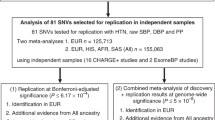
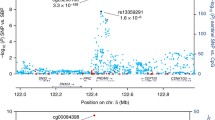
Abstract
Genetic studies of blood pressure (BP) to date have mainly analyzed common variants (minor allele frequency > 0.05). In a meta-analysis of up to ~1.3 million participants, we discovered 106 new BP-associated genomic regions and 87 rare (minor allele frequency ≤ 0.01) variant BP associations (P < 5 × 10−8), of which 32 were in new BP-associated loci and 55 were independent BP-associated single-nucleotide variants within known BP-associated regions. Average effects of rare variants (44% coding) were ~8 times larger than common variant effects and indicate potential candidate causal genes at new and known loci (for example, GATA5 and PLCB3). BP-associated variants (including rare and common) were enriched in regions of active chromatin in fetal tissues, potentially linking fetal development with BP regulation in later life. Multivariable Mendelian randomization suggested possible inverse effects of elevated systolic and diastolic BP on large artery stroke. Our study demonstrates the utility of rare-variant analyses for identifying candidate genes and the results highlight potential therapeutic targets.





Similar content being viewed by others
Data availability
Summary association results for all the traits are available to download from https://app.box.com/s/1ev9iakptips70k8t4cm8j347if0ef2u and from CHARGE dbGaP Summary (https://www.ncbi.nlm.nih.gov/gap/) under accession number phs000930.
Change history
16 March 2021
A Correction to this paper has been published: https://doi.org/10.1038/s41588-021-00832-z
References
Forouzanfar, M. H. et al. Global burden of hypertension and systolic blood pressure of at least 110 to 115 mm Hg, 1990–2015. JAMA 317, 165–182 (2017).
Newton-Cheh, C. et al. Genome-wide association study identifies eight loci associated with blood pressure. Nat. Genet. 41, 666–676 (2009).
Cho, Y. S. et al. A large-scale genome-wide association study of Asian populations uncovers genetic factors influencing eight quantitative traits. Nat. Genet. 41, 527–534 (2009).
Levy, D. et al. Genome-wide association study of blood pressure and hypertension. Nat. Genet. 41, 677–687 (2009).
Kato, N. et al. Meta-analysis of genome-wide association studies identifies common variants associated with blood pressure variation in east Asians. Nat. Genet. 43, 531–538 (2011).
Wain, L. V. et al. Genome-wide association study identifies six new loci influencing pulse pressure and mean arterial pressure. Nat. Genet. 43, 1005–1011 (2011).
International Consortium for Blood Pressure Genome-Wide Association Studies. Genetic variants in novel pathways influence blood pressure and cardiovascular disease risk. Nature 478, 103–109 (2011).
Johnson, A. D. et al. Association of hypertension drug target genes with blood pressure and hypertension in 86,588 individuals. Hypertension 57, 903–910 (2011).
Johnson, T. et al. Blood pressure loci identified with a gene-centric array. Am. J. Hum. Genet. 89, 688–700 (2011).
Tragante, V. et al. Gene-centric meta-analysis in 87,736 individuals of European ancestry identifies multiple blood-pressure-related loci. Am. J. Hum. Genet. 94, 349–360 (2014).
Simino, J. et al. Gene-age interactions in blood pressure regulation: a large-scale investigation with the CHARGE, Global BPgen and ICBP Consortia. Am. J. Hum. Genet. 95, 24–38 (2014).
Kato, N. et al. Trans-ancestry genome-wide association study identifies 12 genetic loci influencing blood pressure and implicates a role for DNA methylation. Nat. Genet. 47, 1282–1293 (2015).
Surendran, P. et al. Trans-ancestry meta-analyses identify rare and common variants associated with blood pressure and hypertension. Nat. Genet. 48, 1151–1161 (2016).
Liu, C. et al. Meta-analysis identifies common and rare variants influencing blood pressure and overlapping with metabolic trait loci. Nat. Genet. 48, 1162–1170 (2016).
Ehret, G. B. et al. The genetics of blood pressure regulation and its target organs from association studies in 342,415 individuals. Nat. Genet. 48, 1171–1184 (2016).
Hoffmann, T. J. et al. Genome-wide association analyses using electronic health records identify new loci influencing blood pressure variation. Nat. Genet. 49, 54–64 (2017).
Warren, H. R. et al. Genome-wide association analysis identifies novel blood pressure loci and offers biological insights into cardiovascular risk. Nat. Genet. 49, 403–415 (2017).
Kraja, A. T. et al. New blood pressure-associated loci identified in meta-analyses of 475,000 individuals. Circ. Cardiovasc. Genet. 10, e001778 (2017).
Wain, L. V. et al. Novel blood pressure locus and gene discovery using genome-wide association study and expression datasets from blood and the kidney. Hypertension 70, e4–e19 (2017).
Evangelou, E. et al. Genetic analysis of over 1 million people identifies 535 new loci associated with blood pressure traits. Nat. Genet. 50, 1412–1425 (2018).
Giri, A. et al. Trans-ethnic association study of blood pressure determinants in over 750,000 individuals. Nat. Genet. 51, 51–62 (2019).
Yang, J., Lee, S. H., Goddard, M. E. & Visscher, P. M. GCTA: a tool for genome-wide complex trait analysis. Am. J. Hum. Genet. 88, 76–−82 (2011).
Bycroft, C. et al. The UK Biobank resource with deep phenotyping and genomic data. Nature 562, 203–209 (2018).
Loh, P. R. et al. Efficient Bayesian mixed-model analysis increases association power in large cohorts. Nat. Genet. 47, 284–290 (2015).
Benner, C. et al. FINEMAP: efficient variable selection using summary data from genome-wide association studies. Bioinformatics 32, 1493–1501 (2016).
Wu, M. C. et al. Rare-variant association testing for sequencing data with the sequence kernel association test. Am. J. Hum. Genet. 89, 82–93 (2011).
McLaren, W. et al. The Ensembl Variant Effect Predictor. Genome Biol. 17, 122 (2016).
Marouli, E. et al. Rare and low-frequency coding variants alter human adult height. Nature 542, 186–190 (2017).
Liu, D. J. et al. Exome-wide association study of plasma lipids in >300,000 individuals. Nat. Genet. 49, 1758–1766 (2017).
Turcot, V. et al. Protein-altering variants associated with body mass index implicate pathways that control energy intake and expenditure in obesity. Nat. Genet. 50, 26–41 (2018).
Finan, C. et al. The druggable genome and support for target identification and validation in drug development. Sci. Transl. Med. 9, eaag1166 (2017).
Foley, C. N. et al. A fast and efficient colocalization algorithm for identifying shared genetic risk factors across multiple traits. Preprint at bioRxiv https://doi.org/10.1101/592238 (2019).
Solovieff, N., Cotsapas, C., Lee, P. H., Purcell, S. M. & Smoller, J. W. Pleiotropy in complex traits: challenges and strategies. Nat. Rev. Genet. 14, 483–495 (2013).
Xu, X. et al. Molecular insights into genome-wide association studies of chronic kidney disease-defining traits. Nat. Commun. 9, 4800 (2018).
Rowland, J. et al. Uncovering genetic mechanisms of kidney aging through transcriptomics, genomics, and epigenomics. Kidney Int. 95, 624–635 (2019).
Staley, J. R. et al. PhenoScanner: a database of human genotype-phenotype associations. Bioinformatics 32, 3207–3209 (2016).
Vishram, J. K. et al. Impact of age on the importance of systolic and diastolic blood pressures for stroke risk: the MOnica, Risk, Genetics, Archiving and Monograph Project. Hypertension 60, 1117–1123 (2012).
Messaoudi, S. et al. Endothelial Gata5 transcription factor regulates blood pressure. Nat. Commun. 6, 8835 (2015).
Kircher, M. et al. A general framework for estimating the relative pathogenicity of human genetic variants. Nat. Genet. 46, 310–315 (2014).
Wang, Z. et al. Phospholipase C beta3 deficiency leads to macrophage hypersensitivity to apoptotic induction and reduction of atherosclerosis in mice. J. Clin. Invest. 118, 195–204 (2008).
Hoeppner, L. H. et al. Revealing the role of phospholipase Cβ3 in the regulation of VEGF-induced vascular permeability. Blood 120, 2167–2173 (2012).
Li, M. et al. SOS2 and ACP1 loci identified through large-scale exome chip analysis regulate kidney development and function. J. Am. Soc. Nephrol. 28, 981–994 (2017).
Tidyman, W. E. & Rauen, K. A. Pathogenetics of the RASopathies. Hum. Mol. Genet. 25, R123–R132 (2016).
Kim, J. H. et al. Functional role of the polymorphic 647 T/C variant of ENT1 (SLC29A1) and its association with alcohol withdrawal seizures. PLoS ONE 6, e16331 (2011).
Best, K. A., Bone, D. B., Vilas, G., Gros, R. & Hammond, J. R. Changes in aortic reactivity associated with the loss of equilibrative nucleoside transporter 1 (ENT1) in mice. PLoS ONE 13, e0207198 (2018).
Zaidi, S. et al. De novo mutations in histone-modifying genes in congenital heart disease. Nature 498, 220–223 (2013).
Jin, S. C. et al. Contribution of rare inherited and de novo variants in 2,871 congenital heart disease probands. Nat. Genet. 49, 1593–1601 (2017).
Nielsen, J. B. et al. Genome-wide study of atrial fibrillation identifies seven risk loci and highlights biological pathways and regulatory elements involved in cardiac development. Am. J. Hum. Genet. 102, 103–115 (2018).
Zhu, X. et al. Meta-analysis of correlated traits via summary statistics from GWAS with an application in hypertension. Am. J. Hum. Genet. 96, 21–36 (2015).
Newton-Cheh, C. et al. Association of common variants in NPPA and NPPB with circulating natriuretic peptides and blood pressure. Nat. Genet. 41, 348–353 (2009).
Tobin, M. D., Sheehan, N. A., Scurrah, K. J. & Burton, P. R. Adjusting for treatment effects in studies of quantitative traits: antihypertensive therapy and systolic blood pressure. Stat. Med. 24, 2911–2935 (2005).
Mahajan, A. et al. Identification and functional characterization of G6PC2 coding variants influencing glycemic traits define an effector transcript at the G6PC2-ABCB11 locus. PLoS Genet. 11, e1004876 (2015).
Grove, M. L. et al. Best practices and joint calling of the HumanExome BeadChip: the CHARGE Consortium. PLoS ONE 8, e68095 (2013).
Willer, C. J., Li, Y. & Abecasis, G. R. METAL: fast and efficient meta-analysis of genomewide association scans. Bioinformatics 26, 2190–2191 (2010).
Fadista, J., Manning, A. K., Florez, J. C. & Groop, L. The (in)famous GWAS P-value threshold revisited and updated for low-frequency variants. Eur. J. Hum. Genet. 24, 1202–1205 (2016).
Flannick, J. et al. Exome sequencing of 20,791 cases of type 2 diabetes and 24,440 controls. Nature 570, 71–76 (2019).
Mahajan, A. et al. Fine-mapping type 2 diabetes loci to single-variant resolution using high-density imputation and islet-specific epigenome maps. Nat. Genet. 50, 1505–1513 (2018).
Mahajan, A. et al. Refining the accuracy of validated target identification through coding variant fine-mapping in type 2 diabetes. Nat. Genet. 50, 559–571 (2018).
Malik, R. et al. Multiancestry genome-wide association study of 520,000 subjects identifies 32 loci associated with stroke and stroke subtypes. Nat. Genet. 50, 524–537 (2018).
van der Harst, P. & Verweij, N. Identification of 64 novel genetic loci provides an expanded view on the genetic architecture of coronary artery disease. Circ. Res. 122, 433–443 (2018).
Burgess, S. et al. Using published data in Mendelian randomization: a blueprint for efficient identification of causal risk factors. Eur. J. Epidemiol. 30, 543–552 (2015).
Bowden, J., Davey Smith, G. & Burgess, S. Mendelian randomization with invalid instruments: effect estimation and bias detection through Egger regression. Int. J. Epidemiol. 44, 512–525 (2015).
Verbanck, M., Chen, C. Y., Neale, B. & Do, R. Detection of widespread horizontal pleiotropy in causal relationships inferred from Mendelian randomization between complex traits and diseases. Nat. Genet. 50, 693–698 (2018).
Pierce, B. L., Ahsan, H. & Vanderweele, T. J. Power and instrument strength requirements for Mendelian randomization studies using multiple genetic variants. Int. J. Epidemiol. 40, 740–752 (2011).
Lawlor, D. A., Tilling, K. & Davey Smith, G. Triangulation in aetiological epidemiology. Int. J. Epidemiol. 45, 1866–1886 (2016).
Sanderson, E., Davey Smith, G., Windmeijer, F. & Bowden, J. An examination of multivariable Mendelian randomization in the single-sample and two-sample summary data settings. Int. J. Epidemiol. 48, 713–727 (2019).
Di Angelantonio, E. et al. Efficiency and safety of varying the frequency of whole blood donation (INTERVAL): a randomised trial of 45,000 donors. Lancet 390, 2360–2371 (2017).
Astle, W. J. et al. The allelic landscape of human blood cell trait variation and links to common complex disease. Cell 167, e19 (2016).
Day, N. et al. EPIC-Norfolk: study design and characteristics of the cohort. European Prospective Investigation of Cancer. Br. J. Cancer 80, 95–103 (1999).
Giambartolomei, C. et al. Bayesian test for colocalisation between pairs of genetic association studies using summary statistics. PLoS Genet. 10, e1004383 (2014).
Iotchkova, V. et al. Discovery and refinement of genetic loci associated with cardiometabolic risk using dense imputation maps. Nat. Genet. 48, 1303–1312 (2016).
Iotchkova, V. et al. GARFIELD classifies disease-relevant genomic features through integration of functional annotations with association signals. Nat. Genet. 51, 343–353 (2019).
Acknowledgements
P. Surendran is supported by a Rutherford Fund Fellowship from the Medical Research Council (grant no. MR/S003746/1). N.L. is supported by the Foundation “De Drie Lichten’ in the Netherlands and the Netherlands Cardiovascular Research Initiative, an initiative supported by the Dutch Heart Foundation (CVON2012-10 PREDICT and CVON2018-30 PREDICT2). S. Karthikeyan and B.P. are funded by a BHF Programme Grant (RG/18/13/33946). J.N.H. was supported by the Vanderbilt Molecular and Genetic Epidemiology of Cancer (MAGEC) Training Program (T32CA160056, PI X.-O.S.). N.F. is supported by the National Institute of Health awards HL140385, MD012765 and DK117445. E.Y.-D. was funded by the Isaac Newton Trust/Wellcome Trust ISSF/University of Cambridge Joint Research Grants Scheme. R.C. is funded by a Medical Research Council-Newton Project Grant to study genetic risk factors of cardiovascular disease among Southeast Asians and a UK Research and Innovation-Global Challenges Research Fund Project Grant (CAPABLE) to study risk factors of non-communicable diseases in Bangladesh. F.W.A. is supported by UCL Hospitals NIHR Biomedical Research Centre. P.D. was supported by the British Heart Foundation (BHF) grant RG/14/5/30893. R.J.F.L. is funded by grants R01DK110113, U01HG007417, R01DK101855 and R01DK107786. C.H. is supported by MRC University Unit Programme grants MC_UU_00007/10 (QTL in Health and Disease) and MC_PC_U127592696. M.I.M* was a Wellcome Senior Investigator (098381; 212259) and an NIHR Senior Investigator (NF-SI-0617-10090). The research was supported by the NIHR Oxford Biomedical Research Centre (BRC) and by the Wellcome Trust (090532, 106130, 098381, 203141 and 212259). The views expressed by C.Y., S.H., J.R. and G.J.P. are those of the authors and do not necessarily represent the views of the National Heart, Lung, and Blood Institute; the National Institutes of Health; or the US Department of Health and Human Services. T.F.* is supported by the NIHR Biomedical Research Centre in Oxford. M.J. is supported by PrevMetSyn/SALVE, H2020 DynaHEALTH action (grant agreement 633595), EU H2020-HCO-2004 iHEALTH Action (grant agreement 643774). K.-H.H. is supported by PrevMetSyn/SALVE, H2020 DynaHEALTH action (grant agreement 633595) and EU H2020-HCO-2004 iHEALTH Action (grant agreement 643774). N.S. is supported by the British Heart Foundation Research Excellence Award (RE/18/6/34217). J.P. is supported by a UKRI Innovation Fellowship at Health Data Research UK. N.F. is supported by NIH awards R01-DK117445, R01-MD012765 and R21-HL140385. G.D.S., T.R.G. and C.L.R. work in the Medical Research Council Integrative Epidemiology Unit at the University of Bristol, which is supported by the Medical Research Council (MC_UU_00011/1, 4 & 5). S.T. holds a Junior 1 Clinical Research Scholar award from the Fonds de Recherche du Québec-Santé (FRQS). M.T. is supported by the BHF (PG/17/35/33001 and PG/19/16/34270) and Kidney Research UK (RP_017_20180302). V.S.R. is supported in part by the Evans Medical Foundation and the Jay and Louis Coffman Endowment from the Department of Medicine, Boston University School of Medicine. J.D.* holds a British Heart Foundation Professorship and a National Institute for Health Research Senior Investigator Award. C.M.L.* is supported by the Li Ka Shing Foundation, WT-SSI/John Fell funds and by the NIHR Biomedical Research Centre, by Widenlife and the National Institutes of Health (5P50HD028138-27). J.M.M.H.* was funded by the NIHR (Cambridge Biomedical Research Centre at the Cambridge University Hospitals NHS Foundation Trust). *The views expressed are those of the authors and not necessarily those of the NHS, the NIHR or the Department of Health and Social Care. Full acknowledgements and full lists of consortia members are provided in the Supplementary Note.
Author information
Authors and Affiliations
Consortia
Contributions
These authors contributed to the drafting of the manuscript: P. Surendran, E.V.F., N.L., I.N., A.C.M., B.M.P., E.B., D.I.C., D.L., P.B.M. and J.M.M.H. The following authors were involved in the central analyses: P. Surendran, E.V.F., N.L., I.N., S. Karthikeyan, J. Cook, D.J.L., F.D., C.N.-C., P.B.M. and J.M.M.H. All authors critically reviewed and approved the final version of the manuscript. The following authors performed bioinformatics analyses: S. Karthikeyan, L.C., B.M., C.Y., A.T.K., J.H.C., B.P.P., I.D.S., C.P.C., J.M.E., A.A., E.B.F., C.N.F., L.A.L., D.S.P., J.R.S., S. Burgess, M.K., J.P., E.Y.-D., M.R.B., M.T., P.B.M. and J.M.M.H. Study analysts: J.N.H., A.G., V.T., G. Thorleifsson, B.P.P., J.M.E., A.A., P.L.A., L.F.B., J.C.B., V.S.B., J.A.B., E.W.D., F.D., S.F.N., J.D.F., C.F., T.F., H.G., O.G., F.G., D.F.G., X.G., S.E.H., A.S.H., A.H., J.E.H., S.H., S. Kanoni, J.K., M.G.L., R.L., J. Lindström, Y.L., J. Luan, A.M., G.M., N.G.M., H.M., C.M., D.M., M. Müller-Nurasyid, G.P., M.P., A.P., R. Rainer, M. Richard, T.G.R., N. Sepúlveda, X.S., A.V.S., J.A.S., A.S., P. Sulem, S. Thériault, U.T., S. Trompet, T.V.V., D.R.V.E., G.V., S.W., S.M.W., J.Y., R.Y., B.Y., W. Zhang, J.Z., W. Zhao (University of Pennsylvania), W. Zhao (University of Michigan), E.Ε., L.L.B., K.C., T.L., I.L., M.N., N.W.R., M. Reedik, T.S., I.T., H.R.W., H.Z., S.F., B.J.H., M.I.M., T.D.S., A.S.B., H.H., C. Liu, A.K.M., A.P.M., A.C.M., D.I.C. and J.M.M.H. Study principal investigators (PIs) or co-PIs: N.F., D.O.M.-K., I.B., P.S.B., C.C., J. Connell, A.F.D., J.M.G., N.G., T.H., F. Karpe, H.A.K., K.K., M. Moitry, C.N.A.P., O. Pedersen, N.P., A.R., F.R., P.J. Sever, E. Tõnu, C.J.W., P. Almgren, P. Amouyel, F.W.A., A.I.B., M.B., M.L.B., E.P.B., J.E.B., J.C.C., Y.-D.I.C., R.C., D.C., A.C., G.D.S., R.A.d.B., I.J.D., G.D., P.D., E.D.A., P.E., S.B.F., J.F., I.F., M.F., P.W.F., P.F., G.G., T.R.G., L.G., V.G., T.B.H., C.H., B.J.H., K.-H.H., E.I., T.J., M.J., J.W.J., S.L.K., F. Kee, J.S.K., C.K., L.J.L., L. Lind, R.J.F.L., A.a.S.M., L.M., M.I.M., O.M., K.L.M., A.D.M., B.G.N., M.O.-M., C.J.P., S.P., W.P., O. Polasek, D.J.P., A.M.P., M.A.P., C.L.R., K.R., P.M.R., O.R., F.R.R., J.I.R., I.R., V.S., N.J.S., N. Sattar, W.H.-H.S., B.H.S., N. Soranzo, T.D.S., J.M.S., S.S., K.D.T., L.A.T., N.J.T., M.D.T., P.v.d.H., P.v.d.M., V.S.R., N.V., J.V., U.V., D.R. Weir, E.Z., F.J.C., N.J.W., C. Langenberg, M.T., A.S.B., M.J.C., J.D., T.L.E., A.M.H., C.M.L., A.P.M., C.O., B.M.P., D.S., K.S., E.B., D.I.C., D.L., P.B.M. and J.M.M.H. Study phenotyping: A.T.K., J.D.F., M.G.L., Y.L., S.A., E.A., S. Blankenberg, R.d.M., M.D., G.E., A.F., M.L.G., G. Hallmans, G. Heiss, P.J., E.K., A.K., K.K., T.L., L. Lannfelt, W.L., L.W.M., M.N., G.J.P., K.L.R., M. Reedik, F.R., R. Rettig, J.R., P.J. Schreiner, E.L.S., J.S., G. Thorgeirsson, E. Trabetti, T.T., S.T.T., I.T., I.V., A.V., P. Amouyel, J.E.B., J.C.C., Y.-D.I.C., R.A.d.B., J.F., G.G., V.G., B.J.H., F. Kee, J.S.K., L. Lind, R.J.F.L., O.M., W.P., O. Polasek, P.M.R., I.R., N. Sattar, W.H.-H.S., T.D.S., J.M.S., P.v.d.H., P.v.d.M., N.V., J.V., D.R. Weir, B.M.P., D.I.C. and D.L.
Corresponding authors
Ethics declarations
Competing interests
The following authors affiliated with deCODE genetics/Amgen are employed by the company: V.T., G. Thorleifsson, A.H., P. Sulem, G. Thorgeirsson, H. H, D.F.G., U.T. and K.S. B.P. serves on the steering committee of the Yale Open Data Access Project funded by Johnson & Johnson. J.D. reports grants, personal fees and nonfinancial support from Merck Sharp & Dohme; grants, personal fees and nonfinancial support from Novartis; grants from Pfizer; and grants from AstraZeneca outside the submitted work. J.D. sits on the International Cardiovascular and Metabolic Advisory Board for Novartis (since 2010); the Steering Committee of UK Biobank (since 2011); the MRC International Advisory Group (ING) member, London (since 2013); the MRC High Throughput Science ‘Omics Panel Member, London (since 2013); the Scientific Advisory Committee for Sanofi (since 2013); the International Cardiovascular and Metabolism Research and Development Portfolio Committee for Novartis; and the Astra Zeneca Genomics Advisory Board (2018). A.B. reports grants outside of this work from AstraZeneca, Biogen, Merck, Novartis and Pfizer and personal fees from Novartis. V.S. has participated in a conference trip sponsored by Novo Nordisk and received an honorarium for participating in an advisor board meeting, outside the present study. V.S. also has ongoing research collaboration with Bayer, outside the present study. The spouse of C.J.W. works at Regeneron. N.S. is partially supported by Fundação para a Ciência e Tecnologia, Portugal (grant ref. UID/MAT/00006/2013 and UID/MAT/00006/2019). M.E.J. has received research grants from AMGEN, Sanofi Aventis, Astra Zeneca and Boehringer Ingelheim. M.E.J. holds shares in Novo Nordisk. J.T. has received honoraria from AstraZeneca, Eli Lilly and Merck, and research funding from Bayer, Boehringer Ingelheim and Merck, and is a stock owner of Orion Pharma. D.O.M.-K. is a part-time clinical research consultant for Metabolon. M.I.M. has served on advisory panels for Pfizer, Novo Nordisk, Zoe Global, has received honoraria from Merck, Pfizer, Novo Nordisk and Eli Lilly and research funding from Abbvie, Astra Zeneca, Boehringer Ingelheim, Eli Lilly, Janssen, Merck, NovoNordisk, Pfizer, Roche, Sanofi Aventis, Servier and Takeda. As of June 2019, M.I.M. is an employee of Genentech and a holder of Roche stock. D.S.P. became a full-time employee of AstraZeneca during the revision of the manuscript. E.B.F. is an employee of and owns stock in Pfizer. M.J.C. is Chief Scientist for Genomics England, a UK Government company. J.M.M.H. became a full-time employee of Novo Nordisk, and I.N. became a full-time employee of Gilead during revision of the manuscript.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Power estimation for stage 2 meta-analyses.
Power calculations were performed assuming that, for any given variant, there were 1,318,884 individuals for EAWAS PA analyses, 1,164,961 participants for EAWAS EA analyses, and 670,472 participants for RV-GWAS analyses. Calculations were performed in R (https://genome.sph.umich.edu/wiki/Power_Calculations:_Quantitative_Traits).
Extended Data Fig. 2 Expression of genes implicated by the rare SNVs in GTEx v7 tissues.
We used FUMA GWAS to perform these analyses. We included genes closest to the identified rare variants from the EAWAS and the RV-GWAS.
Extended Data Fig. 3 Tissue enrichment of rare variant gene expression levels in GTEx v7.
We used FUMA GWAS to perform these analyses. We included genes closest to the identified rare variants from the EAWAS and the RV-GWAS.
Supplementary information
Supplementary Information
Supplementary Note, Figs. 1–5 and Table 26
Rights and permissions
About this article
Cite this article
Surendran, P., Feofanova, E.V., Lahrouchi, N. et al. Discovery of rare variants associated with blood pressure regulation through meta-analysis of 1.3 million individuals. Nat Genet 52, 1314–1332 (2020). https://doi.org/10.1038/s41588-020-00713-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41588-020-00713-x
- Springer Nature America, Inc.
This article is cited by
-
Can Omega-3 prevent the accidence of stroke: a mendelian randomization study
Hereditas (2024)
-
Causal effects of blood pressure and the risk of frailty: a bi-directional two-sample Mendelian randomization study
Journal of Human Hypertension (2024)
-
Germline predictors for bevacizumab induced hypertensive crisis in ECOG-ACRIN 5103 and BEATRICE
British Journal of Cancer (2024)
-
Genome-wide association analysis of hypertension and epigenetic aging reveals shared genetic architecture and identifies novel risk loci
Scientific Reports (2024)
-
Genetic imputation of kidney transcriptome, proteome and multi-omics illuminates new blood pressure and hypertension targets
Nature Communications (2024)