Abstract
Laonice Malmgren, 1867 (Annelida: Spionidae) is a common polychaete genus in the deep-sea. Although most species are quite well studied morphologically, fragmentation and other damage that occurs during sampling often hampers morphological species identification of deep-sea specimens. In this study, we employ three molecular markers (16S, COI and 18S) to study the biodiversity and the distribution patterns of Laonice from the tropical North Atlantic and the Puerto Rico Trench. Based upon different molecular analyses (Automated Barcode Gap Discovery, pairwise genetic distances, phylogenetics, haplotype networks) we were able to identify and differentiate eight Laonice species. Up to four of these species co-occurred sympatrically at the same station. The majority of species were found at multiple stations and two species in the eastern as well as western Atlantic had ranges of up to 4,000 km. Genetic differentiation across these extensive geographic distances was very low. Surprisingly, one 16S haplotype was shared between individuals 2,776 km apart and individuals from the Caribbean and the abyssal plain in the eastern Atlantic (>3,389 km) differed in only a single mutation in 16S. Our results suggest that members of this genus successfully disperse across large geographic distances and are largely unaffected by topographic barriers.
Similar content being viewed by others
Introduction
Spionidae Grube, 18501 is one of the most abundant and diverse groups of polychaetes and occur in almost all marine habitats, from shallow waters to the deep-sea2. All spionids are characterized by a pair of long palps, used for deposit or suspension feeding; most species are tube-dwellers, but free-living or commensal species are also found within the taxon3,4. Like several other annelid taxa, Spionidae are soft-bodied and very fragile and are, therefore, rarely found undamaged in deep-sea samples. These incomplete and fragmented individuals often lack crucial taxonomic characters, hampering their identification5. Nonetheless, the spionid genus Laonice Malmgren, 18676 is well studied, especially species from the deep sea of the North Atlantic7,8,9,10. To facilitate the identification of Laonice species extensive studies on species-specific characters were conducted and four subgenera were suggested based on morphological characters8,11. However, the recently published first molecular phylogenetic study on Laonice rejected two of these four subgenera5. Several Laonice species have been reported from a wide geographical range, and the presumed long planktonic life and planktotrophic larvae would offer the potential for long-distance dispersal12,13,14. However, Laonice cirrata (Sars, 185115), a presumed widespread species, was shown to probably represent several geographically restricted species5,16.
The abyssal Atlantic Ocean is divided by the Mid-Atlantic Ridge (MAR) longitudinally into eastern and western basins17. Due to its geology, the MAR is believed to represent a dispersal barrier for some components of the abyssal benthic fauna18,19,20,21. However, the MAR is not a closed barrier as several Fracture Zones interrupt it. When two tectonic plates passing each other in parallel to their original motions, a so-called transform fault is formed at the offsets of the ridge22. Over geological time the movement results in an extension past the transform fault in opposite directions, the Fracture-Zones23.
Our study area encompasses the abyssal eastern and western basins in the tropical North Atlantic along the Vema Fracture Zone as well as the Puerto Rico Trench. The first morphological studies rejected a barrier effect of the MAR on the distribution of selected widespread spionid species in the abyss of the tropical North Atlantic, though other species were found to be limited to either side of the MAR24. However, the presence of morphologically cryptic species could not be ruled out.
The aim of this study is to investigate the diversity and distribution of Laonice from the tropical North Atlantic and the Puerto Rico Trench with molecular tools and further assess the potential barrier effect of the MAR on abyssal spionid taxa.
Material and Methods
Collection and identification of specimens
All analysed specimens were collected from the tropical North Atlantic and the Puerto Rico Trench during the VEMA-Transit expedition in December 2014–January 2015 (Fig. 1, Supplement 1). Sampling was conducted with a camera-equipped epibenthic sledge at depths between 4918–5736 m, followed by a fixation of either cooled 96% ethanol or 4% buffered formalin. More detailed information about sample treatment and sampling localities are described in Guggolz et al.24 and Devey et al.25. According to the geographical position, four areas were defined as following: the eastern part of the Vema-Fracture Zone (eVFZ), extending eastwards from the MAR in the Cape Verde Basin; the western part of the Vema Fracture Zone (wVFZ), extending westwards from the MAR in the Demerara Basin; the Vema Transform Fault (VTF), located between these two areas in the MAR; the Puerto Rico Trench (PRT), located in the shallower part of the trench near Puerto Rico (Fig. 1). Distances between areas varied between 276 km (wVFZ) and 1,298 km (eVFZ). The eastern-most and western-most studied sites were separated by 4,610 km (Table 1).
All specimens were sorted and identified at least to genus level using stereo zoom and compound microscopes. All specimens identified as Laonice and aff. Lindaspio24 were analysed. The identification of the latter has been revised and reassigned to Laonice (unpublished data). Specimens have been deposited in the collection of the Center of Natural History (Universität Hamburg, Germany) (Supplement 1).
DNA extraction, PCR amplification, sequencing and alignment
DNA was extracted with Chelex 100. Depending on the size of specimens, one or two parapodia were dissected and transferred into 30 µl of 10% Chelex solution in purified water and incubated for 30 minutes at 56 °C and 10 minutes at 99 °C. Polymerase Chain Reactions (PCR) were performed with a total volume of 15 µl consisting of 1.5 µl DNA extract, 7.5 µl AccuStart II PCR ToughMix (Quanta Bio, Germany), 0.6 µl of each primer (10mmol), 0.3 µl of GelTrack loading dye (QuantaBio, Germany) and 4.8 µl Millipore H2O. Fragments of mitochondrial (16S and COI) and nuclear (18S) rRNA genes were amplified (see Table 2 for list of all primers). PCR amplification had an initial denaturation step of 94 °C for 3 min, followed by 35 cycles of 30 s at 94 °C, 45 s at 43 °C and 45 sec at 72 °C, followed by a final elongation step for 5 min at 72 °C. Success of amplification was determined via gel electrophoresis on 1% agarose/TAE gel. For sequencing, 8 µl of the PCR products were purified using FastAP (1.6 µl; 1 U/µl) and Exonuclease I (0.8 µl; 20 U/µl) (Thermo Fisher Scientific, Germany) with an incubation time of 37 °C for 15 min followed by 15 min with 85 °C and a final holding temperature of 14 °C. Purified PCR products were sent to Macrogen Europe, Inc. (Amsterdam-Zuidoost, Netherlands) for sequencing. All in all, 80 specimens were successfully sequenced for 16S, a subset of 27 specimens for COI and 47 specimens for 18S. Sequences were assembled and corrected with Geneious 6.1.8 (http://www.geneious.com)26 and all sequences were deposited in GenBank (for accession numbers see Supplement 1). The obtained sequences of the different gene fragments were aligned separately using MUSCLE27 implemented in Genious 6.1.8.
Initial identification of species, phylogenetic analyses and haplotype networks
To obtain a first estimation of the number of species among Laonice investigated, the Automated Barcode Gap Discovery (ABGD28) was conducted separately for each of the three genes (16S, COI, 18S). The ABGD identifies potential barcoding gaps separating hypothetical species, based on the assumption that interspecific genetic distances are larger than intraspecific distances. The ABGD analysis was run on the web-based version of the software (http://wwwabi.snv.jussieu.fr/public/abgd/abgdweb.html), using uncorrected p distances (Table 3), which were calculated with MEGA729 on all available sequences. Standard settings were kept, except for Pmin (0.005), the numbers of steps (100) and the relative gap width (X = 0.5).
To assess the phylogenetic relationships among the studied specimens and to assess whether the lineages suggested by ABGD are monophyletic, phylogenetic analyses were performed with Bayesian inference. All three gene fragments were analysed separately and concatenated with MrBayes (version 3.230) online with CIPRES Science Gateway V.3.3 (www.phylo.org)31. For the analyses of the 16S and COI genes, Marenzelleria neglecta Sikorski & Bick, 200432, Malacoceros indicus (Fauvel, 1928)33, Polydora hoplura Claparède, 186834 and Spio blakei Maciolek, 199035 were employed as outgroups (Supplement 2). Four chains were run for 107 generations, with sampling every 1200th generation, and discarding the first 25% as burn-in. The GTR + I + G substitution model was identified by MEGA7 as the best fitting model under the AIC criterion.
Guggolz et al.24 studied the same Laonice individuals morphologically. That data was used to identify morphological differences between the herein delimited species and to add another line of evidence for species delimitation.
To assess the genus-wide phylogenetic relationships of the herein studied Laonice and to find out whether any of these species have a wider distribution than anticipated by our own data, a phylogenetic analysis with Laonice sequences available from GenBank was conducted for COI and 16S. Additional data includes: Laonice from expeditions around Iceland (IceAGE I + II5) and other GenBank entries14,36,37,38,39,40,41,42,43 (Supplement 2). The genus-wide analysis was focussed on the COI data, because of the more comprehensive COI data being available (Supplement 2), even if analysis with 16S data was also conducted (Supplement 3).
To better visualize the geographic distribution of the genetic diversity median-joining haplotype networks were generated with Network 5.0.0.344 (http://fluxus-engineering.com/) for each gene fragment. The generated haplotype networks were redrawn with Adobe Illustrator CS6.
Analyses of population differentiation were performed with Arlequin 3.545 for species with sufficiently large specimen numbers (at least four specimens per site). Pairwise Φst was calculated for Laonice sp. D, F, H (16S). For Laonice sp. D areas eVFZ and wVFZ, for Laonice sp. F areas eVFZ and VTF and for Laonice sp. H the areas eVFZ, wVFZ and VTF were compared (Tables 4 and 5).
Results
Alignment
The alignment of the 16S fragment included a total of 79 sequences with a length of 525 bp, of which 223 bp were variable and 165 bp were parsimony informative. The COI alignment featured 26 sequences and had a length of 694 bp, of which 283 bp were variable and 207 bp parsimony informative. The alignment contained no indels and the derived amino acid alignment consisted of 208 amino acids, with 16 variable amino acids and no stop codons. The genus-wide COI alignment featured 134 sequences (including outgroup) and had a length of 683 bp, of which 324 were variable and 301 were parsimony informative. The alignment of the 18S fragment consisted of 46 sequences with 2195 bp, of which only 68 bp were variable and 34 bp were parsimony informative.
Species delimitation
The ABGD analysis of the 16S dataset retrieved eight main lineages when barcode thresholds of 0.5–4.5% were employed. For now, we use the term lineages rather than species, as not all of them necessarily correspond to species. To the eight lineages, we will refer to as Laonice sp. A–H. With higher threshold values several lineages collapsed (4.6–6.5% = 3 lineages), or all lineages collapsed into a single lineage (>6.7%). The analysis of the COI dataset resulted in seven lineages (barcode thresholds 0.5–10%). The seven lineages identified with COI are in full agreement with the lineages derived with 16S, with the same specimens being clustered together. The discrepancy between 16S and COI is due to the absence of one lineage, Laonice sp. E (PVT 471_I), which was not successfully sequenced for COI. Pairwise genetic distances (uncorrected p-distances) between the lineages ranged for 16S from 2.8–23%, for COI from 8.6–23% and for 18S from 0–2.8% (based on the eight lineages derived by 16S) (Table 3). The lowest pairwise distances were found between the lineages F and G (16S: 2.8–4%; COI: 8.6–8.8%; 18S: 0%), whereas all other pairwise distances between the lineages were higher than 4.1% for 16S, 12.2% for COI and 0.1% for 18S. Within lineages, the highest observed pairwise distances were 1.7% for 16S, 0.7% for COI and 0.3% for 18S (Table 3).
The phylogenetic analyses of COI and 16S recovered lineages A-H as reciprocal monophyletic with full support each (Fig. 2). Also the phylogenetic relationships among the lineages were very similar for 16S and COI. Laonice sp. F and G are sister species (in 16S, Laonice sp. E clusters with these two species), as are Laonice sp. C and B as well as Laonice sp. D and H. Differences between the analyses of the 16S and COI data are found in the position of Laonice sp. A. In COI, Laonice sp. A is found to be a sister taxon to Laonice sp. B and C (Fig. 2a), whereas in 16S Laonice sp. A is placed as a sister taxa to all other species (Fig. 2b).
Phylogenetic tree of Laonice specimens from the Vema-Transit expedition based on mitochondrial 16S (a) and COI (b) gene fragments. Posterior probabilities shown next to the nodes (values below 0.8 are not shown). Morphological identification after Guggolz et al. 201824 are color coded (see legend in the middle).
The haplotypes networks of the different gene fragments (16S, COI and 18S) showed slightly different patterns (Fig. 3a–c). For 16S, with the highest number of sequenced individuals, 27 haplotypes (h1-16S-h27-16S) were found with a maximum of eight haplotypes in one lineage (Laonice sp. D, Fig. 3a). Networks of COI dataset showed a total of 18 different haplotypes (h1-COI–h18-COI) with a maximum of six haplotypes within the same lineage as in 16S (Laonice sp. D, Fig. 4b). For 18S the smallest genetic diversity was found with 17 haplotypes (h1-18S–h17-18S; Fig. 3c). The low number of mutational steps between haplotypes, as evidenced in the 18S network (Fig. 3c), is probably responsible for the lower resolution in the phylogenetic analysis of this gene when it comes to species delimitation. The 18S network shows that Laonice sp. A, B and C are well differentiated from each other and the other lineages. Laonice sp. D, E, F, G and H all have very similar haplotypes and do not form well differentiated clusters. Laonice sp. F and G even share their only haplotype.
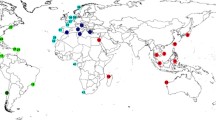
Phylogenetic tree of Laonice specimens from the Atlantic, Antarctic and Pacific Ocean based on the mitochondrial COI gene fragment. Posterior probabilities shown next to the nodes (values below 0.8 are not shown). Sampling localities and depth are colour coded (see legend in upper right-hand corner).
As all investigated specimens were incomplete or damaged and relatively short (maximum 22 segments), the main characters for species identification were the shape of the prostomium, the beginning of the lateral pouches, the beginning of the sabre chaeta and the beginning and number of teeth of the neuropodial hooks, as well as the length of the nuchal organ (Table 6).
Slight morphological variations were observed between the eight species delimitated with molecular analyses. For instance, Laonice sp. F and sp. G differ in the beginning of the lateral pouches (sp. F: 3rd chaetiger; sp. G: 4th chaetiger), the beginning of the sabre chaeta (sp. F: 10th chaetiger; sp. G: 8th chaetiger) as well as the length of the nuchal organ (sp. F: until 9th chaetiger; sp. G: until 8th chaetiger) (Table 6). Furthermore, Laonice sp. B was the only species with the peri- and prostomium fused and in Laonice sp. E the beginning of the neuropodial hooks was observed more posteriorly than in all other species (16th chaetiger). Laonice sp. A differed from all other species, as the nuchal organ reached the end of the available fragments (until 18th chaetiger) and a prominent dorsolateral ridge was present from chaetiger 8 ̶11 (Table 6).
Distribution of species
In the genus-wide phylogenetic analysis of COI with Laonice species from the Atlantic, the Southern Ocean, Russian waters and the North-East Pacific, all Laonice lineages identified herein were recovered as monophyletic, and none of these seemed to be conspecific with any of the published Laonice sequences (Fig. 4). Laonice sp. D, F, G and H constitute a monophylum, within a clade including Laonice blakei Sikorski and Jirkov in Sikorski et al.46 and Laonice sp. b sensu Bogantes et al.5, both sampled from Icelandic waters. Laonice sp. B and C constitute a monophyletic group that is sister to a large clade of Laonice species, including Laonice sp. A, from various localities (Fig. 4).
Five of the lineages were only recorded in one of the four areas: Laonice sp. A and B in the eVFZ, Laonice sp. C and E in the wVFZ and Laonice sp. G in the PRT (Fig. 3). These five lineages were relatively rarely collected with three specimens at most (Supplement 1). In contrast, the other three lineages were recorded at larger geographic scales, in either the eVFZ and VTF (Laonice sp. F) or even in all four areas (Laonice sp. D and H). Even single haplotypes of these lineages exhibited such extensive distributions and were recorded in all of these areas, except PRT (16S: h5, h7, h15, h17, h23; COI: h5, h15; 18S: h7, h13, h14, h15; Fig. 3). For example, Laonice sp. D had one haplotype in each of the three studied genes that occurred in the eVFZ, VTF as well as the wVFZ (Fig. 3: h7-16S, h5-COI, h7-18S).
Population differentiation was not significant, neither between different sites, nor between different areas for the three widely distributed lineages Laonice sp. D, F and H (Tables 4 and 5).
Discussion
Employing mitochondrial markers (16S and COI) we were able to identify eight lineages well supported and consistently delimited. Following a strict DNA barcoding approach (sensu Hebert et al.47), these results might easily be interpreted as eight species. However, mitochondrial markers are linked and thus not independently inherited. Therefore, consistency among these markers does not necessarily equate reproductive isolation among the respective lineages48. Consistency with other marker types - e.g., nuclear markers or morphology - does offer the possibility to delimit species adequately49,50. Taken all data together, lineages A, B, C, D, E and H can be easily delimited as distinct species, even though the differentiation is less pronounced between lineages D, E and H in 18S. The lack of shared haplotypes, despite their sympatric distribution over large geographic scales, is a good indication of reproductive isolation among them. Lineages F and G shared an identical 18S haplotype and also their pairwise uncorrected distances were the lowest for all pairs of lineages (COI: 8.6–8.8%; 16S: 2.8–4.0%). However, a lack of differentiation in 18S may not be surprising for recently diverged species and the levels of differentiation in COI and 16S are comparable to those observed among other polychaete species, which usually exceeded 5–6% for COI51,52,53,54,. Intraspecific distances were always lower than interspecific distances with a maximum of 1.7% within Laonice sp. F for 16S and 0.7% within Laonice sp. H for COI (Table 3), similar to the 0–2% uncorrected distances found within Laonice species from the North-Atlantic5. These results could imply thresholds of about 2% for 16S and 2–8% for COI to distinguish between Laonice species.
The present molecular study reveals inconsistencies with previous morphology based studies24. Guggolz et al.24 identified six species (aff. Lindaspio sp. 1, Laonice sp. 1, 4, 5, 6 and Laonice cf. blakei). The majority of specimens were identified as L. cf. blakei (about 86.5% of the identified Laonice specimens) and the slight variations observed between individuals were interpreted as intraspecific variability within L. cf. blakei. Of the six species identified based on their morphology by Guggolz et al.24, only Laonice sp. A (aff. Lindaspio sp. 1 in Guggolz et al.24) and sp. C (Laonice sp. 6 in Guggolz et al.24) could be confirmed in our molecular analyses. Specimens identified as Laonice cf. blakei by Guggolz et al.24 are here assigned to six different species based on the results from molecular studies: Laonice sp. B, D, E, F, G and H. Furthermore, Laonice sp. 2, 4, and 5 are all included in Laonice sp. D and Laonice sp. 1 included in Laonice sp. F (see Fig. 2). Most of the disagreement between the morphological study and the present results can be explained as misinterpretations of morphological differences as intraspecific variability rather than interspecific variation. The slight differences observed among individuals identified as Laonice cf. blakei probably represent interspecific variation between several species of Laonice. Taken together with the molecular data, these variations lend additional support for differentiating the eight species identified herein. For instance, Laonice sp. F and sp. G, sharing the same 18S haplotype, showed differences in their morphology, supporting a separation at the species level. Comparable morphological differences can be found for Laonice sp. A–E as well.
These morphological patterns support the differentiation of the eight lineages and we therefore propose that these eight lineages represent eight species. The lack of differentiation in 18S is probably caused by a combination of a low substitution rate and incomplete lineage sorting55 rather than ongoing reproduction among these species.
Apart from delimiting species, we were interested in distribution patterns of the species. Even over large geographic distances (>4,000 km; Table 1, Fig. 3), there seems to be no genetic differentiation within some species. This is most obvious for species distributed across the MAR (Laonice sp. D, H), as the same haplotypes are found in the eVFZ and the wVFZ. Species restricted to only one (Laonice sp. A and B in the eVFZ) or two of the areas (Laonice sp. F in the eVFZ and the VTF) exhibited identical haplotypes across distances of hundreds of kilometres. These species might represent rare species and we could have missed them in the other areas due to the sampling design, as we managed to obtain a higher number of individuals from the eVFZ compared to the other sampled areas24,25 (Supplement 1). The present data suggest gene flow over the MAR or potentially through Fracture Zones in the tropical North Atlantic, supported by the low and non-significant levels of differentiation among populations (Laonice sp. D, F and H). Guggolz et al.24 already suggested that the Mid-Atlantic Ridge (MAR) does not represent a physical barrier for some polychaetes based on morphological studies and the lack of significant differences between the eastern and western sides of the ridge. A widespread distribution over 4,000 km was never proven genetically for Laonice, but it was reported for other abyssal taxa like Aurospio dibranchiata Maciolek, 198156, a polychaete species occurring in different oceans37 and Nicomache lokii Kongsrud & Rapp, 201257 and Sclerolinum contortum Smirnov, 200058, polychaetes living in chemosynthetic-based ecosystems distributed from the Arctic to Antarctic59. Larval distribution is suggested to play a major role in the efficiency of the distribution of deep-sea invertebrates, even if the specific larvae are unknown for most species60. The exact types of development of the investigated Laonice specimens from the tropical North Atlantic is unknown, but in general Laonice is supposed to have long-lived larvae and very high dispersal capabilities12,14,24. The development strategies seem to be highly connected with the ability to distribute in the abyss even with potential topographic barriers like ridges, rises or canyons. For instance, different molluscs with planktonic larvae were reported to be able to distribute over such barriers61,62. Contrary, taxa with direct development, such as brooding isopods, were found to have a restricted distribution with limited or no gene flow across the MAR18,63.
None of the eight species recorded in the tropical North Atlantic were found to be conspecific with Laonice species for which published genetic data was available. Bogantes et al.5 recently performed first phylogenetic studies on Laonice and suggested that the Antarctic was colonized several times independently. A comparable pattern can be found in our study. Nonetheless, one should keep in mind that these results are based only on one gene (COI) and only a small proportion of known Laonice species are included.
Until now, around 16 deep-sea Laonice species have been described, mainly based on morphology9. Unfortunately, it is not possible to perform subsequent molecular studies with most of the described material, due to fixation, unless new material is collected from the respective type localities. Identification of Laonice specimens from deep-sea samples is almost always difficult due to fragmentation and the subsequent loss of important characters independently of the fixation method5,24. Therefore, molecular techniques might be of great importance for a correct estimation of their diversity. DNA extraction from fresh material before fixation in formalin takes place would be an appropriate way to combine morphology and molecular studies in soft-bodied animals like spionid polychaetes and should be part of the workflow during sampling.
The present study gives new insights into the phylogeny of Laonice and stresses the importance of molecular analyses for estimates of species diversity, ideally combined with morphological studies. The eight Laonice species identified in the tropical North Atlantic might be new to science, and certainly do not belong to any of the Laonice species investigated with molecular tools to date. Due to the incomplete specimens and thus the absence of important morphological characters, a clear differentiation from all described Laonice species is impossible. Therefore, at present the identified lineages cannot be described as new species. However, molecular data is sparse for the genus and new information would further improve our understanding of the evolution of Laonice and the dynamics of speciation in the deep-sea. Our present study highlights the importance of integrative taxonomy to allow species delimitation in deep-sea spionids.
The genus’ potential to disperse over large geographic distances in the deep-sea and across topographic barriers such as ridges is shown here and support the hypothesis of other studies14,64. We were able to show the occurrence of the same Laonice species from the Caribbean to the abyssal plain near West-Africa, highlighting for the first time such a wide distribution for a species of this genus based on molecular analyses. These dispersal abilities are also notable for annelids in general, showing the relevance of molecular tools for our understanding of their distribution in the deep-sea.
References
Grube, A. E. Die Familien der Anneliden. Archiv für Naturgeschichte. 16, 249–364 (1850).
Blake, J. A., Maciolek, N. J. & Meißner, K. Spionidae Grube, 1850 in Handbook of Zoology, a Natural History of the Phyla of the Animal Kingdom—Annelida, Polychaetes (ed. Schmidt-Rhaesa, A.) 1–109 (De Gruyter, 2017).
Glasby, C. J. et al. Class polychaeta in Polychaetes & Allies: The Southern Synthesis. Fauna of Australia (eds Beesley, P. L., Ross, B., Glasby, C. J.) 1–296 (Melbourne: CSIRO Publishing 2000).
Jumars, P. A., Dorgan, K. M. & Lindsay, S. M. Diet of worms emended: an update of polychaete feeding guilds. Annu. Rev. Mar. Sci. 7, 497–520 (2015).
Bogantes, V. E., Halanych, K. M. & Meißner, K. Diversity and phylogenetic relationships of North Atlantic Laonice Malmgren, 1867 (Spionidae, Annelida) including the description of a novel species. Mar. Biodivers, 1–13 (2018).
Malmgren, A. J. Annulata Polychaeta Spetsbergiæ, Groenlandiæ, Islandiæ et Scandinaviæ. Hactenus Cognita. Ex Officina Frenckelliana, Helsingforslæ. 127 (1867).
Maciolek, N. J. New species and records of Aonidella, Laonice, and Spiophanes (Polychaeta: Spionidae) from shelf and slope depths of the Western North. Atlantic. B. Mar. Sci. 67, 529–548 (2000).
Sikorski, A. V. Laonice (Polychaeta, Spionidae) in the Arctic and the North Atlantic. Sarsia 88, 316–345 (2003).
Sikorski, A. V., Gunton, L. M. & Pavlova, L. Laonice species (Polychaeta, Spionidae) from the Whittard Canyon (NE Atlantic) with descriptions of two new species. J. Mar. Biol. Assoc. U. K. 97, 961–973 (2017).
Sikorski, A. V. & Pavlova, L. Three new species of Laonice (Polychaete: Spionidae) from West and Southwest Africa. Zootaxa 4097, 353–368 (2016).
Sikorski, A. V. Review of Laonice (Spionidae, Annelida) with remarks on several species and a description of a new species from South Africa. Ital. J. Zool. 78, 37–41 (2011).
Blake, J. A. Spionida in Reproductive biology and phylogeny of Annelida (eds Rouse, G. and Pleijel, F.) 565–638 (Science Publishers, Enfield 2006).
Wilson, W. H. Sexual reproductive modes in polychaetes: classification and diversity. B- Mar. Sci. 48, 500–516 (1991).
Meißner, K., Bick, A., Guggolz, T. & Götting, M. Spionidae (Polychaeta: Canalipalpata: Spionida) from seamounts in the NE Atlantic. Zootaxa 3786, 201–245 (2014).
Sars, M. Beretning om en i Sommeren 1849 foretagen zoologisk Reise i Lofoten og Finmarken [1850 date used in Hydrozoa, 1851 in Polychaeta & others]. Nyt Magazin for Naturvidenskaberne 6, 121–211 (1851).
Sikorski, A. V. On distinguishing of Laonice cirrata (Sars, 1851) and Laonice bahusiensis Söderström, 1920 (Polychaeta: Spionidae) – the two morphologically related species. Zool Zhurnal 81(4), 406–419 (2002).
Murray, J. & Hjort, J. The Depths of the Ocean. (McMillan, London 1912).
Bober, S., Brix, S., Riehl, T., Schwentner, M., Brandt, A. Does the Mid-Atlantic Ridge affect the distribution of abyssal benthic crustaceans across the Atlantic Ocean? Deep-Sea Res. Pt II 148, 91-104 (2018).
Brandt, A. et al. Composition of abyssal macrofauna along the Vema Fracture Zone and the hadal Puerto Rico Trench, northern tropical Atlantic. Deep-Sea R es. Pt II 148, 35-44 (2018).
McClain, C. R., Rex, M. A. & Etter, R. J. Deep-sea macroecology in Marine Macroecology (eds. Witman, J. D., Roy, K.) 65–100 (Chicago, IL: University of Chicago Press 2009).
Priede, I. G. et al. Does presence of a mid-ocean ridge enhance biomass and biodiversity? PLoS One 8(5), e61550 (2013).
Ball, M. M. & Harrison, C. A. Crustal plates in the central Atlantic. Science 167(3921), 1128–1129 (1970).
van Andel, T. H., von Herzen, R. P. & Phillips, J. D. The Vema fracture zone and the tectonics of transverse shear zones in oceanic crustal plates. Mar. Geophys. Res. 1(3), 261–283 (1971).
Guggolz, T., Lins, L., Meißner, K. & Brandt, A. Biodiversity and distribution of polynoid and spionid polychaetes (Annelida) in the Vema Fracture Zone, tropical North Atlantic. Deep-Sea Res. Pt II 148, 54–63 (2018).
Devey, C. W. RV SONNE Fahrtbericht/Cruise Report SO237 Vema-TRANSIT: bathymetry of the Vema-Fracture-Zone and Puerto Rico TRench and Abyssal AtlaNtic BiodiverSITy Study, Las Palmas (Spain)-Santo Domingo (Dom. Rep.) 14.12.14–26.01.15 (2015).
Kearse, M. et al. Geneious Basic: an integrated and extendable desktop software platform for the organization and analysis of sequence data. Bioinformatics 28.12, 1647–1649 (2012).
Edgar, R. C. MUSCLE: multiple sequence alignment with high accuracy and high throughput. Nucleic Acids Res. 32, 1792–1797 (2004).
Puillandre, N., Lambert, A., Brouillet, S. & Achaz, G. ABGD, Automatic Barcode Gap Discovery for primary species delimitation. Mol. Ecol. 21(8), 1864–1877 (2012).
Kumar, S., Stecher, G. & Tamura, K. MEGA7: Molecular Evolutionary Genetics Analysis version 7.0 for bigger datasets. Mol. Biol. Evol. 33, 1870–1874 (2016).
Ronquist, F. & Huelsenbeck, J. P. MrBayes 3: Bayesian phylogenetic inference under mixed models. Bioinformatics 19(12), 1572–1574 (2003).
Miller, M. A., Pfeiffer, W. & Schwartz, T. Creating the CIPRES Science Gateway for inference of large phylogenetic trees in Proceedings of the Gateway Computing Environments Workshop (GCE) 1–8. (New Orleans, LA 2010).
Sikorski, A. V. & Bick, A. Revision of Marenzelleria Mesnil, 1896 (Spionidae, Polychaeta). Sarsia 89(4), 253–275 (2004).
Fauvel, P. Annélides Polychètes nouvelles de l’Inde in I. Bulletin du Muséum d’Histoire Naturelle, 34(1), 90–96 (Paris, 1928).
Claparède, É. Les annélides chétopodes du Golfe de Naples. Mémoires de la Société de physique et d’histoire naturelle de Genève 19, 313–584 (1868).
Maciolek, N. J. A. redescription of some species belonging to the genus Spio and Microspio (Polychaeta: Annelida) and descriptions of three new species from Northwestern Atlantic Ocean. J. Nat. Hist. 24, 1109–1141 (1990).
Bastrop, R. & Blank, M. Multiple invasions–a polychaete genus enters the Baltic Sea. Biol. Invasions 8(5), 1195–1200 (2006).
Mincks, S. L., Dyal, P. L., Paterson, G. L. J., Smith, C. R. & Glover, A. G. A new species of Aurospio (Polychaeta, Spionidae) from the Antarctic shelf, with analysis of its ecology, reproductive biology and evolutionary history. Mar. Ecol. 30, 181–197 (2009).
Brasier, M. J. et al. Distributional Patterns of Polychaetes across the West Antarctic Based on DNA Barcoding and Particle Tracking. Analyses. Front. Mar. Sci. 4, 356 (2017).
Carr, C. M., Hardy, S. M., Brown, T. M., Macdonald, T. A. & Hebert, P. D. N. A tri-oceanic perspective: DNA barcoding reveals geographic structure and cryptic diversity in Canadian Polychaetes. PLoS One (2011).
Gallegos, R., Lavery, S. & Sewell, M. A. The meroplankton community of the oceanic Ross Sea during late summer. Antarc. Sci. 26(4), 345–360 (2014).
Williams, L.-G., Karl, S. A., Rice, S. & Simon, C. Molecular identification of polydorid polychaetes (Annelida: Spionidae): is there a quick way to identify pest and alien species? Afr. Zool. 52(2), 105–117 (2017).
Sato-Okoshi, W., Abe, H., Nishitani, G. & Simon, C. A. And then there was one: Polydora uncinata and Polydora hoplura (Annelida: Spionidae), the problematic polydorid pest species represent a single species. J. Mar. Biol. Assoc. U. K. 97(8), 1675–1684 (2017).
Meißner, K. & Götting, M. Spionidae (Annelida:‘Polychaeta’: Canalipalpata) from Lizard Island, Great Barrier Reef, Australia: the genera Malacoceros, Scolelepis, Spio, Microspio, and Spiophanes. Zootaxa 4019(1), 378–413 (2015).
Bandelt, H. J., Forster, P. & Röhl, A. Median-joining networks for inferring intraspecific phylogenies. Mol. Biol. Evol. 16(1), 37–48 (1999).
Excoffier, L. & Lischer, H. E. Arlequin suite ver 3.5: a new series of programs to perform population genetics analyses under Linux and Windows. Mol. Ecol. Resour. 10(3), 564–567 (2010).
Sikorski, A. V., Jirkov, I. A. & Tsetlin, A. B. The genus Laonice (Polychaeta, Spionidae) in the Arctic Ocean: weighing the taxonomic characters and species composition. Zool Zhurnal 67, 826–838 (1988).
Hebert, P. D., Ratnasingham, S. & de Waard, J. R. Barcoding animal life: cytochrome c oxidase subunit 1 divergences among closely related species. Proc. Royal Soc. B 270, 96–99 (2003).
Ballard, J. W. O. & Whitlock, M. C. The incomplete natural history of mitochondria. Molec. Ecol. 13(4), 729–744 (2004).
Schwentner, M., Timms, B. V. & Richter, S. An integrative approach to species delineation incorporating different species concepts: a case study of Limnadopsis (Branchiopoda: Spinicaudata). Biol. J. Linn. Soc. 104, 575–599 (2011).
Schwentner, M., Just, F. & Richter, S. Evolutionary systematics of the Australian Cyzicidae (Crustacea, Branchiopoda, Spinicaudata) with the description of a new genus. Zool. J. Linn. Soc. 173, 271–295 (2014).
Kvist, S. Does a global DNA barcoding gap exist in Annelida? Mitochond. DNA Pt A 27(3), 2241–2252 (2016).
Janssen, A. et al. reverse taxonomic approach to assess macrofaunal distribution patterns in abyssal pacific polymetallic nodule fields. PLoS One 10, e0117790 (2015).
Mahon, A. R., Mahon, H. K., Dauer, D. M. & Halanych, K. M. Discrete genetic boundaries of three Streblospio (Spionidae, Annelida) species and the status of S. shrubsolii. Mar. Biol. Resear. 5, 172–178 (2009).
Meißner, K. & Blank, M. Spiophanes norrisi sp. nov. (Polychaeta: Spionidae) a new species from the NE Pacific coast, separated from the Spiophanes bombyx complex based on both morphological and genetic studies. Zootaxa 2278, 1–25 (2009).
Funk, D. J. & Omland, K. E. Species-level paraphyly and polyphyly: frequency, causes, and consequences, with insights from animal mitochondrial DNA. Annu. Rev. Ecol. Evol. Syst. 34(1), 397–423 (2003).
Maciolek, N. J. A new genus and species of Spionidae (Annelida: Polychaeta) from the North and South Atlantic. P. Biol. Soc. Wash. 94(1), 228–239 (1981).
Kongsrud, J. A. & Rapp, H. T. Nicomache (Loxochona) lokii sp. nov. (Annelida: Polychaeta: Maldanidae) from the Loki’s Castle vent field: an important structure builder in an Arctic vent system. Polar Biol. 35(2), 161–170 (2011).
Smirnov, R. V. Two new species of Pogonophora from the arctic mud volcano off northwestern Norway. Sarsia 85, 141–150 (2000).
Eilertsen, M. H. et al. Genetic connectivity from the Arctic to the Antarctic: Sclerolinum contortum and Nicomache lokii (Annelida) are both widespread in reducing environments. Sci. Rep. 8(1), 4810 (2018).
McClain, C. R. & Hardy, S. M. The dynamics of biogeographic ranges in the deep sea. Proc. Royal Soc. B 277(1700), 3533–3546 (2010).
Linse, K. & Schwabe, E. Diversity of macrofaunal Mollusca of the abyssal Vema Fracture Zone and hadal Puerto Rico Trench, Tropical North Atlantic. Deep-Sea Res. Pt II 148, 45–53 (2018).
Zardus, J. D., Etter, R. J., Chase, M. R., Rex, M. A. & Boyle, E. E. Bathymetric and geographic population structure in the pan-Atlantic deep-sea bivalve Deminucula atacellana (Schenck, 1939). Mol. Ecol. 15, 639–651 (2006).
Brix, S. et al Molecular species delimitation and its implications for species descriptions using desmosomatid and nannoniscid isopods from the VEMA fracture zone as example taxa. Deep-Sea Res. Pt II 148, 180–207 (2018).
Scheltema, R. S. The dispersal of the larvae of shoal-water benthic invertebrate species over long distances by ocean currents in Fourth European Marine Biology Symposium (ed. Crisps, D. J.) 7–28 (London: Cambridge University Press 1971).
Geller, J., Meyer, C., Parker, M. & Hawk, H. Redesign of PCR primers for mitochondrial cytochrome c oxidase subunit I for marine invertebrates and application in all-taxa biotic surveys. Mol. Ecol. Resour. 13(5), 851–861 (2013).
Folmer, O., Black, M., Hoeh, W., Lutz, R. & Vrijenhoek, R. DNA primers for amplification of mitochondrial cytochrome c oxidase subunit I from diverse metazoan invertebrates. Mol. Mar. Biol. Biotech. 3, 294–299 (1994).
Carpenter, J. M. & Wheeler, W. C. Towards simultaneous analysis of morphological and molecular data in Hymenoptera. Zool. Scr. 28, 251–260 (1999).
Palumbi, S. R. Nucleic acids II: the polymerase chain reaction in Molecular Systematics Hillis, (eds Moritz, C., Mable, B. K.) 205–247 (Massachusetts: Sinauer & Associates Inc., Sunderland 1996).
Palumbi, S. R. et al. The Simple Fool’s Guide to PCR, Version 2.0. Privately published, Univ. Hawaii (1991).
Dzikowski, R., Levy, M. G., Poore, M. F., Flowers, J. R. & Paperna, I. Use of rDNA polymorphism for identification of Heterophyidae infecting freshwater fishes. Dis. Aquat. Organ. 59(1), 35–41 (2004).
Acknowledgements
The first author’s work was supported by the Bauer Foundation (Germany). The Vema-TRANSIT Expedition was undertaken with financial support of the PTJ (German Ministry for Science and Education), Grant 03G0227A. Many thanks are owed to the crew of the RV Sonne for support during the Vema-TRANSIT Expedition. GRG are thanked for valuable comments and moral support. Special thanks to the two anonymous reviewers for helpful comments and new issues of consideration. The English has kindly been checked and corrected by Dr. Huw Griffiths, British Antarctic Survey.
Author information
Authors and Affiliations
Contributions
First draft of the manuscript and figures were prepared by Theresa Guggolz. Martin Schwentner and Karin Meißner were proof-reading the manuscript, giving advices for interpreting the data and helping with analyses of data. Angelika Brandt was the head of the expedition and initiated the Vema-Transit project. She was also proof-reading the manuscript. The revision of the manuscript was mainly implemented by Theresa Guggolz with support of the co-authors.
Corresponding author
Ethics declarations
Competing Interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Guggolz, T., Meißner, K., Schwentner, M. et al. Diversity and distribution of Laonice species (Annelida: Spionidae) in the tropical North Atlantic and Puerto Rico Trench. Sci Rep 9, 9260 (2019). https://doi.org/10.1038/s41598-019-45807-7
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-019-45807-7
- Springer Nature Limited
This article is cited by
-
Multi-ocean distribution of a brooding predator in the abyssal benthos
Scientific Reports (2023)
-
Abyssal fauna of polymetallic nodule exploration areas, eastern Clarion-Clipperton Zone, central Pacific Ocean: Annelida: Spionidae and Poecilochaetidae
Marine Biodiversity (2022)
-
High diversity and pan-oceanic distribution of deep-sea polychaetes: Prionospio and Aurospio (Annelida: Spionidae) in the Atlantic and Pacific Ocean
Organisms Diversity & Evolution (2020)
-
Disentangling invasions in the sea: molecular analysis of a global polychaete species complex (Annelida: Spionidae: Pseudopolydora paucibranchiata)
Biological Invasions (2020)