Abstract
Recombinant rabies viral vectors have proven useful for applications including retrograde targeting of projection neurons and monosynaptic tracing, but their cytotoxicity has limited their use to short-term experiments. Here we introduce a new class of double-deletion-mutant rabies viral vectors that left transduced cells alive and healthy indefinitely. Deletion of the viral polymerase gene abolished cytotoxicity and reduced transgene expression to trace levels but left vectors still able to retrogradely infect projection neurons and express recombinases, allowing downstream expression of other transgene products such as fluorophores and calcium indicators. The morphology of retrogradely targeted cells appeared unperturbed at 1 year postinjection. Whole-cell patch-clamp recordings showed no physiological abnormalities at 8 weeks. Longitudinal two-photon structural and functional imaging in vivo, tracking thousands of individual neurons for up to 4 months, showed that transduced neurons did not die but retained stable visual response properties even at the longest time points imaged.






Similar content being viewed by others
References
Wickersham, I. R., Finke, S., Conzelmann, K. K. & Callaway, E. M. Nat. Methods 4, 47–49 (2007).
Wickersham, I. R. et al. Neuron 53, 639–647 (2007).
Wall, N. R., Wickersham, I. R., Cetin, A., De La Parra, M. & Callaway, E. M. Proc. Natl. Acad. Sci. USA 107, 21848–21853 (2010).
Marshel, J. H., Mori, T., Nielsen, K. J. & Callaway, E. M. Neuron 67, 562–574 (2010).
Rancz, E. A. et al. Nat. Neurosci. 14, 527–532 (2011).
Watabe-Uchida, M., Zhu, L., Ogawa, S. K., Vamanrao, A. & Uchida, N. Neuron 74, 858–873 (2012).
Wertz, A. et al. Science 349, 70–74 (2015).
Tian, J. et al. Neuron 91, 1374–1389 (2016).
Wallace, M. L. et al. Neuron 94, 138–152.e5 (2017).
Rompani, S. B. et al. Neuron 93, 1519 (2017).
Liu, K. et al. Nature 548, 582–587 (2017).
Chung, S. et al. Nature 545, 477–481 (2017).
Dölen, G., Darvishzadeh, A., Huang, K. W. & Malenka, R. C. Nature 501, 179–184 (2013).
Kiritani, T., Wickersham, I. R., Seung, H. S. & Shepherd, G. M. J. Neurosci. 32, 4992–5001 (2012).
Kress, G. J. et al. Nat. Neurosci. 16, 665–667 (2013).
Namburi, P. et al. Nature 520, 675–678 (2015).
Rajasethupathy, P. et al. Nature 526, 653–659 (2015).
Yamawaki, N., Suter, B. A., Wickersham, I. R. & Shepherd, G. M. Cold Spring Harb. Protoc. 2016, t090084 (2016).
Shin Yim, Y. et al. Nature 549, 482–487 (2017).
Osakada, F. et al. Neuron 71, 617–631 (2011).
Gong, Y. et al. Science 350, 1361–1366 (2015).
Roland, B. et al. eLife https://doi.org/10.7554/eLife.16335.001 (2016).
Wickersham, I. R., Sullivan, H. A. & Seung, H. S. Nat. Commun. 4, 2332 (2013).
Reardon, T. R. et al. Neuron 89, 711–724 (2016).
Ciabatti, E., González-Rueda, A., Mariotti, L., Morgese, F. & Tripodi, M. Cell 170, 382–392.e14 (2017).
Albertini, A. A., Ruigrok, R. W. & Blondel, D. Adv. Virus. Res. 79, 1–22 (2011).
Madore, H. P. & England, J. M. J. Virol. 22, 102–112 (1977).
Schnell, M. J., McGettigan, J. P., Wirblich, C. & Papaneri, A. Nat. Rev. Microbiol. 8, 51–61 (2010).
Cormack, B. P., Valdivia, R. H. & Falkow, S. Gene 173, 33–38 (1996).
Sauer, B. & Henderson, N. Proc. Natl. Acad. Sci. USA 85, 5166–5170 (1988).
Sadowski, P. D. Prog. Nucleic Acid Res. Mol. Biol. 51, 53–91 (1995).
Raymond, C. S. & Soriano, P. PLoS ONE 2, e162 (2007).
Shaner, N. C. et al. Nat. Biotechnol. 22, 1567–1572 (2004).
Fenno, L. E. et al. Nat. Methods 11, 763–772 (2014).
Schwarz, L. A. et al. Nature 524, 88–92 (2015).
Gomme, E. A., Wirblich, C., Addya, S., Rall, G. F. & Schnell, M. J. PLoS. Pathog. 8, e1002971 (2012).
Ohki, K., Chung, S., Ch’ng, Y. H., Kara, P. & Reid, R. C. Nature 433, 597–603 (2005).
Chen, T. W. et al. Nature 499, 295–300 (2013).
Ohki, K. & Reid, R. C. Cold Spring Harb. Protoc. 2014, 402–416 (2014).
Soudais, C., Laplace-Builhe, C., Kissa, K. & Kremer, E. J. FASEB J. 15, 2283–2285 (2001).
Tervo, D. G. et al. Neuron 92, 372–382 (2016).
Senn, V. et al. Neuron 81, 428–437 (2014).
Kohara, K. et al. Nat. Neurosci. 17, 269–279 (2014).
Xu, C. et al. Cell 167, 961–972.e16 (2016).
Schwarz, L. A. & Luo, L. Curr. Biol. 25, R1051–R1056 (2015).
Wu, Z., Yang, H. & Colosi, P. Mol. Ther. 18, 80–86 (2010).
Dong, B., Nakai, H. & Xiao, W. Mol. Ther. 18, 87–92 (2010).
Finke, S., Cox, J. H. & Conzelmann, K. K. J. Virol. 74, 7261–7269 (2000).
Tian, L. et al. Nat. Methods 6, 875–881 (2009).
Finke, S. & Conzelmann, K. K. J. Virol. 73, 3818–3825 (1999).
Finke, S., Mueller-Waldeck, R. & Conzelmann, K. K. J. Gen. Virol. 84, 1613–1621 (2003).
Paterson, R. G., Russell, C. J. & Lamb, R. A. Virology 270, 17–30 (2000).
Matsuda, T. & Cepko, C. L. Proc. Natl. Acad. Sci. USA 101, 16–22 (2004).
Marr, R. A. et al. J. Mol. Neurosci. 22, 5–11 (2004).
Niwa, H., Yamamura, K. & Miyazaki, J. Gene 108, 193–199 (1991).
Atasoy, D., Aponte, Y., Su, H. H. & Sternson, S. M. J. Neurosci. 28, 7025–7030 (2008).
Shaner, N. C. et al. Nat. Methods 5, 545–551 (2008).
Koresawa, Y. et al. J. Biochem. 127, 367–372 (2000).
Wickersham, I. R., Sullivan, H. A. & Seung, H. S. Nat. Protoc. 5, 595–606 (2010).
Wickersham, I. R. & Sullivan, H. A. Cold Spring Harb. Protoc. 2015, 375–385 (2015).
Wickersham, I. R. et al. Cold Spring Harb. Protoc. 2015, 368–374 (2015).
Madisen, L. et al. Nat. Neurosci. 13, 133–140 (2010).
Madisen, L. et al. Neuron 85, 942–958 (2015).
Tallquist, M. D. & Soriano, P. Genesis 26, 113–115 (2000).
Mayford, M. et al. Science 274, 1678–1683 (1996).
Husson, T. R., Mallik, A. K., Zhang, J. X. & Issa, N. P. J. Neurosci. 27, 8665–8675 (2007).
Andermann, M. L., Kerlin, A. M., Roumis, D. K., Glickfeld, L. L. & Reid, R. C. Neuron 72, 1025–1039 (2011).
Pologruto, T. A., Sabatini, B. L. & Svoboda, K. Biomed. Eng. Online 2, 13 (2003).
Thévenaz, P., Ruttimann, U. E. & Unser, M. IEEE Trans. Image. Process. 7, 27–41 (1998).
Peng, H., Bria, A., Zhou, Z., Iannello, G. & Long, F. Nat. Protoc. 9, 193–208 (2014).
Peirce, J. W. J. Neurosci. Methods 162, 8–13 (2007).
Pnevmatikakis, E. A. et al. Neuron 89, 285–299 (2016).
Hnasko, T. S. et al. Proc. Natl. Acad. Sci. USA 103, 8858–8863 (2006).
Garrett, M. E., Nauhaus, I., Marshel, J. H. & Callaway, E. M. J. Neurosci. 34, 12587–12600 (2014).
Allen Institute for Brain Science. Allen Mouse Brain Connectivity Atlas Overview http://help.brain-map.org/download/attachments/2818171/Connectivity_Overview.pdf (2017).
Oh, S. W. et al. Nature 508, 207–214 (2014).
Acknowledgements
S.C. thanks D. Williams and S. de Vries for software, A. Cheng and N. Orlova for hardware construction, and R. D’Aleo and N. Ouellette for experimental assistance. J.A.H. thanks K.E. Hirokawa and P. Bohn for technical assistance. I.R.W. thanks W. Salmon for assistance with confocal imaging. The authors thank the founders of the Allen Institute for Brain Science, Paul G. Allen and Jody Allen, for their vision, encouragement and support. Research reported in this publication was supported primarily by the National Institute of Mental Health under award number U01MH106018 (BRAIN Initiative) to I.R.W. and in part by the National Institute on Aging under award number R01AG047589 to J.A.H. and the National Eye Institute under award numbers R01EY010115 and R01EY018742 to R.C.R. K.M.T. was supported in part by a New York Stem Cell Foundation Robertson Investigator award and McKnight Scholar Award, and G.G.C. was supported by a JFDP fellowship and a NARSAD Young Investigator Award. The content of this paper is solely the responsibility of the authors and does not necessarily represent the official views of the National Institutes of Health.
Author information
Authors and Affiliations
Contributions
S.C. helped design experiments, performed two-photon imaging and quantified and analyzed the results, assisted by B.J.M. and overseen by R.C.R. H.A.S. produced all RV vectors, assisted by S.D. R.X. and Y.H. cloned all RV vectors. T.K.L. and N.E.L. injected RV vectors in vivo for anatomical experiments in reporter mice and, in combination with FLEX AAV in rats; they also processed the tissue for these experiments, assisted by J.E.M. T.K.L. also conducted confocal imaging. K.R.B. bred mice for the anatomical experiments and assisted with cloning viral vectors. G.A.M., A.B., and G.G.C. performed ex vivo physiological recordings, G.G. injected vectors and retrobeads for these ex vivo experiments, and K.M.T. oversaw and helped plan them. J.D.W. and J.A.H. conducted experiments (injections, histology, imaging and analysis) comparing RV with rAAV2-retro and CAV2-Cre and using RV in combination with FLEX AAV in mice; H.Z. oversaw these comparative and combinatorial experiments, and S.Y. and A.C. produced the AAV2-retro stocks. I.R.W. conceived and designed the study; I.R.W. and S.C. wrote the manuscript with input from other authors.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Integrated supplementary information
Supplementary Figure 1 RVΔGL-Cre can be used in combination with Cre-dependent AAV for intersectional targeting of projection neurons in mice.
(a) RVΔGL-Cre injection site from a paired injection experiment in an Ai75 (Cre-dependent nuclear tdTomato) mouse. Center of injection site is enlarged in panel b. RVΔGL-Cre was injected into the retrosplenial cortex (RSP) so that cells across the brain which provide input to RSP were labeled via Cre-induced nuclear tdTomato expression (magenta). (c) AAV-FLEX-EGFP was injected into anterior cingulate cortex (ACA) so cells that were co-infected with RVΔGL-Cre also express EGFP. Several EGFP+ cell bodies in cortical layers 2/3 and 5 are visible in this section, as well as axonal projections in contralateral ACA and the striatum (STR). EGFP-labeled projections from ACA can be seen in the lateral dorsal (LD) and central lateral (CL) thalamic nuclei while cells retrogradely labeled from RSP are present only in LD (a). Axonal projections from ACA neurons are visible in the RVΔGL-Cre injection site (arrowheads in b indicate some axons). (d) Segmented pixels from this experiment projected onto a cortical surface view of the Allen Institute Common Coordinate Framework show the extent and location of cortical projections labeled with this paired injection strategy. The approximate locations of the AAV-FLEX-EGFP and RVΔGL-Cre injections are indicated, as well as the rostral-caudal location of the coronal sections in panels (a) and (c). Images were acquired by serial 2-photon tomography. Scale bars: a, c = 1 mm; b = 500 μm. This experiment was repeated with similar results for 8 different pairs of injection coordinates (n = 2 for the specific ACA-RSP coordinates shown) in a total of 15 mice.
Supplementary Figure 2 RVΔGL-Cre can be used in combination with Cre-dependent AAV for intersectional targeting of projection neurons in rats.
(a) mCherry-labeled layer 6 corticothalamic cells in barrel cortex labeled via RVΔGL-Cre injected in ventral posteromedial thalamus and AAV-FLEX-mCherry injected in barrel cortex. (b) mCherry-labeled thalamocortical neurons in ventral posteromedial thalamus labeled via RVΔGL-Cre injected in barrel cortex and AAV-FLEX-mCherry injected in ventral posteromedial thalamus. (c) Axons expressing ChrimsonR-tdTomato (Klapoetke, N.C. et al. Independent optical excitation of distinct neural populations. Nat. Methods 11, 338-46 (2014)) in barrel cortex of thalamocortical cells labeled via RVΔGL-Cre injected in barrel cortex and AAV-FLEX-ChrimsonR-tdTomato injected in ventral posteromedial thalamus. (d) ChrimsonR-tdTomato-expressing axons in ventral posteromedial thalamus labeled via RVΔGL-Cre injected in ventral posteromedial thalamus and AAV-FLEX-ChrimsonR-tdTomato injected in barrel cortex. Experiments in this figure were performed four times for each condition with similar results in all cases. Scale bar: 66 μm, applies to all panels.
Supplementary Figure 3 Longitudinal in vivo two-photon structural and functional imaging in RVΔG-tdTomato- and RVΔG-GCaMP6f-injected mice.
(a–f) Structural imaging from layer 2/3 of V1 in a wild-type mouse injected with first-generation vector RVΔG-tdTomato. Panels (a–c) are sequential images of one field of view (FOV) over time; panels (d–f) are sequential images of a different FOV. (a,d) 9 days after injection, most neurons appear to maintain their structural integrity with no outward signs of toxicity. (b,e) By 22 days postinjection, there is a large dropoff in visible cells, and remaining processes are strongly blebbed and assume a more punctate appearance. Somata begin to lose their shape. (c,f) At 34 days postinjection, almost all cells are gone. An example of a surviving cell at this time point (f) shows extremely dysmorphic processes and soma. This was consistent across our data set of 6 FOVs, so structural imaging for the RVΔG-tdTomato condition was discontinued at week 4. Scale bars for (a–f): 20 μm. (g–i) Calcium imaging from layer 2/3 of V1 in a wild-type mouse injected with first-generation vector RVΔG-GCaMP6f. (g) 5 days after injection, neurons are still visually responsive to drifting gratings (mean ∆F/F ± s.e.m., averaged over 15 repeats; see Methods), but mean ∆F/F responses are not as robust as in case of the second-generation vector. (h) The number of surviving cells drops substantially by 11 days postinjection, and those that remain show even weaker orientation selectivity and smaller mean responses. (i) By day 29, only dim puncta and speckle can be seen, with no cell bodies visible. The white box shows where the FOV in (a) overlaps with that in (c). This was consistent across our data set of 6 FOVs, so cells labeled with RVΔG-GCaMP6f were not included in the analysis of long-term activity and visual responses. Labels on leftmost tuning plots apply to all plots. Scale bar for (g–i): 100 μm.
Supplementary Figure 4 Long-term stability of orientation and temporal frequency tuning in RVΔGL-Cre-labeled neurons.
Visual responses of cells taken from Fig. 5a (lower FOV) are plotted for three different time points. Each panel (a–d) shows the orientation tuning of one cell at 7 (top), 54 (middle), and 122 days (bottom row) postinjection, obtained with drifting gratings presented at 8 directions of motion and 5 temporal frequencies (TF) (same stimuli as in Fig. 5; mean ∆F/F ± s.e.m., averaged over 15 repeats). The FOVs of the three imaging sessions (e–g) show the cells circled and labeled with their respective panel letter. Both the orientation tuning curves and temporal frequency preferences remain strikingly similar over four months of imaging. This was consistent across our data set of 9 FOVs. Axis labels in (a) apply to all plots. Scale bar: 100 μm.
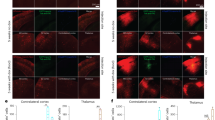
Supplementary Figure 5 Different patterns of cortical input cells labeled by Cre reporter after injections of three retrogradely transported Cre viruses into the anteromedial visual cortex (AM).
CAV2-Cre (a,b), RVΔGL-Cre (c,d), or AAV-retro-EF1a-Cre (e,f) was injected into AM of Ai75 mice. The same brain regions containing retrogradely labeled cells from AM were identified in all three types of virus experiments. However, as with ACA, within these cortical input areas, we again observed differences in the layer location of labeled cells. For example, labeled input cells in contralateral AM (boxes in row i indicate panels enlarged in row ii) were seen predominantly in L5, with scattered labeling in L6 for CAV2-Cre (a(ii), b(ii)). RVΔGL-Cre (c(ii), d(ii)) and rAAV2-retro-EF1a-Cre (e(ii), f(ii)) resulted in a more uniform distribution of labeled cells across layers. Similar differences in layer patterns were observed in most cortical areas, including posteromedial visual cortex (PM, a(iii)–e(iii) and a(iv)–e(iv)) and anterolateral visual cortex (AL; a(iii)–e(iii), a(v)–e(v)). Quantification of the number of labeled cells per layer in each region indicated that, in all cases, CAV2-Cre labeling is biased toward L5 neurons, whereas RVΔGL-Cre results in cells labeled across all layers that are known to contain neurons with long-range inter-areal projections (L2/3, L5, L6; a(vi)–d(vi)). Cre+ input cells were also observed in subcortical structures, including several thalamic nuclei (e.g. lateral posterior nucleus (LP, seen in row i), consistent with known connectivity of AM. The more prominent labeling in CAV2-Cre may be due to true differences in efficiency between viruses, or in the amount of viral uptake for each experiment; further experiments are necessary to explore this observation. Images in a-b were acquired by serial 2-photon tomography. Images in (c–f) were acquired by epifluorescence microscopy and sections counterstained with DAPI (blue). Graphs in (a(vi)–c(vi)) show the mean and full range of values (min to max, n = 2 independent experiments for each virus). Boxplots in (d(vi)) indicate the median and full range (min to max) in all three regions from (a(vi)–c(vi)) (n = 6 independent experiments). Scale bars: rows i,iii = 1 mm; ii, iv, v = 500 μm.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–5
Supplementary Video S1
Volume renderings of mouse visual cortical neurons imaged at 9 days (left) and 22 days (right) following injection of a first-generation rabies viral vector encoding tdTomato. The vast majority of labeled neurons visible at the earlier time point are gone by the later imaging session.
Supplementary Video S2
Volume renderings of mouse visual cortical neurons imaged at 10 days (left) and 19 days (right) following injection of a first-generation rabies viral vector encoding Cre, in a tdTomato reporter mouse line. Although many cells remain at the later time point, many have disappeared.
Supplementary Video S3
Volume renderings of mouse visual cortical neurons imaged at 8 days (left) and 16 days (right) following injection of a second-generation rabies viral vector encoding Cre. All neurons visible at the earlier time point are still present at the later one, and indeed more neurons are clearly visible at the later time point than at the earlier one.
Supplementary Video S4
Volume renderings of tdTomato-labeled mouse visual cortical neurons imaged at 4 weeks (left) and 8 weeks (right) following injection of a second-generation rabies viral vector encoding Cre. All neurons visible at the earlier time point are still present at the later one.
Supplementary Video S5
Stable calcium responses to visual stimuli of GCaMP6f-labeled neurons in layer 2/3 of mouse primary visual cortex, imaged at 18 days (top left), 34 days (top right), 68 days (bottom left), and 124 days (bottom right) following injection of a second-generation rabies viral vector encoding Cre. Neurons visible at the earlier time points are still present at later ones; visual responses remain stable at even the longest time point imaged. Each frame is 250 ms of imaging data, 500 frames total, sped up to 15 fps.
Supplementary Video S6
Abnormal calcium responses to visual stimuli of GCaMP6f-labeled neurons in layer 2/3 of mouse primary visual cortex, imaged at 55 days (top) and 57 days (bottom) following injection of a first-generation rabies viral vector encoding Cre. Neuron indicated by top arrow shows a slow plateau of brightness (comes on around 3 seconds into the video), not calcium transients associated with spiking (same idea as Fig. 5e bottom trace, different cell). Middle arrow, bright-inactive cell. Bottom arrow, bright-inactive cell with obvious morphological signs of unhealthiness (blebbing processes).
Rights and permissions
About this article
Cite this article
Chatterjee, S., Sullivan, H.A., MacLennan, B.J. et al. Nontoxic, double-deletion-mutant rabies viral vectors for retrograde targeting of projection neurons. Nat Neurosci 21, 638–646 (2018). https://doi.org/10.1038/s41593-018-0091-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41593-018-0091-7
- Springer Nature America, Inc.
This article is cited by
-
A flowchart for adequate controls in virus-based monosynaptic tracing experiments identified Cre-independent leakage of the TVA receptor in RΦGT mice
BMC Neuroscience (2024)
-
Current advance of nanotechnology in diagnosis and treatment for malignant tumors
Signal Transduction and Targeted Therapy (2024)
-
New rabies viral resources for multi-scale neural circuit mapping
Molecular Psychiatry (2024)
-
Long-term labeling and imaging of synaptically connected neuronal networks in vivo using double-deletion-mutant rabies viruses
Nature Neuroscience (2024)
-
ENGEP: advancing spatial transcriptomics with accurate unmeasured gene expression prediction
Genome Biology (2023)