Abstract
The APOE4 allele is the strongest genetic risk factor for late-onset Alzheimer’s disease (AD). The contribution of microglial APOE4 to AD pathogenesis is unknown, although APOE has the most enriched gene expression in neurodegenerative microglia (MGnD). Here, we show in mice and humans a negative role of microglial APOE4 in the induction of the MGnD response to neurodegeneration. Deletion of microglial APOE4 restores the MGnD phenotype associated with neuroprotection in P301S tau transgenic mice and decreases pathology in APP/PS1 mice. MGnD–astrocyte cross-talk associated with β-amyloid (Aβ) plaque encapsulation and clearance are mediated via LGALS3 signaling following microglial APOE4 deletion. In the brains of AD donors carrying the APOE4 allele, we found a sex-dependent reciprocal induction of AD risk factors associated with suppression of MGnD genes in females, including LGALS3, compared to individuals homozygous for the APOE3 allele. Mechanistically, APOE4-mediated induction of ITGB8–transforming growth factor-β (TGFβ) signaling impairs the MGnD response via upregulation of microglial homeostatic checkpoints, including Inpp5d, in mice. Deletion of Inpp5d in microglia restores MGnD–astrocyte cross-talk and facilitates plaque clearance in APP/PS1 mice. We identify the microglial APOE4–ITGB8–TGFβ pathway as a negative regulator of microglial response to AD pathology, and restoring the MGnD phenotype via blocking ITGB8–TGFβ signaling provides a promising therapeutic intervention for AD.








Similar content being viewed by others
Data availability
Our Smart-seq2 RNA-seq, scRNA-seq and ChIP–seq data that support the findings of this study have been deposited into Gene Expression Omnibus (GEO) under SuperSeries GSE239603. The raw data from Olah et al. are available through Synapse (https://www.synapse.org/#!Synapse:syn21438358). The raw data from Zhou et al. are also available through Synapse (https://doi.org/10.7303/syn21125841). The raw data from Zhang et al. are available in GEO under accession code GSE52564. The raw data from Lund et al. are available in GEO through accession code GSE111385. The raw data from Wong et al. are available in GEO through accession code GSE84148.
Code availability
Customized code is available at https://github.com/The-Butovsky-Lab/APOE4-impairs-the-microglial-response-in-Alzheimer-disease-by-inducing-TGFB-mediated-checkpoint.
References
Jansen, I. E. et al. Genome-wide meta-analysis identifies new loci and functional pathways influencing Alzheimer’s disease risk. Nat. Genet. 51, 404–413 (2019).
Wightman, D. P. et al. A genome-wide association study with 1,126,563 individuals identifies new risk loci for Alzheimer’s disease. Nat. Genet. 53, 1276–1282 (2021).
Nott, A. et al. Brain cell type-specific enhancer–promoter interactome maps and disease-risk association. Science 366, 1134–1139 (2019).
Krasemann, S. et al. The TREM2–APOE pathway drives the transcriptional phenotype of dysfunctional microglia in neurodegenerative diseases. Immunity 47, 566–581 (2017).
Keren-Shaul, H. et al. A unique microglia type associated with restricting development of Alzheimer’s disease. Cell 169, 1276–1290 (2017).
Farrer, L. A. et al. Effects of age, sex, and ethnicity on the association between apolipoprotein E genotype and Alzheimer disease. A meta-analysis. APOE and Alzheimer Disease Meta Analysis Consortium. JAMA 278, 1349–1356 (1997).
Corder, E. H. et al. Gene dose of apolipoprotein E type 4 allele and the risk of Alzheimer’s disease in late onset families. Science 261, 921–923 (1993).
Strittmatter, W. J. et al. Apolipoprotein E: high-avidity binding to β-amyloid and increased frequency of type 4 allele in late-onset familial Alzheimer disease. Proc. Natl Acad. Sci. USA 90, 1977–1981 (1993).
Raber, J. et al. Isoform-specific effects of human apolipoprotein E on brain function revealed in ApoE knockout mice: increased susceptibility of females. Proc. Natl Acad. Sci. USA 95, 10914–10919 (1998).
Youmans, K. L. et al. APOE4-specific changes in Aβ accumulation in a new transgenic mouse model of Alzheimer disease. J. Biol. Chem. 287, 41774–41786 (2012).
Liu, D. S. et al. APOE4 enhances age-dependent decline in cognitive function by down-regulating an NMDA receptor pathway in EFAD-Tg mice. Mol. Neurodegener. 10, 7 (2015).
Wang, C. et al. Selective removal of astrocytic APOE4 strongly protects against tau-mediated neurodegeneration and decreases synaptic phagocytosis by microglia. Neuron 109, 1657–1674 (2021).
Butovsky, O. et al. Identification of a unique TGF-β-dependent molecular and functional signature in microglia. Nat. Neurosci. 17, 131–143 (2014).
Margeta, M. A. et al. Apolipoprotein E4 impairs the response of neurodegenerative retinal microglia and prevents neuronal loss in glaucoma. Immunity 55, 1627–1644 (2022).
Parkhurst, C. N. et al. Microglia promote learning-dependent synapse formation through brain-derived neurotrophic factor. Cell 155, 1596–1609 (2013).
Goldmann, T. et al. Origin, fate and dynamics of macrophages at central nervous system interfaces. Nat. Immunol. 17, 797–805 (2016).
Cugurra, A. et al. Skull and vertebral bone marrow are myeloid cell reservoirs for the meninges and CNS parenchyma. Science 373, eabf7844 (2021).
Gosselin, D. et al. Environment drives selection and function of enhancers controlling tissue-specific macrophage identities. Cell 159, 1327–1340 (2014).
Huang, K. L. et al. A common haplotype lowers PU.1 expression in myeloid cells and delays onset of Alzheimer’s disease. Nat. Neurosci. 20, 1052–1061 (2017).
Serrano-Pozo, A., Frosch, M. P., Masliah, E. & Hyman, B. T. Neuropathological alterations in Alzheimer disease. Cold Spring Harb. Perspect. Med. 1, a006189 (2011).
Shi, Y. et al. ApoE4 markedly exacerbates tau-mediated neurodegeneration in a mouse model of tauopathy. Nature 549, 523–527 (2017).
Yoshiyama, Y. et al. Synapse loss and microglial activation precede tangles in a P301S tauopathy mouse model. Neuron 53, 337–351 (2007).
Rosenzweig, N. et al. PD-1/PD-L1 checkpoint blockade harnesses monocyte-derived macrophages to combat cognitive impairment in a tauopathy mouse model. Nat. Commun. 10, 465 (2019).
Radde, R. et al. Aβ42-driven cerebral amyloidosis in transgenic mice reveals early and robust pathology. EMBO Rep. 7, 940–946 (2006).
Bennett, F. C. et al. A combination of ontogeny and CNS environment establishes microglial identity. Neuron 98, 1170–1183 (2018).
Van Hove, H. et al. A single-cell atlas of mouse brain macrophages reveals unique transcriptional identities shaped by ontogeny and tissue environment. Nat. Neurosci. 22, 1021–1035 (2019).
Mrdjen, D. et al. High-dimensional single-cell mapping of central nervous system immune cells reveals distinct myeloid subsets in health, aging, and disease. Immunity 48, 380–395 (2018).
Jordao, M. J. C. et al. Single-cell profiling identifies myeloid cell subsets with distinct fates during neuroinflammation. Science 363, eaat7554 (2019).
Habib, N. et al. Disease-associated astrocytes in Alzheimer’s disease and aging. Nat. Neurosci. 23, 701–706 (2020).
Munger, J. S. & Sheppard, D. Cross talk among TGF-β signaling pathways, integrins, and the extracellular matrix. Cold Spring Harb. Perspect. Biol. 3, a005017 (2011).
Arnold, T. D. et al. Impaired αVβ8 and TGFβ signaling lead to microglial dysmaturation and neuromotor dysfunction. J. Exp. Med. 216, 900–915 (2019).
Wolf, Y., Anderson, A. C. & Kuchroo, V. K. TIM3 comes of age as an inhibitory receptor. Nat. Rev. Immunol. 20, 173–185 (2020).
Nakao, A. et al. TGF-β receptor-mediated signalling through Smad2, Smad3 and Smad4. EMBO J. 16, 5353–5362 (1997).
Olah, M. et al. Single cell RNA sequencing of human microglia uncovers a subset associated with Alzheimer’s disease. Nat. Commun. 11, 6129 (2020).
Zhou, Y. et al. Human and mouse single-nucleus transcriptomics reveal TREM2-dependent and TREM2-independent cellular responses in Alzheimer’s disease. Nat. Med. 26, 131–142 (2020).
Travis, M. A. et al. Loss of integrin αvβ8 on dendritic cells causes autoimmunity and colitis in mice. Nature 449, 361–365 (2007).
Nakawesi, J. et al. αvβ8 integrin-expression by BATF3-dependent dendritic cells facilitates early IgA responses to rotavirus. Mucosal Immunol. 14, 53–67 (2021).
Zhang, Y. et al. An RNA-sequencing transcriptome and splicing database of glia, neurons, and vascular cells of the cerebral cortex. J. Neurosci. 34, 11929–11947 (2014).
Liu, Z. et al. Fate mapping via Ms4a3-expression history traces monocyte-derived cells. Cell 178, 1509–1525 (2019).
Wong, K. et al. Mice deficient in NRROS show abnormal microglial development and neurological disorders. Nat. Immunol. 18, 633–641 (2017).
Samuels, J. D. et al. The Alzheimer’s disease risk factor INPP5D restricts neuroprotective microglial responses in amyloid β-mediated pathology. Alzheimer’s Dement. https://doi.org/10.1002/alz.13089 (2023).
Iguchi, A. et al. INPP5D modulates TREM2 loss-of-function phenotypes in a β-amyloidosis mouse model. iScience 26, 106375 (2023).
Dodagatta-Marri, E. et al. Integrin αvβ8 on T cells suppresses anti-tumor immunity in multiple models and is a promising target for tumor immunotherapy. Cell Rep. 36, 109309 (2021).
Mahan, T. E. et al. Selective reduction of astrocyte ApoE3 and ApoE4 strongly reduces Aβ accumulation and plaque-related pathology in a mouse model of amyloidosis. Mol. Neurodegener. 17, 13 (2022).
Rohn, T. T. & Moore, Z. D. Nuclear localization of apolipoprotein E4: a new trick for an old protein. Int. J. Neurol. Neurother. 4, 067 (2017).
Theendakara, V. et al. Direct transcriptional effects of apolipoprotein E. J. Neurosci. 36, 685–700 (2016).
Hollingworth, P. et al. Common variants at ABCA7, MS4A6A/MS4A4E, EPHA1, CD33 and CD2AP are associated with Alzheimer’s disease. Nat. Genet. 43, 429–435 (2011).
Pimenova, A. A. et al. Alzheimer’s-associated PU.1 expression levels regulate microglial inflammatory response. Neurobiol. Dis. 148, 105217 (2021).
Gosselin, D. et al. An environment-dependent transcriptional network specifies human microglia identity. Science 356, eaal3222 (2017).
Liddelow, S. A. et al. Neurotoxic reactive astrocytes are induced by activated microglia. Nature 541, 481–487 (2017).
Katsouri, L. et al. Ablation of reactive astrocytes exacerbates disease pathology in a model of Alzheimer’s disease. Glia 68, 1017–1030 (2020).
Boza-Serrano, A. et al. Galectin-3, a novel endogenous TREM2 ligand, detrimentally regulates inflammatory response in Alzheimer’s disease. Acta Neuropathol. 138, 251–273 (2019).
Tao, C. C. et al. Galectin-3 promotes Aβ oligomerization and Aβ toxicity in a mouse model of Alzheimer’s disease. Cell Death Differ. 27, 192–209 (2020).
Mabuchi, F. et al. The apolipoprotein E gene polymorphism is associated with open angle glaucoma in the Japanese population. Mol. Vis. 11, 609–612 (2005).
Lam, C. Y. et al. Association of apolipoprotein E polymorphisms with normal tension glaucoma in a Chinese population. J. Glaucoma 15, 218–222 (2006).
Margeta, M. A. et al. Association of APOE with primary open-angle glaucoma suggests a protective effect for APOE ε4. Invest. Ophthalmol. Vis. Sci. 61, 3 (2020).
Tesseur, I. et al. Deficiency in neuronal TGF-β signaling promotes neurodegeneration and Alzheimer’s pathology. J. Clin. Invest. 116, 3060–3069 (2006).
Wyss-Coray, T. et al. Amyloidogenic role of cytokine TGF-β1 in transgenic mice and in Alzheimer’s disease. Nature 389, 603–606 (1997).
Kandasamy, M. et al. TGF-β signaling: a therapeutic target to reinstate regenerative plasticity in vascular dementia? Aging Dis. 11, 828–850 (2020).
Yin, Z. et al. Identification of a protective microglial state mediated by miR-155 and interferon-λ signaling in a mouse model of Alzheimer’s disease. Nat. Neurosci. 26, 1196–1207 (2023).
Lund, H. et al. Fatal demyelinating disease is induced by monocyte-derived macrophages in the absence of TGF-β signaling. Nat. Immunol. https://doi.org/10.1038/s41590-018-0091-5 (2018).
Kaiser, T. & Feng, G. Tmem119-EGFP and Tmem119-CreERT2 transgenic mice for labeling and manipulating microglia. eNeuro 6, ENEURO.0448-18.2019 (2019).
Wang, J. W. et al. Influence of SHIP on the NK repertoire and allogeneic bone marrow transplantation. Science 295, 2094–2097 (2002).
Ju, W. et al. Deletion of Smad2 in mouse liver reveals novel functions in hepatocyte growth and differentiation. Mol. Cell. Biol. 26, 654–667 (2006).
Li, Q. et al. Redundant roles of SMAD2 and SMAD3 in ovarian granulosa cells in vivo. Mol. Cell. Biol. 28, 7001–7011 (2008).
Huynh, T. V. et al. Lack of hepatic APOE does not influence early Aβ deposition: observations from a new APOE knock-in model. Mol. Neurodegener. 14, 37 (2019).
Sullivan, P. M. et al. Targeted replacement of the mouse apolipoprotein E gene with the common human APOE3 allele enhances diet-induced hypercholesterolemia and atherosclerosis. J. Biol. Chem. 272, 17972–17980 (1997).
Knouff, C. et al. Apo E structure determines VLDL clearance and atherosclerosis risk in mice. J. Clin. Invest. 103, 1579–1586 (1999).
Iwasaki, H. et al. Distinctive and indispensable roles of PU.1 in maintenance of hematopoietic stem cells and their differentiation. Blood 106, 1590–1600 (2005).
Proctor, J. M., Zang, K., Wang, D., Wang, R. & Reichardt, L. F. Vascular development of the brain requires β8 integrin expression in the neuroepithelium. J. Neurosci. 25, 9940–9948 (2005).
Zhao, X. F. et al. Targeting microglia using Cx3cr1-Cre lines: revisiting the specificity. eNeuro 6, ENEURO.0114-19.2019 (2019).
Sanmarco, L. M. et al. Gut-licensed IFNλ+ NK cells drive LAMP1+TRAIL+ anti-inflammatory astrocytes. Nature 590, 473–479 (2021).
Hao, Y. et al. Integrated analysis of multimodal single-cell data. Cell 184, 3573–3587 (2021).
Ianevski, A., Giri, A. K. & Aittokallio, T. Fully-automated and ultra-fast cell-type identification using specific marker combinations from single-cell transcriptomic data. Nat. Commun. 13, 1246 (2022).
Picelli, S. et al. Smart-seq2 for sensitive full-length transcriptome profiling in single cells. Nat. Methods 10, 1096–1098 (2013).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550 (2014).
Mootha, V. K. et al. PGC-1α-responsive genes involved in oxidative phosphorylation are coordinately downregulated in human diabetes. Nat. Genet. 34, 267–273 (2003).
Subramanian, A. et al. Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc. Natl Acad. Sci. USA 102, 15545–15550 (2005).
Browaeys, R., Saelens, W. & Saeys, Y. NicheNet: modeling intercellular communication by linking ligands to target genes. Nat. Methods 17, 159–162 (2020).
Gu, Z., Gu, L., Eils, R., Schlesner, M. & Brors, B. circlize implements and enhances circular visualization in R. Bioinformatics 30, 2811–2812 (2014).
Langmead, B. & Salzberg, S. L. Fast gapped-read alignment with Bowtie 2. Nat. Methods 9, 357–359 (2012).
Tarasov, A., Vilella, A. J., Cuppen, E., Nijman, I. J. & Prins, P. Sambamba: fast processing of NGS alignment formats. Bioinformatics 31, 2032–2034 (2015).
Li, H. et al. The sequence alignment/map format and SAMtools. Bioinformatics 25, 2078–2079 (2009).
Ramirez, F., Dundar, F., Diehl, S., Gruning, B. A. & Manke, T. deepTools: a flexible platform for exploring deep-sequencing data. Nucleic Acids Res. 42, W187–W191 (2014).
Zhang, Y. et al. Model-based analysis of ChIP–seq (MACS). Genome Biol. 9, R137 (2008).
Bero, A. W. et al. Bidirectional relationship between functional connectivity and amyloid-β deposition in mouse brain. J. Neurosci. 32, 4334–4340 (2012).
Acknowledgements
We thank the NeuroTechnology Studio at Brigham and Women’s Hospital for providing access to the Zeiss LSM710 confocal microscope and Leica DMi8 microscope and L. Ding for consultation on data acquisition and data analysis. We also thank the Broad Institute of Massachusetts Institute of Technology and Harvard for Smart-seq2 RNA-seq and scRNA-seq, M. Jucker for providing the APP/PS1 mice, D. Tenen and J. Zhang for providing Spi1fl/fl mice and D. Sheppard for providing the antibody to ITGB8. This study was supported by the Cure Alzheimer’s Fund (to O.B. and D.M.H.); BrightFocus Foundation 2020A016806 (O.B.); NIH-NIA R01AG051812 (O.B.), R01AG054672 (O.B.), R01AG075509 (O.B.), R21AG076982 (O.B.) and R01AG080992 (O.B.); NIH-NEI R01EY027921 (O.B.), K12EY016335 (M.A.M.) and K08 EY030160 (M.A.M.); NIH-NINDS R01NS088137 (O.B.), R21NS104609 (O.B.) and R21NS101673 (O.B.); NIH-NIGMS R01GM132668 (O.B.); Nancy Davis Foundation Innovative Award (O.B.) and RF1NS090934 (D.M.H.); Alzheimer’s Association Research Fellowship AARF-21-846786 (N.R.); National Multiple Sclerosis Society FG-2108-38372 (W.B.); Department of Defense MS210257 (W.B.); Research to Prevent Blindness Career Development Award (M.A.M.); Glaucoma Research Foundation Catalyst for a Cure Initiative to Prevent and Cure Neurodegeneration (M.A.M.) and Alcon Research Institute Young Investigator Award (M.A.M.).
Author information
Authors and Affiliations
Contributions
O.B. supervised the study, data interpretation and manuscript editing. Z.Y. and N.R. designed the study under the supervision of O.B. Z.Y. and C.M. performed stereotactic injections. Z.Y., N.R. and A.Durao performed immunohistochemistry experiments and imaging. X.Z. performed ChIP–seq and ITGB8 blockade. W.B. and Z.Y. performed RNA isolation. Z.Y. performed image analysis with the assistance of J.-L.B. and J.R. Z.Y. and C.M. performed microglia isolation with the assistance of C.S., S.S., A.Durao, H.S., Y.C., J.-L.B., S.W. and M.A. R.K.K. performed FACS. K.L.K, D.M. and C.G. performed data analysis of bulk RNA-seq and scRNA-seq data. K.L.K. performed ChIP–seq data analysis. S.H., Z.Y. and K.N.S. performed RNAscope under the supervision of T.I. and R.N. A.Deik performed the lipidomic analysis under the supervision of C.C. D.M.H., C.W., N.J. and X.B. assisted with experiments related to tau mice. Z.Y., C.M., C.B., M.A.M., K.M.P., A.Y., J.-L.B. and S.K. developed and procured mouse models. E.G., N.B. and B.J.L.E. provided human brain samples. N.G.S., G.L.M. and T.D.A. provided and performed experiments related to Itgb8-KO mice, Itgb8-tdT mice and Smad2/Smad3-cKO mice. D.G.T. provided Spi1fl/fl mice. T.D.A. and D.S. provided neutralizing antibody to ITGB8. R.E.T. and D.M.H. provided APOE3fl/fl and APOE4fl/fl mice and assisted with data interpretation. The manuscript was written by O.B., N.R., Z.Y. and K.L.K.
Corresponding author
Ethics declarations
Competing interests
O.B., Z.Y. and N.R. are inventors of a patent licensed by Brigham and Women’s Hospital for use of LGALS3 for treatment of neurodegenerative diseases. O.B., Z.Y., N.R., X.Z., T.D.A. and D.S. are inventors for a patent licensed by Brigham and Women’s Hospital and University of California, San Francisco, for use of an ITGB8 blocker for treatment of neurodegenerative diseases. O.B. has a collaboration with GSK and Regulus Therapeutics, receives research funding from Sanofi and GSK and receives honoraria for lectures and consultancy from UCB, Camp4, Ono Pharma USA and General Biophysics. D.M.H. is an inventor on a patent licensed by Washington University to C2N Diagnostics on the therapeutic use of antibodies to tau and on a patent licensed to NextCure on antibodies to APOE. D.M.H. cofounded and is on the scientific advisory board of C2N Diagnostics. D.M.H. is on the scientific advisory board of Denali, Genentech and Cajal Neuroscience and consults for Alector. All other authors have no competing interests.
Peer review
Peer review information
Nature Immunology thanks the anonymous reviewers for their contribution to the peer review of this work. Primary Handling Editor: S. Houston, in collaboration with the Nature Immunology team.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 APOE4 impairs microglial response to acute neurodegeneration.
a, H3K9ac peak-plot heat map of microglia from 4-month-old APOE3-KI or APOE4-KI mice (n = 3 mice/group). b, H3K9ac peaks of key genes. c, Volcano plot of DEGs of phagocytic microglia isolated from APOE4-KI vs. APOE3-KI mice. DEGs were identified using DESeq2 analysis with an LRT (n = 4-6 mice/group, P < 0.05). d, Fold change induction of key MGnD genes expressed in phagocytic microglia relative to non-phagocytic microglia, isolated from APOE3-KI and APOE4-KI mice. DEGs were identified using DESeq2 analysis with an LRT and key MGnD genes were selected from Krasemann et al.4. e, Violin plot of top-300 induced MGnD genes in phagocytic microglia isolated from APOE3-KI and APOE4-KI mice (Fig. 1g). f, Top-affected canonical pathways in phagocytic microglia compared to non-phagocytic microglia from APOE3-KI and APOE4-KI mice (P < 0.05). g, Quantification of phagocytic Iba1+ cell numbers from AN-injected APOE3-KI and APOE4-KI mice (n = 6 APOE3-KI mice, n = 7 APOE4-KI mice). h, Quantification of Iba1+ cells from AN-injected APOE3-KI and APOE4-KI mice (n = 6 APOE3-KI mice, n = 7 APOE4-KI mice). i, Circos plot illustrating ligand-receptor interactions in APOE4-cKO microglia compared to APOE4-KI microglia. Two-tailed Student’s t-test. Data were presented as mean ± s.e.m.
Extended Data Fig. 2 No differences of APOE expression between P301S:APOE3-KI and P301S:APOE4-KI mice.
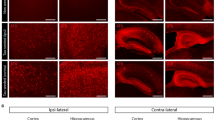
a, Confocal images of GFAP, APOE and Iba1 in P301S:APOE3-KI and P301S:APOE4-KI mice. Scale bar: 50 μm. b, Quantification of APOE in the cortex (n = 5 mice/group). Two-tailed Student’s t-test. Data presented as mean ± s.e.m.
Extended Data Fig. 3 Targeting microglial APOE4 restores neurodegenerative microglia in APP/PS1 mice.
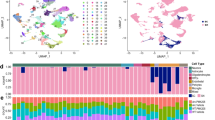
a, UMAP plot of unsupervised seurat clusters from scRNAseq analysis of brain cells isolated from APP/PS1:APOE4-KI and APP/PS1:APOE4-cKO mice (n = 2). b, UMAP plot of labelled cell types. c, Cell-specific genes used to identify cell types from seurat clusters. d, UMAP plot of microglia/myeloid cells re-clustered. e, Feature plot showing expression of border-associated macrophages genes. f, Heat map of DEGs representing the microglia/myeloid cluster. DEGs were identified using FindAllMarker Seurat function (P < 0.05). g, Feature plot showing co-expression of Lgals3 and Clec7a microglia. h, Ridgeplot of Clec7a and Lgals3 in APOE4-cKO and APOE4-KI in MGnD microglia showing gene expression cut-off for Clec7aHi (2.65) and Lgals3Hi (1.68) microglia.
Extended Data Fig. 4 Deletion of microglial APOE4 alleviates lipid dysregulation in APP/PS1 mice.
a, Heat map of lipids significantly altered by genotype in microglia isolated from APOE3-KI, APOE4-KI, APP/PS1:APOE3-KI, and APP/PS1:APOE4-KI. DEGs were identified using DESeq2 analysis with an LRT (n = 2 mice, P < 0.05). b, Confocal images of Plin2, Iba1 and HJ3.4B in APP/PS1:APOE3-KI, APP/PS1:APOE3-cKO, APP/PS1:APOE4-KI, and APP/PS1:APOE4-cKO mice. Scale bar: 50 μm. c, Quantification of Plin2+ area in Iba1+ cells (n = 15 cells from APP/PS1:APOE3-KI mice, n = 8 cells from APP/PS1:APOE3-cKO mice, n = 24 cells from APP/PS1:APOE4-KI mice, n = 16 cells from APP/PS1:APOE4-cKO mice). One-way ANOVA. Data presented as mean ± s.e.m.
Extended Data Fig. 5 Deletion of microglial APOE4 promotes astrocyte activation in APP/PS1 mice.
a, Heat map showing DEGs of clusters 3 and 5 (astrocyte clusters). DEGs were identified using FindMarkers Seurat function (P < 0.05, n = 2). b, UMAP projection of disease-associated astrocyte genes29 in APP/PS1:APOE4-KI and APP/PS1:APOE4-cKO mice. c, Donut charts showing percentage of GfapHi:Serpina3nLo, GfapHi:Serpina3nHi, GfapHi:VimLo, GfapHi:VimHi, GfapHi:Cd9Lo, and GfapHi:Cd9Hi microglia between APP/PS1:APOE4-KI and APP/PS1:APOE4-cKO mice.
Extended Data Fig. 6 Impaired induction of MGnD signature and astrocyte activation in APOE ε4 AD brains.
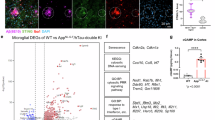
a, DEGs of female APOE ε3/3 carriers (n = 4) vs. APOE ε3/4 (n = 3) carriers analyzed from Olah et al.34. DEGs were identified using FindMarkers Seurat function (P < 0.01). b, UMAP plot of brain scRNAseq showing Seurat clusters of AD:APOE ε3/3 carriers and AD:APOE ε3/4 carriers analyzed from dataset by Zhou et al.35. c, UMAP plot indicating cell type assignment to clusters analyzed from dataset by Zhou et al.35. d, Confocal images of IBA1 and HJ3.4B immunoreactivity and detection of INPP5D gene expression using RNAscope in AD:APOE ε3/4 carriers compared to AD:APOE ε3/3 carriers. e, Quantification of INPP5D fluorescence in IBA1+ microglia in AD:APOE ε3/4 carriers compared to AD:APOE ε3/3 carriers (n = 14 AD:APOE ε3/3 carriers, n = 16 AD:APOE ε3/4 carriers). f, Ridgeplot of GFAPHi vs. GFAPLo astrocytes analyzed from dataset by Zhou et al.35. g, Confocal images of GFAP and HJ3.4B immunoreactivity and detection of ITGB8 gene expression using RNAscope in AD:APOE ε3/4 males compared to AD:APOE ε3/3 males. h,i, Quantification of ITGB8 fluorescence in ROI (h) and in GFAP+ astrocytes (i) in AD:APOE ε3/4 males compared to AD:APOE ε3/3 males (n = 7 AD:APOE ε3/3 carriers, n = 8 AD:APOE ε3/4 carriers). j, Confocal images of Gfap and HJ3.4B immunoreactivity and detection of Itgb8 mRNA expression using RNAscope in APP/PS1:APOE4-cKO mice compared to APP/PS1:APOE4-KI mice. k, Quantification of Itgb8 fluorescence in Gfap+ astrocytes (n = 10 ROIs from 3 APP/PS1:APOE4-KI mice, 18 ROIs from 5 APP/PS1:APOE4-cKO mice). Two-tailed unpaired Student’s t-test. Scale bar: 50 μm. Data were shown as mean ± s.e.m.
Extended Data Fig. 7 Expression of Itgb8 in the adult mouse brain.
Dot plot showing expression level of Itgb8 in multiple CNS cell types (adopted from Zhang et al.38). b, Representative in situ image of Itgb8 in the cortex of adult mouse brain (Allen Brain Atlas: https://mouse.brain-map.org). n = 2 (the Allen Brain Atlas tested the Itgb8 probe on two brains; one sectioned sagitally and another sectioned coronally). Scale bar: 1 mm. c, Schematics of Itgb8−tdT transgenic mouse strain37. d, Representative images of Itgb8-TdT, Sox9, Gfap (top); Itgb8-TdT, Pdgfra, Olig2 (middle); and Itgb8-TdT, NeuN, Olig2 (bottom). Arrowheads indicate magnified ROIs (n = 6). Data are representative of 2 independent experiments. Scale bar: 50 μm.
Extended Data Fig. 8 Blocking Itgb8 signaling enhances MGnD response and reduces AD pathology in APP/PS1 mice.
a–c, Confocal images of Tmem119, Apoe, and Clec7a staining (a); P2ry12, Apoe, and Clec7a staining (b); Lgals3, Iba1, and Clec7a staining (c) in the cortex of Itgb8-cKO mice and littermate controls. Arrows indicate microglia. Data are representative of 2 independent experiments. Scale bar: 50 μm. d, Gene ontology network of increased DEGs in microglia from Itgb8-cKO mice compared to WT littermates. DEGs were identified using DESeq2 analysis with an LRT and gene ontology pathways selected with P < 0.05. e, PCA of each genotype. f, Heat map of DEGs of phagocytic- and non-phagocytic microglia isolated from WT and Itgb8-cKO mice. DEGs were identified using DESeq2 analysis with an LRT (n = 5-6 mice/group, P < 0.05). g, Gene ontology network of top pathways in phagocytic- and non-phagocytic microglia from Itgb8-cKO mice compared with WT mice. DEGs were identified using DESeq2 analysis with an LRT and gene ontology pathways selected with P < 0.05. h, FACS plot of Aβ phagocytosis in WT and Itgb8-cKO mice. Percentage calculated as Aβ42-Alexa Fluor 555+ out of Fcrls+CD11b+Ly6C– microglia. i, Quantification of percentage of Aβ-42 phagocytic microglia in WT and Itgb8-cKO mice (n = 5 WT mice, n = 10 Itgb8-cKO mice). j, Confocal images of MHC II, Iba1, and HJ3.4B in APP/PS1 mice injected with anti-ITGB8 neutralizing antibody and IgG isotype control. Scale bar: 100 μm. k, Quantification of MHC II+ immunoreactivity at the injection site (n = 4 mice/group). Two-tailed Student’s t-test. Data were presented as mean ± s.e.m.
Extended Data Fig. 9 Microglial deletion of Smad2/3 induces MGnD phenotype.
a, Confocal images showing Iba1, Tmem119 and Cd68 staining in control and Smad2/3-cKO mice. Scale bar: 50 μm. b, Quantification of Cd68+ immunoreactivity (n = 5 WT mice, n = 4 Smad2/3-cKO mice). c, Confocal images of Iba1, Apoe and Gfap in control and Smad2/3-cKO mice. Scale bar: 50 μm. d, Quantification of Apoe+ and Gfap+ immunoreactivity (n = 4 mice/group). e–h, Expression of key homeostatic and MGnD genes in microglia from Itgb8-cKO (n = 4-5 mice/group) (e), Smad2/3-cKO (n = 4 mice/group) (f), Tgfbr2-cKO (n = 3 mice/group) (g) and Nrros-KO (n = 5 mice/group) (h) relative to non-transgenic control mice. i–l, Expression levels of key AD-risk factor genes (Inpp5d, Havcr2 and Bin1) in microglia from Itgb8-cKO in microglia from Itgb8-cKO (n = 4-5 mice/group) (i), Smad2/3-cKO (n = 4 mice/group) (j), Tgfbr2-cKO (n = 3 mice/group) (k) and Nrros-KO (n = 5 mice/group) (l), relative to non-transgenic control mice. Two-tailed Student’s t-test. Data were shown as mean ± s.e.m.
Extended Data Fig. 10 Microglial deletion of the homeostatic checkpoint Inpp5d facilitates plaque clearance via the induction of MGnD response.
a, Heat map of top-500 DEGs in WT:control vs. WT:Inpp5d-cKO vs. APP/PS1:control vs. APP/PS1:Inpp5d-cKO microglia. DEGs were identified using DESeq2 analysis with an LRT (n = 2-5 mice/group, P < 0.05). b, Gene ontology analysis of DEGs for IFNγ response, antigen processing and presentation in microglia. DEGs were identified using DESeq2 analysis with an LRT and gene ontology pathways selected with P < 0.05. c, Representative images of Thioflavin-S staining. Scale bar: 1 mm (left), 50 μm (right). d, Quantification of Thioflavin-S staining in APP/PS1 (n = 7) and APP/PS1:Inpp5d-cKO (n = 7) mice. e, Confocal images of Iba1, Clec7a and HJ3.4B staining. Scale bar: 50 μm. f,g, Quantification of Clec7a (f) and Iba1 (g) immunoreactivity in association with Aβ plaques (n = 218 plaques from APP/PS1 mice, n = 115 plaques from APP/PS1:Inpp5d-cKO mice). h, Confocal images of Iba1, Lgals3 and HJ3.4B staining. Scale bar: 50 μm. i,j, Quantification of Lgals3 (i) and HJ3.4B (j) immunoreactivity in association with Aβ plaques (n = 141 plaques from APP/PS1 mice, n = 186 plaques from APP/PS1:Inpp5d-cKO mice for Lgals3, n = 7 mice for HJ3.4B). k, Confocal images of Lamp1 and HJ3.4B staining. Scale bar: 50 μm. l, Quantification of Lamp1 immunoreactivity in association with Aβ plaques (n = 126 plaques from APP/PS1 mice, n = 138 plaques from APP/PS1:Inpp5d-cKO mice). m, Confocal images of Iba1, Gfap and HJ3.4B staining. Scale bar: 50 μm. n, Quantification of Gfap immunoreactivity in association with Aβ plaques (n = 154 plaques from APP/PS1 mice, n = 127 plaques from APP/PS1:Inpp5d-cKO mice). Two-tailed Student’s t-test. Data were presented as mean ± s.e.m.
Supplementary information
Supplementary Table 1
RNA-seq and ChIP–seq data for Fig. 1 and related Extended Data figures.
Supplementary Table 2
RNA-seq data for Fig. 2.
Supplementary Table 3
RNA-seq data for Fig. 3.
Supplementary Table 4
RNA-seq and scRNA-seq data for Fig. 4 and related Extended Data figures.
Supplementary Table 6
RNA-seq data for Fig. 7.
Supplementary Table 7
Information for human materials for Fig. 7.
Supplementary Table 8
RNA-seq for Fig. 8 and related Extended Data figures.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Yin, Z., Rosenzweig, N., Kleemann, K.L. et al. APOE4 impairs the microglial response in Alzheimer’s disease by inducing TGFβ-mediated checkpoints. Nat Immunol 24, 1839–1853 (2023). https://doi.org/10.1038/s41590-023-01627-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41590-023-01627-6
- Springer Nature America, Inc.
This article is cited by
-
Emerging role of senescent microglia in brain aging-related neurodegenerative diseases
Translational Neurodegeneration (2024)
-
Updates in Alzheimer's disease: from basic research to diagnosis and therapies
Translational Neurodegeneration (2024)
-
Deciphering the effect of phytosterols on Alzheimer’s disease and Parkinson’s disease: the mediating role of lipid profiles
Alzheimer's Research & Therapy (2024)
-
Tracking neuroinflammatory biomarkers in Alzheimer’s disease: a strategy for individualized therapeutic approaches?
Journal of Neuroinflammation (2024)
-
Onset of Alzheimer disease in apolipoprotein ɛ4 carriers is earlier in butyrylcholinesterase K variant carriers
BMC Neurology (2024)