Abstract
The lung is inhabited by resident alveolar and interstitial macrophages as well as monocytic cells that survey lung tissues. Each cell type plays distinct functional roles under homeostatic and inflammatory conditions, but mechanisms establishing their molecular identities and functional potential remain poorly understood. In the present study, systematic evaluation of transcriptomes and open chromatin of alveolar macrophages (AMs), interstitial macrophages (IMs) and lung monocytes from two mouse strains enabled inference of common and cell-specific transcriptional regulators. We provide evidence that these factors drive selection of regulatory landscapes that specify distinct phenotypes of AMs and IMs and entrain qualitatively different responses to toll-like receptor 4 signaling in vivo. These studies reveal a striking divergence in a fundamental innate immune response pathway in AMs and establish a framework for further understanding macrophage diversity in the lung.






Similar content being viewed by others
Data availability
The sequencing data reported in this article are deposited in the Gene Expression Omnibus with the following accession number: GSE136916. Additional data supporting the presented findings are available in the manuscript and upon request from the corresponding author.
References
Pollard, J. W. Trophic macrophages in development and disease. Nat. Rev. Immunol. 9, 259–270 (2009).
Ardini-Poleske, M. E. et al. LungMAP: the molecular atlas of lung development program. Am. J. Physiol. Lung Cell. Mol. Physiol. 313, L733–L740 (2017).
Gomez Perdiguero, E. et al. Tissue-resident macrophages originate from yolk-sac-derived erythro-myeloid progenitors. Nature 518, 547–551 (2015).
Hoeffel, G. et al. C-Myb+ erythro-myeloid progenitor-derived fetal monocytes give rise to adult tissue-resident macrophages. Immunity 42, 665–678 (2015).
Guilliams, M. et al. Alveolar macrophages develop from fetal monocytes that differentiate into long-lived cells in the first week of life via GM-CSF. J. Exp. Med. 210, 1977–1992 (2013).
Beck-Schimmer, B. et al. Alveolar macrophages regulate neutrophil recruitment in endotoxin-induced lung injury. Respir. Res. 6, 61 (2005).
Hussell, T. &. Bell, T. J. Alveolar macrophages: plasticity in a tissue-specific context. Nat. Rev. Immunol. 14, 81–93 (2014).
Schneider, C. et al. Alveolar macrophages are essential for protection from respiratory failure and associated morbidity following influenza virus infection. PLoS Pathog. 10, e1004053 (2014).
Cohen, M. et al. Lung single-cell signaling interaction map reveals basophil role in macrophage imprinting. Cell 175, 1031–1044.e18 (2018).
Reyfman, P. A. et al. Single-cell transcriptomic analysis of human lung provides insights into the pathobiology of pulmonary fibrosis. Am. J. Respir. Crit. Care Med. 199, 1517–1536 (2019).
Tan, S. Y. & Krasnow, M. A. Developmental origin of lung macrophage diversity. Development 143, 1318–1327 (2016).
Chakarov, S. et al. Two distinct interstitial macrophage populations coexist across tissues in specific subtissular niches. Science 363, eaau0964 (2019).
Gibbings, S. L. et al. Three unique interstitial macrophages in the murine lung at steady state. Am. J. Respir. Cell Mol. Biol. 57, 66–76 (2017).
Tatham, K. C. et al. Intravascular donor monocytes play a central role in lung transplant ischaemia–reperfusion injury. Thorax 73, 350–360 (2018).
Maus, U. A. et al. Monocytes are potent facilitators of alveolar neutrophil emigration during lung inflammation: role of the CCL2–CCR2 axis. J. Immunol. 170, 3273–3278 (2003).
Misharin, A. V. et al. Monocyte-derived alveolar macrophages drive lung fibrosis and persist in the lung over the life span. J. Exp. Med. 214, 2387–2404 (2017).
Jardine, L. et al. Lipopolysaccharide inhalation recruits monocytes and dendritic cell subsets to the alveolar airspace. Nat. Commun. 10, 1999 (2019).
T’Jonck, W., Guilliams, M. & Bonnardel, J. Niche signals and transcription factors involved in tissue-resident macrophage development. Cell Immunol. 330, 43–53 (2018).
Mass, E. et al. Specification of tissue-resident macrophages during organogenesis. Science 353, aaf4238 (2016).
Lavin, Y. et al. Tissue-resident macrophage enhancer landscapes are shaped by the local microenvironment. Cell 159, 1312–1326 (2014).
Schneider, C. et al. Induction of the nuclear receptor PPAR-γ by the cytokine GM-CSF is critical for the differentiation of fetal monocytes into alveolar macrophages. Nat. Immunol. 15, 1026–1037 (2014).
Baker, A. D. et al. PPARγ regulates the expression of cholesterol metabolism genes in alveolar macrophages. Biochem. Biophys. Res. Commun. 393, 682–687 (2010).
Gosselin, D. et al. Environment drives selection and function of enhancers controlling tissue-specific macrophage identities. Cell 159, 1327–1340 (2014).
Gosselin, D. et al. An environment-dependent transcriptional network specifies human microglia identity. Science 356, eaal3222 (2017).
Zhang, D. X. & Glass, C. K. Towards an understanding of cell-specific functions of signal-dependent transcription factors. J. Mol. Endocrinol. 51, T37–T50 (2013).
Geissmann, F. et al. Development of monocytes, macrophages, and dendritic cells. Science 327, 656–661 (2010).
Mildner, A. et al. Genomic characterization of murine monocytes reveals C/EBPβ transcription factor dependence of Ly6C– cells. Immunity 46, 849–862.e7 (2017).
La Manno, G. et al. RNA velocity of single cells. Nature 560, 494–498 (2018).
Misharin, A. V., Morales-Nebreda, L., Mutlu, G. M., Budinger, G. R. & Perlman, H. Flow cytometric analysis of macrophages and dendritic cell subsets in the mouse lung. Am. J. Respir. Cell Mol. Biol. 49, 503–510 (2013).
Buenrostro, J. D., Wu, B., Chang, H. Y. & Greenleaf, W. J. ATAC-seq: a method for assaying chromatin accessibility genome-wide. Curr. Protoc. Mol. Biol. 109, 21–29 (2015).
Keane, T. M. et al. Mouse genomic variation and its effect on phenotypes and gene regulation. Nature 477, 289–294 (2011).
Whitehead, G. S., Burch, L. H., Berman, K. G., Piantadosi, C. A. & Schwartz, D. A. Genetic basis of murine responses to hyperoxia-induced lung injury. Immunogenetics 58, 793–804 (2006).
Bartalesi, B. et al. Different lung responses to cigarette smoke in two strains of mice sensitive to oxidants. Eur. Respir. J. 25, 15–22 (2005).
Rittling, S. R. Osteopontin in macrophage function. Expert Rev. Mol. Med. 13, e15 (2011).
Shan, M. et al. Cigarette smoke induction of osteopontin (SPP1) mediates TH17 inflammation in human and experimental emphysema. Sci. Transl. Med. 4, 117ra9 (2012).
Russell, C. D. & Schwarze, J. The role of pro-resolution lipid mediators in infectious disease. Immunology 141, 166–173 (2014).
Zhen, A. et al. CD4 ligation on human blood monocytes triggers macrophage differentiation and enhances HIV infection. J. Virol. 88, 9934–9946 (2014).
Adachi, H. & Tsujimoto, M. FEEL-1, a novel scavenger receptor with in vitro bacteria-binding and angiogenesis-modulating activities. J. Biol. Chem. 277, 34264–34270 (2002).
Heinz, S. et al. Simple combinations of lineage-determining transcription factors prime cis-regulatory elements required for macrophage and B cell identities. Mol. Cell 38, 576–589 (2010).
Thomas, G. D. et al. Deleting an Nr4a1 super-enhancer subdomain ablates Ly6Clow monocytes while preserving macrophage gene function. Immunity 45, 975–987 (2016).
Kannan, M. B., Solovieva, V. & Blank, V. The small MAF transcription factors MAFF, MAFG and MAFK: current knowledge and perspectives. Biochim. Biophys. Acta 1823, 1841–1846 (2012).
You, F. et al. ELF4 is critical for induction of type I interferon and the host antiviral response. Nat. Immunol. 14, 1237–1246 (2013).
Link, V. M. et al. Analysis of genetically diverse macrophages reveals local and domain-wide mechanisms that control transcription factor binding and function. Cell 173, 1796–1809.e17 (2018).
Heinz, S. et al. Effect of natural genetic variation on enhancer selection and function. Nature 503, 487–492 (2013).
Link, V. M., Romanoski, C. E., Metzler, D. & Glass, C. K. MMARGE: motif mutation analysis for regulatory genomic elements. Nucleic Acids Res. 46, 7006–7021 (2018).
Matute-Bello, G. et al. An official American Thoracic Society workshop report: features and measurements of experimental acute lung injury in animals. Am. J. Respir. Cell Mol. Biol. 44, 725–738 (2011).
Oishi, Y. et al. SREBP1 contributes to resolution of pro-inflammatory TLR4 signaling by reprogramming fatty acid metabolism. Cell Metab. 25, 412–427 (2017).
Picelli, S. et al. Smart-seq2 for sensitive full-length transcriptome profiling in single cells. Nat. Methods 10, 1096–1098 (2013).
Butler, A., Hoffman, P., Smibert, P., Papalexi, E. & Satija, R. Integrating single-cell transcriptomic data across different conditions, technologies, and species. Nat. Biotechnol. 36, 411–420 (2018).
Dobin, A. et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics 29, 15–21 (2013).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550 (2014).
Langmead, B. & Salzberg, S. L. Fast gapped-read alignment with Bowtie 2. Nat. Methods 9, 357–359 (2012).
Acknowledgements
These studies were supported by National Institutes of Health (NIH) grants (nos. DK091183 and DK063491). E.S. was supported by a NIH grant (no. K08HL140198). C.E.R. was supported by NIH grants (nos. R00HL123485 and R01HL147187). L.S.P. was supported by NIH grants (nos. HL086324, HL126703 and HL143256) and the Gerber Foundation (grant no. 1823-3830). We thank T. Rombaldo for assistance with FACS, J. Collier for technical assistance and L. van Ael for help with preparation of the manuscript. Sequencing of RNA-seq and ATAC-seq libraries was conducted at the IGM Genomics Center, University of California San Diego, La Jolla, CA.
Author information
Authors and Affiliations
Contributions
E.S., N.J.S. and C.K.G. designed the project. E.S., N.J.S., E.W. and G.J.F. performed the experiments. V.M.L, Z.O., C.E.R., E.S. and C.K.G. analyzed the data. E.S., L.S.P. and C.K.G. secured the funding. E.S. and C.K.G. wrote the manuscript. L.S.P, V.M.L. and N.J.S. critically revised the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information L. A. Dempsey was the primary editor on this article and managed its editorial process and peer review in collaboration with the rest of the editorial team.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Single-cell RNA-seq clusters, FACS sorting strategy and quality control of FACS and RNA-seq results.
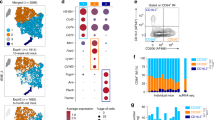
a. Single-cell RNA-seq gene expression in clusters used to determine cluster identities. Heat map is representing expression values for the most significant genes in each cluster. Cells from the lungs of 3 male and 1 female DBA/2J mice were pooled. b. Zoomed-in view of velocity analysis for single-cell RNA-seq from Fig. 1a. c. Flow cytometry analysis and sorting strategy to obtain subsets of lung mononuclear phagocytes (MPs). d. Validation of sorting strategy with gene expression in sorted lung MPs in C57BL/6J (B6) mice (top panel) and DBA/2J (DBA) mice (bottom panel). Bars represent transcripts per million (TPM) for one mouse. Two replicates are shown. e. Spearman correlation heat map of all RNA-seq replicates, N=2 for each cell type in both mouse strains. f. Comparison of gene expression for AM vs. IM and iMo vs. pMo in B6 mice. Scatter plots are showing genes with TPM > 16. Blue dots for AM, orange dots for IM and bordeaux dots for iMo show genes with fold change (FC) 2 or higher. g. Comparison of gene expression in pMo isolated from lung vs pMo isolated from circulating blood. Scatter plots show genes with TPM > 16. Pale blue dots show genes with FC 2 or higher, dark blue dots show genes with FC 4 or higher. h. Ingenuity pathway analysis (IPA) functional pathways for genes differentially regulated in pMo isolated from lung vs pMo isolated from circulating blood.
Extended Data Fig. 2 ATAC-seq quality control and HOMER de novo motif analysis.
a. Spearman correlation heat map of all ATAC-seq replicates, N=2 for each cell type in both mouse strains. b. Comparison of open chromatin regions for iMo vs pMo in B6 mice. Scatter plot shows log2 (tag counts+1) of ATAC-seq peaks, colored dots show ATAC-seq peaks with fold change (FC) 4 or higher, bordeaux for iMo and purple for pMo. c. HOMER de novo motif enrichment analysis for transcription factor (TF) binding sites in distal regions of open chromatin (> 3 kb from a TSS) likely representing enhancers using GC matched background for B6 mice. Boxes display –log10 p-values for enrichment of the motif, rank order in parenthesis and percentage of motif occurrence in peaks vs background, N=2 for each cell type in both mouse strains. d. HOMER de novo motif enrichment analysis for TF binding sites in distal regions of open chromatin (> 3 kb from a TSS) likely representing enhancers using GC matched background for DBA mice. Boxes display –log10 p-values for enrichment of the motif, rank order in parenthesis and percentage of motif occurrence in peaks vs background, N=2 for each cell type in both mouse strains.
Extended Data Fig. 3 Flow cytometry and ingenuity pathway analysis (IPA) of lung mononuclear phagocytes (MPs) after LPS administration.
a. Flow cytometry analysis of lung MP subsets and neutrophils after i.p. LPS administration. Bars represent the average percentage of given cell subset out of CD45+ leukocytes ± s.d. Three independent experiments were pooled, 0h N=11, for all other time points N=8. Non-parametric Wilcoxon signed-rank test with Bonferroni correction was used, * p < 0.05, ** p < 0.01, *** p < 0.001. b. Venn diagram of 4,499 differentially expressed genes in AM, IM and iMo after i.p. LPS administration. c. Flow cytometry analysis of lung MP subsets and neutrophils after i.n. LPS administration. Bars represent the average percentage of given cell subset out of CD45+ leukocytes ± s.d. Two independent experiments were pooled, 0h N= 11, for all other time points N=4. Non-parametric Wilcoxon signed-rank test with Bonferroni correction was used, * p < 0.05, ** p < 0.01, *** p < 0.001. d. Venn diagram of 806 differentially expressed genes in AM, IM and iMo after i.n. LPS administration.
Supplementary information
Supplementary Information
Supplementary Tables 1 and 2.
Rights and permissions
About this article
Cite this article
Sajti, E., Link, V.M., Ouyang, Z. et al. Transcriptomic and epigenetic mechanisms underlying myeloid diversity in the lung. Nat Immunol 21, 221–231 (2020). https://doi.org/10.1038/s41590-019-0582-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41590-019-0582-z
- Springer Nature America, Inc.
This article is cited by
-
Macrophages in tissue repair and regeneration: insights from zebrafish
Cell Regeneration (2024)
-
Single-cell sequencing reveals that endothelial cells, EndMT cells and mural cells contribute to the pathogenesis of cavernous malformations
Experimental & Molecular Medicine (2023)
-
Kupffer cells prevent pancreatic ductal adenocarcinoma metastasis to the liver in mice
Nature Communications (2023)
-
Novel insights into molecular signatures and pathogenic cell populations shared by systemic lupus erythematosus and vascular dementia
Functional & Integrative Genomics (2023)
-
SALL1 enforces microglia-specific DNA binding and function of SMADs to establish microglia identity
Nature Immunology (2023)