Abstract
During interphase, the eukaryotic genome is organized into chromosome territories that are spatially segregated into compartment domains. The extent to which interacting domains or chromosomes are entangled is not known. We analyze series of co-occurring chromatin interactions using multi-contact 3C (MC-3C) in human cells to provide insights into the topological entanglement of chromatin. Multi-contact interactions represent percolation paths (C-walks) through three-dimensional (3D) chromatin space. We find that the order of interactions within C-walks that occur across interfaces where chromosomes or compartment domains interact is not random. Polymer simulations show that such C-walks are consistent with distal domains being topologically insulated, that is, not catenated. Simulations show that even low levels of random strand passage, for example by topoisomerase II, would result in entanglements, increased mixing at domain interfaces and an order of interactions within C-walks not consistent with experimental MC-3C data. Our results indicate that, during interphase, entanglements between chromosomes and chromosomal domains are rare.






Similar content being viewed by others
Data availability
The PacBio dataset is available at the GEO under accession no. GSE146945. Source data are provided with this paper.
Code availability
The C-walk assembly pipeline and scripts necessary to generate all plots are available at https://github.com/dekkerlab/MC-3C_scripts.
References
Cremer, T. & Cremer, C. Chromosome territories, nuclear architecture and gene regulation in mammalian cells. Nat. Rev. Genet. 2, 292–301 (2001).
Branco, M. R. & Pombo, A. Intermingling of chromosome territories in interphase suggests role in translocations and transcription-dependent associations. PLoS Biol. 4, e138 (2006).
Dorier, J. & Stasiak, A. Topological origins of chromosomal territories. Nucleic Acids Res. 37, 6316–6322 (2009).
Rosa, A. & Everaers, R. Structure and dynamics of interphase chromosomes. PLoS Comput. Biol. 4, e1000153 (2008).
Suzuki, J., Takano, A. & Matsushita, Y. Topological effect in ring polymers investigated with Monte Carlo simulation. J. Chem. Phys. 129, 034903 (2008).
Vettorel, T., Grosberg, A. Y. & Kremer, K. Statistics of polymer rings in the melt: a numerical simulation study. Phys. Biol. 6, 025013 (2009).
Müller, M., Wittmer, J. P. & Cates, M. E. Topological effects in ring polymers. II. Influence of persistence length. Phys. Rev. E 61, 4078–4089 (2000).
Branco, M. R., Branco, T., Ramirez, F. & Pombo, A. Changes in chromosome organization during PHA-activation of resting human lymphocytes measured by cryo-FISH. Chromosome Res. 16, 413–426 (2008).
Simonis, M. et al. Nuclear organization of active and inactive chromatin domains uncovered by chromosome conformation capture-on-chip (4C). Nat. Genet. 38, 1348–1354 (2006).
Lieberman-Aiden, E. et al. Comprehensive mapping of long-range interactions reveals folding principles of the human genome. Science 326, 289–293 (2009).
Dekker, J., Rippe, K., Dekker, M. & Kleckner, N. Capturing chromosome conformation. Science 295, 1306–1311 (2002).
Jiang, T. et al. Identification of multi-loci hubs from 4C-seq demonstrates the functional importance of simultaneous interactions. Nucleic Acids Res. 44, 8714–8725 (2016).
Olivares-Chauvet, P. et al. Capturing pairwise and multi-way chromosomal conformations using chromosomal walks. Nature 540, 296–300 (2016).
Allahyar, A. et al. Enhancer hubs and loop collisions identified from single-allele topologies. Nat. Genet. 50, 1151–1160 (2018).
Rubinstein, M. & Colby, R. H. Polymer Physics (Chemistry) (Oxford Univ. Press, 2003).
Li, H. Aligning sequence reads, clone sequences and assembly contigs with BWA-MEM. Preprint at arXiv https://arxiv.org/abs/1303.3997 (2013).
Dekker, J. Mapping in vivo chromatin interactions in yeast suggests an extended chromatin fiber with regional variation in compaction. J. Biol. Chem. 283, 34532–34540 (2008).
Jeon, C., Kim, J., Jeong, H., Jung, Y. & Ha, B.-Y. Chromosome-like organization of an asymmetrical ring polymer confined in a cylindrical space. Soft Matter 11, 8179–8193 (2015).
Doi, M. & Edwards, S. F. The Theory of Polymer Dynamics (Oxford Univ. Press, 1986).
Jost, D., Carrivain, P., Cavalli, G. & Vaillant, C. Modeling epigenome folding: formation and dynamics of topologically associated chromatin domains. Nucleic Acids Res. 42, 9553–9561 (2014).
Falk, M. et al. Heterochromatin drives organization of conventional and inverted nuclei. Nature 570, 395–399 (2019).
Klenin, K. & Langowski, J. Computation of writhe in modeling of supercoiled DNA. Biopolymers 54, 307–317 (2000).
Boettiger, A. N. et al. Super-resolution imaging reveals distinct chromatin folding for different epigenetic states. Nature 529, 418–422 (2016).
Baù, D. et al. The three-dimensional folding of the α-globin gene domain reveals formation of chromatin globules. Nat. Struct. Mol. Biol. 18, 107–114 (2011).
Quinodoz, S. A. et al. Higher-order inter-chromosomal hubs shape 3D genome organization in the nucleus. Cell 174, 744–757 (2018).
Bintu, B. et al. Super-resolution chromatin tracing reveals domains and cooperative interactions in single cells. Science 362, eaau1783 (2018).
Goundaroulis, D., Lieberman Aiden, E. & Stasiak, A. Chromatin is frequently unknotted at the megabase scale. Biophys. J. 118, 2268–2279 (2019).
de Gennes, P.-G. Scaling Theory of Polymer Physics (Cornell Univ. Press, 1979).
Holm, C., Stearns, T. & Botstein, D. DNA topoisomerase II must act at mitosis to prevent nondisjunction and chromosome breakage. Mol. Cell. Biol. 9, 159–168 (1989).
Canela, A. et al. Genome organization drives chromosome fragility. Cell 170, 507–521 (2017).
Gittens, W. H. et al. A nucleotide resolution map of Top2-linked DNA breaks in the yeast and human genome. Nat. Commun. 10, 4846 (2019).
Brahmachari, S. & Marko, J. F. Chromosome disentanglement driven via optimal compaction of loop-extruded brush structures. Proc. Natl Acad. Sci. USA 116, 24956–24965 (2019).
Orlandini, E., Marenduzzo, D. & Michieletto, D. Synergy of topoisomerase and structural-maintenance-of-chromosomes proteins creates a universal pathway to simplify genome topology. Proc. Natl Acad. Sci. USA 116, 8149–8154 (2019).
Racko, D., Benedetti, F., Goundaroulis, D. & Stasiak, A. Chromatin loop extrusion and chromatin unknotting. Polymers 10, 1126 (2018).
Goloborodko, A., Marko, J. F. & Mirny, L. A. Chromosome compaction by active loop extrusion. Biophys. J. 110, 2162–2168 (2016).
Goloborodko, A., Imakaev, M. V., Marko, J. F. & Mirny, L. Compaction and segregation of sister chromatids via active loop extrusion. Elife 5, e14864 (2016).
Rao, S. S. P. et al. A 3D map of the human genome at kilobase resolution reveals principles of chromatin looping. Cell 159, 1665–1680 (2014).
Vietri Rudan, M. et al. Comparative Hi-C reveals that CTCF underlies evolution of chromosomal domain architecture. Cell Rep. 10, 1297–1309 (2015).
de Wit, E. et al. CTCF binding polarity determines chromatin looping. Mol. Cell 60, 676–684 (2015).
Sanborn, A. L. et al. Chromatin extrusion explains key features of loop and domain formation in wild-type and engineered genomes. Proc. Natl Acad. Sci. USA 112, E6456–E6465 (2015).
Fudenberg, G. et al. Formation of chromosomal domains by loop extrusion. Cell Rep. 15, 2038–2049 (2016).
Marsden, M. P. & Laemmli, U. K. Metaphase chromosome structure: evidence for a radial loop model. Cell 17, 849–858 (1979).
Paulson, J. R. & Laemmli, U. K. The structure of histone-depleted metaphase chromosomes. Cell 12, 817–828 (1977).
Riggs, A. D. DNA methylation and late replication probably aid cell memory, and type I DNA reeling could aid chromosome folding and enhancer function. Philos. Trans. R. Soc. Lond. B Biol. Sci. 326, 285–297 (1990).
Nasmyth, K. Disseminating the genome: joining, resolving and separating sister chromatids during mitosis and meiosis. Annu. Rev. Genet. 35, 673–745 (2001).
Dekker, J. & Mirny, L. A. The 3D genome as moderator of chromosomal communication. Cell 164, 1110–1121 (2016).
Ganji, M. et al. Real-time imaging of DNA loop extrusion by condensin. Science 360, 102–105 (2018).
Kim, Y., Shi, Z., Zhang, H., Finkelstein, I. J. & Yu, H. Human cohesin compacts DNA by loop extrusion. Science 366, 1345–1349 (2019).
Davidson, I. F. et al. DNA loop extrusion by human cohesin. Science 366, 1338–1345 (2019).
Golfier, S., Quail, T., Kimura, H. & Brugués, J. Cohesin and condensin extrude DNA loops in a cell cycle-dependent manner. Elife 9, e53885 (2020).
Belaghzal, H., Dekker, J. & Gibcus, J. H. Hi-C 2.0: an optimized Hi-C procedure for high-resolution genome-wide mapping of chromosome conformation. Methods 123, 56–65 (2017).
Naumova, N. et al. Organization of the mitotic chromosome. Science 342, 948–953 (2013).
Abramo, K. et al. A chromosome folding intermediate at the condensin-to-cohesin transition during telophase. Nat. Cell Biol. 21, 1393–1402 (2019).
Bonev, B. et al. Multiscale 3D genome rewiring during mouse neural development. Cell 171, 557–572 (2017).
Norouzi, D. & Zhurkin, V. B. Dynamics of chromatin fibers: comparison of Monte Carlo simulations with force spectroscopy. Biophys. J. 115, 1644–1655 (2018).
Acknowledgements
We thank members of the Dekker and Mirny laboratories for helpful discussions. We acknowledge support from the National Institutes of Health Common Fund 4D Nucleome Program (DK107980) and the National Human Genome Research Institute (HG003143). J.D. is an investigator of the Howard Hughes Medical Institute.
Author information
Authors and Affiliations
Contributions
J.D. designed the project. B.D. and Y.L. performed the experimental MC-3C procedure. F.T.-C. and D.N. analyzed the data. D.N. designed and performed the simulations. F.T.-C., D.N. and J.D. wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Peer reviewer reports are available. Anke Sparmann was the primary editor on this article and managed its editorial process and peer review in collaboration with the rest of the editorial team.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Analysis of DNA sizes in control circular DNA, control linear DNA and 3C library DNA before and after exonuclease digestion.
Agarose gel analysis (0.8% agarose) of DNA size of control circular plasmid DNA, control linear plasmid DNA and of 3C library DNA before and after exonuclease digestion. 3C library and control plasmid DNA were digested with Exonuclease V and T5 exonuclease to determine whether 3C ligation products are linear or circular. In the presence of either exonuclease dsDNA will be digested, and circular DNA will remain present. As control both exonucleases are shown not to digest circular plasmid (pCMV6, 4.6 kb) (lane 1–3), whereas linearized plasmid was digested (lane 4–6). Different concentration of 3C library and ligase did not result in detection of circular ligation products as no DNA was detected after exonuclease treatment (lanes 7–18). Markers: 1 kb ladder (New England Biolabs).
Extended DataFig. 2 Statistics of the inter-chromosomal C-walks.
Additional analyses of inter-chromosomal C-walks. a, Proportion of inter-chromosomal steps for C- walks that visit only two distinct compartment domains, separated on the x-axis by the total number of steps in each C-walk (solid line). These are compared to the inter-chromosomal steps after permutating this set of C-walks (dots). This was done for all the walks that touch two domains (on the left), and separately for those that visit one A and one B domain (A-B), those that visit two A domains (A-A) and those that visit two B domains (B-B). b, Density plot of the maximum intra-chromosomal span (largest intra-chromosomal distance between fragments that make up a C-walk) for inter-chromosomal C-walks.
Extended Data Fig. 3 Ruling out the impact of random ligation events.
Frequency of intra- and inter-domain interactions as a function of the genomic distance. Most long-range interactions are inter-domain steps. We considered the possibility that the inter-domain steps are experimental noise due to random ligation events that do occur with some frequency in 3C-based experiments. To investigate this, here, we determined whether inter-domain steps display the expected inverse relationship with genomic distance that separates the loci. We find that the frequency of inter-domain steps decays rapidly. Such distance-dependent decay would not be expected if these interactions were the result of random ligation. a, Density of interactions corresponding to Fig. 1d with logarithmic binning similar to Bonev et al. 54. This graph shows clearly that indirect interactions are more long range. b, Contact probability profiles for steps in walks with single inter-compartment step (A-A, B-B, A-B). These steps follow a similar distance decay as direct and indirect interactions. This shows that inter-compartment steps are not random ligation events. c, Schematic of inter-domain interactions with (upper) or without (lower) distance decay. d, Observed distance decay scaling for C-walks with inter-compartment steps and the permutated versions of those walks. Left Panel (A-A): All inter-compartment A-A steps (purple; 2,980 interactions from all C-walks that contain one or more inter-compartment steps) and single inter-compartment A-A (red; 936 interactions from C-walks that contain only a single inter-compartment step). Middle Panel (B-B): All B-B steps (purple; 2,720 interactions) and single B-B (red; 882 interactions). Right Panel (A-B): All A-B steps (purple; 5,655 interactions) and single A-B (red; 1,392 interactions). The permutated walks mimic the effect of random ligation between distal domains. Randomized walks generate flat scaling profiles not consistent with the observations.
Extended Data Fig. 4 Details of the polymer simulations.
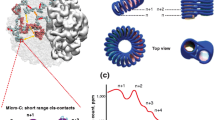
Details of the Monte-Carlo approach used to study the interface of interacting chromosomal domains in contact. a, Coarse-graining is done at the level of chromatin fibers with the diameter of ~25–30 nm. Every ~3 nucleosomes are considered one bead with the bead to bead bond distance of ~15 nm at equilibrium and the bending persistence length of 40 nm55. A hard-core excluded volume and a short-range pairwise interaction with the form Epairwise=q × exp[-2(r - σ)2 / σ2)] is considered, in which, σ is the diameter of the beads and ‘q’ is a small negative/positive charge depending on the attraction/repulsion between the beads. b, Monte Carlo is used because it is much more effective in sampling the phase space of possible conformations. Two types of Monte Carlo moves are possible. Those which don’t change the topology (that is the linking or knotting state) of the system and those which do. Top two moves, that is a single bead random movement and a crank-shaft rotation between two randomly chosen points along the chain, do not cause strand passage and do not move the relative position of the chain ends, hence do not affect the topology. Random rotations around a single point and/or brief random ghosting (no excluded volume) of parts of the chromatin fiber allow for relative movement of the ends and/or strand-passage, and thereby affect the topology. c, Equilibration for the case with topologically closed domains is presented in red. Equilibration for the case with topologically open domains with fixed-ends but with strand-passage is presented in black. Energy and linking number approach equilibrium after a few millions Monte Carlo steps. Linking number or number of entanglements is always an integer and remains zero for the case with topologically closed domains. RMSD values measured between two distal parts of a chain are shown. It takes up to ~50 million steps for the system to relax in a dynamic-globule structure. We call this a dynamic-globule because even at equilibrium, fluctuations in the distance between two regions on the same chain remain considerable (≈±100 nm). d, Simulation snapshots for the case of topologically closed domains with fixed-ends and no strand-passage in ‘Monte Carlo’ time from t0 to t3. The interface between the two domains remains unmixed. e, Simulation snapshots for the case of topologically open domains with fixed-ends but with strand-passage allowed. This mimics the activity of the Topo II enzyme. Mixing increases gradually as a function of the Monte Carlo time. Note that the domain ends, highlighted with red and black ball pairs, do not move. f, Simulation snapshots for the case of topologically open domains with no strand-passage but with freely moving ends. Domains mingle gradually as a function of the Monte Carlo time.
Extended Data Fig. 5 Simulation snapshots for domain interfaces at different solution conditions.
Equilibrated simulation snapshots for domain interfaces at different solution conditions. Two major determinants of polymer mixing are (1) whether polymer prefers to interact with the solvent (good solvent) or to interact with itself (bad solvent). This can be simulated effectively by a repulsive/attractive short-range interaction between the chain’s beads. Both conditions might be present in the nucleus leading to microphase separation of chromatin into euchromatin versus heterochromatin domains; and (2) the density of the polymer mixture, here simulated by the confining cylindrical walls surrounding the two chains. Our simulations at different solvent and under different crowding conditions still show that domains that are topologically unlinked will barely mix and retain their smooth interfaces. The result might be different for the good solvent in ultra-dilute condition, in which, domains will barely touch and won’t generate measurable multi-contact walks. We believe this condition is not relevant to the chromatin in vivo and would not be consistent with our MC-3C data.
Extended Data Fig. 6 Simul-walks statistics not dependent on the selected percolation parameters.
Simul-walks statistics do not change considerably by changing the selected percolation parameter rcutoff (Left: rcutoff=75 nm, Right: rcutoff=60 nm). The left panel is shown in the main Fig. 4 because the rcutoff is chosen in such a way that the maximum number of steps in simul-walks in the histogram is ~3-4 which is similar to the peak in the histogram of C-walks in Fig. 2c. Regardless, the top two histograms show that most walks still involve only 1 or 2 inter-chromosomal or inter-domain interactions. At the bottom, the shift between the simul-walks (black solid lines) and randomly permutated versions (circles) remains the same as well. The rcutoff value cannot be changed much more than this, because the minimum center to center distance of two touching beads is ~50–60 nm and choosing larger cutoffs will result in skipping some neighboring chains in the percolation path search. Therefore, our results are independent of the parameters we used to find the simul-walks.
Supplementary information
Supplementary Table 1
Sum of P values of a chi-square statistics test on the significance of ‘number of inter-chain steps in walks’ versus their ‘permutated versions’ as a measure for degree of mixing in C-walks or simul-walks. Stacked bar plots (for example, main Figs. 2 and 4) represent the distribution of the number of inter-chain steps in walks (observed). These distributions change considerably when the walks are randomized (expected) as described in the Methods. Chi-square goodness of fit test is a non-parametric test that is used to find how the observed value is significantly different from the expected value. It is a good test, especially when the sample data are divided into categories. In this test, the numbers of points that fall into each category (here each category is defined by the number of inter-domain or inter-chromosomal steps) are compared with the expected numbers of points in each category. Analysis is done for all of the walks containing at least three steps, and the calculated P values are summed up for all possible steps (usually a maximum of 6 or 10 steps are counted). The smaller the sum of the P values, the more significant the degree of unmixing is. The sum of P values for the inter-chromosomal C-walks in Fig. 2, <10−7, the topologically insulated simul-walks of Fig. 4, <10−8, and the inter-domain walks of Fig. 3f, <10−5, show considerable unmixing. Simul-walks with strand passage or free ends have large P values, indicating considerable mixing. The P values corresponding to the distal parts within the same compartment domain (related to Fig. 5 data) are in the intermediate range. Overall, the distal parts within the same domain are mixed, and mixing increases if strand passage is allowed. The sum of P values for the experimental C-walks of Fig. 5a do not indicate strand passage. StPs, strand passage; Ins, topologically insulated. Percentage of walks that represent pure percolation paths are shown for the inter-chromosomal C-walks of Fig. 2 and the simul-walks of Fig. 4. Pure percolations are walks with only one inter-chain crossing. Note that, even for the long walks of length 6 to 10, ~30% of the walks correspond to pure percolation paths rather than clusters of loci. The numbers are consistent between the inter-chromosomal C-walks and the topologically insulated simul-walks. This table quantifies the degree of mixing/unmixing and demonstrates the statistical significance of our findings.
Supplementary Table 2
Number of walks that visit each chromosome and compartment type. As the tables make clear, all chromosomes are represented in our data, with larger chromosomes generally being visited more often. A and B compartments are visited by a similar number of walks.
Source data
Source Data Fig. 1
Statistical source data for MC-3C direct and indirect interactions.
Source Data Fig. 2
Statistics of the inter/intra-chromosomal steps and the source data.
Source Data Fig. 3
Statistics of the inter/intra-domain steps and the source data.
Source Data Fig. 4
Statistics of the inter/intra-chain steps from polymer simulations and the source data.
Source Data Fig. 5
Statistics of the inter/intra-chain steps inside single domains and the source data.
Source Data Extended Data Fig. 2
Distribution of the inter-chromosomal C-walks.
Source Data Extended Data Fig. 3
Distance decay profiles as a function of the genomic distance.
Source Data Extended Data Fig. 6
Statistics of simul-walks under different percolation parameters.
Rights and permissions
About this article
Cite this article
Tavares-Cadete, F., Norouzi, D., Dekker, B. et al. Multi-contact 3C reveals that the human genome during interphase is largely not entangled. Nat Struct Mol Biol 27, 1105–1114 (2020). https://doi.org/10.1038/s41594-020-0506-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41594-020-0506-5
- Springer Nature America, Inc.
This article is cited by
-
Regulating epithelial-mesenchymal plasticity from 3D genome organization
Communications Biology (2024)
-
Ambiguities in neural-network-based hyperedge prediction
Journal of Applied and Computational Topology (2024)
-
Multi-feature clustering of CTCF binding creates robustness for loop extrusion blocking and Topologically Associating Domain boundaries
Nature Communications (2023)
-
The spatial organization of transcriptional control
Nature Reviews Genetics (2023)
-
High-throughput Pore-C reveals the single-allele topology and cell type-specificity of 3D genome folding
Nature Communications (2023)