Abstract
While numerous studies have investigated the abundance of ammonia oxidising bacteria and archaea (AOB/AOA) via the ammonia monooxygenase gene amoA, less is known about their small-scale variation and if amoA gene abundance equates to activity. Here we present a spatial and temporal study of ammonia oxidation in two small intertidal bays, Rusheen and Clew bay, Ireland. Potential Nitrification Rate (PNR) was ten-fold higher in Rusheen bay (Clew: 0.27 ± SD 0.55; Rusheen: 2.46 ± SD 3.4 NO2 − µg−1 g−1 day−1, P < 0.001) than in Clew bay but amoA gene abundances were similar between bays, and comparable to those in other coastal ecosystems. Within bays AOB genes increased towards the muddy sediments and were positively correlated with PNR and pH. Less spatial variation was observed in AOA abundances which nevertheless positively correlated with pH and temperature and negatively with salinity and ammonia. Transcriptionally active AOB and AOA were quantified from all sites in Rusheen bay, February 2014, following the general trends observed at DNA level. AOB phylotypes predominantly from the known Nitrosomonas group were distributed across the bay, while Nitrosomonas group B phylotypes were absent from low salinity sites. AOA genes and transcripts were primarily affiliated with Thaumarchaeota group I.1a.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Introduction
Nitrogen is a limited element for primary productivity. While nitrate-limited environments suffer from low primary productivity, excess nitrate in the environment is problematic as it leads to eutrophication and hypoxia. Coastal ecosystems are particularly vulnerable to excess nitrogen loads as they are transition regions between terrestrial and marine environments, where land, streams, groundwater and rivers meet the sea. However, microbial processes within coastal ecosystems mediate the nutrient load entering coastal waters via the various steps of the nitrogen cycle, including nitrification, denitrification, dissimilatory nitrate reduction to ammonia and anammox1,2,3. In fact, it is estimated that greater than 50% of anthropogenic dissolved inorganic nitrogen (DIN) inputs to coastal ecosystems are removed by microbial transformations of the nitrogen cycle4.
Nitrification, the oxidation of ammonia to nitrite and then nitrate, is a central process of the nitrogen cycle. Through coupling with denitrification (the stepwise reduction of nitrate to di-nitrogen gas), it plays an important role in mitigating associated risks of increased DIN5. Traditional understanding of nitrification is a two-step process whereby ammonia oxidising bacteria (AOB) or archaea (AOA) oxidize ammonia to nitrite. Nitrite is then oxidised to nitrate by nitrite oxidising bacteria. The recent discovery of complete ammonia oxidation to nitrate (comammox) by organisms within the genus Nitrospira 6 adds a new dynamic to current understanding.
In two-step nitrification, ammonia oxidation, the first and rate-limiting step, is conducted by AOA and AOB. The ammonia monooxygenase (AMO) enzyme converts ammonia to nitrite7 and is encoded by the amo gene8. amoA is frequently used as a functional marker to detect, quantify and identify AOA and AOB in the environment9. Using this functional gene marker approach, the abundance of AOA and AOB has been extensively surveyed in a range of coastal and estuarine environments to infer the distribution of AOA and AOB10,11,12,13,14,15,16,17. While AOA and AOB are both present, niche differentiation is often apparent18,19,20 and abiotic factors such as ammonia concentration, oxygen, temperature, pH and salinity have been shown to influence abundance and distribution21,22,23,24,25. Within coastal sediments, in some cases AOB have been reported as more abundant13,26,27 whilst other studies report AOA as dominant28,29. Generally a trend of greater AOA at low ammonia18,30 verses AOB at high ammonia concentrations is evident31,32,33,34. What is not understood is if gene abundance equates to activity. For this, the detection and quantification of amoA mRNA transcripts would be more informative than DNA alone35.
Further, to-date, the majority of studies in coastal sediments have been conducted over large spatial scales (e.g. 5 to 330 km26,36,37,38,39). Nitrifier dynamics over smaller spatial scales and active ammonia oxidisers (AO) within intertidal sediments are less well understood. It has been shown that within saltmarshes significant differences in AO diversity occur over local scales (3 cm to 1 km), that are not observed at larger regional (1 to 100 km) or continental (up to 12,500 km apart) spatial scales40. Intertidal bays are dynamic ecosystems that vary on both temporal and spatial scales largely driven by tidal variations. Further within a bay, differences in sediment characteristics occur with organic matter accumulating in low energy areas resulting in muddy sediments verses sandier sediments of the high energy open bay. In addition, any freshwater inputs from rivers, streams and groundwater alter salinity and nutrient concentrations. Thus we hypothesise that such small-scale variation in environmental parameters is likely to drive differences in nitrification activity, diversity and abundances. To this end, we investigated nitrification across a range of sediment types and salinities within and between two small intertidal bays (~1 km2) on the west coast of Ireland over an annual period. Contrasting bays, one located in a rural (Clew bay) and the other in an urban setting (Rusheen bay; approximately 100 km apart) were selected. Two bays were selected to compare and contrast nitrifier abundance and activity across similar environmental gradients (TOC, salinity, sediment type, nutrients, pH) within each to elucidate whether observed differences are greater within or between bays for a given set of environmental drivers. The objectives of the study were to monitor nitrification potential in addition to AOA and AOB amoA gene abundances within and between bays over an annual cycle. Furthermore, we aimed to quantify and identify active ammonia oxidisers in situ by targeting amoA gene transcripts within a single-bay and time-point. Intertidal bays are highly complex and dynamic ecosystems; we therefore hypothesised that nitrifier activity and amoA gene abundances would vary spatially and temporally within and between bays due to fluctuating environmental parameters. Furthermore, we postulated that AOB would be more abundant than AOA due to elevated ammonia concentrations; and that AOA and AOB diversity would vary with changing environmental parameters within intertidal bays.
Results
Site description
Sites representing sandy (RSed_3, RSed_4, RSed_6, RSed_7, CSed_2, CSed_3 and CSed_4) to muddy (RSed_1, RSed_2, RSed_5, CSed_1, CSed_5 and CSed_6) gradients in addition to low salinity (<10 psu) sites (RSed_6, RSed_7 and CSed_6) in each bay, were sampled every three months from May (2013) to February (2014) (Fig. 1). Physical-chemical parameters are reported in Supplementary Table S1A and B. pH ranged between 6.65–8.43, increasing from May to November (One-way ANOVA; P < 0.05). NH4 + concentrations were higher in Rusheen (range: 0–1166.2, average: 283.55 ± SD 38.68 µM−1 g−1 fs−1) and lower in Clew bay in April and November, (range: 0–269.11 µM−1 g−1 fs−1 average, 63.76 ± SD 16.99 µM −1 g−1 fs−1 NH4 +, One-way ANOVA P < 0.05).
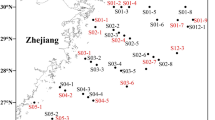
Location of sampling sites within (A) Rusheen Bay (sites RSed_1 to RSed_5) and at the Corrib estuary mouth (RSed_6 & 7), (B) Clew Bay (sites CSed_1 to CSed_6) and (C) Location of Rusheen and Clew Bay on the West Atlantic coast of Ireland. Map was drawn using ArcGis V10 (www.arcgis.com).
Spatial and temporal variation in potential nitrification rates and AOA/AOB amoA gene abundances
PNR in Rusheen (range: 0 to 12.65 µg−1 g−1 d−1) was up to a log-fold greater than Clew bay (range: 0 to 2.14 µg−1 g−1 d−1; Fig. 2; Three-way ANOVA P < 0.001). Within Rusheen bay, PNR varied spatially increasing towards the muddy-sediments (range: 0.06 to 12.65 µg−1 g−1 d−1; P < 0.05), in all months except November. Within Clew bay PNR did not vary spatially with the exception of the low salinity site (CSed_6) which was significantly higher than all other sites in August (0.54 µg−1 g−1 d−1) and November (0.97 µg−1 g−1 d−1; One-way ANOVA P < 0.05). Temporally, within each bay, there was no difference in PNR between April, August and November; however, PNR was higher in Rusheen bay, but lower in Clew bay in February than all other time-points (Fig. 2; Three-way ANOVA; P < 0.05).
Spatial and temporal variation in Potential Nitrification Rates (PNR) (PNR NO2 − µg−1 g−1 day−1) in Rusheen and Clew bay sediments (mean n = 3, standard deviation indicated by error bars). PNR in Rusheen bay are located on the left y-axis, Clew bay is on the right y-axis. Sites are arranged in order of sandy sediments to muddy sediments. Four time points include rates measured in April, August, November 2013 and February 2014. Letters indicate significant differences between sites within a bay (One-way ANOVA, P < 0.05), Greek letters indicate significant differences between time points and the asterisk symbols indicate significant differences between bays (Three-way ANOVA, P < 0.05).
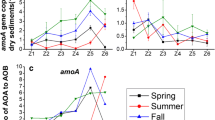
AOB amoA gene abundances ranged from 1.89 × 105 to 2.5 × 107 (mean ± SD 3.81 × 106 ± 1.5 × 107) copies per gram of sediment (Fig. 3). In contrast to the lower PNR in Clew bay (Three-way ANOVA; P < 0.001) there was no difference in AOB amoA gene abundances between bays (Three-way ANOVA; P = 0.180). Within each bay, AOB amoA gene generally increased towards the muddy sites (One-way ANOVA; P < 0.05, Fig. 3). Temporally, within Rusheen bay, AOB amoA gene abundances were similar in the winter months (November and February) but were significantly different between April and August (One-way ANOVA; P < 0.05). No temporal change was observed in Clew bay. AOA amoA gene abundances ranged from 1.21 × 104 to 6.29 × 107 (mean ± SD 2.27 × 106 ± 1.14 × 106) copies per gram of sediment (Fig. 3) with AOA amoA gene abundances statistically higher in Clew bay (Three-way ANOVA; P < 0.05). AOA gene abundances were generally the same within a bay, with any significant increase observed at muddy sediment sites only (One-way ANOVA; P < 0.05).
Spatial and temporal variation in AOA (red) and AOB (black) amoA gene abundances in Rusheen and Clew bay sediments. Significant differences within a bay are indicated by letters above the site (One-way ANOVA, P < 0.05). Significant differences between time-points are indicated by Greek symbols (Two-way ANOVA, P < 0.05).
AOB gene abundances correlated positively with PNR, in situ TOC and in situ pH (r = 0.509, P < 0.001; r = 0.434, P < 0.001; r = 0.315 P < 0.05); and negatively with temperature and NH4 + concentration (r = 0.459 P < 0.001; r = 0.404 P < 0.001). AOA gene abundances positively correlated with in situ pH and temperature (r = 0.69 P < 0.001; r = 0.242 P < 0.05) and negatively with in situ salinity and NH4 + concentration (r = −0.245, P < 0.05; r = −0.438, P < 0.001). PNR positively correlated with in situ TOC and sediment salinity (r = 0.223, P < 0.05; r = 0.265, P < 0.05; Supplementary Table S2).
Abundance and community shift of amoA transcripts across Rusheen bay
Rusheen bay, February 2014 was selected to further investigate the diversity and abundance of amoA DNA (total) and mRNA (active) from AOA and AOB based on the overall higher rates of PNR in Rusheen than Clew bay. Only February sediments were targeted as we wanted to use fresh sediments to investigate transcript activity. AOB amoA transcripts across the bay ranged from 1.93 × 105–1.52 × 107 per gram of sediment with an average 6.02 × 106 ± SD 2.77 × 106 and were greater than AOA amoA transcripts at all sites except for Rsed_3 (9.7 × 104 − 6.2 × 106 per gram of sediment, average 2.49 × 106 ± SD 3.31 × 106; Fig. 4). AOB amoA transcripts increased from the sandier to muddier sediments (One-way ANOVA, P < 0.001). AOA amoA transcripts were quantified from all sites, but no spatial difference was observed (One-way ANOVA, P = 0.126). AOB amoA transcripts correlated positively with PNR (r = 0.868, P < 0.001), while AOA did not; in addition, AOB amoA transcripts and in situ nitrate concentrations were significantly correlated (r = 0.444, P < 0.05) (Spearman rank correlations; Supplementary Table S3).
Log10 AOA and AOB transcript numbers g−1 sediment for February 2014 in Rusheen bay. AOB numbers are indicated by the black squares while AOA are indicated by the red circle. The black letters represent the significant differences between sites across Rusheen bay for AOB transcripts (One-way ANOVA, P < 0.001).
To determine if different active AOB amoA phylotypes were present across the bay, T-RFLP was conducted. Site RSed_4, the coarse sand site, was excluded due to low PCR yield. Three groups of AOB amoA cDNA phylotypes, at 45% similarity, were present (Fig. 5a) – group I the low salinity sites, Rsed_6 and 7 (ANOSIM r = 0.815, P = 0.1); group II the muddy sites, RSed_1 and 2 (ANOSIM, r = 0.37, P = 0.2) and group III the sandy sites, RSed_3 and RSed_5 (ANOSIM r = 0.333, P = 0.1). Furthermore, CCA showed that a combination of ammonium (NH4 +; f = 1.6, P = 0.024) and TOC (f = 2.05, P = 0.004) explained 42.74% of variation of AOB phylotypes present (Fig. 5b). After which, salinity (f = 1.73, P = 0.008) was the next significant variable explaining 32.6% of the distribution of active AOB phylotypes across the bay.
AOB amoA transcript community analysis (a) Cluster dendrogram representing T-RFLP profiles of AOB amoA transcripts from Rusheen bay sediments February 2014. Scale bar: Bray-Curtis similarity coefficient. Clusters highlighted in blue and green indicate salinity below 9 psu and equal to 24 psu respectively. Shapes represent different sediment types from muddy sand to sand. (b) Canonical correspondence analysis illustrates the separation of AOB cDNA amoA gene TRFs based on physical/chemical parameters measured. The different colours represent an individual site. The most significant parameters are represented by the labelled vectors. The abbreviations PNR and TOC represent the potential nitrification rates and total organic carbon respectively.
amoA gene and transcript phylogeny of ammonia oxidisers within Rusheen bay sediments
Based on the cluster dendrogram (Fig. 5a) sites RSed_1, 5, 6 & 7 were selected for amoA AOA/AOB gene and transcript sequencing. In general, DNA PCR amplicons were too weak for Illumina MiSeq analysis so a clone library approach was used. In total, 68 AOB DNA and 197 cDNA amoA sequences and 61 AOA DNA and 32 cDNA sequences, correctly translating to protein were recovered. From these, thirty-six DNA and 41 cDNA AOB OTUs and 19 DNA and 8 cDNA AOA OTUs at 97%41 similarity were identified (Supplementary Table S4). amoA AOB transcripts were retrieved from all sites, but amoA AOA transcripts were only recovered from the sandy sediment sites RSed_5 (high salinity) and RSed_6 (low salinity).
Archaeal amoA grouped into Thaumarchaeota Group I.1a, and Group I.1b. (Fig. 6). The majority of AOA fell into Thaumarchaeota group I.1a, with a single OTU from RSed_5 falling into group I.1b. The sequences affiliating with group I.1b shared high identity to Can. Nitrososphaera evergladensis and gargensis and the sequence from mangrove sediments. One cluster, closely related to Can. Nitrosoarchaeum koreensis MY1, contained the majority of OTUs recovered from all sites. A single DNA phylotype (amoAArchD23) from RSed_1, the high salinity muddy sediment site, clustered with Nitrosopumilus maritimus, and two candidate Nitrosopumilus sp. The remaining sequences in Group I.1a clustered with environmental clones generally from similar coastal environments.
Neighbour-joining phylogenetic tree of AOA amoA genes at protein level (≥ 97% similarity). Bootstrap values shown are greater than 70% of 1000 are shown near nodes. Accession numbers from GenBank are shown from other studies and isolates. Circles indicate DNA sequences, triangles indicate cDNA sequences. The numbers in brackets indicate the number of sequences recovered from that site. The colours maroon, yellow, blue and aqua signify different sampling sites RSed_1, RSed_5, RSed_6 and RSed_7 respectively.
All AOB fell into the Nitrosomonas lineage, which was further divided into three clusters; the known Nitrosomonas group, and Nitrosomonas group A and B (Fig. 7), corresponding to a similar classification as used in O’Mullan and Ward, 2005. The known Nitrosomonas group accounted for 93.4% of sequences, including the dominant OTUs found at all sites (amoABacD4 and D5). These sequences were affiliated with isolates and enrichments N. aestuarii, N. ureae, N. oligotropha and N. marina. Nitrosomonas group A cluster contains OTUs from the high salinity muddy sediment site, RSed_1 only. These were highly similar to amoA sequences from other coastal ecosystems distributed globally. The Nitrosomonas group B cluster contained sequences from three sites (RSed_1, 5 and 6, muddy and sandy sediments) and absent from Rsed_7, the lowest salinity site (<1 psu). These clustered with amoA phylotypes from similar high salinity coastal ecosystems.
Neighbour-joining phylogenetic tree of AOB amoA genes at protein level (≥ 97% similarity). Bootstrap values shown are greater than 70% of 1000 are shown near nodes. Accession numbers from GenBank are shown from other studies and isolates. Circles indicate DNA sequences, triangles specify cDNA sequences. The numbers in brackets indicate the number of sequences recovered from that site. The colours maroon, yellow, blue and aqua signify the sampling sites RSed_1, RSed_5, RSed_6 and RSed_7, respectively.
To identify AOB cDNA TRFs, in silico TRFs were generated from amoA gene sequences and compared to observed TRFs (Table 1). TRFs representing the two dominant amoA gene sequences (amoABacD4 and D5, Fig. 7) from the known Nitrosomonas group, were present at all sites. In addition, TRFs equating to two other phylotypes within the known Nitrosomonas group (amoABacR42 and 47) were ubiquitously present. TRFs representing Nitrosomonas group A (amoABacD1, D7 & D24) were distributed across the bay. However, TRF’s representing the Nitrosomonas group B, were present at the high salinity sites only.
Discussion
Spatial variation in nitrification and drivers of activity, abundance and distribution of AO were investigated within and between intertidal bays. Rusheen bay (0–12.6 µg−1 g−1d−1) had higher nitrifier activity than Clew bay (0–0.972 µg−1 g−1d−1). The reason for the difference in activity between the bays was not revealed in this study. Rusheen bay PNRs were slightly higher compared to similar environments e.g. Western English Channel (1.344–9.408 µg−1ml−1d−1), salt marsh sediments, Skidaway island (0–7.728 µg −1g−1d−1) but lower than the hypernutrified Colne estuary, UK (126–546 µg −1 g−1d−1)27,42,43. Small scale spatial variation in PNR was observed with rates increasing towards the muddy sediments (0.064–12.648 µg−1 g−1 d−1). This is consistent with Zheng et al., (2014) who also reported highest nitrification in muddy sediments and lowest in sandy sediments37. In contrast, no difference in PNR was reported between sandy and muddy sediment types in deep-sea sediments44. In Rusheen bay, intermediate salinities (16.66 and 24 psu) were recorded at sites RSed_1–5, August 2013 and February 2014 respectively. Both time-points had higher nitrification rates compared to the higher salinity (~32 psu) time-points (April and November 2013). While, in general there was no difference in PNR activity across Clew Bay, when significant increases were observed it was always associated with the low salinity site (CSed_6; 0–0.23 psu). Similarly, studies from a number of other temperate estuaries report highest nitrification at the intermediate salinities (5 to 20 psu)27,45,46.
AOB at DNA level, were 1 to 2 orders of magnitude greater than AOA amoA at all four-time points in both bays. amoA gene copies per cell in AOB range from 1 to 347 while a single copy is present in AOA48. Adjusting AOB amoA gene abundances for the maximum amoA gene copies per cell they still outnumbered AOA. Despite the variation in PNR between bays, AOB amoA gene abundances were not different between bays. Indeed AOB amoA gene abundances from both bays in this study were similar in range to other coastal bay and intertidal sediments where AOB were also more abundant than AOA27,37,39,49. In contrast other water column and submerged coastal and estuarine sediment studies have reported AOA as more abundant26,28,38,50,51,52. Further there was a difference in AOA amoA gene abundance between the bays in this study, however they were still within the range of those reported in other coastal sediments14,39.
AOB gene abundances correlated significantly with PNR in both bays, and thus also shared the general trend of increasing in abundance towards muddy sediments. Further AOB abundance was positively correlated with TOC, pH and negatively with NH4 + concentration (Supplementary Table S2). Similar observations were also seen in the Yangtze estuary and bioturbated sediments24,37. The large seasonal variation in TOC (Supplementary Table S1a and b) may have contributed to an increase in ammonia via increased mineralisation. This potentially had a knock on effect on nitrification, explaining the correlation of AOB gene abundances, transcripts and PNR with TOC (Supplementary Tables S2 and S3). No correlation of AOB abundance with salinity was seen. AOA amoA gene abundances also tended to show spatial change across the bays increasing towards the muddy sediment although the trend was less distinct than for AOB. AOA gene abundance was correlated with both NH4 + and salinity. Similar observation have been shown in other coastal sediments and AOA are thought to play a more significant role in low salinity ecosystems26,37,53.
Both AOB and AOA amoA transcripts were quantified in situ from all sites in Rusheen bay, February 2014, indicating the presence of transcriptionally active AOB and AOA within intertidal sediments. A study of ammonia oxidisers in North Sea sediments, characterised by low ammonia concentrations (3 to 53 µM) and dominated by AOA, reported the co-existence of metabolically active AOA and AOB. However, in the North Sea sediments amoA transcripts from AOB were more abundant than AOA, despite AOAs higher gene abundance52. On the other hand in Antarctic coastal waters AOA genes and transcripts were more abundant than AOB genes and transcripts54. Our data suggests that AOB are the primary drivers of AO in Rusheen bay, based on consistently higher spatial and temporal gene abundances, in addition to the higher amoA transcripts numbers, and significant correlation of AOB amoA genes and transcripts with PNR (Supplementary Table S3). While AOA amoA gene abundances also correlated significantly with PNR, AOA transcripts did not, nor with any other parameters (r = 0.515, P < 0.05; Supplementary Table S3).
Within Rusheen bay, the known Nitrosomonas group were the dominant AOB. The two most abundant AOB OTUs within the know Nitrosomonas (amoAbacD5 and D4) were present at both DNA and/or mRNA level and found at all sites. They are highly similar to N. ureae and clustered with similar phylotypes from estuarine environments characterised by high ammonia55,56. N. ureae and N. oligotrophy- like AOB can adapt to high initial ammonium concentrations (up to 10 mM) but also can grow at low ammonia concentrations (limiting concentration of 6 µM)57. This is consistent with ammonia concentrations within Rusheen bay where in situ concentrations never went above 1.2 mM NH4 +.
The Nitrosomonas group A had low identity to known AOBs and were only recovered from muddy sites (Rsed_1, 24 psu), but T-RFLP data linked active phylotypes from this group across the entire bay. O’Mullan and Ward showed Nitrosomonas group A phylotypes associated with brackish water (15–30 psu)58. In contrast, sequences affiliated with Nitrosomonas group B were only present at the high salinity site and were closely related with sequences from marine coastal sediment sites with high salinity located in Monterey bay, California and Sendai bay, Japan, suggesting this group is associated with high salinity sites globally58.
AOA clustered with group I.1a. and were highly similar to Can. Nitrosoarchaeum koreensis MY1 at 99% similarity59 (Fig. 7). N. koreensis MY1, is inhibited by high salinity and grows optimally in 11 µM NH4 + 60. While OTUs from the MY1 cluster were present across the bay, transcripts were only recovered from the low salinity (0 psu), low ammonia (0.39 µM) site (RSed_6). Indeed, the majority of AOA amoA cDNA transcripts were recovered from this site, further, AOA gene abundance negatively correlated with high salinity over four time-points (Supplementary Table S2), indicating this cluster may be more active at low salinity. Nitrosoarchaeum-like sequences are consistently found in estuarine and freshwater sediments globally61,62 In San Francisco’s Sacramento-San Joaquin Delta this group was found in the low salinity sites where AOA tended to be more abundant than AOB13,26.
Overall, this study shows that AOB and AOA co-occur within intertidal sediments. AOB are likely a larger contributor to ammonia oxidation, based on their abundance at both DNA and mRNA level and through their significant correlations with PNR. However transcriptionally active AOA were also present. Further work to explore the co-occurrence and activity of AOA and AOB is needed. We also report evidence of small scale changes in ammonia oxidiser phylotypes across the bay selecting for AOB that are ubiquitously present and those that are only found at high salinity sites.
Materials and Methods
Site description and field sampling
Rusheen (53° 25.5894′N, −9° 11.9532′W) and Clew bay (53° 78.6962′N, −9° 64.9515′W), Ireland (Fig. 1) were sampled over four time-points: May, August, November 2013 and February 2014. Distances between sites ranged from ~200 m to 5 km apart within a site to 100 km apart between bays. At each site, at low tide, three replicates were collected from the top 2 cm of sediment, to ensure the top most layer of oxic sediment was sampled. Each replicate, composed of 10 random samples and was collected within a 10 m2 area, and stored on ice until return to the laboratory from which 0.5–0.7 g aliquots were flash frozen at −80 °C for molecular analysis. Samples for Potential Nitrification Rate (PNR) and physicochemical analysis were stored at 4 °C upon analysis. The Mastersizer 2000 Laser Particle Sizer (Malvern, UK) was used to determine particle sizes <1mm. Particles above 1 mm were analysed using a dry sieving technique63. Sediment was classified using the EUNIS A2 classification scheme. Ammonium (NH4 +) and nitrate (NO3 −) were extracted from the sediment with 1 M KCl and analysed using a colorimetric method64. Nitrite (NO2 −) was extracted with 2 M KCl and analysed using Hach Nitriver 3 (Hach-Lange, Ireland). Total organic carbon (TOC) content was determined by combustion at 450 °C for 12 hours63.
Potential nitrification rates (PNR)
Triplicate PNR containing 5 g sediment (wet-weight), 30 ml site-water amended with 24 µM sodium azide65 (NaN3, nitrite oxidation inhibitor) and (NH4)2SO4 at both 0 and 250 µM concentrations (Sigma Aldrich, Ireland), were incubated at 15 °C in the dark shaking (90 rpm) for 24 h. Nitrite was extracted and measured as described above. PNR rate was calculated as follows: \(\frac{(mg/l\,Nitrite\times 60\,ml)}{(5\,g\,sediment)}\) = PNR per g sediment per day.
DNA and RNA co-extraction
DNA and RNA was co-extracted from 0.5–0.7 g fresh sediment using a bead-beating protocol as detailed in Smith et al.66. Nucleic acids were re-suspended in 50 µl DEPC water of which a 25 µl aliquot was treated with TURBO DNase (Ambion, Ireland) according to the manufacturer’s instructions to prepare RNA. The remaining 25 µl aliquot was left untreated for use as DNA. The absence of DNA in the RNA fraction was confirmed by PCR of the 16 S rRNA (F63 and R518; Supplementary Table S5) using 2 µl RNA (neat to 10−2 dilution) as template. RNA was converted to cDNA using Superscript III (Life Science, USA) as in Smith et al., 2015, using gene specific primers targeting AOA or AOB amoA3. Each 10 µl RT reaction contained 8 µl of RNA, 2 mM of the appropriate reverse primer, ArchamoAR or BacamoA-2R (Supplementary Table S5), and 10 mM of each dNTP.
amoA (RT)-Q-PCR
DNA standard curves were constructed from the target gene (Supplementary Table S5) according to Smith et al.67. A one-in-five dilution series of appropriate standard was used over an eight-point dynamic range from 1010 to 103 gene copies per μl. amoA (Supplementary Table S5) was amplified from triplicate sediments collected from each site. Each 20 µl Q-PCR reaction mixture contained 10 µl EVA Green master mix (Biorad, Ireland), 0.4 µl of each primer (10 mM) (Supplementary Table S5), and 2 µl template DNA (10−1 dilution) or cDNA. Triplicate no-template controls (NTC) were included. Specificity of the amplicons was confirmed by melting curve analysis at the end of each (RT)-Q-PCR experimental run (Supplementary Table S5). The slope, y-intercept, and r2 values of standard curves are reported in Supplementary Table S6.
Terminal restriction fragment length polymorphism (T-RFLP) of AOB amoA transcripts
AOB amoA cDNA from Ruhseen bay sites, February 2014 was amplified with a HEX labelled forward primer (Supplementary Table S5). Each 50 µl reaction mix contained 1 µl amoA AOB gene specific cDNA; 20 µM forward and reverse primers and 25 µl 2X MyTaq mix (Bioline, Ireland) using a touchdown PCR (Supplementary Table S5). PCR products were purified using SureClean Plus (Bioline, Ireland) and re-suspended in 15 µl of sterile water. Five µl of PCR product was independently digested with three enzymes, TaqI, HhaI and AluI, (Thermo Fisher Scientific, Ireland) according to manufacturer’s instructions. Samples were sent to Source Bioscience, Ireland, for T-RFLP analysis. Terminal Restriction Fragments (TRF) were sized against a ROX-genescan 500 internal size standard using Peakscanner software v1.0 (Applied Biosystems, Ireland). T-RFLP profiles per site from individual enzymes were combined into a single profile and aligned using T-Align68.
Cloning and sequence analysis of bacterial and archaeal amoA genes and transcripts
AOA and AOB amoA genes and transcripts from replicate samples from sites RSed_1, RSed_5, RSed_6 & RSed_7 (Fig. 1) Rusheen Bay February 2014 were cloned and sequenced. 5 µl DNA (10−1) or cDNA (neat) was amplified as follows: 10 × 1.5 mM MgCl2 PCR buffer (Sigma Aldrich, Ireland), 0.2 mM of each dNTP (Sigma Aldrich, Ireland), 0.25 µM of each primer ArchamoAF, and ArchamoAR, or BacamoA1F and BacamoA-2R (Supplementary Table S5; Eurofins, UK), and 2.5 units of Taq polymerase (Sigma Aldrich, Ireland) using touchdown PCR (Supplementary Table S5). Amplicons were gel purified using a Gel Purification Kit (Qiagen, Ireland) according to manufacturer’s instructions and re-suspended in 15 µl sterile water. PCR products were cloned using PGem-T cloning kit (Promega, Ireland) according to manufacturer’s instructions. Fifty white colonies were screened using PGEM-T vector specific primers M13F and R (10 mM) (Supplementary Table S5). Clones containing inserts of the correct size were sent for sequencing using vector primers M13F/T7F (Source Bioscience, Ireland). amoA nucleotide sequences were translated into protein sequences using the Translate tool on ExPASy (Expert Protein Analysis System; http://us.expasy.org/tools/dna.html). Nucleotide and protein sequences were compared to entries in GenBank using BlastN and BlastP69. Protein sequence alignments were constructed using Bioedit (version 7.2)70. amoA gene sequences displaying more than 97% similarity were grouped into a single operational taxonomic unit (OTU) using DOTUR software by the furthest neighbour approach71. Distance matrices were calculated using the PROTDIST program in PHYLIP72. Phylogenetic trees were created from the distance matrices using the neighbour-joining method73 and Kimura substitution algorithm74 in MEGA-675. Consensus trees were calculated after bootstrapping (1,000 replicate trees).
Statistical analysis
All data was tested for normality using Kolmogorov-Smirnov test and log-transformed as necessary76. Variation in PNR, gene abundances and environmental variables between sites, bays and time-points were analysed using a three-way analysis of variance (ANOVA) followed by a post-hoc Bonferroni test77 in SPSS v21 (IBM, USA). A one-way ANOVA was used on environmental variables between sites in both bays at each time-point in SPSS v21. Analysis of covariance (ANCOVA) was conducted on gene abundance data by time-point in Graph Pad Prism v6 (GraphPad software, USA). A cluster dendrogram was created in Primer 7 (Quest research limited) using a bray-curtis similarity resemblance matrix, analysis of similarities (ANOSIM) was carried out between sites. Canonical correspondence analysis (CCA) using CANOCO 5 (http://www.canoco5.com) was used to explore relationships between community structure and environmental parameters. Explanatory value of environmental factors was determined using forward selection (tested by 499 Montecarlo permutations). Differences and correlation coefficients were considered significant at P < 0.05 unless otherwise stated in the text. Only significant explanatory variables were plotted.
Data availability
The sequence datasets generated during and/or analysed during the current study are available in the NCBI repository, with the following accession numbers. AOA accession KX664728 - KX664820. AOB DNA accession numbers KX673291 - KX673360. AOB cDNA accession numbers KX690304 - KX690497. All other datasets generated during and/or analysed during the current study are available from the corresponding author on reasonable request.
References
Trimmer, M. & Nicholls, J. C. Production of nitrogen gas via anammox and denitrification in intact sediment cores along a continental shelf to slope transect in the North Atlantic. Limnol. Oceanogr. 54, 577–589 (2009).
Smith, J. M., Mosier, A. C. & Francis, C. A. Spatiotemporal Relationships Between the Abundance, Distribution, and Potential Activities of Ammonia-Oxidizing and Denitrifying Microorganisms in Intertidal Sediments. Microb. Ecol. 69, 13–24 (2014).
Smith, C. J. et al. Seasonal variation in denitrification and dissimilatory nitrate reduction to ammonia process rates and corresponding key functional genes along an estuarine nitrate gradient. Front. Microbiol. 6 (2015).
Lohse, L., Malschaert, J. F. P., Slomp, C. P., Helder, W. & Vanraaphorst, W. Nitrogen cycling in North Sea sediments - Interaction of denitrification and nitrification in offshore and coastal areas. Mar. Ecol. Prog. Ser. 101, 283–296 (1993).
Mosier, A. C. & Francis, C. A. In Methods in Enzymology: Research on Nitrification and Related Processes Part B 486, 205–221 (2011).
Daims, H. et al. Complete nitrification by Nitrospira bacteria. Nature 528, 504–509 (2015).
Dua, R. D., Bhandari, B. & Nicholas, D. J. D. Stable isotope studies on the oxidation of ammonia to hydroxylamine by Nitrosomonas europaea. FEBS Lett. 106, 401–404 (1979).
Kuhn, R., Sinigalliano, C., Kuhn, D. & Jones, R. Amplification of the amoA gene from diverse species of ammonium-oxidizing bacteria and from an indigenous bacterial population from seawater. Appl. Environ. Microbiol. 61, 2702 (1995).
Rotthauwe, J.-H., Witzel, K.-P. & Liesack, W. The Ammonia Monooxygenase Structural Gene amoA as a Functional Marker: Molecular Fine-Scale Analysis of Natural Ammonia-Oxidizing Populations. Appl. Environ. Microbiol. 63, 4704–4712 (1997).
Caffrey, J. M., Bano, N., Kalanetra, K. & Hollibaugh, J. T. Ammonia oxidation and ammonia-oxidizing bacteria and archaea from estuaries with differing histories of hypoxia. ISME J. 1, 660–662 (2007).
Magalhães, C. M., MacHado, A. & Bordalo, A. A. Temporal variability in the abundance of ammonia-oxidizing bacteria vs. archaea in sandy sediments of the Douro River estuary, Portugal. Aquat. Microb. Ecol. 56, 13–23 (2009).
Moin, N. S., Nelson, K. A., Bush, A. & Bernhard, A. E. Distribution and diversity of archaeal and bacterial ammonia oxidizers in salt marsh sediments. Appl. Environ. Microbiol. 75, 7461–7468 (2009).
Mosier, A. C. & Francis, C. A. Relative abundance and diversity of ammonia-oxidizing archaea and bacteria in the San Francisco Bay estuary. Environ. Microbiol. 10, 3002–3016 (2008).
Santoro, A. E., Francis, C. A., Sieyes, N. R. De & Boehm, A. B. Shifts in the relative abundance of ammonia-oxidizing bacteria and archaea across geochemical gradients in a subterranean estuary. Environ. Microbiol. 10, 1068–1079 (2008).
Abell, G. C. et al. Archaeal ammonia oxidizers and nirS-type denitrifiers dominate sediment nitrifying and denitrifying populations in a subtropical macrotidal estuary. ISME J. 4, 286–300 (2010).
Bernhard, A. E. et al. Abundance of ammonia-oxidizing archaea and bacteria along an estuarine salinity gradient in relation to potential nitrification rates. Appl. Environ. Microbiol. 76, 1285–1289 (2010).
Wankel, S. D., Mosier, A. C., Hansel, C. M., Paytan, A. & Francis, C. A. Spatial variability in nitrification rates and ammonia-oxidizing microbial communities in the agriculturally impacted Elkhorn Slough estuary, California. Appl. Environ. Microbiol. 77, 269–280 (2011).
Martens-Habbena, W. et al. Ammonia oxidation kinetics determine niche separation of nitrifying Archaea and Bacteria. Nature 461, 976–979 (2009).
Prosser, J. I. & Nicol, G. W. Archaeal and bacterial ammonia-oxidisers in soil: The quest for niche specialisation and differentiation. Trends in Microbiology 20, 523–531 (2012).
Hatzenpichler, R. Diversity, Physiology, and Niche Differentiation of Ammonia-Oxidizing Archaea. Appl. Environ. Microbiol. 78, 7501–7510 (2012).
Zhang, Y., Chen, L., Dai, T., Tian, J. & Wen, D. The influence of salinity on the abundance, transcriptional activity, and diversity of AOA and AOB in an estuarine sediment: a microcosm study. Appl. Microbiol. Biotechnol. 99, 9825–9833 (2015).
Ke, X., Lu, W. & Conrad, R. High Oxygen Concentration Increases the Abundance and Activity of Bacterial Rather than Archaeal Nitrifiers in Rice Field Soil. Microb. Ecol. 70, 961–970 (2015).
Urakawa, H. et al. Ammonia availability shapes the seasonal distribution and activity of archaeal and bacterial ammonia oxidizers in the Puget Sound Estuary. Limnol. Oceanogr. 59, 1321–1335 (2014).
Laverock, B., Tait, K., Gilbert, J. A., Osborn, A. M. & Widdicombe, S. Impacts of bioturbation on temporal variation in bacterial and archaeal nitrogen-cycling gene abundance in coastal sediments. Environ. Microbiol. Rep. 6, 113–121 (2014).
Freitag, T. E., Chang, L. & Prosser, J. I. Changes in the community structure and activity of betaproteobacterial ammonia-oxidizing sediment bacteria along a freshwater – marine gradient. Environ. Microbiol. https://doi.org/10.1111/j.1462-2920.2005.00947.x (2006).
Damashek, J., Smith, J. M., Mosier, A. C. & Francis, C. A. Benthic ammonia oxidizers differ in community structure and biogeochemical potential across a riverine delta. Front. Microbiol. 6, 1–18 (2015).
Li, J. et al. amoA gene abundances and nitrification potential rates suggest that benthic ammonia-oxidizing bacteria and not archaea dominate N cycling in the Colne estuary, United Kingdom. Appl. Environ. Microbiol. 81, 159–165 (2015).
Lipsewers, Y. A. et al. Seasonality and depth distribution of the abundance and activity of ammonia oxidizing microorganisms in marine coastal sediments (North Sea). Front. Microbiol. 5, 1–12 (2014).
Wuchter, C. et al. Archaeal nitrification in the ocean. Proc. Natl. Acad. Sci. USA 103, 12317–12322 (2006).
Sauder, L. A., Peterse, F., Schouten, S. & Neufeld, J. D. Low-ammonia niche of ammonia-oxidizing archaea in rotating biological contactors of a municipal wastewater treatment plant: Thaumarchaeota in rotating biological contactors. Environ. Microbiol. 14, 2589–2600 (2012).
Verhamme, D. T., Prosser, J. I. & Nicol, G. W. Ammonia concentration determines differential growth of ammonia-oxidising archaea and bacteria in soil microcosms. ISME J. 5, 1067–1071 (2011).
Jia, Z. & Conrad, R. Bacteria rather than Archaea dominate microbial ammonia oxidation in an agricultural soil. Environ. Microbiol. 11, (1658–1671 (2009).
Di, H. J. et al. Ammonia-oxidizing bacteria and archaea grow under contrasting soil nitrogen conditions. FEMS Microbiol. Ecol. 72, 386–394 (2010).
Baolan, H. et al. Effect of Different Ammonia Concentrations on Community Succession of Ammonia-oxidizing Microorganisms in a Simulated Paddy Soil Column. PLoS One 7, e44122 (2012).
Philippot, L., Hallin, S., Sá Nchez, S. & Olson, B. Finding the missing link between diversity and activity using denitrifying bacteria as a model functional community This review comes from a themed issue on Ecology and industrial microbiology Edited. Curr. Opin. Microbiol. 8, 234–239 (2005).
Marton, J. M., Roberts, B. J., Bernhard, A. E. & Giblin, A. E. Spatial and Temporal Variability of Nitrification Potential and Ammonia-Oxidizer Abundances in Louisiana Salt Marshes. Estuaries and Coasts 38, 1824–1837 (2015).
Zheng, Y. et al. Community Dynamics and Activity of Ammonia-Oxidizing Prokaryotes in Intertidal Sediments of the Yangtze Estuary. Appl. Environ. Microbiol. 80, 408–419 (2014).
Zhang, Q. et al. Shifts in the pelagic ammonia-oxidizing microbial communities along the eutrophic estuary of Yong River in Ningbo City, China. Front. Microbiol. 6, (2015).
Yu, S. et al. Diversity, Abundance, and Niche Differentiation of Ammonia-Oxidizing Prokaryotes in Mud Deposits of the Eastern China Marginal Seas. Front. Microbiol. 7, 1–13 (2016).
Martiny, J. B. H., Eisen, J. A., Penn, K., Allison, S. D. & Horner-Devine, M. C. Drivers of bacterial beta-diversity depend on spatial scale. Proc. Natl. Acad. Sci. USA 108, 7850–4 (2011).
Wang, B. et al. Differential contributions of ammonia oxidizers and nitrite oxidizers to nitrification in four paddy soils. ISME J. 9, 1062–75 (2015).
Dollhopf, S. L. et al. Quantification of Ammonia-Oxidizing Bacteria and Factors Controlling Nitrification in Salt Marsh Sediments Quantification of Ammonia-Oxidizing Bacteria and Factors Controlling Nitrification in Salt Marsh Sediments. Appl. Environ. Microbiol. 71(1), 240–246 (2005).
Laverock, B. et al. Bioturbation determines the response of benthic ammonia-oxidizing microorganisms to ocean acidification. Philos. Trans. R. Soc. London B Biol. Sci. 368 (2013).
Henriksen, K., Hansen, J. I. & Blackburn, T. H. Rates of nitrification, distribution of nitrifying bacteria, and nitrate fluxes in different types of sediment from Danish waters. Mar. Biol. 61, 299–304 (1981).
Bernhard, A. E., Tucker, J., Giblin, A. E. & Stahl, D. A. Functionally distinct communities of ammonia-oxidizing bacteria along an estuarine salinity gradient. Environ. Microbiol. 9, 1439–1447 (2007).
Magalhäes, C. M., Wiebe, W. J., Joye, S. B. & Bordalo, A. A. Inorganic nitrogen dynamics in intertidal rocky biofilms and sediments of the Douro River estuary (Portugal). Estuaries 28, 592–607 (2005).
Norton, J. M., Alzerreca, J. J., Suwa, Y. & Klotz, M. G. Diversity of ammonia monooxygenase operon in autotrophic ammonia-oxidizing bacteria. Arch. Microbiol. 177, 139–149 (2002).
Walker, C. B. et al. Nitrosopumilus maritimus genome reveals unique mechanisms for nitrification and autotrophy in globally distributed marine crenarchaea. Proc. Natl. Acad. Sci. USA 107, 8818–23 (2010).
Lisa, J. A., Song, B., Tobias, C. R. & Hines, D. E. Genetic and biogeochemical investigation of sedimentary nitrogen cycling communities responding to tidal and seasonal dynamics in Cape Fear River Estuary. Estuar. Coast. Shelf Sci. 167, A313–A323 (2015).
Mincer, T. J. et al. Quantitative distribution of presumptive archaeal and bacterial nitrifiers in Monterey Bay and the North Pacific Subtropical Gyre. Environ. Microbiol. 9, 1162–75 (2007).
Santoro, A. E., Casciotti, K. L. & Francis, C. A. Activity, abundance and diversity of nitrifying archaea and bacteria in the central California Current: Nitrification in the central California Current. Environ. Microbiol. 12, 1989–2006 (2010).
Zheng, Y. et al. Diversity, abundance, and activity of ammonia-oxidizing bacteria and archaea in Chongming eastern intertidal sediments. Appl. Microbiol. Biotechnol. 97, 8351–8363 (2013).
Fan, H., Bolhuis, H. & Stal, L. J. Nitrification and nitrifying bacteria in a coastal microbial mat. Front. Microbiol. 6, 1–13 (2015).
Tolar, B. B., Ross, M. J., Wallsgrove, N. J. & Liu, Q. Contribution of ammonia oxidation to chemoautotrophy in Antarctic coastal waters. ISME (2016).
Cébron, A., Berthe, T. & Garnier, J. Nitrification and Nitrifying Bacteria in the Lower Seine River and Estuary (France). Appl. Environ. Microbiol. 69, 7091–7100 (2003).
Stehr, G., Böttcher, B., Dittberner, P., Rath, G. & Koops, H. P. The ammonia-oxidizing nitrifying population of the River Elbe estuary. FEMS Microbiol. Ecol. 17, 177–186 (1995).
Bollmann, A. & Laanbroek, H. J. Continuous culture enrichments of ammonia-oxidizing bacteria at low ammonium concentrations. FEMS Microbiol. Ecol. 37, 211–221 (2001).
Mullan, G. D. O. & Ward, B. B. Relationship of Temporal and Spatial Variabilities of Ammonia-Oxidizing Bacteria to Nitrification Rates in Monterey Bay, California Relationship of Temporal and Spatial Variabilities of Ammonia-Oxidizing Bacteria to Nitrification Rates in Monterey Bay. Appl. Environ. Microbiol. 71, 697–705 (2005).
Kim, B. K. et al. Genome Sequence of an Ammonia-Oxidizing Soil Archaeon, ‘Candidatus Nitrosoarchaeum koreensis’ MY1. J. Bacteriol. 193, 5539–5540 (2011).
Jung, M. Y. et al. Enrichment and characterization of an autotrophic ammonia-oxidizing archaeon of mesophilic crenarchaeal group I.1a from an agricultural soil. Appl. Environ. Microbiol. 77, 8635–8647 (2011).
Cao, H., Auguet, J.-C., Gu, J.-D., Cornwell, W. & Morlon, H. Global Ecological Pattern of Ammonia-Oxidizing Archaea. PLoS One 8, e52853 (2013).
Vetterli, A., Hietanen, S. & Leskinen, E. Spatial and temporal dynamics of ammonia oxidizers in the sediments of the Gulf of Finland, Baltic Sea. Mar. Environ. Res. 113, 153–163 (2016).
Rowden, A., Jago, C. & Jones, S. Influence of benthic macrofauna on the geotechnical and geophysical properties of surface sediments, North Sea. Continental Shelf Research 18, 1347–1363 (1998).
Bollmann, A., French, E. & Laanbroek, H. J. Isolation, cultivation, and characterization of ammonia-oxidizing bacteria and archaea adapted to low ammonium concentrations. Methods Enzymol. 486, 55–88 (2010).
Ginestet, P., Audic, J.-M., Urbain, V. & Block, J.-C. Estimation of Nitrifying Bacterial Activities by Measuring Oxygen Uptake in the Presence of the Metabolic Inhibitors Allylthiourea and Azide. Appl. Environ. Microbiol. 64, 2266–2268 (1998).
Smith, C. J. et al. Diversity and Abundance of Nitrate Reductase Genes (narG and napA), Nitrite Reductase Genes (nirS and nrfA), and Their Transcripts in Estuarine Sediments. Appl. Environ. Microbiol. 73, 3612–3622 (2007).
Smith, C. J. & Osborn, A. M. Advantages and limitations of quantitative PCR (Q-PCR) -based approaches in microbial ecology. FEMS Microbiol. Ecol. 67, 6–20 (2009).
Smith, C. J. et al. T-Align, a web-based tool for comparison of multiple terminal restriction fragment length polymorphism profiles. FEMS Microbiol. Ecol. 54, 375–80 (2005).
Altschul, S. F., Gish, W., Miller, W., Myers, E. W. & Lipman, D. J. Basic local alignment search tool. J. Mol. Biol. 215, 403–410 (1990).
Hall, T. BioEdit: a user-friendly biological sequence alignment editor and analysis program for Windows 95/98/NT. Nucleic Acids Symp. Ser. 41, 95–98 (1999).
Schloss, P. D. & Handelsman, J. Introducing DOTUR, a Computer Program for Defining Opera: onal Taxonomic Units and Es: ma: ng Species Richness. Appl. Environ. Microbiol. 71, 1501–1506 (2005).
Felsenstein, J. PHYLIP—phylogeny inference package. Cladistics 5, 164–166 (1989).
Saitou N, N. M. The Neighbor-joining Method: A New Method for Reconstructing Phylogenetic Trees’. Mol. Biol. Evol. 4, 406–425 (1987).
Kimura, M. The neutral theory of molecular evolution: a review of recent evidence. Jpn. J. Genet. 66, 367–86 (1991).
Tamura, K., Stecher, G., Peterson, D., Filipski, A. & Kumar, S. MEGA6: Molecular Evolutionary Genetics Analysis version 6.0. Mol. Biol. Evol. 30, 2725–9 (2013).
BARTLETT, M. S. The use of transformation. (Biometric Bull, 1947).
Dunn, O. J. Multiple Comparisons Among Means. J. Am. Stat. Assoc. 56, 52–64 (1961).
Acknowledgements
This publication has emanated from research conducted with the financial support of Science Foundation Ireland & the Marie-Curie Action COFUND under Grant Number 11/SIRG/B2159 awarded to CJS. We thank Enrico Tatti for training in canoco analysis and review of the manuscript.
Author information
Authors and Affiliations
Contributions
A.D., L.Z. & C.J.S. designed the study. A.D. and L.Z. conducted field work. A.D. performed the laboratory and data analysis. A.D. & C.J.S. wrote the paper. All authors reviewed the manuscript.
Corresponding author
Ethics declarations
Competing Interests
The authors declare that they have no competing interests.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Duff, A.M., Zhang, LM. & Smith, C.J. Small-scale variation of ammonia oxidisers within intertidal sediments dominated by ammonia-oxidising bacteria Nitrosomonas sp. amoA genes and transcripts. Sci Rep 7, 13200 (2017). https://doi.org/10.1038/s41598-017-13583-x
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-017-13583-x
- Springer Nature Limited
This article is cited by
-
The contribution of nirK gene-containing thaumarchaea to denitrification and N2O production across coastal sediment and terrestrial ecosystems
Journal of Soils and Sediments (2022)
-
Long-term greenhouse vegetable cultivation alters the community structures of soil ammonia oxidizers
Journal of Soils and Sediments (2019)