Abstract
Astrocyte diversity is greatly influenced by local environmental modulation. Here we report that the majority of astrocytes across the mouse brain possess a singular primary cilium localized to the cell soma. Comparative single-cell transcriptomics reveals that primary cilia mediate canonical SHH signaling to modulate astrocyte subtype-specific core features in synaptic regulation, intracellular transport, energy and metabolism. Independent of canonical SHH signaling, primary cilia are important regulators of astrocyte morphology and intracellular signaling balance. Dendritic spine analysis and transcriptomics reveal that perturbation of astrocytic cilia leads to disruption of neuronal development and global intercellular connectomes in the brain. Mice with primary ciliary-deficient astrocytes show behavioral deficits in sensorimotor function, sociability, learning and memory. Our results uncover a critical role for primary cilia in transmitting local cues that drive the region-specific diversification of astrocytes within the developing brain.






Similar content being viewed by others
Data availability
scRNA-seq data are deposited at Gene Expression Omnibus and are publicly available (GSE253643). Source data are provided with this paper.
Code availability
Analysis code generated in this study is available through https://github.com/TheGuoLab/AstrocytesPrimaryCilia. Any additional information required to reanalyze the data reported in this paper is available from the corresponding author upon request.
References
Clarke, L. E. & Barres, B. A. Emerging roles of astrocytes in neural circuit development. Nat. Rev. Neurosci. 14, 311–321 (2013).
Liddelow, S. A. & Barres, B. A. Reactive astrocytes: production, function, and therapeutic potential. Immunity 46, 957–967 (2017).
Eroglu, C. & Barres, B. A. Regulation of synaptic connectivity by glia. Nature 468, 223–231 (2010).
Todd, F. W. et al. Neurons diversify astrocytes in the adult brain through sonic hedgehog signaling. Science 351, 849–854 (2016).
Batiuk, M. Y. et al. Identification of region-specific astrocyte subtypes at single cell resolution. Nat. Commun. 11, 1220 (2020).
Chai, H. et al. Neural circuit-specialized astrocytes: transcriptomic, proteomic, morphological, and functional evidence. Neuron 95, 531–549 (2017).
Khakh, B. S. & Deneen, B. The emerging nature of astrocyte diversity. Annu. Rev. Neurosci. 42, 187–207 (2019).
Hill, S. A. et al. Sonic hedgehog signaling in astrocytes mediates cell type-specific synaptic organization. eLife 8, e45545 (2019).
Xie, Y. et al. Astrocyte–neuron crosstalk through hedgehog signaling mediates cortical synapse development. Cell Rep. 38, 110416 (2022).
Garcia, A. D. R., Petrova, R., Eng, L. & Joyner, A. L. Sonic hedgehog regulates discrete populations of astrocytes in the adult mouse forebrain. J. Neurosci. 30, 13597–13608 (2010).
Stogsdill, J. A. et al. Astrocytic neuroligins control astrocyte morphogenesis and synaptogenesis. Nature 551, 192–197 (2017).
Wheway, G., Nazlamova, L. & Hancock, J. T. Signaling through the primary cilium. Front. Cell Dev. Biol. 6, 8 (2018).
Schou, K. B., Pedersen, L. B. & Christensen, S. T. Ins and outs of GPCR signaling in primary cilia. EMBO Rep. 16, 1099–1113 (2015).
Christensen, S. T., Clement, C. A., Satir, P. & Pedersen, L. B. Primary cilia and coordination of receptor tyrosine kinase (RTK) signalling. J. Pathol. 226, 172–184 (2012).
Hilgendorf, K. I., Johnson, C. T. & Jackson, P. K. The primary cilium as a cellular receiver: organizing ciliary GPCR signaling. Curr. Opin. Cell Biol. 39, 84–92 (2016).
Delling, M., DeCaen, P. G., Doerner, J. F., Febvay, S. & Clapham, D. E. Primary cilia are specialized calcium signalling organelles. Nature 504, 311–314 (2013).
Guo, J. et al. Primary cilia signaling shapes the development of interneuronal connectivity. Dev. Cell 42, 286–300 (2017).
Phua, S. C., Lin, Y.-C. & Inoue, T. An intelligent nano-antenna: primary cilium harnesses TRP channels to decode polymodal stimuli. Cell Calcium 58, 415–422 (2015).
Badano, J. L., Mitsuma, N., Beales, P. L. & Katsanis, N. The ciliopathies: an emerging class of human genetic disorders. Annu. Rev. Genomics Hum. Genet. 7, 125–148 (2006).
Hildebrandt, F., Benzing, T. & Katsanis, N. Ciliopathies. N. Eng. J. Med. 364, 1533–1543 (2011).
Novarino, G., Akizu, N. & Gleeson, J. G. Modeling human disease in humans: the ciliopathies. Cell 147, 70–79 (2011).
Guo, J. et al. Developmental disruptions underlying brain abnormalities in ciliopathies. Nat. Commun. 6, 7857 (2015).
Higginbotham, H. et al. ARL13B-regulated cilia activities are essential for polarized radial glial scaffold formation. Nat. Neurosci. 16, 1000–1007 (2013).
Guo, J. et al. Primary cilia signaling promotes axonal tract development and is disrupted in Joubert syndrome-related disorders models. Dev. Cell 51, 759–774 (2019).
Alvarez Retuerto, A. I. et al. Association of common variants in the Joubert syndrome gene (AHI1) with autism. Hum. Mol. Genet. 17, 3887–3896 (2008).
Brancati, F., Dallapiccola, B. & Valente, E. Joubert syndrome and related disorders. Orphanet J. Rare Dis. 5, 20 (2010).
Cantagrel, V. et al. Mutations in the cilia gene ARL13B lead to the classical form of Joubert syndrome. Am. J. Hum. Genet. 83, 170–179 (2008).
Migliavacca, E. et al. A potential contributory role for ciliary dysfunction in the 16p11.2 600 kb BP4–BP5 pathology. Am. J. Hum. Genet. 96, 784–796 (2015).
Guemez-Gamboa, A., Coufal, N. G. & Gleeson, J. G. Primary cilia in the developing and mature brain. Neuron 82, 511–521 (2014).
Green, J. A. & Mykytyn, K. Neuronal ciliary signaling in homeostasis and disease. Cell. Mol. Life Sci. 67, 3287–3297 (2010).
Amador-Arjona, A. et al. Primary cilia regulate proliferation of amplifying progenitors in adult hippocampus: implications for learning and memory. J. Neurosci. 31, 9933–9944 (2011).
Min, P. S., Jin, J. H. & Ho, L. J. Roles of primary cilia in the developing brain. Front. Cell. Neurosci. 13, 1358 (2019).
Sheu, S. H. et al. A serotonergic axon-cilium synapse drives nuclear signaling to alter chromatin accessibility. Cell 185, 3390–3407.e18 (2022).
Guadiana, S. M. et al. Arborization of dendrites by developing neocortical neurons is dependent on primary cilia and type 3 adenylyl cyclase. J. Neurosci. 33, 2626–2638 (2013).
Kang, K. & Song, M.-R. Diverse FGF receptor signaling controls astrocyte specification and proliferation. Biochem. Biophys. Res. Commun. 395, 324–329 (2010).
Savchenko, E. et al. FGF family members differentially regulate maturation and proliferation of stem cell-derived astrocytes. Sci. Rep. 9, 9610 (2019).
Acaz-Fonseca, E., Ortiz-Rodriguez, A., Azcoitia, I., Garcia-Segura, L. M. & Arevalo, M.-A. Notch signaling in astrocytes mediates their morphological response to an inflammatory challenge. Cell Death Discov. 5, 85 (2019).
Su, C. Y., Bay, S. N., Mariani, L. E., Hillman, M. J. & Caspary, T. Temporal deletion of Arl13b reveals that a mispatterned neural tube corrects cell fate over time. Development 139, 4062–4071 (2012).
Haycraft, C. J. et al. Intraflagellar transport is essential for endochondral bone formation. Development 134, 307–316 (2007).
Humbert, M. C. et al. ARL13B, PDE6D, and CEP164 form a functional network for INPP5E ciliary targeting. Proc. Natl Acad. Sci. USA 109, 19691–19696 (2012).
Pazour, G. J. et al. Chlamydomonas IFT88 and its mouse homologue, polycystic kidney disease gene Tg737, are required for assembly of cilia and flagella. J. Cell Biol. 151, 709–718 (2000).
Freeman, M. R. Specification and morphogenesis of astrocytes. Science 330, 774–778 (2010).
Caspary, T., Larkins, C. E. & Anderson, K. V. The graded response to sonic hedgehog depends on cilia architecture. Dev. Cell 12, 767–778 (2007).
Matusova, Z., Hol, E. M., Pekny, M., Kubista, M. & Valihrach, L. Reactive astrogliosis in the era of single-cell transcriptomics. Front. Cell Neurosci. 17, 1173200 (2023).
Corbit, K. C. et al. Vertebrate Smoothened functions at the primary cilium. Nature 437, 1018–1021 (2005).
Xie, J. et al. Activating Smoothened mutations in sporadic basal-cell carcinoma. Nature 391, 90–92 (1998).
Zhuo, L. et al. hGFAP‐cre transgenic mice for manipulation of glial and neuronal function in vivo. Genesis 31, 85–94 (2001).
Siehler, S. Regulation of RhoGEF proteins by G12/13‐coupled receptors. Br. J. Pharmacol. 158, 41–49 (2009).
Gigante, E. D., Taylor, M. R., Ivanova, A. A., Kahn, R. A. & Caspary, T. ARL13B regulates sonic hedgehog signaling from outside primary cilia. eLife 9, e50434 (2020).
Ferent, J. et al. The ciliary protein ARL13B functions outside of the primary cilium in SHH-mediated axon guidance. Cell Rep. 29, 3356–3366 (2019).
Nakagawa, N. et al. Memo1-mediated tiling of radial glial cells facilitates cerebral cortical development. Neuron 103, 836–852 (2019).
Gutkind, J. S. The pathways connecting G protein-coupled receptors to the nucleus through divergent mitogen-activated protein kinase cascades. J. Biol. Chem. 273, 1839–1842 (1998).
Aibar, S. et al. SCENIC: single-cell regulatory network inference and clustering. Nat. Methods 14, 1083–1086 (2017).
Sprenkle, N. T., Sims, S. G., Sánchez, C. L. & Meares, G. P. Endoplasmic reticulum stress and inflammation in the central nervous system. Mol. Neurodegener. 12, 42 (2017).
Chung, W.-S. et al. Astrocytes mediate synapse elimination through MEGF10 and MERTK pathways. Nature 504, 394–400 (2013).
Morizawa, Y. M. et al. Synaptic pruning through glial synapse engulfment upon motor learning. Nat. Neurosci. 25, 1458–1469 (2022).
Feng, G. et al. Imaging neuronal subsets in transgenic mice expressing multiple spectral variants of GFP. Neuron 28, 41–51 (2000).
Jin, S. et al. Inference and analysis of cell–cell communication using CellChat. Nat. Commun. 12, 1088 (2021).
Thomas, S. et al. Identification of a novel ARL13B variant in a Joubert syndrome-affected patient with retinal impairment and obesity. Eur. J. Hum. Genet 23, 621–627 (2015).
Miertzschke, M., Koerner, C., Spoerner, M. & Wittinghofer, A. Structural insights into the small G-protein ARL13B and implications for Joubert syndrome. Biochem. J. 457, 301–311 (2014).
Rafiullah, R. et al. A novel homozygous ARL13B variant in patients with Joubert syndrome impairs its guanine nucleotide-exchange factor activity. Eur. J. Hum. Genet 25, 1324–1334 (2017).
Lawal, O., Severino, F. P. U. & Eroglu, C. The role of astrocyte structural plasticity in regulating neural circuit function and behavior. Glia 70, 1467–1483 (2022).
Bronzuoli, M. R. et al. Neuroglia in the autistic brain: evidence from a preclinical model. Mol. Autism 9, 66 (2018).
Petrelli, F., Pucci, L. & Bezzi, P. Astrocytes and microglia and their potential link with autism spectrum disorders. Front. Cell. Neurosci. 10, 21 (2016).
Wang, Y. et al. Melanocortin 4 receptor signals at the neuronal primary cilium to control food intake and body weight. J. Clin. Invest. 131, e142064 (2021).
Higginbotham, H. et al. ARL13B in primary cilia regulates the migration and placement of interneurons in the developing cerebral cortex. Dev. Cell 23, 925–938 (2012).
DeMars, K. M., Ross, M. R., Starr, A. & McIntyre, J. C. Neuronal primary cilia integrate peripheral signals with metabolic drives. Front. Physiol. 14, 1150232 (2023).
Ma, R., Kutchy, N. A., Chen, L., Meigs, D. D. & Hu, G. Primary cilia and ciliary signaling pathways in aging and age-related brain disorders. Neurobiol. Dis. 163, 105607 (2021).
Karunakaran, K. B., Chaparala, S., Lo, C. W. & Ganapathiraju, M. K. Cilia interactome with predicted protein–protein interactions reveals connections to Alzheimer’s disease, aging and other neuropsychiatric processes. Sci. Rep. 10, 15629 (2020).
Schmidt, S. et al. Primary cilia and SHH signaling impairments in human and mouse models of Parkinson’s disease. Nat. Commun. 13, 4819 (2022).
Sofroniew, M. V. Astrocyte reactivity: subtypes, states, and functions in CNS innate immunity. Trends Immunol. 41, 758–770 (2020).
Bouvier, D. S. et al. The multifaceted neurotoxicity of astrocytes in ageing and age-related neurodegenerative diseases: a translational perspective. Front. Physiol. 13, 814889 (2022).
Koike, K. et al. Danger perception and stress response through an olfactory sensor for the bacterial metabolite hydrogen sulfide. Neuron 109, 2469–2484 (2021).
Long, F., Zhang, X. M., Karp, S., Yang, Y. & McMahon, A. P. Genetic manipulation of hedgehog signaling in the endochondral skeleton reveals a direct role in the regulation of chondrocyte proliferation. Development 128, 5099–5108 (2001).
Ganat, Y. M. et al. Early postnatal astroglial cells produce multilineage precursors and neural stem cells in vivo. J. Neurosci. 26, 8609–8621 (2006).
Madisen, L. et al. A robust and high-throughput Cre reporting and characterization system for the whole mouse brain. Nat. Neurosci. 13, 133–140 (2010).
Jeong, J., Mao, J., Tenzen, T., Kottmann, A. H. & McMahon, A. P. Hedgehog signaling in the neural crest cells regulates the patterning and growth of facial primordia. Genes Dev. 18, 937–951 (2004).
Zhang, Y. et al. Purification and characterization of progenitor and mature human astrocytes reveals transcriptional and functional differences with mouse. Neuron 89, 37–53 (2016).
Chen, J. K., Taipale, J., Cooper, M. K. & Beachy, P. A. Inhibition of hedgehog signaling by direct binding of cyclopamine to Smoothened. Genes Dev. 16, 2743–2748 (2002).
Byun, Y. G. & Chung, W. S. A novel in vitro live-imaging assay of astrocyte-mediated phagocytosis using pH indicator-conjugated synaptosomes. J. Vis. Exp. 5, 56647 (2018).
Auguste, Y. S. S. et al. Oligodendrocyte precursor cells engulf synapses during circuit remodeling in mice. Nat. Neurosci. 25, 1273–1278 (2022).
Wheway, G. & Mitchison, H. M. Opportunities and challenges for molecular understanding of ciliopathies—The 100,000 Genomes Project. Front. Genet. 10, 127 (2019).
Zheng, G. X. et al. Massively parallel digital transcriptional profiling of single cells. Nat. Commun. 8, 14049 (2017).
Hafemeister, C. & Satija, R. Normalization and variance stabilization of single-cell RNA-seq data using regularized negative binomial regression. Genome Biol. 20, 296 (2019).
Korsunsky, I. et al. Fast, sensitive and accurate integration of single-cell data with Harmony. Nat. Methods 16, 1289–1296 (2019).
Wilk, A. J. et al. Multi-omic profiling reveals widespread dysregulation of innate immunity and hematopoiesis in COVID-19. J. Exp. Med. 218, e20210582 (2021).
Alquicira-Hernandez, J. & Powell, J. E. Nebulosa recovers single-cell gene expression signals by kernel density estimation. Bioinformatics 37, 2485–2487 (2021).
Vorhees, C. V. & Williams, M. T. Morris water maze: procedures for assessing spatial and related forms of learning and memory. Nat. Protoc. 1, 848–858 (2006).
Acknowledgements
J.G. is a New York Stem Cell Foundation Robertson Investigator. This research was supported by The New York Stem Cell Foundation (NYSCF-R-N163 to J.G.), the Canadian Institutes of Health Research (RN418797-438409 to J.G.) and the National Science and Engineering Research Council (RGPIN-2019-04820 to J.G.). We thank the support from the Harley N. Hotchkiss Samuel Weiss Postdoctoral Fellowship (L.W.), Cumming School of Medicine Postdoctoral Scholarship (L.W.) and ACHRI STEP Postdoctoral Fellowship (L.W.).
Author information
Authors and Affiliations
Contributions
Conceptualization: L.W. and J.G. Mouse experiments and husbandry: L.W., Q.G., X.Z., V.H., B.W., M.M., S.C., C.C. and J.G. Laboratory experiments: L.W., Q.G., X.Z., V.H., B.W., A.Y., N.R., E.L., C.C., M.M., S.C., K.G., J.H., D.A.E., T.R., H.K. and J.G. Bioinformatics: L.W., S.A. and N.R. Paper editing and review: L.W., Q.Z., G.G., J.B. and J.G. Paper writing: L.W. and J.G.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Neuroscience thanks Markus Delling, Steven Sloan and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Tamoxifen mediated Cre recombination and ciliary gene deletion in astrocytes.
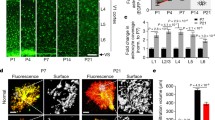
(a, b) Tamoxifen mediated highly efficient and consistent Cre recombination in all 5 mouse lines. (a) Representative images of astrocytic tdTom (Ai9) expression in S100β+ astrocytes in the cerebral cortex in control brains. Mice were injected with tamoxifen from P7 to P9, and brains were collected at P35. (b) Quantifications of the % of S100β+ astrocytes that expressed tdTom in all 5 mouse lines. N = 4 mice per group. (c) Primary cilia (labeled by anti-Arl13b) are absent in Ift88lox/lox; Aldh1l1CreERT2; Ai9 (Ift88cKO) mice. Mice were injected with tamoxifen from P7 to P9, and brains were collected at P35. This experiment was repeated in 3 independent mice. (d, e) Cilia morphology of control and Arl13bcKO cultured primary astrocytes stained by anti-acetylated tubulin antibody. N = 30–33 cells per group. Comparison was performed using unpaired two-tailed Student’s t-test. ***P < 0.001. Scale bar in (a), 50 µm; scale bars, 10 µm (c), 5 µm (d).
Extended Data Fig. 2 Single cell RNAseq analyses of astrocyte clusters.
(a) Astrocyte sorting using flow cytometry. Cortices and cerebellums were dissected and were digested to isolate single cells. Cells were stained with viability dye E780 and ACSA2-PE. ACSA2 positive live cells were sorted to construct the single cell RNA seq libraries. (b) UMAP plots of all cells annotated with main cell types. (c) UMAP plots of all cells grouped by tissue types from which the cells are isolated from. (d) Expression of representative cell type markers used for cell annotation. (e) Expression patterns of regional markers and representative cluster marker genes of cortical astrocytes. (f) Heatmap of top cortical astrocyte cluster markers and the related GO terms. (g) GSEA plot of enriched key GO pathways in cortical astrocyte clusters.
Extended Data Fig. 3 Single cell RNAseq analyses of Ift88cKO cortical astrocytes.
(a) Kernel density estimates depicting magnitude of the molecular changes in Ift88cKO calculated by summing DEG FCs for each cortical astrocyte cluster. (b) Venn diagrams of DEGs found in Arl13bcKO and Ift88cKO groups (compared to Control) across cortical astrocyte Cluster 0, 1, 2. (c) Visualization of top DEGs in Ift88cKO cortical astrocyte clusters. DEGs found using the function of FindMarkers in Seurat R package and were defined significant when P_adj < 0.05. (d-f) Top GO pathways significantly altered (P < 0.05) in Arl13bcKO and Ift88cKO cortical astrocyte clusters. The insignificant GO terms are indicated as P > 0.05. (g) The influence of Ift88 deletion on the key GO pathways in cortical astrocyte clusters. GSEA was used for statistical analysis in (d-g).
Extended Data Fig. 4 Single cell RNAseq analyses of cerebellar astrocytes.
(a) Violin plot of the expression of genes used to distinguish Bergmann glia (BG) and velate astrocytes (VA). (b) GSEA plot of enriched key GO pathways in cerebellar astrocyte clusters. (c-f) Top GO pathways altered in Arl13bcKO cerebellar astrocyte clusters. (g) Gene signature expression of pan-reactive, A1-reactive, A2-reactive and Alzheimer’s disease (AD)-associated astrocytes in control and Arl13bcKO cerebellar astrocyte clusters BG_I, BG_II, BG_III, and VA. A2 reactive astrocyte signature is elevated in Arl13bcKO BG_III cluster.
Extended Data Fig. 5 Shh signaling related changes in Arl13bcKO, Ift88cKO, and SmocKO groups.
(a, b) SmoM2 ciliary localization in astrocytes in SmoM2-eYFP; Aldh1l1-CreERT2; Ai9 mice. 82.14% of SmoM2+ astrocytes showed SmoM2 primary ciliary localization. N = 36 cells from 3 mice. (c, d) Kir4.1 expression patterns in Control and Ift88cKO mouse cortex. Kir4.1 expression is pseudocolored based on intensity. (e) Kir4.1 expression patterns in Aldh1l1-CreERT2, Arl13blox/lox; Aldh1l1-CreERT2, Arl13blox/lox; hGFAP-Cre, Ift88lox/lox; Aldh1l1-CreERT2, and Smolox/lox; Aldh1l1-CreERT2 mouse cerebellum. (f) Kir4.1 expression patterns in Aldh1l1-CreERT2, Arl13blox/lox; Aldh1l1-CreERT2, Ift88lox/lox; Aldh1l1-CreERT2, and Smolox/lox; Aldh1l1-CreERT2 mouse hippocampus. (f’) Kir4.1 expression pseudocolored based on intensity in (f). (g) GluA1 expression patterns in Aldh1l1-CreERT2, Arl13blox/lox; Aldh1l1-CreERT2, Ift88lox/lox; Aldh1l1-CreERT2, and Smolox/lox; Aldh1l1-CreERT2 mouse cerebellum. (h) GluA1 expression patterns in Aldh1l1-CreERT2, Arl13blox/lox; Aldh1l1-CreERT2, Ift88lox/lox; Aldh1l1-CreERT2, and Smolox/lox; Aldh1l1-CreERT2 mouse hippocampus. (h’) GluA1 expression pseudocolored based on intensity in (h). All brains were collected at the age of 2 months. (i, j) Cre recombination efficiency in Arl13blox/lox; SmoM2-eYFP; Aldh1l1-CreERT2; Ai9 mice. 97.6% of all Cre+ (tdTom+) astrocytes expressed SmoM2-eYFP. All mice were administered with tamoxifen from P7 to P9 and brains were collected at P35. (k, l) SmoM2-eYFP expressing astrocytes show complete loss of Arl13b protein in SmoM2-Arl13bcKO astrocytes. SmoM2-eYFP was labeled by anti-GFP antibody. Scale bars, 5 µm (a, i, k, l), 50 µm (c-h). Experiments in (a, e-i, k, l) were repeated at least three times using independent mice. (m) Cell percentage of cortical astrocyte clusters in all samples. (n) Top Cilia-cShh GO biological processes in Arl13bcKO were similarly changed in Ift88cKO cortical astrocyte clusters. (o) Expression of representative BGIII marker genes in cerebellar astrocytes. (p) Bubble plots reflecting the expression of BGIII marker genes across sample groups. (q) Visualization of the Cilia-cShh related GO terms that are commonly up- or down-regulated in BG and VA in Arl13bcKO and SmocKO groups. Adjusted P values (P_adj) and normalized enrichment scores (NES) shown in (n, q) were derived from statistical tests of GSEA. When P_adj <0.05 and NES < 0, it is defined as downregulated (down); when P_adj <0.05 and NES > 0, it is defined as upregulated (up); when P_adj > 0.05, it is defined as unchanged.
Extended Data Fig. 6 Reduced actin dynamics and morphological defects in cilia mutant astrocytes.
(a) Circos plot of top GO biological processes enriched for actin dynamics regulation in Ift88cKO cortical Cluster 1. (b) The expression of mArl13bWT and mArl13bV358A in Arl13bcKO astrocytes. AAV viruses AAV5-GAG-DIO-mArl13bWT-mCh or AAV5-CAG-DIO-mArl13bV358A-mCh, together with AAV5-GFAP-GFP was injected in the lateral ventricles of Arl13blox/lox; Aldh1l1-CreERT2 (Arl13bcKO) brains at P1. Tamoxifen was administered from P7 to P9. Brain sections were collected at P35. Scale bar, 5 µm. This experiment was repeated three times using individual mice. (c) Morphological defects in ciliary mutants. Surface area of upper- (control, n = 59; Ift88cKO, n = 30; Arl13bcKO, n = 28; SmoM2-Arl13bcKO, n = 23; mArl13b_rescue-Arl13bcKO, n = 28; mArl13bV358A_rescue-Arl13bcKO, n = 32 astrocytes) and deeper-cortical (control, n = 49; Ift88cKO, n = 30; Arl13bcKO, n = 51; SmoM2-Arl13bcKO, n = 25; mArl13b_rescue-Arl13bcKO, n = 34; mArl13bV358A_rescue-Arl13bcKO, n = 29 astrocytes) astrocytes was measured after morphology reconstruction using IMARIS. All data of astrocyte morphology analysis were derived from 3 male mice per group. Comparisons of groups were performed using one-way ANOVA followed by Tukey’s test. *P < 0.05, **P < 0.01, ***P < 0.001.
Extended Data Fig. 7 Arl13bcKO but not SmocKO astrocytes show increased GPCR signaling activity, elevated ER stress response and distinct gene regulatory networks.
(a) Representative circos plot of top GO biological processes enriched for GPCR signaling in Arl13bcKO cortical astrocyte cluster 1. (b) Expression patterns of M3 (Chrm3) in cultured control, Arl13bcKO, and SmocKO cortical astrocytes. This experiment was repeated three times. (c, d) Western blotting of phospho-MAPK using cultured control and Arl13bcKO astrocytes (n = 3 biological replicates per group). Data are shown as means ± SEM. Comparison of three groups was performed using one-way-ANOVA followed by Tukey’s test. *P < 0.05. (e) Representative circos plot of cShh independent pathways enriched for ER stress in Arl13bcKO cortical astrocyte cluster 1. (f-h) Rank plot of Arl13bcKO and SmocKO top regulons ordered by regulon specificity score (RSS) in cortical astrocyte clusters. (i) Immunostaining of XBP1 and ATF6 in cultured control, Arl13bcKO, and SmocKO cortical astrocytes. This experiment was repeated twice. Scale bars, 10 µm. (j, k) Rank plot of Arl13bcKO and SmocKO top regulons ordered by regulon specificity score (RSS) in cerebellar astrocyte clusters. Scale bars, 10 µm.
Extended Data Fig. 8 Primary ciliary dysfunction resulted in astrocytic pruning and neuronal spine defects.
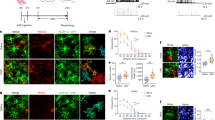
(a) Illustration of astrocytic engulfment analysis. (b-c) IMARIS volumetric reconstructions of astrocytes (green) containing Vglut1 (magenta) in the hippocampus in male control and Arl13bcKO brains. (b’, c’) Enlarged region of (b) and (c) indicated by red squares. (b’, c’) cross-section images from the yellow line indicated locations in (b) and (c). Scale bars in (a-c), 5 µm. (d) Quantifications of the Vglut1+ puncta within astrocytes in the hippocampus of control (n = 6 astrocytes) and Arl13bcKO (n = 9 astrocytes) from 3 male mice per group. (e) Quantifications of the Vglut1+ puncta within astrocytes in the upper (control, n = 17; Arl13bcKO, n = 18 astrocytes) and deeper (control, n = 16; Arl13bcKO, n = 16 astrocytes) layers of cortex, and hippocampus (control, n = 13; Arl13bcKO, 17 astrocytes) from 3 female mice per group. (f, g) Quantifications of the Vglut1+ puncta engulfed by upper- (control, n = 19; Ift88cKO, n = 20; Arl13bcKO, n = 21; SmoM2-Arl13bcKO, n = 20; mArl13b_rescue-Arl13bcKO, n = 19; mArl13bV358A_rescue-Arl13bcKO, n = 14 astrocytes) and deeper- cortical astrocytes (control, n = 18; Ift88cKO, n = 18; Arl13bcKO, n = 18; SmoM2-Arl13bcKO, n = 20; mArl13b_rescue-Arl13bcKO, n = 14; mArl13bV358A_rescue-Arl13bcKO, n = 17 astrocytes) from 3 male mice per group. Arl13bcKO mice in the mArl13bWT or mArl13bV358A groups were injected with AAV5-CAG-DIO-Arl13bWT-mCh or AAV5-CAG-DIO-Arl13bV358A-mCh viruses, respectively, together with AAV5-GFAP-GFP viruses, at P1. All mice were injected with tamoxifen from P7 to P9 and brains were collected at P35 for analysis. Vglut1+ puncta engulfed by astrocytes were analyzed by IMARIS. (h, i) Quantifications of spine volume, area, length, diameter in apical (control, n = 18 dendrites; Arl13bcKO, n = 13 dendrites) and basal (control, n = 18 dendrites; Arl13bcKO, n = 22 dendrites) dendrites from 3 male mice per group. (j, k) Quantifications of spine characteristics of M1 apical and basal dendrites from female mice. (l, m) Fraction of spine types in M1 cortex of female control and Arl13bcKO mice. Data shown in (j-m) were analyzed from apical (control, n = 31 dendrites; Arl13bcKO, n = 23 dendrites) and basal (control, n = 33 dendrites; Arl13bcKO, n = 21 dendrites) dendrites from 3 female mice per group. Comparisons of two groups were performed using unpaired two-tailed Student’s t-test and comparisons of six groups were performed using one-way ANOVA followed by Tukey’s test. *P < 0.05, **P < 0.01, ***P < 0.001.
Extended Data Fig. 9 Single nucleus RNA sequencing of control and Arl13bcKO cortices.
(a) UMAP plot of all cells from single nucleus RNA sequencing. (b) Density plot of representative marker genes used for cell type annotation. (c) Density plot of markers used to identify upper layers (Layer II/III, Layer IV) and deeper layers (Layer V, Layer VI) excitatory neurons and inhibitory neurons. (d) CellChat predicted number of interactions from astrocytes to other cell types in control and Arl13bcKO groups. (e) CellChat generated dot plot showing the significant ligand-receptor pairs between Control and Arl13bcKO brains, which contribute to the signaling from astrocytes to other cell types. Dot color reflects communication probabilities and dot size represents P values computed using a one-sided permutation test. (f) CellChat predicted altered signaling interaction patterns in Arl13bcKO brains.
Extended Data Fig. 10 Behavior tests in control, Arl13bcKO, and SmoM2-Arl13bcKO mice.
(a) Quantifications of the latency time to fall in the rotarod test from control (n = 21), Arl13bcKO (n = 18), SmoM2-Arl13bcKO (n = 8) male mice. (b) Quantifications of the number of foot slips in the beam walk test from control (n = 17), Arl13bcKO (n = 14), SmoM2-Arl13bcKO (n = 8) male mice. (c) Quantifications of the time in chamber and sniffing time in the 3-chamber sociability test from control (n = 20), Arl13bcKO (n = 16), SmoM2-Arl13bcKO (n = 8) male mice. (d) Quantifications of % of freezing time in the fear conditioning test from control (n = 22), Arl13bcKO (n = 15), SmoM2-Arl13bcKO (n = 8) male mice. (e) Quantifications for the Morris water maze test from control (n = 14), Arl13bcKO (n = 10), SmoM2-Arl13bcKO (n = 8) male mice. Data in (a, e) are shown as means ± SEM, and data in (b-d) are represented as box-and-whisker plots, presenting the upper and lower quartiles, mean and median. Comparisons were performed using one-way ANOVA followed by Tukey’s test. *P in red, Arl13bcKO vs. Control; * P in purple, SmoM2-Arl13bcKO vs. Control. *P < 0.05, **P < 0.01, ***P < 0.001.
Supplementary information
Source data
Source Data Fig. 4
Unprocessed western blots.
Source Data Extended Data Fig. 7
Unprocessed western blots.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Wang, L., Guo, Q., Acharya, S. et al. Primary cilia signaling in astrocytes mediates development and regional-specific functional specification. Nat Neurosci 27, 1708–1720 (2024). https://doi.org/10.1038/s41593-024-01726-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41593-024-01726-z
- Springer Nature America, Inc.
This article is cited by
-
Astrocyte antennae
Nature Reviews Neuroscience (2024)