Abstract
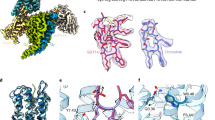
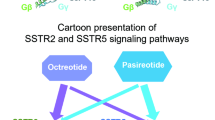
G protein-coupled receptors (GPCRs) modulate every aspect of physiological functions mainly through activating heterotrimeric G proteins. A majority of GPCRs promiscuously couple to multiple G protein subtypes. Here we validate that in addition to the well-known Gi/o pathway, somatostatin receptor 2 and 5 (SSTR2 and SSTR5) couple to the Gq/11 pathway and show that smaller ligands preferentially activate the Gi/o pathway. We further determined cryo-electron microscopy structures of the SSTR2‒Go and SSTR2‒Gq complexes bound to octreotide and SST-14. Structural and functional analysis revealed that G protein selectivity of SSTRs is not only determined by structural elements in the receptor–G protein interface, but also by the conformation of the agonist-binding pocket. Accordingly, smaller ligands fail to stabilize a broader agonist-binding pocket of SSTRs that is required for efficient Gq/11 coupling but not Gi/o coupling. Our studies facilitate the design of drugs with selective G protein signaling to improve therapeutic efficacy.






Similar content being viewed by others
References
Fredriksson, R., Lagerstrom, M. C., Lundin, L. G. & Schioth, H. B. The G-protein-coupled receptors in the human genome form five main families. Phylogenetic analysis, paralogon groups, and fingerprints. Mol. Pharmacol. 63, 1256–1272 (2003).
Pierce, K. L., Premont, R. T. & Lefkowitz, R. J. Seven-transmembrane receptors. Nat. Rev. Mol. Cell Biol. 3, 639–650 (2002).
Wilkie, T. M. et al. Evolution of the mammalian G protein α subunit multigene family. Nat. Genet. 1, 85–91 (1992).
DeWire, S. M., Ahn, S., Lefkowitz, R. J. & Shenoy, S. K. Beta-arrestins and cell signaling. Annu Rev. Physiol. 69, 483–510 (2007).
Olsen, R. H. J. et al. TRUPATH, an open-source biosensor platform for interrogating the GPCR transducerome. Nat. Chem. Biol. 16, 841–849 (2020).
Inoue, A. et al. Illuminating G-protein-coupling selectivity of GPCRs. Cell 177, 1933–1947 (2019).
Smith, J. S., Lefkowitz, R. J. & Rajagopal, S. Biased signalling: from simple switches to allosteric microprocessors. Nat. Rev. Drug Disco. 17, 243–260 (2018).
Bermudez, M., Nguyen, T. N., Omieczynski, C. & Wolber, G. Strategies for the discovery of biased GPCR ligands. Drug Disco. Today 24, 1031–1037 (2019).
Rajagopal, K. et al. β-arrestin2-mediated inotropic effects of the angiotensin II type 1A receptor in isolated cardiac myocytes. Proc. Natl Acad. Sci. USA 103, 16284–16289 (2006).
Zhu, W. Z. et al. Dual modulation of cell survival and cell death by β(2)-adrenergic signaling in adult mouse cardiac myocytes. Proc. Natl Acad. Sci. USA 98, 1607–1612 (2001).
Li, Y. Q. et al. Gq/11α and Gsα mediate distinct physiological responses to central melanocortins. J. Clin. Invest. 126 (2016).
Wootten, D., Christopoulos, A., Marti-Solano, M., Babu, M. M. & Sexton, P. M. Mechanisms of signalling and biased agonism in G protein-coupled receptors. Nat. Rev. Mol. Cell Biol. 19, 638–653 (2018).
Chaturvedi, M., Maharana, J. & Shukla, A. K. Terminating G-protein coupling: structural snapshots of GPCR-β-arrestin complexes. Cell 180, 1041–1043 (2020).
Theodoropoulou, M. & Stalla, G. K. Somatostatin receptors: from signaling to clinical practice. Front Neuroendocrinol. 34, 228–252 (2013).
Patel, Y. C. Molecular pharmacology of somatostatin receptor subtypes. J. Endocrinol. Invest 20, 348–367 (1997).
de Lecea, L. et al. A cortical neuropeptide with neuronal depressant and sleep-modulating properties. Nature 381, 242–245 (1996).
Stueven, A. K. et al. Somatostatin analogues in the treatment of neuroendocrine tumors: past, present and future. Int. J. Mol. Sci. 20, 3049 (2019).
Ben-Shlomo, A. & Melmed, S. Somatostatin agonists for treatment of acromegaly. Mol. Cell. Endocrinol. 286, 192–198 (2008).
Maecke, H. R. & Reubi, J. C. Somatostatin receptors as targets for nuclear medicine imaging and radionuclide treatment. J. Nucl. Med. 52, 841–844 (2011).
Jiang, W. & Zheng, S. Structural insights into galanin receptor signaling. Proc. Natl Acad. Sci. USA 119, e2121465119 (2022).
Wan, Q. et al. Mini G protein probes for active G protein-coupled receptors (GPCRs) in live cells. J. Biol. Chem. 293, 7466–7473 (2018).
Akbar, M. et al. Phospholipase C activation and Ca2+ mobilization by cloned human somatostatin receptor subtypes 1–5, in transfected COS-7 cells. FEBS Lett. 348, 192–196 (1994).
Wilkinson, G. F., Feniuk, W. & Humphrey, P. P. Homologous and heterologous desensitisation of somatostatin-induced increases in intracellular Ca2+ and inositol 1,4,5-trisphosphate in CHO-K1 cells expressing human recombinant somatostatin sst5 receptors. Eur. J. Pharmacol. 340, 277–285 (1997).
Takasaki, J. et al. A novel Gαq/11-selective inhibitor. J. Biol. Chem. 279, 47438–47445 (2004).
Yang, L. et al. Spiro[1H-indene-1,4′-piperidine] derivatives as potent and selective non-peptide human somatostatin receptor subtype 2 (sst2) agonists. J. Med. Chem. 41, 2175–2179 (1998).
Harris, J. A. et al. Selective G protein signaling driven by substance P-neurokinin receptor dynamics. Nat. Chem. Biol. 18, 109–115 (2022).
Erlandson, S. C. et al. The relaxin receptor RXFP1 signals through a mechanism of autoinhibition. Preprint at bioRxiv https://doi.org/10.1101/2022.01.22.477343 (2022).
Teng, X. et al. Structural insights into G protein activation by D1 dopamine receptor. Sci. Adv. 8, eabo4158 (2022).
Maeda, S. et al. Development of an antibody fragment that stabilizes GPCR/G-protein complexes. Nat. Commun. 9, 3712 (2018).
Melacini, G., Zhu, Q. & Goodman, M. Multiconformational NMR analysis of sandostatin (octreotide): equilibrium between β-sheet and partially helical structures. Biochemistry 36, 1233–1241 (1997).
Spiliopoulou, M. et al. New perspectives in macromolecular powder diffraction using single-photon-counting strip detectors: high-resolution structure of the pharmaceutical peptide octreotide. Acta Crystallogr. A Found. Adv. 77, 186–195 (2021).
Strnad, J. & Hadcock, J. R. Identification of a critical aspartate residue in transmembrane domain three necessary for the binding of somatostatin to the somatostatin receptor SSTR2. Biochem. Biophys. Res. Commun. 216, 913–921 (1995).
Kaupmann, K. et al. Two amino acids, located in transmembrane domains VI and VII, determine the selectivity of the peptide agonist SMS 201-995 for the SSTR2 somatostatin receptor. EMBO J. 14, 727–735 (1995).
Robertson, M. J., Meyerowitz, J. G., Panova, O., Borrelli, K. & Skiniotis, G. Plasticity in ligand recognition at somatostatin receptors. Nat. Struct. Mol. Biol. 29, 210–217 (2022).
Chen, L. N. et al. Structures of the endogenous peptide- and selective non-peptide agonist-bound SSTR2 signaling complexes. Cell Res. 32, 785–788 (2022).
Zhao, W. et al. Structural insights into ligand recognition and selectivity of somatostatin receptors. Cell Res. 32, 761–772 (2022).
Liu, Q. et al. Ligand recognition and G-protein coupling selectivity of cholecystokinin A receptor. Nat. Chem. Biol. 17, 1238–1244 (2021).
Wang, J., Hua, T. & Liu, Z. J. Structural features of activated GPCR signaling complexes. Curr. Opin. Struct. Biol. 63, 82–89 (2020).
Maeda, S., Qu, Q., Robertson, M. J., Skiniotis, G. & Kobilka, B. K. Structures of the M1 and M2 muscarinic acetylcholine receptor/G-protein complexes. Science 364, 552–557 (2019).
Duan, J. et al. Molecular basis for allosteric agonism and G protein subtype selectivity of galanin receptors. Nat. Commun. 13, 1364 (2022).
Moro, O., Lameh, J., Hogger, P. & Sadee, W. Hydrophobic amino acid in the i2 loop plays a key role in receptor-G protein coupling. J. Biol. Chem. 268, 22273–22276 (1993).
Rasmussen, S. G. et al. Crystal structure of the beta2 adrenergic receptor-Gs protein complex. Nature 477, 549–555 (2011).
Krishna Kumar, K. et al. Structure of a Signaling Cannabinoid Receptor 1-G Protein Complex. Cell 176, 448–458 (2019).
Chen, X. P. et al. Structural determinants in the second intracellular loop of the human cannabinoid CB1 receptor mediate selective coupling to G(s) and G(i). Br. J. Pharmacol. 161, 1817–1834 (2010).
Skorpen, F. et al. The rare Arg181Cys mutation in the mu opioid receptor can abolish opioid responses. Acta Anaesthesiol. Scand. 60, 1084–1091 (2016).
Koehl, A. et al. Structure of the µ-opioid receptor–Gi protein complex. Nature 558, 547–552 (2018).
DeVree, B. T. et al. Allosteric coupling from G protein to the agonist-binding pocket in GPCRs. Nature 535, 182–186 (2016).
Sharp, A. J., Hayes, A. R. & Grossman, A. High-dose somatostatin analogues for progressive neuroendocrine tumours. Eur. Endocrinol. 16, 93–95 (2020).
Cordoba-Chacon, J. et al. Somatostatin dramatically stimulates growth hormone release from primate somatotrophs acting at low doses via somatostatin receptor 5 and cyclic AMP. J. Neuroendocrinol. 24, 453–463 (2012).
Holze, J. et al. Ligand-specific allosteric coupling controls G-protein-coupled receptor signaling. ACS Pharm. Transl. Sci. 3, 859–867 (2020).
Kroeze, W. K. et al. PRESTO-Tango as an open-source resource for interrogation of the druggable human GPCRome. Nat. Struct. Mol. Biol. 22, 362–369 (2015).
Nehme, R. et al. Mini-G proteins: Novel tools for studying GPCRs in their active conformation. PLoS ONE 12, e0175642 (2017).
Zheng, S. Q. et al. MotionCor2: anisotropic correction of beam-induced motion for improved cryo-electron microscopy. Nat. Methods 14, 331–332 (2017).
Punjani, A., Rubinstein, J. L., Fleet, D. J. & Brubaker, M. A. cryoSPARC: algorithms for rapid unsupervised cryo-EM structure determination. Nat. Methods 14, 290–296 (2017).
Sanchez-Garcia, R. et al. DeepEMhancer: a deep learning solution for cryo-EM volume post-processing. Commun. Biol. 4, 874 (2021).
Kim, K. et al. Structure of a hallucinogen-activated Gq-coupled 5-HT2A serotonin receptor. Cell 182, 1574–1588 (2020).
Adams, P. D. et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr D. Biol. Crystallogr. 66, 213–221 (2010).
Acknowledgements
We thank S. Rajagopal for his suggestion on interpretation of bias signaling. We thank staff at Shuimu BioSciences for their help with cryo-EM data collection. All electron microscopy images were collected at Shuimu BioSciences. This work was supported by the Chinese Ministry of Science and Technology, Beijing Municipal Science & Technology Commission (Z201100005320012, to S.Z.) and Tsinghua University.
Author information
Authors and Affiliations
Contributions
S.C. and X.T. purified the protein complex, collected cryo-EM data and performed cryo-EM data processing and model building and performed all functional assays with the supervision by S.Z. S.Z. and S.C. wrote the manuscripts.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Chemical Biology thanks Asuka Inoue and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 SSTR2 and SSTR5 activate the Gq/11 pathway.
a, Primary sequences of SST-14, CST-17, octreotide and seglitide, and chemical structure of L-054,264. b-d, Dose-response curves of cAMP inhibition in HEK293 cells expressing SSTR1 (b), SSTR3 (c), or SSTR4 (d) upon stimulation with octreotide and SST-14. e, NanoBiT mini-Gq recruitment assays demonstrated that mini-Gq is recruited to the SST-14 activated SSTR2 and SSTR5. Measurements were performed with three independently biological replicates. f-h, Dose-response curves of IP1 production in cells expressing SSTR1 (f), SSTR3 (g) or SSTR4 (h) treated with SST-14 or octreotide. i-j, Effects of a specific Gq/11 inhibitor YM-254890 on activation of Gq/11(i) and Gi/o (j) by SSTR2. Each data point of dose-response curves is shown as mean ± SEM from three independent experiments. All IP1 data are normalized as percentage of maximum response of SSTR2 treated with SST-14.
Extended Data Fig. 2 Smaller ligands lead to the selective Gi/o coupling of SSTR2 and SSTR5.
a-b, Summary of the maximum effects (Emax), potencies (pEC50) and bias factors of different ligands at SSTR2 (a) and SSTR5 (b) from the cAMP inhibition assay and IP1 assay. Emax represents maximum effects of ligands. pEC50 is the logarithm of the concentration of ligands that produces 50% of Emax. Bias factor is calculated by comparing the Gi/o pathway to the Gq/11 pathway with SST-14 as a reference ligand. Positive value of bias factor indicates Gi/o-biased. n, number of independent repeats. ND, not determined because of no response. c-d, BRET assay for monitoring Go (c) and Gq (d) activation by SSTR2 stimulated by different ligands. All data represent mean ± SEM from three independent repeats. e, Summary of the maximum effects (Emax), potencies (pEC50) and bias factors of different ligands at SSTR2 from the BRET assay. ND, not determined because of low response and poor fit.
Extended Data Fig. 3 Cryo-EM data collection and processing of the octreotide-bound SSTR2-mini-Gαo/Gβ1γ2/scFv16 complex.
a, Size-exclusion chromatography profile. b, SDS-PAGE of the purified complex. c, Representative micrograph from 2349 micrographs. Scale bar represents 50 nm. d, Representative 2D classes. e, Cryo-EM data workflow. f, Gold standard FSC curves.
Extended Data Fig. 4 Cryo-EM data collection and processing of the octreotide- and SST-14-bound SSTR2-mini-Gαq/Gβ1γ2/scFv16 complex.
a, Size-exclusion chromatography profile. b, SDS-PAGE of the purified complex. c-d, Representative micrograph from 6480 micrographs (c) and 2D classes (d) of the octreotide-bound SSTR2-mini-Gq/Gβ1γ2/scFv16 complex. Scale bar represents 50 nm. e, Representative micrograph of the SST-14-bound SSTR2-mini-Gq/Gβ1γ2/scFv16 complex from 4881 micrographs. Scale bar represents 50 nm. f-g, Cryo-EM data workflow (f) and gold standard FSC curves (g) of the octreotide-bound SSTR2-mini-Gq/Gβ1γ2/scFv16 complex. h-i, Cryo-EM data workflow (h) and gold standard FSC curves (i) of the SST-14-bound SSTR2-mini-Gαq/Gβ1γ2/scFv16 complex.
Extended Data Fig. 5 Characterization of ligand binding of SSTR2.
a, Solution structures of octreotide alone in two different states. b, Crystal structure of octreotide alone in two different states. c-d, NanoBiT assay shows that SSTR2 equally activates Gi1 and Go when stimulated by octreotide (c) or SST-14 (d). Each data point is presented as mean +/− SEM from three independent experiments. e, Sequence alignment of SSTR subtypes from Homo sapiens. Secondary structures are shown above the alignment. Residues that are important for ligand binding and G protein activation of SSTR2 based on the structure and the NanoBiT Gi1 dissociation assay are denoted with orange circles below the alignment.
Extended Data Fig. 6 Comparison of structures of CCKAR and NK1R bound to different G protein subtypes.
a, Structural overlay of the CCKAR-Gi and CCKAR-Gq complexes from two orthogonal views. b, Structural overlay of the NK1R-miniGs and NK1R-miniGsq complexes from two orthogonal views.
Extended Data Fig. 7 Interaction interfaces between SSTR2 and the C-terminal helix of Gα.
a, Structural superposition of the octreotide-bound SSTR2-Go and SSTR2-Gq complex with the receptor aligned. b, Detailed interactions between TM3, TM5 and TM6 of SSTR2 and the extreme C terminus of Gαo. c, Detailed interactions between TM3, TM5 and TM6 of SSTR2 and the extreme C terminus of Gαq. d-e, Dose-response curves of cAMP inhibition (c) and IP1 production (d) in cells expressing SSTR2 or N186ECL2G mutant upon stimulation by SST-14. Each data point is presented as mean +/− SEM from three independent experiments.
Supplementary information
Supplementary Information
Supplementary Tables 1–5 and Supplementary Note.
Source data
Source Data Fig. 1
Statistical source data.
Source Data Fig. 3
Statistical source data.
Source Data Fig. 4
Statistical source data.
Source Data Fig. 5
Statistical source data.
Source Data Extended Data Fig. 1
Statistical source data.
Source Data Extended Data Fig. 2
Statistical source data.
Source Data Extended Data Fig. 3
Unprocessed gel.
Source Data Extended Data Fig. 4
Unprocessed gel.
Source Data Extended Data Fig. 5
Statistical source data.
Source Data Extended Data Fig. 7
Statistical source data.
Rights and permissions
Springer Nature or its licensor holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Chen, S., Teng, X. & Zheng, S. Molecular basis for the selective G protein signaling of somatostatin receptors. Nat Chem Biol 19, 133–140 (2023). https://doi.org/10.1038/s41589-022-01130-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41589-022-01130-3
- Springer Nature America, Inc.
This article is cited by
-
Structural basis of tethered agonism and G protein coupling of protease-activated receptors
Cell Research (2024)
-
Balancing G protein selectivity and efficacy in the adenosine A2A receptor
Nature Chemical Biology (2024)
-
Molecular simulations of SSTR2 dynamics and interaction with ligands
Scientific Reports (2023)