Abstract
N6-methyladenosine (m6A) is the most prevalent modification of messenger RNA in mammals. To interrogate its functions and dynamics, there is a critical need to quantify m6A at three levels: site, gene and sample. Current approaches address these needs in a limited manner. Here we develop m6A-seq2, relying on multiplexed m6A-immunoprecipitation of barcoded and pooled samples. m6A-seq2 allows a big increase in throughput while reducing technical variability, requirements of input material and cost. m6A-seq2 is furthermore uniquely capable of providing sample-level relative quantitations of m6A, serving as an orthogonal alternative to mass spectrometry-based approaches. Finally, we develop a computational approach for gene-level quantitation of m6A. We demonstrate that using this metric, roughly 30% of the variability in RNA half life in mouse embryonic stem cells can be explained, establishing m6A as a main driver of RNA stability. m6A-seq2 thus provides an experimental and analytic framework for dissecting m6A-mediated regulation at three different levels.



Similar content being viewed by others
Data availability
All the m6A-seq/2 datasets that were generated by us are stored at GSE178832. The GSE accession number of datasets that were not generated by us are listed in Supplementary Table 11. Source data are provided with this paper.
Code availability
The used in-house python script for demultiplexing as the script (bam2ReadEnds.R) used for paired-end transcript coverage are stored at https://github.com/SchwartzLab.
References
Perry, R. P. & Kelley, D. E. Existence of methylated messenger RNA in mouse L cells. Cell 1, 37–42 (1974).
Desrosiers, R., Friderici, K. & Rottman, F. Identification of methylated nucleosides in messenger RNA from Novikoff hepatoma cells. Proc. Natl Acad. Sci. USA 71, 3971–3975 (1974).
Dominissini, D. et al. Topology of the human and mouse m6A RNA methylomes revealed by m6A-seq. Nature 485, 201–206 (2012).
Meyer, K. D. et al. Comprehensive analysis of mRNA methylation reveals enrichment in 3′ UTRs and near stop codons. Cell 149, 1635–1646 (2012).
Wang, X. et al. N6-methyladenosine-dependent regulation of messenger RNA stability. Nature 505, 117–120 (2014).
Wang, X. et al. N(6)-methyladenosine modulates messenger RNA translation efficiency. Cell 161, 1388–1399 (2015).
Roundtree, I. A. et al. YTHDC1 mediates nuclear export of N6-methyladenosine methylated mRNAs. eLife 6, e31311 (2017).
Shi, H. et al. YTHDF3 facilitates translation and decay of N6-methyladenosine-modified RNA. Cell Res. 27, 315–328 (2017).
Ke, S. et al. m6A mRNA modifications are deposited in nascent pre-mRNA and are not required for splicing but do specify cytoplasmic turnover. Genes Dev. 31, 990–1006 (2017).
Li, A. et al. Cytoplasmic m6A reader YTHDF3 promotes mRNA translation. Cell Res. 27, 444–447 (2017).
Liu, N. et al. N(6)-methyladenosine-dependent RNA structural switches regulate RNA-protein interactions. Nature 518, 560–564 (2015).
Batista, P. J. et al. m(6)A RNA modification controls cell fate transition in mammalian embryonic stem cells. Cell Stem Cell 15, 707–719 (2014).
Cui, Q. et al. m6A RNA methylation regulates the self-renewal and tumorigenesis of glioblastoma stem cells. Cell Rep. 18, 2622–2634 (2017).
Rosa-Mercado, N. A., Withers, J. B. & Steitz, J. A. Settling the m6A debate: methylation of mature mRNA is not dynamic but accelerates turnover. Genes Dev. 31, 957–958 (2017).
Darnell, R. B., Ke, S. & Darnell, J. E. Jr. Pre-mRNA processing includes N6 methylation of adenosine residues that are retained in mRNA exons and the fallacy of ‘RNA epigenetics’. RNA 24, 262–267 (2018).
Zhao, B. S., Nachtergaele, S., Roundtree, I. A. & He, C. Our views of dynamic N6-methyladenosine RNA methylation. RNA 24, 268–272 (2018).
Zaccara, S. & Jaffrey, S. R. A unified model for the function of YTHDF proteins in regulating m6A-modified mRNA. Cell 181, 1582–1595.e18 (2020).
Molinie, B. et al. m(6)A-LAIC-seq reveals the census and complexity of the m(6)A epitranscriptome. Nat. Methods 13, 692–698 (2016).
Legrand, C. et al. Statistically robust methylation calling for whole-transcriptome bisulfite sequencing reveals distinct methylation patterns for mouse RNAs. Genome Res. 27, 1589–1596 (2017).
Sas-Chen, A. et al. Dynamic RNA acetylation revealed by quantitative cross-evolutionary mapping. Nature 583, 638–643 (2020).
Schwartz, S. m1A within cytoplasmic mRNAs at single nucleotide resolution: a reconciled transcriptome-wide map. RNA 24, 1427–1436 (2018).
Garcia-Campos, M. A. et al. Deciphering the ‘m6A code’ via antibody-independent quantitative profiling. Cell 178, 731–747.e16 (2019).
Schwartz, S. et al. Perturbation of m6A writers reveals two distinct classes of mRNA methylation at internal and 5′ sites. Cell Rep. 8, 284–296 (2014).
McIntyre, A. B. R. et al. Limits in the detection of m6A changes using MeRIP/m6A-seq. Sci. Rep. 10, 6590 (2020).
Schwartz, S. et al. High-resolution mapping reveals a conserved, widespread, dynamic mRNA methylation program in yeast meiosis. Cell 155, 1409–1421 (2013).
Agarwala, S. D., Blitzblau, H. G., Hochwagen, A. & Fink, G. R. RNA methylation by the MIS complex regulates a cell fate decision in yeast. PLoS Genet. 8, e1002732 (2012).
Clancy, M. J., Shambaugh, M. E., Timpte, C. S. & Bokar, J. A. Induction of sporulation in Saccharomyces cerevisiae leads to the formation of N6-methyladenosine in mRNA: a potential mechanism for the activity of the IME4 gene. Nucleic Acids Res. 30, 4509–4518 (2002).
Zhuang, F., Fuchs, R. T., Sun, Z., Zheng, Y. & Robb, G. B. Structural bias in T4 RNA ligase-mediated 3′-adapter ligation. Nucleic Acids Res. 40, e54 (2012).
Fuchs, R. T., Sun, Z., Zhuang, F. & Robb, G. B. Bias in ligation-based small RNA sequencing library construction is determined by adaptor and RNA structure. PLoS ONE 10, e0126049 (2015).
Shishkin, A. A. et al. Simultaneous generation of many RNA-seq libraries in a single reaction. Nat. Methods 12, 323–325 (2015).
Roth, S. H., Levanon, E. Y. & Eisenberg, E. Genome-wide quantification of ADAR adenosine-to-inosine RNA editing activity. Nat. Methods 16, 1131–1138 (2019).
Geula, S. et al. m6A mRNA methylation facilitates resolution of naïve pluripotency toward differentiation. Science 347, 1002–1006 (2015).
Ries, R. J. et al. m6A enhances the phase separation potential of mRNA. Nature 571, 424–428 (2019).
Anders, M. et al. Dynamic m6A methylation facilitates mRNA triaging to stress granules. Life Sci. Alliance 1, e201800113 (2018).
Lasman, L. et al. Context-dependent functional compensation between Ythdf m6A reader proteins. Genes Dev. 34, 1373–1391 (2020).
Gokhale, N. S. et al. Altered m6A modification of specific cellular transcripts affects flaviviridae infection. Mol. Cell 77, 542–555.e8 (2020).
Mauer, J. et al. Reversible methylation of m6Am in the 5′ cap controls mRNA stability. Nature 541, 371–375 (2016).
Akichika, S. et al. Cap-specific terminal N6-methylation of RNA by an RNA polymerase II-associated methyltransferase. Science 363, eaav0080 (2019).
Boulias, K. et al. Identification of the m6Am methyltransferase PCIF1 reveals the location and functions of m6Am in the transcriptome. Mol. Cell 75, 631–643.e8 (2019).
Sun, H., Zhang, M., Li, K., Bai, D. & Yi, C. Cap-specific, terminal N6-methylation by a mammalian m6Am methyltransferase. Cell Res. 29, 80–82 (2019).
Sendinc, E. et al. PCIF1 catalyzes m6Am mRNA methylation to regulate gene expression. Mol. Cell 75, 620–630.e9 (2019).
Cowling, V. H. CAPAM: the mRNA cap adenosine N6-methyltransferase. Trends Biochem. Sci. 44, 183–185 (2019).
Zhao, B. S., Roundtree, I. A. & He, C. Post-transcriptional gene regulation by mRNA modifications. Nat. Rev. Mol. Cell Biol. 18, 31–42 (2017).
Zaccara, S., Ries, R. J. & Jaffrey, S. R. Reading, writing and erasing mRNA methylation. Nat. Rev. Mol. Cell Biol. 20, 608–624 (2019).
Schwartz, S. Cracking the epitranscriptome. RNA 22, 169–174 (2016).
Linder, B. et al. Single-nucleotide-resolution mapping of m6A and m6Am throughout the transcriptome. Nat. Methods 12, 767–772 (2015).
Grozhik, A. V., Linder, B., Olarerin-George, A. O. & Jaffrey, S. R. Mapping m6A at individual-nucleotide resolution using crosslinking and immunoprecipitation (miCLIP). Methods Mol. Biol. 1562, 55–78 (2017).
Liu, H. et al. Accurate detection of m6A RNA modifications in native RNA sequences. Nat. Commun. 10, 4079 (2019).
Meyer, K. D. DART-seq: an antibody-free method for global m6A detection. Nat. Methods 16, 1275–1280 (2019).
Wang, Y., Xiao, Y., Dong, S., Yu, Q. & Jia, G. Antibody-free enzyme-assisted chemical approach for detection of N6-methyladenosine. Nat. Chem. Biol. 16, 896–903 (2020).
Shu, X. et al. A metabolic labeling method detects m6A transcriptome-wide at single base resolution. Nat. Chem. Biol. 16, 887–895 (2020).
Zhang, Z. et al. Single-base mapping of m6A by an antibody-independent method. Sci. Adv. 5, eaax0250 (2019).
Drexler, H. L., Choquet, K. & Churchman, L. S. Splicing kinetics and coordination revealed by direct nascent RNA sequencing through nanopores. Mol. Cell 77, 985–998.e8 (2020).
Lorenz, D. A., Sathe, S., Einstein, J. M. & Yeo, G. W. Direct RNA sequencing enables m6A detection in endogenous transcript isoforms at base-specific resolution. RNA 26, 19–28 (2020).
Zhang, Z. et al. RADAR: differential analysis of MeRIP-seq data with a random effect model. Genome Biol. 20, 294 (2019).
Wen, J. et al. Zc3h13 regulates nuclear RNA m6A methylation and mouse embryonic stem cell self-renewal. Mol. Cell 69, 1028–1038.e6 (2018).
Longtine, M. S. et al. Additional modules for versatile and economical PCR-based gene deletion and modification in Saccharomyces cerevisiae. Yeast 14, 953–961 (1998).
Chia, M. et al. High-resolution analysis of cell-state transitions in yeast suggests widespread transcriptional tuning by alternative starts. Genome Biol. 22, 34 (2021).
Nishimura, K., Fukagawa, T., Takisawa, H., Kakimoto, T. & Kanemaki, M. An auxin-based degron system for the rapid depletion of proteins in nonplant cells. Nat. Methods 6, 917–922 (2009).
Chu, S. & Herskowitz, I. Gametogenesis in yeast is regulated by a transcriptional cascade dependent on Ndt80. Mol. Cell 1, 685–696 (1998).
Levi-Galibov, O. et al. Heat Shock Factor 1-dependent extracellular matrix remodeling mediates the transition from chronic intestinal inflammation to colon cancer. Nat. Commun. 11, 6245 (2020).
Ran, F. A. et al. Genome engineering using the CRISPR-Cas9 system. Nat. Protoc. 8, 2281–2308 (2013).
Stelzer, Y., Shivalila, C. S., Soldner, F., Markoulaki, S. & Jaenisch, R. Tracing dynamic changes of DNA methylation at single-cell resolution. Cell 163, 218–229 (2015).
Safra, M. et al. The m1A landscape on cytosolic and mitochondrial mRNA at single-base resolution. Nature 551, 251–255 (2017).
Dobin, A. et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics 29, 15–21 (2013).
Li, H. et al. The sequence alignment/map format and SAMtools. Bioinformatics 25, 2078–2079 (2009).
Heinz, S. et al. Simple combinations of lineage-determining transcription factors prime cis-regulatory elements required for macrophage and B cell identities. Mol. Cell 38, 576–589 (2010).
Martin, M. Cutadapt removes adapter sequences from high-throughput sequencing reads. EMBnet. J. 17, 10–12 (2011).
Li, B. & Dewey, C. N. RSEM: accurate transcript quantification from RNA-seq data with or without a reference genome. BMC Bioinformatics 12, 323 (2011).
Daley, T. & Smith, A. D. Predicting the molecular complexity of sequencing libraries. Nat. Methods 10, 325–327 (2013).
Acknowledgements
D.D., M.A.G.C., A.U., M.S., S.E. and S.S. have received funding from the Israel Science Foundation (grant no. 543165), the European Research Council (ERC) under the European Union’s Horizon 2020 research and innovation programme (grant no. 714023 and ERC-POC 899122), the Estate of Emile Mimran. S.S. is the incumbent of the Robert Edward and Roselyn Rich Manson Career Development Chair in Perpetuity. R.S.S. is supported by ISF grant nos. 401/17 and 1384/1, ERC grant no. 754320, and the Laura Gurwin Flug Family Fund. R.S.S. is the incumbent of the Ernst and Kaethe Ascher Career Development Chair in Life Sciences. F.v.W., T.S., R.A.V. and A.R. are supported by the Francis Crick Institute, which receives its core funding from Cancer Research UK (grant no. FC001203), the UK Medical Research Council (grant no. FC001203) and the Wellcome Trust (grant no. FC001203).
Author information
Authors and Affiliations
Contributions
D.D. and S.S. conceived the project. M.S., S.E., A.U. and D.D. conducted all genomic experiments. A.B. performed the conducted mass-spectroscopy analysis. F.v.W., T.S., R.A.V. and A.R. generated and conducted the experiment of yeast inducible perturbations. A.U. and Y.S. conducted the experiments of the partial KO experiments of m6A writers in mESC. M.A.G.C. provided and created the in-house generated scripts that were used to perform the analysis. D.D. and S.S. performed the data analysis. R.S.S. and S.S. acquired funding. D.D., R.S.S. and S.S. wrote the paper, with input from all authors.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Nature Methods thanks Christopher Mason and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Lei Tang was the primary editor on this article and managed its editorial process and peer review in collaboration with the rest of the editorial team.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 m6A-seq2 reduces batch-induced variability while maintaining m6A-enrichment capacities.
m6A-seq2 reduces batch-induced variability while maintaining m6A-enrichment capacities a) Sequencing complexity metric based on the ratio of distinct- to total read numbers for 500,000 sampled reads (Supplementary Table 11) b) m6A-IP and Input normalized coverage profile of MEI5 for the 12 m6A-seq2 replicates and the 3 m6A-seq replicates c) Metagene coverage densities of m6A-IP and Input for the 12 m6A-seq2 replicates (red) and 3 m6A-seq replicates (black). d) Motif analysis showing the most significant HOMER (Supplementary Table 11) de novo motif analysis, based on detected peaks (see Methods) of the 12 m6A-seq2 replicates (red) and 3 m6A-seq replicates (black) e) m6A-seq2 number of reads of the 12 technical replicates. f) Identity of nucleotide of the first position of read 2 (top), which resembles the 3′ terminus of a sequenced RNA fragment. Stacked barplot of the mean nucleotide identity of the first 30 positions (bottom). g) Principal component analysis of m6A-site score 2 of 486 m6A sites. h Cumulative plot showing the fraction of false-positive detected ‘differential-methylation’ cases and the effect-size as the absolute fold-change of the mean score1 between the sampled groups.
Extended Data Fig. 2 m6A-seq2 reduces batch-induced variability in technical replicates of mouse embryonic fibroblasts.
m6A-seq2 reduces batch-induced variability in technical replicates of mouse embryonic fibroblasts. a) Library complexity metric as the ratio of distinct- to total read numbers based on one million reads of the m6A-seq2 and m6A-seq MEF dataset (preseq70) b) Motif analysis showing the most significant HOMER (Supplementary Table 11) de novo motif analysis, based on detected peaks (see Methods) of the m6A-seq2 technical replicates (left) and m6A-seq technical replicates (right). c) Metagene m6A-peak distribution (see Methods) of the m6A-seq technical replicates (top) and m6A-seq2 replicates (bottom). d) m6A-site score 1 comparison between technical replicates generated via m6A-seq or m6A-seq2. score2 for 3416 m6A-sites passing the coverage thresholds for high-confidence m6A site estimation (see Methods). e) Principal component analysis of m6A-site score 1 of 3416 high-confidence m6A sites. f) Comparison of the percentage coefficient of variation (%CV) of score1 (left) and score2 (right) estimates across the technical replicates measured via m6A-seq and m6A-seq2. Wilcoxon two-sided test p value annotated. Comparison of n = 3416 m6A-sites for score1 and score 2. Boxplot: Center is the median, lower and upper hinges depict the first and third quartile and the whiskers stretch to 1.5 times the interquartile range from the corresponding hinge. g) Cumulative plot showing the fraction of false-positive detected ‘differential-methylation’ cases and the effect-size as the absolute fold-change of the mean score between the sampled groups.
Extended Data Fig. 3 m6A-methylome profiling upon genetic perturbations during yeast meiosis.
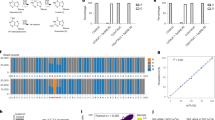
m6A-methylome profiling upon genetic perturbations during yeast meiosis a) m6A-SI based on m6A-Seq2 measurements on individual genetic perturbations strains with ndt80Δ/Δ background. Barplot for each of the m6A-seq2 batches. In total 3 gene deletion strains (pho92Δ/Δ, not3Δ/Δ, gis2Δ/Δ), 12 genes with an auxin-inducible degrons (AID), 5 positive controls (only ndt80Δ/Δ, red) and 5 negative controls (ime4Δ/Δ & ndt80Δ/Δ, blue) (noted in legend) and AID-control (TIR1). U: untreated samples and T: treatment with CU and IAA (see Methods). b) Raf1 m6A-IP fragment coverage normalized by sample library size and Raf1 expression level (RPM) based on the input sample for all 24 samples (% amount of WT mESC RNA per triplicate is annotated at the right). The Y-axis is fixed to an identical range across all samples. c) Heatmap of log2 transformed m6A-site score 1 of 4505 annotated high-confidence m6A-sites (scaled by row) of the samples (merged triplicates).
Extended Data Fig. 4 m6A-GI infers with half-life and steady-state expression.
m6A-GI infers with half-life and steady-state expression a) m6A-GI of the mESC WT cells plotted against mRNA half-life in Mettl3-KO mESC b) HPLC-MS abundances of m6A across WT and m6A-writer perturbed mESC clones plotted against the ‘expression-vs-methylation index’, defined as the spearman correlation between normalized expression (TPM, see Methods) and m6A-GIs calculated based on 3 previously published m6A-seq datasets23. Annotated is the Pearson’s R coefficient and p-value (two-sided analysis) and 95 % confidence interval.
Extended Data Fig. 5 m6A-gene index correlates with the increase of m6A.
m6A-gene index correlates with the increase of m6A a) m6A-gene index (m6A-GI) of 100 % WT sample (merged triplicates) plotted against the other sample concentrations (merged triplicates) with a linear model fit (blue line). b) Scatterplot of the slope of the linear modeling fit of all unique combinations of m6A-GIs derived from different % WT mESC (as in a)), plotted against the log2 transformed fold-change (FC) of the corresponding unique combination out of the 7 samples (14 % WT to 100 % WT). Annotated is the Pearson’s R coefficient and p-value (two-sided analysis) and 95 % confidence interval c) Heatmap of the log2 transformed m6A-GIs (scaled by gene) for 6572 genes of the different samples (merged triplicates). d) Scatter plots of the m6A index of five distinct in vitro transcribed spikes (IS1-5, see Supplementary Table 1) with a defined amount of m6A ranging from 0% m6A (in red) over 7 increasing concentrations to 100% (in black). Linear-modeling and Pearson’s R determined and p-value (two-sided analysis) for all data points with m6A (black) annotated. Spikes with 0% methylation are plotted in red, and were not taken into account for the linear model.
Supplementary information
Supplementary Tables 1–12.
Contains all Supplementary Tables in different tabs.
Rights and permissions
About this article
Cite this article
Dierks, D., Garcia-Campos, M.A., Uzonyi, A. et al. Multiplexed profiling facilitates robust m6A quantification at site, gene and sample resolution. Nat Methods 18, 1060–1067 (2021). https://doi.org/10.1038/s41592-021-01242-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41592-021-01242-z
- Springer Nature America, Inc.