Abstract
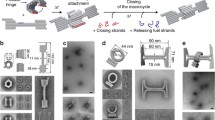
Mechanically interlocked DNA nanostructures are useful as flexible entities for operating DNA-based nanomachines. Interlocked structures made of double-stranded (ds) DNA components can be constructed by irreversibly threading them through one another to mechanically link them. The interlocked components thus remain bound to one another while still permitting large-amplitude motion about the mechanical bond. The construction of interlocked dsDNA architectures is challenging because it usually involves the synthesis and modification of small dsDNA nanocircles of various sizes, dependent on intrinsically curved DNA. Here we describe the design, generation, purification, and characterization of interlocked dsDNA structures such as catenanes, rotaxanes, and daisy-chain rotaxanes (DCRs). Their construction requires precise control of threading and hybridization of the interlocking components at each step during the assembly process. The protocol details the characterization of these nanostructures with gel electrophoresis and atomic force microscopy (AFM), including acquisition of high-resolution AFM images obtained in intermittent contact mode in liquid. Additional functionality can be conferred on the DNA architectures by incorporating proteins, molecular switches such as photo-switchable azobenzene derivatives, or fluorophores for studying their mechanical behavior by fluorescence quenching or fluorescent resonance energy transfer experiments. These modified interlocked DNA architectures provide access to more complex mechanical devices and nanomachines that can perform a variety of desired functions and operations. The assembly of catenanes can be completed in 2 d, and that of rotaxanes in 3 d. Addition of azobenzene functionality, fluorophores, anchor groups, or the site-specific linkage of proteins to the nanostructure can extend the time line.
















Similar content being viewed by others
Data availability
All data generated or analyzed during this study are included in this published article and its supplementary information files.
References
Krishnan, Y. & Simmel, F. C. Nucleic acid based molecular devices. Angew. Chem. Int. Ed. Engl. 50, 3124–3156 (2011).
Bath, J. & Turberfield, A. J. DNA nanomachines. Nat. Nanotechnol. 2, 275–284 (2007).
Heckel, A. & Famulok, M. Building objects from nucleic acids for a nanometer world. Biochimie 90, 1096–1107 (2008).
Seeman, N. C. Nanomaterials based on DNA. Annu. Rev. Biochem. 79, 65–87 (2010).
Endo, M. & Sugiyama, H. DNA origami nanomachines. Molecules 23, E1766 (2018).
Erbas-Cakmak, S., Leigh, D. A., McTernan, C. T. & Nussbaumer, A. L. Artificial molecular machines. Chem. Rev. 115, 10081–10206 (2015).
Jester, S. S. & Famulok, M. Mechanically interlocked DNA nanostructures for functional devices. Acc. Chem. Res. 47, 1700–1709 (2014).
Lu, C. H., Cecconello, A. & Willner, I. Recent advances in the synthesis and functions of reconfigurable interlocked DNA nanostructures. J. Am. Chem. Soc. 138, 5172–5185 (2016).
Valero, J., Lohmann, F. & Famulok, M. Interlocked DNA topologies for nanotechnology. Curr. Opin. Biotechnol. 48, 159–167 (2017).
Rasched, G. et al. DNA minicircles with gaps for versatile functionalization. Angew. Chem. Int. Ed. Engl. 47, 967–970 (2008).
Schmidt, T. L. et al. Polyamide struts for DNA architectures. Angew. Chem. Int. Ed. Engl. 46, 4382–4384 (2007).
Li, T., Lohmann, F. & Famulok, M. Interlocked DNA nanostructures controlled by a reversible logic circuit. Nat. Commun. 5, 4940 (2014).
Lohmann, F., Valero, J. & Famulok, M. A novel family of structurally stable double stranded DNA catenanes. Chem. Commun. 50, 6091–6093 (2014).
Ackermann, D., Jester, S. S. & Famulok, M. Design strategy for DNA rotaxanes with a mechanically reinforced PX100 axle. Angew. Chem. Int. Ed. Engl. 51, 6771–6775 (2012).
Ackermann, D. et al. A double-stranded DNA rotaxane. Nat. Nanotechnol. 5, 436–442 (2010).
Cecconello, A., Lu, C. H., Elbaz, J. & Willner, I. Au nanoparticle/DNA rotaxane hybrid nanostructures exhibiting switchable fluorescence properties. Nano Lett. 13, 6275–6280 (2013).
List, J., Falgenhauer, E., Kopperger, E., Pardatscher, G. & Simmel, F. C. Long-range movement of large mechanically interlocked DNA nanostructures. Nat. Commun. 7, 12414 (2016).
Lohmann, F., Weigandt, J., Valero, J. & Famulok, M. Logic gating by macrocycle displacement using a double-stranded DNA [3]rotaxane shuttle. Angew. Chem. Int. Ed. Engl. 53, 10372–10376 (2014).
Powell, J. T., Akhuetie-Oni, B. O., Zhang, Z. & Lin, C. DNA origami rotaxanes: tailored synthesis and controlled structure switching. Angew. Chem. Int. Ed. Engl. 55, 11412–11416 (2016).
Ulanovsky, L., Bodner, M., Trifonov, E. N. & Choder, M. Curved DNA: design, synthesis, and circularization. Proc. Natl. Acad. Sci. USA 83, 862–866 (1986).
Centola, M., Valero, J. & Famulok, M. Allosteric control of oxidative catalysis by a DNA rotaxane nanostructure. J. Am. Chem. Soc. 139, 16044–16047 (2017).
Lohmann, F., Ackermann, D. & Famulok, M. Reversible light switch for macrocycle mobility in a DNA rotaxane. J. Am. Chem. Soc. 134, 11884–11887 (2012).
Valero, J., Pal, N., Dhakal, S., Walter, N. G. & Famulok, M. A bio-hybrid DNA rotor-stator nanoengine that moves along predefined tracks. Nat. Nanotechnol. 13, 496–503 (2018).
Fang, L. et al. Mechanically bonded macromolecules. Chem. Soc. Rev. 39, 17–29 (2010).
Stoddart, J. F. The chemistry of the mechanical bond. Chem. Soc. Rev. 38, 1802–1820 (2009).
Sauvage, J. & Amabilino, D. The beauty of knots at the molecular level. Top. Curr. Chem 323, 107–125 (2012).
Schalley, C. A., Beizai, K. & Vögtle, F. On the way to rotaxane-based molecular motors: studies in molecular mobility and topological chirality. Acc. Chem. Res. 34, 465–476 (2001).
Gil-Ramirez, G., Leigh, D. A. & Stephens, A. J. Catenanes: fifty years of molecular links. Angew. Chem. Int. Ed. Engl. 54, 6110–6150 (2015).
Kay, E. R. & Leigh, D. A. Rise of the molecular machines. Angew. Chem. Int. Ed. Engl. 54, 10080–10088 (2015).
Feringa, B. L. The art of building small: from molecular switches to molecular motors. J. Org. Chem. 72, 6635–6652 (2007).
Sauvage, J. P. From chemical topology to molecular machines (nobel lecture). Angew. Chem. Int. Ed. Engl. 56, 11080–11093 (2017).
Stoddart, J. F. Mechanically interlocked molecules (MIMs)-molecular shuttles, switches, and machines (nobel lecture). Angew. Chem. Int. Ed. Engl. 56, 11094–11125 (2017).
Lu, C. H. et al. Switchable reconfiguration of a seven-ring interlocked DNA catenane nanostructure. Nano Lett. 15, 7133–7137 (2015).
Lu, C. H. et al. Switchable reconfiguration of an interlocked DNA olympiadane nanostructure. Angew. Chem. Int. Ed. Engl. 53, 7499–7503 (2014).
Lu, C. H., Cecconello, A., Elbaz, J., Credi, A. & Willner, I. A three-station DNA catenane rotary motor with controlled directionality. Nano Lett. 13, 2303–2308 (2013).
MacDonald, D., Herbert, K., Zhang, X., Pologruto, T. & Lu, P. Solution structure of an A-tract DNA bend. J. Mol. Biol. 306, 1081–1098 (2001).
Valero, J., Lohmann, F., Keppner, D. & Famulok, M. Single-stranded tile stoppers for interlocked DNA architectures. ChemBioChem 17, 1146–1149 (2016).
Weigandt, J., Chung, C.-L., Jester, S.-S. & Famulok, M. A daisy chain rotaxane interlocked DNA nanostructure. Angew. Chem. Int. Ed. Engl. 55, 5512–5516 (2016).
Haydell, M. W., Centola, M., Adam, V., Valero, J. & Famulok, M. Temporal and reversible control of a DNAzyme by orthogonal photoswitching. J. Am. Chem. Soc. 140, 16868–16872 (2018).
Škugor, M. et al. Orthogonally photocontrolled non-autonomous DNA walker. Angew. Chem. Intl. Edn. 58, 6948–6951 (2019).
Zadeh, J. N. et al. NUPACK: analysis and design of nucleic acid systems. J. Comput. Chem. 32, 170–173 (2011).
Brinkers, S., Dietrich, H. R. C., de Groote, F. H., Young, I. T. & Rieger, B. The persistence length of double stranded DNA determined using dark field tethered particle motion. J. Chem. Phys. 130, 215105 (2009).
Shen, Z., Yan, H., Wang, T. & Seeman, N. C. Paranemic crossover DNA: a generalized Holliday structure with applications in nanotechnology. J. Am. Chem. Soc. 126, 1666–1674 (2004).
Ackermann, D. & Famulok, M. Pseudo-complementary PNA actuators as reversible switches in dynamic DNA nanotechnology. Nucleic Acids Res 41, 4729–4739 (2013).
Rotzler, J. & Mayor, M. Molecular daisy chains. Chem. Soc. Rev. 42, 44–62 (2013).
Asanuma, H., Liang, X., Yoshida, T. & Komiyama, M. Photocontrol of DNA duplex formation by using azobenzene-bearing oligonucleotides. ChemBioChem 2, 39–44 (2001).
Adam, V. et al. Expanding the toolbox of photoswitches for DNA nanotechnology using arylazopyrazoles. Chemistry 24, 1062–1066 (2018).
Nishioka, H., Liang, X., Kashida, H. & Asanuma, H. 2′,6′-Dimethylazobenzene as an efficient and thermo-stable photo-regulator for the photoregulation of DNA hybridization. Chem. Commun. (Camb) 2007, 4354-4356 (2007).
Asanuma, H. et al. Synthesis of azobenzene-tethered DNA for reversible photo-regulation of DNA functions: hybridization and transcription. Nat. Protoc. 2, 203–212 (2007).
Daubendiek, S. L., Ryan, K. & Kool, E. T. Rolling-circle RNA synthesis: circular oligonucleotides as efficient substrates for T7 RNA polymerase. J. Am. Chem. Soc. 117, 7818–7819 (1995).
Nakata, E. et al. Zinc-finger proteins for site-specific protein positioning on DNA-origami structures. Angew. Chem. Int. Ed. Engl. 51, 2421–2424 (2012).
Acknowledgements
This work was made possible through funding from the European Research Council (ERC; grant no. 267173), the Max-Planck-Society (Max-Planck fellowship to M.F.), the Alexander von Humboldt Foundation (postdoctoral grants to J.V. and Y.M.), the German Academic Exchange Service (doctoral grant to M.Š.), and the China Scholarship Council CSC (doctoral scholarship to Z.Y.). M.F. extends his most sincere thanks to all the co-workers and collaborators who have contributed to this project over the past 10 years. Their names appear throughout the references. We also thank V. Adam for the synthesis of azobenzene derivatives.
Author information
Authors and Affiliations
Contributions
J.V., M.C., Y.M., M.Š., Z.Y., M.W.H., and M.F. conceived and designed the protocol and wrote the manuscript. J.V., M.C., Y.M., M.Š., Z.Y., M.W.H., and D.K. carried out the experiments.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information: Nature Protocols thanks Yamuna Krishnan, Francesco Ricci and other anonymous reviewer(s) for their contribution to the peer review of this work.
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Related links
Key references using this protocol
Rasched, G. et al. Angew. Chem. Int. Ed. Engl. 47, 967–970 (2008): https://onlinelibrary.wiley.com/doi/abs/10.1002/anie.200704004
Ackermann, D. et al. Nat. Nanotechnol. 5, 436–442 (2010): https://www.nature.com/articles/nnano.2010.65
Lohmann, F., Weigandt, J., Valero, J., & Famulok, M. Angew. Chem. Int. Ed. Engl. 53, 10372–10376 (2014): https://onlinelibrary.wiley.com/doi/pdf/10.1002/anie.201405447
Valero, J. Pal, N., Dhakal, S., Walter, N. G. & Famulok, M. Nat. Nanotechnol. 13, 496–503 (2018): https://www.nature.com/articles/s41565-018-0109-z
Supplementary Information
Supplementary Information
Supplementary Figures 1–25 and Supplementary Tables 1–20
Rights and permissions
About this article
Cite this article
Valero, J., Centola, M., Ma, Y. et al. Design, assembly, characterization, and operation of double-stranded interlocked DNA nanostructures. Nat Protoc 14, 2818–2855 (2019). https://doi.org/10.1038/s41596-019-0198-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41596-019-0198-7
- Springer Nature Limited
This article is cited by
-
Structure and dynamics of an archetypal DNA nanoarchitecture revealed via cryo-EM and molecular dynamics simulations
Nature Communications (2023)
-
External stimulation-controlled dynamic DNA devices for biosensing and biomedical applications
Science China Chemistry (2023)
-
Direct imaging of antigen–antibody binding by atomic force microscopy
Applied Nanoscience (2021)