Abstract
Apomixis is an asexual reproduction system without fertilization, which is an important proliferation strategy for plants and algae. Here, we report on the apomeiosis in the green seaweed Ulva prolifera, which has sexual and obligate asexual populations. Genomic PCR of mating type (MT)-locus genes revealed asexual thalli carrying both MT genomes. Observation of the chromosomes during the formation of each type of reproductive cell revealed that cells in asexual thalli performed apomeiosis without chromosome reduction. Moreover, genotyping revealed that laboratory-cultured sporophytic thalli produced not only each type of gametophyte but also diploid thalli carrying the mt− and mt+ genome (mt± thallus strains). The mt± thallus strain released diploid biflagellate zoids, with ultrastructure and behavior similar to mt+ gametes. Additionally, a transcriptomic analysis revealed that some meiosis-related genes (Mei2L and RAD1) were highly expressed in the quadriflagellate zoosporoids. Our results strongly suggest that asexual thalli originally evolved via apomeiosis in sporophytic thalli.
Similar content being viewed by others
Introduction
Plants, including flowering plants, ferns, and bryophytes, produce their progenies via sexual or asexual reproduction systems. Apomixis is an asexual reproduction in which mother plants produce genetically identical progenies in sexual organs without fertilization. The process of apomictic seed development in land plants consists of three distinct steps: apomeiosis (avoidance of meiosis), parthenogenesis (development of an embryo without fertilization), and functional endosperm development1,2. The elucidation of the mechanism of apomixis is important not only for biological interests, but also for agricultural technology, because it may be possible to add this trait to crops3,4.
The green seaweed Ulva belongs to the class Ulvophyceae and primarily exists in marine habitats, but is also found in brackish water and even in freshwater5,6,7,8,9. In the lifecycle of Ulva, gametophytes of two mating types – minus (mt−) and plus (mt+) – release biflagellate gametes with positive phototaxis; mt− gametes then conjugate with mt+ gametes and vice versa; the zygote develops into the sporophytic phase10. Gametes that fail to conjugate develop parthenogenetically into gametophytes9. Sporophytes generate and release quadriflagellate meiospores through meiosis, which then develop into genetically separate mt− or mt+ gametophytes9 (Supplementary Fig. 1). Thus, each sexual individual has a lifecycle that consists of a gametophytic stage and a sporophytic stage9.
Instead of undergoing the ordinary sexual lifecycle, several Ulva species have obligately asexual lifecycles that occur without sexual reproduction via meiosis and conjugation. Sexual and asexual populations coexist in phylogenetically independent lineages, suggesting that each asexual population evolved separately in each lineage11,12,13. There are two types of asexual cycles, one via biflagellate zoids, and the other via quadriflagellate zoids (Supplementary Fig. 1)14. Both biflagellate and quadriflagellate zoids develop directly into asexual thalli. The asexual zoids are termed ‘zoosporoids’14. Like other Ulva species, Ulva prolifera has both types of asexual populations, and the amount of DNA in the cells of an asexual thallus in this species is similar to that in the cells the sporophytic thallus11. Although previous research has indicated that asexual thalli are diploid, it has not been examined whether each asexual thallus was generated by diploidization of the gametophyte or by apomixis in the sporophyte.
In the 2010s, macroalgal genomics became an active area of research, and the genomes of some brown and red marine alga have been sequenced15,16,17. The genome of the model brown alga Ectocarpus was released in 201015 and was recently improved by deep sequencing using next-generation sequencing technology18. Sex-determining regions (SDRs) of Ectocarpus were also identified19, and a candidate male reproductive gene, which is similar to a receptor involved in sperm-egg recognition in sea urchins, was found by RNA sequencing (RNA-Seq) analysis of mature male and female gametophytes20. Although genomic analysis in Ulvophyceae has lagged behind that of other taxa, the genome of Ulva mutabilis has recently been published21. Our recent analysis of the Ulva partita genome also revealed a mating-type (MT) locus22. The MT locus contains MT-specific genes and gametologs that are shared, but with differentiated sequences, between the two MTs. One of the gametologs, the PRA1 (proliferation-associated protein 1) gene, has been isolated from individual MT strains of other Ulva species, and the molecular phylogenetic tree of the Ulva species reveals that these isolated genes are clearly classified into two distinct clades associated with the MTs, which suggests that the MT locus is conserved in the genus Ulva and evolved independently within the MTs of each species. The availability of genomic data and the discovery of the MT locus now provide a molecular basis for studying the origin and evolution of sex, the sex-determination system, and the pathway by which asexual populations emerged in the Ulvophyceae.
In this study, we explored the evolutionary process by which asexual thalli (biflagellate type and quadriflagellate type) arose from the sexual population in the green seaweed Ulva. We used MT locus information and DNA content analysis to determine whether each asexual thallus was produced by diploidization of the gametophyte or by apomixis in the sporophyte. We also observed chromosomal dynamics during reproductive cell formation to investigate the nuclear division pattern in the cells. Finally, we performed RNA-Seq analysis to search for the causative genes for apomixis.
Results
Genotyping of culture strains using genes in the MT locus
Linkages between Ulva MT locus–specific marker genes and each MT of U. prolifera were examined in culture strains established from thalli collected in various regions in Japan and Denmark (Table 1). Two mt− MT locus–specific markers were detected in all mt− gametophytic thalli, and two mt+ MT locus–specific markers were detected in all mt+ gametophytic thalli. Each MT locus–specific marker was not detected in gametophytic thalli of the opposite MT. Thus, the MT locus–specific marker genes were completely linked to the MT of each gametophytic thallus (Fig. 1a). Conversely, both MT locus–specific markers were detected in each strain of asexual thallus (Fig. 1b), suggesting that the asexual thalli produced via a biflagellate zoosporoid and the asexual thalli produced via a quadriflagellate zoosporoid are carrying both the mt− and mt+ MT loci.
Genomic PCR of the MT-locus gene homologs for distinct culture strains. The presence of orthologous genes of the U. partita MT-locus genes (mt− genes: UpRWP1, UpPAR1m, mt+ genes: Up00832, UpPAR1f) was confirmed in 21 strains (10 gametophytic thallus strains and 11 asexual thallus strains) isolated from different areas along the Japanese coast and Denmark: (a) gametophytic thallus strains, (b) asexual thallus strains. Details of each strain are summarized in Table 1.
Comparing the DNA content of gametes, biflagellate zoosporoids, and quadriflagellate zoosporoids
The nuclear DNA contents of mt− gametes, mt+ gametes, biflagellate zoosporoids, and quadriflagellate zoosporoids were measured by fluorescence microscopy for the determination of the ploidy. After calibrating the fluorescence of nuclear DNA using the fluorescence of standard beads, the nuclear DNA content of each reproductive cell was measured. The relative nuclear DNA content of the mt− gametes was consistent with that of mt+ gametes, and each of the values were designated 1 C (Fig. 2a,b). Comparison of the DNA contents of gametes with those of the biflagellate or quadriflagellate zoosporoids yielded the estimate that the DNA content of the biflagellate and quadriflagellate zoosporoids were 2 C (Fig. 2c,d). These results indicate that the two types of zoosporoids are cytologically diploids.
Histogram comparing the relative nuclear DNA content among gametes and zoosporoids: (a) mt− gametes released by Up01 gametophytic thallus, (b) mt+ gametes released by Up02 gametophytic thallus, (c) biflagellate zoosporoids released by Up13 asexual thallus, (d) quadriflagellate zoosporoids released by Up17 asexual thallus.
Morphological observation of gametes, biflagellate zoosporoids, and quadriflagellate zoosporoids by field-emission scanning electron microscopy (FE-SEM)
The mt− and mt+ gametes, the biflagellate zoosporoids, and the quadriflagellate zoosporoids were observed by FE-SEM to determine their morphological differences. The mt− gametes had a mating structure (i.e., cell fusion apparatus) on the opposite side of the eyespot (Fig. 3a,b). In contrast, the mating structure of the mt+ gamete was located on the same side of the eyespot (Fig. 3c,d). The biflagellate zoosporoids had a larger eyespot than those of the gametes (Fig. 3e,f), and no mating structure was found on either the same or opposite side from the eyespot. The quadriflagellate zoosporoids without a mating structure had the largest cell size and had four cruciately arranged flagella (Fig. 3g,h).
Field emission scanning electron micrographs of gametes and zoosporoids of Ulva prolifera: (a,b) mt− gametes released by Up01 gametophytic thallus, (c,d) mt+ gametes released by Up02 gametophytic thallus, (e,f) biflagellate zoosporoids released by Up15 asexual thallus, (g,h) quadriflagellate zoosporoids released by Up17 asexual thallus. Arrow indicates the mating structure. E, eyespot. Scale bar, 1 μm.
Observation of chromosomal dynamics during the formation of reproductive cells
To understand the mechanism of gamete, meiospore, and diploid zoosporoid formation, the chromosomal dynamics were observed during each stage of reproductive cell formation. First, the cells of the mt− gametophytic thallus during mitotic prophase were observed to reveal the chromosome number of the gametophyte, and seven chromosomes were clearly observed (Fig. 4a). Fourteen chromosomes were observed during the mitotic prophase in cells of the sporophytic thallus; those of the asexual thallus produced via biflagellate zoosporoids and those of the asexual thallus produced via quadriflagellate zoosporoids (Fig. 4b–d). Additionally, 14 chromosomes were located on the equatorial plate in cells of the asexual thallus produced via quadriflagellate zoosporoids during mitotic mid-metaphase (Supplementary Fig. 2).
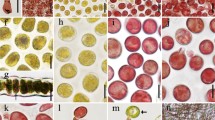
Chromosome dynamics during mitosis and each reproductive cell formation. (a–d) Chromosome dynamics in mitosis: (a) an mt− gametophytic thallus (Up01), (b) a sporophytic thallus (Up01 × Up02), (c) an asexual thallus produced via a biflagellate zoosporoid (Up13), (d) an asexual thallus produced via a quadriflagellate zoosporoid (Up17). Seven chromosomes were observed in a gametophytic thallus (a) and 14 chromosomes were observed in a sporophytic thallus and asexual thalli (b–d). (e–q) Chromosome dynamics during the formation of each reproductive cell. (e–g) Mitotic nuclear division for gamete formation in an mt− gametophytic thallus (Up01). (h–q) Meiotic or apomeiotic nuclear division for meiospores or zoosporoids formation. (h–j) A sporophytic thallus (Up01 × Up02). (k–m) An asexual thallus produced via a biflagellate zoosporoid (Up13); (n–p) an asexual thallus produced via a quadriflagellate zoosporoid (Up17). Seven bivalent chromosomes were observed during meta-phase of a sporophytic thallus and two types of an asexual thallus. Mt−gt, mt− gametophytic thallus; St, sporophytic thallus; Atbz, asexual thallus produced via a biflagellate zoosporoid; Atqz, asexual thallus produced via a quadriflagellate zoosporoid. Scale bar, 3 μm.
During the first nuclear division before gamete formation, seven chromosomes were found in cells of the mt− gametophytic thallus (Fig. 4e). In metaphase, the seven chromosomes were arranged at the equatorial plate (Fig. 4f), and then the homologous chromosomes were segregated toward opposite poles (Fig. 4g). In cells of the sporophytic thallus, the chromosomes were entangled in meiotic prophase I (Fig. 4h). The seven bivalent chromosomes were arranged at the equatorial plate in meiotic metaphase I (Fig. 4i), and the seven chromosomes were observed at both polar areas after the first meiotic division (Fig. 4j). The chromosome number reduced from 14 to 7 in this first meiotic phase. In the cells of each asexual thallus produced via biflagellate zoosporoids and quadriflagellate zoosporoids, entangled chromosomes were observed in prophase I (Fig. 4k,n). In metaphase I, the seven bivalent chromosomes aligned at the equatorial plate (Fig. 4l,o), and after the segregation of chromosomes toward opposite poles, the number of chromosomes was unreduced (Fig. 4m,p). The reduction in the chromosome number was not observed in the cells of the two types of asexual thalli in this first meiotic phase.
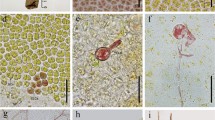
The apomeiotic process, which was similar to the phenomena that occurred in both of the asexual thalli, was also observed in the cells of the sporophytic thallus. Normal first meiotic division was observed in 42 cells, whereas apomeiotic division was observed in 14 cells (n = 56; Fig. 5a–c). In the latter case, the reduction in the chromosome number was not observed, and the 14 chromosomes existed in telophase I (Fig. 5b). To confirm that the apomeiotic process occurred in the sporophytic thallus, the genotypes of 66 thalli that developed from meiospores were examined using the above MT locus–specific markers. Genotyping revealed that 59 thalli were haploid gametophytic thalli (mt−: 19, mt+: 40). However, both of the MT markers were detected in seven thalli (which were defined as the mt± thallus strain). This result also provided evidence for the occurrence of apomeiosis during reproductive cell formation in sporophytic thalli. Additionally, the biflagellate zoids released from the mt± thalli (F1-3, F1-7, F1-14) had the round mating structure on the same side as the eyespot (Fig. 5d,e) and the amount of DNA in the biflagellate zoids was 2 C, which was the same as that of the biflagellate or quadriflagellate zoosporoids (Fig. 5f). The biflagellate zoids released from the F1-14 strain were crossed with mt− or mt+ gametes, resulting in conjugation between the biflagellate zoids and the mt− gametes, but not the mt+ gametes (Supplementary Fig. 3).
Apomeiosis in the cells of a sporophytic thallus (Up01 × Up02) and observation of diploid biflagellate zoids released from the mt± thallus. (a) Meiotic telophase I in normal meiosis. (b) Meiotic telophase I in apomeiosis; chromosome numbers were not reduced. (c) Cells number of progressing meiosis or apomeiosis in a sporophytic thallus. n = 56. (d,e) Morphology of biflagellate zoids released from the mt± thallus. (d) Light micrograph. (e) Field-emission scanning electron micrograph. Arrow indicates the mating structure. E, eyespot. (f) Histogram of relative nuclear DNA content in diploid biflagellate zoids. Scale bar, 3 μm (a,b), 10 μm (d), 1 μm (e).
Expression of the MT-locus genes and meiosis-related genes in reproductive cells
To understand the molecular basis of the morphological, cytological, and genetic differences between the reproductive cells, RNA-Seq analysis was conducted (Supplementary Table 1); this focused on the MT-locus genes and the meiosis-related genes. Genes that were homologous to the MT-locus genes of U. partita were searched using BLASTX querying a database of U. prolifera contigs assembled by RNA-Seq, and their expression levels were estimated (Fig. 6). 60% of the mt− specific genes (14/23), 45% of the mt+ specific genes (20/44), and 91% of the gametologs (21/23) were found to be homologous to the contigs of U. prolifera (Supplementary Tables 2 and 3). Many gametologs were highly expressed in the gametes of the individual MTs (Fig. 6a,b). All sets of gametologs derived from both MTs were expressed in the biflagellate and quadriflagellate zoosporoids, and the expression patterns of gametologs were clearly different when comparing both gametes and zoosporoids (Fig. 6a,b; Supplementary Tables 4 and 5). The expression pattern of MT-specific genes did not show a distinct tendency when comparing the gametes and zoosporoids (Supplementary Tables 6 and 7). Differentially expressed genes (DEGs) were not detected among mt− gametologs or mt- specific genes, while there were some DEGs in mt+ gametologs (elF1, PRP1, SLG1, 01506 and MET1) and in mt+ specific genes (02393 f and 02227 f).
Changes in the expression of the mating type (MT) locus homologous genes and the meiosis-related genes in gametes and zoosporoids: (a) mt− gametologs, (b) mt+ gametologs, (c) mt−-specific genes, (d) mt+-specific genes, (e) meiosis-related genes. Numbers under the gene names in (e) indicate the meiotic process category: 1. entry into meiosis, 2. recombination, 3. sister chromatid cohesion, 4. cell cycle control: spindle–cytokinesis. The blue heatmap shows the fragments per kilobase of exon per million fragments (FPKM) values. High FPKM values (>200) are shown in the same color as values of 200. The red heat map shows z-score calculated from FPKM values. Differentially expressed genes are indicated by an asterisk on the right side of or above the heat maps (*p < 0.05). mt−, mt− gamete; mt+, mt+ gamete; bz, biflagellate zoosporoid; qz, quadriflagellate zoosporoid.
Thirty-six meiosis-related genes from land plants were found in the U. prolifera RNA-Seq contigs (Supplementary Table 8). The expression level of each homologous gene is shown as a heatmap in Fig. 6e. Clustering analysis revealed that the gene expression tendencies of the mt− and mt+ gametes were similar, and that those of the quadriflagellate zoosporoids and the mt+ gametes were markedly different (Fig. 6e; Supplementary Table 9). The expression of Mei2L (Category 1) and RAD1 (Category 2) genes differed significantly between each type of reproductive cell.
Discussion
This study revealed that the asexual thalli of Ulva prolifera are diploid and have the genes of both of the MT loci, regardless of the number of flagella on the zoosporoids. Although the diploidy of asexual thalli has previously been suggested11, the cytological, DNA content, chromosome number, and genotype results presented here strongly suggest that the asexual thalli contain both the mt− and mt+ genomes and originated from sporophytes.
The mating structure at the tip of each gamete was observed and these structures corresponded to previous findings23. The conclusion that the function of the mating structure in the Ulvophyceae is for fertilization was also supported by the localization of membrane fusion protein HAP2/GCS124. Conversely, in the biflagellate zoosporoids, no mating structure could be found on the eyespot side or the side opposite from the eyespot, even in the samples collected in the field or from isolated cultures. The biflagellate zoosporoids were completely lacking in the mating structure and the ability to conjugate, although this study did not elucidate the mechanism by which this occurred. The diploid biflagellate zoids with a mating structure on the same side as the eyespots (Fig. 5e) released from the diploid mt± thalli were obtained from a sporophytic thallus and conjugated with mt− gametes (Supplementary Fig. 3). These phenotypes indicated that mt+ is dominant over mt− in Ulva, which is contrary to the results for Chlamydomonas. In Chlamydomonas, heterozygous diploid cells displayed the characteristics of mt−, indicating that mt− is dominant over mt+ 25. A discrepancy between the Ulvophyceae and Chlamydomonas in the relationship between sex and cell fusion site has been also reported23,26,27. This incongruence between two green algal lineages may be the result of differences in the genetic pathways that determine MT.
In this study, we observed chromosome dynamics during the formation of reproductive cells in the green algae U. prolifera. Observation by fluorescent microscopy revealed that normal meiosis I occurred in a major portion of the cells in the sporophytes, but some of the sporophyte cells, and cells of the asexual thalli produced via biflagellate and quadriflagellate zoosporoids, carried out apomeiosis without a reduction in the number of chromosomes (Supplementary Fig. 4). Bivalent chromosomes were clearly observed during the formation of biflagellate and quadriflagellate zoosporoids, indicating that cells in asexual thalli begin the meiosis I process, but do not complete it. These chromosomal dynamics indicate that zoosporoid formation in asexual thalli does not occur via somatic cells like the mitotic process, but rather by the apomeiotic process. The observed pattern of apomeiosis, in which bivalent chromosomes are formed, but sister chromatids segregate in the first division, is similar to the nuclear division pattern observed in the Arabidopsis MiMe (osd1/Atspo11-1/Atrec8) mutant28. In the MiMe mutant, Atspo11-1 and Atrec8 mutations instigate a mitotic-like first meiotic division, while an osd1 mutation prevents the second meiotic division. In the U. prolifera asexual thallus, it is unclear that the second meiotic division is skipped (as it is in the MiMe mutant); understanding the whole meiotic or apomeiotic process that occurs in each reproductive cell would require flowcytometric analysis for measuring ploidy before cells begin the meiotic division process. Meiosis without a decrease in the number of the chromosomes in the first meiotic division has been also reported in land plants having holocentromeric chromosomes29,30. There are no reports of holocentromeres in the Ulvophyceae, and thus a more detailed analysis of centromere structure in this group would also help to elucidate the meiosis process in U. prolifera.
Apomictic plants can be produced through outcrossing between genetically distant strains in the rhodophyta Caloglossa31. In U. prolifera, some asexual thalli were heterogeneous in an exon region of Hsp90, a single-copy gene, and a similar process may contribute to the generation of asexual thalli in Ulva13. In this study, the Up01 × Up02 sporophyte was found to produce asexual thalli through the apomeiotic process (Fig. 5; Supplementary Fig. 5). However, the Up01 and Up02 strains were isolated from meiospores released by a single sporophyte (E21)6,32. This result suggests that apomeiosis may occur readily in the sporophytic thallus of U. prolifera regardless of the genetic distance between male and female gametes. In U. mutabilis, sporophytic thalli rarely released diploid quadriflagellate zoids (0.5%), which grow into mt−/mt−, mt+/mt+, or mt+/mt− thalli10. In comparison, our results revealed a high production rate of mt± (mt+/mt−) thalli. This difference may be due to species characteristics or experimental conditions. The proportion of normal embryos and embryos produced by apomixis varies in land plants depending on environmental stressors33,34. In the alpine plant Ranunculus kuepferi, which is a facultative apomict, cold stress increases the frequency of apomictic seed formation35. These results indicate that environmental stresses affect the determination of the reproductive mode in plants. The gametophytes and sporophytes used in the present study were originally collected from a river estuarine area with low salinity6. Our experiment was carried out under seawater conditions, which might have induced a high rate of apomeiosis in the sporophytes due to salinity stress (Fig. 5).
Two mt+-specific genes were slightly up-regulated in mt+ gametes and quadriflagellate zoosporoids, but these genes show no homology to known proteins. Some gametologs (e.g., mt+ MAPKKK1 and mt− HAR1) and MT-specific genes (GTB1m1 and 00368f1) were expressed in one MT gamete but not in the opposite one (Fig. 6a,b). This finding suggests either that these genes have already been translocated from the MT locus in U. prolifera to an autosomal region or that the U. partita homologous gene was translocated from the autosome after the two species diverged from each other. The observation of chromosome dynamics during reproductive cell formation indicated that biflagellate and quadriflagellate zoosporoids were produced through the meiotic process rather than the mitotic process. The Mei2-Like gene homolog and RAD1 homolog were especially highly expressed in quadriflagellate zoosporoids. Expression and mutant analysis of the AML (Arabidopsis-mei2-like) gene in Arabidopsis thaliana suggests that AML is involved in meiosis as well as vegetative growth, indicating that Mei2 homologs are meiosis-related genes that are widely conserved from yeasts to land plants36. High levels of expression of the Mei2-Like (Mei2L) homolog in quadriflagellate zoosporoids suggests that the Mei2L homolog has a function during meiosis in Ulva (Fig. 6e).
Regardless of the number of flagella, each asexual thallus has both mt− and mt+ genomes, thus, each asexual thallus was originally produced through apomixis via apomeiosis in a sporophytic thallus (Supplementary Fig. 4). Transcriptome analysis revealed that the MT-locus genes were expressed in the biflagellate and quadriflagellate zoosporoids, and the Mei2L homolog was upregulated in the quadriflagellate zoosporoids. In the next step, transcriptomic analysis will be conducted during the meiotic stage to find genes related to the apomeiotic pathway. We also found that apomeiosis in sporophytic thalli produce mt± thalli, which release diploid biflagellate zoids. Although there is a morphological and behavioral difference between the diploid biflagellate zoids and the biflagellate zoosporoids released from the asexual thalli (e.g., the existence of the mating structure and conjugation ability), asexual populations of U. prolifera most likely originated from this apomeiotic pathway in sporophytic thalli. Moreover, the distribution of the U. prolifera population in the Shimanto River37 suggests that the sexual and asexual populations have different environmental tolerances of stress factors, such as salinity, nutrient levels, and temperature. In addition to molecular biological analysis for uncovering the genetic basis of apomixis, ecological research will help clarify how sexual and asexual reproduction in U. prolifera is involved in environmental adaptation.
Methods
Culture conditions
Table 1 summarizes strain information. Although the U. prolifera MT has previously been described as male or female based on the slight tendency toward anisogamy11, we used the term ‘mt−’ for male and ‘mt+’ for female, consistent with the usage in a previous study in U. partita22,27. Unialgal cultures were established from meiospores, gametes, or zoosporoids using the induction method38. Released meiospores, gametes, or zoosporoids were cultured in petri dishes at 20 °C with a 14:10-h light-dark (LD) cycle under fluorescent light at 100 µmol m−2 s−1 in artificial seawater (ASW; MARINE ART SF-1, Tomita Pharmaceutical Co., Ltd., Tokushima, Japan) supplemented with PES medium stock solution39. Upon reaching a size of 1 cm, some thalli were placed in 1-L aerated flasks and cultured for 2–3 weeks.
Genotyping of field materials and F1 progeny of U. prolifera using MT-locus genes
To isolate orthologs of the MT-locus genes of U. partita in U. prolifera22, all the MT-locus genes of U. partita were queried against contigs derived from assembled RNA-Seq data of the mt− and mt+ gametes and the asexual biflagellate and quadriflagellate zoosporoids of U. prolifera using the TBLASTX program. UpRWP1 (mt−) and Up0832 (mt+)22 were used as MT markers among the MT-locus gene orthologs found in the contigs of U. prolifera. A gametolog, UpPRA22, which was used for molecular phylogenetic analysis in Ulva to indicate the conserved differentiation of the MT locus in the genus, has also been used as an MT marker. To identify the genotype of individual strains using the MT-locus genes, DNA was extracted from each culture strain and from thalli developed from meiospores released by a single sporophyte (Up01 × Up02) using the DNeasy Plant Mini Kit (Qiagen, Valencia, CA, USA) or CicaGeneus DNA Extraction Regents ST (Kanto Chemical Co., Inc., Tokyo, Japan). The primer sequences are listed in Supplementary Table 10. To amplify each of the marker fragments, the Kapa Taq PCR Kit (Kapa Biosystems, Woburn, MA, USA) was used following the manufacturer’s protocol. The PCR program for each gene consisted of an initial denaturation step of 3 min at 95 °C, followed by 45 cycles of denaturation for 30 s at 94 °C, annealing for 30 s at 58 °C, and extension for 90 s at 68 °C. The obtained DNA fragments were separated by electrophoresis and visualized by staining with GelRed (Biotium, Fremont, CA, USA).
Measurement of the relative amounts of DNA
To compare the relative amounts of DNA, the DNA content of each gamete and asexual zoosporoid was measured. Gamete and zoosporoid formation were induced by the punching method38. Gamete and asexual zoosporoids were collected using positive or negative phototaxis and fixed with 1% glutaraldehyde (GA) in ASW at 4 °C for 2 hr. After fixation, the cells were washed with PBS three times. Each cell was stained with 0.5 μg/L DAPI diluted with PBS for 10 min at room temperature. After washing with PBS, the cells were observed with a fluorescence microscope (DM6000B; Leica Microsystems, Wetzlar, Germany). The fluorescence intensity was measured using ImageJ software and calibrated using InSpeck™ Blue (350/440) Microscope Image Intensity Calibration Kit (Thermo Fisher, Waltham, MA, USA).
Observation of the ultrastructure of gametes and zoosporoids by FE-SEM
Meiospores, gametes, and zoosporoids were prepared using the induction procedure38. Individual gametes, zoosporoids, and mt± biflagellate zoids were fixed using a previously described methodology23. Crossings between gametes and biflagellate zoids released from the F1-14 strain were performed at room temperature. The cells were fixed23 2 min after mixing. Post-fixation was for 2 h at 4 °C in 1% OsO4 dissolved in 0.05 M phosphate buffer (pH 7.4). After post-fixation, samples were treated in 1% tannic acid dissolved in 0.05 M phosphate buffer (pH 7.4) for 15 min at room temperature, and then fixed for 2 h at 4 °C in 1% OsO4 dissolved in 0.05 M phosphate buffer (pH 7.4). The samples were dehydrated through a graded series of ethanol solutions, infiltrated with t-butyl alcohol and freeze-dried (ES-2030; Hitachi, Tokyo, Japan). Samples were coated with platinum–palladium using ion sputter (E1045; Hitachi Corp., Tokyo, Japan). Observations were made under a FE-SEM (JSM6330F; JEOL Ltd., Tokyo, Japan) at 5 kV.
Chromosome analysis
Thalli containing the cells in meiotic phase were collected at 30–32 h after the induction of gametogenesis or sporogenesis. Mitosis of Ulva occurs during the dark period40. Therefore, each thallus was collected during the dark period of the LD cycles. Thalli were fixed in 1% GA in ASW at 4 °C for 2 h, and then washed with PBS three times. Thalli were treated with an enzyme mixture41 at 30 °C for 3 h to digest their cell walls. Thalli were washed with PBS three times and stained with SYBR Green I. Chromosomes were observed using a fluorescence microscope (DM6000B; Leica Microsystems). Florescence images of SYBR Green I were captured by a CCD camera (DFC360 FX; Leica Microsystems) as 0.5-μm-thick stacks in the z-axis and each image was deconvoluted by LAS AF ver. 2.6.0 software (Leica Microsystems).
RNA-Seq analyses
We conducted transcriptome analysis to clarify the molecular basis of the morphological and biological differences between the mt− and mt+ gametes and asexual biflagellate and quadriflagellate zoosporoids. Total RNA was extracted from mt− and mt+ gametes and asexual biflagellate and quadriflagellate zoosporoids using the RNeasy Plant Mini Kit (Qiagen, Valencia, CA, USA). Contaminating DNA was removed using RNase-Free DNase I (Qiagen). cDNA library construction and sequencing with the Illumina Hiseq. 2500 platform (Illumina, San Diego, CA, USA) were carried out by BGI (Shenzhen, China). The short-reads data were merged and then assembled using Trinity software42. The contigs obtained were clustered using Cd-hit software43; homologous genes located in the MT locus of U. partita22 and related to meiosis in eukaryotes44,45 were identified by BLAST X from the database for U. prolifera clustered contigs. Each of the short reads was mapped onto each of the clustered contigs using Bowtie software46, and the FPKM value of each contig was calculated by eXpress47 software on the Maser (Management and Analysis System for Enormous Reads) platform (https://cell-innovation.nig.ac.jp). Calculation of z-scores and clustering analysis based on the FPKM values for each gene were carried out using the gplots (2.16.0) package with R software (3.5.1; http://www.R-project.org/). Statistical testing for gene expression was performed in R with DESeq2 (3.8) using the no replicate method48. A Wilcoxon signed-rank test was conducted using Python 3.6.1/SciPy 0.19.1 software to assess the statistical significance between the gene expression patterns of the mt− and mt+ gametes and asexual biflagellate and quadriflagellate zoosporoids.
References
Koltunow, A. M. & Koltunow, A. M. Apomixis: Embryo sacs and embryos formed without meiosis or fertilization in ovules. Plant Cell 5, 1425–1437, https://doi.org/10.1105/tpc.5.10.1425 (1993).
Koltunow, A. M. & Grossniklaus, U. Apomixis: A developmental perspective. Annual Review of Plant Biology 54, 547–574, https://doi.org/10.1146/annurev.arplant.54.110901.160842 (2003).
Koltunow, A. M., Bicknell, R. A. & Chaudhury, A. M. Apomixis: Molecular strategies for the generation of genetically identical seeds without fertilization. Plant Physiology 108, 1345–1352, https://doi.org/10.1104/pp.108.4.1345 (1995).
Spillane, C., Curtis, M. D. & Grossniklaus, U. Apomixis technology development: Virgin births in farmers’ fields? Nature Biotechnolgy 22, 687–691, https://doi.org/10.1038/nbt976 (2004).
Reed, R. H. & Russell, G. Adaptation to salinity stress in populations of Enteromorpha intestinalis (L.) Link. Estuarine and Coastal Marine Science. 8, 251–258, https://doi.org/10.1016/0302-3524(79)90095-1 (1979).
Shimada, S., Yokoyama, N., Arai, S. & Hiraoka, M. Phylogeography of the genus Ulva (Ulvophyceae, Chlorophyta), with special reference to the Japanese freshwater and brackish taxa. Journal of Applied Phycology 20, 979–989, https://doi.org/10.1007/s10811-007-9296-y (2008).
Ichihara, K. et al. New species of freshwater Ulva, Ulva limnetica (Ulvales, Ulvophyceae) from the Ryukyu Islands, Japan. Phycological Research 57, 94–103, https://doi.org/10.1111/j.1440-1835.2009.00525.x (2009).
Rybak, A. S. Ecological preferences of freshwater Ulva flexuosa (Ulvales; Ulvophyceae): Development of macroalgal mats in a Tulce fishpond (Wielkopolska Region, Poland). Oceanological and Hydrobiological. Studies 45, 100–111, https://doi.org/10.1515/ohs-2016-0010 (2016).
van den Hoek C., Mann D. G. & Jahns, H. M. Algae. An Introduction to Phycology, 391–408, (Cambridge University Press, 1995).
Fjeld, A. & Lovlie, A. “Genetics of multicellular marine algae,” in The Genetics of Algae. (ed. Lewin, R. A.), 219–235, (University California Press, 1976).
Hiraoka, M. et al. Different life histories of Enteromorpha prolifera (Ulvales, Chlorophyta) from four rivers on Shikoku Island, Japan. Phycologia 42, 275–284, https://doi.org/10.2216/i0031-8884-42-3-275.1 (2003).
Hiraoka, M., Shimada, S., Ohno, M. & Serisawa, Y. Asexual life history by quadriflagellate swarmers of Ulva spinulosa (Ulvales, Ulvophyceae). Phycological Research 51, 29–34, https://doi.org/10.1046/j.1440-1835.2003.00290.x (2003).
Ogawa, T., Ohki, K. & Kamiya, M. High heterozygosity and phenotypic variation of zoids in apomictic Ulva prolifera (Ulvophyceae) from brackish environments. Aquatic Bototany 120, 185–192, https://doi.org/10.1016/j.aquabot.2014.05.015 (2015).
Bliding, C. A critical survey of European taxa in Ulvales. Part I: Capsosiphon, Percursaria, Blidingia, Enteromorpha. Opera Botanica 8, 45–148 (1963).
Cock, J. M. et al. The Ectocarpus genome and the independent evolution of multicellularity in brown algae. Nature 465, 617–621, https://doi.org/10.1038/nature09016 (2010).
Collen, J. et al. Genome structure and metabolic features in the red seaweed Chondrus crispus shed light on evolution of the Archaeplastida. Proceedings of the National Academy of Sciences of the United States of America 110, 5247–5252, https://doi.org/10.1073/pnas.1221259110 (2013).
Ye, N. et al. Saccharina genomes provide novel insight into kelp biology. Nature Communications 6, 6986, https://doi.org/10.1038/ncomms7986 (2015).
Cormier, A. et al. Re-annotation, improved large-scale assembly and establishment of a catalogue of noncoding loci for the genome of the model brown alga Ectocarpus. New Phytologist 214, 219–232, https://doi.org/10.1111/nph.14321 (2016).
Ahmed, S. et al. A haploid system of sex determination in the brown alga Ectocarpus sp. Current Biology 24, 1945–1957, https://doi.org/10.1016/j.cub.2014.07.042 (2014).
Lipinska, A. P., Van Damme, E. J. M. & De Clerck, O. Molecular evolution of candidate male reproductive genes in the brown algal model. Ectocarpus. BMC Evolutionary Biology 16, 5, https://doi.org/10.1186/s12862-015-0577-9 (2016).
De Clerck, O. et al. Insights into the evolution of multicellularity from the sea lettuce genome. Current Biology 28, 2921–2933. e5, https://doi.org/10.1016/j.cub.2018.08.015 (2018).
Yamazaki, T. et al. Genomic structure and evolution of the mating type locus in the green seaweed Ulva partita. Scientific Reports 7, 11679, https://doi.org/10.1038/s41598-017-11677-0 (2017).
Mogi, Y. et al. Asymmetry of eyespot and mating structure positions in Ulva compressa (Ulvales, Chlorophyta) revealed by a new field emission scanning electron microscopy method. Journal of Phycology 44, 1290–1299, https://doi.org/10.1111/j.1529-8817.2008.00573.x (2008).
Yamazaki, T. et al. HAP2/GCS1 is involved in the sexual reproduction system of the marine macroalga Ulva compressa (Ulvales, Chlorophyta). Cytologia (Tokyo). 79, 575–584, https://doi.org/10.1508/cytologia.79.575 (2014).
Ebersold, W. T. Chlamydomonas reinhardi: Heterozygous diploid strains. Science 157, 447–449, https://doi.org/10.1126/science.157.3787.447 (1967).
Miyamura, S. & Nagumo, T. Inheritance pattern of chloroplast DNA is correlated with gamete types based on sex-specific arrangement of the cell fusion site in Caulerpa (Ulvophyceae, Chlorophyta). Phycological Research 55, 47–57, https://doi.org/10.1111/j.1440-1835.2006.00444.x (2007).
Kagami, Y. et al. Sexuality and uniparental inheritance of chloroplast DNA in the isogamous green alga Ulva compressa (Ulvophyceae). Journal of Phycology 44, 691–702, https://doi.org/10.1111/j.1529-8817.2008.00527.x (2008).
d’Erfurth, I. et al. Turning meiosis into mitosis. PLoS Biology 7, e1000124, https://doi.org/10.1371/journal.pbio.1000124 (2009).
Heckmann, S. et al. Alternative meiotic chromatid segregation in the holocentric plant Luzula elegans. Nature. Communications 5, 4979, https://doi.org/10.1038/ncomms5979 (2014).
Cabral, G., Marques, A., Schubert, V., Pedrosa-Harand, A. & Schlögelhofer, P. Chiasmatic and achiasmatic inverted meiosis of plants with holocentric chromosomes. Nature Communications 5, 5070, https://doi.org/10.1038/ncomms6070 (2014).
Kamiya, M., West, J. A. & Hara, Y. Induction of apomixis by outcrossing between genetically divergent entities of Caloglossa leprieurii (ceramiales, rhodophyta) and evidence of hybrid apomicts in nature. Journal of Phycology 47, 753–762, https://doi.org/10.1111/j.1529-8817.2011.01016.x (2011).
Hiraoka, M., Ichihara, K., Zhu, W., Ma, J. & Shimada, S. Culture and hybridization experiments on an ulva clade including the qingdao strain blooming in the yellow sea. PLoS One 6, e19371, https://doi.org/10.1371/journal.pone.0019371 (2011).
Barcaccia, G., Tavoletti, S., Falcinelli, M. & Veronesi, F. Environmental influences on the frequency and viability of meiotic and apomeiotic cells of a diploid mutant of alfalfa. Crop Science 37, 72–76, https://doi.org/10.2135/cropsci1997.0011183X003700010011x (1997).
Klatt, S. et al. Photoperiod extension enhances sexual megaspore formation and triggers metabolic reprogramming in facultative apomictic Ranunculus auricomus. Frontiers in Plant Science 7, 278, https://doi.org/10.3389/fpls.2016.00278 (2016).
Klatt, S., Schinkel, C. C. F., Kirchheimer, B., Dullinger, S. & Horandl, E. Effects of cold treatments on fitness and mode of reproduction in the diploid and polyploid alpine plant Ranunculus kuepferi (Ranunculaceae). Annals of Botany 121, 1287–1298, https://doi.org/10.1093/aob/mcy017 (2018).
Kaur, J., Sebastian, J. & Siddiqi, I. The Arabidopsis - mei2 - like genes play a role in meiosis and vegetative growth in Arabidopsis. Plant Cell 18, 545–559, https://doi.org/10.1105/tpc.105.039156 (2006).
Hiraoka, M. & Higa, M. Novel distribution pattern between coexisting sexual and obligate asexual variants of the true estuarine macroalga Ulva prolifera. Ecology and Evolution 6, 3658–3671, https://doi.org/10.1002/ece3.2149 (2016).
Hiraoka, M. & Enomoto, S. The induction of reproductive cell formation of Ulva pertusa Kjellman (Ulvales, Ulvophyceae). Phycological Research 46, 199–203, https://doi.org/10.1111/j.1440-1835.1998.tb00114.x (1998).
Provasoli, L. Media and prospects for the cultivation of marine algae. In Watanabe, A and Hattori, A (Eds). Culture and Collections of Algae. in Proceedings of the U.S.-Japan Conference, Hakone, September 1966, Japanese Society of Plant Physiologist 63–75 (1968).
Titlyanov, E., Titlyanova, T. & Luning, K. Diurnal and circadian periodicity of mitosis and growth in marine macroalgae. II. The green alga Ulva pseudocurvata. European Journal of Phycology 31, 181–188, https://doi.org/10.1080/09670269600651361 (1996).
Reddy, C. R. K. et al. An improved enzyme preparation for rapid mass production of protoplasts as seed stock for aquaculture of macrophytic marine green algae. Aquaculture 260, 290–297, https://doi.org/10.1016/j.aquaculture.2006.06.034 (2006).
Grabherr, M. G. et al. Full-length transcriptome assembly from RNA-Seq data without a reference genome. Nature Biotechnology 29, 644–52, https://doi.org/10.1038/nbt.1883 (2011).
Li, W. & Godzik, A. Cd-hit: A fast program for clustering and comparing large sets of protein or nucleotide sequences. Bioinformatics 22, 1658–1659, https://doi.org/10.1093/bioinformatics/btl158 (2006).
Mercier, R., Mézard, C., Jenczewski, E., Macaisne, N. & Grelon, M. The molecular biology of meiosis in plants. Annual Review of Plant Biology 66, 297–327, https://doi.org/10.1146/annurev-arplant-050213-035923 (2015).
Nasmyth, K. A meiotic mystery: How sister kinetochores avoid being pulled in opposite directions during the first division. BioEssays 37, 657–665, https://doi.org/10.1002/bies.201500006 (2015).
Langmead, B. Aligning short sequencing reads with Bowtie. Curr. Protoc. Bioinforma. 32, 11.7.1–11.7.14, https://doi.org/10.1002/0471250953.bi1107s32 (2010).
Roberts, A. & Pachter, L. Streaming fragment assignment for real-time analysis of sequencing experiments. Nature Methods 10, 71–73, https://doi.org/10.1038/nmeth.2251 (2013).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-Seq data with DESeq2. Genome Biology 15, 550, https://doi.org/10.1186/s13059-014-0550-8 (2014).
Cui, J. et al. Taxonomic reassessment of Ulva prolifera (Ulvophyceae, Chlorophyta) based on specimens from the type locality and Yellow Sea green tides. Phycologia 57, 692–704, https://doi.org/10.2216/17-139.1 (2018).
Acknowledgements
We thank Dr. Shuhei Ota for the advice about microscopic observation. This study was funded by a Grant-in-Aid for JSPS Research Fellow (No. 14J08330) to K.I. and JSPS KAKENHI (no. 25291070) to S.K. This research was partially supported by the Platform Project for Supporting Drug Discovery and Life Science Research (Basis for Supporting Innovative Drug Discovery and Life Science Research [BINDS]) from AMED under Grant Number JP17am0101001.
Author information
Authors and Affiliations
Contributions
K.I. and S.K. conceived and designed the research, and S.K. supervised the study. M.H. and K.I. maintained and cultured the strains and M.H. conducted crossing analysis and revealed the mating type of each culture strain. S.M. observed reproductive cells by FE-SEM and K.I. carried out other microscopic analysis. K.I. extracted genomic DNA and RNA samples. K.I. performed the genotyping analysis. T.Y. assembled RNA-Seq data and clustered contigs. K.I. and T.Y. analyzed the transcriptome. K.I. prepared the figures and tables. K.I. wrote the manuscript, and T.Y., S.M., M.H., and S.K. assisted in writing the manuscript.
Corresponding authors
Ethics declarations
Competing Interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Ichihara, K., Yamazaki, T., Miyamura, S. et al. Asexual thalli originated from sporophytic thalli via apomeiosis in the green seaweed Ulva. Sci Rep 9, 13523 (2019). https://doi.org/10.1038/s41598-019-50070-x
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-019-50070-x
- Springer Nature Limited
This article is cited by
-
Genome-wide SNP markers provided insights into the reproductive strategy and genetic diversity of the green tide causative species Ulva prolifera in China
Journal of Oceanology and Limnology (2024)
-
Gamete dimorphism of the isogamous green alga (Chlamydomonas reinhardtii), is regulated by the mating type-determining gene, MID
Communications Biology (2022)