Abstract
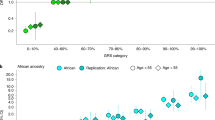
The transferability and clinical value of genetic risk scores (GRSs) across populations remain limited due to an imbalance in genetic studies across ancestrally diverse populations. Here we conducted a multi-ancestry genome-wide association study of 156,319 prostate cancer cases and 788,443 controls of European, African, Asian and Hispanic men, reflecting a 57% increase in the number of non-European cases over previous prostate cancer genome-wide association studies. We identified 187 novel risk variants for prostate cancer, increasing the total number of risk variants to 451. An externally replicated multi-ancestry GRS was associated with risk that ranged from 1.8 (per standard deviation) in African ancestry men to 2.2 in European ancestry men. The GRS was associated with a greater risk of aggressive versus non-aggressive disease in men of African ancestry (P = 0.03). Our study presents novel prostate cancer susceptibility loci and a GRS with effective risk stratification across ancestry groups.





Similar content being viewed by others
Data availability
The full summary statistics resulting from this investigation are available in the GWAS Catalog (https://www.ebi.ac.uk/gwas/) under accession codes as follows: cross-ancestry (GCST90274713), European (GCST90274714), African (GCST90274715), Asian (GCST90274716) and Hispanic (GCST90274717). Genotype and covariate data used in this study are deposited in dbGaP under accession codes phs001391.v1.p1, phs000306.v4.p1, phs001120.v2.p2phs001221.v1.p1, phs000812.v1.p1 and phs000838.v1.p1. The variants and weights for the GRS269 and GRS451 are available on the PGS Catalog under accession codes PGP000122 and PGP000488, respectively (https://www.pgscatalog.org/). Publicly available data described in this manuscript can be found from the following websites: 1000 Genomes Project (http://ftp.1000genomes.ebi.ac.uk/vol1/ftp/phase3/); Human Genome Diversity Project (https://www.internationalgenome.org/data-portal/data-collection/hgdp); SEER (https://seer.cancer.gov/); National Center for Health Statistics, CDC (https://www.cdc.gov/nchs/index.htm); Cistrome Data Browser (http://cistrome.org/db/); RefZ (https://www.ncbi.nlm.nih.gov/projects/gap/cgi-bin/study.cgi?study_id=phs000985.v1.p1); GTEx (https://gtexportal.org/home/datasets); TCGA (https://portal.gdc.cancer.gov); CancerSplicingQTL database (http://www.cancersplicingqtl-hust.com/); and EnTEx/ENCODE (http://entex.encodeproject.org/).
Code availability
Imputation was performed using IMPUTE2, MACH 1.0, Beagle 4.1, Beagle 5.1, EAGLE v2.4, Minimac3 and Minimac4. Association testing was performed using PLINK 1.07 and 2.0, SNPtest v2.5.2, SAIGE v.0.20 and R v3.6.3. Meta-analyses were conducted using METAL v2011-03-25 and fine-mapping with mJAM (https://github.com/USCbiostats/hJAM/). Genome-wide PRS was derived from PRS-CSx v1.0.0 (https://github.com/getian107/PRScsx). Variant annotation was performed with wANNOVAR (https://wannovar.wglab.org/, accessed 20 May 2022) and R package rtracklayer v1.42.2. TWAS was performed with FUSION (https://github.com/gusevlab/fusion_twas, accessed 20 May 2022; TWAS weights: GTEx v8 and TCGA: http://gusevlab.org/projects/fusion/, RefZ: https://www.mancusolab.com/prostate-twas/, INTERVAL: https://www.mancusolab.com/pwas/) and GCTA v1.94.0beta. Data visualization was performed using ggplot2 v3.4.2 and gwasforest v1.0.0 packages in R software (v3.6.3).
References
Bray, F. et al. Global cancer statistics 2018: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J. Clin. 68, 394–424 (2018).
Conti, D. V. et al. Trans-ancestry genome-wide association meta-analysis of prostate cancer identifies new susceptibility loci and informs genetic risk prediction. Nat. Genet. 53, 65–75 (2021).
Schumacher, F. R. et al. Association analyses of more than 140,000 men identify 63 new prostate cancer susceptibility loci. Nat. Genet. 50, 928–936 (2018).
Dadaev, T. et al. Fine-mapping of prostate cancer susceptibility loci in a large meta-analysis identifies candidate causal variants. Nat. Commun. 9, 2256 (2018).
Conti, D. V. et al. Two novel susceptibility loci for prostate cancer in men of African ancestry. J. Natl Cancer Inst. 109, djx084 (2017).
Wang, M. et al. Large-scale association analysis in Asians identifies new susceptibility loci for prostate cancer. Nat. Commun. 6, 8469 (2015).
Hoffmann, T. J. et al. A large multiethnic genome-wide association study of prostate cancer identifies novel risk variants and substantial ethnic differences. Cancer Discov. 5, 878–891 (2015).
Al Olama, A. A. et al. A meta-analysis of 87,040 individuals identifies 23 new susceptibility loci for prostate cancer. Nat. Genet. 46, 1103–1109 (2014).
Eeles, R. A. et al. Identification of 23 new prostate cancer susceptibility loci using the iCOGS custom genotyping array. Nat. Genet. 45, 391e1–392e1 (2013). 385-91.
Gudmundsson, J. et al. A study based on whole-genome sequencing yields a rare variant at 8q24 associated with prostate cancer. Nat. Genet. 44, 1326–1329 (2012).
Takata, R. et al. Genome-wide association study identifies five new susceptibility loci for prostate cancer in the Japanese population. Nat. Genet. 42, 751–754 (2010).
Gudmundsson, J. et al. Common sequence variants on 2p15 and Xp11.22 confer susceptibility to prostate cancer. Nat. Genet. 40, 281–283 (2008).
Amundadottir, L. T. et al. A common variant associated with prostate cancer in European and African populations. Nat. Genet. 38, 652–658 (2006).
Wojcik, G. L. et al. Genetic analyses of diverse populations improves discovery for complex traits. Nature 570, 514–518 (2019).
Newcombe, P. J., Conti, D. V. & Richardson, S. JAM: a scalable Bayesian framework for joint analysis of marginal SNP effects. Genet. Epidemiol. 40, 188–201 (2016).
Chen, F. et al. Evidence of novel susceptibility variants for prostate cancer and a multiancestry polygenic risk score associated with aggressive disease in men of African ancestry. Eur. Urol. 84, 13–21 (2023).
Barfeld, S. J., East, P., Zuber, V. & Mills, I. G. Meta-analysis of prostate cancer gene expression data identifies a novel discriminatory signature enriched for glycosylating enzymes. BMC Med Genomics 7, 513 (2014).
Halvorsen, O. J. et al. Increased expression of SIM2-s protein is a novel marker of aggressive prostate cancer. Clin. Cancer Res. 13, 892–897 (2007).
Dhanasekaran, S. M. et al. Delineation of prognostic biomarkers in prostate cancer. Nature 412, 822–826 (2001).
Linn, D. E. et al. Differential regulation of androgen receptor by PIM-1 kinases via phosphorylation-dependent recruitment of distinct ubiquitin E3 ligases. J. Biol. Chem. 287, 22959–22968 (2012).
Wang, J. et al. Pim1 kinase synergizes with c-MYC to induce advanced prostate carcinoma. Oncogene 29, 2477–2487 (2010).
Sun, X. et al. Frequent somatic mutations of the transcription factor ATBF1 in human prostate cancer. Nat. Genet. 37, 407–412 (2005).
Chandler, J. D., Williams, E. D., Slavin, J. L., Best, J. D. & Rogers, S. Expression and localization of GLUT1 and GLUT12 in prostate carcinoma. Cancer 97, 2035–2042 (2003).
White, M. A. et al. GLUT12 promotes prostate cancer cell growth and is regulated by androgens and CaMKK2 signaling. Endocr. Relat. Cancer 25, 453–469 (2018).
Mi, Y. et al. Down-regulation of Barx2 predicts poor survival in colorectal cancer. Biochem. Biophys. Res. Commun. 478, 67–73 (2016).
Zhang, J. et al. Plasma proteome analyses in individuals of European and African ancestry identify cis-pQTLs and models for proteome-wide association studies. Nat. Genet. 54, 593–602 (2022).
Gusev, A. et al. Integrative approaches for large-scale transcriptome-wide association studies. Nat. Genet. 48, 245–252 (2016).
Gamazon, E. R. et al. A gene-based association method for mapping traits using reference transcriptome data. Nat. Genet. 47, 1091–1098 (2015).
The GTEx Consortium. The GTEx Consortium atlas of genetic regulatory effects across human tissues. Science 369, 1318–1330 (2020).
Thibodeau, S. N. et al. Identification of candidate genes for prostate cancer-risk SNPs utilizing a normal prostate tissue eQTL data set. Nat. Commun. 6, 8653 (2015).
The Cancer Genome Atlas Research Network. The molecular taxonomy of primary prostate cancer. Cell 163, 1011–1025 (2015).
Sun, B. B. et al. Genomic atlas of the human plasma proteome. Nature 558, 73–79 (2018).
Mancuso, N. et al. Large-scale transcriptome-wide association study identifies new prostate cancer risk regions. Nat. Commun. 9, 4079 (2018).
Liu, D. et al. A transcriptome-wide association study identifies novel candidate susceptibility genes for prostate cancer risk. Int J. Cancer 150, 80–90 (2022).
Amin Al Olama, A. et al. Multiple novel prostate cancer susceptibility signals identified by fine-mapping of known risk loci among Europeans. Hum. Mol. Genet. 24, 5589–5602 (2015).
Pencina, M. J., D’Agostino, R. B. Sr. & Steyerberg, E. W. Extensions of net reclassification improvement calculations to measure usefulness of new biomarkers. Stat. Med 30, 11–21 (2011).
Chen, F. et al. Validation of a multi-ancestry polygenic risk score and age-specific risks of prostate cancer: a meta-analysis within diverse populations. eLife 11, e78304 (2022).
Darst, B. F. et al. Evaluating approaches for constructing polygenic risk scores for prostate cancer in men of African and European ancestry. Am. J. Hum. Genet 110, 1200–1206 (2023).
Das, S. et al. Next-generation genotype imputation service and methods. Nat. Genet. 48, 1284–1287 (2016).
Howie, B. N., Donnelly, P. & Marchini, J. A flexible and accurate genotype imputation method for the next generation of genome-wide association studies. PLoS Genet. 5, e1000529 (2009).
Loh, P.-R. et al. Reference-based phasing using the Haplotype Reference Consortium panel. Nat. Genet. 48, 1443–1448 (2016).
Kurki, M. I. et al. FinnGen provides genetic insights from a well-phenotyped isolated population. Nature 613, 508–518 (2023).
Auton, A. et al. A global reference for human genetic variation. Nature 526, 68–74 (2015).
Taliun, D. et al. Sequencing of 53,831 diverse genomes from the NHLBI TOPMed Program. Nature 590, 290–299 (2021).
Bycroft, C. et al. The UK Biobank resource with deep phenotyping and genomic data. Nature 562, 203–209 (2018).
de Bakker, P. I. et al. Practical aspects of imputation-driven meta-analysis of genome-wide association studies. Hum. Mol. Genet. 17, R122–R128 (2008).
Zawistowski, M. et al. The Michigan Genomics Initiative: a biobank linking genotypes and electronic clinical records in Michigan Medicine patients. Cell Genom. 3, 100257 (2023).
Karlson, E. W., Boutin, N. T., Hoffnagle, A. G. & Allen, N. L. Building the Partners HealthCare Biobank at Partners Personalized Medicine: informed consent, return of research results, recruitment lessons and operational considerations. J. Pers. Med. 6, 2 (2016).
Plym, A. et al. Evaluation of a multiethnic polygenic risk score model for prostate cancer. J. Natl Cancer Inst. 114, 771–774 (2022).
Andrews, C. et al. Development, evaluation, and implementation of a pan-African cancer research network: men of African descent and carcinoma of the prostate. J. Glob. Oncol. 4, 1–14 (2018).
Leitsalu, L. et al. Cohort Profile: Estonian Biobank of the Estonian Genome Center, University of Tartu. Int. J. Epidemiol. 44, 1137–1147 (2015).
Ruan, Y. et al. Improving polygenic prediction in ancestrally diverse populations. Nat. Genet. 54, 573–580 (2022).
Ge, T., Chen, C. Y., Ni, Y., Feng, Y. A. & Smoller, J. W. Polygenic prediction via Bayesian regression and continuous shrinkage priors. Nat. Commun. 10, 1776 (2019).
The International HapMap 3 Consortium Integrating common and rare genetic variation in diverse human populations. Nature 467, 52–58 (2010).
1000 Genomes Project Consortiumet al. A global reference for human genetic variation. Nature 526, 68–74 (2015).
Kachuri, L. et al. Genetically adjusted PSA levels for prostate cancer screening. Nat. Med. 29, 1412–1423 (2023).
Amin Al Olama, A. et al. Risk analysis of prostate cancer in PRACTICAL, a multinational consortium, using 25 known prostate cancer susceptibility loci. Cancer Epidemiol. Biomark. Prev. 24, 1121–1129 (2015).
Antoniou, A. C. et al. Common breast cancer susceptibility alleles and the risk of breast cancer for BRCA1 and BRCA2 mutation carriers: implications for risk prediction. Cancer Res. 70, 9742–9754 (2010).
Antoniou, A. C. et al. Evidence for further breast cancer susceptibility genes in addition to BRCA1 and BRCA2 in a population-based study. Genet. Epidemiol. 21, 1–18 (2001).
Kuchenbaecker, K. B. et al. Evaluation of polygenic risk scores for breast and ovarian cancer risk prediction in BRCA1 and BRCA2 mutation carriers. J. Natl Cancer Inst. 109, djw302 (2017).
Surveillance, Epidemiology, and End Results (SEER) Program. SEER*Stat Database: Incidence - SEER Research Plus Data, 18 Registries, Nov 2020 Sub (2000–2018) - Linked To County Attributes - Total U.S., 1969–2019 Counties National Cancer Institute https://seer.cancer.gov/statistics-network/explorer/ (2021).
Surveillance, Epidemiology, and End Results (SEER) Program. SEER*Stat Database: Mortality - All COD, Aggregated With State, Total U.S. (1990–2019) <Katrina/Rita Population Adjustment> National Cancer Institute https://seer.cancer.gov/statistics-network/explorer/ (2021).
Chang, X. & Wang, K. wANNOVAR: annotating genetic variants for personal genomes via the web. J. Med. Genet. 49, 433–436 (2012).
Mei, S. et al. Cistrome Data Browser: a data portal for ChIP–seq and chromatin accessibility data in human and mouse. Nucleic Acids Res. 45, D658–d662 (2017).
Lawrence, M., Gentleman, R. & Carey, V. rtracklayer: an R package for interfacing with genome browsers. Bioinformatics 25, 1841–1842 (2009).
Gong, J. et al. PancanQTL: systematic identification of cis-eQTLs and trans-eQTLs in 33 cancer types. Nucleic Acids Res. 46, D971–D976 (2018).
Tian, J. et al. CancerSplicingQTL: a database for genome-wide identification of splicing QTLs in human cancer. Nucleic Acids Res. 47, D909–D916 (2018).
Bergstrom, A. et al. Insights into human genetic variation and population history from 929 diverse genomes. Science 367, eaay5012 (2020).
Rozowsky, J. et al. The EN-TEx resource of multi-tissue personal epigenomes & variant-impact models. Cell 186, 1493–1511 e40 (2023).
Stegle, O., Parts, L., Durbin, R. & Winn, J. A Bayesian framework to account for complex non-genetic factors in gene expression levels greatly increases power in eQTL studies. PLoS Comput. Biol. 6, e1000770 (2010).
Yang, J., Lee, S. H., Goddard, M. E. & Visscher, P. M. GCTA: a tool for genome-wide complex trait analysis. Am. J. Hum. Genet 88, 76–82 (2011).
Wang, G., Sarkar, A., Carbonetto, P. & Stephens, M. A simple new approach to variable selection in regression, with application to genetic fine mapping. J. R. Stat. Soc. Ser. B 82, 1273–1300 (2020).
Acknowledgements
This project was supported by the US National Institutes of Health (NIH) grants R01CA257328 (C.A.H.), U19CA214253 (C.A.H.), U01CA261339 (D.V.C.), P01CA196569 (D.V.C.) and R00CA246063 (B.F. Darst), and the Prostate Cancer Foundation grants 20CHAS03 (CAH) and 21YOUN11 (B.F. Darst). We acknowledge support from The National Institute of Health Research to The Biomedical Research Centre at The Royal Marsden NHS Foundation Trust and The Institute of Cancer Research, the DJ Fielding Medical Trust and the Joseph Fraser Trust via The Royal Marsden Cancer Charity. The funders had no role in study design, data collection and analysis, decision to publish or preparation of the manuscript. This research has been conducted using the UK Biobank Resource under application number 42195. This research is based on data from the Million Veteran Program, Office of Research and Development, and the Veterans Health Administration. This publication does not represent the views of the Department of Veteran Affairs or the US Government. A full description of funding and acknowledgements for each of the contributing studies can be found in the Supplementary Note.
Author information
Authors and Affiliations
Consortia
Contributions
C.A.H., D.V.C., R.A.E. and Z.K.-J. contributed to study conception. A. Wang, C.A.H., D.V.C., E.J.S. and N.M. wrote the manuscript. E.J.S., Y.X., X.S., P.W., M.B., A.A.R., R.K.M. and T.D. provided data management and bioinformatics support. A. Wang, J. Shen, A.A.R., D.V.C. and C.A.H. contributed to data analysis and interpretation. All authors contributed data to the study, revised, critically reviewed and approved the final version of the manuscript: A. Wang, J. Shen, A.A.R., E.J.S., F. Chen, R. Janivara, B.F. Darst, X.S., Y.X., A.J.C., S.B., T.D., M.N.B., A.P., A.S., T.J.H., A. Takahashi, K. Matsuda, Y. Momozawa, M.F., T.L., J.F., K. Muir, S.I., X.L., Y. Yamanashi, Y.F., T.M., Y. Murakami, K. Muto, A.N., W.O., K. Yamaji, K.T., S. Asai, Y. Takahashi, T. Suzuki, N.S., H.Y., S. Minami, S. Murayama, K. Yoshimori, S. Nagayama, D.O., M.H., A. Masumoto, Y. Koretsune, Y.U., M. Kubo, Y. Kamatani, A. Lophatananon, P.W., C.A., A. Lori, P.P.C., J. Schleutker, T.L.J.T., C. Sipeky, A. Auvinen, G.G.G., M.C. Southey, R.J.M., C.C., D.W., J. Lubinski, C.T.R., K.C., B.H.M., D.E.N., J.L.D., F.C.H., R.M. Martin, B.G.N., S.F.N., M.W., S.E.B., M.A.R., H.V.S., J.B., S.C., L.H., J.A.C., W. Tilly, G.P.R., H.G., M.A., R.S., M.E., T.N., N.P., A.M.D., M. Ghoussaini, R.C.T., T.J.K., E.R., J.Y.P., T.A.S., H.-Y.L., D.A., S. Weinstein, M.B.C., L.A.M., E.G., S. Lindstrom, P.K., D.J.H., K.L.P., C. Turman, C.M.T., P.J.G., I.M.T.J., R.J.H., N.E.F., A.F., M.-É.P., J.L.S., E.A.O., S.K., L.E.B.F., M. Stampfer, A. Wolk, N.H., G.L.A., R.N.H., M.J.M., K.D.S., M.B., W.J.B., W.Z., E.D.Y., J.E.M., Y.-J.L., H.-W.Z., N.F., X.M., Y. Wu, S.-C.Z., Z.S., S.N.T., S.K.M.D., D.J.S., C.M.L.W., G.B., C.M., T. Schnoeller, M.L., A.S.K., B.F. Drake, O.C., G.C.-T., F.M., T.T., Y.A.K., E.M.J., E.M.G., L. Maehle, K.-T.K., S.A.I., M.C. Stern, A.V., A.G.-C., L. Fachal, B.S.R., S.L.K., H.O., M.R.T., P. Paulo, A.B., S. Watya, A. Lubwama, J.T.B., E.N.B., J.L.M., J.A.T., M. Kogevinas, T.D.-S., G.C.-V., L.C.-A., C.C.T., C.D.H., P. Pilie, Y. Yu, R.J.B., J.G., S.S.S.,. L. Multigner, P.B., L.B., R.K., C. Slavov, V.M., R.J.L., H.B., X.C., B.H., B.S., E.A.K., A.W.H., R.A.K., A.B.M., C.J.L., J.K., S.L.N., L.S., Y.C.D., W.B.I., B.N., A.J.M.H., J. Carpten, H.P., A. Michael, K.D.R., G.D.M., P.O., J.X., A.R., J. Lim, S.-H.T., L.F.N., D.W.L., J.H.F., C.M.N.-D., B.A.R., M. Gamulin, D.L., T.K., N.U., A. Abraham, S. Singhal, M.P., F. Claessens, S.J., T.V.B., M.G.-D., J.E.C., M.E.M., S. Larkin, P.A.T., C.A.-H., W.S.B., M.C.A., D.C.C., S. Srivastava, J. Cullen, G.P., G. Casey, Y. Wang, Y. Tettey, J. Lachance, W. Tang, R.B.B., A.A.A., E. Tay, A. Truelove, S. Nielsen, K. Yamoah, K.G., A.P.C., J.M.K., J.N.H., P.E.C., M.J., S.M. Gueye, L.N., O.O., O.S., O.A., A.O.A., O.I.A.-S., H.O.A., M.A.J., O.P.O., M.N., B.A., S. Mante, A.D.-A., H.D., S.M. Gundell, M.J.R., G.J., R.H.N.S., J.J.H., M. Sanderson, L.K., R.V., R.M.K.-C., M.T., M.H.P., R.J.F.L., M.Z., S.Z., Z.L., S.K.V.D.E., D.F.E., S. Ambs, T.L.E., R.M., T.R.R., L. Fritsche, S.J.C., S.I.B., F.W., H.N., J.S.W., J.M.G., A.C.J., N.M., C. Terao, R.A.E., Z.K.-J., R.K.M., D.V.C. and C.A.H. C.A.H. and R.K.M. had full access to the data in the study and take responsibility for the integrity of the data and the accuracy of the data analysis.
Corresponding author
Ethics declarations
Competing interests
R.A.E. declares the following conflicts of interest: Honoraria from GU-ASCO, Janssen, University of Chicago, Dana Farber Cancer Institute USA as a speaker. Educational honorarium from Bayer and Ipsen, member of external expert committee to Astra Zeneca UK. She undertakes private practice as a sole trader at The Royal Marsden NHS Foundation Trust, London, UK. The other authors declare no competing interests.
Peer review
Peer review information
Nature Genetics thanks Mark Rubin and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Peer reviewer reports are available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Venn diagram of prostate cancer risk variants common (MAF > 1%) among European, African, Asian and Hispanic populations.
The plot illustrates the distribution of 451 prostate cancer risk variants, highlighting the number of variants that are either unique to or shared among European, African, Asian and Hispanic populations. Five variants with a minor allele frequency (MAF) of ≤1% across all populations are included under the European population, where they are most frequent. Numbers in parentheses denote the total count of variants common in each respective population.
Extended Data Fig. 2 The associations of GRS451 with total prostate cancer risk in GWAS discovery and replication sub-studies and meta-analyses within and across ancestry groups.
Odds ratios and 95% confidence intervals for one SD increase in GRS451 and total prostate cancer risk were calculated from logistic regression. The columns ‘Case’ and ‘Control’ show the case and control sample sizes, respectively. ‘META’ refers to the meta-analyzed results using the inverse-variance weighted method. The y-axis shows each sub-study (details of each sub-study are available in Supplementary Tables 1 and 2) and their corresponding meta-analyzed results by ancestry and study phase (GWAS discovery or replication), as well as the overall meta-analyzed results.
Supplementary information
Supplementary Information
Supplementary Figs. 1–7 and additional acknowledgements.
Supplementary Tables 1–19
Supplementary Tables 1–19.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Wang, A., Shen, J., Rodriguez, A.A. et al. Characterizing prostate cancer risk through multi-ancestry genome-wide discovery of 187 novel risk variants. Nat Genet 55, 2065–2074 (2023). https://doi.org/10.1038/s41588-023-01534-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41588-023-01534-4
- Springer Nature America, Inc.
This article is cited by
-
Using gene and gene-set association tests to identify lethal prostate cancer genes
Prostate Cancer and Prostatic Diseases (2024)
-
The broad impact of cell death genes on the human disease phenome
Cell Death & Disease (2024)