Abstract
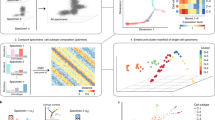
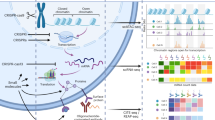
Cells can be perturbed by various chemical and genetic treatments and the impact on gene expression and morphology can be measured via transcriptomic profiling and image-based assays, respectively. The patterns observed in these high-dimensional profile data can power a dozen applications in drug discovery and basic biology research, but both types of profiles are rarely available for large-scale experiments. Here, we provide a collection of four datasets with both gene expression and morphological profile data useful for developing and testing multimodal methodologies. Roughly a thousand features are measured for each of the two data types, across more than 28,000 chemical and genetic perturbations. We define biological problems that use the shared and complementary information in these two data modalities, provide baseline analysis and evaluation metrics for multi-omic applications, and make the data resource publicly available (https://broad.io/rosetta/).




Similar content being viewed by others
Data availability
Preprocessed profiles that are augmented with gene and compound annotation are freely available through the ‘Registry of Open Data on AWS’ on a public S3 bucket. Documentation on the folder structure, dataset details and instructions for accessing the data are available at https://broad.io/rosetta/. Datasets are described and referenced in Supplementary Data 1. Source data are provided with this paper.
Code availability
Source code to reproduce and build upon the presented results is available at https://broad.io/rosetta/. We licensed the source code as BSD 3-Clause, and licensed the data, results and figures as CC0 1.0.
References
Moshkov, N. et al. Predicting compound activity from phenotypic profiles and chemical structures. Preprint at bioRxiv https://www.biorxiv.org/content/10.1101/2020.12.15.422887v4 (2022).
Breinig, M., Klein, F. A., Huber, W. & Boutros, M. A chemical–genetic interaction map of small molecules using high-throughput imaging in cancer cells. Mol. Syst. Biol. 11, 846 (2015).
Rohban, M. H. et al. Systematic morphological profiling of human gene and allele function via Cell Painting. Elife 6, e24060 (2017).
Meng, T., Jing, X., Yan, Z. & Pedrycz, W. A survey on machine learning for data fusion. Inf. Fusion 57, 115–129 (2020).
Baldwin, E. et al. On fusion methods for knowledge discovery from multi-omics datasets. Comput. Struct. Biotechnol. J. 18, 509–517 (2020).
Patel-Murray, N. L. et al. A multi-omics interpretable machine learning model reveals modes of action of small molecules. Sci. Rep. 10, 954 (2020).
Subramanian, A. et al. A next generation connectivity map: L1000 platform and the first 1,000,000 profiles. Cell 171, 1437–1452 (2017).
Bray, M. -A. et al. Cell Painting, a high-content image-based assay for morphological profiling using multiplexed fluorescent dyes. Nat. Protoc. 11, 1757–1774 (2016).
McQuin, C. et al. CellProfiler 3.0: next-generation image processing for biology. PLoS Biol. 16, e2005970 (2018).
Wawer, M. J. et al. Toward performance-diverse small-molecule libraries for cell-based phenotypic screening using multiplexed high-dimensional profiling. Proc. Natl Acad. Sci. USA 111, 10911–10916 (2014).
Berger, A. H. et al. High-throughput phenotyping of lung cancer somatic mutations. Cancer Cell 30, 214–228 (2016).
Haftbaradaran Esfahani, P. & Knöll, R. Cell shape: effects on gene expression and signaling. Biophys. Rev. 12, 895–901 (2020).
Drareni, K., Gautier, J.-F., Venteclef, N. & Alzaid, F. Transcriptional control of macrophage polarisation in type 2 diabetes. Semin. Immunopathol. 41, 515–529 (2019).
Mota de Sá, P., Richard, A. J., Hang, H. & Stephens, J. M. Transcriptional regulation of adipogenesis. Compr. Physiol. 7, 635–674 (2017).
Way, G. P. et al. Morphology and gene expression profiling provide complementary information for mapping cell state. Preprint at bioRxiv https://www.biorxiv.org/content/10.1101/2021.10.21.465335 (2022).
Nassiri, I. & McCall, M. N. Systematic exploration of cell morphological phenotypes associated with a transcriptomic query. Nucleic Acids Res. 46, e116 (2018).
Spiering, D. & Hodgson, L. Dynamics of the Rho-family small GTPases in actin regulation and motility. Cell Adh. Migr. 5, 170–180 (2011).
Dai Yang, K. et al. Multi-domain translation between single-cell imaging and sequencing data using autoencoders. Nat. Commun. 12, 31 (2021).
Gundersen, G., Dumitrascu, B. & Ash, J. T. End-to-end training of deep probabilistic CCA on paired biomedical observations. In Proceedings of PMLR pp.945–955 (2019).
He, B. et al. Integrating spatial gene expression and breast tumour morphology via deep learning. Nat. Biomed. Eng. 4, 827–834 (2020).
Pasquer, Q. T. L., Tsakoumagkos, I. A. & Hoogendoorn, S. From phenotypic hit to chemical probe: Chemical biology approaches to elucidate small molecule action in complex biological systems. Molecules 25, 5702 (2020).
Rohban, M. H. et al. Virtual screening for small-molecule pathway regulators by image-profile matching. Cell Syst. 13, 724–736 (2022).
Ljosa, V. et al. Comparison of methods for image-based profiling of cellular morphological responses to small-molecule treatment. J. Biomol. Screen. 18, 1321–1329 (2013).
Warchal, S. J., Dawson, J. C. & Carragher, N. O. Evaluation of machine learning classifiers to predict compound mechanism of action when transferred across distinct cell lines. SLAS Discov. 24, 224–233 (2019).
Aliper, A. et al. Deep learning applications for predicting pharmacological properties of drugs and drug repurposing using transcriptomic data. Mol. Pharm. 13, 2524–2530 (2016).
Lapins, M. & Spjuth, O. Evaluation of gene expression and phenotypic profiling data as quantitative descriptors for predicting drug targets and mechanisms of action. Preprint at bioRxiv https://doi.org/10.1101/580654 (2019).
Cantini, L. et al. Benchmarking joint multi-omics dimensionality reduction approaches for the study of cancer. Nat. Commun. 12, 124 (2021).
Tenenhaus, M., Tenenhaus, A. & Groenen, P. J. F. Regularized generalized canonical correlation analysis: a framework for sequential multiblock component methods. Psychometrika https://doi.org/10.1007/s11336-017-9573-x (2017).
Roselle, C., Verch, T. & Shank-Retzlaff, M. Mitigation of microtiter plate-positioning effects using a block randomization scheme. Anal. Bioanal. Chem. 408, 3969–3979 (2016).
Lill, J. R., Mathews, W. R., Rose, C. M. & Schirle, M. Proteomics in the pharmaceutical and biotechnology industry: a look to the next decade. Expert Rev. Proteom. 18, 503–526 (2021).
Chandrasekaran, S. N., Ceulemans, H., Boyd, J. D. & Carpenter, A. E. Image-based profiling for drug discovery: due for a machine learning upgrade? Nat. Rev. Drug Discov. 20, 145–159 (2021).
Tandon, G., Chan, P. & Mitra, D. MORPHEUS: motif oriented representations to purge hostile events from unlabeled sequences. in Proceedings of the 2004 ACM Workshop on Visualization and Data Mining for Computer Security https://doi.org/10.1145/1029208.1029212 (2004).
Broad Institute. Guide to LINCS data release into NCBI GEO—L1000. Connectopedia. https://clue.io/connectopedia/guide_to_geo_l1000_data
Huang, D. W., Sherman, B. T. & Lempicki, R. A. Systematic and integrative analysis of large gene lists using DAVID bioinformatics resources. Nat. Protoc. 4, 44–57 (2009).
Acknowledgements
We thank all the researchers who created and shared the data, who are mentioned in their respective publications cited in the paper. Funding was provided by grants (2018-183451 to A.E.C. and 2020-225720 to B.A.C.) from the Chan Zuckerberg Initiative DAF, an advised fund of the Silicon Valley Community Foundation and the National Institutes of Health NIGMS (R35 GM122547, to A.E.C.).
Author information
Authors and Affiliations
Contributions
M.H., S.S., B.A.C. and A.E.C. contributed to drafting the manuscript and designing the research. J.C.C. initiated the project and performed early explorations of the LUAD dataset. M.H. analyzed and explored the data with inputs from the other co-authors.
Corresponding authors
Ethics declarations
Competing interests
S.S. and A.E.C. serve as scientific advisors for companies that use image-based profiling and Cell Painting (A.E.C.: Recursion, S.S.: Waypoint Bio, Dewpoint Therapeutics), and receive honoraria for occasional talks at pharmaceutical and biotechnology companies.
Peer review
Peer review information
Nature Methods thanks Haiquan Li, Matthew McCall, and the other, anonymous, reviewer for their contribution to the peer review of this work. Primary Handling Editor: Rita Strack, in collaboration with the Nature Methods team. Peer reviewer reports are available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Generalizability of the prediction model across datasets.
Prediction of each L1000 mRNA level by Cell Painting features in dataset A, using a model trained on dataset B. We have trained Lasso and MLP models on each of LUAD and LINCS datasets and checked the prediction results on the other dataset which was not used in model training. Distribution of R2 prediction scores for all landmark genes are shown. Comparison of the results here with Fig. 2 indicates weakness of the prediction model in generalizability across datasets. This is an indication of dataset-specific technical variations (batch effects) that need exploration of experimental alignment techniques (batch-effect correction), which is an active area of research. We also observe that the model’s prediction power is stronger when the model is trained on the LINCS dataset and tested on the LUAD dataset. This is expected as the LUAD dataset is limited to a narrow set of genes associated with lung adenocarcinoma cancer; however, the LINCS dataset contains a wide variety of compounds with different mechanisms and known phenotypes. The y-axis is trimmed at −1 for clarity. Distributions are presented as boxplots, with center line being median, box limits being upper and lower quartiles and whiskers being 1.5× interquartile range; n = 978 landmark genes for each boxplot.
Extended Data Fig. 2 Gene group names for top 100 predictable landmark genes in LUAD dataset.
Top 100 predictable landmark genes by MLP model are shown along with their gene group names (based on HGNC Database41) for the LUAD dataset, finding a diverse array represented, though we note the perturbations in this experiment included only genes found mutated in lung cancers.
Extended Data Fig. 3 Over-Representation Analysis (ORA) of highly predictable (top 100) landmark genes in LUAD dataset.
Over-Representation Analysis of top 100 highly predictable landmark genes according to the MLP model applied on the LUAD dataset. ORA analysis was performed by WebGestalt analysis toolkit 42. Nineteen enriched categories (FDR < 0.2) are labeled in the volcano plot.
Extended Data Fig. 4 Visualization of cells in a cluster of landmark genes that are tightly correlated with RNA texture category of morphological features.
For the cluster of landmark genes shown in the top heatmap, which is a partial snapshot of Fig. 2d, we have shown example cell images for perturbations that have high and low predicted values for each gene in that cluster. We have filtered perturbations to those that have low prediction errors prior to that selection. We can observe that cells that are predicted to have (and actually do have) high levels of these five genes’ mRNA all are associated with visible changes in the staining for mitochondria, even though only half of these genes already have functional annotations related to the mitochondria.
Extended Data Fig. 5 Validation of the observed GE-CP relationship by GO-terms search analysis.
Landmark genes highly predictable according to morphological features in each specific Cell Painting channel are more likely to have GO annotation related to that channel compared to the rest of CP channels. For each channel in the rows of the table, the first column shows the Odds Ratio (OR) derived from the Fisher’s exact test for associations between the landmark genes being highly predictable (R2 > 0.6) by CP features in a channel and having GO annotations for that channel. The second column shows the association between the same set of highly predictable genes and having GO annotation for any channel but not the target row channel. Higher values in the first column compared to the second column show that highly predictable genes according to features in a CP channel are more likely to have GO annotations for that channel compared to the rest of the channels. This pattern holds for DNA and ER channels but not for the rest of CP channels. The third and fourth columns show the same associations but for low-predictability genes (R2 < 0). Lower values in the third column compared to the fourth column show that non-predictable genes according to features in a CP channel are less likely to have GO annotations for that channel compared to the rest of the channels. This pattern holds for all CP channels except for RNA. The CP channel specific predictability map used for this analysis was derived from the result of the experiment and results presented partially in Fig. 2d. As we can observe from the map, usually multiple categories of morphological features contribute to the predictability of a gene, which explains the lack of a simple relationship between a given channel’s predictability and GO term associations presented in this table.
Extended Data Fig. 6 Association between landmark gene predictability and having gene ontology annotations related to Cell-Painting stains.
Landmark genes that are predictable according to at least three of the four datasets (59 genes shown in Fig. 2c) are more likely to have GO annotations related to any of the stains in the Cell Painting assay compared to a random subset of landmark genes.
Supplementary information
Supplementary Information
Supplementary Data 1–3 and 5, Supplementary Table 1 and Supplementary Figs. 1–3
Supplementary Data 4
Median prediction scores for each landmark gene across each dataset and model.
Source data
Source Data Fig. 2
Fig. 2a: Prediction score values per gene in each dataset and model. Single scores in rows of the table indicate the median of the predictability scores across all cross-validation folds. Fig. 2b: Extracted predictability threshold for each dataset and number of predictable genes. Fig. 2c: List of the genes for each area of the Venn diagram. Fig. 2d: Source numbers for predictability score, shown as a heat map.
Source Data Fig. 3
Fig. 3a: Prediction score values for each single morphological CP feature in each dataset and for MLP models. Each score in each row is the median of the predictability scores across all cross-validation folds in its corresponding experiment. Fig. 3b: Ratio of predictable features to total number of features in each cell-paining feature category. Fig. 3d: Values for relative importance of each feature, shown as bar plots.
Source Data Fig. 4
Fig. 4a: Jaccard Index values for each of MOA clusters for each model and each dataset and data modality. Fig. 4b: F1-score values for MOA classification experiments for each modality, integration method, model, dataset and cross-validation fold. Fig. 4c: MoA class-specific classification F1-scores, shown partially as a bar plot.
Source Data Extended Data Fig. 1
Prediction scores for each individual gene resulted from a model trained on one dataset and used to predict landmark GEs in another dataset. LUAD and LINCS datasets were used.
Source Data Extended Data Fig. 2
A table as shown in Extended Data Fig. 2.
Source Data Extended Data Fig. 3
ORA analysis results (set size, enrichment ratio, P value and false discovery rate), which are partially shown in Extended Data Fig. 3.
Source Data Extended Data Fig. 5
Numbers as shown in Extended Data Table. 5.
Source Data Extended Data Fig. 6
Numbers as shown in Extended Data Table. 6.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Haghighi, M., Caicedo, J.C., Cimini, B.A. et al. High-dimensional gene expression and morphology profiles of cells across 28,000 genetic and chemical perturbations. Nat Methods 19, 1550–1557 (2022). https://doi.org/10.1038/s41592-022-01667-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41592-022-01667-0
- Springer Nature America, Inc.
This article is cited by
-
High-dimensional phenotyping to define the genetic basis of cellular morphology
Nature Communications (2024)
-
Orientation-invariant autoencoders learn robust representations for shape profiling of cells and organelles
Nature Communications (2024)
-
Progress in toxicogenomics to protect human health
Nature Reviews Genetics (2024)
-
VIBRANT: spectral profiling for single-cell drug responses
Nature Methods (2024)
-
Autocorrelation analysis of a phenotypic screen reveals hidden drug activity
Scientific Reports (2024)