Abstract
Histopathology image evaluation is indispensable for cancer diagnoses and subtype classification. Standard artificial intelligence methods for histopathology image analyses have focused on optimizing specialized models for each diagnostic task1,2. Although such methods have achieved some success, they often have limited generalizability to images generated by different digitization protocols or samples collected from different populations3. Here, to address this challenge, we devised the Clinical Histopathology Imaging Evaluation Foundation (CHIEF) model, a general-purpose weakly supervised machine learning framework to extract pathology imaging features for systematic cancer evaluation. CHIEF leverages two complementary pretraining methods to extract diverse pathology representations: unsupervised pretraining for tile-level feature identification and weakly supervised pretraining for whole-slide pattern recognition. We developed CHIEF using 60,530 whole-slide images spanning 19 anatomical sites. Through pretraining on 44 terabytes of high-resolution pathology imaging datasets, CHIEF extracted microscopic representations useful for cancer cell detection, tumour origin identification, molecular profile characterization and prognostic prediction. We successfully validated CHIEF using 19,491 whole-slide images from 32 independent slide sets collected from 24 hospitals and cohorts internationally. Overall, CHIEF outperformed the state-of-the-art deep learning methods by up to 36.1%, showing its ability to address domain shifts observed in samples from diverse populations and processed by different slide preparation methods. CHIEF provides a generalizable foundation for efficient digital pathology evaluation for patients with cancer.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Main
Histopathology image evaluation is integral to the diagnosis of cancers and cancer subtype classification. Previous studies on artificial intelligence (AI)-based histopathology image analysis primarily rely on training task-specific models optimized for each use case1,2. For example, specialized deep neural networks have been developed for cancer cell identification4,5, histological and molecular subtype classification6,7,8,9,10, prognosis evaluation11,12,13,14 and treatment response prediction using gigapixel whole-slide images (WSIs)15,16,17. Moreover, state-of-the-art computational pathology analyses have revealed quantitative morphological signals indicative of clinically important molecular markers18,19, demonstrating the potential of AI methods in identifying cellular features imperceptible to the human eyes20. Although these advances offer promising avenues for improving cancer evaluation, several limitations continue to plague quantitative pathology image analyses. To begin with, standard deep learning methods require a large amount of data to train a performing model for each task. As it is difficult to obtain comprehensive pathology representations that cover the heterogeneity of diverse tissue microenvironments, existing approaches mainly focus on solving each narrow diagnostic task individually1,7. In addition, most AI models for pathology imaging analyses are tailored from general computer vision models designed for classifying macroscopic objects (for example, animals, cars and buses)2. These conventional approaches do not leverage the general tissue pathology patterns when training specialized diagnostic models. Furthermore, AI models trained by images from a single source tend to overfit the training data distribution and suffer from substantial performance deterioration when applied to images processed by different pathology laboratories3,21. These limitations have hindered the effective application of state-of-the-art AI models for reliable pathology evaluation.
Self-supervised learning has emerged as a promising approach for obtaining robust image feature representation useful for a wide range of prediction tasks using samples collected in diverse settings22,23. As diverse unlabelled training data are relatively straightforward to collect and the model training process is task-agnostic, self-supervised learning has achieved robust performance across different tasks and data distributions, such as image retrieval24,25,26 and weakly supervised WSI analysis27. Recent advancements in self-supervised learning for pathology image analyses further utilized both images and their text descriptions to augment the performance of computer vision models28,29. However, these methods have two major limitations. First, they primarily focus on individual image tiles in the WSIs, without considering the interactions of different regions of the same tissue. Second, previous studies focused on narrow diagnostic tasks and did not evaluate the generalizability of the extracted quantitative imaging features in different prediction tasks across cancer types and samples from several sources. As pathologists often face a variety of disease samples and need to assimilate contextual information from the tissue microenvironment, developing a general-purpose pathology AI system capable of accommodating a wide range of tissue types and evaluation tasks is of paramount importance.
To address these pressing clinical needs, we established the CHIEF model, a general-purpose machine learning framework that provides the foundation for various pathology diagnosis and prediction tasks (Fig. 1a). We leveraged two complementary forms of AI model pretraining: self-supervised pretraining using 15 million pathology image tiles for tile-level feature representation and weakly supervised pretraining on 60,530 WSIs across 19 anatomical sites for tissue context representation. In addition, we devised an efficient framework for tile-level feature aggregation in large-scale WSI analysis. We further validated CHIEF’s capability in cancer detection, tumour origin characterization, genomic mutation identification and survival prediction using 32 independent datasets consisting of 19,491 weakly annotated WSIs. Our approach challenges conventional attention-based tile-aggregation methods, offering a holistic representation of WSI features. CHIEF enables systematic microscopic feature identification and lays the groundwork for reliable pathology evaluation.
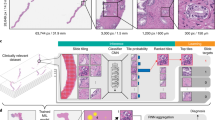
a, CHIEF is a generalizable machine learning framework for weakly supervised histopathological image analysis. CHIEF extracts pathology imaging representations useful for cancer classification, tumour origin prediction, genomic profile prediction and prognostic analyses. We pretrained CHIEF in a weakly supervised manner using 60,530 WSIs representing 19 anatomical sites. During the pretraining process, we cropped the WSIs into non-overlapping imaging tiles, and we encoded the anatomic site information of each WSI using the contrastive language–image pretraining (CLIP) embedding method to obtain a feature vector for each anatomic site. We merged the text and image embeddings to represent the heterogeneous pathology information from the training data. We then used the pathology imaging features extracted by CHIEF to infer cancer types directly. In the genomic profile and prognostic prediction tasks, CHIEF features served as the foundation for fine-tuning models for each specific task. b, A summary of the 60,530 slides for training the CHIEF model. We collected these pathology slides belonging to 19 anatomical sites from 14 cohorts. c, CHIEF significantly outperformed state-of-the-art methods in cancer classification, genomic profile identification and survival prediction tasks. The left panel summarizes the AUROCs for cancer classification and genomic profile prediction tasks. Overall, CHIEF outperformed state-of-the-art deep learning methods by up to 36.1% in these tasks. The right panel outlines the c-index of survival prediction. On average, CHIEF performed 9% better than conventional methods. Supplementary Tables 1–3 show detailed performance comparisons. DFCI, Dana–Farber Cancer Institute; PAIP, Pathology AI Platform; PLCO, the Prostate, Lung, Colorectal and Ovarian study. The graphics of the human and DNA in a were created with BioRender.com.
An overview of CHIEF
We established the CHIEF model, a general-purpose machine learning framework for weakly supervised histopathological image analyses. Unlike commonly used self-supervised feature extractors27,30, CHIEF leveraged two types of pretraining procedure: unsupervised pretraining on 15 million unlabelled tile images and weakly supervised pretraining on more than 60,000 WSIs. Tile-level unsupervised pretraining established a general feature extractor30 for haematoxylin–eosin-stained histopathological images collected from heterogeneous publicly available databases, which captured diverse manifestations of microscopic cellular morphologies. Subsequent WSI-level weakly supervised pretraining constructed a general-purpose model by characterizing the similarities and differences between cancer types. We evaluated the performance of CHIEF in a wide range of pathology evaluation tasks, including cancer detection, tumour origin prediction, genomic profile identification and survival prediction (Fig. 1a). The details of model design and implementation are described in the Methods.
CHIEF augmented cancer cell detection
Detecting malignant cells from pathological images is crucial for cancer diagnoses4,5. State-of-the-art AI methods for cancer cell detection predominantly concentrate on training models for specific cancer types, without leveraging the commonalities of malignant cell morphology across cancers. The resulting models are not easily extensible to other cancer categories. To address this gap, we built a weakly supervised cancer detection platform using CHIEF and evaluated its generalizability across cancers. We conducted an extensive external validation using 15 independent datasets with a total of 13,661 WSIs. These datasets encompass both public (for example, Clinical Proteomic Tumor Analysis Consortium (CPTAC), Diagset-B31, Dataset-patient-level-test (Dataset-PT)32, the Diagnostic Reference Oncology Imaging Database (DROID)-Breast and TissueNet33 cohorts) and institutional (for example, samples from Shenzhen Maternity & Child Healthcare Hospital (SMCH) and Chongqing University Cancer Hospital (CUCH)) data sources, contain biopsy and surgical resection slides and span 11 different primary cancer sites, including the breast, uterus–endometrium, oesophagus, stomach, cervix, colon, prostate, kidney, skin, pancreas and lung. To better assess the performance of CHIEF, we compared it with three weakly supervised WSI classification methods: clustering-constrained-attention multiple instance learning (CLAM)6, attention-based deep multiple instance learning (ABMIL)34 and dual-stream multiple instance learning networks (DSMIL)35.
CHIEF consistently attained superior performance in a variety of cancer identification tasks using either biopsy or surgical resection slides (Fig. 2a). CHIEF achieved a macro-average area under the receiver operating characteristic curve (AUROC) of 0.9397 across 15 datasets representing 11 cancer types (Fig. 2a), which is approximately 10% higher than that attained by DSMIL (a macro-average AUROC of 0.8409), 12% higher than that of ABMIL (a macro-average AUROC of 0.8233) and 14% higher than that of CLAM (a macro-average AUROC of 0.8016). In all five biopsy datasets collected from independent cohorts, CHIEF possessed AUROCs of greater than 0.96 across several cancer types, including oesophagus (CUCH-Eso), stomach (CUCH-Sto), colon (CUCH-Colon) and prostate (Diagset-B and CUCH-Pros). On independent validation with seven surgical resection slide sets spanning five cancer types (that is, colon (Dataset-PT), breast (DROID-Breast), endometrium (SMCH-Endo and CPTAC-uterine corpus endometrial carcinoma (UCEC)), lung (CPTAC-lung squamous cell carcinoma (LUSC)) and cervix (SMCH-Cervix and TissueNet)), CHIEF attained AUROCs greater than 0.90. Both CHIEF and the set of baseline methods had lower performance in CPTAC. Nonetheless, CHIEF significantly outperformed all other methods in cancer cell identification in these datasets (DeLong test P value < 0.001). These results demonstrated CHIEF’s generalizability across diverse cancer tissues and samples obtained from heterogeneous sources internationally.
a,b, We validated CHIEF’s capability of cancer detection using 15 independent datasets collected from several hospitals worldwide. Our test datasets encompassed 13,661 WSIs from 11 sites of origin (breast, endometrium–uterus, oesophagus, stomach, cervix, colon, prostate, kidney, skin, pancreas and lung). a, CHIEF attained up to 0.9943 in the AUROCs across 15 independent test datasets and consistently outperformed (two-sided Wilcoxon signed-rank test P value = 0.000061) three deep learning methods (that is, CLAM, ABMIL and DSMIL). The receiver operating characteristic curves of CHIEF and baseline methods are shown. The mean AUROC and its 95% CIs, calculated using the non-parametric bootstrapping method (n = 1,000 replicates), are presented. The diagonal dashed line in each plot represents the performance of a null model. CCRCC, clear cell renal cell carcinoma; CM, cutaneous melanoma; PDA, pancreatic ductal adenocarcinoma. b, Visualization of model attention scores showed CHIEF accurately identified cancerous regions in WSIs. For each cancer type, the left image panel represents the ground truth annotations labelled by experienced pathologists. As CHIEF uses a weakly supervised approach that requires only slide-level annotations, these region-level annotations were not revealed to the model during the training phase. The middle panel visualizes the amount of attention CHIEF paid to each region in the WSIs. The right panel shows the zoomed-in view of regions receiving high (image tiles with red outlines) and low (image tiles with black outlines) attention scores. Extended Data Fig. 2 and Supplementary Fig. 1 show additional result visualization of this classification task. The original WSIs and their corresponding heat maps are available at https://yulab.hms.harvard.edu/projects/CHIEF/CHIEF.htm. Scale bars, 2 mm (left images for Cervix (TissueNet), Prostate (Diagset-B) and Colon (Dataset-PT)), 3 mm (left image for Breast (DROID-Breast)) and 50 μm (bottom right magnifications).
We used whole-slide attention visualization to identify diagnostic signals utilized by the CHIEF models. Figure 2b, Extended Data Fig. 2 and Supplementary Fig. 1 show the original WSIs, pixel-level ground truth annotated by pathologists (Methods) and attention maps output by CHIEF. CHIEF directed most of its attention to cancerous regions, exhibiting a remarkable alignment with ground truth annotations at the pixel level despite being trained only on slide-level labels. Notably, tiles receiving high attention from CHIEF contained tissue with typical cytologic and architectural patterns of malignancy (for example, increased nuclear/cytoplasmic ratio, irregularly shaped nuclei, cellular pleomorphism and disorganized architecture), showing the model’s capacity to identify key diagnostic features using a weakly supervised approach.
CHIEF predicted genomic profiles
Genomic profiles of cancer samples indicate patients’ treatment responses and are crucial for formulating treatment plans19. The comprehensive genomic profiling of patients with cancer is not routinely conducted worldwide owing to the additional cost and time involved18. Identifying quantitative morphological patterns indicative of genomic profiles from routine haematoxylin–eosin-stained slides offers an instantaneous and cost-effective alternative to genomic sequencing. We examined CHIEF’s capability to systematically predict molecular profiles of cancer samples. We focused on four clinically important prediction tasks: systematic prediction of prevalent genetic mutations across cancer types; identification of mutations related to targeted therapies; isocitrate dehydrogenase (IDH) status prediction for the new WHO (World Health Organization) classification of glioma; and microsatellite instability (MSI) prediction for assessing the benefits of immune checkpoint blockade in patients with colorectal cancer (CRC).
Prevalent genetic mutations
We conducted a systematic analysis that associated prevalent genetic mutations with histopathology images (Fig. 3 and Extended Data Fig. 3). Our study involved 13,432 WSIs across 30 cancer types and 53 genes with the top five highest mutation rates in each cancer type.
CHIEF predicted prevalent somatic mutations (n = 11,483) and mutations related to targeted therapies (n = 6,013) in several cancer types using histopathology images alone. We stratified our analyses by cancer types and organized the prediction results by genes. The detailed sample counts for each cancer type can be found in Supplementary Tables 17 and 18. Owing to differences in the tumour microenvironment in different cancer types, variations in the prediction performance were observed. The mean ± 95% CI for each prediction task is shown. Error bars represent the 95% CIs estimated by fivefold cross-validation. ACC, adrenocortical carcinoma; BLCA, bladder urothelial carcinoma; CESC, cervical squamous cell carcinoma and endocervical adenocarcinoma; COADREAD, colon adenocarcinoma and rectum adenocarcinoma; DLBC, diffuse large B cell lymphoma; ESCA, oesophageal carcinoma; HNSC, head and neck squamous cell carcinoma; KICH, chromophobe renal cell carcinoma; KIRC (also known as CCRCC), clear cell renal cell carcinoma; KIRP, papillary renal cell carcinoma; LIHC, liver hepatocellular carcinoma; MESO, mesothelioma; OV, ovarian serous cystadenocarcinoma; PAAD, pancreatic adenocarcinoma; PCPG, pheochromocytoma and paraganglioma; PRAD, prostate adenocarcinoma; SARC, sarcoma; SKCM, skin cutaneous melanoma; STAD, stomach adenocarcinoma; TGCT, testicular germ cell tumors; THCA, thyroid carcinoma; THYM, thymoma; UCS, uterine carcinosarcoma; UVM, uveal melanoma.
CHIEF predicted the mutation status of nine genes with AUROCs greater than 0.8 in our systematic pan-cancer genetic mutation analyses (Fig. 3). Consistent with previous studies18,36, pathology images contain strong signals related to TP53 mutation across 19 cancer types, with high AUROCs in low-grade glioma (LGG; 0.8756; 95% confidence interval (CI) 0.8624–0.8888), adrenal carcinoma (0.8119; 95% CI 0.7488–0.8751) and UCEC (0.8115; 95% CI 0.7971–0.8259). CHIEF also identified mutations in GTF2I, which occur in 43.4% of patients with thymic epithelial tumours37, with an AUROC of 0.9111 (95% CI 0.8935–0.9287). Furthermore, CHIEF predicted BAP1 mutation in uveal melanoma (AUROC = 0.817; 95% CI 0.7668–0.8672), which is observed in approximately 45% of uveal melanoma cases38.
We tested CHIEF in an independent patient cohort from CPTAC. CHIEF consistently maintained similar AUROCs for various genes in these new patient cohorts (Extended Data Fig. 4). Compared with the state-of-the-art method for histopathology-based genomic mutation prediction (that is, the pan-cancer computational histopathology (PC-CHiP) method36; Supplementary Fig. 2), CHIEF showed significantly higher performance (Wilcoxon signed-rank test P value < 0.001), with a macro-average AUROC of 0.7043 (range 0.51–0.89). By contrast, the PC-CHiP method attained a macro-average AUROC of 0.6523 (range 0.39–0.92).
Mutations related to targeted therapies
We further used CHIEF to predict genes associated with FDA (Food and Drug Administration)-approved targeted therapies presented in OncoKB39 (www.oncokb.org) across 18 genes spanning 15 cancer types (Fig. 3). CHIEF predicted the mutation status of all 18 genes with AUROCs greater than 0.6 (Fig. 3). Mutations with high prediction performance included EZH2 in diffuse large B-cell lymphoma (AUROC = 0.9571; 95% CI 0.9321–0.9822), NTRK1 in stomach adenocarcinoma (AUROC = 0.8192; 95% CI 0.7767–0.8618), BRCA2 in prostate adenocarcinoma (AUROC = 0.8938; 95% CI 0.8310–0.9567), BRAF in thyroid carcinoma (AUROC = 0.8889; 95% CI 0.8715–0.9064), ERBB2 in lung squamous cell carcinoma (LUSC; AUROC = 0.8211; 95% CI 0.7597–0.8826) and FGFR3 in bladder urothelial carcinoma (AUROC = 0.8161; 95% CI 0.7921–0.8402). On independent validation, CHIEF achieved a similar level of performance in the CPTAC cohorts (Extended Data Fig. 4). Among these genes, ESR1 in breast cancer (BRCA), EGFR in lung adenocarcinoma (LUAD) and BRAF in colon adenocarcinoma and rectum adenocarcinoma (COADREAD) all exhibited AUROCs greater than 0.7 in both held-out and independent test sets.
IDH status prediction
The fifth edition of the WHO Classification of Tumors of the Central Nervous System distinguished glioblastoma (GBM) from LGG on the basis of IDH status instead of conventional histological features8,40. Thus, it is crucial to identify patients’ IDH status at the time of diagnosis. To identify IDH mutation-related signals independent of histological grades, we stratified our study cohorts by histological grade and used CHIEF to predict IDH status in each stratum. We conducted IDH status prediction analyses on six datasets: The Cancer Genome Atlas (TCGA)-LGG, TCGA-GBM, Medical University of Vienna (MUV)-LGG41, MUV-GBM41, Harvard Medical School and the University of Pennsylvania (HMS)-LGG and HMS-GBM, including a total of 2,718 WSIs. The CHIEF model demonstrated superior performance compared to other baseline methods in both the held-out and independent test sets (Wilcoxon signed-rank test P value < 0.01; Fig. 4a and Supplementary Fig. 3). To increase interpretability, we visualized the quantitative image feature vectors and examined the distribution of attention scores determined by CHIEF (Extended Data Figs. 5 and 9b). Results showed that necrotic regions received significantly higher attention when identifying gliomas with IDH-wild-type status (Mann–Whitney U-test P < 0.0001; Extended Data Fig. 9b).
a, CHIEF successfully identified IDH mutation status in low histological grade groups (n = 1,289). These results indicated that CHIEF characterized IDH-related morphological signals independent of histological grades. As the fifth edition of the WHO Classification of Tumors of the Central Nervous System 40 incorporated IDH mutation status in the definition of GBM and LGG, CHIEF provides molecular profile predictions that enable fast cancer classification based on the new clinical guidelines. The left panels show the mean receiver operating characteristic curves of tenfold cross-validations using the TCGA-LGG (n = 842) dataset. The middle and right panels show the validation results in the independent datasets (MUV-LGG (n = 365) and HMS-LGG (n = 82)). b, CHIEF identified patient with MSI-high status with AUROCs of 0.869–0.875. The left panel represents the MSI prediction performance in the TCGA-COADREAD dataset (n = 437) using fourfold cross-validation. The middle and right panels illustrate the performance of two independent test sets (that is, PAIP2020 (n = 77) and CPTAC-COAD (n = 221)). Results in a,b are presented as mean ± s.d. across cross-validation.
MSI status prediction
MSI is a well-established biomarker for responses to immune checkpoint blockade in CRCs27. To enable rapid treatment personalization at the time of diagnosis, we examined the performance of CHIEF in predicting MSI status using histopathological images. CHIEF significantly outperformed the best-performing baseline method (DSMIL) in the TCGA-COADREAD dataset and two independent cohorts (PAIP202042 and CPTAC-COAD), with an AUROC improvement of approximately 12%, 15% and 26%, respectively (Fig. 4b). Attention analyses showed that regions containing solid tumours, luminal necrosis and tumour-infiltrating lymphocytes received high attention from CHIEF (Extended Data Fig. 6).
CHIEF predicted survival outcomes
Owing to differential responses to standard treatments, patients with cancer have varying disease-specific survival outcomes after their initial diagnoses43. Although many clinical and genomic biomarkers have been proposed, they do not fully predict the prognosis of every patient. To address this challenge, we extended our CHIEF framework to establish stage-stratified survival prediction models for each cancer type under study. We used a total of 9,404 WSIs in 17 datasets (from both publicly available and institutional sample sources) and focused on 7 cancer types (COADREAD, LUSC, BRCA, GBM, UCEC, LUAD and renal cell carcinoma (RCC)) with reliable prognostic information in the independent cohorts.
CHIEF successfully predicted patients’ survival outcomes using the histopathology images obtained at the time of initial diagnosis. In all cancer types and all study cohorts, CHIEF distinguished patients with longer-term survival from those with shorter-term survival (log-rank test P < 0.05; Fig. 5 shows the prediction results of patients with stage I and stage II cancer). In comparison, state-of-the-art deep learning methods (for example, pathology-omics research platform for integrative survival estimation (PORPOISE)12 and DSMIL35) cannot reliably differentiate patients with different survival outcomes in the same settings (log-rank test P > 0.05 in 11 out of 15 cohorts; Supplementary Fig. 4). In addition, the Kaplan–Meier curves produced by CHIEF possessed narrower CIs than other methods. Overall, CHIEF attained an average concordance index (c-index) of 0.74 across cancer types in the held-out test set (Supplementary Table 3), which was 12% and 7% higher than those of PORPOISE and DSMIL (0.62 and 0.67, respectively). Notably, the performance difference between CHIEF and baseline methods was even more pronounced in the independent cohorts not participating in the model development process. In these patient populations, CHIEF attained an average c-index of 0.67 (9% better than all baseline models) and distinguished patients with different survival outcomes in all datasets, whereas PORPOISE and DSMIL yielded average c-indices of 0.54 and 0.58, respectively.
a, CHIEF distinguished longer-term survivors from shorter-term survivors among patients with stage I and stage II cancer (n = 4,147). Kaplan–Meier curves for CHIEF-based predictions are shown. The red and blue lines represent patients with high and low mortality risks, respectively. In comparison, PORPOISE and DSMIL did not generalize well to 11 out of 15 independent patient cohorts (Supplementary Fig. 4). Two-sided log-rank test without adjustment is used to compare the survival distributions between the high-risk and low-risk groups (P = 0.0005 in TCGA-BRCA, P = 0.0189 in DFCI-BRCA, P = 0.0013 in PLCO-BRCA, P < 0.0001 in TCGA-RCC, P = 0.0495 in CPTAC-RCC, P = 0.0293 in Brigham and Women’s Hospital (BWH)-RCC, P = 0.0006 in TCGA-LUAD, P = 0.035 in DFCI-LUAD, P = 0.011 in PLCO-LUAD, P = 0.0144 in TCGA-LUSC, P < 0.0001 in CPTAC-LUSC, P = 0.0004 in TCGA-UCEC, P = 0.0176 in CPTAC-UCEC, P = 0.0003 in TCGA-COADREAD, and P = 0.0008 in PLCO-COADREAD). Error bands represent 95% CIs. b, CHIEF significantly outperformed other methods in predicting survival outcomes of patients with cancer. c-indices of held-out (n = 2,593) and independent cohorts (n = 1,554) are shown. Box plots were generated on the basis of fivefold cross-validation. Dashed lines represent the mean c-indices across datasets. In these box plots, the central line is the median, box bounds are the 25th and 75th percentiles, and whiskers extend to 1.5 times the interquartile range. These statistics included samples from TCGA-BRCA (n = 760), TCGA-COADREAD (n = 294), TCGA-LUAD (n = 344), TCGA-LUSC (n = 334), TCGA-RCC (n = 507), TCGA-UCEC (n = 354), DFCI-BRCA (n = 48), PLCO-BRCA (n = 647), DFCI-LUAD (n = 235), PLCO-LUAD (n = 139), CPTAC-LUSC (n = 81), CPTAC-RCC (n = 124), BWH-RCC (n = 49), CPTAC-UCEC (n = 183) and PLCO-COADREAD (n = 48). The detailed sample count for each patient cohort is available in Supplementary Table 21.
We observed similar performance trends in patients with stage III (Supplementary Fig. 6) and stage IV cancers (Supplementary Fig. 7), with CHIEF outperforming other methods by up to 10%. As some previously published methods focused on mixed-stage results, we computed the results from mixed-stage analyses and showed that CHIEF outperformed baseline methods in these study settings (Extended Data Fig. 7 and Supplementary Fig. 5). In addition, we conducted a multivariate analysis that incorporated model-derived risk score, patient age, sex and stage (Supplementary Tables 9 and 10). Results showed that CHIEF-derived risk score is a significant prognostic factor independent of known indicators of survival outcomes. Furthermore, our univariate analysis showed that CHIEF-derived risk scores are statistically significantly associated with survival outcomes across all cancer types in all patient cohorts under investigation (Supplementary Tables 11 and 12). In comparison, the risk scores predicted by other pathology imaging-based methods cannot differentiate patients’ survival outcomes in most patient cohorts using either multivariate or univariate analyses.
To better understand the histological features indicative of patients’ survival outcomes, four attending pathologists independently reviewed the attention heat maps generated by CHIEF (Methods). In both longer-term survivors and shorter-term survivors, high-attention areas contained malignant tissues across cancer types (Extended Data Figs. 8 and 9 and Supplementary Figs. 8 and 9). High-attention areas for longer-term survivors had more infiltrating immune cells than those for patients with higher mortality risks. In cancer samples from shorter-term survivors, high-attention regions exhibited larger nuclear/cytoplasmic ratios, more pronounced nuclear atypia, less stromal fibrosis, and weak intercellular adhesion.
Discussion
We developed CHIEF as a general-purpose, pan-cancer foundation deep learning framework for quantitative pathology evaluation. CHIEF leveraged unsupervised tile-level pretraining, weakly supervised WSI-level pretraining and 44 terabytes of histopathology imaging data from several countries for robust pathology image analysis. The CHIEF framework successfully characterized tumour origins, predicted clinically important genomic profiles, and stratified patients into longer-term survival and shorter-term survival groups. Furthermore, our approach established a general pathology feature extractor capable of a wide range of prediction tasks even with small sample sizes. Our results showed that CHIEF is highly adaptable to diverse pathology samples obtained from several centres, digitized by various scanners, and obtained from different clinical procedures (that is, biopsy and surgical resection). This new framework substantially enhanced model generalizability, a critical barrier to the clinical penetrance of conventional computational pathology models1,3.
CHIEF effectively leveraged anatomic site information as a source of prior knowledge and considered the contextual interactions across different image regions in the WSIs, contributing to substantially better generalizability than standard approaches. We successfully used the CHIEF framework in various WSI-level prediction tasks, and our models achieved superior performance compared to state-of-the-art methods. For example, CHIEF exhibited a robust ability to recognize the origins of the primary tumours in patient cohorts not involved in the training process.
In addition, CHIEF substantially outperformed baseline methods in predicting genomic variations using pathology imaging profiles36. In particular, CHIEF predicted the mutation status of several oncogenes and tumour suppressors with higher performance (AUROCs > 0.8), such as TP53, GTF2I, BTG2, CIC, CDH1, IGLL5 and NRAS. As the updated WHO diagnostic guidelines incorporated molecular markers in tumour classifications, we further showed that CHIEF predicted key mutations related to major diagnostic categories and validated the results in several patient populations. CHIEF also accurately predicted the MSI status of patients with CRC, which may facilitate clinical decisions regarding the administration of immune checkpoint inhibitors18,19,27. Finally, imaging features extracted by CHIEF served as the foundation for survival outcome prediction models. These models stratified patients into high- and low-mortality risk groups across all cancer types under study, and the results were validated in 17 cohorts.
We further interpreted CHIEF models by visualizing imaging regions that received high attention from the model. CHIEF used a weakly supervised machine learning approach, which identified the regions of interest automatically by comparing positive and negative examples, thereby eliminating the need for pixel-level or region-level annotations. This approach made it possible to leverage large-scale publicly available and institutional datasets to capture the heterogeneity of pathology manifestations across thousands of samples. For example, visualization of survival outcome prediction models indicated that samples from patients with cancer with lower mortality risks contain more infiltrating immune cells and abundant stroma with clear glandular and cribriform structures.
Last, we showed that CHIEF outperformed recently released general-purpose foundation models and patch-based pathology foundation models with statistically significant performance differences26,44,45,46 (Supplementary Fig. 10 and Supplementary Tables 25 and 26). The additional weakly supervised pretraining approach leveraging large-scale WSI datasets probably contributed to its enhanced performance.
Our study has a few limitations. First, although CHIEF was trained with a large number of samples collected from several hospitals and study cohorts worldwide, the inclusion of a larger number of non-malignant slides and slides from rare diseases could further improve the performance of our general-purpose pathology feature extractor. In addition, our prognostic prediction models focused on the disease-specific and overall survival prediction of patients receiving standard care. Future research can extend our methods to study the predicted benefits and adverse effects of new cancer treatments.
In conclusion, CHIEF is a foundation model useful for a wide range of pathology evaluation tasks across several cancer types. We have demonstrated the generalizability of this foundation model across several clinical applications using samples collected from 24 hospitals and patient cohorts worldwide. CHIEF required minimal image annotations and extracted detailed quantitative features from WSIs, which enabled systematic analyses of the relationships among morphological patterns, molecular aberrations and important clinical outcomes. Accurate, robust and rapid pathology sample assessment provided by CHIEF will contribute to the development of personalized cancer management.
Methods
Datasets for CHIEF pretraining
The CHIEF model was pretrained using 60,530 WSIs from 14 study cohorts, including eight large study consortia (TCGA47, Genotype-Tissue Expression (GTEx)48, PAIP, Prostate Cancer Grade Assessment (PANDA)49, Basal Cell Carcinomas (BCC)50, Early Breast Cancer Core-Needle Biopsy WSI (BCNB)51, Automatic Registration of Breast Cancer Tissue (ACROBAT)52 and Treatment Effectiveness to Ovarian Cancer (TOC)53) and six institutional cohorts (Yuhuangding Hospital (YH)-Breast, YH-Eso, YH-Colon, YH-Sto, YH-Cervix and YH-Endo) from YH, Yantai, China. The training datasets included cancers from 19 anatomic sites, including brain, breast, bladder, kidney, prostate, testis, lung, pancreas, liver, skin, ovary, cervix, uterus, colon, oesophagus, stomach, thyroid, adrenal gland and soft tissues. We obtained formalin-fixed paraffin-embedded haematoxylin and eosin (H&E)-stained tissues from these patient cohorts. Figure 1b summarizes the breakdowns of the slide counts across these cohorts. Below we describe these cohorts in detail.
Datasets from large research consortia
We first obtained 46,340 publicly available H&E-stained WSIs. These included 29,001 slides of 19 anatomical sites from TCGA and GTEx48. In addition, we acquired 2,405 WSIs of 5 cancer types from PAIP. These WSIs contained cancers from the liver (558 WSIs), colon (894 WSIs), prostate (399 WSIs), kidney (390 WSIs) and pancreas (164 WSIs). We further incorporated data from PANDA49, BCC50, BCNB51, ACROBAT52 and TOC53. Each of these research consortia focuses on a single cancer type (for example, prostate, skin, breast by two consortia, and ovary). Among these datasets, PANDA is the largest publicly available prostate histopathological image set, containing 10,616 WSIs of prostate biopsies from 2,113 patients. Two pathologists (J.J. and F.W.) reviewed these digital pathology slides in PANDA and removed 13 low-quality slides. The BCC dataset50 contained 1,832 WSIs of basal cell carcinomas. The BCNB51 and ACROBAT52 datasets contained 1,058 and 1,153 H&E-stained WSIs obtained by BRCA biopsy and surgical resection, respectively. The TOC53 dataset contained 288 H&E-stained pathology slides from patients with ovarian cancer. Supplementary Table 13 summarizes the detailed information for each patient cohort.
Institutional datasets
As most participants in the large research consortia are Caucasian, we further included six institutional datasets from a wide range of demographic groups for model pretraining. Specifically, we collected an additional 14,190 slides from six patient cohorts from YH, Yantai, China. This sample set contains pathology slides of breast, oesophagus, stomach, cervix, uterus and colon cancers. Supplementary Table 13 summarizes the detailed information of these study cohorts.
CHIEF model architecture
CHIEF is pretrained with a two-stage process to capture pathology manifestations useful for a wide range of evaluation tasks. First, we used self-supervised pretraining to obtain patch-level feature representations from unlabelled data. Second, we integrated patch-level features using weakly supervised learning and an attention module, thereby generating global pathology representations of WSIs. The second stage requires only WSI-level labels, enabling CHIEF to construct a holistic understanding of pathology images from global features.
Figure 1 shows an architecture overview of the CHIEF model. CHIEF integrated multi-modality information from microscopic imaging and anatomical site information to enhance feature representation for quantitative pathology analyses. By incorporating both histological images and text information, the CHIEF pretraining strategy enhances the model’s capability to account for anatomical information and optimize structural feature embeddings, thereby enhancing the model’s feature representations. In short, we established a histopathological image branch for image encoding and another text branch for anatomic site encoding. The image encoder used the self-supervised CTransPath backbone30 for extracting histopathology image feature representations. We aggregated these features using attention-based feature fusion, with assistance from instance-level feature identification and WSI-level contrastive learning (Supplementary Fig. 13). The text encoder adopted the pretrained text encoder from the CLIP model54, which was obtained by pretraining on diverse datasets of images and their captions to learn the rich and multimodal representations that capture the relationships between images and text descriptions. Below we elaborate on our methods in detail.
Anatomical site information encoding
The anatomic site information for each WSI is often available but rarely utilized to improve machine learning models for pathology image evaluation. To address this gap, we added text information on the anatomic sites into the feature representation to enhance supervision during the training process of CHIEF. To ensure the effectiveness of the text feature representation, we leveraged the text encoder of the CLIP model for text embedding extraction. This text encoder is a transformer-based model and pretrained with a dataset of 400 million image–text pairs54.
As pathology samples from many large research consortia lack detailed text descriptions, we used simple text prompts as the input of the text encoding branch. Our prompt took the form of “This is a histopathological image of the [CLS]”, in which the [CLS] was the anatomic site of the samples, such as the brain, stomach or other organs. Mathematically, let \({T}_{n}\) and \(\widetilde{{F}_{n}}\) be the text embedding (CLIP embedding) and image embedding of the nth slides, respectively. The text embedding is further passed through a multilayer perceptron (MLP) with two fully connected layers and then concatenated with the visual features on the image branch; that is, \({F}_{n}^{{\rm{fusion}}}={\rm{MLP}}({T}_{n})+\widetilde{{F}_{n}}\). Through the pretraining process, the CHIEF model learned to associate visual features with corresponding text descriptions, thereby identifying their semantic relevance across organs.
Histopathological image feature encoding
As most histopathology images from clinical sources do not come with detailed region-level annotations, we designed an image processing branch for weakly supervised WSI analysis. Our approaches effectively learn the relationships between WSIs and labels assigned to these slides, without requiring region-level annotations from pathologists. Two key elements of our image feature encoding branch are data preprocessing and a weakly supervised feature aggregation network. During data preprocessing, we processed each WSI using the Otsu thresholding method55 to remove the image background not representing any tissues. Next, we cropped the WSIs into non-overlapping tiles with a size of 256 × 256 pixels at a magnification of ×10 with a resolution of 1.0 μm per pixel. We used CTransPath pretrained on 15 million image patches to obtain the quantitative representation of each tile. We further designed the feature aggregator network to integrate the context information across tiles in each WSI. This core element of the histopathological image branch used the attention-based pooling strategy and consisted of three modules. First, the main module is a deep attention aggregation method with class-specific attention computation, which generates a learnable attention score for each tile in WSIs. To enhance the efficiency of these attention scores, we included two auxiliary modules to perform the inter-WSI and intra-WSI feature learning, respectively. Specifically, the instance branch assigned an attention score of 1 for tiles receiving the highest attention levels and a score of 0 for tiles obtaining the lowest attention. The WSI branch performed WSI-level contrastive learning to facilitate information integration across regions in WSIs, enabling robust separation for each category labelled at the WSI level. The Supplementary Methods describes these three modules in greater detail.
CHIEF pretraining details
We pretrained CHIEF with 60,530 WSIs from 14 cohorts that were split into 90% training data and 10% validation data. We split the training data at the patient level and ensured that samples from different anatomic sites were represented in the training and validation sets proportionally. In the training phase, the memory banks in the WSI-level contrastive learning module were constructed separately for different cancer types. In the validation phase, we calculated the AUROC, sensitivity, specificity and other validation set performance metrics for each anatomic site individually. We optimized the model hyperparameters to maximize the average AUROC across sites. The weakly supervised learning adopted a batch size of 1 WSI and a maximum epoch number of 50. We used the Adam optimizer56 with an initial learning rate of 3.0 × 10−4. We used the cosine annealing method57 to determine the learning rate schedule. We exploited the early stop strategy to mitigate overfitting, which terminated network training when the validation AUROC no longer increased in ten consecutive epochs. CHIEF was pretrained using eight NVIDIA V100 32-GB GPUs.
Evaluation
We evaluated the performance and generalizability of the pretrained CHIEF models using four different WSI-level prediction tasks (that is, cancer cell detection, tumour origin identification, genomic profile characterization and survival outcome prediction). We conducted external validation using samples from 24 hospitals and study cohorts, including 5 collaborating medical centres worldwide (DFCI, BWH, MUV, SMCH and CUCH), 11 study cohorts from CPTAC (CPTAC-CCRCC, CPTAC-LUSC, CPTAC-PDA, CPTAC-CM, CPTAC-UCEC, CPTAC-HNSC, CPTAC-COAD, CPTAC-OV, CPTAC-GBM, CPTAC-LUAD and CPTAC-BRCA), 3 National Cancer Institute-sponsored study cohorts (PLCO-BRCA, PLCO-COADREAD and PLCO-LUAD) and 5 publicly available pathology image datasets (TissueNet, DROID-Breast, Dataset-PT, Diagset-B and PAIP). Below we present the detailed evaluation settings for these tasks.
Cancer cell detection task
We first evaluated the performance of CHIEF in detecting cancer cells in WSIs. We examined the performance of CHIEF on 11 primary cancer sites (endometrium, breast, oesophagus, stomach, prostate, cervix, colon, pancreas, lung, kidney and skin) with available data. These cancer types were represented by 13,661 WSIs from 15 datasets. We included nine publicly available datasets from large research consortia (that is, CPTAC-CCRCC, CPTAC-LUSC, CPTAC-PDA, CPTAC-CM, CPTAC-UCEC, TissueNet, Dataset-PT, DROID-Breast and Diagset-B) and six institutional datasets (that is, SMCH-Endo, SMCH-Cervix, CUCH-Sto, CUCH-Eso, CUCH-Colon and CUCH-Pros) from several hospitals as independent test sets to evaluate the robustness of our model. Below are the details of these sample sets.
We first obtained 9,686 publicly available WSIs from CPTAC, Diagset-B, Dataset-PT, DROID-Breast and TissueNet. Specifically, we included 3,712 WSIs from 5 working groups (kidney (CPTAC-CCRCC), lung (CPTAC-LUSC), pancreas (CPTAC-PDA), melanoma (CPTAC-CM), and endometrium (CPTAC-UCEC)) of the CPTAC. We further included pathology images of prostate, colon, breast and cervix cancer samples from Diagset-B (4,626 WSIs)31, Dataset-PT (498 WSIs)32, DROID-Breast (361 WSIs)58 and TissueNet (489 WSIs)33. Supplementary Table 13 summarizes the detailed descriptions of these patient cohorts.
To increase the diversity of our validation datasets, we further included 3,975 WSIs from 2 hospitals (that is, SMCH and CUCH). SMCH provided 2 datasets, SMCH-Endo (164 WSIs) and SMCH-Cervix (290 WSIs), from patients with endometrium and cervix cancer, respectively. CUCH provided 4 datasets (CUCH-Sto (550 WSIs), CUCH-Eso (385 WSIs), CUCH-Colon (1,742 WSIs) and CUCH-Pros (844 WSIs)) from patients with stomach, oesophagus, colon and prostate cancer.
Tumour origin identification task
We further examined the performance of CHIEF in identifying the primary sites of tumour origin using WSIs. We first used formalin-fixed paraffin-embedded slides from primary tumours in TCGA to fine-tune the CHIEF model for tumour origin prediction (Supplementary Table 14). We focused on pathology slides obtained from 18 anatomical sites to enhance comparability with a previous study59. After removing WSIs without magnification information in their metadata, we retained 9,432 slides, which were split into training, validation and held-out test sets in a ratio of 7:1:2. We processed the test set only after we finalized all model parameters.
To objectively evaluate our model’s generalizability, we used slides of primary tumours from CPTAC for independent validation. These slides represented nine types of primary cancer. After removing WSIs without magnification information, a total of 3,019 slides remained from CPTAC, which included 853 slides with lung cancers, 277 with endometrial cancers, 328 with BRCAs, 287 with head and neck cancers, 192 with colon cancers, 116 with ovarian cancers, 239 with gliomas, 331 with renal cancers, and 396 with pancreatic cancers.
Genomic profile prediction task
We next evaluated the performance of CHIEF in predicting genomic profiles using whole-slide pathology images. We focused on four clinically important prediction tasks: systematic prediction of prevalent genetic mutations across cancer types; identification of mutations related to targeted therapies; IDH status prediction for WHO classification of gliomas; and MSI prediction for immunotherapy administration in CRCs. We summarize each of these tasks and their implementation details below.
We used the TCGA dataset to train machine learning models for predicting prevalent genetic mutations across cancer types. For each cancer type, we selected the top five genes with the highest mutational prevalence for this prediction task. The TCGA training data included a total of 11,483 WSIs and covered 30 cancer types (Supplementary Table 17). In total, we investigated CHIEF’s capability of predicting the mutational status of 53 genes across these cancer types. We developed separate models for each mutation prediction task. To evaluate CHIEF models’ generalizability to patient populations not included in the model development process, we conducted independent validations using the CPTAC datasets, which contained 1,949 WSIs from 7 cancer types (Supplementary Table 19).
We used TCGA and CPTAC datasets as the training and independent test sets for predicting genetic mutations related to FDA-approved targeted therapies. Our training dataset included 6,013 WSIs (Supplementary Table 18), covering 15 cancer types and 18 genes related to targeted therapies. These genes included ALK, BRAF, BRCA1, BRCA2, EGFR, ERBB2, ESR1, EZH2, FGFR2, FGFR3, KRAS, MET, NTRK1, NTRK2, NTRK3, PIK3CA, RET and ROS1. Our independent test set contained 1,705 WSIs (Supplementary Table 20) and covered 6 cancer types and 14 different genes.
To predict IDH mutation status in patients with brain cancer from H&E-stained pathology images, we collected WSIs from three study cohorts: MUV41, HMS (with data from BWH and the Hospital of the University of Pennsylvania) and TCGA. We obtained samples from LGGs and GBMs. We stratified these samples by their histological grade to identify additional IDH-related morphological signals independent of histological grade. We trained our CHIEF model using TCGA cohorts (that is, TCGA-LGG with 842 WSIs and TCGA-GBM with 834 WSIs) and then externally evaluated the models using MUV and HMS cohorts (that is, MUV-LGG with 365 WSIs, HMS-LGG with 82 WSIs, MUV-GBM with 507 WSIs, and HMS-GBM with 88 WSIs; Supplementary Table 15).
MSI status in CRCs is a well-established predictor of responses to immune checkpoint blockade. To enable real-time MSI identification at the time of diagnosis, we used H&E-stained pathology specimens of CRCs for MSI prediction (Supplementary Table 16). We collected our training data from TCGA, which contained 437 WSIs (63 WSIs with MSI-high and 374 WSIs with MSI-low status), and we split this dataset into four folds for cross-validation. We further validated our models using independent patient cohorts from PAIP202042 and CPTAC-COAD, which contained 77 WSIs (19 WSIs with MSI-high and 58 WSIs with MSI-low status) and 221 WSIs (53 WSIs with MSI-high and 168 WSIs with MSI-low status), respectively.
Survival prediction task
Last, we evaluated the performance of CHIEF for predicting survival outcomes of patients with cancer. We conducted this analysis for seven cancer types with extensive survival information: COADREAD, LUSC, BRCA, GBM, endometrioid cancer (UCEC), LUAD and RCC. We collected 17 datasets, consisting of 9,404 WSIs from 6,464 patients (Supplementary Table 21). We used 7 publicly available TCGA cohorts with a total of 4,749 WSIs to train the model, and we used 4 publicly available CPTAC datasets (1,541 WSIs) and 6 institutional datasets (3,114 WSIs) for independent validations. The CPTAC datasets included in this analysis included CPTAC-GBM (244 WSIs), CPTAC-LUSC (292 WSIs), CPTAC-RCC (459 WSIs) and CPTAC-UCEC (546 WSIs). Three additional consortium datasets were obtained from the PLCO study60 (that is, PLCO-COADREAD with 333 WSIs, PLCO-LUAD with 176 WSIs, and PLCO-BRCA with 1,893 WSIs). We further included 3 institutional datasets collected from DFCI and BWH: the DFCI-BRCA (152 WSIs), DFCI-LUAD (486 WSIs) and BWH-RCC (74 WSIs) datasets. In this prediction task, we used all available overall survival data from CPTAC, PLCO-LUAD and PLCO-COADREAD and disease-specific survival information from all other datasets. Supplementary Table 21 summarizes the detailed demographic information for each patient cohort.
CHIEF fine-tuning details
We fine-tuned CHIEF models for various histopathological image analytical tasks. We fine-tuned these models by using CHIEF’s pretrained weights as the initial weights and added a task-specific fully connected layer. We implemented the tumour origin prediction task as an 18-class weakly supervised classification task by changing the prediction head into an 18-way classifier. We formulated genetic mutation predictions as two-class weakly supervised WSI classification tasks. We enabled prognostic predictions by appending a regression model as a head to CHIEF’s backbone, which output an estimated mortality risk score by a single neuron in the last layer of the neural network. For each dataset, we used the median value of the predicted risk scores to divide samples into longer-term and shorter-term survival groups. We then tested the difference between these two groups using the log-rank test. During fine-tuning, we set the mini-batch size to 1 for all tasks except the prognostic prediction task, which had a mini-batch size of 32 to increase efficiency. We fine-tuned models for all prediction tasks using the Adam optimizer with an initial learning rate of 0.0003. The learning rate is adjusted using the cosine annealing strategy. We fine-tuned all weakly supervised prediction tasks on one NVIDIA V100 32-GB GPU.
Model visualization
To enhance model interpretability, we visualized the prediction for each WSI by highlighting the image regions of relatively high importance in the prediction6. To generate fine-grained attention heat maps, we cropped WSIs into highly overlapped tiles (85% overlap ratio) and computed the attention scores for these tiles in each WSI. We scaled these scores between 0.0 (low attention) and 1.0 (high attention). To identify regions with high prediction confidence, we multiplied these attention scores with the prediction probability obtained from the instance-level classification branch. Finally, we overlaid the weighted attention score heat maps with their corresponding original H&E images. We used a transparency value of 0.5 for the heat maps to facilitate visualization of both the spatial distribution of attention scores and the associated pathology patterns. J.J., F.W., Y.P., C.R.J., J.A.G. and M.P.N. independently evaluated the highlighted regions from the heat maps. To objectively compare model attention and regions occupied by cancer cells, J.J. and F.W. annotated the pixel-level ground truth of cancerous regions independently without viewing the model output.
Comparative analysis
In the cancer detection task, we compared CHIEF with three state-of-the-art weakly supervised WSI classification methods: CLAM6, ABMIL34 and DSMIL35. We reproduced these three baseline methods using their officially released codes. We used the same pretraining data to train our CHIEF model and these alternative methods to ensure the comparability of our results. In the tumour origin identification task, we implemented three baseline methods (that is, TOAD59, TransMIL61 and DSMIL35) using their released codes and compared them with CHIEF. In the genetic profile prediction tasks, we compared CHIEF with PC-CHiP36. As PC-CHiP used the same training data for genetic mutation prediction, we directly compared our results with the reported performance of this baseline method. In the patient prognosis prediction task, we compared CHIEF, DSMIL35 and the histopathology branch of PORPOISE12 to ensure fair comparisons. We further compared CHIEF with other recently released foundation models in comparable tasks. The Supplementary Information includes the detailed methods for these comparisons. To simplify the result presentation, we reported the absolute percentage point differences of AUROCs in all comparisons.
Inclusion and ethics statement
The research included local researchers throughout the research process. The research is determined locally relevant in all of our research sites. The roles and responsibilities were agreed on among collaborators ahead of the research, and capacity-building plans for local researchers were discussed. The research does not result in stigmatization, incrimination, discrimination or otherwise personal risk to participants. The research does not involve health, safety, security or other risks to researchers. Benefit-sharing measures have been discussed. We have taken local and regional research relevant to this study into account in citations.
Ethics oversight
The study was approved by the institutional review boards of Harvard Medical School (IRB20-1509), the Ethical Review Committee of CUCH (CZLS2023279-A), the Ethics Committee of SMCH (SFYLS [2021]045) and the Ethics Committee of Yantai Yuhuangding Hospital (2023-328). Informed consent for this project was waived by the respective ethics commissions because this study only involves a retrospective anonymized analysis of archival pathology samples.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
This work utilized 16 pathology datasets from large research consortia, including TCGA (https://portal.gdc.cancer.gov), GTEx (https://www.gtexportal.org/home/), PAIP (http://www.wisepaip.org/paip), PANDA (https://www.kaggle.com/c/prostate-cancer-grade-assessment), BCC (https://datahub.aida.scilifelab.se/10.23698/aida/bccc), ACROBAT (https://doi.org/10.48723/w728-p041), BCNB (https://bcnb.grand-challenge.org/), TOC (https://www.cancerimagingarchive.net/collection/ovarian-bevacizumab-response/), CPTAC (https://portal.gdc.cancer.gov), DROID-Breast (https://datahub.aida.scilifelab.se/10.23698/aida/drbr), Dataset-PT (https://github.com/CSU-BME/pathology_SSL), Diagset-B (https://github.com/michalkoziarski/DiagSet), MUV (https://doi.org/10.25493/WQ48-ZGX) and PLCO (https://cdas.cancer.gov/plco/). The other two datasets, PAIP2020 and TissueNet, can be requested from the respective data science challenge organizers: PAIP2020 (https://paip2020.grand-challenge.org/) and TissueNet (https://www.drivendata.org/competitions/67/competition-cervical-biopsy/). Supplementary Table 22 provides the links to the raw data from these sources. We obtained institutional data for CHIEF pretraining and validation from DFCI, BWH, YH, SMCH, CUCH and the Hospital of the University of Pennsylvania. These data are not publicly available owing to patient privacy obligations and institutional review board and data use agreement requirements. Researchers may obtain de-identified data directly from DFCI, BWH, YH, SMCH, CUCH and the Hospital of the University of Pennsylvania by reasonable request and subject to institutional ethical approvals. Data access enquiries should be directed to K.-H.Y. We aim to forward all requests to the managers of these institutional datasets in 2 weeks, and these requests will be evaluated according to their institutional policies. Data are strictly only for non-commercial academic use. This study relies on retrospective analysis of anonymized pathology slides. Source data are provided with this paper.
Code availability
All code was implemented in Python using PyTorch as the primary deep learning package. The source codes for CHIEF are available at https://github.com/hms-dbmi/CHIEF. Our Docker images are available at https://hub.docker.com/r/chiefcontainer/chief.
References
Van der Laak, J., Litjens, G. & Ciompi, F. Deep learning in histopathology: the path to the clinic. Nat. Med. 27, 775–784 (2021).
Shmatko, A., Ghaffari Laleh, N., Gerstung, M. & Kather, J. N. Artificial intelligence in histopathology: enhancing cancer research and clinical oncology. Nat. Cancer 3, 1026–1038 (2022).
Song, A. H. et al. Artificial intelligence for digital and computational pathology. Nat. Rev. Bioeng. 1, 930–949 (2023).
Campanella, G. et al. Clinical-grade computational pathology using weakly supervised deep learning on whole slide images. Nat. Med. 25, 1301–1309 (2019).
Bejnordi, B. E. et al. Diagnostic assessment of deep learning algorithms for detection of lymph node metastases in women with breast cancer. JAMA 318, 2199–2210 (2017).
Lu, M. Y. et al. Data-efficient and weakly supervised computational pathology on whole-slide images. Nat. Biomed. Eng. 5, 555–570 (2021).
Coudray, N. et al. Classification and mutation prediction from non–small cell lung cancer histopathology images using deep learning. Nat. Med. 24, 1559–1567 (2018).
Nasrallah, M. P. et al. Machine learning for cryosection pathology predicts the 2021 WHO classification of glioma. Med 4, 526–540 (2023).
Tsai, P.-C. et al. Histopathology images predict multi-omics aberrations and prognoses in colorectal cancer patients. Nat. Commun. 14, 2102 (2023).
Yu, K.-H. et al. Classifying non-small cell lung cancer types and transcriptomic subtypes using convolutional neural networks. J. Am. Med. Inform. Assoc. 27, 757–769 (2020).
Yu, K.-H. et al. Association of omics features with histopathology patterns in lung adenocarcinoma. Cell Syst. 5, 620–627 (2017).
Chen, R. J. et al. Pan-cancer integrative histology-genomic analysis via multimodal deep learning. Cancer Cell 40, 865–878 (2022).
Marostica, E. et al. Development of a histopathology informatics pipeline for classification and prediction of clinical outcomes in subtypes of renal cell carcinoma. Clin. Cancer Res. 27, 2868–2878 (2021).
Yu, K.-H. et al. Predicting non-small cell lung cancer prognosis by fully automated microscopic pathology image features. Nat. Commun. 7, 12474 (2016).
Vanguri, R. S. et al. Multimodal integration of radiology, pathology and genomics for prediction of response to PD-(L)1 blockade in patients with non-small cell lung cancer. Nat. Cancer 3, 1151–1164 (2022).
Yu, K.-H. et al. Deciphering serous ovarian carcinoma histopathology and platinum response by convolutional neural networks. BMC Med. 18, 236 (2020).
Foersch, S. et al. Multistain deep learning for prediction of prognosis and therapy response in colorectal cancer. Nat. Med. 29, 430–439 (2023).
Kather, J. N. et al. Pan-cancer image-based detection of clinically actionable genetic alterations. Nat. Cancer 1, 789–799 (2020).
Echle, A. et al. Deep learning in cancer pathology: a new generation of clinical biomarkers. Br. J. Cancer 124, 686–696 (2021).
Ektefaie, Y. et al. Integrative multiomics-histopathology analysis for breast cancer classification. NPJ Breast Cancer 7, 147 (2021).
Yu, K.-H., Beam, A. L. & Kohane, I. S. Artificial intelligence in healthcare. Nat. Biomed. Eng. 2, 719–731 (2018).
Krishnan, R., Rajpurkar, P. & Topol, E. J. Self-supervised learning in medicine and healthcare. Nat. Biomed. Eng. 6, 1346–1352 (2022).
Zhou, Y. et al. A foundation model for generalizable disease detection from retinal images. Nature 622, 156–1638 (2023).
Chen, C. et al. Fast and scalable search of whole-slide images via self-supervised deep learning. Nat. Biomed. Eng. 6, 1420–1434 (2022).
Wang, X. et al. RetCCL: clustering-guided contrastive learning for whole-slide image retrieval. Med. Image Anal. 83, 102645 (2023).
Chen, R. J. et al. Towards a general-purpose foundation model for computational pathology. Nat. Med. 30, 850–862 (2024).
Wagner, S. J. et al. Transformer-based biomarker prediction from colorectal cancer histology: a large-scale multicentric study. Cancer Cell 41, 1650–1661 (2023).
Huang, Z., Bianchi, F., Yuksekgonul, M., Montine, T. J. & Zou, J. A visual–language foundation model for pathology image analysis using medical twitter. Nat. Med. 29, 2307–2316 (2023).
Lu, M. Y. et al. A visual-language foundation model for computational pathology. Nat. Med. 30, 863–874 (2024).
Wang, X. et al. Transformer-based unsupervised contrastive learning for histopathological image classification. Med. Image Anal. 81, 102559 (2022).
Koziarski, M. et al. Diagset: a dataset for prostate cancer histopathological image classification. Sci. Rep. 14, 6780 (2024).
Yu, G. et al. Accurate recognition of colorectal cancer with semi-supervised deep learning on pathological images. Nat. Commun. 12, 6311 (2021).
Loménie, N. et al. Can AI predict epithelial lesion categories via automated analysis of cervical biopsies: the TissueNet challenge? J. Pathol. Inform. 13, 100149 (2022).
Ilse, M., Tomczak, J. & Welling, M. Attention-based deep multiple instance learning. In Proc. 35th International Conference on Machine Learning (eds Dy, J. & Krause, A.) 2127–2136 (PMLR, 2018).
Li, B., Li, Y. & Eliceiri, K. W. Dual-stream multiple instance learning network for whole slide image classification with self-supervised contrastive learning. In Proc. 2021 IEEE/CVF Conference on Computer Vision and Pattern Recognition 14313–14323 (IEEE, 2021).
Fu, Y. et al. Pan-cancer computational histopathology reveals mutations, tumor composition and prognosis. Nat. Cancer 1, 800–810 (2020).
Petrini, I. et al. A specific missense mutation in GTF2I occurs at high frequency in thymic epithelial tumors. Nat. Genet. 46, 844–849 (2014).
Carbone, M. et al. Biological mechanisms and clinical significance of BAP1 mutations in human cancer. Cancer Discov. 10, 1103–1120 (2020).
Chakravarty, D. et al. OncoKB: a precision oncology knowledge base. JCO Precision Oncology 1, 1–16 (2017).
Louis, D. N. et al. The 2021 WHO Classification of Tumors of the Central Nervous System: a summary. Neuro-Oncology 23, 1231–1251 (2021).
Roetzer-Pejrimovsky, T. et al. The Digital Brain Tumour Atlas, an open histopathology resource. Sci. Data 9, 55 (2022).
Kim, K. et al. PAIP 2020: microsatellite instability prediction in colorectal cancer. Med. Image Anal. 89, 102886 (2023).
Amin, M. B. et al. The Eighth Edition AJCC Cancer Staging Manual: continuing to build a bridge from a population-based to a more “personalized” approach to cancer staging. CA Cancer J. Clin. 67, 93–99 (2017).
Achiam, J. et al. GPT-4 technical report. Preprint at https://doi.org/10.48550/arXiv.2303.08774 (2023).
Team, G. et al. Gemini: a family of highly capable multimodal models. Preprint at https://doi.org/10.48550/arXiv.2312.11805 (2023).
Azizi, S. et al. Robust and data-efficient generalization of self-supervised machine learning for diagnostic imaging. Nat. Biomed. Eng. 7, 756–779 (2023).
Cancer Genome Atlas Research Network, J. et al. The Cancer Genome Atlas Pan-Cancer analysis project. Nat. Genet. 45, 1113–1120 (2013).
Lonsdale, J. et al. The genotype-tissue expression (GTEx) project. Nat. Genet. 45, 580–585 (2013).
Bulten, W. et al. Artificial intelligence for diagnosis and Gleason grading of prostate cancer: the PANDA challenge. Nat. Med. 28, 154–163 (2022).
Yacob, F. et al. Weakly supervised detection and classification of basal cell carcinoma using graph-transformer on whole slide images. Sci Rep. 13, 7555 (2023).
Xu, F. et al. Predicting axillary lymph node metastasis in early breast cancer using deep learning on primary tumor biopsy slides. Front. Oncol. 11, 4133 (2021).
Weitz, P. et al. A multi-stain breast cancer histological whole-slide-image data set from routine diagnostics. Sci. Data 10, 562 (2023).
Wang, C.-W. et al. Histopathological whole slide image dataset for classification of treatment effectiveness to ovarian cancer. Sci. Data 9, 25 (2022).
Radford, A. et al. Learning transferable visual models from natural language supervision. In Proc. 38th International Conference on Machine Learning (eds Meila, M. & Zhang, T.) 8748–8763 (PMLR, 2021).
Otsu, N. A threshold selection method from gray-level histograms. IEEE Trans. Syst. Man Cybern. Syst. 9, 62–66 (1979).
Kingma, D. P. & Ba, J. L. Adam: a method for stochastic optimization. In Proc. 3rd International Conference on Learning Representations (eds Bengio, Y. & LeCun, Y.) (ICLR, 2015).
Loshchilov, I. & Hutter, F. SGDR: stochastic gradient descent with warm restarts. In Proc. 5th International Conference on Learning Representations 1769–1784 (ICLR, 2017).
Stadler, C. B. et al. Proactive construction of an annotated imaging database for artificial intelligence training. J. Digit. Imaging 34, 105–115 (2021).
Lu, M. Y. et al. AI-based pathology predicts origins for cancers of unknown primary. Nature 594, 106–110 (2021).
Black, A. et al. PLCO: evolution of an epidemiologic resource and opportunities for future studies. Rev. Recent Clin. Trials 10, 238–245 (2015).
Shao, Z. et al. TransMIL: Transformer based correlated multiple instance learning for whole slide image classification. Adv. Neural Inf. Process. Syst. 34, 2136–2147 (2021).
Liang, J. et al. Deep learning supported discovery of biomarkers for clinical prognosis of liver cancer. Nat. Mach. Intell. 5, 408–420 (2023).
Courtiol, P. et al. Deep learning-based classification of mesothelioma improves prediction of patient outcome. Nat. Med. 25, 1519–1525 (2019).
Graham, S. et al. Hover-Net: simultaneous segmentation and classification of nuclei in multi-tissue histology images. Med. Image Anal. 58, 101563 (2019).
Acknowledgements
We thank C. Burroughs, M. Kapanadze and F. McDonald for administrative support; and the AWS Cloud Credits for Research programme, the Microsoft Azure for Research Award programme, the NVIDIA GPU Grant Program and the Extreme Science and Engineering Discovery Environment at the Pittsburgh Supercomputing Center (allocation TGBCS180016) for computational support. K.-H.Y. is in part supported by the National Institute of General Medical Sciences grant R35GM142879, the Department of Defense Peer Reviewed Cancer Research Program Career Development Award HT9425-231-0523, the Research Scholar Grant RSG-24-1253761-01-ESED (grant DOI: https://doi.org/10.53354/ACS.RSG-24-1253761-01-ESED.pc.gr.193749) from the American Cancer Society, a Google Research Scholar Award, the Harvard Medical School Dean’s Innovation Award and the Blavatnik Center for Computational Biomedicine Award. K.L.L. is in part supported by the National Institutes of Health award P50CA165962 and the 3000 Miles to the Cure Foundation. The PAIP data were provided by the Seoul National University Hospital and funded by the Ministry of Health and Welfare, Republic of Korea (grant number HI18C0316).
Author information
Authors and Affiliations
Contributions
X.W., J. Zhao, S.Y. and K.-H.Y. conceived and designed the study. J. Zhao, E.M., D.D., N.U.L., L.S., T.D., D.M., K.L.L., S.S., S.O., J.A.G., M.P.N., K.-H.Y., F.W., H.T., Jing Zhang, K.W. and Y.L. curated the data from their respective institutes. X.W., J. Zhao, S.Y., W.Y., Jiayu Zhang and K.-H.Y. developed, validated and evaluated the models. J.J., F.W., K.W., Y.L., Y.P., J. Zhu, C.R.J., J.A.G., M.P.N. and K.-H.Y. interpreted the pathological images. Jun Zhang, Jing Zhang, X.H. and R.L. contributed to the technical discussion. X.W., J. Zhao, E.M., C.R.J., J.A.G, J.J., F.W., S.Y. and K.-H.Y interpreted the analytical results. X.W., J. Zhao, S.Y. and K.-H.Y wrote the manuscript. All authors contributed to the edits of the manuscript. K.-H.Y supervised the project.
Corresponding authors
Ethics declarations
Competing interests
Jun Zhang and X.H. were employees of Tencent AI Lab. K.-H.Y. is an inventor on US patent 16/179,101 (patent assigned to Harvard University) and was a consultant for Curatio.DL (not related to this work). K.L.L. was a consultant for Travera, BMS, Servier, Integragen, LEK and Blaze Bioscience, received equity from Travera, and has research funding from BMS and Lilly (not related to this work). C.R.J is an inventor on US patent applications 17/073,123 and 63/528,496 (patents assigned to Dartmouth Hitchcock Medical Center and ViewsML) and is a consultant and CSO for ViewsML, none of which is related to this work.
Peer review
Peer review information
Nature thanks the anonymous reviewers for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 CHIEF accurately identified the origins of tumors, with results validated in independent patient cohorts from the Clinical Proteomic Tumor Analysis Consortium (CPTAC).
a. The confusion matrix of CHIEF’s prediction in the held-out test sets. The overall macro-averaged accuracy of CHIEF is 0.895. b. CHIEF achieved high prediction performance and generalizability to independent cohorts in tumor origin prediction (AUROC = 0.9853 ± 0.0245). Micro-averaged one-versus-rest ROC curves for tumor origin classification are shown. We presented the AUROC±s.d. calculated across 18 tumor origins. In comparison, state-of-the-art methods have substantially lower performance in the independent cohorts (two-sided Wilcoxon signed-rank test P-value = 0.000015). c. CHIEF attained higher accuracy than state-of-the-art deep learning methods in tumor origin prediction. Overall accuracies for the held-out (n = 1,895) and independent test sets (n = 3,019) for CHIEF and other deep learning methods are shown. d. CHIEF attained higher AUROC, sensitivity, and specificity for each tumor origin in the held-out test sets (n = 1,895) compared with other methods. The model performance for all 18 tumor origins is shown. e. CHIEF possessed significantly higher AUROC, sensitivity, and specificity for each origin in the independent test sets (n = 3,019, P-value = 0.003906, two-sided Wilcoxon signed-rank test). In contrast, standard machine learning approaches suffer from substantial performance drops when applied to patient cohorts not involved in model development. In c-e, error bars represent 95% confidence intervals computed by the bootstrap method (n = 1,000 replicates), and the centers represent the values of various performance metrics specified in these figure panels. The detailed sample size for each cancer type shown in d-e can be found in Supplementary Table 14.
Extended Data Fig. 2 Visualization of model attention scores showed CHIEF accurately identified cancerous regions of melanoma, lung, and kidney cancers.
For each cancer type, the left image panel represented the ground truth annotations labeled by experienced pathologists. Because CHIEF employs a weakly supervised approach that only requires slide-level annotations, these region-level annotations were not used during the training phase. The middle panel visualized the amount of attention CHIEF paid to each region in the WSIs. The right panel showed the zoomed-in view of regions receiving high (image tiles with red outlines) and low (image tiles with black outlines) attention scores. The original WSIs and their corresponding heatmaps are available at https://yulab.hms.harvard.edu/projects/CHIEF/CHIEF.htm.
Extended Data Fig. 3 Detailed genetic mutation prediction results organized by cancer types.
Prediction performance of prevalent genetic mutations (n = 11,483) and targeted-therapy-associated genetic mutations (n = 6,013) is shown. The detailed sample counts for each genetic mutation are available in Supplementary Tables 17, 18. CHIEF predicted several prevalent mutations (e.g., TP53 in ACC, LGG, and UCEC) with AUROCs > 0.80. The mean ± 95% confidence interval is shown for each prediction task. Error bars represent the 95% confidence intervals estimated by 5-fold cross-validation (5 independent runs).
Extended Data Fig. 4 CHIEF attained a high performance in predicting genetic mutation status from histopathology images across cancer types.
Prediction performance in the held-out test set (TCGA) and independent test set (CPTAC) were shown side by side. These results were grouped by the genes to highlight the prediction performance of the same genes across cancer types. The red and blue horizontal lines represent the average AUROCs in the held-out and independent test sets, respectively. Top, CHIEF’s performance in predicting mutation status for frequently mutated genes across cancer types. Supplementary Tables 17 and 19 show the detailed sample count for each cancer type. Bottom, CHIEF’s performance in predicting genetic mutation status related to FDA-approved targeted therapies. Supplementary Tables 18 and 20 show the detailed sample count for each cancer type. In a and b, results are presented as mean ± 95% confidence interval. Error bars represent the 95% confidence intervals estimated by 5-fold cross-validation.
Extended Data Fig. 5 CHIEF predicted IDH status of glioma samples in several patient cohorts.
CHIEF classified glioma samples with and without IDH mutation. Here, we showed that CHIEF successfully predicted IDH mutation status in both high and low histological grade groups defined by conventional visual-based histopathology assessment. a. Regions with increased cellularity and perinuclear halos received high model attention in IDH-mutant samples, while regions showing poorer cell adhesion received high attention in IDH-wildtype slides. We used samples from the MUV-GBM dataset as an example for this visualization. The bottom figures show the corresponding image tiles. Six experienced pathologists (see Methods) examined these tiles independently and annotated the morphological patterns correlated with regions receiving high and low attention. b. IDH-mutant gliomas from the six cohorts exhibit a similar bi-modal distribution along the attention score axis. In contrast, IDH-wildtype gliomas display an unimodal distribution with mostly low-attention image regions. We normalized the attention scores to a range from 0 to 1, representing the importance of each image tile to the prediction output by CHIEF. These analyses included samples from TCGA-GBM (n = 834), MUV-GBM (n = 507), HMS-GBM (n = 88), TCGA-LGG (n = 842), MUV-LGG (n = 365), and HMS-LGG (n = 82). In these violin plots, the central white dots represent the median, the thick black bars indicate the interquartile range (IQR), and the thin black lines (whiskers) extend to 1.5 times the IQR from the first and third quartiles. The width of the violin represents the density of data at different values.
Extended Data Fig. 6 CHIEF predicted MSI status in several colorectal cancer patient cohorts.
a. Solid tumor regions of MSI-high samples received high attention scores, while adjacent benign mucosal epithelium regions received low attention scores. In MSI-low samples, most regions received low attention scores. Example images from the PAIP2020 dataset were shown in this visualization. The bottom portion of this figure panel showed image tiles receiving high and low attention scores. Malignant regions were highly attended in both MSI-low and MSI-high samples. Solid tumors, intraluminal and extraluminal mucin, and signet ring cells received high attention in MSI-high samples. In MSI-low samples, infiltrative malignant glands interfacing with fibroblasts, luminal necrosis, and lymphocytic infiltrates received relatively high attention. Adjacent benign colonic epithelium receives low attention in both MSI-high and MSI-low patients. b. CHIEF paid high levels of attention to 30% of regions in MSI-high samples, while more regions in MSI-low samples received low attention scores. Attention score distributions of the three patient cohorts (n = 437 in TCGA-COADREAD, n = 77 in PAIP2020, and n = 221 in CPTAC-COAD) are shown. In these violin plots, the central white dots represent the median, the thick black bars indicate the interquartile range (IQR), and the thin black lines (whisker) extend to 1.5 times the IQR from the first and third quartiles. The width of the violin represents the density of data at different values.
Extended Data Fig. 7 Survival prediction results for patients with all stages.
Previous methods pooled patients with all stages in their survival outcome prediction12,62,63. To facilitate comparisons with these previous reports, we compared CHIEF with baseline methods in this study setting, using 9,404 whole slide images from 6,464 patients. CHIEF attained substantially better survival prediction performance (unadjusted two-sided log-rank test P-value < 0.05 in all patient cohorts under study) and distinguished patients with different survival outcomes using histopathology images alone. Supplementary Fig. 5 shows results from two baseline methods (PORPOISE and DSMIL). Error bands represent 95% confidence intervals.
Extended Data Fig. 8 Visualization of model attention showed regions of importance in survival prediction for lung cancer patients.
In patients with shorter-term survival, CHIEF paid high levels of attention to lesional regions with high tumor cellularity and strands of fibrosis in lung adenocarcinoma, tumor budding in squamous cell carcinoma, and necrotic regions in both types of lung cancers. In contrast, highly attended regions in patients with lower mortality risks highlighted dyskeratosis in lung squamous cell carcinoma. The original WSIs and their corresponding heatmaps are available at https://yulab.hms.harvard.edu/projects/CHIEF/CHIEF_survival.htm.
Extended Data Fig. 9 Quantitative analyses of regions receiving high attention revealed pathology microenvironments predictive of molecular profiles and survival outcomes.
For each WSI, we selected the top 1% of patches with the highest attention from CHIEF at 40× magnification. We excluded WSIs with fewer than 100 image patches. We employed Hover-Net64 trained with pathologists’ annotations in the PanNuke dataset (including tumor cells, lymphocytes, stromal cells, necrotic cells, and epithelial cells) for cell segmentation and classification. We compared the cell type compositions across different patient groups. a. Colorectal cancer samples with MSI-high status have significantly more tumor-infiltrating lymphocytes in the high-attention regions (unadjusted two-sided Mann-Whitney U test P-value = 0.00052 in PAIP2020, P-value = 0.00016 in CPTAC-COAD). b. IDH wild-type glioma samples have significantly more necrotic cells (unadjusted two-sided Mann-Whitney U test P-value = 0.00006 in TCGA-GBM and P-value = 0.000001 in TCGA-LGG). c. Samples from longer-term colorectal cancer survivors have a larger number of stromal cells, more tumor-infiltrating lymphocytes, and fewer tumor cells in the high-attention regions, compared with those with shorter-term survival. Samples from shorter-term lung squamous cell carcinoma survivors have a larger fraction of tumor cells and smaller fractions of lymphocytes and epithelial cells in the high-attention regions, compared with those with longer-term survival. These analyses included samples from PAIP2020 (n = 77), CPTAC-COAD (n = 221), TCGA-GBM (n = 825), TCGA-LGG (n = 834), TCGA-COADREAD (n = 520), and TCGA-LUSC (n = 400). In these box plots, the central lines indicate the median, box bounds are the 25th and 75th percentiles, and whiskers extend to 1.5 times the interquartile range. In these figures, one star (*), two stars (**), three stars (***), and four stars (****) represent P-value < 0.05, P-value < 0.01, P-value < 0.001, and P-value < 0.0001, respectively.
Supplementary information
Supplementary Information
Descriptions of additional results, including Supplementary Figs. 1–14 and Tables 1–25, and methods.
Source data
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Wang, X., Zhao, J., Marostica, E. et al. A pathology foundation model for cancer diagnosis and prognosis prediction. Nature (2024). https://doi.org/10.1038/s41586-024-07894-z
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41586-024-07894-z
- Springer Nature Limited