Abstract
Measures of selective constraint on genes have been used for many applications, including clinical interpretation of rare coding variants, disease gene discovery and studies of genome evolution. However, widely used metrics are severely underpowered at detecting constraints for the shortest ~25% of genes, potentially causing important pathogenic mutations to be overlooked. Here we developed a framework combining a population genetics model with machine learning on gene features to enable accurate inference of an interpretable constraint metric, shet. Our estimates outperform existing metrics for prioritizing genes important for cell essentiality, human disease and other phenotypes, especially for short genes. Our estimates of selective constraint should have wide utility for characterizing genes relevant to human disease. Finally, our inference framework, GeneBayes, provides a flexible platform that can improve the estimation of many gene-level properties, such as rare variant burden or gene expression differences.






Similar content being viewed by others
Data availability
Posterior means and 95% credible intervals for shet are available in Supplementary Table 1. Data sources for pLOF annotations, CpG methylation levels, exome sequencing coverage, variant frequencies and mappability/segmental duplication annotations are available in Supplementary Table 5. A description of the gene features is available in Supplementary Table 8. Posterior densities for shet, likelihoods for shet, LOF variants with misannotation probabilities and gene feature tables are available in ref. 83. Additional publicly available datasets used in this study are described in Methods and Supplementary Information and are accessible at IMPC essential genes (https://www.ebi.ac.uk/mi/impc/essential-genes-search/); pLOF annotations (gs://gnomad-public/papers/2019-tx-annotation/pre_computed/all.possible.snvs.tx_annotated.GTEx.v7.021520.tsv); mean methylation for CpG sites (gs://gcp-public-data–gnomad/resources/methylation); exome sequencing coverage (gs://gcp-public-data–gnomad/release/2.1/coverage/exomes/gnomad.exomes.coverage.summary.tsv.bgz); variant frequencies (gs://gcp-public-data–gnomad/release/2.1.1/vcf/exomes/gnomad.exomes.r2.1.1.sites.vcf.bgz); low mappability and segmental duplications (https://ftp-trace.ncbi.nlm.nih.gov/ReferenceSamples/giab/release/genome-stratifications/v3.1/GRCh37/Union/GRCh37_alllowmapandsegdupregions.bed.gz); ClinVar variants (https://ftp.ncbi.nlm.nih.gov/pub/clinvar/vcf_GRCh38/); DepMap 22Q2 release (https://depmap.org/portal/download/all/); DDD annotations (https://www.deciphergenomics.org/ddd/ddgenes); HPO phenotype-to-gene annotations (http://purl.obolibrary.org/obo/hp/hpoa/phenotype_to_genes.txt); DNMs from developmental disorder patients5; UK Biobank summary statistics (https://nealelab.github.io/UKBB_ldsc); RNA-seq from chimpanzee/human cortical models28; GTEx v8 release29.
Code availability
GeneBayes and code for estimating shet are available at https://github.com/tkzeng/GeneBayes and in ref. 84. Analysis code is available in ref. 85. All analyses were performed using Python v3.8, Python v3.9 or R v4.2. To train models, we used a modified version of NGBoost (v0.3.12)16,86 (https://github.com/tkzeng/ngboost), XGBoost (v2.0.2)87 and PyTorch (v1.12.1)88. Likelihoods were computed with fastDTWF (v.0.0.3)15 (https://github.com/jeffspence/fastDTWF). For hyperparameter tuning, we used shap-hypetune v0.2 (https://github.com/cerlymarco/shap-hypetune). For heritability enrichment analyses, we used ldsc (v1.0.1)89. For additional analyses, we used NumPy (v1.26.0)90, SciPy (v1.8.1)91, Pandas (v2.1.3)92, Scikit-learn (1.3.0)93 and Statsmodels (v0.14.0)94.
References
Cassa, C. A. et al. Estimating the selective effects of heterozygous protein-truncating variants from human exome data. Nat. Genet. 49, 806–810 (2017).
Weghorn, D. et al. Applicability of the mutation–selection balance model to population genetics of heterozygous protein-truncating variants in humans. Mol. Biol. Evol. 36, 1701–1710 (2019).
Fuller, Z. L., Berg, J. J., Mostafavi, H., Sella, G. & Przeworski, M. Measuring intolerance to mutation in human genetics. Nat. Genet. 51, 772–776 (2019).
Agarwal, I., Fuller, Z. L., Myers, S. R. & Przeworski, M. Relating pathogenic loss-of-function mutations in humans to their evolutionary fitness costs. eLife 12, e83172 (2023).
Kaplanis, J. et al. Evidence for 28 genetic disorders discovered by combining healthcare and research data. Nature 586, 757–762 (2020).
Fu, J. M. et al. Rare coding variation provides insight into the genetic architecture and phenotypic context of autism. Nat. Genet. 54, 1320–1331 (2022).
Whiffin, N. et al. The effect of LRRK2 loss-of-function variants in humans. Nat. Med. 26, 869–877 (2020).
Gazal, S. et al. Combining SNP-to-gene linking strategies to identify disease genes and assess disease omnigenicity. Nat. Genet. 54, 827–836 (2022).
Wang, X. & Goldstein, D. B. Enhancer domains predict gene pathogenicity and inform gene discovery in complex disease. Am. J. Hum. Genet. 106, 215–233 (2020).
Mostafavi, H., Spence, J. P., Naqvi, S. & Pritchard, J. K. Systematic differences in discovery of genetic effects on gene expression and complex traits. Nat. Genet. 55, 1866–1875 (2023).
Lek, M. et al. Analysis of protein-coding genetic variation in 60,706 humans. Nature 536, 285–291 (2016).
Karczewski, K. J. et al. The mutational constraint spectrum quantified from variation in 141,456 humans. Nature 581, 434–443 (2020).
Gillespie, J. H. Population Genetics: A Concise Guide (JHU Press, 2004).
LaPolice, T. M. & Huang, Y. F. An unsupervised deep learning framework for predicting human essential genes from population and functional genomic data. BMC Bioinformatics 24, 347 (2023).
Spence, J. P., Zeng, T., Mostafavi, H. & Pritchard, J. K. Scaling the discrete-time Wright–Fisher model to biobank-scale datasets. Genetics 225, iyad168 (2023).
Duan, T. et al. Ngboost: natural gradient boosting for probabilistic prediction. In Proc. International Conference on Machine Learning (eds Daumé, H. III & Singh, A.) 2690–2700 (PMLR, 2020).
Ewens, W. J. Mathematical Population Genetics: Theoretical Introduction Vol. 27 (Springer, 2004).
Agarwal, I. & Przeworski, M. Mutation saturation for fitness effects at human CpG sites. eLife 10, e71513 (2021).
Huang, Y. F. Unified inference of missense variant effects and gene constraints in the human genome. PLoS Genet. 16, e1008922 (2020).
Da Costa, L., Leblanc, T. & Mohandas, N. Diamond–Blackfan anemia. Blood 136, 1262–1273 (2020).
Berger, W. et al. Mutations in the candidate gene for Norrie disease. Hum. Mol. Genet. 1, 461–465 (1992).
Howard, T. D. et al. Mutations in TWIST, a basic helix–loop–helix transcription factor, in Saethre–Chotzen syndrome. Nat. Genet. 15, 36–41 (1997).
Ghouzzi, V. E. et al. Mutations of the TWIST gene in the Saethre–Chotzene syndrome. Nat. Genet. 15, 42–46 (1997).
Meyers, R. M. et al. Computational correction of copy number effect improves specificity of CRISPR–Cas9 essentiality screens in cancer cells. Nat. Genet. 49, 1779–1784 (2017).
Ghandi, M. et al. Next-generation characterization of the cancer cell line encyclopedia. Nature 569, 503–508 (2019).
Wright, C. F. et al. Genomic diagnosis of rare pediatric disease in the United Kingdom and Ireland. N. Engl. J. Med. 388, 1559–1571 (2023).
Köhler, S. et al. The Human Phenotype Ontology in 2021. Nucleic Acids Res. 49, D1207–D1217 (2021).
Agoglia, R. M. et al. Primate cell fusion disentangles gene regulatory divergence in neurodevelopment. Nature 592, 421–427 (2021).
GTEx Consortium The GTEx Consortium atlas of genetic regulatory effects across human tissues. Science 369, 1318–1330 (2020).
Basha, O. et al. Differential network analysis of multiple human tissue interactomes highlights tissue-selective processes and genetic disorder genes. Bioinformatics 36, 2821–2828 (2020).
Gao, S. et al. Tracing the temporal-spatial transcriptome landscapes of the human fetal digestive tract using single-cell RNA-sequencing. Nat. Cell Biol. 20, 721–734 (2018).
Charlesworth, B. et al. Evolution in Age-Structured Populations Vol. 2 (Cambridge University Press, 1994).
Barrio-Hernandez, I. et al. Network expansion of genetic associations defines a pleiotropy map of human cell biology. Nat. Genet. 55, 389–398 (2023).
Van Dam, S., Vosa, U., van der Graaf, A., Franke, L. & de Magalhaes, J. P. Gene co-expression analysis for functional classification and gene–disease predictions. Brief. Bioinform. 19, 575–592 (2018).
Nasser, J. et al. Genome-wide enhancer maps link risk variants to disease genes. Nature 593, 238–243 (2021).
Wieder, N. et al. Differences in 5′ untranslated regions highlight the importance of translational regulation of dosage sensitive genes. Genome Biol. 25, 111 (2024).
Sella, G. & Barton, N. H. Thinking about the evolution of complex traits in the era of genome-wide association studies. Annu. Rev. Genomics Hum. Genet. 20, 461–493 (2019).
Charlesworth, B. Effective population size and patterns of molecular evolution and variation. Nat. Rev. Genet. 10, 195–205 (2009).
Simons, Y. B., Mostafavi, H., Smith, C. J., Pritchard, J. K. & Sella, G. Simple scaling laws control the genetic architectures of human complex traits. Preprint at bioRxiv https://doi.org/10.1101/2022.10.04.509926 (2022).
Mathieson, I. & Terhorst, J. Direct detection of natural selection in Bronze Age Britain. Genome Res. 32, 2057–2067 (2022).
Emdin, C. A. et al. Phenotypic characterization of genetically lowered human lipoprotein(a) levels. J. Am. Coll. Cardiol. 68, 2761–2772 (2016).
Langsted, A., Nordestgaard, B. G. & Kamstrup, P. R. Low lipoprotein(a) levels and risk of disease in a large, contemporary, general population study. Eur. Heart J. 42, 1147–1156 (2021).
Rausell, A. et al. Common homozygosity for predicted loss-of-function variants reveals both redundant and advantageous effects of dispensable human genes. Proc. Natl Acad. Sci. USA 117, 13626–13636 (2020).
Reyes-Soffer, G. et al. Lipoprotein(a): a genetically determined, causal, and prevalent risk factor for atherosclerotic cardiovascular disease: a scientific statement from the American Heart Association. Arterioscler. Thromb. Vasc. Biol. 42, e48–e60 (2022).
Millar, D. S. et al. Molecular genetic analysis of severe protein C deficiency. Hum. Genet. 106, 646–653 (2000).
Romeo, G. et al. Hereditary thrombophilia: identification of nonsense and missense mutations in the protein C gene. Proc. Natl Acad. Sci. USA 84, 2829–2832 (1987).
O’Connor, L. J. et al. Extreme polygenicity of complex traits is explained by negative selection. Am. J. Hum. Genet. 105, 456–476 (2019).
Benton, M. L. et al. The influence of evolutionary history on human health and disease. Nat. Rev. Genet. 22, 269–283 (2021).
Gulko, B., Hubisz, M. J., Gronau, I. & Siepel, A. A method for calculating probabilities of fitness consequences for point mutations across the human genome. Nat. Genet. 47, 276–283 (2015).
Huang, Y. F., Gulko, B. & Siepel, A. Fast, scalable prediction of deleterious noncoding variants from functional and population genomic data. Nat. Genet. 49, 618–624 (2017).
Huang, Y. F. & Siepel, A. Estimation of allele-specific fitness effects across human protein-coding sequences and implications for disease. Genome Res. 29, 1310–1321 (2019).
Chen, S. et al. A genomic mutational constraint map using variation in 76,156 human genomes. Nature 625, 92–100 (2024).
Satterstrom, F. K. et al. Large-scale exome sequencing study implicates both developmental and functional changes in the neurobiology of autism. Cell 180, 568–584 (2020).
Gardner, E. J. et al. Reduced reproductive success is associated with selective constraint on human genes. Nature 603, 858–863 (2022).
He, X. et al. Integrated model of de novo and inherited genetic variants yields greater power to identify risk genes. PLoS Genet. 9, e1003671 (2013).
Zhu, X. & Stephens, M. Bayesian large-scale multiple regression with summary statistics from genome-wide association studies. Ann. Appl. Stat. 11, 1561–1592 (2017).
Boyeau, P. et al. An empirical Bayes method for differential expression analysis of single cells with deep generative models. Proc. Natl Acad. Sci. USA 120, e2209124120 (2023).
Des Portes, V. et al. A novel CNS gene required for neuronal migration and involved in X-linked subcortical laminar heterotopia and lissencephaly syndrome. Cell 92, 51–61 (1998).
Nascimento, R. M., Otto, P. A., de Brouwer, A. P. & Vianna-Morgante, A. M. UBE2A, which encodes a ubiquitin-conjugating enzyme, is mutated in a novel X-linked mental retardation syndrome. Am. J. Hum. Genet. 79, 549–555 (2006).
Stevenson, R. E. et al. Renpenning syndrome comes into focus. Am. J. Med. Genet. A 134, 415–421 (2005).
Esmailpour, T. et al. A splice donor mutation in NAA10 results in the dysregulation of the retinoic acid signalling pathway and causes Lenz microphthalmia syndrome. J. Med. Genet. 51, 185–196 (2014).
Laumonnier, F. et al. Transcription factor SOX3 is involved in X-linked mental retardation with growth hormone deficiency. Am. J. Hum. Genet. 71, 1450–1455 (2002).
Faundes, V. et al. Impaired eIF5A function causes a Mendelian disorder that is partially rescued in model systems by spermidine. Nat. Commun. 12, 833 (2021).
Hatada, I. et al. An imprinted gene p57 KIP2 is mutated in Beckwith–Wiedemann syndrome. Nat. Genet. 14, 171–173 (1996).
Cacciagli, P. et al. Mutations in BCAP31 cause a severe X-linked phenotype with deafness, dystonia, and central hypomyelination and disorganize the Golgi apparatus. Am. J. Hum. Genet. 93, 579–586 (2013).
Fantes, J. et al. Mutations in SOX2 cause anophthalmia. Nat. Genet. 33, 462–463 (2003).
Nichols, K. E. et al. Inactivating mutations in an SH2 domain-encoding gene in X-linked lymphoproliferative syndrome. Proc. Natl Acad. Sci. USA 95, 13765–13770 (1998).
Garg, V. et al. GATA4 mutations cause human congenital heart defects and reveal an interaction with TBX5. Nature 424, 443–447 (2003).
Bione, S. et al. A novel X-linked gene, G4. 5. is responsible for Barth syndrome. Nat. Genet. 12, 385–389 (1996).
Amberger, J. S., Bocchini, C. A., Schiettecatte, F., Scott, A. F. & Hamosh, A. OMIM.org: Online Mendelian Inheritance in Man (OMIM®), an online catalog of human genes and genetic disorders. Nucleic Acids Res. 43, D789–D798 (2015).
Schiffels, S. & Durbin, R. Inferring human population size and separation history from multiple genome sequences. Nat. Genet. 46, 919–925 (2014).
Cummings, B. B. et al. Transcript expression-aware annotation improves rare variant interpretation. Nature 581, 452–458 (2020).
McLaren, W. et al. The Ensembl Variant Effect Predictor. Genome Biol. 17, 122 (2016).
Frankish, A. et al. GENCODE: reference annotation for the human and mouse genomes in 2023. Nucleic Acids Res. 51, D942–D949 (2023).
Olson, N. D. et al. PrecisionFDA Truth Challenge V2: calling variants from short and long reads in difficult-to-map regions. Cell Genom. 2, 100129 (2022).
Blake, J. A. et al. Mouse Genome Database (MGD): knowledgebase for mouse–human comparative biology. Nucleic Acids Res. 49, D981–D987 (2021).
Groza, T. et al. The International Mouse Phenotyping Consortium: comprehensive knockout phenotyping underpinning the study of human disease. Nucleic Acids Res. 51, D1038–D1045 (2023).
Gudmundsson, S. et al. Variant interpretation using population databases: lessons from gnomAD. Hum. Mutat. 43, 1012–1030 (2022).
Hart, T., Brown, K. R., Sircoulomb, F., Rottapel, R. & Moffat, J. Measuring error rates in genomic perturbation screens: gold standards for human functional genomics. Mol. Syst. Biol. 10, 733 (2014).
Blomen, V. A. et al. Gene essentiality and synthetic lethality in haploid human cells. Science 350, 1092–1096 (2015).
Samocha, K. E. et al. A framework for the interpretation of de novo mutation in human disease. Nat. Genet. 46, 944–950 (2014).
Finucane, H. K. et al. Partitioning heritability by functional annotation using genome-wide association summary statistics. Nat. Genet. 47, 1228–1235 (2015).
Zeng, T., Spence, J. P., Mostafavi, H. & Pritchard, J. K. s_het estimates from GeneBayes and other supplementary datasets. Zenodo https://doi.org/10.5281/zenodo.10403680 (2023).
Zeng, T. tkzeng/GeneBayes: GeneBayes v1.0. Zenodo https://doi.org/10.5281/zenodo.10939506 (2024).
Zeng, T. Code and data to reproduce GeneBayes figures. Zenodo https://doi.org/10.5281/zenodo.11141460 (2024).
Schuler, A. et al. tkzeng/ngboost: NGBoost for GeneBayes v1.0. Zenodo https://doi.org/10.5281/zenodo.10944711 (2024).
Chen, T. & Guestrin, C. Xgboost: a scalable tree boosting system. In Proc. 22nd ACM SIGKDD International Conference on Knowledge Discovery and Data Mining 785–794 (Association for Computing Machinery, 2016).
Paszke, A. et al. Pytorch: an imperative style, high-performance deep learning library. In Proc. Advances in Neural Information Processing Systems (eds Wallach, H. M., Larochelle, H., Beygelzimer, A., d'Alché-Buc, F. & Fox, E. B.) 32 (Curran Associates Inc., 2019).
Finucane, H. K. et al. Heritability enrichment of specifically expressed genes identifies disease-relevant tissues and cell types. Nat. Genet. 50, 621–629 (2018).
Harris, C. R. et al. Array programming with NumPy. Nature 585, 357–362 (2020).
Virtanen, P. et al. SciPy 1.0: fundamental algorithms for scientific computing in Python. Nat. Methods 17, 261–272 (2020).
Van der Walt, S. & Millman, J. (eds). Data structures for statistical computing in Python. In Proc. 9th Python in Science Conference 56–61 (SciPy, 2010).
Pedregosa, F. et al. Scikit-learn: machine learning in Python. J. Mach. Learn. Res. 12, 2825–2830 (2011).
Van der Walt, S. & Millman, J. (eds). Statsmodels: econometric and statistical modeling with Python. In Proc. 9th Python in Science Conference 92–96 (SciPy, 2010).
Acknowledgements
We would like to thank I. Agarwal, M. Przeworski, J. Engreitz and members of the Pritchard Lab for valuable feedback and discussions. This work was supported by the National Institutes of Health (NIH; grants R01HG011432, R01HG008140 and U01HG009431 to J.K.P. and R01AG066490 to S. Montgomery). The funders had no role in study design, data collection and analysis, decision to publish or preparation of the paper.
Author information
Authors and Affiliations
Contributions
J.P.S., H.M. and J.K.P. conceived and designed the study. T.Z. and J.P.S. performed all data analyses and developed the model. H.M. provided intellectual contributions to all aspects of the study. T.Z., J.P.S., H.M. and J.K.P. wrote the paper. J.K.P. supervised the study and acquired funding.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Genetics thanks Zilin Li and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Peer reviewer reports are available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Performance of shet estimates from a model with some features removed.
a, Scatterplot of posterior mean shet estimated from a model trained without missense constraint or cross-species conservation features (y axis) against shet estimated from the full model (x axis). b, Precision–recall curves comparing the performance of shet estimated from the full model (blue) and from the model without missense/conservation features (orange) in classifying essential genes. c, Precision–recall curves comparing the performance of shet estimated from the two models in classifying developmental disorder genes.
Extended Data Fig. 2 Comparison of shet estimates from models trained on subsets of gnomAD.
a, Scatterplot of posterior mean shet estimated from a model trained with non-NFE individuals (y axis) against shet estimated from the full model (x axis). NFE, Non-Finnish European. This subset consists of 56,000 individuals or 45% of the total dataset. b, Scatterplot of posterior mean shet estimated from a model trained with NFE individuals (y axis) against shet estimated from the full model (x axis). This subset consists of 67,000 individuals or 55% of the total dataset.
Extended Data Fig. 3 shet distributions for additional example genes.
Left: posterior distributions and rescaled likelihoods for genes with few expected LOFs (genes in the bottom quartile). Right: posterior distributions and rescaled likelihoods for genes with many expected LOFs (genes in the top quartile).
Extended Data Fig. 4 Additional validation analyses.
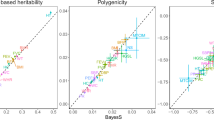
a, Precision–recall curves comparing the performance of shet estimates from GeneBayes against LOEUF from gnomAD v4.0.0 (731k exomes) or LOEUF from gnomAD v2.1.1 (125k exomes) in classifying essential genes. b, Precision–recall curves comparing the performance of shet estimates from GeneBayes against other constraint metrics in classifying nonessential genes. c, Precision–recall curves comparing the performance of shet against other constraint metrics in classifying developmental disorder genes. d, Enrichment of de novo mutations in patients with developmental disorders, calculated as the observed number of mutations over the expected number under a null mutational model (n = 31,058 parent–offspring trios). We plot the enrichment of synonymous, missense, splice and nonsense variants in the 10% of genes considered most constrained by shet (blue) and the enrichment of these variants in all other genes (gray), including (left) and excluding (right) known developmental disorder genes. Bars represent 95% confidence intervals, centered around the mean. e, Scatterplot of the enrichment of common variant heritability in the 10% of genes considered most constrained by shet (y axis) or LOEUF (x axis), normalized by the enrichment of heritability in all genes. Each point represents one trait.
Extended Data Fig. 5 Performance of shet and LOEUF for genes with differing numbers of expected LOFs.
Left: precision–recall curves comparing the performance of shet against LOEUF in classifying essential genes for groups of genes binned by their expected number of LOFs. Right: precision–recall curves comparing the performance of shet against LOEUF in classifying developmental disorder genes for binned genes.
Extended Data Fig. 6 Correlation of gene features with gene length.
a, Histogram of the Spearman ρ between gene features and coding sequence (CDS) length. b, Histogram of the Spearman ρ between gene features and CDS length for gene expression features, colored by category. c, Spearman ρ between gene features and CDS length for additional features of interest. d, Scatterplot of the Spearman ρ between gene features and posterior mean shet (y axis) against the partial Spearman ρ (x axis) after controlling for the effect of gene (CDS) length.
Supplementary information
Supplementary Information
Supplementary Note and Figs. 1–4.
Supplementary Tables
Supplementary Table 1: Posterior means and 95% credible intervals for GeneBayes estimates of shet. Supplementary Table 2: LOEUF and shet for ribosomal proteins associated with Diamond–Blackfan anemia. Supplementary Table 3: Terms used to define tissues for expression features. Supplementary Table 4: Filtering criteria for LOF variant curation. Supplementary Table 5: Sources for the LOF data. Supplementary Table 6: Parameters for fitting the gradient-boosted trees. Supplementary Table 7: Parameters for fitting the gradient-boosted trees for models trained on feature subsets. Supplementary Table 8a–k: Gene feature descriptions.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Zeng, T., Spence, J.P., Mostafavi, H. et al. Bayesian estimation of gene constraint from an evolutionary model with gene features. Nat Genet 56, 1632–1643 (2024). https://doi.org/10.1038/s41588-024-01820-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41588-024-01820-9
- Springer Nature America, Inc.
This article is cited by
-
Improving estimates of loss-of-function constraint for short genes
Nature Genetics (2024)