Abstract
Downy mildew of spinach is caused by the obligate oomycete pathogen, Peronospora effusa. The disease causes significant economic losses, especially in the organic sector of the industry where the use of synthetic fungicides is not permitted for disease control. New pathotypes of this pathogen are increasingly reported which are capable of breaking resistance. In this study, we took advantage of new spinach genome resources to conduct RNA-seq analyses of transcriptomic changes in leaf tissue of resistant and susceptible spinach cultivars Solomon and Viroflay, respectively, at an early stage of pathogen establishment (48 hours post inoculation, hpi) to a late stage of symptom expression and pathogen sporulation (168 hpi). Fold change differences in gene expression were recorded between the two cultivars to identify candidate genes for resistance. In Solomon, the hypersensitive inducible genes such as pathogenesis-related gene PR-1, glutathione-S-transferase, phospholipid hydroperoxide glutathione peroxidase and peroxidase were significantly up-regulated uniquely at 48 hpi and genes involved in zinc finger CCCH protein, glycosyltransferase, 1-aminocyclopropane-1-carboxylate oxidase homologs, receptor-like protein kinases were expressed at 48 hpi through 168 hpi. The types of genes significantly up-regulated in Solomon in response to the pathogen suggests that salicylic acid and ethylene signaling pathways mediate resistance. Furthermore, many genes involved in the flavonoid and phenylpropanoid pathways were highly expressed in Viroflay compared to Solomon at 168 hpi. As anticipated, an abundance of significantly down-regulated genes was apparent at 168 hpi, reflecting symptom development and sporulation in cultivar Viroflay, but not at 48 hpi. In the pathogen, genes encoding RxLR-type effectors were expressed during early colonization of cultivar Viroflay while crinkler-type effector genes were expressed at the late stage of the colonization. Our results provide insights on gene expression in resistant and susceptible spinach-P. effusa interactions, which can guide future studies to assess candidate genes necessary for downy mildew resistance in spinach.
Similar content being viewed by others
Introduction
The demand for prepackaged ready to eat salad mixes has resulted in increased production of leafy greens in recent years. Currently, spinach is cultivated in more than 60 countries globally, with over 53 million tons of total production annually1. In the US, nearly four hundred thousand tons of spinach are produced every year1. There has also been a rapid increase in the demand for spinach in the US, beginning in the 1990s2,3. The Salinas Valley of California is sometimes referred to as ‘Salad Bowl of America’ since this region produces the majority of leafy greens grown in the US as well as nearly half of the total spinach in California4,5.
Downy mildew disease of spinach is caused by the obligate oomycete pathogen Peronospora effusa (Phylum Oomycota, Kingdom Stramenopila)6. Downy mildew on spinach can sometimes cause 100% yield loss in organic production systems, though the disease can be effectively managed by the applications of synthetic fungicides in conventional production. The use of resistant cultivars is the most promising control measure to minimize the downy mildew damage in organic spinach as well as to help reduce chemical usage for disease control in conventional spinach production. P. effusa is limited in its host range to spinach, and therefore spinach must be present for either sexual or asexual reproduction7,8,9. Given what is known about related species of Peronospora, the thick-walled, sexually produced oospores of P. effusa may be viable for several years in the environment despite adverse conditions6. Recently, oospore infestations were reported in about 19% of commercial spinach seed lots10. The movement of pathogen through infested seeds as oospores introduces novel recombinant isolates into spinach production sites and can spread the disease rapidly over large geographical regions9. Additionally, P. effusa produces tens of thousands of asexual spores per plant, which disperse aerially and initiate secondary infections within a field, or primary infections at another site, in distant fields9,11.
Increasing numbers of races, or new virulent pathotypes of P. effusa, have been detected using a set of differential spinach cultivars and reported in recent years in the US and other countries12,13. The emergence of virulent isolates capable of overcoming resistance genes is more common in pathogens such as downy mildews since they produce many asexual spores and retain high recombination rates14,15, but their emergence is exacerbated by sexual reproduction, as noted by the presence of oospores in the population10. Research efforts on P. effusa have focused on race phenotyping and understanding the basic epidemiology of the disease9. However, the recent public release of the genome sequences of P. effusa isolates of races 12, 13, and 14 have provided additional tools for genetic analysis of the pathogen16,17.
With the availability and affordability of next generation sequencing technology, transcriptome analyses between compatible (host susceptible) and incompatible (host resistant) plant-pathogen interactions have been useful to understand the molecular mechanisms that underlie the outcomes of these interactions in various pathosystems18,19,20,21,22,23. In plants, extracellular receptors serve as gatekeepers and recognize incoming pathogen-associated molecular patterns (PAMPs)24. Detection of PAMPs activates a basal defense system called pattern-triggered immunity (PTI)24,25,26. PTI can confer broad spectrum resistance through a basal defense network but pathogens coevolve with their host plants and can often breach the basal defense barrier by secreting effectors that are transported into the host cells24,27. Plant resistance (R) genes confer resistance against particular strains or races of a pathogen based on the recognition of specific effector molecules24,27. Plant R genes have been utilized as resources to develop resistant crop cultivars for several plant pathogens. In spinach, R genes at the RPF1 locus mediated resistance have been reported but functional roles of these genes conferring resistance to different races of P. effusa have not been thoroughly evaluated and validated28.
In this study, we performed genome-wide expression profiling of resistant and susceptible spinach-P. effusa interactions to investigate the defense mechanisms that confer resistance against P. effusa infection. For this purpose, we used resistant and susceptible spinach cultivars Solomon (also known as Lion) and Viroflay, respectively, which were inoculated with P. effusa and monitored for differential gene expression at two different time points, at an early stage of pathogen establishment and at the late stage of symptom expression and sporulation. We assembled a high-quality spinach reference genome sequence with annotation to facilitate this current analysis. In-depth analysis of genes differentially expressed during infection or in response to P. effusa yielded insights into the genetic basis of resistance and susceptibility in spinach and expression profiling of putative effector genes in P. effusa that are required for virulence and proliferation.
Materials and methods
Plant materials, pathogen, and pathogen inoculation
Seeds of spinach differentials (Viroflay and Solomon) for detection of distinct pathotypes of P. effusa were sown in Sunshine potting mix (Sun Gro Horticulture) and allowed to grow for two weeks in the glasshouse under ambient conditions12. Viroflay is a semi-smooth leaf cultivar and susceptible to all known races of P. effusa while Solomon is a smooth leaf cultivar resistant to race numbers 1–9 and 11–1629 Two-week old seedlings were spray-inoculated with freshly harvested sporangia of P. effusa with the concentration of nearly 1 ×104 sporangia/ml or remained un-inoculated as control plants. (Fig. 1). All seedlings were incubated in the dark for 24 hours in a dew chamber under high humidity (> 90%) at 18 oC. The seedlings were moved into a mist chamber maintained at 19 oC fixed with water misters to provide high humidity. Both P. effusa inoculated and un-inoculated control plants were remained in the mist chamber under a 12:12-h light-dark photoperiod for five days and were returned to the dew chamber for 24 hours in the dark at 18 oC to stimulate sporulation12.
Experimental design and sample collection for RNA sequencing analyses of resistant and susceptible spinach cultivars in response to the spinach downy mildew pathogen, Peronospora effusa. Two-week-old spinach seedlings of the differential cultivars were inoculated with P. effusa sporangial suspensions. Leaves of the downy mildew resistant cultivar Solomon and susceptible cultivar Viroflay were collected at 48 and 168 hpi, and immediately transferred in liquid nitrogen. Un-inoculated control seedlings were also incubated overnight in the dew chamber, prior to collecting leaf samples. Three leaves per plant per treatment were pooled for total RNA extraction. cDNA libraries were prepared using three separate biological replicates per treatment.
Sample collections and experimental design
The experimental design is outlined in Fig. 1. Briefly, pathogen-inoculated leaf samples from cultivars Viroflay and Solomon were each collected at 48, and at 168 hours post-inoculation period (hpi). The un-inoculated control leaf samples were collected after 24-hr incubation in the dew chamber. There were three biological replicates (leaves harvested from three individual plants) per treatment (Fig. 1). Three true leaves were harvested manually from each plant at the respective time points, transferred immediately into liquid nitrogen, and stored at −80 oC until further processing.
RNA extraction, library preparation, and sequencing
Total RNA was extracted from frozen leaf samples using the RNeasy Mini Kit (Qiagen, Valencia, CA) according to manufacturer’s instructions on 100 mg of tissue for each sample, and yields were quantified using a Qubit 2.0 Fluorimeter (Invitrogen, Carlsbad, CA). cDNA libraries were constructed following the protocol as described earlier by Zhong et al.30. Libraries were sequenced on an Illumina HiSeq 4000 (Illumina, San Diego, CA) at the DNA Technologies and Expression Analysis Core Laboratory, University of California, Davis to yield 364,359,078 total sequence reads (single-end 50 bp) from all samples (Supplementary Table 1).
Reference genomes used in this study
The reference genome of P. effusa used in this study was derived from a race 13 isolate from Monterey County, California (National Center for Biotechnology Information accession: QKXF00000000.1). The sequencing and annotation of 8607 genes in this P. effusa genome is described in detail by Fletcher et al.16.
The reference spinach genome of cultivar Viroflay used in this study was sequenced using Pacific Biosciences (Menlo Park, CA) technology to 70X coverage and assembled de novo using a hierarchical genome-assembly process and Celera assembler software31. The genome sequence, 890,266,198 bp in length, was annotated by Beijing Genomics Institute (Beijing, China) using MAKER32 and yielded a total of 34,878 genes in this spinach reference genome.
Mapping sequence reads to the reference genomes and identification of differentially expressed genes
The mRNA-sequence reads obtained from each of the experimental spinach leaf samples were trimmed to remove adapter sequences using CLC Genomics Workbench 11.0.1 (http://www.clcbio.com). Trimmed reads were mapped to the spinach and P. effusa reference genomes using the default setting of RNA-Seq analysis tool in CLC Genomics. To ascertain the expression values, gene and mRNA tracks were created with CLC using both spinach and P. effusa genomes and their respective generic feature format files. Total gene reads, or in other words the total number of reads mapped to genes in the reference genome, were used to assess differential expression (Supplementary Table 1). An equally mapped read to both genic and intergenic regions was also considered as a mapped gene and included in the total counts. Reads that mapped outside of the annotated genes were counted as intergenic hits only and were excluded from the total gene reads. A two-dimensional heatmap was developed by clustering the expression values of individual genes across the samples at 48 and 168 hpi, respectively. The normalized log count per million (CPM) was calculated for each gene as an expression value. The Manhattan distance was used to calculate pairwise distances between all clusters where two closest clusters were merged into a single new cluster33. Furthermore, a three-dimensional principal component analysis (PCA) was performed using the default setting in CLC Genomics where normalized log CPM values were used as expression values. In the PCA analysis, PC1, PC2 and PC3 displayed the direction with the maximum, intermediate and minimum variability respectively, in the data.
In the RNA-seq tool in CLC Genomics Workbench (version 11.0.1; http://www.clcbio.com), we used the differential expression setting to determine the statistical significance of expressed genes in Solomon versus Viroflay at un-inoculated control and inoculated: 48 hpi and 168 hpi. We uploaded gene expression tracks of each samples containing total exon reads which were chosen to calculate the fold change and p-value statistics for the individual gene. Subsequently, we selected the metadata file containing all information including sample name, genotype (Solomon/Viroflay), time point of sample collection, and lane used while sequencing. For the experimental design and comparisons, we selected ‘genotype’ to test the differential expression, ‘lane’ for the control factor, and ‘all group pairs’ for the comparison. The analysis was conducted separately for each time point. Each expressed gene was modeled by using a generalized linear model (GLM)34, and calculated the dispersion estimate by a linear combination of the probability for that particular gene and its neighboring genes with similar average expression amounts. The GLM assumes the negative binomial distribution of the expression data. In the negative binomial distribution, dispersion of expression values serves as a free parameter indicating the homogeneity or heterogeneity i.e. smaller or greater variation in expression levels for a particular gene. We used Wald test34 for statistical testing and identified the significantly expressed genes at 48 hpi and 168 hpi between cultivars Solomon and Viroflay. The Wald test provided the p-value statistics of fold change of each gene between treatments by testing a non-zero coefficient. Species distribution of all BLAST35 hits of spinach genes was performed using the default setting of Blast2GO 5 PRO (https://www.blast2go.com/) program.
Functional and pathway enrichment analysis of differentially expressed genes
Functional annotations of 34,878 spinach genes were generated through BLAST searches using CloudBlast in Blast2GO 5 Pro (https://www.blast2go.com/) program. The Blasted sequences were then mapped, annotated, and assigned the gene ontology (GO) IDs and functional description. The Fisher’s Exact Test was used in Blast2GO to perform a GO enrichment analysis to identify enrichments of genes involved in incompatible P. effusa-spinach interactions. Differentially expressed genes with a False Discovery Rate (FDR) threshold of 0.1% and a fold change> 8.0 were used for GO enrichment analysis. The Enzyme Code and KEGG (Kyoto Encyclopedia of Genes and Genomes; Kanehisa Laboratories)36,37,38 analysis was performed using KEGG maps to find the biochemical pathways that are regulated by significantly expressed genes in the different treatments.
Single nucleotide polymorphism calling
The occurrence of single nucleotide polymorphisms (SNPs) in the expressed genes may modify the functional signature of these genes39,40. Therefore, we assessed SNPs in differentially expressed genes (FDR p-value = <0.01 from cultivars Solomon and Viroflay with minimum absolute fold change = 4) in spinach cultivars Solomon and Viroflay at 48 and 168 hpi, respectively. The variant detection toolbox in CLC Genomics Workbench 12 (http://www.clcbio.com) was used where the required variant probability was set to 90% with ploidy set to 2.
Results
Summary of transcript sequencing and mapping of sequence reads to the reference genome
A total of 364,359,078 sequence reads were generated from all samples included in this study using an Illumina sequencing platform (Supplementary Table 1). More than 95% reads were mapped to at least one position in the spinach reference genome for the majority of the samples, except for the 2nd and 3rd replications of cultivar Viroflay at 168 hpi, due to increased presence of pathogen transcripts sequenced in those samples. Furthermore, nearly 70% of total mapped reads from each sample were mapped to annotated genes within the spinach genome. Total gene reads include reads that span partly or entirely within an intron, an exon or an exon-exon junction, reads mapping fully within an exon, and reads that mapped to an exon junction. All other reads mapped partly or entirely between genes and were reported as mapped to the intergenic region. About 30% of total mapped reads from all samples were mapped to the intergenic regions in the genome.
The PCA showed clustering of expression values according to treatments i.e. un-inoculated control, P. effusa-inoculated 48 hpi, and P. effusa-inoculated 168 hpi, respectively (Fig. 2A). PC1, PC2, and PC3 specified varied gene counts from 26%, 16.7%, and 7.5%, respectively, from the replicated samples. A heatmap of hierarchical clustering showed that most genes expressed at higher levels in cultivar Solomon were expressed at lower levels in cultivar Viroflay and vice-versa, at 48 hpi and 168 hpi (Fig. 2B), clearly reflecting changes in metabolism and defense-associated gene expression during a late stage of pathogen colonization at 168 hpi, when leaf symptoms were present.
RNA-sequencing analyses of gene expression patterns in resistant and susceptible cultivars Solomon and Viroflay, respectively, inoculated with Peronospora effusa, or in the un-inoculated control treatment. (A) The principal component analysis of expression values across samples from cultivars Solomon and Viroflay in response to infection. (B) Heat map of expression values clustered in Solomon (Solo) and Viroflay (Viro) at 48 hours post-inoculation period (hpi) and 168 hpi, respectively. The expression value refers to normalized log counts per million, calculated for each gene.
Homologous species distribution
The reference spinach genome from cultivar Viroflay used in our analysis was 890,266,198 bp long and comprises of 34,878 genes (see materials and methods). The output received from the BLAST of the spinach gene set against the National Center Biotechnology Information database was used to develop the species distribution as determined by BLAST hits to validate transcriptome sequence identity (Supplementary Fig. 1). As anticipated, the maximum number of BLAST hits was associated primarily with Spinacia oleracea (84,705), followed by those of the two closely related species of Chenopodium quinoa (45, 614), and then Beta vulgaris subsp. vulgaris (29,727) (Supplementary Fig. 1).
Detection of single nucleotide polymorphisms in expressed genes in cultivars Solomon and Viroflay
In our experiments, we detected SNPs in differentially expressed genes at a frequency of 0.0003% and 0.0006%, at 48 hpi and 0.002% and 0.006% at 168 hpi in spinach cultivars Solomon and Viroflay, respectively. At 48 hpi, the frequencies of 0.5% to 0.3% SNPs and 0.4% to 0.1% SNPs at 168 hpi in cultivars Solomon and Viroflay, respectively, detected in the same locations in the genome. Though SNPs in expressed genes may modify their functional signatures39,40, the extent of SNPs in significantly differentially expressed genes in cultivars Solomon and Viroflay was negligible. The SNP frequencies detected in our studies were not anticipated to have significant effects on the expression profiles of the genes or their functions.
Overview of differential expression
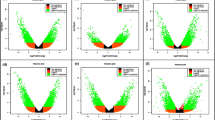
To determine the host response in resistant cultivar Solomon and susceptible cultivar Viroflay to P. effusa infection, differentially expressed genes were examined at an early stage of infection 48 hpi and at the commencement of disease symptoms and sporulation of pathogen at 168 hpi. At 168 hpi, no disease symptoms were observed in cultivar Solomon, but chlorosis together with sporulation was observed on the upper surface and underside of leaves, respectively, in cultivar Viroflay. A total of 23,063 and 23,364 genes were differentially expressed (Fold change = > ±1.0) at 48 and 168 hpi in cultivars Solomon and Viroflay, respectively (Supplementary File 1). However, only 416, 1,848, and 1,605 genes were expressed significantly (FDR p-value = <0.01 and minimum absolute fold change = 2) in accordance with the infection process (Fig. 3A; Venn diagram). A higher number of genes, 1387, was expressed uniquely at an early infection establishment stage at 48 hpi versus 1157 at 168 hpi. At 48 hpi, 310 genes were significantly up-regulated in Solomon versus Viroflay while 343 genes were down-regulated in Solomon versus Viroflay (Fig. 3B) (FDR p-value = <0.01 and minimum absolute fold change = 4). There was a marked shift in the proportions of up- and down-regulated genes at 168 hpi, reflecting the fact that Viroflay tissue was colonized and symptomatic at 168 hpi. At 168 hpi, 220 genes were significantly up-regulated while 890 genes were down-regulated (FDR p-value = <0.01 and minimum absolute fold change = 4) in Solomon versus Viroflay (Fig. 3B).
Venn diagram and Volcano plots of differentially expressed genes (DEGs), as determined by RNA-sequencing analysis in spinach cultivars Solomon (Peronospora effusa-resistant) and Viroflay (P. effusa-susceptible) at different time points after inoculation with the spinach downy mildew pathogen, P. effusa. (A) The Venn diagram displays significant differential gene expression values (FDR p-value = <0.01) with the minimum absolute fold change = 2. (B) Volcano plots of DEGs at 48 hours post-inoculation period (hpi) and 168 hpi, red circles indicate significant DEGs (FDR p-value = <0.01) with minimum absolute fold change = 4.
Among significantly expressed genes at 48 and 168 hpi in cultivar Solomon versus Viroflay, 85 were up-regulated and 112 were down-regulated at both time points while 214 genes that were up-regulated and 227 that were down-regulated were only expressed at 48 hpi. Similarly, there were 131 up-regulated and 767 down-regulated genes at 168 hpi in cultivar Solomon (Table 1). The list of significantly expressed genes at both 48 and 168 hpi time points or at either time point and their putative functions are presented in Supplementary File 2. To investigate the overall gene expression of the P. effusa-resistant cultivar Solomon and compare these values with the P. effusa-susceptible cultivar Viroflay, we combined expression values of each time point, and for each treatment of cultivar Solomon, and compared these individually with expression values in susceptible cultivar Viroflay at 48 hpi and 168 hpi, respectively. The up- or down-regulated expression patterns of the majority of the genes were similar to our results showing pairwise comparisons between the two cultivars at each time point.
Differential expression of defense-associated genes
Plants have evolved specialized defense mechanisms which can recognize and discriminate between different environmental stresses or even different biotic threats24. Plant pathogens trigger up-regulation of defense genes, which may produce products that are directly antimicrobial or stimulate the biochemical pathways capable of producing antimicrobial metabolites26,40. Up-regulated genes significantly expressed at both time points, 48 and 168 hpi, may have roles in plant immunity, including the receptor-like protein kinase FERONIA, leucine-rich repeat (LRR) receptor-like serine/threonine-protein kinase, serine/threonine-protein kinase TIO (Supplementary File 2). Additionally, several genes encoding for transcription factor basic-helix-loop-helix (bHLH)-like or proteins with unknown function were also significantly up-regulated across both time points.
Furthermore, potential defense-related up-regulated genes expressed only at 48 hpi included homologs of the serine/threonine-protein kinase EDR10, LRR receptor-like serine/threonine-protein kinase, receptor-like serine/threonine-protein kinase ALE2, LRR receptor-like serine/threonine-protein kinase FLS2, and defensin-like protein. Similarly, up-regulated genes expressed uniquely at 168 hpi and probably crucial for the defense during late stages of downy mildew infection were ankyrin repeat-containing protein, 1-deoxy-D-xylulose-5-phosphate synthase, and defensin-like protein AX1. Six genes associated with early response to the pathogen were significantly expressed across both time points; however, these were induced at 48 hpi and repressed later at 168 hpi (Table 2). We also assessed the expression profile of common sets of genes relating to plant immunity such as protein kinases, pathogenesis related (PR) proteins, WRKY transcription factors, and ethylene- esponsive transcription factors or ethylene synthesizing enzymes. When compared to the fold change differences in expression values of resistant cultivar Solomon versus susceptible cultivar Viroflay, most of these genes were up-regulated at 48 hpi but down-regulated at 168 hpi (Supplementary file 3). Specifically, genes encoding for 1-aminocyclopropane-1-carboxylic acid (ACC) oxidase and ethylene transcription factors were significantly induced at 48 hpi but only a few of these genes were induced at 168 hpi. Two terminal enzymes in the phenylpropanoid pathway are chalcone synthase and cinnamoyl-CoA reductase, which are responsible for biosynthesis of flavonoids and lignin, respectively, in plants41. Genes encoding for cinnamoyl-CoA reductase 1-like were induced at 48 hpi but both cinnamoyl-CoA reductase and chalcone synthase-like genes were highly repressed at 168 hpi (Table 3).
A total of 100 and 78 significantly up-regulated genes (FDR p-value = <0.001 and Fold change = > 8.00) at 48 and 168 hpi, respectively, were used to assign GO terms and summarize their functional description. At 48 hpi, 155 GO terms were assigned, of which 60 were associated with biological process, 34 cellular component, and 61 molecular function. At 168 hpi, 126 GO terms were assigned, of which 45 were assigned to the biological process, 27 cellular component, and 54 molecular function (Supplementary File 4). GO enrichment analysis was performed using up-regulated genes at 48 and 168 hpi. At 48 hpi, specific GO terms were associated with up-regulated genes that may have roles in defense responses, wounding, toxin activity, pathogenesis, systemic acquired resistance, glutathione transferase activity, etc. (Fig. 4A). At 168 hpi, significantly enriched GO terms associated with host defense responses at late stages of infection were terpenoid biosynthetic process, alpha-glucan, water dikinase activity, response to wounding etc. (Fig. 4B). The GO terms including negative regulation of endopeptidase activity, serine-type endopeptidase inhibitor activity, response to wounding, DNA repair, and sulfotransferase activity were enriched at both the 48 and 168 hpi time points.
Gene ontology (GO) term enrichment derived from Peronospora effusa-spinach interactions, in which both resistant and susceptible cultivars were compared. (A) GO term enrichment at 48 hours post-inoculation period (hpi), and (B) GO term enrichment at 168 hpi resistant cultivar Solomon versus susceptible Viroflay. GO term enrichment was computed using significantly up-regulated DEGs at 48 and 168 hpi, respectively. Histograms with gold color indicate GO terms enriched across both time points.
KEGG pathway analysis was performed using genes significantly up- and down-regulated (FDR p-value = <0.001 and minimum absolute fold change = 8.00) at 48 and 168 hpi, respectively. The pathway analysis identified 13 and 5 up-regulated genes involved in 14 metabolic pathways at 48 and 168 hpi, respectively. Likewise, 21 and 91 down-regulated genes in 33 metabolic pathways, at 48 and 168 hpi, respectively (Supplementary Table 2). At both 48 and 168 hpi, but particularly at 168 hpi, there were multiple down-regulated genes that were identified as involved in thiamine metabolism and phenylpropanoid pathways.
Expression profiling of the P. effusa transcriptome during incompatible and compatible interactions
The RNA-seq reads mapped to the genome of P. effusa race 13 were used primarily to examine the temporal expression patterns of putative effector proteins, which may influence how the pathogen infects and colonizes spinach. At 48 hpi, less than 1% of total reads retrieved from susceptible cultivar Viroflay were mapped to the P. effusa genome, while at 168 hpi, nearly 25 to 48% of total reads were mapped to the P. effusa genome. This is in contrast to the resistant cultivar Solomon, where the mapping percentage was approximately the same at both 48 hpi and 168 hpi but less than 1% of total reads. In the Viroflay samples, several P. effusa genes were strongly expressed at 168 hpi but only a few genes were activated and weakly expressed in the samples derived from the cultivar Solomon (Heatmap; Fig. 5A). Differential expression of genes encoding homologs of RxLR and crinklers, two major effector protein families in oomycetes, was calculated in Viroflay using expression values at 168 and 48 hpi. Interestingly, only effector proteins of the crinkler family were significantly up-regulated (FDR p-value = <0.05) while RxLR effectors were significantly down-regulated when expression values were compared at 168 hpi versus 48 hpi (Fig. 5B). We performed BLASTX35 searches of sequences encoding these putative effectors and checked for homologs in the Phytophthora infestants T30–4 database42. These putative effector proteins from P. effusa race 13 shared a substantial homology with RxLR and crinkler proteins of P. infestants T30-4. The RxLR and crinkler effectors from P. effusa shared 30 to 82% homology with RxLR proteins from P. infestans and 50% homology with the identified Crinklers of P. infestans.
Expression profile and differential expression of Peronospora effusa transcriptomes/translated proteins during incompatible and compatible interactions with resistant spinach cultivar Solomon and susceptible cultivar Viroflay, respectively. (A) Heat map of expression values clustered in resistant cultivar Solomon and susceptible cultivar Viroflay at 168 hours post-inoculation period (hpi). (B) Fold change of putative P. effusa effectors (168 hpi versus 48 hpi) in response to infection in susceptible cultivar Viroflay.
Additional pathogen genes of interest included those encoding peroxidase/catalases, peroxide reductases, and several NPP1 type necrosis inducing-like proteins that were identified at the late infection stage of 168 hpi (Supplementary File 5). Each of these types of proteins may contribute to host colonization since catalase-peroxidases are important for virulence in some plant pathogens by protecting against hydrogen peroxide produced in the interaction with the host43.
Infectivity of P. effusa in susceptible cultivar Viroflay at 48 hpi and 168 hpi
Sporangia of P. effusa germinated overnight, producing germ tubes that can potentially invade the leaf surface (Supplementary Fig. 2A). The germ tube differentiated and grew rapidly, generating networks of infectious hyphae in the susceptible cultivar Viroflay (Supplementary Fig. 2B).
Discussion
The recent advances in RNA sequencing have enabled unprecedented depth and accuracy for transcriptome analyses. Using this powerful approach, differential gene expression associated with resistant and susceptible crop genotypes has been examined in various pathosystems to understand the molecular basis of plant defense responses19,44,45,46,47,48. In this study, we applied this tool along with the newly available spinach and P. effusa genome resources, to analyze gene expression during incompatible and compatible spinach- P. effusa interactions. We used resistant and susceptible spinach cultivars Solomon and Viroflay, respectively, and inoculated with P. effusa and monitored the temporal pattern of gene expression at 48 and 168 hpi, respectively. We examined genes expressed in both host (spinach cultivars) and pathogen (P. effusa) sides during host-microbe interactions, at 48 and 168 hpi.
Diverse and distinct spinach gene sets were expressed during incompatible interactions between P. effusa and cultivar Solomon versus compatible interactions between P. effusa and cultivar Viroflay. Genes significantly up-regulated at 48 and 168 hpi in cultivar Solomon were of particular interest since these were hypothesized as critical for resistance to pathogen invasion. Plant immunity, which may occur during an incompatible interaction, operates at two levels: a basal immunity that is regulated through pattern recognition receptors upon perception of pathogen-associated molecular patterns and a race or cultivar-specific resistance mediated through resistance genes that recognize and inactivate pathogen-derived effector molecules26,41. In spinach, race-specific resistance has been reported to the downy mildew but the functional mechanisms of host resistance remain undetermined28. During incompatible plant-pathogen interactions, plant R proteins recognize and inactivate pathogen-derived effector molecules26,41, which often leads to the hypersensitive cell death at the site of pathogen attack27,49. The hypersensitive response is often associated with extensive transcriptional reprogramming and activation of several defense genes to sustain the immunity27,50. Plant R genes may encode a combination of conserved domains, including LRRs, protein kinases, nucleotide binding site (NBS)s, and in at least one case, a WRKY transcription factor domain47,51. These domains confer the potential to perceive and signal inputs from external stimuli27,49,52. Furthermore, there is increasing evidence that receptor-like kinases regulate plant growth and development and also modulate defense signaling pathways by sensing modifications in plant cell wall architecture53,54,55,56. Also, phytohormones, mainly salicylic acid, ethylene and jasmonic acid mediated signaling networks, have been explored regarding their significance in the induction and maintenance of plant immunity22.
In our study, we found significant upregulation of genes encoding a zinc finger CCCH protein, ACC oxidase, receptor-like protein kinase FERONIA, LRR receptor-like serine/threonine-protein kinase, and serine/threonine-protein kinase TIO during early (48 hpi) and late (168 hpi) stages of infections in the resistant cultivar Solomon. The occurrence of zinc finger domains has been documented in many disease resistance genes that are required for host resistance during incompatible interactions57. The receptor-like kinase FERONIA is not only important for normal plant growth and reproduction, but also induces the liberation of reactive oxygen species in response to the changes in the cell wall58,59. We speculate that the encoded FERONIA homolog in spinach may be a first responder to P. effusa as sporangia germinate and initially invade through the cell walls in spinach leaves by direct penetration9. The instantaneous increase of reactive oxygen species at the infection site, possibly activated the enzyme glycosyltransferase, and involves the biosynthesis and strengthening of plant cell walls60. The upregulation of genes associated with glycosyltransferase activity has been reported upon inoculation of Arabidopsis plants with plant pathogens Botrytis cinerea and P. infestans61. The enriched GO term ‘response to wounding’ at both 48 and 168 hpi also suggested that preserving cell wall integrity is an important step to retain the resistance over P. effusa invasion.
The significant upregulation of genes associated with PR-1 protein, glutathione-S-transferase, phospholipid hydroperoxide glutathione peroxidase, peroxidase was observed particularly at 48 hpi which are reported to minimize the oxidative damage in plants in response to biotic and abiotic stresses62,63. Homologs of these gene products restrict the oxidative damage in host tissue and initiate several succeeding downstream signaling cascades62,64. The PR-1 gene has been thoroughly studied in the past for its role in defense to different pathogens, including P. infestans65,66. Additionally, genes encoding R-proteins such as LRR receptor-like serine/threonine-protein kinase, ankyrin repeat-containing protein NPR4-like, G-type lectin S-receptor-like serine/threonine-protein kinase, and nucleoside diphosphate kinase 2 were highly expressed at 48 hpi in cultivar Solomon compared to Viroflay. It is possible that extracellular domains of these receptors physically interact with the P. effusa specific PAMP molecules and activate the downstream defense signals that are critical to subsequently suppress the infection process. Some of these genes may be targeted by inhibitory RNAs or CRISPR-Cas9 approaches to assess their functions during a resistant response.
Certain host genes, including those encoding transcription factors, were highly differentially expressed in this study, in susceptible and resistant spinach-P. effusa interactions. The members of the bHLH family are involved in cooperating, through various phytohormone-regulated signaling pathways to regulate plant immunity67,68. We found significant up-regulation of genes encoding transcription factor bHLH3-like proteins in the resistant interaction at both 48 and 168 hpi. Spinach homologs of those genes encoding plant defensin-like proteins were also strongly expressed during the incompatible interaction, indicating their involvement in defense in this interaction. A number of studies have shown the inhibitory effect of plant defensin peptides to various fungal and bacterial pathogens69,70,71,72. Genes encoding ACC oxidase homologs in spinach were up-regulated across the infection times in our experiments. ACC oxidase is involved in the biosynthesis of the plant hormone ethylene and the presence of ethylene subsequently activates the downstream genes which can inactive the effector proteins and restore the plant immunity73,74. The increased synthesis of ethylene has been reported as one of the earliest responses of plants to a wide variety of plant pathogens, including downy mildew pathogens4875,76. A gene encoding for AP2-like ethylene-responsive transcription factor was also significantly up-regulated at 48 hpi and was reported to have an important role in conferring resistance to plant pathogens77,78. The expression of AP2 transcription factor is activated by plant hormones including ethylene, which induces pathogenesis-related genes and activates defense signaling pathways74,79.
The flavonoid and phenylpropanoid pathways play a central role in plant defense by providing structural barriers through lignification of cell walls and in establishing chemical barriers by activating defense genes involving in signaling networks for resistance to pathogen infection80,81. In the case of grape downy mildew, a thiamine-induced phenylpropanoid pathway was reported as a crucial mechanism to confer resistance to the Plasmopara viticola infection82. At 168 hpi, there was notable up-regulation in the resistant interaction of gene Spiol06Chr03650, encoding a probable 1-deoxy-D-xylulose-5-phosphate synthase (DXS). DXS catalyzes an early step in the isoprenoid biosynthetic pathway, which results in carotenoid and chlorophyll production in plants. In earlier studies, DXS transcripts were negatively correlated with late blight symptoms in potato, indicating its potential role in host defense against plant pathogens83. The gene encoding enzyme cinnamoyl-CoA reductase, a key enzyme in the lignin biosynthesis, was significantly up-regulated in the resistant cultivar Solomon at 48 hpi, which may have a role in lignification to confer tolerance to pathogen invasion. Conversely, at the 168 hpi, down-regulated genes associated with the phenlypropanoid biosynthesis pathway were highly expressed in the susceptible cultivar Viroflay versus cultivar Solomon. Down-regulation of key genes involved in phenlypropanoid biosynthesis may assist in pathogen proliferation. This down-regulation may be due to the delivery of pathogen effector proteins that target and suppress the expression of these host genes.
Examination of P. effusa revealed highly expressed genes encoding for RxLR and crinkler effectors, which were especially prevalent in the interaction with the susceptible cultivar Viroflay. A recent study revealed that the P. effusa genomes comprise various potential PAMP and effector molecules which can suppress the host immunity16. These effector proteins share significant sequence similarity with known effectors of P. infestans42. Our results suggested that RxLR effectors were actively expressed at the early stage of the infection while Crinkler effectors were active at the late stage of the infection. In this current study, RxLR effectors were associated with the vegetative invasive stage in the pathogen, likely facilitating host colonization and acquisition of nutrients from the host. The significant expression of RxLR effector genes was also reported at the early stage of infection (12 hpi) when susceptible potato plants were inoculated with virulent strains of P. infestans84. Thus, the expression of Crinkler effectors identified in this study at the late stage of infection may affect the host physiology to initiate the disease symptoms and the expansion of infected areas. Additional data may be mined from this data set generated in this study to further study mechanisms of infectious hyphal growth during spinach-P. effusa interactions. Though this awaits further validation, the peroxidase/catalases, peroxide reductases, and the NPP1 homolog expressed in the pathogen during late stages of infection may reflect the mounting effort on the part of the pathogen to combat high levels of certain reactive oxygen species produced by the plant, and also to inflict cell death at the late stage of infection when symptoms including both chlorosis and necrosis may be present. In Arabidopsis and parsley, NPP1 from Phytophthora can induce host defenses gene expression and the production of reactive oxygen species, and cause cell death85,86.
Mechanisms of downy mildew resistance in spinach were identified in this study, along with the candidate genes that may be useful in projects that aim to breed downy mildew resistance. Hypersensitive response-inducible genes such as PR-1 and glutathione-S-transferase and protein kinase FERONIA were strongly up-regulated in the resistant cultivar Solomon in response to P. effusa, and thus defense responses that occur via salicylic acid signaling and reactive oxygen species are likely similar to those described in other plant species87,88,89. Furthermore, multiple genes related to phenylpropanoid pathways were significantly down-regulated in the susceptible interaction and genes encoding ACC oxidase and AP2-like ethylene-responsive transcription factor were up-regulated in the resistant interaction. Thus, it is plausible that crosstalk between salicylic acid and ethylene signaling pathways and cell wall lignification undergirds the resistant interaction, but this requires further validation. In future studies, we will investigate the potential role of ethylene-based signaling in spinach cultivars, especially in relation to ethylene-modulated kinase signaling events conferring tolerance or resistance to pathogen invasion. This study and other studies that attempt to unravel the molecular basis of disease or resistance in the spinach-P. effusa interaction may enable development of effective management strategies for spinach downy mildew.
Data availability
The RNA-seq data generated in this study have been deposited at National Center for Biotechnology Information under the BioProject identification number PRJNA603356. The reference spinach genome can be accessed through the repository at https://github.com/USDA-ARS-GBRU/Spinach_Peffusa.
References
FAOSTAT. Production domain. In: Crops. FAO, Rome; retrieved from, http://www.fao.org/faostat/en/#data/QC, (2018).
Atallah, Z. K. et al. Population analyses of the vascular plant pathogen Verticillium dahliae detect recombination & transcontinental gene flow. Fungal Genet. Biol. 47, 416–422 (2010).
Klosterman, S. J. Spinach downy mildew: threat, prevention & control. Prog. Crop Consult. 1, 13–15 (2016).
CDFA, California Agricultural Statistics Review, 2017-2018; retrieved from, https://www.cdfa.ca.gov/statistics/PDFs/2017-18AgReport.pdf, (2018).
Farnsworth, K. L. & Milliman, J. D. Effects of climatic and anthropogenic change on small mountainous rivers: the Salinas River example. Glob. Planet. Change 39, 53–64 (2003).
Thines, M. & Choi, Y.-J. Evolution, diversity, & taxonomy of the Peronosporaceae, with focus on the genus Peronospora. Phytopathology 106, 6–18 (2016).
Byford, W. J. Host specialization of Peronospora farinosa on Beta, Spinacia and Chenopodium. Trans. Br. Mycol. Soc. 50, 603–607 (1967).
Klosterman, S. J. et al. Coupling spore traps and quantitative PCR assays for detection of the downy mildew pathogens of spinach (Peronospora effusa) and beet (P. schachtii). Phytopathology 104, 1349–1359 (2014).
Kandel, S. L. et al. Spinach Downy Mildew: Advances in our understanding of the disease cycle and prospects for disease management. Plant Dis. 103, 791–803 (2019).
Kunjeti, S. G., Anchieta, A., Subbarao, K., Koike, S. T. & Klosterman, S. J. Plasmolysis and vital staining reveal viable oospores of Peronospora effusa in spinach seed lots. Plant Dis. 100, 59–65 (2016).
Choudhury, R. A. et al. Season-long dynamics of spinach downy mildew determined by spore trapping & disease incidence. Phytopathology 106, 1311–1318 (2016).
Feng, C. et al. New races & novel strains of the spinach downy mildew pathogen Peronospora effusa. Plant Dis. 102, 613–618 (2018).
Correll, J. C. et al. Spinach: Better management of downy mildew & white rust through genomics. Eur. J. Plant Pathol. 129, 193–205 (2011).
Ojiambo, P. S., Gent, D. H., Quesada-Ocampo, L. M., Hausbeck, M. K. & Holmes, G. J. Epidemiology & population biology of Pseudoperonospora cubensis: A model system for management of downy mildews. Annu. Rev. Phytopathol. 53, 223–246 (2015).
Salgado Salazar, C., Shishkoff, N., Daughtrey, M. L., Palmer, C. & Crouch, J. A. Downy mildew: a serious disease threat to rose health worldwide. Plant Dis. 10, 1873–1882 (2018).
Fletcher, K. et al. Comparative genomics of downy mildews reveals potential adaptations to biotrophy. BMC genomics, 19 (2018).
Feng, C. et al. Genome sequences of three races of Peronospora effusa: A resource for studying the evolution of the spinach downy mildew pathogen. MPMI 31, 1230–1231 (2018).
Adhikari, B. N. et al. Expression profiling of Cucumis sativus in response to infection by Pseudoperonospora cubensis. PloS One 7, e34954 (2012).
Bhattarai, K., Wang, W., Cao, Z. & Deng, Z. Comparative analysis of impatiens leaf transcriptomes reveal candidate genes for resistance to downy mildew caused by Plasmopara obducens. Int. J. Mol. Sci. 19, 2057 (2018).
Chen, T. et al. Comparative transcriptome profiling of a resistant vs. susceptible tomato (Solanum lycopersicum) cultivar in response to infection by tomato yellow leaf curl virus. PloS One 8, e80816 (2013).
Joshi, R. K., Megha, S., Rahman, M. H., Basu, U. & Kav, N. N. A global study of transcriptome dynamics in canola (Brassica napus L.) responsive to Sclerotinia sclerotioruminfection using RNA-Seq. Gene 590, 57–67 (2016).
Mine, A. et al. The defense phytohormone signaling network enables rapid, high-amplitude transcriptional reprogramming during effector-triggered immunity. Plant Cell 30, 1199–1219 (2018).
Pombo, M. A. et al. Transcriptome-based identification & validation of reference genes for plant-bacteria interaction studies using Nicotiana benthamiana. Sci. Rep. 9, 1632 (2019).
Jones, J. D. & Dangl, J. L. The plant immune system. Nature 444, 323–329 (2006).
Dodds, P. N. & Rathjen, J. P. Plant immunity: towards an integrated view of plant–pathogen interactions. Nat. Rev. Genet. 11, 539 (2010).
Zhang, Y., Lubberstedt, T. & Xu, M. The genetic & molecular basis of plant resistance to pathogens. J. Genet. Genomics 40, 23–35 (2013).
Bent, A. F. & Mackey, D. Elicitors, effectors, and R genes: the new paradigm and a lifetime supply of questions. Annu. Rev. Phytopathol. 45, 399–436 (2007).
She, H. et al. Fine mapping and candidate gene screening of the downy mildew resistance gene RPF1 in Spinach. Theor. Appl. Genet. 131, 2529–2541 (2018).
Feng, C., Correll, J. C., Kammeijer, K. E. & Koike, S. T. Identification of new races and deviating strains of the spinach downy mildew pathogen Peronospora farinosa f. sp. spinaciae. Plant Dis. 98, 145–152 (2014).
Zhong, S. et al. High-throughput illumina strand-specific RNA sequencing library preparation. Cold Spring Harb. Protoc, https://doi.org/10.1101/pdb.prot5652 (2011).
Myers, E. et al. A whole-genome assembly of Drosophila. Science 287, 2196–2204 (2000).
Cantarel, B. L. et al. MAKER: an easy-to-use annotation pipeline designed for emerging model organism genomes. Genome Res. 18, 188–196 (2008).
Liu, J. J. et al. Transcriptome analysis of the white pine blister rust pathogen Cronartium ribicola: de novo assembly, expression profiling, and identification of candidate effectors. BMC Genomics 16, 1–16 (2015).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change & dispersion for RNA-seq data with DESeq 2. Genome Biol. 15, 550 (2014).
Altschul, S. F., Gish, W., Miller, W., Myers, E. W. & Lipman, D. J. Basic local alignment search tool. J. Mol. Biol. 215, 403–410 (1990).
Kanehisa, M. & Goto, S. KEGG: Kyoto Encyclopedia of Genes and Genomes. Nucleic Acids Res. 28, 27–30 (2000).
Kanehisa, M., Sato, Y., Furumichi, M., Morishima, K. & Tanabe, M. New approach for understanding genome variations in KEGG. Nucleic Acids Res. 47, D590–D595 (2019).
Kanehisa, M. Toward understanding the origin and evolution of cellular organisms. Protein Sci. 28, 1947–1951 (2019).
Neums, L. et al. VaDiR: an integrated approach to Variant Detection in RNA. Gigascience 7, 1–13 (2017).
Zhao, Y. et al. A high-throughput SNP discovery strategy for RNA-seq data. BMC Genomics 20, 160 (2019).
Cui, H., Tsuda, K. & Parker, J. E. Effector-triggered immunity: from pathogen perception to robust defense. Annu. Rev. of Plant Biol. 66, 487–511 (2015).
Deng, Y. & Lu, S. Biosynthesis and regulation of phenylpropanoids in plants. Crit. Rev. Plant Sci. 36, 257–290 (2017).
Haas, B. J. et al. Genome sequence and analysis of the Irish potato famine pathogen Phytophthora infestans. Nature 461, 393 (2009).
Jittawuttipoka, T., Buranajitpakorn, S., Vattanaviboon, P. & Mongkolsuk, S. The catalase-peroxidase KatG is required for virulenceof Xanthomonas campestris pv. campestris in a host plant by providing protection against low levels of H2O2. J. Bacteriol. 191, 7372–7377 (2009).
Goyer, A., Hamlin, L., Crosslin, J. M., Buchanan, A. & Chang, J. H. RNA-Seq analysis of resistant and susceptible potato varietiesduring the early stages of potato virus Y infection. BMC Genomics 16, 472 (2015).
Kamber, T. et al. Fire blight disease reactome: RNA-seq transcriptional profile of apple host plant defense responses to Erwiniaamylovora pathogen infection. Sci. Rep. 6, 21600 (2016).
Li, X., Kapos, P. & Zhang, Y. NLRs in plants. Curr. Opin Immunl. 32, 114–121 (2015).
Toffolatti, S. L. et al. Unique resistance traits against downy mildew from the center of origin of grapevine (Vitis vinifera). Sci. Rep. 8, 12523 (2018).
Oh, S. K. et al. In planta expression screens of Phytophthora infestans RXLR effectors reveal diverse phenotypes, including activation of the Solanum bulbocastanum disease resistance protein Rpi-blb2. Plant Cell 21, 2928–2947 (2009).
Adachi, H. & Tsuda, K. Convergence of cell-surface and intracellular immune receptor signaling. New Phytol. 221, 1676–1678 (2019).
Nurnberger, T. & Kemmerling, B. Receptor protein kinases–pattern recognition receptors in plant immunity. Trends Plant Sci. 11, 519–522 (2006).
Wu, C. H., Derevnina, L. & Kamoun, S. Receptor networks underpin plant immunity. Science 360, 1300–1301 (2018).
Afzal, A. J., Wood, A. J. & Lightfoot, D. A. Plant receptor-like serine threonine kinases: roles in signaling and plant defense. MPMI 21, 507–517 (2008).
De Smet, I., Voss, U., Jurgens, G. & Beeckman, T. Receptor-like kinases shape the plant. Nat. Cell Biol. 11, 1166 (2009).
Pogorelko, G., Lionetti, V., Bellincampi, D. & Zabotina, O. Cell wall integrity: targeted post-synthetic modifications to reveal its role in plant growth and defense against pathogens. Plant Signal Behav. 8, 1–8 (2013).
Savatin, D. V., Gramegna, G., Modesti, V. & Cervone, F. Wounding in the plant tissue: the defense of a dangerous passage. Front. Plant Sci. 5, 1–11 (2014).
Gupta, S. K., Rai, A. K., Kanwar, S. S. & Sharma, T. R. Comparative analysis of zinc finger proteins involved in plant disease resistance. PLoS One 7, e42578 (2012).
Cheung, A. Y. & Wu, H. M. THESEUS 1, FERONIA and relatives: a family of cell wall-sensing receptor kinases? Curr. Opin. Plant Biol. 14, 632–641 (2011).
Li, C., Wu, H. M. & Cheung, A. Y. FERONIA and her pals: functions & mechanisms. Plant Physiol. 171, 2379–2392 (2016).
Persson, S. et al. The Arabidopsis irregular xylem8 mutant is deficient in glucuronoxylan and homogalacturonan, which are essential for secondary cell wall integrity. Plant Cell 19, 237–255 (2007).
Langlois-Meurinne, M., Gachon, C. M. & Saindrenan, P. Pathogen-responsive expression of glycosyltransferase genes UGT73B3 & UGT73B5 is necessary for resistance to Pseudomonas syringae pv. tomato in Arabidopsis. Plant Physiol. 139, 1890–1901 (2005).
Nianiou-Obeidat, I. et al. Plant glutathione transferase-mediated stress tolerance: functions and biotechnological applications. Plant Cell Rep. 36, 791–805 (2017).
Gullner, G., Komives, T., Kiraly, L. & Schroder, P. Glutathione S-transferase enzymes in plant-pathogen interactions. Front. Plant Sci. 9, 1–19 (2018).
Agrawal, G. K., Rakwal, R., Jwa, N. S. & Agrawal, V. P. Effects of signaling molecules, protein phosphatase inhibitors and blast pathogen (Magnaporthe grisea) on the mRNA level of a rice (Oryza sativa L.) phospholipid hydroperoxide glutathione peroxidase (OsPHGPX) gene in seedling leaves. Gene 283, 227–236 (2002).
Breen, S., Williams, S. J., Outram, M., Kobe, B. & Solomon, P. S. Emerging insights into the functions of pathogenesis-related protein 1. Trends Plant Sci. 22, 871–879 (2017).
Saubeau, G., Perrin, F., Marnet, N., Andrivon, D. & Val, F. Hormone signalling pathways are differentially involved in quantitative resistance of potato to Phytophthora infestans. Plant Pathol. 65, 342–352 (2016).
Xu, F. et al. NLR-associating transcription factor bHLH84 and its paralogs function redundantly in plant immunity. PLoS Pathogens 10, 1–14 (2014).
Tsuda, K. & Somssich, I. E. Transcriptional networks in plant immunity. New Phytol. 206, 932–947 (2015).
Sathoff, A. E., Velivelli, S., Shah, D. M. & Samac, D. A. Plant defensin peptides have antifungal and antibacterial activity against human and plant pathogens. Phytopathology 109, 402–408 (2019).
Odintsova, T. I. et al. Defensin-like peptides in wheat analyzed by whole-transcriptome sequencing: a focus on structural diversity and role in induced resistance. PeerJ 7, e6125 (2019).
Van der Weerden, N. L. & Anderson, M. A. Plant defensins: common fold, multiple functions. Fungal Biol. Rev. 26, 121–131 (2013).
Wong, J. H. & Ng, T. B. Limenin, a defensin-like peptide with multiple exploitable activities from shelf beans. J. Pept. Sci. 12, 341–346 (2006).
Dubois, M., Van den Broeck, L. & Inze, D. The pivotal role of ethylene in plant growth. Trends Plant Sci. 23, 311–323 (2018).
Wang, K. L. C., Li, H. & Ecker, J. R. Ethylene biosynthesis and signaling networks. Plant Cell 14, S131–S151 (2002).
Nie, X., Singh, R. P. & Tai, G. C. Molecular characterization & expression analysis of 1-aminocyclopropane-1-carboxylate oxidase homologs from potato under abiotic and biotic stresses. Genome 45, 905–913 (2002).
Abiri, R. et al. 2017. Role of ethylene and the APETALA 2/ethylene response factor superfamily in rice under various abiotic and biotic stress conditions. Environ. Exper. Bot. 134, 33–44 (2017).
Gutterson, N. & Reuber, T. L. Regulation of disease resistance pathways by AP2/ERF transcription factors. Curr. Opin. Plant Biol. 7, 465–471 (2004).
Park, J. M. et al. Overexpression of the tobacco Tsi1 gene encoding an EREBP/AP2–type transcription factor enhances resistance against pathogen attack and osmotic stress in tobacco. Plant Cell 13, 1035–1046 (2001).
Licausi, F., Ohme-Takagi, M. & Perata, P. APETALA 2/Ethylene responsive factor (AP 2/ERF) transcription factors: Mediators of stress responses and developmental programs. New Phytol. 199, 639–649 (2013).
Fraser, C. M. & Chapple, C. The phenylpropanoid pathway in Arabidopsis. The Arabidopsis Book 9, American Society of Plant Biologists, https://doi.org/10.1199/tab.0152, (2011).
Dixon, R. A. et al. The phenylpropanoid pathway and plant defense - a genomics perspective. Mol. Plant Pathol. 3, 371–390 (2002).
Boubakri, H. et al. Thiamine modulates metabolism of the phenylpropanoid pathway leading to enhanced resistance to Plasmopara viticola in grapevine. BMC Plant Biol. 13, 1–15 (2013).
Henriquez, M. A. et al. Molecular cloning, functional characterization & expression of potato (Solanum tuberosum) 1-deoxy-d-xylulose 5-phosphate synthase 1 (StDXS1) in response to Phytophthora infestans. Plant Sci. 243, 71–83 (2016).
Yin, J. et al. Conserved RXLR effector genes of Phytophthora infestans expressed at the early stage of potato infection are suppressive to host defense. Front. Plant Sci. 8, 1–11 (2017).
Galiana, E. et al. Plant-induced cell death in the oomycete pathogen Phytophthora parasitica. Cell Microbiol. 7, 1365–1378 (2005).
Fellbrich, G. et al. NPP1, a Phytophthora-associated trigger of plant defense in parsley and Arabidopsis. Plant J. 32, 375–390 (2002).
Derksen, H., Rampitsch, C. & Daayf, F. Signaling cross-talk in plant disease resistance. Plant Sci. 207, 79–87 (2013).
Herrera-Vasquez, A., Salinas, P. & Holuigue, L. Salicylic acid and reactive oxygen species interplay in the transcriptional control of defense genes expression. Front. Plant Sci. 6, 1–9 (2015).
Zander, M., Thurow, C. & Gatz, C. TGA transcription factors activate the salicylic acid-suppressible branch of the ethylene-induced defense program by regulating ORA59 expression. Plant Physiol. 165, 1671–1683 (2014).
Acknowledgements
We thank the University of California Seed Biotechnology Center, the USDA-NIFA-AFRI Food Security grant through Michigan State University (Award No. 2016-68004-24931), and the USDA-NIFA Specialty Crop Research Initiative (Award No. 2017-51181-26830) through University of Arkansas for funding. We also thank Roberto Ornelas (California State University, Monterey Bay) for analyses of RNA-Seq data prior to completion of the final spinach reference genome annotation. Mention of trade names or commercial products in this publication is solely for the purpose of providing specific information and does not imply recommendation or endorsement by the United States Department of Agriculture (USDA). USDA is an equal opportunity provider and employer.
Author information
Authors and Affiliations
Contributions
S.J.K. and A.V. conceived the study and designed experiments; S.L.K. wrote the manuscript, conducted the RNA-seq analyses, and prepared all of the figures, tables, and supplemental files; S.T.K. directed the differential screening to identify downy mildew resistant and susceptible spinach with assistance from S.J.K., K.S. isolated RNA and prepared cDNA libraries; A.M.H.-K. annotated the spinach genome used in this study; S.J.K., B.M., and A.S. contributed to the writing of the manuscript. All authors commented on and approved the final version of the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Kandel, S.L., Hulse-Kemp, A.M., Stoffel, K. et al. Transcriptional analyses of differential cultivars during resistant and susceptible interactions with Peronospora effusa, the causal agent of spinach downy mildew. Sci Rep 10, 6719 (2020). https://doi.org/10.1038/s41598-020-63668-3
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-020-63668-3
- Springer Nature Limited
This article is cited by
-
Comparative Transcriptome Analysis of Resistant and Susceptible Genotypes of Isabgol (Plantago ovata) During Interactions with Peronospora plantaginis, the Causal Agent of Downy Mildew Disease
Plant Molecular Biology Reporter (2024)
-
Transcriptomics: illuminating the molecular landscape of vegetable crops: a review
Journal of Plant Biochemistry and Biotechnology (2024)
-
Plant biomarkers as early detection tools in stress management in food crops: a review
Planta (2024)
-
Integration of early disease-resistance phenotyping, histological characterization, and transcriptome sequencing reveals insights into downy mildew resistance in impatiens
Horticulture Research (2021)