Abstract
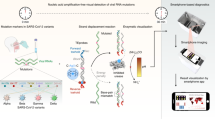
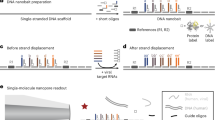
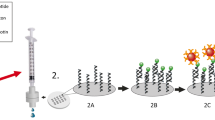
The evolution and mutation of severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) are urgent concerns as they pose the risk of vaccine failure and increased viral transmission. However, affordable and scalable tools allowing rapid identification of SARS-CoV-2 variants are not readily available, which impedes diagnosis and epidemiological surveillance. Here we present a colorimetric nucleic acid assay named MARVE (multiplexed, preamplification-free, single-nucleotide-resolved viral evolution) that is convenient to perform and yields single-nucleotide resolution. The assay integrates nucleic acid strand displacement reactions with enzymatic amplification to colorimetrically sense viral RNA using a metal ion-incorporated DNA probe (TEprobe). We provide detailed guidelines to design TEprobes for discriminating single-nucleotide variations in viral RNAs, and to fabricate a test paper for the detection of SARS-CoV-2 variants of concern. Compared with other nucleic acid assays, our assay is preamplification-free, single-nucleotide-resolvable and results are visible via a color change. Besides, it is smartphone readable, multiplexed, quick and cheap ($0.30 per test). The protocol takes ~2 h to complete, from the design and preparation of the DNA probes and test papers (~1 h) to the detection of SARS-CoV-2 or its variants (30–45 min). The design of the TEprobes requires basic knowledge of molecular biology and familiarity with NUPACK or the Python programming language. The fabrication of the origami papers requires access to a wax printer using the CAD and PDF files provided or requires users to be familiar with AutoCAD to design new origami papers. The protocol is also applicable for designing assays to detect other pathogens and their variants.
Key points
-
MARVE is a paper-based, preamplification-free assay for the detection of viral variants using toehold exchange (TE) DNA probes. The TEprobe binds to the target viral RNA and starts an enzymatic reaction, which results in low-pH conditions and a color change visible to the eye.
-
The high specificity of a nucleic acid test, combined with low cost and ease of use, makes MARVE suitable for home testing.













Similar content being viewed by others
Data availability
The data supporting the findings of this study are available within the source data and the supporting primary research papers16,17. Source data are provided with this paper.
Code availability
The mobile web apps for COVID-19 self-testing and viral contamination monitoring in food can be visited at http://47.109.38.99/origami1/ or at http://47.109.38.99/origami2/, respectively. The code for designing TEprobes is available on GitHub at https://github.com/Nelson233/TEprobe-design. The code for the two web apps is provided on GitHub at https://github.com/Nelson233/origami1 and https://github.com/Nelson233/origami2, respectively. The code for image analysis using MATLAB software is available on GitHub at https://github.com/Nelson233/MARVEL.
References
Carabelli, A. M. et al. SARS-CoV-2 variant biology: immune escape, transmission and fitness. Nat. Rev. Microbiol. 21, 162–177 (2023).
Amicone, M. et al. Mutation rate of SARS-CoV-2 and emergence of mutators during experimental evolution. Evol. Med. Public Health 10, 142–155 (2022).
Meng, B. et al. Altered TMPRSS2 usage by SARS-CoV-2 Omicron impacts infectivity and fusogenicity. Nature 603, 706–714 (2022).
Escalera, A. et al. Mutations in SARS-CoV-2 variants of concern link to increased spike cleavage and virus transmission. Cell Host Microbe 30, 373–387.e377 (2022).
DeGrace, M. M. et al. Defining the risk of SARS-CoV-2 variants on immune protection. Nature 605, 640–652 (2022).
Tomalka, J. A., Suthar, M. S., Deeks, S. G. & Sekaly, R. P. Fighting the SARS-CoV-2 pandemic requires a global approach to understanding the heterogeneity of vaccine responses. Nat. Immunol. 23, 360–370 (2022).
Garcia-Beltran, W. F. et al. Multiple SARS-CoV-2 variants escape neutralization by vaccine-induced humoral immunity. Cell 184, 2372–2383.e2379 (2021).
Petherick, A. Developing antibody tests for SARS-CoV-2. Lancet 395, 1101–1102 (2020).
Peeling, R. W., Heymann, D. L., Teo, Y.-Y. & Garcia, P. J. Diagnostics for COVID-19: moving from pandemic response to control. Lancet 399, 757–768 (2022).
Vogels, C. B. F. et al. Analytical sensitivity and efficiency comparisons of SARS-CoV-2 RT–qPCR primer–probe sets. Nat. Microbiol. 5, 1299–1305 (2020).
Bokelmann, L. et al. Point-of-care bulk testing for SARS-CoV-2 by combining hybridization capture with improved colorimetric LAMP. Nat. Commun. 12, 1467 (2021).
Qian, J. et al. An enhanced isothermal amplification assay for viral detection. Nat. Commun. 11, 5920 (2020).
Osório, N. S. & Correia-Neves, M. Implication of SARS-CoV-2 evolution in the sensitivity of RT–qPCR diagnostic assays. Lancet Infect. Dis. 21, 166–167 (2021).
Robishaw, J. D. et al. Genomic surveillance to combat COVID-19: challenges and opportunities. Lancet Microbe 2, e481–e484 (2021).
Li, P., de Vries, A. C., Kamar, N., Peppelenbosch, M. P. & Pan, Q. Monitoring and managing SARS-CoV-2 evolution in immunocompromised populations. Lancet Microbe 3, e325–e326 (2022).
Zhang, T. et al. A paper-based assay for the colorimetric detection of SARS-CoV-2 variants at single-nucleotide resolution. Nat. Biomed. Eng. 6, 957–967 (2022).
Zhang, T. et al. Precise in-field molecular diagnostics of crop diseases by smartphone-based mutation-resolved pathogenic RNA analysis. Nat. Commun. 14, 4327 (2023).
Dao Thi, V. L. et al. A colorimetric RT–LAMP assay and LAMP-sequencing for detecting SARS-CoV-2 RNA in clinical samples. Sci. Transl. Med. 12, eabc7075 (2020).
Wu, Q. et al. INSIGHT: a population-scale COVID-19 testing strategy combining point-of-care diagnosis with centralized high-throughput sequencing. Sci. Adv. 7, eabe5054 (2021).
Yelagandula, R. et al. Multiplexed detection of SARS-CoV-2 and other respiratory infections in high throughput by SARSeq. Nat. Commun. 12, 3132 (2021).
Garg, A. et al. Evaluation of seven commercial RT–PCR kits for COVID-19 testing in pooled clinical specimens. J. Med. Virol. 93, 2281–2286 (2021).
Panpradist, N. et al. Harmony COVID-19: a ready-to-use kit, low-cost detector, and smartphone app for point-of-care SARS-CoV-2 RNA detection. Sci. Adv. 7, eabj1281 (2021).
Alafeef, M., Moitra, P., Dighe, K. & Pan, D. RNA-extraction-free nano-amplified colorimetric test for point-of-care clinical diagnosis of COVID-19. Nat. Protoc. 16, 3141–3162 (2021).
Mohammadniaei, M. et al. A non-enzymatic, isothermal strand displacement and amplification assay for rapid detection of SARS-CoV-2 RNA. Nat. Commun. 12, 5089 (2021).
Broughton, J. P. et al. CRISPR–Cas12-based detection of SARS-CoV-2. Nat. Biotechnol. 38, 870–874 (2020).
Casati, B. et al. Rapid, adaptable and sensitive Cas13-based COVID-19 diagnostics using ADESSO. Nat. Commun. 13, 3308 (2022).
Patchsung, M. et al. Clinical validation of a Cas13-based assay for the detection of SARS-CoV-2 RNA. Nat. Biomed. Eng. 4, 1140–1149 (2020).
Joung, J. et al. Detection of SARS-CoV-2 with SHERLOCK one-pot testing. N. Engl. J. Med. 383, 1492–1494 (2020).
Xun, G., Lane, S. T., Petrov, V. A., Pepa, B. E. & Zhao, H. A rapid, accurate, scalable, and portable testing system for COVID-19 diagnosis. Nat. Commun. 12, 2905 (2021).
Ding, X. et al. Ultrasensitive and visual detection of SARS-CoV-2 using all-in-one dual CRISPR–Cas12a assay. Nat. Commun. 11, 4711 (2020).
Chandrasekaran, S. S. et al. Rapid detection of SARS-CoV-2 RNA in saliva via Cas13. Nat. Biomed. Eng. 6, 944–956 (2022).
De Puig, H. et al. Minimally instrumented SHERLOCK (miSHERLOCK) for CRISPR-based point-of-care diagnosis of SARS-CoV-2 and emerging variants. Sci. Adv. 7, eabh2944 (2021).
Welch, N. L. et al. Multiplexed CRISPR-based microfluidic platform for clinical testing of respiratory viruses and identification of SARS-CoV-2 variants. Nat. Med. 28, 1083–1094 (2022).
Arizti-Sanz, J. et al. Simplified Cas13-based assays for the fast identification of SARS-CoV-2 and its variants. Nat. Biomed. Eng. 6, 932–943 (2022).
Metsky, H. C. et al. Designing sensitive viral diagnostics with machine learning. Nat. Biotechnol. 40, 1123–1131 (2022).
Fozouni, P. et al. Amplification-free detection of SARS-CoV-2 with CRISPR–Cas13a and mobile phone microscopy. Cell 184, 323–333.e329 (2021).
Wang, D. et al. Rapid lateral flow immunoassay for the fluorescence detection of SARS-CoV-2 RNA. Nat. Biomed. Eng. 4, 1150–1158 (2020).
Phillips, E. A. et al. Detection of viral RNAs at ambient temperature via reporter proteins produced through the target-splinted ligation of DNA probes. Nat. Biomed. Eng. https://doi.org/10.1038/s41551-41023-01028-y (2023).
Puhach, O., Meyer, B. & Eckerle, I. SARS-CoV-2 viral load and shedding kinetics. Nat. Rev. Microbiol. 21, 147–161 (2023).
Yuasa, S. et al. Viral load of SARS-CoV-2 Omicron is not high despite its high infectivity. J. Med. Virol. 94, 5543–5546 (2022).
La Scola, B. et al. Viral RNA load as determined by cell culture as a management tool for discharge of SARS-CoV-2 patients from infectious disease wards. Eur. J. Clin. Microbiol. 39, 1059–1061 (2020).
Gallichotte, E. N. et al. Early adoption of longitudinal surveillance for SARS-CoV-2 among staff in long-term care facilities: prevalence, virologic and sequence analysis. Microbiol. Spectr. 9, e01003–e01021 (2021).
Wölfel, R. et al. Virological assessment of hospitalized patients with COVID-2019. Nature 581, 465–469 (2020).
Fu, E. & Downs, C. Progress in the development and integration of fluid flow control tools in paper microfluidics. Lab Chip 17, 614–628 (2017).
Wu, K. & Green, A. A. Sensitive detection of SARS-CoV-2 on paper. Nat. Biomed. Eng. 6, 928–929 (2022).
Sakuma, T., Barry, Michael, A. & Ikeda, Y. Lentiviral vectors: basic to translational. Biochem. J. 443, 603–618 (2012).
Myhrvold, C. et al. Field-deployable viral diagnostics using CRISPR–Cas13. Science 360, 444–448 (2018).
Weickmann, J. L. & Glitz, D. G. Human ribonucleases. Quantitation of pancreatic-like enzymes in serum, urine, and organ preparations. J. Biol. Chem. 257, 8705–8710 (1982).
Laue, T., Emmerich, P. & Schmitz, H. Detection of dengue virus RNA in patients after primary or secondary dengue infection by using the TaqMan automated amplification system. J. Clin. Microbiol. 37, 2543–2547 (1999).
Simmel, F. C., Yurke, B. & Singh, H. R. Principles and applications of nucleic acid strand displacement reactions. Chem. Rev. 119, 6326–6369 (2019).
Chen, S. X. & Seelig, G. An engineered kinetic amplification mechanism for single nucleotide variant discrimination by DNA hybridization probes. J. Am. Chem. Soc. 138, 5076–5086 (2016).
Martinez, A. W., Phillips, S. T., Butte, M. J. & Whitesides, G. M. Patterned paper as a platform for inexpensive, low-volume, portable bioassays. Angew. Chem. Int. Ed. 46, 1318–1320 (2007).
Martinez, A. W., Phillips, S. T., Wiley, B. J., Gupta, M. & Whitesides, G. M. FLASH: a rapid method for prototyping paper-based microfluidic devices. Lab Chip 8, 2146–2150 (2008).
Dungchai, W., Chailapakul, O. & Henry, C. S. A low-cost, simple, and rapid fabrication method for paper-based microfluidics using wax screen-printing. Analyst 136, 77–82 (2011).
Yamada, K., Henares, T. G., Suzuki, K. & Citterio, D. Paper-based inkjet-printed microfluidic analytical devices. Angew. Chem. Int. Ed. 54, 5294–5310 (2015).
Apilux, A., Ukita, Y., Chikae, M., Chailapakul, O. & Takamura, Y. Development of automated paper-based devices for sequential multistep sandwich enzyme-linked immunosorbent assays using inkjet printing. Lab Chip 13, 126–135 (2013).
Ghosh, R., Gopalakrishnan, S., Savitha, R., Renganathan, T. & Pushpavanam, S. Fabrication of laser printed microfluidic paper-based analytical devices (LP-µPADs) for point-of-care applications. Sci. Rep. 9, 7896 (2019).
Almeida, M. I. G. S., Jayawardane, B. M., Kolev, S. D. & McKelvie, I. D. Developments of microfluidic paper-based analytical devices (μPADs) for water analysis: a review. Talanta 177, 176–190 (2018).
Smereka, M. & Dulęba, I. Circular object detection using a modified hough transform. Int. J. Appl. Math. Comput. Sci. 18, 85–91 (2008).
Hoo, Z. H., Candlish, J. & Teare, D. What is an ROC curve? Emerg. Med. J. 34, 357–359 (2017).
Acknowledgements
We thank all authors of the primary research paper. The work was funded by the National Key Research and Development Program of China (2023YFB3208302 and 2021YFA1200104), the New Cornerstone Investigator Program, National Natural Science Foundation of China (22074100 and 22027807), the Strategic Priority Research Program of the Chinese Academy of Sciences (XDB36000000) and Tsinghua-Vanke Special Fund for Public Health and Health Discipline Development (2022Z82WKJ003).
Author information
Authors and Affiliations
Contributions
J.L. and R.D. conceived and designed the protocol. R.D. and T.Z. developed the protocol. T.Z., R.D., Y.W. and X.T. performed the experiments and analyzed the data. R.D. and T.Z. wrote the manuscript. J.L. edited the manuscript. All authors read, commented on and accepted the final manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Protocols thanks Changqing Yi, and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Related links
Key references that have used this protocol
Zhang, T. et al. Nat. Biomed. Eng. 6, 957–967 (2022): https://doi.org/10.1038/s41551-022-00907-0
Zhang, T. et al. Nat. Commun. 14, 4327 (2023): https://doi.org/10.1038/s41467-023-39952-x
Supplementary information
Supplementary Information
Supplementary Figs. 1–7 and Tables 1 and 2.
Supplementary Video 1
The illustration of paper folding using the enclosure.
Supplementary Data 1
The original CAD file of eight-site origami paper.
Supplementary Data 2
The PDF file of eight-site origami paper.
Supplementary Data 3
The original CAD file of four-site origami paper
Supplementary Data 4
The PDF file of four-site origami paper.
Supplementary Data 5
The original file of the enclosure.
Supplementary Data 6
The original CAD file of 100-site origami paper.
Supplementary Data 7
The PDF file of 100-site origami paper.
Source data
Source Data Fig. 10
Statistical source data.
Source Data Fig. 11
Statistical source data.
Source Data Fig. 12
Statistical source data.
Source Data Fig. 13
Statistical source data.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Zhang, T., Wang, Y., Teng, X. et al. Preamplification-free viral RNA diagnostics with single-nucleotide resolution using MARVE, an origami paper-based colorimetric nucleic acid test. Nat Protoc (2024). https://doi.org/10.1038/s41596-024-01022-x
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41596-024-01022-x
- Springer Nature Limited