Abstract
The Streptococcus mutans Cid/Lrg system represents an ideal model to study how this organism withstands various stressors encountered in the oral cavity. Mutation of lrgAB renders S. mutans more sensitive to oxidative, heat, and vancomycin stresses. Here, we have performed a comprehensive proteomics experiment using label-free quantitative mass spectrometry to compare the proteome changes of wild type UA159 and lrgAB mutant strains in response to these same stresses. Importantly, many of identified proteins showed either a strikingly large fold-change, or were completely suppressed or newly induced in response to a particular stress condition. Notable stress proteome changes occurred in a variety of functional categories, including amino acid biosynthesis, energy metabolism, protein synthesis, transport/binding, and transcriptional/response regulators. In the non-stressed growth condition, mutation of lrgAB significantly altered the abundance of 76 proteins (a fold change >1.4, or <0.6, p-value <0.05) and several of these matched the stress proteome of the wild type strain. Interestingly, the statistical correlation between the proteome changes and corresponding RNA-seq transcriptomic studies was relatively low (rho(ρ) <0.16), suggesting that adaptation to a new environment may require radical proteome turnover or metabolic remodeling. Collectively, this study reinforces the importance of LrgAB to the S. mutans stress response.
Similar content being viewed by others
Introduction
The pathogenic potential of Streptococcus mutans, a major etiological agent of human dental caries1, is intimately linked to its ability to efficiently and rapidly adjust to the adverse environment of the oral cavity, thus improving its survival and persistence in dental plaque biofilm2,3. In this regard, considerable work has focused on the S. mutans Cid/Lrg system, consisting of two dicistronic operons lrgAB (SMU.575c/574c) and cidAB (SMU.1701c/1700c)4,5,6,7. Expression of cid and lrg is highly responsive to complex and unfavorable environmental signals that integrate into the regulatory networks modulating S. mutans virulence. For example, lrg and cid genes display opposite patterns of expression in response to growth phase, glucose concentration, oxygenation, and blood plasma6,8 and are transcriptionally cross-regulated5,6. Regulation of the lrg and cid operons is further complicated by involvement of multiple major regulators, including CcpA (carbon catabolite protein A) and the TCSs (two-component signal transduction systems) LytST and VicKR4,5,6. Consequently, the Cid/Lrg system affects comprehensive virulence traits such as autolysis, biofilm development, oxidative and heat stress response, antibiotic resistance and genetic competence4,5,6 required for successful colonization in the oral cavity. For these reasons, the Cid/Lrg system has potential as an attractive target for development of anti-caries treatments.
Another value in studying the Cid/Lrg system is that lrgA and cidA encode membrane proteins with predicted similarity to bacteriophage holin: antiholin proteins that modulate cell lysis. Holins are known to control the timing and onset of host cell lysis during bacteriophage lytic infection, and thus it has been hypothesized that Cid and Lrg may contribute to inducing death and/or lysis in a subpopulation of the bacterial community for the altruistic benefit of the entire population, as a survival strategy for adapting to environmental conditions9,10,11. Conclusive data proving that the Cid/Lrg systems of S. mutans and S. aureus are functionally analogous to holins/antiholin pairs is still lacking. However, published S. mutans data strongly suggest that Cid and Lrg proteins are functionally interrelated5,7. For example, we recently found that the lack of either CidB or LrgAB results in very similar phenotypes with respect to ultrasensitivity to aeration, heat, and vancomycin stress5,7. More interestingly, our recent RNA-seq data has also demonstrated almost identical transcriptome changes between cidB and lrgAB mutants, compared to wild type, underlining likely functional and mechanistic similarities between CidB and LrgAB5,7. However, molecular mechanisms for how these encoded proteins modulate S. mutans virulence and its stress response are still not well understood. As well, clues to their specific cellular functionality are relatively scarce due in part to the inherent difficulties associated with studying membrane proteins.
“-Omics” technologies have provided a powerful means of broadly and globally assessing the cellular responses and adaptation of bacteria to stresses. We recently performed a comprehensive RNA-seq experiment to assess the transcriptomic changes of wild-type and isogenic lrgAB mutants under anaerobic (control) and stress-inducing culture conditions (aerobic, heat and vancomycin stress), as a way to connect known stress-sensitive phenotypic aspects to information regarding specific changes in gene expression7. These environmental stresses and lrgAB mutation both influenced the transcriptome in S. mutans, with implications for bacterial cell death/lysis, adaptation and virulence. Nevertheless, understanding transcriptomic changes alone may be insufficient for defining the exact dynamic cellular changes of S. mutans in response to environmental stresses.
This present study investigates the changes in proteome profiles that occurred in the same set of S. mutans wild-type and lrgAB mutant samples used in our recently published RNA-seq study7. To this end, mass spectrometry-based label-free quantitative proteomics12,13,14,15, a technology known to enable comprehensive identification and quantification of complete bacterial proteomes, was adapted to study changes in S. mutans intracellular protein levels in response to aeration, heat, and vancomycin stress. Furthermore, the degree of correlation between S. mutans protein abundance profiles and gene expression changes was determined. Further understanding of these combined “-omics” data at the cellular or molecular level will enhance our knowledge of Cid/Lrg-mediated cellular responses of S. mutans to adverse environments. Furthermore, these “-omics” data will serve as a valuable resource that can be mined to help clarify the role of the S. mutans stress response and physiological activity to its dynamic survival in the oral cavity.
Materials and Methods
Bacterial strains and media
The bacterial strains used in this study were S. mutans UA159, a serotype c strain16, and a previously published isogenic lrgAB mutant6, which was created using the PCR ligation mutagenesis technique17. Both S. mutans strains were grown and maintained on Brain Heart Infusion (BHI) agar plates (containing 1,000 µg/ml kanamycin for the lrgAB mutant), at 37 °C and 5% CO2.
Bacterial cultures for protein preparation
The identical set of bacterial cultures analyzed in our recently-published RNA-seq work7 were also used in this present study. Briefly, wild type strain UA159 (designated “WT” in all subsequent figures and tables) and ΔlrgAB (designated “AB” in all subsequent figures and tables) overnight cultures were each diluted 1:50 in 50 ml sterile BHI broth and grown to mid-exponential growth phase (OD600 = 0.4) under four different environmental conditions (anaerobic, (control); aerobic, (a); heat stress, (h); and vancomycin, (v) stress), as follows: For anaerobic growth, including heat and vancomycin stress conditions, sterile mineral oil was placed on top of the cultures. For aerobic growth, each culture was grown in a 250-ml conical flask at a 1:5 volume to flask ratio, and incubated at 115 rpm and 37 °C. For heat stress growth, cultures were incubated at 40 °C. For vancomycin growth, cultures were supplemented with 1 µg/ml vancomycin and incubated at 37 °C. An aliquot (5 ml) of each culture was previously used for the RNA-seq experiment7, and the rest (45 ml) of the culture was subjected to the mass spectrometry experiments in this study. Growth curves of these strains under each condition can be found in our previous publication7.
Protein Extraction and Quantification
Biological pentaplicates (i.e. n = 5 replicates per S. mutans strain per growth condition, including the n = 3 replicates analyzed previously by RNA-seq.7) of each S. mutans culture were processed for protein extraction according to Fujiki et al.18 with the following modifications: Samples were ground into fine powder in liquid nitrogen and agitated in extraction buffer (0.1 M Tris-HCl pH 8.8, 10 mM EDTA, 0.2 M DTT, 0.9 M sucrose) with an equal volume of phenol (pH 8.0) for 2 hours at room temperature. The phenol phase was precipitated by adding five volumes of 0.1 M ammonium acetate in methanol. After washing twice with 0.1 M ammonium acetate in methanol and twice with 80% acetone, the dried pellet was dissolved with 50 mM ammonium bicarbonate buffer. The mixtures were incubated on ice before centrifugation at 4 °C for 20 minutes at 12,000 × g, separating the proteins into soluble and insoluble phases. The insoluble phase was incubated with protein buffer (8 M urea, 2 M thiourea, 2% ASB-14 (amidosulfobetaine 14)) for at least one hour with occasional vortexing. The proteins were quantified using an EZQ Protein Quantitation Kit (Invitrogen, Carlsbad, CA, USA) and SoftMax Pro Software v5.3 (Molecular Devices, Downingtown, PA, USA).
Protein Digestion and LC-MS/MS
Proteins were precipitated in 25 mM ammonium bicarbonate, pH 8.0 with Amicon Ultra-0.5 ml Centrifugal filters (EMD Millipore Inc., Billerica, MA, USA). For each sample, a total of 30 μg of protein was reduced with 40 mM DTT, alkylated with 100 mM of iodoacetamide, and trypsin-digested. Trypsin-digested peptides were then desalted with C18-solid phase extraction. A hybrid quadrupole Orbitrap (Q Exactive Plus) mass spectrometry (MS) system (Thermo Fisher Scientific, Bremen, Germany) was used with high energy collision dissociation (HCD) in each MS and MS/MS cycle. The instrument was run in data-dependent mode with a full MS (400–2000 m/z) resolution of 70 000 and five MS/MS experiments (HCD NCE = 28%, isolation width = 3 Th, first mass = 105 Th, 5% underfill ratio, peptide match set to “preferred”, and an AGC target of 1e6). Dynamic exclusion for 10 s was used to prevent repeated analysis of the same peptides, and a lock mass of m/z 445.12003 (polysiloxane ion) was used for real-time internal calibration. The MS system was interfaced with an automated Easy-nLC 1000 system (Thermo Fisher Scientific, Bremen, Germany). Each sample fraction was loaded onto an Acclaim Pepmap 100 precolumn (20 mm × 75 μm; 3 μm-C18) and separated on an Easy-Spray analytical column (500 mm × 75 μm; 2 μm-C18) at a flow rate at 300 nL/min during a linear gradient from solvent A (0.1% formic acid (v/v)) to 25% solvent B (0.1% formic acid (v/v), 99.9% acetonitrile (v/v)) for 280 min, followed by ramping up to 98% solvent B for an additional 20 min. Peptides were sprayed into the orifice of the mass spectrometer, which was operated in an information-dependent data acquisition mode.
Proteomics Data Analysis
Raw MS/MS data files were processed using a thorough database search, considering biological modification and amino acid substitution against a nonredundant Streptococcus mutans database with decoy sequences (4,064 entries) using Scaffold Q + S (Proteome Software Inc., Portland, OR, USA) and MASCOT 2.4 (Matrix Science Inc., Boston, MA, USA). The following parameters were used for all the searching: Peptide tolerance at 10 ppm, tandem MS tolerance at ± 0.01 Da, peptide charges of 2+ to 5+ , trypsin as the enzyme, allowing one missed cleavage, Carbamidomethyl (C) as fixed modifications, and oxidation (M) and phosphorylation (S, T, Y) as variable modifications. Peptide and protein were filtered using Scaffold Q + S with strict peptide and protein probabilities, 0.9 and 0.95, respectively. Peptide probability was applied to filter peptide assignments obtained from MS/MS database searching results using predictable false identification error rate. Protein probability was used to filter proteins with the null hypothesis that the database matching is random and consideration of the peptide probability for all the peptides apportioned to that protein. For protein quantification, only MS/MS spectra were normalized by spectral abundance factor (NSAF). To avoid analyzing abundance differences that were unlikely to be biologically relevant, proteins with extremely low abundance (average MS/MS spectral count <2) were excluded from further analysis. Differentially expressed proteins were identified by student t-test, and a protein should be quantified with at least three of the biological pentaplicate, and a fold change >1.4 or <0.6 with p-value < 0.05. Protein annotation/functional categories presented in all Tables and Supplemental Files were obtained using UniProt19, the Los Alamos S. mutans genome data base (http://www.oralgen.lanl.gov/) and/or the NCBI S. mutans UA159 reference genome (NC_004350.2). To maximize the number of identified proteins in this analysis, we extensively annotated the detected peptides using the combined outputs of multiple search engines at a stringent FDR.
Data availability statement
The mass spectrometry proteomics data have been deposited to the ProteomeXchange Consortium via the PRIDE20 partner repository with the dataset identifier PXD006735 and 10.6019/PXD006735.
Correlation analysis with RNA-seq transcriptomic data
To perform correlation analysis with our recently published transcriptomic data7, the Reads Per Kilobase of transcript per Million mapped reads (RPKM) were used for all RNA-seq data sets. The n = 3 proteomic replicates and triplicate RNA-seq replicates used in the correlation analysis were derived from the same n = 3 biological samples. The programming language R was used for all statistical analyses. First, to determine the correlation of the RNA-seq and proteomic datasets, Pearson (linear) correlation was calculated for each two matched samples from RNA-seq and proteomic data using all the genes shared between the two datasets. The expression values were log 2 transformed. The significantly correlated samples were called at thresholds of a false discovery rate (FDR) <0.1. Second, to determine the correlation of the genes between RNA-seq and proteomic datasets across all samples, the Spearman correlation between RNA-seq and proteomic datasets for each gene was calculated using all the samples. The significantly correlated genes were called at thresholds of FDR <0.05 and Spearman’s rho >0.5.
Results and Discussion
We have started to develop an understanding of how Cid and Lrg membrane proteins are related to the ability of S. mutans to efficiently and rapidly adjust to the ever-changing oral cavity environment, consequently contributing to many key virulence traits of S. mutans 5,6,7. Since stress responses are closely integrated into bacterial physiology, we also envision that studying the Cid/Lrg system may provide novel insights into the mechanisms that regulate S. mutans cell homeostasis. The present study provides a valuable opportunity to detect proteomic changes in the same biological set of S. mutans wild-type and lrgAB cultures as those used in our previous transcriptome study7, using a label-free mass spectrometry-based quantitative proteomics. The rationale behind this study is that modulating the degradation and/or regulation of proteins may allow the cell to more rapidly and specifically adjust to stress, compared to transcriptomic changes. This more rapid proteome-based response may function to conserve resources by favoring only those cellular processes required for each particular stress condition21.
Overview of changes in protein abundance
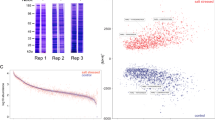
By using LC-MS/MS and a spectral counting based label-free approach, a total of 1049 proteins in the wild type UA159 were identified at a 95% confidence level, and 535 (51%) of proteins identified were differentially accumulated (a fold change >1.4, or <0.6, p-value < 0.05) in response to aeration, heat, or vancomycin stress, compared to the wild-type control samples grown anaerobically (Table 1). In the wild type, more downregulated proteins displayed larger magnitude fold changes as well as higher statistical significance compared to the upregulated proteins (Fig. 1a). In contrast, in the lrgAB strain, a total of 1039 proteins were identified at a 95% confidence level, and 474 (46%) of proteins identified were differentially accumulated (a fold change >1.4, or <0.6, p-value < 0.05) in response to the same conditions (Table 2). Similar to the wild type strain, more proteins were downregulated with larger magnitude fold changes and higher statistical significance (Fig. 1b). These data suggest that nearly half of the proteins identified by the label-free mass spectrometry differentially responded to the stressful conditions tested in both wild type and lrgAB mutant strains. Tables 1 and 2 also summarize the 23 possible protein abundance patterns of S. mutans wild type (A through W) and lrgAB mutant (I through XXIII) strains, respectively, in response to aeration (a), heat (h) and vancomycin (v) stress, relative to control (anaerobic), showing a holistic view for how the proteomes of the organisms shift in response to each stressor or between the stressors. To observe global patterns in the protein abundance data, principal component analysis (PCA) of the complete (both wild-type and lrgAB mutant) protein dataset was performed. As shown in Supplemental Fig. S1, the five biological replicates of each strain/growth condition grouped very closely together, except for the anaerobic samples of lrgAB. This analysis also suggested that the overall abundance trends in lrgAB samples obtained from the aerobic and vancomycin growth conditions were most similar to each other, while the heat lrgAB sample cluster was relatively separated from the other lrgAB samples. In the wild type strain, each stress sample was greatly separated from the control anaerobic sample cluster. Overall, these data indicate that S. mutans reorganizes its proteome composition upon exposure to different environmental stressors and possibly through the involvement of LrgAB.
Volcano plots showing differentially accumulated proteins in the wild type (a) and lrgAB (b) strain in response to all three environmental stressors (aeration, heat, and vancomycin stress), relative to non-stress (anaerobic) condition. Cutoff for significantly differential protein abundance, >1.4-fold change in protein abundance and p-value < 0.05. Red dots, proteins with increased abundance; green dots, proteins with decreased abundance; black dots, proteins without significant change in abundance. Vertical dotted line, protein abundance cutoff; horizontal dotted line, p-value cutoff. WT, wild type strain; AB, lrgAB mutant strain.
Stress proteomic changes in the wild type strain
The greatest number of proteins (n = 381) were differentially accumulated in response to vancomycin stress (Fig. 2a, top), and a considerable number of proteins also changed in abundance when the wild type responded to either aeration (n = 252) or heat (n = 234), reflecting that roughly 22–36% of the identified S. mutans proteome was abundantly reorganized to cope with these stressors. A high percentage of the proteins were largely associated with amino acid biosynthesis, energy metabolism, protein synthesis, and transport/binding (Supplemental Table S1), suggesting a substantial reprogramming of bacterial metabolism and physiology in coping with stress. Many proteins that increased or decreased in abundance also had no distinctive function, belonging to hypothetical, unassigned, or unknown categories.
Charts (top) and Venn diagrams (bottom) showing the number of proteins that exhibited differential abundance in the wild type (a) and lrgAB (b) strain in response to aeration (a), heat (h), and vancomycin (v) stress, relative to non-stress (anaerobic) condition. Cutoff for significantly differential protein abundance, > 1.4-fold change in protein abundance and p-value < 0.05. WTx, wild type strain in the stress (a, v, or v) condition; WT, wild type strain in the non-stress (anaerobic) condition; ABx, lrgAB mutant strain in the stress (a, v, or v) condition; AB, lrgAB mutant strain in the non-stress (anaerobic) condition.
To determine the degree of overlap in protein abundance among all WTx/WT (x = aerobic (a), heat (h) or vancomycin stress (v)), and to better understand the general and specific proteomic responses of the wild-type strain (WT) to each environmental stress, proteins that increased and decreased significantly (p < 0.05) in abundance in response to all three stressors were depicted as Venn diagrams (Fig. 2a, bottom). First, significant changes in 64 proteins were common to all three stressors (Fig. 2a, bottom). Among them, more proteins showed decreased (green) expression in response to aeration and vancomycin, while an equal number of proteins showed either an increase (red) or decrease (green) in response to heat (Supplemental Table S2). Notably, the increased amount of HtrA (high-temperature requirement A) in response to all three stress conditions is in line with the primary role of this enzyme in regulation of stress tolerance, mainly against elevated temperature and oxidative stress22,23,24, and virulence25,26. 25 proteins had opposite patterns of protein abundance between aeration/vancomycin and high temperature groups. Only one protein (SMU_1090; uncharacterized protein) was upregulated in response to both aeration and heat, but downregulated to vancomycin stress. Thus, unlike the response to aeration and vancomycin, response to heat may involve the contribution of additional or different physiological mechanisms in the wild type strain. The common 64 proteins were assigned to 16 different functional categories and alterations in protein abundance were enriched in the categories of amino acid biosynthesis (n = 10) and transport/binding proteins (n = 9) (Supplemental Table S2), suggesting that these functions are altered in S. mutans as an adaptive response to environmental stress.
We also identified protein signatures representing unique responses to each stress in the wild type (n = 45 for aeration; n = 91 for heat stress; n = 131 for vancomycin; Fig. 2a, bottom). Notable protein abundance changes unique to aeration include upregulated accumulation of putative chorismate mutase (SMU_531), cell division protein FtsX, and putative mannose specific EIID component (SMU_1957), which were newly produced only in response to aeration, but not without aeration (Supplemental Table S3). It is also noteworthy that SMU_1904c (uncharacterized protein), previously shown to be one of the most oxygen-sensitive genes7,27, was also more than 15-fold upregulated at the protein level, reinforcing the role of this gene/protein in coping with oxidative stress. Other highly upregulated proteins included GshR (glutathione reductase) and CopZ (putative copper chaperone), which were more than 8-fold and 4-fold upregulated, respectively, and have been previously reported to be important in protection of S. mutans against oxidative stress28,29. Particularly, CopZ has also been shown to be important in modulating a variety of virulence traits, including biofilm formation, stress tolerance, genetic competence, and competitiveness against commensal streptococci30. In contrast, the production of two putative ABC transporters (SMU_238c and SMU_370) and possible transmembrane efflux protein (SMU_1605) was completely suppressed in response to aeration (Supplemental Table S3). Interestingly, no downregulated proteins uniquely responding to aeration were differentially expressed at the transcriptional level, except for SMU_1340 (putative surfactin/bacitracin synthetase)7, which is part of TnSmu2, shown to enhance aerobic growth and tolerance to H2O2 challenge in S. mutans through nonribosomal peptide and polyketide (NRP/PK) biosynthesis31. Next, it is noteworthy that proteins uniquely responding to heat in the wild type, including multiple transcriptional (CpsY, Rex, NrdR, SMU_1398, SMU_677, and SMU_1585c) and global (CcpA and FruR) regulators, suggesting that response to heat stress may require a tighter coordination of both gene and protein expression and/or modification, probably to avoid unwanted cellular and physiological changes (Supplemental Table S4). Multiple ABC transporters, including OpuBa, SMU_922, SMU_1194, and SMU_1568 also showed significantly (p < 0.05) increased abundance. Major functional categories that were uniquely upregulated in response to heat stress included energy metabolism and PTSs, whereas DNA metabolism and protein synthesis-related proteins showed significantly decreased abundance, suggesting possible metabolic rewiring for production of extra energy (ATP) to cope with this stress32,33. It is also notable that among the heat-shock related molecular chaperones and proteases, only DnaK and Clp-like proteases (SMU_956 and SMU_2029) showed increased abundance uniquely in response to heat stress, suggesting that these proteins may be especially important for S. mutans cells living in environments with continuous heat. Interestingly, MazF, the predicted toxin component of the MazEF toxin/antitoxin (TA) module, was newly produced and uniquely responded to heat stress. This likely contributes to cell vialbility during heat stress, as overexpression of MazF presumably results in growth inhibition34. In contrast, the production of Cas9 (SMU_1405c), a CRISPR-associated endonuclease, was completely suppressed (Supplemental Table S4). Although how Cas proteins are involved in the response to heat stress is unclear, mutations in CRISPR1-cas and CRISPR2-cas in S. mutans were both previously shown to confer increased sensitivity to heat stress35. Overall, these data suggest that the response to heat stress in the S. mutans wild type strain may be regulated by a greater number of cellular processes than previously thought. Protein abundance changes unique to vancomycin stress occurred in multiple uncharacterized proteins and ABC transporters (Supplemental Table S5). In fact, tolerance to various peptide antibiotics, including vancomycin, has been reported to coevolve with ABC transporters and neighboring TCSs36. It is also notable that a great number of proteins involved in translation showed significantly decreased abundance. Given that vancomycin targets cell wall biogenesis and metabolism, it is not surprising that proteins involved in these processes were upregulated in response to this stress, including GtfC, WapA, WapE, AtlA (SMU_689), DacA and PtsH.
Finally, Fig. 2a (bottom), also showed that many proteome changes overlapped between two stressors. The largest overlapped proteins (n = 125) were observed between aeration and vancomycin proteome changes, while the smallest overlapped proteins (n = 18) were observed between aeration and heat proteome changes. Overall, it is notable that the proteome changes of wild-type were quite different from its transcriptome changes7 in response to the three stressors (Fig. 2a). In particular, S. mutans responded to vancomycin primarily through proteome readjustment, as very few changes were observed at the transcriptional level7. In contrast, the response of the organism to heat was more profound at the transcriptional level (n = 425)7, compared to its corresponding proteomic changes (n = 234). Aeration moderately influenced both the proteome and transcriptome of the organism. Therefore, these data suggest that proteome adjustment may provide an alternative way to respond to stress as a counterpoint to transcriptional modulation, and be intertwined with expression of virulence-related molecular elements.
Stress proteomic changes in the lrgAB mutant strain
Unlike the wild type strain, the proteome changes of the lrgAB mutant were most affected by heat (n = 335), and least by vancomycin stress (n = 147) (Fig. 2b, top). 177 proteins were also altered in abundance in the lrgAB mutant when cultured aerobically (Fig. 2b, top). Overall, stress responsive proteins corresponded to 14–32% of the identified lrgAB mutant proteome (n = 1039) (Table 2). Notably, except for the uncharacterized proteins (belonging to “hypothetical”, “unassigned” or “unknown” categories), many proteins associated with amino acid biosynthesis were consistently downregulated in response to all three stresses (Supplemental Table S6). Another major functional category with substantial changes in protein abundance was energy metabolism, which was largely upregulated in response to aeration, and downregulated to heat and vancomycin stress. However, the functional alteration profile was similar to that of the wild type in response to these stresses.
As shown in a Venn diagram displaying the degree of overlap in protein abundance among all ABx/AB (Fig. 2b; bottom), 39 proteins showed significant (p < 0.05) changes in common under all three stressors. Unlike the wild type, the lrgAB mutant showed relatively similar protein abundance patterns in response to all three stressors, with the exception of 7 proteins (Supplemental Table S7). Among the common 39 proteins, 11 proteins, including GbpA, SMU_218 (putative transcriptional regulator), Ddl (SMU_599), Obg (SMU_801), MsmK, SecA, SMU_1717c, Pnp (SMU_155), SMU_1416c, SerB, and HisD, were also contained in the overlapping set of proteins in the wild type (Supplemental Tables S7), which could be a potential core stress regulon common to all three stressors in S. mutans. Other notable upregulated proteins included Ftf, SMU_609 (putative 40 K cell wall protein), SMU_1287 (putative transcriptional regulator) and SMU_769. Particualrly, SMU_769 is one of the late competence genes regulated by ComX (or SigX), an alternative sigma factor of S. mutans.37,38. Other notable downregulated proteins included AtlA, SerB, DexA, amd SMU_1661c (putative signal peptidase). Interestingly, two putative cell wall hydrolases, SMU_609 and AtlA, responded to all three stresses in an opposite manner in the lrgAB strain (Supplemental Table S7), suggesting different role of the putative hydrolases in cell wall turnover and stress tolerance. The most obvious alterations in protein abundance belonged to amino acid biosynthesis (n = 6) and energy metabolism (n = 6).
In the lrgAB mutant, the greatest number of proteins (n = 182) uniquely responded to heat (Fig. 2b; bottom), and the majority of the proteins (n = 145) showed a significant decrease in abundance (Supplemental Table S8). Particularly, production of 25 proteins unique to this stress was completely suppressed in the lrgAB strain, and over 40 proteins were more than 5-fold downregulated, suggesting that adaptation to heat may require radical proteome turnover. Notable downregulated proteins included CovX, RelR, ScrK, GbpD, Pbp2b, RecR, RecX, SloR, and ComR (Supplemental Table S8). Given that ComR is a proximal regulator of ComX (SigX)39, decreased genetic competence of the lrgA strain may be related to altered ComR levels4. Proteins with increased abundance showed relatively lower fold-changes and included PtsH, SpxA1, FruI, GbpC, MazE, WapA, CiaR, DnaJ and GrpE. In the lrgAB mutant strain, interestingly, abundance of MazE, the putative antitoxin protein of the MAzEF T/A module, was increased, whereas MazF showed increased abundance in the wild type in response to heat (Supplemental Table S5). This suggests that MazEF, a proposed mediator of bacterial programmed cell death40,41, is closely linked to the heat stress response, as well as LrgAB. Another notable protein is SpxA1, which was previously shown to play an important role in stress tolerance, survival, and virulence in S. mutans with SpxA242,43. However, interestingly, SpxA2 did not significantly respond to any stress in both wild type and lrgAB-deficient strains, supporting a more dominant role of SpxA1 than SpxA243. Similar numbers of proteins uniquely responded to aeration (n = 57) and vancomycin stress (n = 55), which are shown in Supplemental Tables S9 and S10, respectively. Seven uncharacterized proteins were newly produced or >10-fold upregulated uniquely in response to aeration, while the downregulated proteins showed a relatively smaller fold-change (Supplemental Table S9). It is interesting that the production of SMU_571, located downstresm of lrgAB, was completely suppressed in the lrgAB strain uniquely in response to aeration and forms a three gene-operon with two putative ferrous iron transport proteins (FeoA/B, SMU_569/570). Given that iron plays a role in protection against oxidative stress44,45, SMU_571 may contribute to enhanced sensitivity of lrgAB to aeration, possiblly as a signaling peptide associated with FeoA/B transporters. It is also noteworthy that MvaK, a putative mevalonate kinase, was newly produced only when the lrgAB mutant responded to aeration (Supplemental Table S9), suggesting a possible functional linkage of LrgAB with the mevalonate pathway, which is involved in a variety of vital biological functions in bacteria, including cell wall biosynthesis46,47. In response to vancomycin stress, the fold-change of upregulated proteins (largely ribosomal proteins) in the lrgAB mutant was moderate, while downregulated proteins showed greater fold-change or were not accumulated (Supplemental Table S10). Notable proteins with decreased abundance include RcrR (SMU_921), PknB, GtfB, and MazF, which are related to virulence traits such as genetic competence, surface attachment, and stress response of S. mutans 34,48,49,50,51,52. Overall, these results suggest that the absence of LrgAB elicits extensive rewiring of the S. mutans proteome, particularly related to stress responses and virulence traits.
Comparison of UA159 and ΔlrgAB proteome changes in response to each stressor
To better understand how LrgAB is involved in coping with each stress, we compared the proteome changes of wild type (WTx/WT) and lrgAB (ABx/AB) in response to each stressor with a focus on the lrgAB-specific proteome changes, using Venn diagrams (Fig. 3). In response to aeration, 88 proteins demonstrated substantial changes in common between the WTa/WT and ABa/AB proteomes (Fig. 3a; Supplemental Fig. S2; Supplemental Table S11), representing the S. mutans general response to aerobic growth. One interesting finding is that two uncharacterized proteins (SMU_1209c and SMU_2152c) showed an opposite abundance pattern between the wild type and lrgAB mutant in response to aeration. 89 proteins responded to aeration only in the lrgAB strain, one or more of which may be responsible for the growth defect of the strain under the aerobic condition (Fig. 3a, Supplemental Fig. S2, Supplemental Table S12). 122 proteins showed significant (p < 0.05) abundance changes common to both WTh/WT and ABh/AB proteome comparisons (Fig. 3b, Supplemental Fig. S3, Supplemental Table S13). 20 proteins showed an opposite abundance pattern between wild type and lrgAB mutant in response to heat stress, and all of them were upregulated in the wild type and downregulated in the lrgAB strain. Among them, notable proteins showing increased abundance by more than 3-fold in the wild type included CpsY, TrpB, MurD, PyrK, SMU_1194 (putative ABC transporter), and SMU_521 (uncharacterized protein). Particularly, CpsY, a conserved transcriptional regulator, is known to activate transcription of genes involved in methionine biosynthesis and uptake in S. mutans 53 and S. agalactiae 54. CpsY has been also characterized as as virulence determinant that regulates amino acid metabolism54, methionine transport55, and cell wall modifications necessary for systemic infection in S. iniae 56,57,58. Therefore, CpsY may play a role in the Cid/Lrg system that relates the environmental/stress status of S. mutans to the control of growth and expression of virulence traits. A great number of proteins (n = 213) differentially responded to heat in the lrgAB mutant only (Fig. 3b, Supplemental Fig. S3, Supplemental Table S14). More proteins (n = 165) showed a significantly (p < 0.05) decreased abundance, and among them 82 proteins showed a fold change by more than 5-fold or were not accumulated at all. SloR, a DtxR family metalloregulator, known to modulate S. mutans metal ion homeostasis, biofilm formation, oxidative stress tolerance, and antibiotic gene regulation59,60,61,62,63,64, was uniquely suppressed in the lrgAB mutant in response to heat stress. SloR was also recently shown to crosstalk with VicKR and GcrR, major regulatory systems controlling critical S. mutans cellular processes and virulence65,66,67,68,69, through manganese availability60,70. Since VicKR was shown to regulate expression of lrgAB 5, SloR may be one of the intimate regulators collaborating with LrgAB. Lastly, 84 proteins demonstrated significant (p < 0.05) changes common to both WTv/WT and ABv/AB proteomes (Fig. 3a, Supplemental Fig. S4, Supplemental Table S15). More proteins were downregulated (n = 64) than upregulated (n = 20), and among them twelve proteins showed an opposite abundance pattern between wild type and lrgAB mutant in response to vancomycin stress. Interestingly, one uncharacterized protein (SMU_843) was newly induced in the wild type, and not accumulated in the lrgAB strain. This protein is predicted to play a role in cell wall/envelope biosynthesis as a putative poly-gamma-glutamate biosynthesis protein. In response to vancomycin stress, less proteome abundance changes were uniquely elicited in the lrgAB strain (n = 63) than wild-type (n = 297) (Fig. 3c, Supplemental Fig. S4, Supplemental Table S16). One interesting finding is that accumulation of GtfB, a key adhesin of S. mutans, was completely inhibited in the lrgAB mutant in response to vancomycin stress, supporting the potential for LrgAB as an anti-biofilm target.
Venn diagrams showing the comparison of proteome changes of wild type (WTx/WT) and lrgAB (ABx/AB) in response to aeration (a), heat (b), and vancomycin (c) stress. Cutoff for significantly differential protein abundance, >1.4-fold change in protein abundance and p-value < 0.05. WTx, wild type strain in the stress (a, v, or v) condition; WT, wild type strain in the non-stress (anaerobic) condition; ABx, lrgAB mutant strain in the stress (a, v, or v) condition; AB, lrgAB mutant strain in the non-stress (anaerobic) condition. The numbers in parenthesis indicate (up/down)-regulated proteins.
Proteome changes between wild type and lrgAB strains during non-stressed growth
When we directly compared the proteome of wild-type and lrgAB mutant in an optimal growth condition (anaerobic; at 37 °C), loss of LrgAB significantly (p < 0.05) altered the expression of 76 proteins (a fold change >1.4, or <0.6, p-value < 0.05) with 17 downregulated and 59 upregulated (Supplemental Fig. S5a; Table 3). While the previously-published corresponding RNA-seq analysis showed that most of the 54 differentially-expressed genes belonged to very few distinct functional groups (genomic islands (GIs) TnSmu1 and TnSmu2, the CRISPR (clustered regularly interspaced short palindromic repeats)-Cas system, bacteriocin production, energy metabolism, and amino acid ABC transporters)7, altered proteins belonged to many more diverse functional categories, including cell envelope, cellular processes, regulatory functions, and signal transduction (Supplemental Fig. S5b; Table 3). Nevertheless, SMU_209 (down-regulated) and SMU_1348c (up-regulated), overlapped between the transcriptome and proteome profiles, belonged to TnSmu1 and TnSmu2, respectively, which were two major functional groups with most differentially expression genes in the transcriptome profile, reinforcing the involvement of these genomic islands with the Cid/Lrg system. Other notable upregulated proteins included two putative transcriptional regulators (CpsY and SMU_1977c), two fatty acid biosynthesis-related proteins (AccC and FabZ), MazF, NaoX (H2O-forming NADH oxidase), GtfC, and GlgC, which were newly induced or had a fold-change by more than 4-fold. These data suggest that mutation of lrgAB may impose inherent stress on S. mutans cells, as well as shows potential functional partners of LrgAB and related molecular processes.
Correlation with RNA-seq transcriptomic profiles
We attempted to determine the relationship between the proteomic profiles and the gene expression profiles, recently obtained by high-throughput RNA-seq technology on corresponding aliquots of the same cultures used for this proteome experimentation7. As the transcriptome datasets contain three biological replicates per condition, only the three matched samples from each condition were used. In other words, correlation analysis was performed on a total of 24 samples from all conditions for RNA-seq and proteomic datasets, respectively. First, Pearson correlation was performed for the 24 matched samples to assess sample level correlation using all the shared genes/proteins for the matched RNA-seq and proteomic datasets. All of the 24 samples had a relative low positive correlation coefficient (rho(ρ) <0.16) between the RNA-seq and proteome datasets (Supplemental Table S17). Although this correlation was not high, the correlation coefficients for the 15 datasets representing AB, ABa, ABv, WT and WTh samples (in triplicates) are statistically significant (p < 0.05) with q-values (multiple testing corrected p-value; FDR) less than 10% (Supplemental Table S17). Therefore, the proteome changes that occur in the lrgAB mutant in response to heat stress, and in the wild-type in response to aeration and vancomycin stress, may be potentially elicited by non-transcriptional mechanisms such as mRNA decay, translation, and protein degradation and modification. In order to find the specific genes/proteins that contributed to the correlation between the transcriptome and proteome datasets, we also performed Spearman (monotonic) correlation analysis across all samples. At thresholds of q-value 0.05 and rho >0.5, a total of 46 genes (~6%) out of 776 tested genes showed significant (p < 0.05) correlation. Of these, 42 represent significantly differentially-expressed proteins (Supplemental Table S18). We also counted the number of differentially-expressed genes that overlapped between the RNA-seq and proteome datasets, and found that only a small proportion of these genes ( < 9%) were overlapped (Supplemental Table S19). The small proportion of correlated genes is consistent with the low sample level correlation results described above. Possibly, the low correlaton between transcriptomic and proteomic changes in this study may be due to continuous growth of the bacteria in the presence of each stress; this may have facilitated degradation or modification of damaged or unfolded proteins, to avoid deleterious consequences such as cell death or growth inhibition71,72. Another factor that may have lowered our ability to detect a higher correlation is that the transcriptome and proteome datasets were substantially different in size. Our comparison between transcriptome and proteome profiles was limited to the genes for which corresponding proteins were identified by mass spectrometry. The size difference was even bigger per stress condition7. Therefore, gaps in our proteomic dataset could have led to a comparison of a biased selection of only highly abundant proteins with unbiased expression data. A third factor may be the limited number of samples used for the correlation analysis. We did correlation analysis for each gene/protein using the three biological samples within each condition. In this case, perfect correlation were identified (rho = 1) in 707 genes/proteins but none is statistically significant (P value > 0.3). In contrast, when used the 24 biological samples across all the conditions, we identified 46 significantly (p < 0.05) correlated genes/proteins. Therefore, it is possible that with more biological samples, more significantly correlated genes can be potentially identified. Finally, from a technical standpoint, the approach using mass spectrometry would not completely cover the identification of rather hydrophobic proteins, although several cell membrane and surface proteins were identified here. Also, the current experimental approach did not not allow the quantitative analysis of secreted proteins, consequently generating a less representative proteome than its corresponsing transcriptome.
In summary, to our knowledge, this is the first comparative study on the intracellular proteome of S. mutans under multiple stress conditions. The data suggest that S. mutans adjusts its stress response through substantial and dynamic remodeling of the cellular proteome, and LrgAB has an important influence on this reorganization. This study also provided a valuable opportunity to compare transcriptomic and proteomic data generated on corresponding aliquots of the same cultures. It is possible that the overall low correlation between two ‘-omics’ datasets may be due to potential analytical and technical limitations, as well as unknown cellular mechanisms in the presence of rapid or continuous stress shock. These two integrative omics studies have provided a large number of transcripts and proteins that could be used in targeted studies on the molecular mechanisms triggering the persistence phenotype of S. mutans against unfavorable stresses. The data may also give an integrative view of a specific physiological state in coping with an external stress. More detailed biological interpretation of the altered proteins will be an interesting future direction. Given that within biofilms, oral bacteria become more resistant to adverse environments, including antibiotic challenge, compared to planktonic growth, it would be interesting to evaluate the proteomic changes elicited by these environmental stresses on S. mutans biofilms. This approach could provide more ecological insights into how this bacterium accommodates the heterogeneous microenvironments that develop in biofilm, as well as accelerate phenotypic and genotypic diversity within the population. Furthermore, studying proteomic shifts in multispecies oral biofilms by S. mutans in response to other types of environmental challenges, such as nutrient limitation and toxic metabolites, could be another interesting follow-up. Consequently, all these data will improve our integrated models and understanding of how bacterial cells adjust to a new challenging environment, an initial step in initiation of disease.
References
Loesche, W. J. Role of Streptococcus mutans in human dental decay. Microbiol Rev 50, 353–380 (1986).
Lemos, J. A. & Burne, R. A. A model of efficiency: stress tolerance by Streptococcus mutans. Microbiology 154, 3247–3255, https://doi.org/10.1099/mic.0.2008/023770-0 (2008).
Smith, E. G. & Spatafora, G. A. Gene regulation in S. mutans: complex control in a complex environment. J Den Res 91, 133–141, https://doi.org/10.1177/0022034511415415 (2012).
Ahn, S. J., Qu, M. D., Roberts, E., Burne, R. A. & Rice, K. C. Identification of the Streptococcus mutans LytST two-component regulon reveals its contribution to oxidative stress tolerance. BMC Microbiol 12, 187, https://doi.org/10.1186/1471-2180-12-187 (2012).
Ahn, S. J. & Rice, K. C. Understanding the Streptococcus mutans Cid/Lrg System through CidB Function. Appl Environ Microbiol 82, 6189–6203, https://doi.org/10.1128/AEM.01499-16 (2016).
Ahn, S. J., Rice, K. C., Oleas, J., Bayles, K. W. & Burne, R. A. The Streptococcus mutans Cid and Lrg systems modulate virulence traits in response to multiple environmental signals. Microbiology 156, 3136–3147, https://doi.org/10.1099/mic.0.039586-0 (2010).
Rice, K. C., Turner, M. E., Carney, O. V., Gu, T. & Ahn, S. J. Modification of the Streptococcus mutans transcriptome by LrgAB and environmental stressors. Microb Genom 3, e000104, https://doi.org/10.1099/mgen.0.000104 (2017).
Jung, C. J., Zheng, Q. H., Shieh, Y. H., Lin, C. S. & Chia, J. S. Streptococcus mutans autolysin AtlA is a fibronectin-binding protein and contributes to bacterial survival in the bloodstream and virulence for infective endocarditis. Mol Microbiol 74, 888–902, https://doi.org/10.1111/j.1365-2958.2009.06903.x (2009).
Bayles, K. W. Are the molecular strategies that control apoptosis conserved in bacteria? Trends Microbiol 11, 306–311 (2003).
Bayles, K. W. The biological role of death and lysis in biofilm development. Nature reviews. Microbiology 5, 721–726, https://doi.org/10.1038/nrmicro1743 (2007).
Bayles, K. W. Bacterial programmed cell death: making sense of a paradox. Nature reviews. Microbiology 12, 63–69, https://doi.org/10.1038/nrmicro3136 (2014).
Otto, A., Becher, D. & Schmidt, F. Quantitative proteomics in the field of microbiology. Proteomics 14, 547–565, https://doi.org/10.1002/pmic.201300403 (2014).
Van Oudenhove, L. & Devreese, B. A review on recent developments in mass spectrometry instrumentation and quantitative tools advancing bacterial proteomics. Appl Microbiol Biotechnol 97, 4749–4762, https://doi.org/10.1007/s00253-013-4897-7 (2013).
Bantscheff, M., Schirle, M., Sweetman, G., Rick, J. & Kuster, B. Quantitative mass spectrometry in proteomics: a critical review. Anal Bioanal Chem 389, 1017–1031, https://doi.org/10.1007/s00216-007-1486-6 (2007).
Schulze, W. X. & Usadel, B. Quantitation in mass-spectrometry-based proteomics. Annu Rev Plant Biol 61, 491–516, https://doi.org/10.1146/annurev-arplant-042809-112132 (2010).
Caufield, P. W. & Shah, G. Transformation of naturally competent Streptococcus mutans with replicative and non-replicative Tn916-containing plasmids: implications for a mechanism of transposition. Dev Biol Stand 85, 19–25 (1995).
Lau, P. C., Sung, C. K., Lee, J. H., Morrison, D. A. & Cvitkovitch, D. G. PCR ligation mutagenesis in transformable streptococci: application and efficiency. J Microbiol Methods 49, 193–205 (2002).
Fujiki, Y., Hubbard, A. L., Fowler, S. & Lazarow, P. B. Isolation of intracellular membranes by means of sodium carbonate treatment: application to endoplasmic reticulum. J Cell Biol 93, 97–102 (1982).
UniProt, C. UniProt: a hub for protein information. Nucleic Acids Res 43, D204–212, https://doi.org/10.1093/nar/gku989 (2015).
Vizcaino, J. A. et al. 2016 update of the PRIDE database and its related tools. Nucleic Acids Res 44, D447–456, https://doi.org/10.1093/nar/gkv1145 (2016).
Scott, M., Gunderson, C. W., Mateescu, E. M., Zhang, Z. & Hwa, T. Interdependence of cell growth and gene expression: origins and consequences. Science 330, 1099–1102, https://doi.org/10.1126/science.1192588 (2010).
Pallen, M. J. & Wren, B. W. The HtrA family of serine proteases. Mol Microbiol 26, 209–221 (1997).
Schlieker, C., Mogk, A. & Bukau, B. A. PDZ switch for a cellular stress response. Cell 117, 417–419 (2004).
Clausen, T., Southan, C. & Ehrmann, M. The HtrA family of proteases: implications for protein composition and cell fate. Mol Cell 10, 443–455 (2002).
Frees, D., Brondsted, L. & Ingmer, H. Bacterial proteases and virulence. Subcell Biochem 66, 161–192, https://doi.org/10.1007/978-94-007-5940-4_7 (2013).
Ingmer, H. & Brondsted, L. Proteases in bacterial pathogenesis. Res Microbiol 160, 704–710, https://doi.org/10.1016/j.resmic.2009.08.017 (2009).
Ahn, S. J., Wen, Z. T. & Burne, R. A. Effects of oxygen on virulence traits of Streptococcus mutans. J Bacteriol 189, 8519–8527, https://doi.org/10.1128/jb.01180-07 (2007).
Singh, K., Senadheera, D. B., Levesque, C. M. & Cvitkovitch, D. G. The copYAZ Operon Functions in Copper Efflux, Biofilm Formation, Genetic Transformation, and Stress Tolerance in Streptococcus mutans. J Bacteriol 197, 2545–2557, https://doi.org/10.1128/JB.02433-14 (2015).
Yamamoto, Y., Kamio, Y. & Higuchi, M. Cloning, nucleotide sequence, and disruption of Streptococcus mutans glutathione reductase gene (gor). Biosci Biotechnol Biochem 63, 1056–1062, https://doi.org/10.1271/bbb.63.1056 (1999).
Garcia, S. S., Du, Q. & Wu, H. Streptococcus mutans copper chaperone, CopZ, is critical for biofilm formation and competitiveness. Mol Oral Microbiol 31, 515–525, https://doi.org/10.1111/omi.12150 (2016).
Wu, C. et al. Genomic island TnSmu2 of Streptococcus mutans harbors a nonribosomal peptide synthetase-polyketide synthase gene cluster responsible for the biosynthesis of pigments involved in oxygen and H2O2 tolerance. Appl Environ Microbiol 76, 5815–5826, https://doi.org/10.1128/aem.03079-09 (2010).
Liu, C. et al. Streptococcus mutans copes with heat stress by multiple transcriptional regulons modulating virulence and energy metabolism. Sci Rep 5, 12929, https://doi.org/10.1038/srep12929 (2015).
Fleury, B. et al. Transcriptomic and metabolic responses of Staphylococcus aureus exposed to supra-physiological temperatures. BMC Microbiol 9, 76, https://doi.org/10.1186/1471-2180-9-76 (2009).
Syed, M. A. et al. The chromosomal mazEF locus of Streptococcus mutans encodes a functional type II toxin-antitoxin addiction system. J Bacteriol 193, 1122–1130, https://doi.org/10.1128/JB.01114-10 (2011).
Serbanescu, M. A. et al. Role of the Streptococcus mutans CRISPR-Cas systems in immunity and cell physiology. J Bacteriol 197, 749–761, https://doi.org/10.1128/JB.02333-14 (2015).
Dintner, S. et al. Coevolution of ABC transporters and two-component regulatory systems as resistance modules against antimicrobial peptides in Firmicutes Bacteria. J Bacteriol 193, 3851–3862, https://doi.org/10.1128/JB.05175-11 (2011).
Dufour, D., Cordova, M., Cvitkovitch, D. G. & Levesque, C. M. Regulation of the competence pathway as a novel role associated with a streptococcal bacteriocin. J Bacteriol 193, 6552–6559, https://doi.org/10.1128/JB.05968-11 (2011).
Eaton, R. E. & Jacques, N. A. Deletion of competence-induced genes over-expressed in biofilms caused transformation deficiencies in Streptococcus mutans. Mol Oral Microbiol 25, 406–417, https://doi.org/10.1111/j.2041-1014.2010.00589.x (2010).
Mashburn-Warren, L., Morrison, D. A. & Federle, M. J. A novel double-tryptophan peptide pheromone controls competence in Streptococcus spp. via an Rgg regulator. Mol Microbiol 78, 589–606, https://doi.org/10.1111/j.1365-2958.2010.07361.x (2010).
Erental, A., Sharon, I. & Engelberg-Kulka, H. Two programmed cell death systems in Escherichia coli: an apoptotic-like death is inhibited by the mazEF-mediated death pathway. PLoS Biol 10, e1001281, https://doi.org/10.1371/journal.pbio.1001281 (2012).
Ramisetty, B. C., Natarajan, B. & Santhosh, R. S. mazEF-mediated programmed cell death in bacteria: “what is this?”. Crit Rev Microbiol 41, 89–100, https://doi.org/10.3109/1040841X.2013.804030 (2015).
Kajfasz, J. K. et al. Two Spx proteins modulate stress tolerance, survival, and virulence in Streptococcus mutans. J Bacteriol 192, 2546–2556, https://doi.org/10.1128/JB.00028-10 (2010).
Kajfasz, J. K. et al. Transcription of Oxidative Stress Genes Is Directly Activated by SpxA1 and, to a Lesser Extent, by SpxA2 in Streptococcus mutans. J Bacteriol 197, 2160–2170, https://doi.org/10.1128/jb.00118-15 (2015).
Andrews, S. C., Robinson, A. K. & Rodriguez-Quinones, F. Bacterial iron homeostasis. FEMS Microbiol Rev 27, 215–237 (2003).
McHugh, J. P. et al. Global iron-dependent gene regulation in Escherichia coli. A new mechanism for iron homeostasis. J Biol Chem 278, 29478–29486, https://doi.org/10.1074/jbc.M303381200 (2003).
Perez-Gil, J. & Rodriguez-Concepcion, M. Metabolic plasticity for isoprenoid biosynthesis in bacteria. Biochem J 452, 19–25, https://doi.org/10.1042/BJ20121899 (2013).
Heuston, S., Begley, M., Gahan, C. G. & Hill, C. Isoprenoid biosynthesis in bacterial pathogens. Microbiology 158, 1389–1401, https://doi.org/10.1099/mic.0.051599-0 (2012).
Banu, L. D. et al. The Streptococcus mutans serine/threonine kinase, PknB, regulates competence development, bacteriocin production, and cell wall metabolism. Infect Immun 78, 2209–2220, https://doi.org/10.1128/IAI.01167-09 (2010).
Kaspar, J. et al. A unique open reading frame within the comX gene of Streptococcus mutans regulates genetic competence and oxidative stress tolerance. Mol Microbiol 96, 463–482, https://doi.org/10.1111/mmi.12948 (2015).
Kaspar, J., Kim, J. N., Ahn, S. J. & Burne, R. A. An Essential Role for (p)ppGpp in the Integration of Stress Tolerance, Peptide Signaling, and Competence Development in Streptococcus mutans. Front Microbiol 7, 1162, https://doi.org/10.3389/fmicb.2016.01162 (2016).
Reck, M. et al. The biofilm inhibitor carolacton disturbs membrane integrity and cell division of Streptococcus mutans through the serine/threonine protein kinase PknB. J Bacteriol 193, 5692–5706, https://doi.org/10.1128/JB.05424-11 (2011).
Bowen, W. H. & Koo, H. Biology of Streptococcus mutans-derived glucosyltransferases: role in extracellular matrix formation of cariogenic biofilms. Caries Res 45, 69–86, https://doi.org/10.1159/000324598 (2011).
Sperandio, B. et al. Control of methionine synthesis and uptake by MetR and homocysteine in Streptococcus mutans. J Bacteriol 189, 7032–7044, https://doi.org/10.1128/JB.00703-07 (2007).
Bryan, J. D., Liles, R., Cvek, U., Trutschl, M. & Shelver, D. Global transcriptional profiling reveals Streptococcus agalactiae genes controlled by the MtaR transcription factor. BMC Genomics 9, 607, https://doi.org/10.1186/1471-2164-9-607 (2008).
Shelver, D., Rajagopal, L., Harris, T. O. & Rubens, C. E. MtaR, a regulator of methionine transport, is critical for survival of group B streptococcus in vivo. J Bacteriol 185, 6592–6599 (2003).
Allen, J. P. & Neely, M. N. The Streptococcus iniae transcriptional regulator CpsY is required for protection from neutrophil-mediated killing and proper growth in vitro. Infect Immun 79, 4638–4648, https://doi.org/10.1128/IAI.05567-11 (2011).
Allen, J. P. & Neely, M. N. CpsY influences Streptococcus iniae cell wall adaptations important for neutrophil intracellular survival. Infect Immun 80, 1707–1715, https://doi.org/10.1128/IAI.00027-12 (2012).
Koskiniemi, S., Sellin, M. & Norgren, M. Identification of two genes, cpsX and cpsY, with putative regulatory function on capsule expression in group B streptococci. FEMS Immunol Med Microbiol 21, 159–168 (1998).
Crepps, S. C. et al. The SloR metalloregulator is involved in the Streptococcus mutans oxidative stress response. Mol Oral Microbiol 31, 526–539, https://doi.org/10.1111/omi.12147 (2016).
Downey, J. S. et al. In vitro manganese-dependent cross-talk between Streptococcus mutans VicK and GcrR: implications for overlapping stress response pathways. PLoS One 9, e115975, https://doi.org/10.1371/journal.pone.0115975 (2014).
Merchant, A. T. & Spatafora, G. A. A role for the DtxR family of metalloregulators in gram-positive pathogenesis. Mol Oral Microbiol 29, 1–10, https://doi.org/10.1111/omi.12039 (2014).
Paik, S., Brown, A., Munro, C. L. & Cornelissen, C. N. & Kitten, T. The sloABCR operon of Streptococcus mutans encodes an Mn and Fe transport system required for endocarditis virulence and its Mn-dependent repressor. J Bacteriol 185, 5967–5975 (2003).
Rolerson, E. et al. The SloR/Dlg metalloregulator modulates Streptococcus mutans virulence gene expression. J Bacteriol 188, 5033–5044, https://doi.org/10.1128/JB.00155-06 (2006).
O’Rourke, K. P. et al. Genome-wide characterization of the SloR metalloregulome in Streptococcus mutans. J Bacteriol 192, 1433–1443, https://doi.org/10.1128/JB.01161-09 (2010).
Idone, V. et al. Effect of an orphan response regulator on Streptococcus mutans sucrose-dependent adherence and cariogenesis. Infect Immun 71, 4351–4360 (2003).
Sato, Y., Yamamoto, Y. & Kizaki, H. Construction of region-specific partial duplication mutants (merodiploid mutants) to identify the regulatory gene for the glucan-binding protein C gene in vivo in Streptococcus mutans. FEMS Microbiol Lett 186, 187–191 (2000).
Senadheera, D. B. et al. Regulation of bacteriocin production and cell death by the VicRK signaling system in Streptococcus mutans. J Bacteriol 194, 1307–1316, https://doi.org/10.1128/JB.06071-11 (2012).
Senadheera, M. D. et al. A VicRK signal transduction system in Streptococcus mutans affects gtfBCD, gbpB, and ftf expression, biofilm formation, and genetic competence development. J Bacteriol 187, 4064–4076, https://doi.org/10.1128/JB.187.12.4064-4076.2005 (2005).
Ahn, S. J. & Burne, R. A. Effects of oxygen on biofilm formation and the AtlA autolysin of Streptococcus mutans. J Bacteriol 189, 6293–6302, https://doi.org/10.1128/JB.00546-07 (2007).
Dunning, D. W. et al. SloR modulation of the Streptococcus mutans acid tolerance response involves the GcrR response regulator as an essential intermediary. Microbiology 154, 1132–1143, https://doi.org/10.1099/mic.0.2007/012492-0 (2008).
Gur, E., Biran, D. & Ron, E. Z. Regulated proteolysis in Gram-negative bacteria–how and when? Nat Rev Microbiol 9, 839–848, https://doi.org/10.1038/nrmicro2669 (2011).
Mogk, A., Huber, D. & Bukau, B. Integrating protein homeostasis strategies in prokaryotes. Cold Spring Harb Perspect Biol 3, https://doi.org/10.1101/cshperspect.a004366 (2011).
Acknowledgements
This work was supported by the National Institute fro Dental and Craniofacial Research grant R01 DE025237 (SJA). Label-free quantitative mass spectrometry experiments were performed in the Proteomics Core of ICBR (Interdisciplinary Center for Biotechnology Research) at University of Florida. We thank Prof. Robert A. Burne for providing all the resources needed.
Author information
Authors and Affiliations
Contributions
S.J.A. and J.K. performed experiments and analysed the results. T.G. performed the correlation analyses of proteomic and transcriptomic data. S.J.A. designed this study and wrote this manuscript. T.G., J.K., and K.C.R. contributed to writing/editing the manuscript. All authors discussed the results.
Corresponding author
Ethics declarations
Competing Interests
The authors declare that they have no competing interests.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Ahn, SJ., Gu, T., Koh, J. et al. Remodeling of the Streptococcus mutans proteome in response to LrgAB and external stresses. Sci Rep 7, 14063 (2017). https://doi.org/10.1038/s41598-017-14324-w
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-017-14324-w
- Springer Nature Limited