Abstract
In addition to its motor functions, the cerebellum is involved in emotional regulation, anxiety and affect. We found that suppressing the firing of cerebellar Purkinje cells (PCs) rapidly excites forebrain areas that contribute to such functions (including the amygdala, basal forebrain and septum), but that the classic cerebellar outputs, the deep cerebellar nuclei, do not directly project there. We show that PCs directly inhibit parabrachial nuclei (PBN) neurons that project to numerous forebrain regions. Suppressing the PC–PBN pathway influences many regions in the forebrain and is aversive. Molecular profiling shows that PCs directly inhibit numerous types of PBN neurons that control diverse behaviors that are not involved in motor control. Therefore, the PC–PBN pathway allows the cerebellum to directly regulate activity in the forebrain, and may be an important substrate for cerebellar disorders arising from damage to the posterior vermis.





Similar content being viewed by others
Data availability
Analysis relating to the RNA-seq dataset has been deposited on Zenodo (https://doi.org/10.5281/zenodo.6653204). Other data that support the findings of this study are available from the corresponding author upon reasonable request.
References
Jackman, S. L. et al. Cerebellar Purkinje cell activity modulates aggressive behavior. eLife 9, e53229 (2020).
Chabrol, F. P., Blot, A. & Mrsic-Flogel, T. D. Cerebellar contribution to preparatory activity in motor neocortex. Neuron 103, 506–519.e4 (2019).
Gao, Z. et al. A cortico-cerebellar loop for motor planning. Nature 563, 113–116 (2018).
Rochefort, C. et al. Cerebellum shapes hippocampal spatial code. Science 334, 385–389 (2011).
Joyal, C. C. et al. Effects of midline and lateral cerebellar lesions on motor coordination and spatial orientation. Brain Res. 739, 1–11 (1996).
Frontera, J. L. et al. Bidirectional control of fear memories by cerebellar neurons projecting to the ventrolateral periaqueductal grey. Nat. Commun. 11, 5207 (2020).
Vaaga, C. E., Brown, S. T. & Raman, I. M. Cerebellar modulation of synaptic input to freezing-related neurons in the periaqueductal gray. eLife 9, e54302 (2020).
Krook-Magnuson, E., Szabo, G. G., Armstrong, C., Oijala, M. & Soltesz, I. Cerebellar directed optogenetic intervention inhibits spontaneous hippocampal seizures in a mouse model of temporal lobe epilepsy. eNeuro 1, ENEURO.0005-14.2014 (2014).
Schmahmann, J. D. & Sherman, J. C. The cerebellar cognitive affective syndrome. Brain 121, 561–579 (1998).
Anand, B. K., Malhotra, C. L., Singh, B. & Dua, S. Cerebellar projections to limbic system. J. Neurophysiol. 22, 451–457 (1959).
Whiteside, J. A. & Snider, R. S. Relation of cerebellum to upper brain stem. J. Neurophysiol. 16, 397–413 (1953).
Heath, R. G. & Harper, J. W. Ascending projections of the cerebellar fastigial nucleus to the hippocampus, amygdala, and other temporal lobe sites: evoked potential and histological studies in monkeys and cats. Exp. Neurol. 45, 268–287 (1974).
Snider, R. S. & Maiti, A. Cerebellar contributions to the Papez circuit. J. Neurosci. Res. 2, 133–146 (1976).
Newman, P. P. & Reza, H. Functional relationships between the hippocampus and the cerebellum: an electrophysiological study of the cat. J. Physiol. 287, 405–426 (1979).
Saper, C. B. & Loewy, A. D. Efferent connections of the parabrachial nucleus in the rat. Brain Res. 197, 291–317 (1980).
Han, S., Soleiman, M. T., Soden, M. E., Zweifel, L. S. & Palmiter, R. D. Elucidating an affective pain circuit that creates a threat memory. Cell 162, 363–374 (2015).
Norris, A. J., Shaker, J. R., Cone, A. L., Ndiokho, I. B. & Bruchas, M. R. Parabrachial opioidergic projections to preoptic hypothalamus mediate behavioral and physiological thermal defenses. eLife 10, e60779 (2021).
Chiang, M. C. et al. Parabrachial complex: a hub for pain and aversion. J. Neurosci. 39, 8225–8230 (2019).
Barik, A. et al. A spinoparabrachial circuit defined by Tacr1 expression drives pain. eLife 10, e61135 (2021).
Jarvie, B. C., Chen, J. Y., King, H. O. & Palmiter, R. D. Satb2 neurons in the parabrachial nucleus mediate taste perception. Nat. Commun. 12, 224 (2021).
Norgren, R. & Leonard, C. M. Taste pathways in rat brainstem. Science 173, 1136–1139 (1971).
Hashimoto, M. et al. Anatomical evidence for a direct projection from Purkinje cells in the mouse cerebellar vermis to medial parabrachial nucleus. Front. Neural Circuits 12, 6 (2018).
Nisimaru, N. Cardiovascular modules in the cerebellum. Jpn J. Physiol. 54, 431–448 (2004).
Nisimaru, N. & Katayama, S. Projection of cardiovascular afferents to the lateral nodulus-uvula of the cerebellum in rabbits. Neurosci. Res. 21, 343–350 (1995).
Nisimaru, N. et al. Orexin-neuromodulated cerebellar circuit controls redistribution of arterial blood flows for defense behavior in rabbits. Proc. Natl Acad. Sci. USA 110, 14124–14131 (2013).
Oh, S. W. et al. A mesoscale connectome of the mouse brain. Nature 508, 207–214 (2014).
Kebschull, J. M. Cerebellar nuclei evolved by repeatedly duplicating a conserved cell-type set. Science 370, eabd5059 (2020).
Fujita, H., Kodama, T. & du Lac, S. Modular output circuits of the fastigial nucleus for diverse motor and nonmotor functions of the cerebellar vermis. eLife 9, e58613 (2020).
Schwarz, L. A. et al. Viral-genetic tracing of the input-output organization of a central noradrenaline circuit. Nature 524, 88–92 (2015).
Bradley, D. J., Ghelarducci, B., La Noce, A. & Spyer, K. M. Autonomic and somatic responses evoked by stimulation of the cerebellar uvula in the conscious rabbit. Exp. Physiol. 75, 179–186 (1990).
Paton, J. F. & Spyer, K. M. Brain stem regions mediating the cardiovascular responses elicited from the posterior cerebellar cortex in the rabbit. J. Physiol. 427, 533–552 (1990).
Sadakane, K., Kondo, M. & Nisimaru, N. Direct projection from the cardiovascular control region of the cerebellar cortex, the lateral nodulus-uvula, to the brainstem in rabbits. Neurosci. Res. 36, 15–26 (2000).
Khan, M. M., Wu, S., Chen, C. H. & Regehr, W. G. Unusually slow spike frequency adaptation in deep cerebellar nuclei neurons preserves linear transformations on the subsecond timescale. J. Neurosci. 42, 7581–7593 (2022).
Carta, I., Chen, C. H., Schott, A. L., Dorizan, S. & Khodakhah, K. Cerebellar modulation of the reward circuitry and social behavior. Science 363, eaav0581 (2019).
Zingg, B. et al. AAV-mediated anterograde transsynaptic tagging: mapping corticocollicular input-defined neural pathways for defense behaviors. Neuron 93, 33–47 (2017).
Huang, D., Grady, F. S., Peltekian, L. & Geerling, J. C. Efferent projections of Vglut2, Foxp2, and Pdyn parabrachial neurons in mice. J. Comp. Neurol. 529, 657–693 (2021).
Huang, D., Grady, F. S., Peltekian, L., Laing, J. J. & Geerling, J. C. Efferent projections of CGRP/Calca-expressing parabrachial neurons in mice. J. Comp. Neurol. 529, 2911–2957 (2021).
Onat, F. & Cavdar, S. Cerebellar connections: hypothalamus. Cerebellum 2, 263–269 (2003).
Jung, S. J. et al. Novel cerebello-amygdala connections provide missing link between cerebellum and limbic system. Front. Syst. Neurosci. 16, 879634 (2022).
Loughlin, S. E., Foote, S. L. & Bloom, F. E. Efferent projections of nucleus locus coeruleus: topographic organization of cells of origin demonstrated by three-dimensional reconstruction. Neuroscience 18, 291–306 (1986).
Steindler, D. A. Locus coeruleus neurons have axons that branch to the forebrain and cerebellum. Brain Res. 223, 367–373 (1981).
Lein, E. S. et al. Genome-wide atlas of gene expression in the adult mouse brain. Nature 445, 168–176 (2007).
Palmiter, R. D. The parabrachial nucleus: CGRP neurons function as a general alarm. Trends Neurosci. 41, 280–293 (2018).
Chizhikov, V. V. et al. The roof plate regulates cerebellar cell-type specification and proliferation. Development 133, 2793–2804 (2006).
Karthik, S. et al. Molecular ontology of the parabrachial nucleus. J. Comp. Neurol. 530, 1658–1699 (2022).
Mogul, A. S. et al. Cre recombinase driver mice reveal lineage-dependent and -independent expression of Brs3 in the mouse brain. eNeuro 8, ENEURO.0252-21.2021 (2021).
Herbert, H., Guthmann, A., Zafra, F. & Ottersen, O. P. Glycine, glycine receptor subunit and glycine transporters in the rat parabrachial and Kolliker-Fuse nuclei. Anat. Embryol. 201, 259–272 (2000).
Heiney, S. A., Kim, J., Augustine, G. J. & Medina, J. F. Precise control of movement kinematics by optogenetic inhibition of Purkinje cell activity. J. Neurosci. 34, 2321–2330 (2014).
Baek, S. J., Park, J. S., Kim, J., Yamamoto, Y. & Tanaka-Yamamoto, K. VTA-projecting cerebellar neurons mediate stress-dependent depression-like behaviors. eLife 11, e72981 (2022).
Frontera, J. L. et al. The cerebellum regulates fear extinction through thalamo-prefrontal cortex interactions in male mice. Nat. Commun. 14, 1508 (2023).
Hwang, K. D. et al. Cerebellar nuclei neurons projecting to the lateral parabrachial nucleus modulate classical fear conditioning. Cell Rep. 42, 112291 (2023).
Park, E., Ai, J. & Baker, A. J. Cerebellar injury: clinical relevance and potential in traumatic brain injury research. Prog. Brain Res. 161, 327–338 (2007).
Mac Donald, C. L. et al. Detection of blast-related traumatic brain injury in U.S. military personnel. N. Engl. J. Med. 364, 2091–2100 (2011).
Vasterling, J. J., Verfaellie, M. & Sullivan, K. D. Mild traumatic brain injury and posttraumatic stress disorder in returning veterans: perspectives from cognitive neuroscience. Clin. Psychol. Rev. 29, 674–684 (2009).
DelRosso, L. M. & Hoque, R. The cerebellum and sleep. Neurol. Clin. 32, 893–900 (2014).
Moreno-Rius, J. The cerebellum in fear and anxiety-related disorders. Prog. Neuropsychopharmacol. Biol. Psychiatry 85, 23–32 (2018).
Mothersill, O., Knee-Zaska, C. & Donohoe, G. Emotion and theory of mind in schizophrenia-investigating the role of the cerebellum. Cerebellum 15, 357–368 (2016).
Adamaszek, M. et al. Consensus paper: cerebellum and emotion. Cerebellum 16, 552–576 (2017).
Florez-Paz, D., Bali, K. K., Kuner, R. & Gomis, A. A critical role for Piezo2 channels in the mechanotransduction of mouse proprioceptive neurons. Sci. Rep. 6, 25923 (2016).
Grady, F., Peltekian, L., Iverson, G. & Geerling, J. C. Direct parabrachial-cortical connectivity. Cereb. Cortex 30, 4811–4833 (2020).
Paxinos, G. & Franklin, K. B. J. The Mouse Brain in Stereotaxic Coordinates Compact 2nd edn (Elsevier Academic Press, 2004).
Al-Juboori, S. I. et al. Light scattering properties vary across different regions of the adult mouse brain. PLoS ONE 8, e67626 (2013).
Yizhar, O., Fenno, L. E., Davidson, T. J., Mogri, M. & Deisseroth, K. Optogenetics in neural systems. Neuron 71, 9–34 (2011).
Han, K. S., Chen, C. H., Khan, M. M., Guo, C. & Regehr, W. G. Climbing fiber synapses rapidly and transiently inhibit neighboring Purkinje cells via ephaptic coupling. Nat. Neurosci. 23, 1399–1409 (2020).
Conte, W. L., Kamishina, H. & Reep, R. L. Multiple neuroanatomical tract-tracing using fluorescent Alexa Fluor conjugates of cholera toxin subunit B in rats. Nat. Protoc. 4, 1157–1166 (2009).
Guo, C., Rudolph, S., Neuwirth, M. E. & Regehr, W. G. Purkinje cell outputs selectively inhibit a subset of unipolar brush cells in the input layer of the cerebellar cortex. eLife 10, e68802 (2021).
Guo, C. et al. Purkinje cells directly inhibit granule cells in specialized regions of the cerebellar cortex. Neuron 91, 1330–1341 (2016).
Witter, L., Rudolph, S., Pressler, R. T., Lahlaf, S. I. & Regehr, W. G. Purkinje cell collaterals enable output signals from the cerebellar cortex to feed back to Purkinje cells and interneurons. Neuron 91, 312–319 (2016).
Cicconet, M., Hochbaum, D. R., Richmond, D. L. & Sabatini, B. L. 2017 IEEE International Conference on Computer Vision Workshops (ICCVW) (IEEE, 2017).
Turecek, J. & Regehr, W. G. Cerebellar and vestibular nuclear synapses in the inferior olive have distinct release kinetics and neurotransmitters. eLife 9, e61672 (2020).
Turecek, J., Jackman, S. L. & Regehr, W. G. Synaptic specializations support frequency-independent Purkinje cell output from the cerebellar cortex. Cell Rep. 17, 3256–3268 (2016).
Morgan, J. I., Cohen, D. R., Hempstead, J. L. & Curran, T. Mapping patterns of c-fos expression in the central nervous system after seizure. Science 237, 192–197 (1987).
Sheng, M. & Greenberg, M. E. The regulation and function of c-fos and other immediate early genes in the nervous system. Neuron 4, 477–485 (1990).
Marzullo, T. C. & Gage, G. J. The SpikerBox: a low cost, open-source bioamplifier for increasing public participation in neuroscience inquiry. PLoS ONE 7, e30837 (2012).
Kowalczyk, M. S. et al. Single-cell RNA-seq reveals changes in cell cycle and differentiation programs upon aging of hematopoietic stem cells. Genome Res. 25, 1860–1872 (2015).
Acknowledgements
We thank members of the Regehr laboratory and D. Ginty for comments on the manuscript. In particular, we thank B. Miano for assistance with RNAscope experiments. This work was supported by grants from the National Institutes of Health: R01NS032405 and R35NS097284 to W.G.R., K99NS110978 and F32NS101889 to C.H.C., R01DK075632 to B.B.L., U19MH114821 and R01MH130468 to E.Z.M. and NINDS P30 Core Center (NS072030) to the Neurobiology Imaging Center at Harvard Medical School.
Author information
Authors and Affiliations
Contributions
C.H.C. and W.G.R. conceived of the project. C.H.C., L.N.N., A.P.S., K.E.B., B.B.L., E.Z.M. and W.G.R. designed experiments. C.H.C., L.N.N., A.P.S. and D.Z. performed and analyzed in vivo electrophysiology experiments. C.H.C. performed in vitro slice electrophysiology experiments. C.H.C., A.P.S., L.N.N., Z.Y. and I.F. performed and analyzed anatomical/histological experiments. C.H.C., A.P.S., Z.Y. and K.E.B. conducted and analyzed c-Fos functional anatomy experiments. C.H.C., K.E.B., K.M., Z.Y. and L.N.N. performed and analyzed behavioral experiments. C.H.C., A.P.S. and K.E.B. prepared samples for RNA-seq experiments. C.R.V. and N.M.N. isolated and processed samples for RNA-seq, and S.N. analyzed these samples. C.H.C. and W.G.R. wrote the paper with input from all authors.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Neuroscience thanks the anonymous reviewers for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Suppressing PC firing evoked short latency responses in multiple brain regions.
Single-unit, multielectrode array recordings were made from awake, head-restrained PCP2-cre/Halo mice across several areas in the brain. Example single cell responses are shown for the thalamus (a), amygdala (b), septum (c), and basal forebrain (d). Pauses in activity after stimulation were observed in the septum and basal forebrain in a fraction of recorded cells. Example pauses are shown in black. Recording sites are indicated in the gray circles within the corresponding panels on the lower panels. Blue dots indicate responding cells at each site, black dots indicated cells with pauses in firing after stimulation, and open circles indicate nonresponding cells. Sites represent different independent experiments. e, f. Boxplots representing normalized change in firing rate (e) and average latency (f) for each region. For all boxplots: central mark of each box is the median, the edges represent the 25th and 75th percentiles, and the whiskers represent the range of data. LD: laterodorsal, VL: ventrolateral, VP: ventral posterior, CeA: central amygdala, BLA: basolateral amygdala, LS: lateral septum, MS: medial septum, BST: bed nucleus of the stria terminalis, VP: ventral pallidum, SI: substantia innominata, and DB: diagonal band of broca.
Extended Data Fig. 2 Purkinje cell synapses within the PBN.
a. Method for registering PC synapses and identifying the PBN in the PCP2-cre/Synaptophysin-tdTomato mouse. The confocal image (left) showing tdTomato fluorescence, the brightfield image (middle) with the brachium conjunctivum noted, and identified presynaptic boutons (right, yellow) are shown with the bounds of the PBN delineated. b. Average heatmaps of identified PC Synaptophysin-tdT puncta in the PBN. Individual slices were binned at 30 μm2 and averaged by aligning the center of the brachium. Each heatmap is an average of 4 slices (from 2 animals, 2 hemispheres per animal). The bold white line outlines the brachium, and the dotted line the PBN. c. Immunohistochemistry was used to identify inhibitory synapses (vGAT, white), and the locus coeruleus (tyrosine hydroxylase, orange) in a PCP2-cre/tdTomato mouse where PC axons and boutons were labeled (tdT, red). Very few PC boutons were apparent in the locus coeruleus.
Extended Data Fig. 3 PBN neurons rapidly increase firing in response to suppression of PC firing.
a. Single-unit, multielectrode array recordings were made in the PBN or DCN in awake, head-restrained PCP2-cre/Halo mice (n = 6). The posterior cerebellar cortex was stimulated through a thinned skull (20 ms, red light). b. Recording sites were recovered by coating the silicon probe with DiI. An example electrode tract is shown. scp: superior cerebellar peduncle, 4 V: 4th ventricle. C. left, Firing evoked in a rapidly responding PBN neuron ( < 30 ms latency). right, Summary of rapidly responding PBN neurons (13/28 neurons). Shaded area indicates standard error. d. Same as (c) but for slower responding PBN neurons (6/28 neurons). e. Same as (c) but for DCN neurons (16/16 neurons). f. Latencies of PBN neurons (blue) and DCN neurons (red) as a function of evoked firing rate.
Extended Data Fig. 4 Specificity and strategy for suppression of the PC-PBN or PC-DCN pathway.
a. Awake, head-restrained single-unit recordings in the PBN were made using an optrode in a /PCP2-cre/Halo mouse. b. Two example cells showing increases in firing evoked by 100 ms light pulses (red lines). c. Images for an example c-Fos stimulation experiment shown in Fig. 3c. PC inputs to the PBN were suppressed by activating halorhodopsin. In this experiment, unilateral c-Fos upregulation was observed in medial side of the PBN near the tip of the optical fiber(red arrow). Brachium is indicated in the solid gray lines, and bounds of the PBN in dotted lines. d-f. Hindbrain c-Fos densities for (d) PC-PBN suppression, (e) wildtype, and (f) PC-DCN stimulated animals. Average numbers of c-Fos+ cells per mouse/experiment plotted in black lines for the stimulated (i, ipsilateral) and unstimulated (c, contralateral) sides. Individual areas plotted in gray circles. g. Recordings were made from PBN neurons with an optrode. Light was delivered to the immediate region with the attached optical fiber. h. (top) Example single cell response to continuous light delivered to PC-PBN axons. (bottom) Example response of another cell to pulsed (10 Hz, 50 ms pulses) light delivered to PC-PBN axons. i. Summary data showing the firing rate of individual cells in the first and last 5 s of the train for continuous (top) or pulsed (bottom) trains. We used 10 Hz, 50 ms pulses to suppress the PC-PBN in behavioral experiments due to the superior stability of PBN neuron firing. Data are presented as mean values +/- SEM.
Extended Data Fig. 5 Effects of PC-PBN and PC-DCN suppression on motor behaviors.
a-c. PCP2-cre/Halo and wildtype control mice with bilateral optical fiber implants targeting their PBN or DCN were subcutaneously implanted with wires in their backs to measure a field EMG (n = 5). Mice were head-restrained over a freely moving wheel. Speed of the wheel (top) and field EMG (bottom) during stimulation (10 Hz, 5 s), were measured (n = 5). (PBN implant EMG/Wheel n = 5/5, DCN implant EMG/Wheel n = 4/7, control animals EMG/Wheel n = 5/5). d-f. Mice in those groups were also tested in the open field, where 10 Hz optical stimuli were delivered in 60 s intervals. Average position heat maps are shown for each interval (stimulus on and stimulus off). g. Fraction of time in center and average velocity during the open field for the total duration of the test and for the durations while the stimulus was off and on for wildtype control animals. h. As in g, but for PCP2-cre/Halo animals implanted in the PBN. i. As in h, but for PCP2-cre/Halo animals implanted in the DCN. * p < 0.05, see Supplementary Table 4. For all boxplots, central mark of each box is the median, the edges represent the 25th and 75th percentiles, and the whiskers represent the range of data.
Extended Data Fig. 6 Place preference for animals with one optical fiber implanted correctly.
a. Attempted bilateral implants often resulted in one optical fiber implanted in a target region, and another implanted completely elsewhere devoid of halorhodopsin expression. We examined these mice’s behavior in the place preference protocol as in Fig. 3. b. Example (left) and average (right) position heatmaps for corresponding days of the place preference protocol for mice with unilateral implants in the PBN (b, n = 5). c. As in (b), but for unilateral implants in the DCN (c, n = 3). d. Implant locations for all mice with color indicating bottom chamber preference (Test/Baseline; D2/D1). e. Summary of the % time spent in the bottom chamber across all test days for PBN (red) and DCN (blue) unilaterally implanted PCP2-cre/Halo mice. (see Supplementary Table 3).
Extended Data Fig. 7 labeling of the PC-PBN projection pathway.
a. An anterograde AAV-cre was injected into the posterior cerebellar vermis, and AAVs with cre-dependent tdTomato were injected into the PBN. This led to tdT-expression in PC-recipient PBN neurons. The PBN of mouse C was also injected with AAV1-Synaptophysin-YFP to label the presynaptic boutons of all PBN neurons. b. A series of sections shows that tdT expression was restricted to the PBN of the three mice injected as in (a). Approximate location of the LC and PBN is indicated, as determined by the mouse brain atlas and DAPI staining. c. tdT-expressing axons are shown in the amygdala of each mouse. d. tdT-expressing axons are shown in the indicated regions for the indicated mice. e. To visualize PC-PBN projections and determine synaptophysin overlap, 20 μm confocal stacks of each region were taken (left panel), and synaptophysin and axon signals were segmented out (right panels). Puncta associated with a tdTomato-expressing axon are displayed in the examples for the indicated regions.
Extended Data Fig. 8 Suppressing the PC-PBN pathway rapidly increases firing in the septum.
a. In awake, head-restrained mice expressing halorhodopsin in PCs, the PC-PBN pathway was optically suppressed and single-unit recordings were made in the septum. b. Two example cells showing increases in firing during the 100 ms light pulse (red line). c. Optical fiber implant into the parabrachial nuclei. Shown is a sagittal slice of the PBN with cerebellum on the left. Optical fiber implant is indicated with the white arrow. Boundaries of the PBN are indicated with white dotted lines. d. Example recording site through the medial septum (MS) and lateral septum (LS). Probes were coated with different color dyes to enable post-hoc verification of recording sites. Recording sites were determined by matching the dye with the final position of the probe inside the brain. Dye is shown in white. e. Recording sites for all experiments shown above. Every gray circle indicates a recording site. Blue dots indicate responding cells and black dots indicate nonresponding cells for each site.
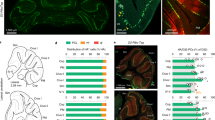
Extended Data Fig. 9 Upregulation of c-Fos following unilateral suppression of the PC-PBN or PC-DCN pathway.
a. Distributions of c-Fos positive cells following inhibition of PC-PBN or PC-DCN pathways. Data corresponds to that shown in Fig. 3a-f and Fig. 4h-m. For wildtype mice hemispheres were combined because there was no apparent difference between hemispheres (see b). * p < 0.05, ** p < 0.01, ***p < 0.001. Gray indicates a decrease in the highlighted comparison. “c” and “i” correspond to contralateral and ipsilateral. For all violin, central mark of each box is the median, the edges represent the 25th and 75th percentiles, and the whiskers represent the range of data. b. Corresponding regions and c-Fos densities for PC-PBN stimulated (top), PC-DCN stimulated (middle), and wildtype (bottom) animals. The average number of c-Fos+ cells in different regions are plotted for each mouse (black lines). Cell counts are plotted for individual slices (gray symbols). See Supplementary Table 1.
Extended Data Fig. 10 RNAseq of brainstem neurons labeled following injections of AAV1-Syn-Cre into the posterior cerebellum of CAG-Sun1/sfGFP mice.
a. tSNE visualization of all cells following fluorescence activated nuclei sorting. Clusters were sorted into cell types based off of expression of markers indicated in b. b. Dot plot of scaled expression for indicated clusters. c. tSNE visualization of just neurons from a. Cluster numbers correspond with those in d. d. Dot plot of scaled expression for indicated clusters ordered from largest to smallest, with number of cells in each cluster indicated in the axis label. Shown are 3 of the most significantly expressed genes for each cluster, along with Slc17a6 (glutamatergic neurons) and Slc32a1 (GABAergic neurons). Clusters with < 100 cells were categorically excluded. Cluster 9 contains Gabra6, a known marker for cerebellar granule cells, thus likely reflects cerebellar contamination, and was excluded from subsequent analysis. e. AAV1-Syn-Cre and AAV1-CAG-tdTomato were coinjected into the posterior vermis of a CAG-Sun1/sfGFP mouse. Slices of the PBN region were made to visualize trans-synaptically labeled neurons with GFP, PC axons with tdTomato, and the locus coeruleus by staining for tyrosine hydroxylase (TH). Two experiments shown in f, g. labeled cells in the LC were far from PC axons, as opposed to labeled PBN neurons, which were close to PC axons. This indicates that most of the labeling of LC neurons is due to retrograde rather than anterograde labeling.
Supplementary information
Supplementary Information
Supplementary Tables 1–4: Statistics related to c-Fos experiments, heart rate experiments, place preference tests and open-field tests.
Supplementary Data 1
Gene expression and clustering for neurons identified in RNA-seq analysis (Fig. 5). Associated code: https://zenodo.org/record/6653204.
Supplementary Data 2
Gene expression and clustering for all cells identified in RNA-seq analysis (Fig. 5). Associated code: https://zenodo.org/record/6653204.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Chen, C.H., Newman, L.N., Stark, A.P. et al. A Purkinje cell to parabrachial nucleus pathway enables broad cerebellar influence over the forebrain. Nat Neurosci 26, 1929–1941 (2023). https://doi.org/10.1038/s41593-023-01462-w
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41593-023-01462-w
- Springer Nature America, Inc.
This article is cited by
-
Neural circuit basis of placebo pain relief
Nature (2024)
-
Markers of positive affect and brain state synchrony discriminate melancholic from non-melancholic depression using naturalistic stimuli
Molecular Psychiatry (2024)
-
Consensus Paper: Cerebellum and Reward
The Cerebellum (2024)