Abstract
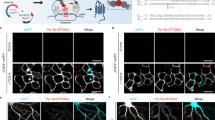
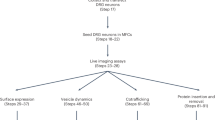
Population behavior of signaling molecules on the cell surface is key to their adaptive function. Live imaging of proteins tagged with fluorescent molecules has been an essential tool in understanding this behavior. Typically, genetic or chemical tags are used to target molecules present throughout the cell, whereas antibody-based tags label the externally exposed molecular domains only. Both approaches could potentially overlook the intricate process of in–out membrane recycling in which target molecules appear or disappear on the cell surface. This limitation is overcome by using a pH-sensitive fluorescent tag, such as Super-Ecliptic pHluorin (SEP), because its emission depends on whether it resides inside or outside the cell. Here we focus on the main glial glutamate transporter GLT1 and describe a genetic design that equips GLT1 molecules with SEP without interfering with the transporter’s main function. Expressing GLT1-SEP in astroglia in cultures or in hippocampal slices enables monitoring of the real-time dynamics of the cell-surface and cytosolic fractions of the transporter in living cells. Whole-cell fluorescence recovery after photobleaching and quantitative image-kinetic analysis of the resulting time-lapse images enables assessment of the rate of GLT1-SEP recycling on the cell surface, a fundamental trafficking parameter unattainable previously. The present protocol takes 15–20 d to set up cell preparations, and 2–3 d to carry out live cell experiments and data analyses. The protocol can be adapted to study different membrane molecules of interest, particularly those proteins whose lifetime on the cell surface is critical to their adaptive function.





Similar content being viewed by others
Data availability
The original experimental data are available as Source Data files in the supporting primary research article1.
References
Michaluk, P., Heller, J. P. & Rusakov, D. A. Rapid recycling of glutamate transporters on the astroglial surface. eLife 10, 64714 (2021).
Zhang, J., Campbell, R. E., Ting, A. Y. & Tsien, R. Y. Creating new fluorescent probes for cell biology. Nat. Rev. Mol. Cell Biol. 3, 906–918 (2002).
Lippincott-Schwartz, J. & Patterson, G. H. Development and use of fluorescent protein markers in living cells. Science 300, 87–91 (2003).
Axelrod, D., Koppel, D. E., Schlessinger, J., Elson, E. & Webb, W. W. Mobility measurement by analysis of fluorescence photobleaching recovery kinetics. Biophys. J. 16, 1055–1069 (1976).
Lippincott-Schwartz, J., Altan-Bonnet N. & Patterson G. H. Photobleaching and photoactivation: following protein dynamics in living cells. Nat. Cell Biol. Suppl, S7–14 (2003).
Lambert, N. A. Uncoupling diffusion and binding in FRAP experiments. Nat. Methods 6, 183–184 (2009).
Choquet, D. & Triller, A. The role of receptor diffusion in the organization of the postsynaptic membrane. Nat. Rev. Neurosci. 4, 251–265 (2003).
Bannai, H., Levi, S., Schweizer, C., Dahan, M. & Triller, A. Imaging the lateral diffusion of membrane molecules with quantum dots. Nat. Protoc. 1, 2628–2634 (2006).
Betzig, E. et al. Imaging intracellular fluorescent proteins at nanometer resolution. Science 313, 1642–1645 (2006).
Rossier, O. et al. Integrins β1 and β3 exhibit distinct dynamic nanoscale organizations inside focal adhesions. Nat. Cell Biol. 14, 1057–1067 (2012).
Manley, S. et al. High-density mapping of single-molecule trajectories with photoactivated localization microscopy. Nat. Methods 5, 155–157 (2008).
Sydor, A. M., Czymmek, K. J., Puchner, E. M. & Mennella, V. Super-resolution microscopy: from single molecules to supramolecular assemblies. Trends Cell Biol. 25, 730–748 (2015).
Giannone, G. et al. Dynamic superresolution imaging of endogenous proteins on living cells at ultra-high density. Biophys. J. 99, 1303–1310 (2010).
Carion, O., Mahler, B., Pons, T. & Dubertret, B. Synthesis, encapsulation, purification and coupling of single quantum dots in phospholipid micelles for their use in cellular and in vivo imaging. Nat. Protoc. 2, 2383–2390 (2007).
Livet, J. et al. Transgenic strategies for combinatorial expression of fluorescent proteins in the nervous system. Nature 450, 56–62 (2007).
Barnes, S. A. et al. Convergence of hippocampal pathophysiology in Syngap+/− and Fmr1−/y mice. J. Neurosci. 35, 15073–15081 (2015).
Politi, A. Z. et al. Quantitative mapping of fluorescently tagged cellular proteins using FCS-calibrated four-dimensional imaging. Nat. Protoc. 13, 1445–1464 (2018).
Heal, W. P., Wright, M. H., Thinon, E. & Tate, E. W. Multifunctional protein labeling via enzymatic N-terminal tagging and elaboration by click chemistry. Nat. Protoc. 7, 105–117 (2011).
Kamiyama, D. et al. Versatile protein tagging in cells with split fluorescent protein. Nat. Commun. 7, 11046 (2016).
Miesenbock, G., De Angelis, D. A. & Rothman, J. E. Visualizing secretion and synaptic transmission with pH-sensitive green fluorescent proteins. Nature 394, 192–195 (1998).
Sankaranarayanan, S., De Angelis, D., Rothman, J. E. & Ryan, T. A. The use of pHluorins for optical measurements of presynaptic activity. Biophys. J. 79, 2199–2208 (2000).
Sankaranarayanan, S. & Ryan, T. A. Calcium accelerates endocytosis of vSNAREs at hippocampal synapses. Nat. Neurosci. 4, 129–136 (2001).
Matz, J., Gilyan, A., Kolar, A., McCarvill, T. & Krueger, S. R. Rapid structural alterations of the active zone lead to sustained changes in neurotransmitter release. Proc. Natl Acad. Sci. USA 107, 8836–8841 (2010).
Ashby, M. C. et al. Removal of AMPA receptors (AMPARs) from synapses is preceded by transient endocytosis of extrasynaptic AMPARs. J. Neurosci. 24, 5172–5176 (2004).
Ashby, M. C., Maier, S. R., Nishimune, A. & Henley, J. M. Lateral diffusion drives constitutive exchange of AMPA receptors at dendritic spines and is regulated by spine morphology. J. Neurosci. 26, 7046–7055 (2006).
Tanaka, K. et al. Epilepsy and exacerbation of brain injury in mice lacking the glutamate transporter GLT-1. Science 276, 1699–1702 (1997).
Danbolt, N. C. Glutamate uptake. Progr. Neurobiol. 65, 1–105 (2001).
Lehre, K. P. & Danbolt, N. C. The number of glutamate transporter subtype molecules at glutamatergic synapses: chemical and stereological quantification in young adult rat brain. J. Neurosci. 18, 8751–8757 (1998).
Diamond, J. S. & Jahr, C. E. Transporters buffer synaptically released glutamate on a submillisecond time scale. J. Neurosci. 17, 4672–4687 (1997).
Wadiche, J. I., Arriza, J. L., Amara, S. G. & Kavanaugh, M. P. Kinetics of a human glutamate transporter. Neuron 14, 1019–1027 (1995).
Lozovaya, N. A., Kopanitsa, M. V., Boychuk, Y. A. & Krishtal, O. A. Enhancement of glutamate release uncovers spillover-mediated transmission by N-methyl-d-aspartate receptors in the rat hippocampus. Neuroscience 91, 1321–1330 (1999).
Scimemi, A., Fine, A., Kullmann, D. M. & Rusakov, D. A. NR2B-containing receptors mediate cross talk among hippocampal synapses. J. Neurosci. 24, 4767–4777 (2004).
Zheng, K., Scimemi, A. & Rusakov, D. A. Receptor actions of synaptically released glutamate: the role of transporters on the scale from nanometers to microns. Biophys. J. 95, 4584–4596 (2008).
Henneberger, C. et al. LTP induction boosts glutamate spillover by driving withdrawal of perisynaptic astroglia. Neuron 108, 919–936 e911 (2020).
Maragakis, N. J. & Rothstein, J. D. Glutamate transporters: animal models to neurologic disease. Neurobiol. Dis. 15, 461–473 (2004).
Fontana, A. C. Current approaches to enhance glutamate transporter function and expression. J. Neurochem. 134, 982–1007 (2015).
Kruyer, A., Scofield, M. D., Wood, D., Reissner, K. J. & Kalivas, P. W. Heroin cue-evoked astrocytic structural plasticity at nucleus accumbens synapses inhibits heroin seeking. Biol. Psychiat. 86, 811–819 (2019).
Al Awabdh, S. et al. Neuronal activity mediated regulation of glutamate transporter GLT-1 surface diffusion in rat astrocytes in dissociated and slice cultures. Glia 64, 1252–1264 (2016).
Murphy-Royal, C. et al. Surface diffusion of astrocytic glutamate transporters shapes synaptic transmission. Nat. Neurosci. 18, 219–226 (2015).
Sylantyev, S. & Rusakov, D. A. Sub-millisecond ligand probing of cell receptors with multiple solution exchange. Nat. Protoc. 8, 1299–1306 (2013).
Shen, Y., Rosendale, M., Campbell, R. E. & Perrais, D. pHuji, a pH-sensitive red fluorescent protein for imaging of exo- and endocytosis. J. Cell Biol. 207, 419–432 (2014).
Liu, A. Y. et al. pHmScarlet is a pH-sensitive red fluorescent protein to monitor exocytosis docking and fusion steps. Nat. Commun. 12, 1413 (2021).
Lazarenko, R. M., DelBove, C. E., Strothman, C. E. & Zhang, Q. Ammonium chloride alters neuronal excitability and synaptic vesicle release. Sci. Rep. 7, 5061 (2017).
Reits, E. A. J. & Neefjes, J. J. From fixed to FRAP: measuring protein mobility and activity in living cells. Nat. Cell Biol. 3, E145–E147 (2001).
Jensen, T. P., Zheng, K., Tyurikova, O., Reynolds, J. P. & Rusakov, D. A. Monitoring single-synapse glutamate release and presynaptic calcium concentration in organised brain tissue. Cell Calcium 64, 102–108 (2017).
Jensen, T. P. et al. Multiplex imaging relates quantal glutamate release to presynaptic Ca2+ homeostasis at multiple synapses in situ. Nat. Commun. 10, 1414 (2019).
Gabriel, L., Stevens Z. & Melikian H. Measuring plasma membrane protein endocytic rates by reversible biotinylation. J. Vis. Exp. https://doi.org/10.3791/1669 (2009).
Turvy, D. N. & Blum J. S. Biotin labeling and quantitation of cell-surface proteins. Curr. Protoc. Immunol. Chapter 18, Unit 18 17 (2001).
Holton, K. L., Loder, M. K. & Melikian, H. E. Nonclassical, distinct endocytic signals dictate constitutive and PKC-regulated neurotransmitter transporter internalization. Nat. Neurosci. 8, 881–888 (2005).
Scott, D. B., Michailidis, I., Mu, Y., Logothetis, D. & Ehlers, M. D. Endocytosis and degradative sorting of NMDA receptors by conserved membrane-proximal signals. J. Neurosci. 24, 7096–7109 (2004).
Rizzolio, S. & Tamagnone, L. Antibody-feeding assay: a method to track the internalization of neuropilin-1 and other cell surface receptors. Methods Mol. Biol. 1493, 311–319 (2017).
Chiu, A. M., Barse L., Hubalkova P. & Sanz-Clemente A. An antibody feeding approach to study glutamate receptor trafficking in dissociated primary hippocampal cultures. J. Vis. Exp. https://doi.org/10.3791/59982 (2019).
Kim, S., Bell, K., Mousa, S. A. & Varner, J. A. Regulation of angiogenesis in vivo by ligation of integrin α5β1 with the central cell-binding domain of fibronectin. Am. J. Pathol. 156, 1345–1362 (2000).
Zilberstein, A., Snider, M. D., Porter, M. & Lodish, H. F. Mutants of vesicular stomatitis virus blocked at different stages in maturation of the viral glycoprotein. Cell 21, 417–427 (1980).
Pepperkok, R. et al. Imaging platforms for measurement of membrane trafficking. Methods Enzymol. 404, 8–18 (2005).
Passafaro, M., Piech, V. & Sheng, M. Subunit-specific temporal and spatial patterns of AMPA receptor exocytosis in hippocampal neurons. Nat. Neurosci. 4, 917–926 (2001).
Hein, L., Ishii, K., Coughlin, S. R. & Kobilka, B. K. Intracellular targeting and trafficking of thrombin receptors. A novel mechanism for resensitization of a G protein-coupled receptor. J. Biol. Chem. 269, 27719–27726 (1994).
Hopp, T. P. et al. A short polypeptide marker sequence useful for recombinant protein identification and purification. Nat. Biotechnol. 6, 1204–1210 (1988).
Leake, M. C. et al. Stoichiometry and turnover in single, functioning membrane protein complexes. Nature 443, 355–358 (2006).
Willig, K. I., Rizzoli, S. O., Westphal, V., Jahn, R. & Hell, S. W. STED microscopy reveals that synaptotagmin remains clustered after synaptic vesicle exocytosis. Nature 440, 935–939 (2006).
Luo, N., Yan, A. & Yang, Z. B. Measuring exocytosis rate using corrected fluorescence recovery after photoconversion. Traffic 17, 554–564 (2016).
Chen, X., Zaro, J. L. & Shen, W. C. Fusion protein linkers: property, design and functionality. Adv. Drug Deliv. Rev. 65, 1357–1369 (2013).
Beaudoin, G. M. 3rd et al. Culturing pyramidal neurons from the early postnatal mouse hippocampus and cortex. Nat. Protoc. 7, 1741–1754 (2012).
Kaech, S. & Banker, G. Culturing hippocampal neurons. Nat. Protoc. 1, 2406–2415 (2006).
Lein, P. J., Barnhart, C. D. & Pessah, I. N. Acute hippocampal slice preparation and hippocampal slice cultures. Methods Mol. Biol. 758, 115–134 (2011).
Jiang, M. & Chen, G. High Ca2+-phosphate transfection efficiency in low-density neuronal cultures. Nat. Protoc. 1, 695–700 (2006).
Chen, C. C. et al. Patch-clamp technique to characterize ion channels in enlarged individual endolysosomes. Nat. Protoc. 12, 1639–1658 (2017).
Rathje, M. et al. AMPA receptor pHluorin-GluA2 reports NMDA receptor-induced intracellular acidification in hippocampal neurons. Proc. Natl Acad. Sci. USA 110, 14426–14431 (2013).
Savtchenko, L. P. et al. Disentangling astroglial physiology with a realistic cell model in silico. Nat. Commun. 9, 3554 (2018).
Drobizhev, M., Makarov, N. S., Tillo, S. E., Hughes, T. E. & Rebane, A. Two-photon absorption properties of fluorescent proteins. Nat. Methods 8, 393–399 (2011).
Hafner, G. et al. Mapping brain-wide afferent inputs of parvalbumin-expressing GABAergic neurons in barrel cortex reveals local and long-range circuit motifs. Cell Rep. 28, 3450–3461 e3458 (2019).
Valenti, M. T. et al. The effect of bisphosphonates on gene expression: GAPDH as a housekeeping or a new target gene? BMC Cancer 6, 49 (2006).
Nifosi, R. & Luo, Y. Predictions of novel two-photon absorption bands in fluorescent proteins. J. Phys. Chem. B 111, 14043–14050 (2007).
Rueden, C. T. et al. ImageJ2: ImageJ for the next generation of scientific image data. BMC Bioinformatics 18, 529 (2017).
Acknowledgements
The study was supported by: Wellcome Trust (212251_Z_18_Z), MRC (MR/W019752/1), ERC (323113) and European Commission NEUROTWIN (857562) to D.A.R.; National Science Centre Poland (2017/26/D/NZ3/01017) to P.M.
Author information
Authors and Affiliations
Contributions
P.M. suggested and implemented genetic designs, planned and carried out experiments, and analyzed the results; D.A.R. narrated the study, designed imaging methods and performed theoretical data analyses; D.A.R. and P.M. wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Protocols thanks Michael Ashby, Alex Verkhratsky and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Related links
Key reference using this protocol
Michaluk, P. et al. eLife 10, 64714 (2021): https://doi.org/10.7554/eLife.64714
Rights and permissions
Springer Nature or its licensor holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Michaluk, P., Rusakov, D.A. Monitoring cell membrane recycling dynamics of proteins using whole-cell fluorescence recovery after photobleaching of pH-sensitive genetic tags. Nat Protoc 17, 3056–3079 (2022). https://doi.org/10.1038/s41596-022-00732-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41596-022-00732-4
- Springer Nature Limited
This article is cited by
-
Spatially mapping the diffusivity of proteins in live cells based on cumulative area analysis
Science China Chemistry (2023)