Abstract
Cyclic diadenylate monophosphate (c-di-AMP) is secreted by bacteria as a secondary messenger. How immune cells detect c-di-AMP and initiate anti-bacterial immunity remains unknown. We found that the endoplasmic reticulum (ER) membrane adaptor ERAdP acts as a direct sensor for c-di-AMP. ERAdP-deficient mice were highly susceptible to Listeria monocytogenes infection and exhibited reduced pro-inflammatory cytokines. Mechanistically, c-di-AMP bound to the C-terminal domain of ERAdP, which in turn led to dimerization of ERAdP, resulting in association with and activation of the kinase TAK1. TAK1 activation consequently initiated activation of the transcription factor NF-κB to induce the production of pro-inflammatory cytokines in innate immune cells. Moreover, double-knockout of ERAdP and TAK1 resulted in heightened susceptibility to L. monocytogenes infection. Thus, ERAdP-mediated production of pro-inflammatory cytokines is critical for controlling bacterial infection.






Similar content being viewed by others
References
Orzalli, M. H. & Knipe, D. M. Cellular sensing of viral DNA and viral evasion mechanisms. Annu. Rev. Microbiol. 68, 477–492 (2014).
Cao, X. Self-regulation and cross-regulation of pattern-recognition receptor signaling in health and disease. Nat. Rev. Immunol. 16, 35–50 (2016).
Keestra-Gounder, A. M. et al. NOD1 and NOD2 signaling links ER stress with inflammation. Nature 532, 394–397 (2016).
Liu, B. et al. Toll-receptor-mediated Hippo signaling controls innate immunity in Drosophila. Cell 164, 406–419 (2016).
Xia, P., Wang, S., Gao, P., Gao, G. & Fan, Z. DNA sensor cGAS-mediated immune recognition. Protein Cell 7, 777–791 (2016).
Gao, P. et al. Cyclic [G(2′,5′)pA(3′,5′)p] is the metazoan second messenger produced by DNA-activated cyclic GMP-AMP synthase. Cell 153, 1094–1107 (2013).
Sun, L., Wu, J., Du, F., Chen, X. & Chen, Z. J. Cyclic GMP-AMP synthase is a cytosolic DNA sensor that activates the type I interferon pathway. Science 339, 786–791 (2013).
Watson, R. O. et al. The cytosolic sensor cGAS detects Mycobacterium tuberculosis DNA to induce type I interferons and activate autophagy. Cell Host Microbe 17, 811–819 (2015).
Ishikawa, H. & Barber, G. N. STING is an endoplasmic reticulum adaptor that facilitates innate immune signaling. Nature 455, 674–678 (2008).
Wu, J. et al. Cyclic GMP-AMP is an endogenous second messenger in innate immune signaling by cytosolic DNA. Science 339, 826–830 (2013).
Zhang, X. et al. Cyclic GMP-AMP containing mixed phosphodiester linkages is an endogenous high-affinity ligand for STING. Mol. Cell 51, 226–235 (2013).
Zhong, B. et al. The adaptor protein MITA links virus-sensing receptors to IRF3 transcription factor activation. Immunity 29, 538–550 (2008).
Jin, L. et al. MPYS is required for IFN response factor 3 activation and type I IFN production in the response of cultured phagocytes to bacterial second messengers cyclic-di-AMP and cyclic-di-GMP. J. Immunol. 187, 2595–2601 (2011).
Surpris, G. et al. Cutting edge: novel Tmem173 allele reveals importance of STING N terminus in trafficking and type I IFN production. J. Immunol. 196, 547–552 (2016).
Fitzgerald, K. A. et al. IKKε and TBK1 are essential components of the IRF3 signaling pathway. Nat. Immunol. 4, 491–496 (2003).
Ishikawa, H., Ma, Z. & Barber, G. N. STING regulates intracellular DNA-mediated, type I interferon-dependent innate immunity. Nature 461, 788–792 (2009).
Tanaka, Y. & Chen, Z. J. STING specifies IRF3 phosphorylation by TBK1 in the cytosolic DNA signaling pathway. Sci. Signal. 5, ra20 (2012).
Tamayo, R., Pratt, J. T. & Camilli, A. Roles of cyclic diguanylate in the regulation of bacterial pathogenesis. Annu. Rev. Microbiol. 61, 131–148 (2007).
Woodward, J. J., Iavarone, A. T. & Portnoy, D. A. c-di-AMP secreted by intracellular Listeria monocytogenes activates a host type I interferon response. Science 328, 1703–1705 (2010).
Corrigan, R. M., Abbott, J. C., Burhenne, H., Kaever, V. & Gründling, A. c-di-AMP is a new second messenger in Staphylococcus aureus with a role in controlling cell size and envelope stress. PLoS Pathog. 7, e1002217 (2011).
Oppenheimer-Shaanan, Y., Wexselblatt, E., Katzhendler, J., Yavin, E. & Ben-Yehuda, S. c-di-AMP reports DNA integrity during sporulation in Bacillus subtilis. EMBO Rep. 12, 594–601 (2011).
Burdette, D. L. et al. STING is a direct innate immune sensor of cyclic di-GMP. Nature 478, 515–518 (2011).
Chen, J. et al. The endoplasmic reticulum adaptor protein ERAdP initiates NK cell activation via the Ubc13-mediated NF-κB pathway. J. Immunol. 194, 1292–1303 (2015).
Han, S. et al. Nuclear envelope phosphatase 1-regulatory subunit 1 (formerly TMEM188) is the metazoan Spo7p ortholog and functions in the lipin activation pathway. J. Biol. Chem. 287, 3123–3137 (2012).
Xia, P. et al. Glutamylation of the DNA sensor cGAS regulates its binding and synthase activity in antiviral immunity. Nat. Immunol. 17, 369–378 (2016).
Wing, E. J. & Gregory, S. H. Listeria monocytogenes: clinical and experimental update. J. Infect. Dis. 185, S18–S24 (2002).
McFarland, A. P. et al. Sensing of bacterial cyclic dinucleotides by the oxidoreductase RECON promotes NF-κB activation and shapes a proinflammatory antibacterial state. Immunity 46, 433–445 (2017).
Midde, K. K. et al. Multimodular biosensors reveal a novel platform for activation of G proteins by growth factor receptors. Proc. Natl. Acad. Sci. USA 112, E937–E946 (2015).
Hennessy, E. J., Parker, A. E. & O’Neill, L. A. Targeting Toll-like receptors: emerging therapeutics? Nat. Rev. Drug Discov. 9, 293–307 (2010).
Hengge, R. Principles of c-di-GMP signalling in bacteria. Nat. Rev. Microbiol. 7, 263–273 (2009).
Whiteley, A. T., Pollock, A. J. & Portnoy, D. A. The PAMP c-di-AMP is essential for Listeria monocytogenes growth in rich but not minimal media due to a toxic increase in (p)ppGpp. [corrected]. Cell Host Microbe 17, 788–798 (2015).
Parvatiyar, K. et al. The helicase DDX41 recognizes the bacterial secondary messengers cyclic di-GMP and cyclic di-AMP to activate a type I interferon immune response. Nat. Immunol. 13, 1155–1161 (2012).
Chen, Q., Sun, L. & Chen, Z. J. Regulation and function of the cGAS-STING pathway of cytosolic DNA sensing. Nat. Immunol. 17, 1142–1149 (2016).
Gao, P. et al. Structure-function analysis of STING activation by c[G(2′,5′)pA(3′,5′)p] and targeting by antiviral DMXAA. Cell 154, 748–762 (2013).
Ouyang, S. et al. Structural analysis of the STING adaptor protein reveals a hydrophobic dimer interface and mode of cyclic di-GMP binding. Immunity 36, 1073–1086 (2012).
Yin, Q. et al. Cyclic di-GMP sensing via the innate immune signaling protein STING. Mol. Cell 46, 735–745 (2012).
Corr, S. C. & O’Neill, L. A. Listeria monocytogenes infection in the face of innate immunity. Cell. Microbiol. 11, 703–709 (2009).
Cossart, P. Illuminating the landscape of host-pathogen interactions with the bacterium Listeria monocytogenes. Proc. Natl. Acad. Sci. USA 108, 19484–19491 (2011).
Pamer, E. G. Immune responses to Listeria monocytogenes. Nat. Rev. Immunol. 4, 812–823 (2004).
O’Connell, R. M. et al. Type I interferon production enhances susceptibility to Listeria monocytogenes infection. J. Exp. Med. 200, 437–445 (2004).
Auerbuch, V., Brockstedt, D. G., Meyer-Morse, N., O’Riordan, M. & Portnoy, D. A. Mice lacking the type I interferon receptor are resistant to Listeria monocytogenes. J. Exp. Med. 200, 527–533 (2004).
Xia, P. et al. IRTKS negatively regulates antiviral immunity through PCBP2 sumoylation-mediated MAVS degradation. Nat. Commun. 6, 8132 (2015).
Ran, F. A. et al. Genome engineering using the CRISPR-Cas9 system. Nat. Protoc. 8, 2281–2308 (2013).
Xia, P. et al. Sox2 functions as a sequence-specific DNA sensor in neutrophils to initiate innate immunity against microbial infection. Nat. Immunol. 16, 366–375 (2015).
Xia, P. et al. WASH is required for the differentiation commitment of hematopoietic stem cells in a c-Myc-dependent manner. J. Exp. Med. 211, 2119–2134 (2014).
Xia, P. et al. WASH inhibits autophagy through suppression of Beclin 1 ubiquitination. EMBO J. 32, 2685–2696 (2013).
Xia, P. et al. Insulin-InsR signaling drives multipotent progenitor differentiation toward lymphoid lineages. J. Exp. Med. 212, 2305–2321 (2015).
Wang, S. et al. Transient activation of autophagy via Sox2-mediated suppression of mTOR is an important early step in reprogramming to pluripotency. Cell Stem Cell 13, 617–625 (2013).
Dancz, C. E., Haraga, A., Portnoy, D. A. & Higgins, D. E. Inducible control of virulence gene expression in Listeria monocytogenes: temporal requirement of listeriolysin O during intracellular infection. J. Bacteriol. 184, 5935–5945 (2002).
Xia, P. et al. RNF2 is recruited by WASH to ubiquitinate AMBRA1 leading to downregulation of autophagy. Cell Res. 24, 943–958 (2014).
Acknowledgements
We thank J. Jia and Y. Teng for technical support; D. Zhou (Xiamen University) for Vav-Cre mice; J. Bennink (National Institute of Allergy and Infectious Diseases) for VSV and VACV viral strains; H. Peng (Institute of Biophysics, Chinese Academy of Sciences) for Vero cells and herpes simplex virus strain HSV-1f; and D. Fan for mouse model preparation. Supported by the National Natural Science Foundation of China (91640203, 31530093 and 31429001 to Z.F.; 81601361, 31770939 and 31570872 to P.X.; 81722023, 31671531 and 31471386 to S.W.; 8157243 and 81772646 to Y.T.; and 31601189 to X.Z.), the Strategic Priority Research Programs of the Chinese Academy of Sciences (XDB19030203 to Z.F.). Youth Innovation Promotion Association of CAS (to S.W.) and the China Postdoctoral Science Foundation (2015M571141 to P.X.).
Author information
Authors and Affiliations
Contributions
P.X. designed and performed the experiments, analyzed the data and wrote the paper; S.W. and Z.X. performed the experiments and analyzed the data; X.Z., S.M. and Y.T. generated knockout mouse models and analyzed the data; B.Y., Y.D., Y.Q. and J.L. performed some experiments; G.G. analyzed data; and Z.F. initiated the study and organized, designed and wrote the paper.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Figure 1 ERAdP-deficient mice have no impact on HSV or VSV infection.
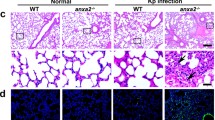
(a) Exon compositions of Cnep1r1 transcript (NM_029074.3, lower panel) and Eradp transcript (upper panel). Primers used for RT-PCR analysis of these two transcripts were annotated. (b–e) RT-PCR analyses of indicated tissues and cell types were performed using primers #1 (forward: 5’-ATCAGCCTAGCTGCTCGC-3’, reverse: 5’-GCCGACAGCGCCAACTCCC-3’) (b, c); and primers #2 (forward: 5’-AGCTGCTCGATGTCGAACGGTA-3’, reverse: 5’-CCGTCACATGTCCGACTCC AAA-3’) (d, e). n = 3 biologically independent samples. (f, g) Cnep1r1 fl/fl and Cnep1r1 fl/fl Lyz2-Cre mice were injected intravenously with HSV (1x106 pfu for each mouse) (f) or VSV (1x106 pfu for each mouse) (g), followed by survival calculation at the indicated days. n = 10 biologically independent animals. (h, i) Cnep1r1 fl/fl and Cnep1r1 fl/fl Lyz2-Cre mice were injected intravenously with HSV (1x106 pfu for each mouse) (h) or VSV (1x106 pfu for each mouse) (i), followed by brain homogenization and viral titer determination 3 days post infection. n = 5 biologically independent animals. (j, k) Cnep1r1 fl/fl and Cnep1r1 fl/fl Lyz2-Cre mice were injected intravenously with HSV (1x106 pfu for each mouse) (j) or VSV (1x106 pfu for each mouse) (k), followed by examination of serum IFN-β through ELISA 3 days post infection. n = 5 biologically independent animals. (l) Peritoneal macrophages were isolated from Cnep1r1 fl/fl and Cnep1r1 fl/fl Lyz2-Cre mice. Cells were infected with L. monocytogenes at an MOI of 0.1 for 30 min, followed by addition of gentamicin (50 μg/ml) to eliminate extracellular bacteria. Ifnb expression was examined through RT-PCR 4 h later. n = 3 biologically independent samples. (m) Peritoneal macrophages were collected from untreated Cnep1r1 fl/fl and Cnep1r1 fl/fl Lyz2-Cre mice. Macrophages were pre-stimulated with or without 100 ng/ml IFN-γ for 20 h and incubated with PBS-washed WT or DacA overexpressing L. monocytogenes (DacA-LM) at an MOI of 0.1 for 30 min in wells of 6 well plates, followed by addition of gentamicin (50 μg/ml) to eliminate extracellular bacteria. CFU was determined through clonal examination at the indicated times. n = 3 biologically independent samples for each data point. Data are shown as means±SD. *, P < 0.01. Data are representative of at least three independent experiments.
Supplementary Figure 2 ERAdP deficiency does not affect cGAMP-induced IRF3 activation.
(a–c) Peritoneal macrophages were isolated from Cnep1r1 fl/fl and Cnep1r1 fl/fl Vav-Cre mice. Cells were transfected with 2 μg/ml c-di-AMP through Lipofectamine 2000 for 4 h, followed by Tnf (a), Il6 (b) and Il1b (c) expression level determination through RT-PCR. (d–f) BMDMs from Cnep1r1 fl/fl and Cnep1r1 fl/fl Vav-Cre mice were transfected with cGAMP or c-di-AMP for 4 h, followed by immunoblotting with the indicated antibodies (d), and examination of Tnf (e) and Ifnb (f) expression via RT-PCR. (g–i) Peritoneal macrophages were isolated from WT, Cnep1r1 fl/fl Lyz2-Cre, Tmem173 –/–, Myd88 fl/fl Lyz2-Cre and Cnep1r1 fl/fl Myd88 fl/fl Lyz2-Cre mice. Cells were transfected with 2 μg/ml c-di-AMP through Lipofectamine 2000 for 4 h, followed by Tnf (g), Il6 (h) and Il1b (i) expression level determination through RT-PCR. (j–l) Peritoneal macrophages were isolated from WT, Cnep1r1 fl/fl Lyz2-Cre, Tmem173 –/–, Myd88 fl/fl Lyz2-Cre and Cnep1r1 fl/fl Myd88 fl/fl Lyz2-Cre mice. Cells were incubated with PBS-washed L. monocytogenes at an MOI of 0.1 for 30 min, followed by addition of gentamicin (50 μg/ml) to eliminate extracellular bacteria. Expression levels of Tnf (j), Il6 (k) and Il1b (l) were determined through RT-PCR. Data are shown as means±SD. *, P < 0.01; **, P < 0.001. For (a–c, e–l), n = 3 biologically independent samples. Data are representative of at least three independent experiments.
Supplementary Figure 3 The transmembrane domains of ERAdP are required for its interaction with c-di-AMP.
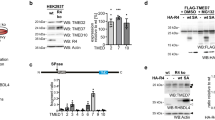
(a) Representation of ERAdP peptide sequences were shown. (b) Identified peptides and coverage of ERAdP through mass spectrometry were shown. (c) Identified peptides and coverage of STING through mass spectrometry were shown. (d, e) 1 μg recombinant GST-ERAdP was incubated with 1 μCi radiolabeled c-di-[32P]AMP with different doses of recombinant GST-STING (d) or GST-RECON (e) and separated through SDS-PAGE, followed by phosphor visualization using Typhoon FLA 7000. 1 μg recombinant GST was used as a negative control. (f) WT BMDMs were mock treated, followed by immunostaining with antibodies against ERAdP and biotin. Nuclei were counterstained with DAPI. Scale bar, 10 μm. (g) 50 ng/μl biotin-c-di-AMP or biotin-c-di-GMP was incubated with various amounts of GST or GST-ERAdP, followed by addition of anti-GST-conjugated AlphaLISA Acceptor Beads. Streptavidin-coated Alpha Donor Beads were then added to reaction mixtures for measurement by EnVision Xcite Multilabel Reader. (h) Scheme for ERAdP truncations. N, N terminal domain; TM, transmembrane domain; C, C terminal domain. (i) GST and GST-ERAdP truncation proteins were incubated with WT BMDM lysates in the presence of biotin-c-di-AMP, followed by pulldown assay of streptavidin-coupled beads. Precipitates were immunoblotted with anti-GST antibody. (j) WT BMDMs were infected with lentiviruses encoding Flag-tagged full length ERAdP and its truncations, followed by puromycin selection for stable expression of exogenous genes. Cell lysates were incubated with biotin-c-di-AMP, followed by further incubation with streptavidin-coupled beads. Beads were then washed and immunoblotted with the indicated antibodies. (k) ERAdP deficient BMDMs were infected with lentiviruses encoding Flag-tagged full length ERAdP and its truncations, followed by puromycin selection for stable expression of exogenous genes. Cells were immunoblotted with the indicated antibodies. (l) ERAdP deficient BMDMs were rescued with Flag-tagged full length ERAdP and its truncations as in (k). Rescued cells were transfected with biotin-c-di-AMP for 4 h, followed by immunostaining with antibodies against Flag and biotin. Nuclei were counterstained with DAPI. Scale bar, 10 μm. (m, n) Cnep1r1 fl/fl and Cnep1r1 fl/fl Lyz2-Cre mice were injected intravenously with 5x105 WT or DacA-LM, followed by clonal examination of CFUs in liver (m) and spleen (n) 3 days post infection. n = 7 biologically independent animals. Data are shown as means±SD. *, P < 0.05; **, P < 0.01. Data are representative of at least three independent experiments.
Supplementary Figure 4 c-di-AMP is required for the interaction of ERAdP with TAK1.
(a)WT BMDMs were mock transfected, followed by immunostaining with antibodies against ERAdP, TAK1 and biotin. Nuclei were counterstained with DAPI. Scale bar, 10 μm. (b) BMDM cells from Cnep1r1 fl/fl and Cnep1r1 fl/fl Vav-Cre mice were incubated with PBS-washed L. monocytogenes at an MOI of 0.1 for 30 min, followed by addition of gentamicin (50 μg/ml) to eliminate extracellular bacteria. Cells were further incubated for 4 h, followed by immunoprecipitation with antibody against ERAdP and immunoblotting with the indicated antibodies. (c, d) WT BMDMs were incubated with PBS-washed CFP-expressing L. monocytogenes at an MOI of 0.1 for 30 min, followed by addition of gentamicin (50 μg/ml) to eliminate extracellular bacteria. Cells were further incubated for 4 h, followed by immunostaining with antibodies against ERAdP and TAK1. Nuclei were counterstained with DAPI (c). Colocalization of ERAdP and TAK1 were calculated (d). For (d), n = 3 biologically independent experiments with at least 200 cells counted for each experiment. Scale bar, 10 μm. (e) GST and GST-ERAdP variants were incubated with WT BMDM lysates in the presence of c-di-AMP, followed by GST-pulldown assay. Precipitates were immunoblotted with the indicated antibodies. (f) BiFC assay was performed to verify the self-association of ERAdP after c-di-AMP exposure. WT BMDMs were infected with lentiviruses carrying VN-ERAdP and ERAdP-VC for 24 h, and then transfected with c-di-AMP for 4 h, followed by confocal microscopy. (g) Scheme of confocal examination for GFP-tagged and mCherry-tagged TAK1 by an in vitro incubation system. (h) HEK293T cells stably expressing a NF-κB reporter were transfected with various amounts of plasmids encoding ERAdP or STING for 24 h, followed by transfection of 2 μg/ml c-di-AMP for 16 h. NF-κB activation was determined through luciferase assay. n = 3 biologically independent samples. Data are shown as means±SD. *, P < 0.001. Data are representative of at least three independent experiments.
Supplementary Figure 5 TAK1-deficient mice are highly susceptible to L. monocytogenes infection.
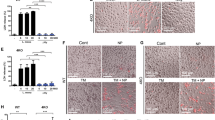
(a) BMDMs from Map3k7 fl/fl and Map3k7 fl/fl Lyz2-Cre mice were immunoblotted with the indicated antibodies. (b, c) Map3k7 fl/fl and Map3k7 fl/fl Lyz2-Cre mice were injected intravenously with 5x105 L. monocytogenes, followed by examination of serum TNF (b) and IL-6 (c) through ELISA analysis 3 days post infection. n = 5 biologically independent animals. (d) Map3k7 fl/fl and Map3k7 fl/fl Lyz2-Cre mice were injected intravenously with 1x106 L. monocytogenes, followed by survival calculation at the indicated days. n = 10 biologically independent animals. (e, f) BMDM cells from Cnep1r1 fl/fl Map3k7 fl/fl, Cnep1r1 fl/fl Lyz2-Cre, Map3k7 fl/fl Lyz2-Cre and Cnep1r1 fl/fl Map3k7 fl/fl Lyz2-Cre mice were treated with Pam3CSK4 (100 ng/ml) (e) or with heat killed L. monocytogenes (HKLM) (1x108 cells/ml) (f) for 4 h, followed by Tnf expression level determination through RT-PCR. n = 3 biologically independent samples. Data are shown as means±SD. *, P < 0.001. Data are representative of at least three independent experiments.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1-5
Rights and permissions
About this article
Cite this article
Xia, P., Wang, S., Xiong, Z. et al. The ER membrane adaptor ERAdP senses the bacterial second messenger c-di-AMP and initiates anti-bacterial immunity. Nat Immunol 19, 141–150 (2018). https://doi.org/10.1038/s41590-017-0014-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41590-017-0014-x
- Springer Nature America, Inc.
This article is cited by
-
Transdifferentiation of tumor infiltrating innate lymphoid cells during progression of colorectal cancer
Cell Research (2020)
-
Redox homeostasis maintained by GPX4 facilitates STING activation
Nature Immunology (2020)
-
Global transcriptional changes in response to cGAMP depend on STING in human THP-1 cells
Cellular & Molecular Immunology (2018)
-
ERAdP standing in the shadow of STING innate immune signaling
Nature Immunology (2018)
-
Sensing bacterial messages
Nature Reviews Immunology (2018)