΔK = 2) when the Evanno method was employed. However, using the Bayesian clustering procedure, the highest likelihoods were obtained for Prob(K) = 16 (Fig. S3). In the first case, the two groups were represented by the two species (Fig. 4). In the second case, with the highest posterior probability Prob(K) = 16, populations of A. bihariensis were assigned to two clusters, while the populations of A. torrentium were assigned to other 14 different clusters. Overall, 211 individuals out of 411 were assigned to one of the clusters at Q ≥ 0.8, and the remaining 200 proved to be admixed (0.2 < Q < 0.8). The first cluster (for abbreviations, we refer readers to Table S1) was comprised of individuals from N1 (Q > 0.9), and the second cluster was comprised mainly of individuals from N6 and N5 (Q > 0.8). The remaining A. bihariensis populations had individuals belonging to both clusters (N3, N2) or were admixed (N4). In the A. torrentium populations, only S14, S10 and S15 populations had a higher Q (Q = 0.793, 0.768 and 0.713, respectively), each representing a different cluster. Populations S6, S1, S2, S3 and S4 had all the individuals admixed, and there was a high percentage of admixed individuals in S13 (21 out of 22). The rest of the remaining populations had a proportion of admixed individuals between 28.5 and 77.8%. The two species showed little evidence of mixing.
Evanno method of determining ΔK is best identify the highest level of genetic structure and may not be particularly reliable under complex evolutionary scenarios59. And although it is an elegant method from the statistic point of view, it always produces a “solution” for the given data and the solution may not be the best one60. The alterative assessment of clusters performed by DAPC revealed KDAPC = 14 inferred clusters, after 40 principal components and 4 discriminant functions were retained (Fig. S4A). The DAPC assignments of individuals from our populations to groups is illustrated in figure S4B. The DAPC scatter plot also clearly delineates the two Austropotamobius species populations (see Fig. S4C). The number of clusters inferred in DAPC is smaller than the one found in STRUCTURE (KDAPC = 14 vs Prob(K) = 16). A. bihariensis populations are assigned in DAPC to 5 distinct groups instead of 2 as in STRUCTURE Prob K results, with populations N1, N2, N5, N6 genetically differentiated and N3 and N4 with admixed individuals. In the A. torrentium populations the number of clusters identified in DAPC is smaller (KDAPC = 9) than in the analysis performed in STRUCTURE (Prob(K) = 14), but the results partially overlap the results performed by the Bayesian clustering aproach. We can see a separation between cluster 1 (k1—including the majority of the samples of S1, S2 and S3), cluster 6 (k6—S5, and S6) and the rest of the clusters with the rest of the A. torrentium individuals.
When we performed the multilocus maximum likelyhood proceedure in FLOCK on all populations we reached a “stopping” condition at KF = 2 (with a very long plateau of 50), recognising thus the two species. When the two metapopulations were analysed separately, we obtained an optimum KF = 4 for the ones belonging to A. bihariensis and an optimum KF = 2 for the A. torrentium populations (see Fig. S5).
Hierarchical AMOVA identified significant geographical structuring and showed an elevated percentage of variation among populations (25%, p = 0.001). Genetic structuring was supported when the STRUCTURE results, 2 and 13 groups, respectively (the number of clusters is lower due to admixed populations) were considered. We also observed a strong genetic structure when we took the genetic variation according to the drainage basins into account (Table S3). Mantel tests of the relationships between genetic and geographic distances proved a moderate correlation and revealed an isolation-by-distance pattern (r = 0.525; p < 0.001).
Discussion
The current status of populations
Generally, the results of this study revealed a moderate to high genetic diversity in comparison to other studies on populations of A. torrentium from Austria, Switzerland and Germany (e.g. ref.61). We observed a strong genetic structuring with no hybridisation between two groups of Austropotamobius species populations, suggesting separation that occurred a very long time ago. This result is in agreement with the recent documentation that these populations no longer belong to a single species18. The separation was, most likely, driven by a tectonic displacement of the Tisza-Dacia microplate (including the current Apuseni Mountains in NW of Romania) during the early Miocene (ca. 15 Ma) (see ref.20).
The clade of A. bihariensis revealed high genetic diversity and genetic structuring with up to 4 populations recognised as distinct clusters. The molecular data is supporting the argument for the endemic nature of this species. A homogenous population of A. bihariensis was found in the N1 river, presenting lower heterozygosity than other populations. The spatial analysis of this watercourse showed a large ecological distance from the rest of the A. bihariensis populations (see Fig. 2). However, genetically, the N1 crayfish were highly similar to the populations in the neighbouring basin (see Fig. 4, populations N1 versus N2, N3). Considering the topography of the area inhabited by populations N1 and N2, namely the fact that the headwaters of both rivers are situated in the same plateau a few hundred meters apart, with human dwellings nearby, we suspect that the N1 population is a result of human (or otherwise mediated) translocations from N2.
The strong genetic differentiation between A. torrentium populations indicates both ecological fragmentation and very limited geneflow. The results from the different k-clustering methods suggest a complex structure related to a complicated population history in relation to geographical river basins. Considering that those populations belong to a more recent haplogroup (CSE, sensu ref.22) which presumably colonised the Danube basin before and after the Pleistocene glaciations20,22, the genetic pattern observed in this study supports the assumption of multiple colonisation episodes in the past, which explains the legacy of high diversity and divergence between neighbouring basins and populations. Spatial analysis of the river network shows a mixture of sectors with zero ecological cost, mainly found in the upper sections of rivers, and sectors with positive ecological cost found especially on the main courses. The RWQ (i.e. water quality) was identified as the most important contributor to the ecological cost, which supports the assumption that the fragmentation pattern is a recent one, most likely resulting from anthropic development62.
An unexpected situation was found in a group of populations with a low level of genetic differentiation but considerable ecological separation (see Fig. 4). This is the case for six populations of A. torrentium belonging to three different river basins (see Fig. 2): S1, S2, S3 (Timiș River basin), S4 (Strei River basin) and S5 and S6 (northern Jiu River basin). At the same time, a branch of the Jiu River, situated further south, hosts only populations (i.e. S7 and S8) expressing high genetic differentiation from those in the previously mentioned cluster (see Fig. 4, S4). A possible scenario explaining this situation might be that the Carpathians are acting as a natural barrier that historically impeded the colonisation route between the southern and northern populations of the Jiu River. In this respect, we found that there are several cells where the FFP greatly exceeds the species optimum in the Jiu River sector traversing the Carpathians. The molecular cluster of the six populations from the three different basins might be explained by the regressive erosion, documented between the Strei River and the neighbouring tributaries of the Timiș and the Jiu rivers63. This phenomenon occurred most likely between Pleistocene glaciation episodes and created temporary connections between tributaries of those river basins64, thus leading to the unexpected distribution. An alternative hypothesis, such as the crayfish traveling by land between rivers, is unlikely, as this behaviour was never documented for any Austropotamobius species. Expansion driven by human-mediated transport cannot be excluded, although the genetic structure of these populations implies this must have occurred repeatedly in the past for all the three river basins.
Remarks on conservation
Conservation strategies targeting crayfish are priorities worldwide17,65, inside and outside of PAs. The results of this study show the current situation for the two Austropotamobius species as being fragmented both ecologically and with respect to gene flow. A deeper analysis of the variables causing ecological fragmentation suggests water quality as a main reason (see Table 2), which is in agreement with other studies21,66,67. Water quality deterioration was caused by recent anthropogenic regional development62, which is why its effects are not clearly expressed in the observed genetic patterns (see Fig. 4). The nowadays unsuitability of large rivers could be an important issue for the current conservation initiatives, since these rivers were migration pathways in the past. The species distribution in the region is documented as being a result of intense colonisation events after the Paratethys retreat20, corresponding to the development of the paleo‐Danube system within the Pannonian Basin68.
Fragmentation causes inbreeding and thus homozygosity69, which can further increase the chances of offspring being affected by recessive or deleterious traits, leading to a decreased biological fitness of populations and the decline of long-term survival70. The artificial gene flow within populations by translocating individuals could be a solution for the improvement of genetic diversity. Still criticised, this method must be carefully applied using the most appropriate gene pool for repopulation to minimise the impact on the original haplotypes’ distribution71 and to avoid spread of diseases72,73. With respect to habitat adequacy, PAs were found to provide reasonably good conditions for both species. However, the degree of suitability to the species’ requirements drops dramatically on the network connecting the population clusters. Thus, a substantial impediment is found against the communication between these crayfish populations. Even though PAs offer good habitat conditions locally, the long-term protection strategy fails to provide interconnectivity, ensuring the communication corridors between population clusters being a challenging task74,75.
Another factor that can influence species distribution in an area is the presence of invasive species and the spread of their associated diseases76. As around 20% of the faunal extinctions could be tracked down to invasive species77, identification of the main pathways of colonisation for freshwater alien species in Europe is of utmost importance78. Our method could help in species risk assessment and identification of probable river pathways (as the ones with the lowest ecological cost) of the highly successful invasive species78. In this context, there is a good side of the fragmented pattern found in A. torrentium and A. bihariensis populations. In the worst scenario of upcoming invasions (see e.g. refs.31,79), the fragmented habitats could offer “ark sites” (sensu ref.80) of isolated populations, turning into an advantage against unwanted colonisers (see e.g. refs.16,81).
Conclusion and recommendations
Quantifying habitat connectivity by considering the dendritic nature of freshwater networks has been a key element in evaluating the current status in conservation of two sensitive and endangered Austropotamobius species in the Carpathians. Combining ecological and genetic approaches, we provide a more complete picture. In this study, without ecological analysis, the fragmentation would have been underestimated, and also without genetic analyses, the translocations and atypical colonisation routes would not have been documented and revealed.
In the light of IUCN Red List of Threatened Species relevant criteria 82 such as decreasing population trend83, the results of this study might be used to reconsider the “data deficient”83 status for A. torrentium. Moreover, for the still unrated A. bihariensis, a newly described species with few populations and a scarce gene pool, a conservation status assessment is urgently needed.
References
Erős, T., O’Hanley, J. R. & Czeglédi, I. A unified model for optimizing riverscape conservation. J. Appl. Ecol. 55, 1871–1883 (2018).
Ruggeri, P., Pasternak, E. & Okamura, B. To remain or leave: Dispersal variation and its genetic consequences in benthic freshwater invertebrates. Ecol. Evol. 9, 12069–12088 (2019).
Baguette, M., Blanchet, S., Legrand, D., Stevens, V. M. & Turlure, C. Individual dispersal, landscape connectivity and ecological networks. Biol. Rev. 88, 310–326 (2013).
Geist, J. Seven steps towards improving freshwater conservation. Aquat. Conserv. Mar. Freshw. Ecosyst. 25, 447–453 (2015).
Kujala, H., Lahoz-Monfort, J. J., Elith, J. & Moilanen, A. Not all data are equal: Influence of data type and amount in spatial conservation prioritisation. Methods Ecol. Evol. 9, 2249–2261 (2018).
Johnson, J. B., Peat, S. M. & Adams, B. J. Where’s the ecology in molecular ecology?. Oikos 118, 1601–1609 (2009).
Janse, J. H. et al. GLOBIO-aquatic, a global model of human impact on the biodiversity of inland aquatic ecosystems. Environ. Sci. Policy 48, 99–114 (2015).
Newbold, T. et al. Global effects of land use on local terrestrial biodiversity. Nature 520, 45–50 (2015).
Moore, D., Cranston, G., Reed, A. & Galli, A. Projecting future human demand on the Earth’s regenerative capacity. Ecol. Indic. 16, 3–10 (2012).
Yawson, D. O., Adu, M. O. & Armah, F. A. Impacts of climate change and mitigation policies on malt barley supplies and associated virtual water flows in the UK. Sci. Rep. 10, 1–12 (2020).
Naidoo, R. et al. Global mapping of ecosystem services and conservation priorities. Proc. Natl. Acad. Sci. USA 105, 9495–9500 (2008).
Hermoso, V., Villero, D., Clavero, M. & Brotons, L. Spatial prioritisation of EU’s LIFE-Nature programme to strengthen the conservation impact of Natura 2000. J. Appl. Ecol. 55, 1575–1582 (2018).
Hermoso, V., Morán-Ordóñez, A., Canessa, S. & Brotons, L. Realising the potential of Natura 2000 to achieve EU conservation goals as 2020 approaches. Sci. Rep. 9, 1–10 (2019).
Lobera, G., Pardo, I., García, L. & García, C. Disentangling spatio-temporal drivers influencing benthic communities in temporary streams. Aquat. Sci. 81, 1–17 (2019).
Richman, N. I. et al. Multiple drivers of decline in the global status of freshwater crayfish (Decapoda: Astacidea). Philos. Trans. R. Soc. B Biol. Sci. 370, 20140060 (2015).
Manenti, R. et al. Causes and consequences of crayfish extinction: Stream connectivity, habitat changes, alien species and ecosystem services. Freshw. Biol. 64, 284–293 (2019).
Kozák, P., Füreder, L., Kouba, A., Reynolds, J. & Souty-Grosset, C. Current conservation strategies for European crayfish. Knowl. Manag. Aquat. Ecosyst. 01, https://doi.org/10.1051/kmae/2011018 (2011).
Pârvulescu, L. Introducing a new Austropotamobius crayfish species (Crustacea, Decapoda, Astacidae): A miocene endemism of the Apuseni Mountains, Romania. Zool. Anz. 279, 94–102 (2019).
Kouba, A., Petrusek, A. & Kozák, P. Continental-wide distribution of crayfish species in Europe: Update and maps. Knowl. Manag. Aquat. Ecosyst. 413, 05–31 (2014).
Pârvulescu, L. et al. A journey on plate tectonics sheds light on European crayfish phylogeography. Ecol. Evol. 9, 1957–1971 (2019).
Pârvulescu, L. & Zaharia, C. Current limitations of the stone crayfish distribution in Romania: Implications for its conservation status. Limnologica 43, 143–150 (2013).
Klobučar, G. I. V. et al. Role of the Dinaric Karst (western Balkans) in shaping the phylogeographic structure of the threatened crayfish Austropotamobius torrentium. Freshw. Biol. 58, 1089–1105 (2013).
Qian, S. S., Cuffney, T. F., Alameddine, I., McMahon, G. & Reckhow, K. H. On the application of multilevel modeling in environmental and ecological studies. Ecology 91, 355–361 (2010).
Manning, P. et al. Redefining ecosystem multifunctionality. Nat. Ecol. Evol. 2, 427–436 (2018).
Koizumi, I., Usio, N., Kawai, T., Azuma, N. & Masuda, R. Loss of genetic diversity means loss of geological information: The endangered Japanese crayfish exhibits remarkable historical footprints. PLoS ONE 7, e33986 (2012).
McNyset, K. M. Use of ecological niche modelling to predict distributions of freshwater fish species in Kansas. Ecol. Freshw. Fish 14, 243–255 (2005).
Henrys, P. A. & Jarvis, S. G. Integration of ground survey and remote sensing derived data: Producing robust indicators of habitat extent and condition. Ecol. Evol. 9, 8104–8112 (2019).
Pârvulescu, L., Zaharia, C., Satmari, A. & Drăguţ, L. Is the distribution pattern of the stone crayfish in the Carpathians related to karstic refugia from Pleistocene glaciations?. Freshw. Sci. 32, 1410–1419 (2013).
Longshaw, M. & Stebbing, P. Biology and Ecology of Crayfish. (CRC Press, 2015).
Chucholl, C. The bad and the super-bad: Prioritising the threat of six invasive alien to three imperilled native crayfishes. Biol. Invasions 18, 1967–1988 (2016).
Chucholl, C. & Schrimpf, A. The decline of endangered stone crayfish (Austropotamobius torrentium) in southern Germany is related to the spread of invasive alien species and land-use change. Aquat. Conserv. Mar. Freshw. Ecosyst. 26, 44–56 (2016).
Pârvulescu, L. et al. Flash-flood potential: A proxy for crayfish habitat stability. Ecohydrology 9, 1507–1516 (2016).
Farr, T. G. et al. The shuttle radar topography mission. Rev. Geophys. 45, RG2004 (2007).
Şandric, I. et al. Integrating catchment land cover data to remotely assess freshwater quality: A step forward in heterogeneity analysis of river networks. Aquat. Sci. 81, 26 (2019).
Burkhard, B., Kroll, F., Nedkov, S. & Müller, F. Mapping ecosystem service supply, demand and budgets. Ecol. Indic. 21, 17–29 (2012).
Zeller, K. A., McGarigal, K. & Whiteley, A. R. Estimating landscape resistance to movement: A review. Landsc. Ecol. 27, 777–797 (2012).
Liaw, A. & Wiener, M. Classification and regression by randomForest. R News 2, 18–22 (2002).
R Core Team. R: A Language and Environment for Statistical Computing. (R Foundation for Statistical Computing, 2017).
Freeman, E. A. & Moisen, G. G. A comparison of the performance of threshold criteria for binary classification in terms of predicted prevalence and kappa. Ecol. Modell. 217, 48–58 (2008).
Iorgu, E. I., Popa, O. P., Petrescu, A.-M. & Popa, L. O. Cross-amplification of microsatellite loci in the endangered stone-crayfish Austropotamobius torrentium (Crustacea: Decapoda). Knowl. Manag. Aquat. Ecosyst. 08, https://doi.org/10.1051/kmae/2011021 (2011).
Peakall, R. & Smouse, P. E. genalex 6: Genetic analysis in Excel. Population genetic software for teaching and research. Mol. Ecol. Notes 6, 288–295 (2006).
Goudet, J. FSTAT (Version 1.2): A computer program to calculate F-statistics. J. Hered. 86, 485–486 (1995).
Rousset, F. genepop’007: A complete re-implementation of the genepop software for Windows and Linux. Mol. Ecol. Resour. 8, 103–106 (2008).
Van Oosterhout, C., Hutchinson, W. F., Wills, D. P. & Shipley, P. micro-checker: Software for identifying and correcting genotyping errors in microsatellite data. Mol. Ecol. Notes 4, 535–538 (2004).
Dempster, A. P., Laird, N. M. & Rubin, D. B. Maximum likelihood from incomplete data via the EM algorithm. J. R. Stat. Soc. Ser. B 39, 1–22 (1977).
Chapuis, M. P. & Estoup, A. Microsatellite null alleles and estimation of population differentiation. Mol. Biol. Evol. 24, 621–631 (2007).
Weir, B. S. & Cockerham, C. C. Estimating F‐statistics for the analysis of population structure. Evolution (N. Y). 38, 1358–1370 (1984).
Hammer, D. A. T., Ryan, P. D., Hammer, Ø. & Harper, D. A. T. Past: Paleontological Statistics Software Package for Education and Data Analysis. Palaeontologia Electronica vol. 4 https://palaeo-electronica.orghttp//palaeo-electronica.org/2001_1/past/issue1_01.htm. (2001).
Nei, M., Tajima, F. & Tateno, Y. Accuracy of estimated phylogenetic trees from molecular data. J. Mol. Evol. 19, 153–170 (1983).
Langella, O. Populations, 1.2. 30. https://bioinformatics.org/~tryphon/populations (1999).
Pritchard, J. K., Stephens, M., Rosenberg, N. A. & Donnelly, P. Association mapping in structured populations. Am. J. Hum. Genet. 67, 170–181 (2000).
Evanno, G., Regnaut, S. & Goudet, J. Detecting the number of clusters of individuals using the software STRUCTURE: A simulation study. Mol. Ecol. 14, 2611–2620 (2005).
Kopelman, N. M., Mayzel, J., Jakobsson, M., Rosenberg, N. A. & Mayrose, I. Clumpak: A program for identifying clustering modes and packaging population structure inferences across K. Mol. Ecol. Resour. 15, 1179–1191 (2015).
Vähä, J. P. & Primmer, C. R. Efficiency of model-based Bayesian methods for detecting hybrid individuals under different hybridization scenarios and with different numbers of loci. Mol. Ecol. 15, 63–72 (2005).
Bergl, R. A. & Viglant, L. Genetic analysis reveals population structure and recent migration within the highly fragmented range of the Cross River gorilla (Gorilla gorilla diehli). Mol. Ecol. 16, 501–516 (2006).
Jombart, T., Devillard, S. & Balloux, F. Discriminant analysis of principal components: A new method for the analysis of genetically structured populations. BMC Genet. 11, 1–15 (2010).
Paetkau, D., Calvert, W., Stirling, I. & Strobeck, C. Microsatellite analysis of population structure in Canadian polar bears. Mol. Ecol. 4, 347–354 (1995).
Duchesne, P. & Turgeon, J. FLOCK Provides Reliable Solutions to the ‘“Number of Populations”’ Problem. https://doi.org/10.1093/jhered/ess038.
Janes, J. K. et al. The K = 2 conundrum. Mol. Ecol. 26, 3594–3602 (2017).
Funk, S. M. et al. Major inconsistencies of inferred population genetic structure estimated in a large set of domestic horse breeds using microsatellites. Ecol. Evol. 10, 4261–4279 (2020).
Berger, C., Štambuk, A., Maguire, I., Weiss, S. & Füreder, L. Integrating genetics and morphometrics in species conservation—A case study on the stone crayfish, Austropotamobius torrentium. Limnologica 69, 28–38 (2018).
Iojă, C. I. et al. The efficacy of Romania’s protected areas network in conserving biodiversity. Biol. Conserv. 143, 2468–2476 (2010).
Rabăgia, T. & Maţenco, L. Tertiary tectonic and sedimentological evolution of the South Carpathians foredeep: Tectonic vs eustatic control. Mar. Pet. Geol. 16, 719–740 (1999).
Rãdoane, M., Rãdoane, N. & Dumitriu, D. Geomorphological evolution of longitudinal river profiles in the Carpathians. Geomorphology 50, 293–306 (2003).
Helms, B., Loughman, Z. J., Brown, B. L. & Stoeckel, J. Recent advances in crayfish biology, ecology, and conservation. Freshw. Sci. 32, 1273–1275 (2013).
Svobodová, J. et al. The relationship between water quality and indigenous and alien crayfish distribution in the Czech Republic: Patterns and conservation implications. Aquat. Conserv. Mar. Freshw. Ecosyst. 22, 776–786 (2012).
Pöckl, M. & Streissl, F. Austropotamobius torrentium as an indicator for habitat quality in running waters? Bull. Français la Pêche la Piscic. 743–758, https://doi.org/10.1051/kmae:2005030 (2005).
Magyar, I. et al. Progradation of the paleo-Danube shelf margin across the Pannonian Basin during the Late Miocene and Early Pliocene. Glob. Planet. Change 103, 168–173 (2013).
Zhang, Y., Luan, P., Ren, G., Hu, G. & Yin, J. Estimating the inbreeding level and genetic relatedness in an isolated population of critically endangered Sichuan taimen (Hucho Bleekeri) using genome-wide SNP markers. Ecol. Evol. 10, 1390–1400 (2020).
Hoarau, G. et al. Low effective population size and evidence for inbreeding in an overexploited flatfish, plaice (Pleuronectes platessa L.). Proc. Biol. Sci. 272, 497–503 (2005).
Jourdan, J. et al. Reintroduction of freshwater macroinvertebrates: Challenges and opportunities. Biol. Rev. https://doi.org/10.1111/brv.12458 (2018).
Oidtmann, B., Heitz, E., Rogers, D. & Hoffmann, R. Transmission of crayfish plague. Dis. Aquat. Organ. 52, 159–167 (2002).
Rusch, J. C. et al. Simultaneous detection of native and invasive crayfish and Aphanomyces astaci from environmental DNA samples in a wide range of habitats in Central Europe. NeoBiota (2020).
Hall, Q. A., Curtis, J. M., Williams, J. & Stunz, G. W. The importance of newly-opened tidal inlets as spawning corridors for adult Red Drum (Sciaenops ocellatus). Fish. Res. 212, 48–55 (2019).
Stewart, F. E. C., Darlington, S., Volpe, J. P., McAdie, M. & Fisher, J. T. Corridors best facilitate functional connectivity across a protected area network. Sci. Rep. 9, 10852 (2019).
Strauss, A., White, A. & Boots, M. Invading with biological weapons: The importance of disease-mediated invasions. Funct. Ecol. 26, 1249–1261 (2012).
Clavero, M. & García-Berthou, E. Invasive species are a leading cause of animal extinctions. Trends Ecol. Evol. 20, 110 (2005).
Nunes, A. L., Tricarico, E., Panov, V. E., Cardoso, A. C. & Katsanevakis, S. Pathways and gateways of freshwater invasions in Europe. Aquat. Invasions 10, 359–370 (2015).
Zeng, Y. & Yeo, D. C. J. Assessing the aggregated risk of invasive crayfish and climate change to freshwater crabs: A Southeast Asian case study. Biol. Conserv. 223, 58–67 (2018).
Alonso, F., Temino, C. & Diéguez-Uribeondo, J. Status of the white-clawed crayfish, Austropotamobius pallipes (Lereboullet, 1858), in Spain: Distribution and legislation. 31–53 (2000).
Van Dyck, H. & Baguette, M. Dispersal behaviour in fragmented landscapes: Routine or special movements?. Basic Appl. Ecol. 6, 535–545 (2005).
Rodrigues, A. S. L., Pilgrim, J. D., Lamoreux, J. F., Hoffmann, M. & Brooks, T. M. The value of the IUCN Red List for conservation. Trends Ecol. Evol. 21, 71–76 (2006).
Füreder, L., Gherardi, F. & Souty-Grosset, C. Austropotamobius torrentium. The IUCN Red List of Threatened Species 2010 e.T2431A9439449 https://doi.org/10.2305/IUCN.UK.2010-3.RLTS.T2431A9439449.en (2010).
Acknowledgements
This work was partially supported by a grant of the Romanian National Authority for Scientific Research and Innovation, CNCS-UEFISCDI, project no. PN-II-ID-PCE-2008-2-1458. MCI contribution was supported by the Institute of Biology Bucharest of Romanian Academy, project no. RO1567-IBB04/2020. The authors thank Andrei Heidelbacher for his help in the management of large datasets, to Ionuț-Ștefan Iorgu for his help with figures and to Tiberiu Sahlean and Andrei Ștefan for their help with some software analysis. We thank the three anonymous reviewers whose valuable suggestions significantly improved the manuscript.
Author information
Authors and Affiliations
Contributions
L.P. conceived the idea. L.P. provided field samples collection. E.I.I., A.M.K., O.P.P. and L.O.P. performed genetic data and population genetic analyses. A.S. and M.C.I. performed geospatial analyses. C.Z. and K.M. performed statistical analyses. I.P. leads the project providing funds. L.P. and E.I.I. led the writing of the manuscript. All authors contributed and approved publication.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Pârvulescu, L., Iorgu, EI., Zaharia, C. et al. The future of endangered crayfish in light of protected areas and habitat fragmentation. Sci Rep 10, 14870 (2020). https://doi.org/10.1038/s41598-020-71915-w
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-020-71915-w
- Springer Nature Limited