Abstract
DNA replication is initiated at replication origins on chromosomes at their scheduled time during S phase of the cell cycle. Replication timing control is highly conserved among eukaryotes but the underlying mechanisms are not fully understood. Recent studies have revealed that some telomere-binding proteins regulate replication timing at late-replicating origins throughout the genome. To investigate the molecular basis of this process, we analyzed the effects of excessive elongation of telomere DNA on replication timing by deleting telomere-associated shelterin proteins in Schizosaccharomyces pombe. We found that rap1∆ and poz1∆ cells showed abnormally accelerated replication at internal late origins but not at subtelomere regions. These defects were suppressed by removal of telomere DNA and by deletion of the telomere-binding protein Taz1. Furthermore, Sds21—a counter protein phosphatase against Dbf4-dependent kinase (DDK)—accumulated at elongated telomeres in a Taz1-dependent manner but was depleted at internal late origins, indicating that highly elongated telomeres sequester Sds21 at telomeres and perturb replication timing at internal regions. These results demonstrate that telomere DNA length is an important determinant of replication timing at internal regions of chromosomes in eukaryotes.
Similar content being viewed by others
Introduction
DNA replication is one of the most fundamental biological processes for preserving genomic information and is initiated from multiple replication origins on chromosomes during S phase of the cell cycle. However, these events do not occur simultaneously; instead, each origin has specific replication timing that is regulated by multiple factors such as chromosomal position, location in the nucleus, and differentiation state1,2,3.
Recent studies in the fission yeast Schizosaccharomyces pombe have revealed the mechanisms regulating replication timing at late-replicating origins (late origins). Rif1 and Taz1, which mainly associate with telomeres at chromosome ends4,5 (Fig. 1A), maintain replication timing at late origins not only at telomere-adjacent subtelomeres but also at internal sites6,7. Rif1, the primary replication timing regulator, associates with telomeres via Taz1, which directly interacts with telomere DNA4,5. Interestingly, Rif1 localization near late origins is both Taz1-dependent and -independent; in the former instance, Taz1 binds to internal telomere repeat DNA and recruits Rif1 whereas in the latter, Rif1 interacts with chromatin through its affinity with G-quadruplex DNA structures7,8,9. Rif1 also directly binds protein phosphatase 1 (PP1) that counteracts the phosphorylation events by Dbf4-dependent kinase (DDK), which is essential for initiation of DNA replication10, to preserve the pre-replication complex (pre-RC) in an inactive state until late S phase5,11. Furthermore, Taz1-dependent late origins are regulated spatiotemporally during the cell cycle; they are tethered around telomeres via mutual interaction of telomere protein complexes especially during G1/S phase, and their replication timings are maintained at late S phase by PP1 that is recruited to telomeres by Rif13.
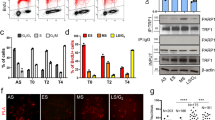
Rap1 deletion accelerates DNA replication from internal late origins. (A) Schematic illustration of telomere-binding proteins in S. pombe. (B) Replication timing of late origins is affected by Rap1 deletion. The locations of replication origins on chromosome 2 are schematically illustrated at the top. AT2024 and ars2004 are early origins (red squares), and AT2035, AT2080, AT2088, ars727, tel-60.0, and tel-0.2 are late origins (blue squares). The subtelomeric late origins tel-60.0 and tel-0.2 are located at 60.0 and 0.2 kb, respectively, from the telomere of the right arm of chromosome 2. The non-origin region (non-ori) is located at 30 kb from ars2004. Wild type, taz1∆, rap1∆, and rif1∆ cells were grown in EMM medium and released from G2/M arrest, and their DNA was labeled with BrdU for 90 min in the presence of 10 mM HU, purified by immunoprecipitation, and analyzed by qPCR. Each value was normalized to that of ars2004. Error bars indicate the s.d. (n = 3). Asterisks indicate a significant change as compared with the wild-type strain (p < 0.05, Student’s t-test).
Telomeres are specialized chromatin structures at the ends of eukaryotic linear chromosomes. Telomere protein complexes maintain genome integrity by protecting chromosome ends, regulating telomere DNA length, and tethering telomeres near the nuclear envelope4,5,12,13,14,15,16. Among S. pombe telomere-binding proteins, Rap1 plays a central role in the formation of telomere protein complexes such as shelterin, which bridges double- and single-stranded telomere DNAs at chromosome ends13 (Fig. 1A). Although recent studies have shown that Taz1 and Rif1 are important for replication timing at late origins, the precise roles of shelterin components and telomere DNA in replication timing control remain obscure.
To address this question, in this study we analyzed the effects of deleting the shelterin components Rap1 and Poz1 on replication timing. We demonstrate that excessively long telomere DNA perturbs replication timing at internal late origins but not near telomeres. Sds21, an S. pombe PP1, accumulated at these elongated telomeres in rap1∆ and poz1∆ cells in a Taz1-/Rif1-dependent manner, resulting in its disappearance from some of the internal late origins. Thus, telomere DNA length is an important determinant of replication timing at internal chromosomal regions.
Results
Rap1 deletion perturbs replication timing at internal late origins
To investigate the role of telomere-binding proteins in replication timing control, we evaluated the effect of deleting Rap1—a shelterin component (Fig. 1A) that contributes to various telomere functions in S. pombe5,13,17,18—on replication timing. We first analyzed the effect of Rap1 deletion on replication during early S phase by treating cells with hydroxyurea (HU), which depletes deoxynucleoside triphosphate pools and restricts further replication after it has been initiated at early-replicating origins (early origins) (Supplemental Fig. S1). In wild-type cells, late origins of chromosome 2 (AT2035, AT2080, AT2088, ars727, tel-60.0, and tel-0.2) did not replicate efficiently compared to early origins of the same chromosome (ars2004 and AT2024) (Fig. 1B). However, internal late origins (AT2035, AT2080, AT2088, and ars727) in rap1∆ cells replicated with greater efficiency than those in wild-type cells, whereas replication at subtelomeric late origins (tel-60.0 and tel-0.2) was largely unaffected (Fig. 1B). In contrast, deletion of Rif1—which is recruited to telomeres via association with the double-stranded telomere DNA-binding protein Taz14,5—increased replication levels at all of the late origins examined while deletion of Taz1 had the similar effect at a subset of internal late origins (AT2035 and AT2088) as well as at subtelomeric late origins (tel-60.0 and tel-0.2) (Fig. 1B). These data demonstrate that Rap1 is required for the maintenance of replication timing at some internal late origins but not at subtelomeres, and that effects of Rap1 deletion differ from those of Taz1 or Rif1 deletion. Importantly, effects of Rap1 deletion were observed even at Taz1-independent internal late origins—i.e., AT2080 and ars727, suggesting that replication timing defects at these sites in rap1∆ were not directly caused by disassembly of the shelterin complex.
Excessive elongation of telomere DNA accelerates replication at internal late origins
As described above, Rap1 is involved in the replication timing control of internal but not subtelomeric late origins; this is in stark contrast to the other telomere-binding proteins Taz1 and Rif1, the deletion of which causes defects in replication timing control at subtelomeres. Rap1 associates with Taz1 and regulates telomere length as well as various telomere functions; thus, its deletion results in abnormal elongation of telomere DNA for more than 10-fold as compared with the wild-type5,17 (Supplemental Fig. S2A). To explore whether absence of Rap1 per se or abnormally elongated telomere DNA in rap1∆ cells induced the observed defects in replication timing control, we analyzed the effects of Rap1 deletion on replication timing in the absence of telomere DNA. Deletion of the trt1+ gene encoding a catalytic subunit of telomerase causes gradual shortening of telomere DNA and is lethal in most cells, but some survive via self-circularization of chromosomes or recombination of chromosome end regions19,20. The rap1∆ trt1∆ strain with circular chromosomes and lacking telomere DNA showed no acceleration in replication timing compared to the control trt1∆ strain, whereas the rif1∆ trt1∆ strain showed accelerated replication at late but not early origins (Fig. 2 and Supplemental Fig. S2A,B). Thus, removal of telomere DNA from the rap1∆ strain suppressed the defects in replication timing. These data indicate that excessive elongation of telomere DNA and not Rap1 itself is important for replication timing control.
The rap1∆ strain shows replication timing defects in a telomere DNA-dependent manner. The rap1∆ trt1∆ strain showed replication timing similar to that observed in the trt1∆ strain at internal late origins in the absence of telomere DNA, whereas the rif1∆ trt1∆ strain showed accelerated replication at late origins. Cells with circular chromosomes were grown in YES medium, and the BrdU incorporation assays were performed as in Fig. 1B. Each value was normalized to that of ars2004 and then to the trt1∆ value.
We next investigated whether excessive telomere elongation caused by deletion of telomere proteins other than Rap1 also results in defects in replication timing at internal late origins. Poz1 is a subunit of the shelterin complex and as in the case of Rap1, its deletion causes abnormal telomere elongation as Rap1 deletion13 (Supplemental Fig. S2A). Accordingly, poz1∆ cells showed replication timing defects similar to those in rap1∆ cells—i.e., accelerated replication at the internal (AT2035, AT2080, AT2088, and ars727) but not at subtelomeric (tel-60.0 and tel-0.2) late origins (Fig. 3). These data confirm that abnormally elongated telomere DNA accelerates replication specifically at internal late origins in the presence of Taz1 and Rif1.
Replication timing defects in rap1∆ and poz1∆ are Taz1-dependent. The poz1∆ strain exhibited replication timing defects similar to those observed in the rap1∆ strain. These defects were suppressed by Taz1 deletion. The BrdU incorporation assays were performed as in Fig. 1B. Asterisks indicate a significant change as compared with the wild-type strain (p < 0.05, Student’s t-test).
Replication timing defects in rap1∆ and poz1∆ are Taz1-dependent
Elongated telomere DNA has more spaces to accommodate Taz1, a protein that directly binds to telomere DNA. We therefore examined whether the replication timing defects in rap1∆ and poz1∆ cells were caused by accumulation of Taz1 at telomeres. Deletion of Taz1 in rap1∆ or poz1∆ cells suppressed the replication timing defects at Taz1-independent internal late origins (AT2080 and ars727) while accelerating replication at subtelomeric late origins (tel-60.0 and tel-0.2) compared to the wild type (Fig. 3). Thus, excess Taz1 bound to abnormally long telomere DNA negatively and positively regulates late replication timing at internal and subtelomeric late origins, respectively.
Sequestering of Sds21 protein phosphatase 1 at telomeres causes accelerated replication at internal late origins
The previous data indicate that the accumulation of Taz1 at telomeres affects replication timing at internal late origins such as AT2080 and ars727; a normal timing was maintained in taz1∆ cells (Figs 1B and 3), suggesting that Taz1 accumulation at telomeres indirectly influences replication timing at these sites. In S. pombe, Rif1—which binds to telomeres by interacting with Taz1 (Fig. 1A)—recruits PP1 to late origins to counteract the phosphorylation events by DDK5,11. Of the two S. pombe PP1 proteins Sds21 and Dis2, the former primarily binds to telomeres and the internal late origin ars727 in a Rif1-dependent manner11.
We examined whether the localization of Rif1 or Sds21 at late origins is altered by excessive elongation of telomere DNA caused by deletion of telomere-binding proteins (Supplemental Fig. S2A). Chromatin immunoprecipitation (ChIP) of Rif1-12myc showed that Rif1 enrichment at ars727 was unaltered in taz1∆ and rap1∆ cells but was moderately increased and markedly reduced near telomeres (i.e., at tel-0.2) in the rap1∆ and taz1∆ strains, respectively (Fig. 4A). These data suggest that replication timing defects at ars727 in the rap1∆ mutant is not due to reduced association of Rif1, which associates with telomeres via Taz1 as previously reported5.
Sds21 is sequestered to excessively elongated telomeres in a Taz1-dependent manner. (A) Localization of Rif1 at ars727 was unaffected by Rap1 or Poz1 deletion. ChIP analyses of Rif1-12Myc localization were performed with cells at 60 min after released from G2/M arrest. Co-purified genomic DNA was analyzed by qPCR. Each value was normalized to that of ars2004. Error bars indicate the s.d. (n = 3). Asterisks indicate a significant change as compared with the wild-type strain (p < 0.05, Student’s t-test). (B) Sds21 localization at ars727 was decreased by Rap1 deletion. ChIP analyses of Sds21-3Flag were performed with cells at 60 min after released from G2/M arrest. Each value was normalized to that of ars2004. Error bars indicate the s.d. (n = 3). Asterisks indicate a significant change as compared with the wild-type strain (p < 0.05, Student’s t-test). (C) Sds21 accumulates at telomeres in rap1∆ cells. DNA co-precipitated with Sds21-3Flag was analyzed by Southern blotting using telomere and rDNA (control) probes. Left panel, representative images cropped from the same Southern blots (the original blots are displayed in Supplemental Fig. S3). Each signal compared with its background was analyzed by ImageJ software (NIH, Bethesda, MD, USA) and was normalized to that of rDNA and then to the wild-type value (right). Error bars indicate the s.d. (n = 3). Asterisks indicate a significant change as compared with the wild-type strain (p < 0.05, Student’s t-test). (D) Localization of Sds21 at ars727 in rap1∆ cells was restored by Taz1 deletion. ChIP was performed as in (B). Asterisks indicate a significant change between the indicated strains (p < 0.05, Student’s t-test).
We next examined the localization of Sds21-3Flag in the telomere-elongated mutants. In contrast to Rif1, Sds21 enrichment at ars727 was decreased in rap1∆ cells as in rif1∆ cells, whereas no decrease was detected near telomeres (at tel-0.2) (Fig. 4B). Furthermore, ChIP–Southern blot analyses showed that Sds21 was enriched at telomeres in rap1∆ cells relative to the wild type (Fig. 4C and Supplemental Fig. S3). This accumulation was abrogated by Taz1 deletion (rap1∆ taz1∆), thereby restoring Sds21 localization at ars727 (Fig. 4C,D).
The above results suggest that the number of Sds21 molecules that can regulate replication timing is limited. To determine whether additional Sds21 proteins can rescue the enrichment of Sds21 at ars727, Sds21 was overexpressed (Sds21-OE) in wild-type and rap1∆ cells. This caused severe defects in cell cycle progression as previously reported21 in both wild-type and rap1∆ cells and decreased Sds21 localization at ars727 (Fig. 5). One possible explanation is that excessive de-phosphorylation of target proteins by Sds21 blocks the interaction between Rif1 and Sds21. Alternatively, Sds21 may require post-translational modification for stable localization at some internal late origins, and an excess amount of unmodified Sds21 interferes with the stable localization of modified Sds21.
Overexpression of Sds21 causes severe defects in cell cycle progression. (A) Expression level of Sds21-3Flag protein in each strain. Wild type, Sds21-overexpressing wild type (wild type-Sds21OE), rap1∆, and Sds21-overexpressing rap1∆ (rap1∆-Sds21OE) cells were grown in EMM. The whole-cell extracts were analyzed by immunoblotting using anti-Flag (M2 F3165; Sigma-Aldrich) for Sds21-3Flag and anti-PSTAIRE (P7962; Sigma-Aldrich) for Cdc2 (loading control). (B) Representative septation index after release from G2/M block. (C) Wild type, wild type-Sds21OE, rap1∆, and rap1∆-Sds21OE cells were grown in EMM medium, and ChIP analyses of Sds21-3Flag localization were performed with cells in early S phase, i.e., at 60 min (for wild type and rap1∆), 130 min (for wild type-Sds21OE), and 80 min (for rap1∆-Sds21OE) after released from G2/M arrest. Co-purified genomic DNA was analyzed by qPCR. Each value was normalized to that of ars2004. Error bars indicate the s.d. (n = 3).
Discussion
In S. pombe, Rif1 is a primary regulator of replication timing at late origins6,7. Rif1 recruits PP1 phosphatases (Sds21 or Dis2) to counteract the activity of DDK to maintain replication timing of late origins11. Rif1-dependent late origins are categorized into two groups, Taz1-dependent and Taz1-independent origins7. At least some of the Taz1-dependent internal late origins are tethered to telomeres by association of shelterin complexes, and their replication timings are regulated by PP1 that is recruited to telomeres by the Taz1-Rif1 complex3. Our results suggest that Sds21 is sequestered to elongated telomeres via the Taz1–Rif1 complex in rap1∆ cells, preventing its localization at some Taz1-independent internal late origins such as ars727 and accelerating replication at these sites. The release of Sds21 from these long telomeres by Taz1 deletion restores Sds21 localization and normal replication timing at the Taz1-independent late origins (Fig. 6). Thus, we propose a new regulatory mechanism of replication timing at late origins; i.e., telomere DNA length is an important determinant of replication timing especially at Taz1-independent internal late origins.
Model of replication timing control based on telomere DNA length. (A) In wild-type cells, the PP1 Sds21 is recruited to telomeres and late origins such as ars727, a Taz1-independent origin, through association with Rif1. Localization of Sds21 at ars727 maintains replication timing at this site in late S phase. Rif1 associates with telomeres via Taz1. (B) In rap1∆ cells, telomere DNA is excessively elongated, resulting in the accumulation of Sds21 via Taz1–Rif1 complexes. Sequestration of Sds21 at telomeres reduces Sds21 localization at ars727 and accelerates replication timing in early S phase. (C) In taz1∆ rap1∆ cells, Sds21 is released from elongated telomeres, which restores Sds21 localization at ars727 and normal replication timing in late S phase. In contrast, the absence of Sds21 at telomeres accelerates replication timing at tel-0.2.
Rif1 and Taz1 are localized at proximal regions but not at exact sites of late origins6,7, suggesting that PP1 that is recruited by Rif1 or Taz1-Rif1 influences a relatively broad range of late origins on chromosomes. Taz1-dependent internal late origins are clustered to telomeres especially at G1/S phase3. Our data that replication at Taz1-independent late origins (ars727 and AT2035), but not at tel-0.2, was accelerated in rap1∆ cells suggest that ars727 and AT2035 are located far from telomeres in the nucleus. Hi-C or 3C analyses of interacting sites of chromosomes especially during G1/S phase will clarify spacio-temporal regulations of replication origins.
The results of this study demonstrate that telomere DNA length is a determinant of replication timing at internal late origins. In S. pombe, telomere DNA length is maintained by an active telomerase and the shelterin complex so the replication timing is basically unchanged over the lifespan of the organism. However, in higher eukaryotes telomere DNA becomes progressively shorter due to the decrease in telomerase activity after fertilization. Interestingly, replication timing at some chromosomal regions in mammals changes during differentiation in association with alterations in transcription levels and nuclear localization22. Thus, it is possible that telomere DNA-dependent replication timing control is dynamic over the course of differentiation, which is a possibility that merits further investigation.
Methods
Strains, media, and general techniques for S. pombe
S. pombe strains used in this study are listed in Supplemental Table S1. Experiments with S. pombe were performed as previously described23,24,25.
Replication timing assay
Chromosomal DNA replication was analyzed based on bromodeoxyuridine (BrdU) incorporation as previously described26,27, with some modifications. Derivatives of the cdc25-22 Pnmt1-TK Padh1-hENT strain expressing the herpes simplex virus thymidine kinase (TK) gene and the human equilibrative nucleoside transporter (ENT) gene were grown in Edinburgh minimal medium (EMM) at 25 °C to induce TK expression and incubated at 35.5 °C for 4 h for G2/M arrest. (Note that the trt1∆ strains were grown in YES medium to maintain their cell growth.) Cells were released at 25 °C to resume cell cycle progression in the presence of 200 µM BrdU (Sigma-Aldrich) with or without 10 mM HU (Sigma-Aldrich) and were fixed with 0.1% sodium azide at pre-determined times. Genomic DNA was prepared by vigorously vortexing of the cells in S&G buffer (10 mM Tris-HCl [pH 8.0], 1 mM EDTA, 100 mM NaCl, 1% SDS, 2% Triton X-100) in the presence of phenol/chloroform, followed by ethanol precipitation. The DNA was fragmented by sonication and heat-denatured, and then used for immunoprecipitation with anti-BrdU antibody (3D4; Becton Dickinson/Pharmingen) bound to Dynabeads M-280 anti-mouse IgG (Thermo Fisher). The DNA was recovered by treatment with 1% SDS and 250 µg/ml proteinase K, and was evaluated by quantitative PCR (qPCR) with the primers listed in Supplemental Table S2.
Flow cytometry
Cells were subjected to 70% ethanol fixation, followed by 200 µg/ml RNaseA treatment (37 °C for 4 h) and 5 µg/ml propidium iodide staining. DNA content was measured by flow cytometry of the propidium iodide-stained cells using FACSCaliburTM (BD), and the data were analyzed by the CellQuestTM (BD) software.
Pulsed-field gel electrophoresis (PFGE)
PFGE of NotI-digested chromosomal DNA was performed using a CHEF-DR III Pulsed Field Electrophoresis Systems (Bio-Rad) under the following conditions: 1% SeaKem® Gold Agarose (Lonza) or Certified™ Megabase Agarose (Bio-Rad) in 0.5 × TBE; temperature, 10 °C; initial switch time, 40 s; final switch time, 80 s; run time, 18 h; voltage gradient, 6.8 V/cm; and angle, 120°.
Southern blotting
Restriction-digested genomic DNA was separated by conventional agarose gel electrophoresis or PFGE and subjected to Southern blotting. For the telomere probe, telomeric DNA was excised from pNSU7028. To detect telomere-containing NotI restriction fragments of the S. pombe genome (designated L, I, M, and C), the gti1+, mcp3+, fbp1+, and amo1+ gene loci, respectively, were amplified by PCR as previously described29. To generate the rDNA probe for slot blot analyses, an rDNA fragment was amplified by PCR from S. pombe genomic DNA using the following primer set:
st144: 5′-CGCTAACCATTATTTACTGAGGAGAAC-3′
st150: 5′-ATCACCATATCCATATCCAATG-3′
The fragments were labeled with digoxigenin (DIG) using DIG High Prime DNA Labeling and Detection Starter Kit II (Roche), and signals were detected according to the manufacturer’s protocol.
ChIP
Cells harboring a cdc25-22 mutation were grown in YES medium at 25 °C to early-log phase and incubated at 35.5 °C for 4 h. They were then released from G2/M arrest at 25 °C to resume cell cycle progression for 60 min, fixed for 10 min in 1% formaldehyde at 25 °C and further fixed on ice for 50 min (for Sds21-Flag) or for 30 min in 3% paraformaldehyde at 25 °C (for Rif1-12myc). Crude extracts were prepared by breaking the fixed cells in lysis buffer (50 mM HEPES-KOH [pH 7.5], 140 mM NaCl, 1 mM EDTA, 1% Triton X-100, 0.1% sodium deoxycholate, 1xcOmplete [Roche]) using a Multi-beads Shocker (Yasui Kikai), after which the chromatin was sheared by sonication using Bioruptor UCD-250 (BM Equipment) to an average genomic DNA length of <0.5 kb. The insoluble fraction of the crude extracts was removed by centrifugation, and the supernatant was immunoprecipitated with anti-Flag (M2 F3165; Sigma-Aldrich) or anti-Myc (9E10; Santa Cruz Biotechnology) antibodies using Dynabeads® M-280 anti-Mouse IgG (Thermo Fisher) as a carrier. The beads were washed twice with Buffer 1 (50 mM HEPES-KOH [pH 7.5], 140 mM NaCl, 1 mM EDTA, 1% Triton X-100, 0.1% sodium deoxycholate), twice with Buffer 1′ (50 mM HEPES-KOH [pH 7.5], 500 mM NaCl, 1 mM EDTA, 1% Triton X-100, 0.1% sodium deoxycholate), twice with Buffer 2 (10 mM Tris-HCl [pH 8.0], 250 mM LiCl, 1 mM EDTA, 0.5% NP-40, 0.5% sodium deoxycholate), and twice with TE buffer (10 mM Tris-HCl [pH 8.0] and 1 mM EDTA). Immunoprecipitants and input extracts were treated with 10 µg/ml RNase A in TE buffer for 30 min at 37 °C, then with 250 µg/ml proteinase K in 0.25% sodium lauryl sulfate (SDS) at 37 °C overnight, followed by the reverse cross-linking reaction at 65 °C for 6 h. Associated DNA fragments were purified by phenol chloroform extraction and ethanol precipitation, and then analyzed by quantitative PCR (q-PCR) using StepOneTM real-time PCR system (Thermo Fisher). Sequences of the primer sets for subsequent q-PCR are listed in Supplemental Table S2.
References
MacAlpine, D. M. & Bell, S. P. A genomic view of eukaryotic DNA replication. Chromosome Res. 13, 309–326, https://doi.org/10.1007/s10577-005-1508-1 (2005).
Gilbert, D. M. et al. Space and time in the nucleus: developmental control of replication timing and chromosome architecture. Cold Spring Harb. Symp. Quant. Biol. 75, 143–153, https://doi.org/10.1101/sqb.2010.75.011 (2010).
Ogawa, S. et al. Shelterin promotes tethering of late replication origins to telomeres for replication-timing control. EMBO J. 37, e98997, https://doi.org/10.15252/embj.201898997 (2018).
Cooper, J. P., Nimmo, E. R., Allshire, R. C. & Cech, T. R. Regulation of telomere length and function by a Myb-domain protein in fission yeast. Nature 385, 744–747, https://doi.org/10.1038/385744a0 (1997).
Kanoh, J. & Ishikawa, F. spRap1 and spRif1, recruited to telomeres by Taz1, are essential for telomere function in fission yeast. Curr. Biol. 11, 1624–1630 (2001).
Hayano, M. et al. Rif1 is a global regulator of timing of replication origin firing in fission yeast. Genes Dev. 26, 137–150, https://doi.org/10.1101/gad.178491.111 (2012).
Tazumi, A. et al. Telomere-binding protein Taz1 controls global replication timing through its localization near late replication origins in fission yeast. Genes Dev. 26, 2050–2062, https://doi.org/10.1101/gad.194282.112 (2012).
Kanoh, Y. et al. Rif1 binds to G quadruplexes and suppresses replication over long distances. Nat. Struct. Mol. Biol. 22, 889–897, https://doi.org/10.1038/nsmb.3102 (2015).
Zofall, M., Smith, D. R., Mizuguchi, T., Dhakshnamoorthy, J. & Grewal, S. I. S. Taz1-shelterin promotes facultative heterochromatin assembly at chromosome-internal sites containing late replication origins. Mol. Cell 62, 862–874, https://doi.org/10.1016/j.molcel.2016.04.034 (2016).
Duncker, B. P. & Brown, G. W. Cdc7 kinases (DDKs) and checkpoint responses: lessons from two yeasts. Mutat Res 532, 21–27 (2003).
Dave, A., Cooley, C., Garg, M. & Bianchi, A. Protein phosphatase 1 recruitment by Rif1 regulates DNA replication origin firing by counteracting DDK activity. Cell Rep. 7, 53–61, https://doi.org/10.1016/j.celrep.2014.02.019 (2014).
Ferreira, M. G. & Cooper, J. P. The fission yeast Taz1 protein protects chromosomes from Ku-dependent end-to-end fusions. Mol. Cell 7, 55–63 (2001).
Miyoshi, T., Kanoh, J., Saito, M. & Ishikawa, F. Fission yeast Pot1-Tpp1 protects telomeres and regulates telomere length. Science 320, 1341–1344, https://doi.org/10.1126/science.1154819 (2008).
Chikashige, Y. et al. Membrane proteins Bqt3 and -4 anchor telomeres to the nuclear envelope to ensure chromosomal bouquet formation. J. Cell Biol. 187, 413–427, https://doi.org/10.1083/jcb.200902122 (2009).
de Lange, T. How telomeres solve the end-protection problem. Science 326, 948–952, https://doi.org/10.1126/science.1170633 (2009).
Fujita, I. et al. Telomere-nuclear envelope dissociation promoted by Rap1 phosphorylation ensures faithful chromosome segregation. Curr. Biol. 22, 1932–1937, https://doi.org/10.1016/j.cub.2012.08.019 (2012).
Chikashige, Y. & Hiraoka, Y. Telomere binding of the Rap1 protein is required for meiosis in fission yeast. Curr. Biol. 11, 1618–1623 (2001).
Fujita, I., Tanaka, M. & Kanoh, J. Identification of the functional domains of the telomere protein Rap1 in Schizosaccharomyces pombe. PLoS One 7, e49151, https://doi.org/10.1371/journal.pone.0049151 (2012).
Nakamura, T. M., Cooper, J. P. & Cech, T. R. Two modes of survival of fission yeast without telomerase. Science 282, 493–496 (1998).
Jain, D., Hebden, A. K., Nakamura, T. M., Miller, K. M. & Cooper, J. P. HAATI survivors replace canonical telomeres with blocks of generic heterochromatin. Nature 467, 223–227, https://doi.org/10.1038/nature09374 (2010).
Yamano, H., Ishii, K. & Yanagida, M. Phosphorylation of dis2 protein phosphatase at the C-terminal cdc2 consensus and its potential role in cell cycle regulation. EMBO J 13, 5310–5318 (1994).
Rivera-Mulia, J. C. & Gilbert, D. M. Replication timing and transcriptional control: beyond cause and effect-part III. Curr. Opin. Cell Biol. 40, 168–178, https://doi.org/10.1016/j.ceb.2016.03.022 (2016).
Moreno, S., Klar, A. & Nurse, P. Molecular genetic analysis of fission yeast Schizosaccharomyces pombe. Methods Enzymol. 194, 795–823 (1991).
Bahler, J. et al. Heterologous modules for efficient and versatile PCR-based gene targeting in Schizosaccharomyces pombe. Yeast 14, 943–951, 10.1002/(SICI)1097-0061(199807)14:10<943::AID-YEA292>3.0.CO;2-Y (1998).
Forsburg, S. L. & Rhind, N. Basic methods for fission yeast. Yeast 23, 173–183, https://doi.org/10.1002/yea.1347 (2006).
Hayashi, M. et al. Genome-wide localization of pre-RC sites and identification of replication origins in fission yeast. EMBO J. 26, 1327–1339, https://doi.org/10.1038/sj.emboj.7601585 (2007).
Katou, Y. et al. S-phase checkpoint proteins Tof1 and Mrc1 form a stable replication-pausing complex. Nature 424, 1078–1083, https://doi.org/10.1038/nature01900 (2003).
Sugawara, N. F. DNA sequences at the telomeres of the fission yeast S. pombe. Ph.D. Thesis, Harvard University (1988).
Tashiro, S., Nishihara, Y., Kugou, K., Ohta, K. & Kanoh, J. Subtelomeres constitute a safeguard for gene expression and chromosome homeostasis. Nucleic Acids Res. 45, 10333–10349, https://doi.org/10.1093/nar/gkx780 (2017).
Acknowledgements
We thank H. Masukata and S. Ogawa for strains, experimental protocols and discussion, T. Sakuno for technical assistance and discussion for flow cytometric analyses, A. Oda, N. Takemata and K. Ohta for experimental support, National BioResource Project (NBRP) in Japan for strains, and all former and present Kanoh laboratory members for discussion and support. This study was supported by JSPS KAKENHI Grant Numbers 17H03606, 26290061 and 19H05262, research grants from Ohsumi Frontier Science Foundation, and the publication support from Initiative for the Implementation of the Diversity Research Environment of Osaka University to J.K.
Author information
Authors and Affiliations
Contributions
J.K. conceived the project, Y.H. and J.K. designed the experiments, and Y.H., M.Y., J.M. and J.K. conducted the experiments, and J.K. wrote the manuscript. All authors analyzed the data and reviewed the manuscript.
Corresponding author
Ethics declarations
Competing Interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Hasegawa, Y., Yamamoto, M., Miyamori, J. et al. Telomere DNA length-dependent regulation of DNA replication timing at internal late replication origins. Sci Rep 9, 9946 (2019). https://doi.org/10.1038/s41598-019-46229-1
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-019-46229-1
- Springer Nature Limited