Abstract
Entomopathogenic nematodes are widely used as biopesticides1,2. Their insecticidal activity depends on symbiotic bacteria such as Photorhabdus luminescens, which produces toxin complex (Tc) toxins as major virulence factors3,4,5,6. No protein receptors are known for any Tc toxins, which limits our understanding of their specificity and pathogenesis. Here we use genome-wide CRISPR–Cas9-mediated knockout screening in Drosophila melanogaster S2R+ cells and identify Visgun (Vsg) as a receptor for an archetypal P. luminescens Tc toxin (pTc). The toxin recognizes the extracellular O-glycosylated mucin-like domain of Vsg that contains high-density repeats of proline, threonine and serine (HD-PTS). Vsg orthologues in mosquitoes and beetles contain HD-PTS and can function as pTc receptors, whereas orthologues without HD-PTS, such as moth and human versions, are not pTc receptors. Vsg is expressed in immune cells, including haemocytes and fat body cells. Haemocytes from Vsg knockout Drosophila are resistant to pTc and maintain phagocytosis in the presence of pTc, and their sensitivity to pTc is restored through the transgenic expression of mosquito Vsg. Last, Vsg knockout Drosophila show reduced bacterial loads and lethality from P. luminescens infection. Our findings identify a proteinaceous Tc toxin receptor, reveal how Tc toxins contribute to P. luminescens pathogenesis, and establish a genome-wide CRISPR screening approach for investigating insecticidal toxins and pathogens.
Similar content being viewed by others
Main
Tc toxin family members were initially identified in P. luminescens3. These bacteria live in the gut of entomopathogenic nematodes, which search for and colonize a wide range of insect larvae in the soil6,7. Once the nematode enters the larval body cavity, P. luminescens is released by regurgitation to repress the immune system of the insect, proliferate and subsequently kill the larvae. After converting the interior of the insect cadaver into P. luminescens biomass, the bacterium itself then serves as the food source for the nematode in a mutualistic relationship6,7. Among an array of toxins and virulence factors released by P. luminescens, Tc toxins have a key role3,6. Growing numbers of Tc and Tc-like toxins have been identified in both entomopathogenic bacteria and human pathogens4,8,9,10,11.
Tc toxins are extremely large protein complexes formed by three subunits named TcA, TcB and TcC (also known as A, B and C subunits, respectively)4,6,12. TcA assembles into a homopentamer with a total molecular weight of around 1.4 MDa8,13,14,15, and is responsible for receptor binding and membrane translocation of the cargo protein. TcB and TcC together form an approximately 250 kDa cocoon, with the cytotoxic carboxy-terminal hypervariable region (HVR, about 30 kDa) of TcC autoproteolytically cleaved and encapsulated inside this cocoon14,16,17. The TcA pentamer and the cocoon assemble into a heterotrimeric holotoxin, with the HVR enzyme as the cargo protein pre-loaded into the TcA translocation channel17.
Tc toxins target and enter cells through receptor-mediated endocytosis. The low pH in endosomes induces conformational changes in TcA that lead to the insertion of the α-helical channel into the endosomal membrane like the needle of a syringe4,13,14,18,19. Subsequently, the HVR translocates through this channel into the cytosol of cells17,18, where it covalently modifies its cellular targets. For instance, TccC3 and TccC5, two TcC units found in P. luminescens, contain HVRs that encode ADP-ribosyl transferases. TccC3 causes cytoskeletal clustering through the ADP-ribosylation of actin, whereas TccC5 targets and modifies small Rho GTPases20.
Our understanding of the cell tropism and species selectivity of Tc toxins remains limited. Recent studies that used glycan array screening revealed that negatively charged sulfated glycosaminoglycans (sGAGs), such as heparins and heparan sulfates, and N-linked glycans act as common receptors for various Tc toxins21. In addition, TcdA1 from the P. luminescens subsp. akhurstii strain recognizes various Lewis oligosaccharides, albeit with low affinity21,22, and the Tc toxin from Yersinia entomophaga can interact broadly with various glycans15. The contribution of N-linked glycans and sGAGs has been further confirmed using CRISPR–Cas9-mediated genetic screens in human cell lines23.
Genome-wide CRISPR–Cas9-mediated knockout (KO) screening in mammalian cells is a powerful approach to identify host factors targeted by pathogens and toxins24,25,26. However, it has been challenging to perform similar screens on insect cells because the delivery of genome-wide libraries encoding single-guide RNAs (sgRNAs) relies on lentiviral transduction, a technique that is inefficient in insect cells. We recently overcame this obstacle by developing plasmid-mediated transfection of sgRNA libraries that achieve stable integration through site-specific recombination. This enabled genome-wide CRISPR–Cas9-mediated KO screening in D. melanogaster S2R+ (S2) cells27. Here, we utilize this approach to carry out genome-wide screens for an archetypal P. luminescens Tc toxin (designated pTc; also known as PTC3), and identify the mucin-like protein Vsg as its receptor.
Genome-wide CRISPR screens in S2 cells
pTc is assembled from one set of TcA (TcdA1), TcB (TcdB2) and TcC (TccC3) subunits from P. luminescens20. The HVR of TccC3 ADP-ribosylates actin, which causes actin to cluster and leads to cell rounding and eventual cell death in mammalian cells20 (Extended Data Fig. 1a). The sensitivity of three human cell lines, HeLa (a cervical carcinoma cell line), U2OS (a bone osteosarcoma cell line) and 5637 (a bladder carcinoma cell line), to pTc was assessed. Toxin concentrations that induced 50% of the cells to become round (defined as CR50) ranged from 1 to 5 nM (Extended Data Fig. 1a,b). In contrast to mammalian cells, actin clustering did not result in cell death in S2 cells but disrupted cell division, which led to enlarged multinucleated cells (Extended Data Fig. 1c). A toxin concentration of about 5 pM induced 50% of the cells to become enlarged (defined as CE50) (Extended Data Fig. 1d).
S2 cells stably expressing Cas9 endonuclease were transfected with a genome-wide library of sgRNA plasmids that targeted 13,685 D. melanogaster genes (6–8 sgRNA per gene, 84,563 sgRNA in total). A plasmid encoding phiC31 integrase was co-transfected to induce site-specific recombination (Fig. 1a). Cells were then treated with pTc, and resistant cells were separated from enlarged cells using cell strainers (30-µm filtration size), which allowed normal S2 cells but not enlarged ones to pass through. Following selection, resistant cells were collected, and target genes were identified through next-generation sequencing (Fig. 1a).
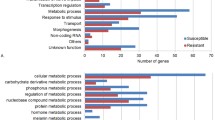
a, Schematic of the pooled screen using pTc or mTc toxins in the D. melanogaster PT5/S2R+ cell line. b, Genes identified in two independent pTc screens (screen 1-high and screen 2 as described in the Methods) ranked on the basis of the robust rank aggregation (RRA) score (analysed with MAGeCK)41. The top-ranking genes are colour-coded and grouped. c, Comparison of the genes identified in the pTc and mTc screens.
Multiple independent screens were carried out for pTc, which produced overlapping top hits (Fig. 1b and Extended Data Fig. 2a,b). The strongest hit was the gene encoding Vsg, a small single-pass membrane protein. Other top hits included genes encoding subunits of protein complexes that are involved in endosomal trafficking, such as ESCRT-0, HOPS and CORVET (Fig. 1b). We then utilized RNA interference (RNAi) to evaluate 29 genes within the top hits (Extended Data Fig. 3a). The sensitivity of S2 cells to pTc was estimated by the measuring the cell size or the Hoechst signal using flow cytometry (Extended Data Fig. 3b–e). Many of the top hits were validated, as knockdown of them reduced S2 cell sensitivity to pTc.
We also carried out a parallel screen with another Tc toxin (designated mTc), which was assembled with the same TcdB2 and TccC3 units as in pTc but with a distinct TcA (TcdA4 from the human pathogen Morganella morganii)8. The top hits from the mTc screen included sugarless (sgl; Fig. 1c and Extended Data Fig. 2a,b), which encodes a homologue of human UDP-glucose 6-dehydrogenase that is involved in sGAG biosynthesis28. Other top hits were genes that encode components of the PIKfyve complex, which is associated with endosomal trafficking. The top hits were clearly different between pTc and mTc (Fig. 1c), which indicated that different TcA subunits utilize distinct receptors.
Vsg mediates the binding and entry of pTc
We next focused on the top hit, Vsg, as a candidate receptor for pTc. Three stable vsg KO S2 cell lines were generated using the CRISPR–Cas9 approach (Extended Data Fig. 4a–c). They showed reductions in sensitivity (CE50) to pTc compared with S2 cells that expressed scrambled sgRNAs (Fig. 2a,b), and they did not show changes in cell size over 5 days after exposure to pTc (30 pM). By contrast, S2 cells that expressed scrambled sgRNAs showed enlarging cell size over time (Fig. 2c). Consistently, S2 cells that expressed scrambled sgRNAs showed strong ADP-ribosylation of actin after exposure to pTc20,29, whereas vsg KO cells showed much reduced ADP-ribosylation of actin (Fig. 2d). We further tested two additional Tc toxins. Both contained the same TcdB2 and TccC3 units as in pTc but had a distinct TcA subunit (XptA1 from Xenorhabdus nematophila or TcdA4 from P. luminescens). Both toxins induced cell enlargement in vsg KO cells and showed no difference in potency in control cells compared with vsg KO cells (Extended Data Fig. 4d).
a,b, The sensitivity to pTc of vsg KO S2 cell lines or control S2 cell lines expressing scrambled sgRNAs was assessed by exposure to a titration of pTc for 4 days. a, Representative images of cells after 15.6 pM pTc treatment. Cell nuclei were marked with Hoechst dye (blue). DIC, differential interference contrast. b, The percentage of enlarged cells was quantified and plotted over pTc concentrations. c, Cell sizes were recorded and plotted over time after exposure to pTc (30 pM) for vsg KO S2 cells compared with control S2 cells expressing a scrambled sgRNA. d, Cells were incubated with pTc for 24 h, and ADP-ribosylation of actin was detected by immunoblot analysis of cell lysates. Tubulin served as a loading control. For gel source data, see Supplementary Fig. 1. e,f, The sensitivity of U2OS cell lines expressing TMEM123, CD164 or Vsg to pTc was assessed by exposure to a titration of pTc for 24 h. e, Representative images of cells after 1.44 nM pTc treatment. f, Left: the percentage of rounded cells was quantified and plotted. Right: the toxin concentration resulting in 50% cell rounding, defined as the CR50, was plotted (numbers above bars indicate exact value). g, Ectopic expression of Vsg–GFP fusion proteins in U2OS cells through transient transfection mediated the binding of the TcdA1 pentamer (15 nm for 3 h) to transfected cells (about 96% TcdA1-bound cells were GFP-positive, n = 165). Biotinylated TcdA1 was labelled with Alexa Fluor 555-conjugated streptavidin. Cells transfected with GFP served as a control. h, Characterization of TcdA1 binding to immobilized biotin-conjugated Vsg-ECD using BLI. i, Pre-incubation of Alexa Fluor-488-conjugated TcdA1 (pentamer, 20 nM) with Vsg-ECD (1 μM) for 2 h at 4 °C reduced TcdA1 binding to S2 cells, as detected by confocal microscopy (top) or flow cytometry (bottom). Cells were labelled with Hoechst (blue) and TcdA1 (green). Representative images and sensorgrams are from one of three independent experiments. Data were analysed from the total number of images indicated in the source data from three experiments and shown as the mean ± s.d., CR50 values in f are displayed as the best-fit value ± 95% confidence interval (CI)). Scale bars, 50 μm (a,e,g,i).
Vsg is composed of an amino-terminal signal peptide, a mucin-like extracellular domain (residues 41–134), a transmembrane domain and a short cytoplasmic domain. Its orthologue in humans, known as endolyn or CD164, is a mucin-like protein localized predominantly on endosomes and can also be transported onto the cell membrane30,31. We next applied gain-of-function approaches to stably express D. melanogaster Vsg in U2OS cells through lentiviral transduction. These U2OS cells were about 14-fold more sensitive to pTc (Fig. 2e,f). By contrast, overexpression of human CD164 or TMEM123, another potential orthologue of Vsg, did not change the sensitivity of U2OS cells (Fig. 2e,f). Consistently, transient transfection of a Vsg–GFP fusion protein in U2OS cells enhanced the binding of fluorescently labelled TcdA1 (Fig. 2g). Overexpression of Vsg in S2 cells also increased the binding of TcdA1 to cells (Extended Data Fig. 5a).
We next expressed and purified the Fc-tagged extracellular domain of Vsg (Vsg-ECD) using S2 cells (Extended Data Fig. 5b). Biotinylated TcdA1 pulled down Vsg-ECD-Fc, but not the control IgG1-Fc (Extended Data Fig. 5c). Biolayer interferometry (BLI) analysis with biotinylated Vsg-ECD-Fc immobilized on the probe further detected binding of TcdA1 (Fig. 2h). As controls, TcdA1 did not bind to empty probes, and TcdA4 from mTc and a Tc toxin from Y. entomophaga (YenTc)8 did not show binding to Vsg-ECD in a BLI analysis (Extended Data Fig. 5d). Consistently, pre-incubation of TcdA1 with Vsg-ECD reduced its binding to both U2OS cells transfected with Vsg (Extended Data Fig. 5e) and to S2 cells (Fig. 2i).
pTc recognizes mosquito and beetle Vsg
We next compared Vsg orthologues in several insect pest species, including two different mosquitoes (Aedes aegypti (aVsg) and Anopheles gambiae (agVsg)), a beetle (Tribolium castaneum (tVsg)) and two moth species (Galleria mellonella (gVsg) and Spodoptera frugiperda (sVsg)) (Fig. 3a and Extended Data Fig. 6a). Their ability to mediate functional entry of pTc was first analysed through transient expression with a GFP tag in U2OS cells (Fig. 3b). Mosquito and beetle orthologues (aVsg, agVsg and tVsg) facilitated pTc entry into U2OS cells. Moth orthologues (gVsg and sVsg) and human CD164 and TMEM123 did not sensitize U2OS cells to pTc (Fig. 3b,c), even though they were expressed on the cell surface (Extended Data Fig. 7a). Consistently, U2OS cells that stably expressed aVsg or tVsg become more sensitive to pTc than control cells, whereas U2OS cells that stably expressed sVsg showed a sensitivity similar to control cells (Fig. 3d,e). Transfection of aVsg in vsg KO S2 cells also restored sensitivity to pTc, similar to transfection with Vsg (Extended Data Fig. 7b,c).
a, The maximum-likelihood tree of Vsg orthologues in the indicated insect species. b,c, Vsg and the indicated Vsg orthologues, each fused with GFP, were expressed in U2OS cells through transient transfection. b, Images of cells after treatment with 2.5 nM pTc for 24 h. c, The percentage of rounded GFP-positive cells was quantified and plotted. d,e, U2OS cells stably expressing aVsg, tVsg or sVsg through lentiviral transduction were exposed to pTc. d, Representative images of cells after exposure to 1.44 nM pTc. e, Percentage of rounded cells was recorded and plotted (left), together with CR50 values (right; numbers above bars indicate exact value). f,g, Two agvsg KO mosquito Sua5B cell lines and the control Sua5B cell line expressing a scrambled sgRNA were exposed to pTc. f, Representative images of cells after exposure to 312.5 pM pTc. g, The percentage of enlarged cells was quantified and plotted (left), together with CE50 values (right; numbers above bars indicate exact value). h, Schematic of Vsg truncation mutants. SP, signal peptide; TM, transmembrane domain; AR, artificial receptor. i,j, The indicated Vsg mutants fused with GFP were expressed in U2OS cells. i, Cells were exposed to pTc (2.5 nM) and representative images were collected. j, Quantification of the percentage of rounded cells among transfected cells. k, O-glycan moieties in Vsg-ECD and sVsg-ECD identified by mass spectrometry analysis are shown as the relative profile intensity in a pie chart. Representative images are from one of three independent experiments. Data were analysed from the total number of images indicated in the source data or bar graphs (n values above bars in c,j) from three experiments and shown as the mean ± s.d. or the best-fit value ± 95% CI. Scale bars, 50 μm (b,d,f,i).
To further confirm the role of Vsg orthologues in mosquitoes, we generated two independent agvsg KO cell lines using the A. gambiae cell line Sua5B and the CRISPR–Cas9 approach (Fig. 3f). Exposure of Sua5B to pTc resulted in enlarged multinucleated cells. The two agvsg KO cell lines showed approximately 29-fold reduction in sensitivity to pTc compared with the control cells that expressed scrambled sgRNA (Fig. 3f,g).
pTc recognizes the HD-PTS region of Vsg
We next constructed a series of GFP-tagged truncation mutants of Vsg and expressed them in U2OS cells (Fig. 3h). Residues 118–148 may not be important for pTc binding as a mutant that contained 1–117 of Vsg-ECD still functioned as a receptor (Fig. 3h–j). A Vsg truncation mutant that contained residues 55–117 of its ECD still partially mediated pTc entry into U2OS cells (Fig. 3h,i). The 55–117 region is within the mucin domain enriched with repeated PTS residues (Extended Data Fig. 6a). As with other mucin-like proteins, O-glycosidase treatment reduced but did not eliminate O-glycans from Vsg-ECD, possibly because many sites are masked by dense O-glycans (Extended Data Fig. 8a). Vsg also contains two potential N-glycosylation sites, but mutating these two sites did not affect its function as a pTc receptor (Extended Data Fig. 8b). Consistently, reducing N-glycosylation with PNGase F did not affect TcdA1 binding to Vsg-ECD. By contrast, O-glycosidase treatment reduced TcdA1 binding to Vsg-ECD (Extended Data Fig. 8c). We further profiled and compared glycan moieties between Vsg-ECD and sVsg-ECD by mass spectrometry (Extended Data Fig. 8d). Typical insect O-glycan moieties were readily identified and there were no major differences in O-glycan moieties between Vsg-ECD that binds to pTc and sVsg-ECD that does not mediate pTc binding (Fig. 3k and Supplementary Data 1). Thus, although O-glycans are involved in interactions, Vsg does not contain specialized glycan moieties that it uses to function as a receptor.
A comparison of the PTS region (residues 41–134) among Vsg orthologues indicated that only Vsg proteins with an extremely high density of threonine and serine residues, designated HD-PTS, can function as pTc receptors (Extended Data Fig. 6b). The mucin regions in CD164, gVsg and sVsg only contain 23–35% of threonine and serine residues (Extended Data Fig. 6b). Vsg is predicted to contain 70 O-glycosylation sites, whereas sVsg contains only 20 such sites (Extended Data Fig. 8e).
The HD-PTS region is expected to form a rigid string saturated with densely packed O-glycans, which indicates that pTc recognition may require highly occupied O-glycosylation sites spatially configured by the HD-PTS sequences. To test this hypothesis, we designed an artificial receptor composed of a HD-PTS segment derived from a mucin protein, human mucin 2 (MUC2; GenBank identifier AAB95295.1, residues 1,400–1,513, with 56% threonine and serine residues) fused with the transmembrane and cytoplasmic domain of sVsg (Fig. 3h and Extended Data Fig. 6a). The artificial receptor mediated entry of pTc when expressed in U2OS cells (Fig. 3h–j).
pTc targets haemocytes through Vsg
We next examined Vsg expression in Drosophila larvae, using a transgenic fly line that expresses GFP–Vsg (Extended Data Fig. 9a). Previous studies using this line reported that Vsg is expressed and clustered around cell surface ring canal structures in germline, follicle and imaginal disc cells32,33. Here, we found that GFP–Vsg appeared as bright puncta on cell membranes between adjacent cells, probably around ring canals, in ovarian tissues, eye discs, brain and lymph glands (Extended Data Fig. 9b). Besides ring canals, GFP–Vsg was found in haemocytes and fat body cells (Extended Data Fig. 9c), two major cell types responsible for innate immunity. Haemocytes clear invading bacteria and parasites through phagocytosis and nodule formation, whereas fat body cells produce antimicrobial peptides34,35.
We then generated two vsg KO D. melanogaster lines using the CRISPR–Cas9 approach (Fig. 4a) and then isolated primary haemocytes from control and KO lines. These haemocytes were exposed to pTc, and toxin-induced actin clustering was detected using phalloidin staining (Fig. 4b). Compared with control haemocytes, vsg KO haemocytes showed around 55-fold reduction in sensitivity to pTc (Fig. 4b,c and Extended Data Fig. 10a,b). We further generated a rescue fly line that expresses aVsg in haemocytes based on our vsg KO fly line. Expression of aVsg in vsg KO haemocytes restored their sensitivity to pTc (Fig. 4d and Extended Data Fig. 10c,d).
a, Schematic of the vsg KO strategy and the genotyping results of two different vsg KO strains. b,c, Haemocytes from control (CTRL, nos-Cas9;attP2), vsg KO or a wild-type strain (w1118) of D. melanogaster were exposed to the indicated concentrations of pTc for 18 h. Cells were fixed and stained with phalloidin (red) and Hoechst dye (blue). b, Representative images. c, Percentages of cells with actin clustering were quantified and plotted (numbers above bars at right indicate exact value). d, Haemocytes from a vsg KO fly line (Hml-Gal4,UAS-EGFP; KO2) and a rescue line that expresses aVsg in haemocytes (Hml-Gal4,UAS-EGFP/UAS-avsg;KO2) were exposed to 20 pM pTc for 18 h. Cells were fixed and stained with phalloidin (red), and imaged (left) and quantified (right). e,f, Haemocytes from control (w1118) or vsg KO2 D. melanogaster were treated with pTc (25 pM or 50 pM for 18 h) and then co-incubated with pHrodo Red-labelled E. coli bioparticles (red) for 30 min. e, Cells were washed, stained with Hoechst (blue) and imaged. f, Internalized E. coli fluorescence intensity per cell was counted and plotted. g,h, Control (nos-Cas9;attP2) and vsg KO D. melanogaster were infected with the control bacterium, Y. entomophaga MH96 through pricking with an insect pin. g, Survival curves. h, Flies were collected and homogenized, and bacterial loads (colony formation unit (CFU)) were assessed by serial dilution plating. i,j, Control (nos-Cas9;attP2) and vsg KO D. melanogaster were infected with P. luminescens FRG04 through pin prick. i, Survival curves. j, Bacterial loads. Data in c, d and f were analysed from the total number of images indicated in the source data or bar graphs (n values above bars in d,f) from three experiments and shown as the mean ± s.d. or the best-fit value ± 95% CI. Data in h and j were analysed from the total number of flies indicated in the Methods. Centre values represent the median. The box plots in h and j show all points. The bounds are minimum to maximum values, 25th and 75th percentiles, respectively. Two-tailed test was used for comparing between control-microbe and vsg KO-microbe group in h and j. log-rank (Mantel–Cox) test was used for comparing between control-microbe group and vsg KO-microbe group in i. P = 0.29 (h), P = 0.008 (i) and P = 0.003 (j). **P < 0.01. Representative images are from one of three independent experiments. Scale bars, 10 μm (b), 20 μm (d,e).
To further assess the impact of pTc on phagocytosis of haemocytes, we utilized pHrodo Red-labelled Escherichia coli bioparticles, which show increased fluorescence signals under low pH conditions, as in endosomes. Incubation with pTc blocked phagocytosis of E. coli bioparticles in control haemocytes. By contrast, phagocytosis in vsg KO haemocytes was not affected by pTc. That is, both the fluorescence intensity and the number of internalized E. coli bioparticles was similar to that in cells not exposed to pTc (Fig. 4e,f and Extended Data Fig. 10e,f).
P. luminescens infection in vsg KO flies
Finally, we assessed the role of Vsg during P. luminescens infection in vivo using adult D. melanogaster, which has been widely utilized as a model for bacterial infection36. The flies were pricked with an insect pin that had been dipped in a bacterial suspension. There was no difference in effects between vsg KO and control flies when infected with the insect pathogen Y. entomophaga (Fig. 4g,h). By contrast, vsg KO flies showed reduced lethality compared with control flies when infected with P. luminescens subsp. akhurstii (Fig. 4i). Quantification of P. luminescens colony forming units in homogenized flies showed that bacterial loads in vsg KO flies were reduced by 30% compared with control flies (Fig. 4j). P. luminescens subsp. akhurstii possesses many other insecticidal toxins and virulence factors6, which may contribute to its sustained lethality in vsg KO flies.
Discussion
Here, we identified Vsg as a receptor for a prototypical Tc toxin and revealed how Tc toxin contributes to P. luminescens pathogenesis. Our findings suggest that pTc recognition requires densely packed sites of O-glycosylation along a HD-PTS region in Vsg. However, the precise molecular mechanism of pTc–Vsg interactions remains to be established, which would require future studies to resolve a high-resolution structure of the pTc–Vsg complex.
Our findings also revealed a previously underappreciated insect species-specificity of pTc. This finding may explain why previous studies of pTc using cells from moth models often required the use of nanomolar pTc concentrations, similar to the potency for mammalian cells. pTc binding to those cell types is probably mediated by lower affinity interactions with sGAGs, N-glycans and other carbohydrate moieties rather than by a particular receptor protein21,22,23. Thus, instead of being a nondiscriminatory toxin, pTc affects a specific subset of insects. This insight may explain the diversification of Tc toxins within P. luminescens, which may target different receptors and act on distinct cell types and/or insect species4. The selectivity of Tc toxins also offers the opportunity to engineer tailor-made biopesticides that target only certain harmful insect species.
The diversification of TcA subunits suggests that their key role in P. luminescens infection cannot be fulfilled with more commonly presented cell surface glycan moieties. This might reflect selective pressure to maximize their potency towards the insect immune system at the early stage of infection, and high potency can only be achieved by evolving the capability to bind highly specific receptors. Vsg is expressed in haemocytes and fat body cells, both of which are vital components of the insect innate immune system. Notably, CD164 has been recently identified as an essential entry factor for lymphocytic choriomeningitis virus37,38. It will be interesting to investigate whether Vsg may serve as a receptor for additional insect toxins and pathogens.
Suppression of the immune system allows rapid replication of P. luminescens39,40. With the increased toxin concentrations in the later stage of infection, various Tc toxins and other toxins or virulence factors in P. luminescens may then work synergistically and attack a broad range of cells through their interactions with common cell surface glycans. This could be the reason why vsg KO flies were still lethally susceptible to P. luminescens. Diversification also occurs for the cytotoxic TcC HVR moieties, which may target various cellular functions beyond phagocytosis.
Genome-wide CRISPR–Cas9-mediated pooled screens in mammalian cells have been widely used. However, this approach remained challenging in insect cells owing to the inefficient transduction of such systems by typically used lentiviruses. With the current study, we provide a successful example of carrying out genome-wide CRISPR–Cas9 screens in insect cells for characterizing insect-targeting toxins and pathogens relevant to agriculture and vector-borne infectious diseases.
Methods
Cell lines and fly strains
D. melanogaster S2R+ cells and A. gambiae Sua5B cells were provided by the Drosophila RNAi Screening Center (Harvard Medical School). A subline of S2R+ cells and Sua5B cells expressing SpCas9 and containing an attP integration site, S2R+/NPT005/MT-Cas9 (PT5/Cas9) and Sua5B-IE8-Cas9, have been previously described27,42 and are available at the Drosophila Genomics Resource Center (DGRC). All cells were grown in Schneider’s medium (21720-024, Thermo Fisher Scientific) containing 10% FBS (16140-071, Thermo Fisher Scientific) and 1× penicillin–streptomycin (15070063, Thermo Fisher Scientific). To maintain the Cas9 transgene, PT5/Cas9 cells were grown in 200 ng ml–1 hygromycin (40051, Calbiochem) and Sua5B/Cas9 cells were grown in 500 μg ml–1 geneticin (11811031, Thermo Fisher Scientific). HeLa, 5637 and U2OS cells were obtained from the American Type Culture Collection. nos-Cas9;attP2 flies were a gift from J. Ni at Tsinghua University, Beijing, China. The w1118 strain is from the Perrimon Laboratory. The UAS-avsg strain was constructed by inserting the A. aegypti Vsg coding sequence (protein identifier XP_021708011.1) into EcoRI-XbaI-digested pWalium10-moe (1470, DGRC). Transgenic flies were generated using PhiC31 integration by injecting the pWalium10-moe-avsg plasmid into flies carrying an attP docking site on chromosome II (attP40). The w1118;P{Hml-GAL4.Δ}2,P{UAS-2xEGFP}AH2 (30140) and y1w*;P{PTT-GA}vsgCA07004 (50812) strains were obtained from the Bloomington Drosophila Stock Center.
Antibodies
Goat anti-human IgG Fc secondary antibody (A18817, Invitrogen), mouse anti-Hts antibody (1B1, DSHB), rabbit anti-GFP antibody (A11122, Invitrogen), anti-pan-ADP-ribose binding reagent (MABE1016, Millipore), donkey anti-rabbit lgG (H+L) Alexa Fluor-488 (R37118, Invitrogen), donkey anti-mouse lgG (H+L) Alexa Fluor-568 (A10042, Invitrogen), Alexa Fluor 555-conjugated phalloidin (8953S, Cell Signaling), Alexa Fluor-488-conjugated phalloidin (A12379, Thermo Fisher Scientific), Alexa Fluor 647-conjugated phalloidin (A22287, Invitrogen), anti-α-tubulin (T5168, Sigma-Aldrich), HRP-conjugated goat anti-rabbit IgG (H+L) secondary antibody (31460, Thermo Fisher) and HRP-conjugated goat anti-mouse IgG (H+L) secondary antibody (31430, Thermo Fisher) were purchased from the indicated vendors.
Tc toxin production
P. luminescens TcdA1 and M. morganii TcdA4 were produced as previously described8. Expression was carried out in E. coli BL21 CodonPlus(DE3) RIPL cells (Agilent) using a pET-19b vector (Novagen) with the toxin genes cloned in-frame with an N-terminal 6×His-tag and a PreScission protease cleavage site. After transformation and growth to OD600 = 0.6 in 10 litres of LB medium, protein production was induced with 25 µM IPTG. Protein expression was carried out at 20 °C for 20 h, and cells were lysed in 50 mM Tris-HCl pH 8.0, 300 mM NaCl, 0.05% Tween-20 supplemented with benzonase endonuclease and AEBSF using a pre-chilled microfluidizer. Cell debris was spun down at 167,000g using a type 45 Ti rotor (Beckman Coulter Life Sciences) for 30 min at 4 °C. Next, 25 mM imidazole pH 8.0 was added to the supernatant, which was subsequently loaded onto an equilibrated 5 ml HisTrap HP column (Cytiva). After washing until the baseline was stable, the bound protein was eluted using a linear gradient of 25–500 mM imidazole. The elution fractions of the Tc toxins were pooled, concentrated and further purified using a Sephacryl S400 size-exclusion column (Cytiva) equilibrated in 20 mM HEPES pH 8.0, 100 mM NaCl, 0.05% Tween-20. The purified protein was pooled, concentrated and stored at −80 °C once supplemented with 40% glycerol. Production and purification of YenTc from Y. entomophaga was performed as previously described43. P. luminescens TcdB2-TccC3 was produced in E. coli BL21 CodonPlus(DE3) RIPL as previously described17 using pET-28a with the tcdB2 and tccC3 genes cloned as a fusion protein in-frame with an N-terminal 6×His-tag and a thrombin cleavage site.
Biotinylation and deglycosylation of Vsg-ECD
To prepare biotinylated Vsg-ECD-Fc, it was first desalted into 20 mM HEPES pH 8.0, 100 mM NaCl, 0.05% Tween-20. A sixfold molar excess of EZ-Link NHS-PEG12-biotin (Thermo Fisher Scientific) was added, and the mixture was incubated at 4 °C for 14 h. Excess biotin was removed by desalting the biotinylated protein into 20 mM HEPES pH 8.0, 100 mM NaCl, 0.05% Tween-20 (when no deglycosylation was performed) or 20 mM MES pH 6.5, 150 mM NaCl (when deglycosylation was performed). When needed, Vsg-ECD-Fc was deglycosylated for 14 h at 4 °C through the addition of 500 U PNGase F (New England Biolabs) and 40,000 U O-glycosidase (New England Biolabs) per 30 µl of 7.5 µM biotinylated Vsg. This was followed by desalting into 20 mM HEPES pH 8.0, 100 mM NaCl, 0.05% Tween-20 and dilution to 3.1 µg ml–1.
BLI assay
Biotinylated Vsg-ECD-Fc protein was immobilized on the surface of streptavidin sensors. In reference experiments, TcdA1, TcdA4 or YenTc were used as ligands against an independent set of streptavidin sensors not exposed to biotinylated Vsg-ECD-Fc. To obtain referenced data that corrects for TcdA1 binding to streptavidin sensors, the values of reference wells were first subtracted from the original data. The values from the Vsg-free reference sensors were then subtracted from those of the Vsg-bound sensors. BSA from a 100× stock was added to both Vsg and the Tc toxins before use to a final concentration of 0.3 mg ml–1. The buffer used for the relevant BLI washing steps and for preparing Tc toxin dilutions was 20 mM HEPES pH 8.0, 100 mM NaCl, 0.05% Tween-20 and 0.3 mg ml–1 BSA. A dilution series of the toxins (5,000–78 nM, plus a toxin-free condition as a reference) were prepared directly before measurement. BLI sensorgrams were acquired at 25 °C with an initial 30 s sensor check step, followed by 300 s loading time, a 300 s baseline step and 100 s association and 400 s dissociation times.
Genome-wide CRISPR–Cas9 screening with pTc or mTc
The Drosophila CRISPR genome-wide KO library has been previously described and is available from Addgene (134582-4)27. In brief, 1 × 108 PT5/S2R+ cells were transfected with an equal-parts mixture of pLib6.4 containing a genome-wide library (84,563 sgRNAs targeting 13,685 Drosophila genes) and pBS130 (26290, Addgene) using Effectene (301427, Qiagen) per the manufacturer’s instructions. After 4 days, the transfected cell library was selected with 5 μg ml–1 puromycin (540411, Calbiochem) for 12 additional days, subculturing every 4 days. After the stable sgRNA-expressing cell library was established, 2 × 107 cells were plated onto four 15-cm cell culture dishes to ensure sufficient sgRNA coverage, with each sgRNA being represented around 1,000 times. These cells were exposed to pTc for 1 week. Cells were then filtered through 30-μm cell strainers, only collecting the cells that passed through the strainer. The collected cells were further cultured in toxin-free medium to ultra-confluence and then subjected to subsequent rounds of screening with higher concentrations of toxins. Three independent screens were performed for pTc with the following conditions (in pM): screen 1: 6, 12 (sequenced as round 1), followed with either 100 (screen 1-low, round 2) or 125 (screen 1-high, round 2); screen 2: 12.5, 12.5 (round 1), 20 (round 2), 40 (round 3); screen 3: 10, 10 (round 1), 25 (round 2), 50 (round 3). For mTc, 550, 1,100 and 3,600 pM were used. Aliquots of cell libraries following each selection round were collected, and their genomic DNA was extracted using a Zymo Quick-gDNA Miniprep kit (D3025, Zymo Research). DNA fragments containing the sgRNA sequences were amplified by PCR using at least 1,000 genomes per sgRNA as templates for each sample. PCR fragments were in-line barcoded using a previously described approach27. Next-generation sequencing (Illumina NextSeq) was performed at the Biopolymers Facility at Harvard Medical School. Fastq sequence files were demultiplexed using in-line barcodes as sequence tags, generating a distinct file for each treatment sample. All further processing used MAGeCK (v.0.5.4 or 0.5.9.4) to generate readcount files and to perform robust rank aggregation analysis directly on the generated readcount files using default parameters. RRA values presented used plasmid readcounts for normalization.
Secondary screening by RNAi
All pre-synthesized dsRNA templates for the top genes from genome-wide pTc CRISPR screen 1 were obtained from the Drosophila RNAi Screening Center and in vitro transcribed to produce dsRNAs using a T7 Megascript kit (Thermo Fisher Scientific). Template sequences were obtained by searching sequence identifiers at https://www.flyrnai.org/up-torr/. S2R+ cells were washed into serum-free medium and 10,000 cells were transferred to each well of 384-well dishes containing 25 ng dsRNA for 35 min followed by outgrowth in serum-containing medium for 3 days. Then, cells were subjected to 100 pM pTc for 2 days and expanded to 96-well dishes and grown further for 7 days. Live cells were stained with 10 mM Hoechst 33342 (B2261, Sigma) for 15 min before being subjected to flow cytometry using a LSRII with a 96-well autoloader. Data analysis was carried out using FlowJo v.10.8.1 software. FACSDiva v.9.0 software was used for data collection in all flow cytometry experiments. The percentage of all events passing through a gate corresponding to the BV-421 intensity of most (>86%) of untreated cells was measured. Note that for the Hoechst analysis, because pTc alters cell size, we did not carry out gating using the forward/side scatter profile. For a separate analysis of cell size, the percentage of all events passing through a gate corresponding to an enlarged forward scatter/side scatter profile (>89% of untreated cells) was measured.
Plasmid construction
avsg (protein identifier XP_021708011.1), tvsg (XP_972460.1), svsg (KAF9794377.1), vsg (NP_729535.1), agvsg (XP_001688320.1), gvsg (XP_026748881.1), TMEM123 (NP_443164.2) and CD164 (NP_006007.2) were synthesized by Twist Bio. Their entire coding sequences, excluding the final 15 bp (deletion of four residues from the C terminus, which constitute the endocytic sorting motif), were cloned into a pEGFP-N1 vector between the sites NheI and HindIII to build the Vsg–GFP fusion constructs for transient expression in mammalian cells. For all the pCMV-SP-GFP–Vsg constructs, GFP was inserted immediately after the signal peptide (SP) of Vsg proteins and amplified SP-GFP–Vsg fragments were inserted into the pEGFP-N1 vector between the sites NheI and HindIII. The entire coding sequences of avsg, tvsg, svsg, vsg, TMEM123 and CD164 were subcloned into the sites BamHI and XbaI in the vector pLenti CMV/TO Hygro empty (w214-1) (17484, Addgene) to establish stable cell lines. For S2-cell-based rescue experiments, entire coding sequences of avsg and vsg were subcloned into the sites EcoRI and XhoI in the vector Ac5-stable2-Neo (32426, Addgene). To build the Vsg-ECD-Fc expression construct, two fragments, vsg (67–444 bp) or sVsg-ECD (52–546 bp) and human Fc-his, were amplified from vsgs and pHL-FcHis vector (99846, Addgene), respectively, and cloned into the vector pMT-Bip-GFP:V5:KDEL (69917, Addgene) between the KpnI and PmeI sites through Gibson assembly (E2621, NEB).
Vsg-ECD protein purification
pMT-Bip-Vsgs-ECD-Fc-his plasmids were transfected into S2R+ cells using PolyJet in vitro DNA transfection reagent (SL100688, SignaGen). Cells were selected with 2 μg ml–1 puromycin (A1113803, Gibco) for 2 weeks to establish stable cell lines. Cells were seeded into six 15-cm plates with 60% confluency, and kept in serum free Schneider’s medium (21720024, Gibco) with 500 μM Cu2+ for 5 days. Cell medium was collected, and proteins were then purified through immobilized metal ion-affinity chromatography (17-0575-01, GE Healthcare).
ADP-ribosylation detection
vsg KO cells and control cells were seeded onto 6-well plates with around 20% confluency, and incubated with 0, 25 or 50 pM pTc for 24 h at 25 °C. Cells were collected, washed by PBS and collected in 100 μl lysis buffer (pH 7.4, 20 mM Tris-HCl, 150 mM NaCl, 0.5% Triton X-100 and 0.5% SDS) for 10 min on ice. Next, 10 μl of each sample was subjected to SDS–PAGE and immunoblot analysis with 1:1,000 anti-pan-ADP-ribose binding reagent and 1:1,000 anti-α-tubulin as the primary antibody and 1:5,000 HRP-conjugated goat anti-rabbit IgG or goat anti-mouse IgG as the secondary antibody.
Pull-down experiments
Dynabeads M-280 streptavidin beads (50 μl) were equilibrated in wash buffer (0.1% Triton X-100, 20 mM Tris-HCl and 150 mM NaCl, pH 7.5). Biotinylated TcdA1 (10 μg) and equilibrated beads were added to 300 μl wash buffer, and incubated at 4 °C for 2 h, and then Vsg-ECD-Fc (3 μg) or human IgG1-Fc (10702-HNAH, Sinobiological) was added to the buffer and incubated overnight. Beads were washed, collected, boiled and subjected to SDS–PAGE and immunoblot analysis.
Generating vsg KO cell lines
CRISPR KOs in D. melanogaster PT5/Cas9 or A. gambiae Sua5B cells were carried out by transfecting cells with pLibAgam containing the following sgRNAs: Dmel_vsg1: 5′-TCGAAATGACCGCACGGCAC-3′; Dmel_vsg2: 5′-CTAATCTTGTGCCTCTTTGT-3′; Dmel_vsg3: 5′-GTGGTGTCCGGCGGAGTGG-3′). Acol_vsg1: 5′-GTAGTAGTATTAGAACTTG-3′; Acol_vsg2: 5′-CGACGTCGTAGGCGTAGCTG-3′. Transfection was performed with Effectene along with pBS130 and selected in 5 μg ml–1 puromycin. Control sgRNAs for PT5/Cas9 cells were scrambled 1: 5′-GATCATGCACGTGGTCAATA-3′; scrambled 2: 5′- GCTCTACAACCGACCGCTGC-3′; and scrambled 3: 5′-AGTCAGTGGTCCGTTACAGG-3′. Control sgRNA for Sua5B cells was 5′- CTAGACTCACTAGAGGGCAC-3′.
Generating vsg KO flies
The vsgKO1 and vsgKO2 alleles were generated using the CRISPR–Cas9 approach. Three recognition sequences of guide RNA to the vsg locus were designed. sgRNA1: 5′-CTAATCTTGTGCCTCTTTGT-3′; sgRNA2: 5′-GTGCCGTGCGGTCATTTCGA-3′; and sgRNA3: 5′-AGTGGCCTACAAGTTCTACA-3′. These were cloned into the U6b-sgRNA-short vector, and pU6-sgRNA1 and pU6-sgRNA2 were used to generate the vsgKO1 flies, whereas pU6-sgRNA1 and pU6-sgRNA3 were used for the vsgKO2 flies. In brief, plasmids were co-injected into the embryos of (nos-Cas9) attP2 flies, then the F1 progeny were screened by PCR to identify the corresponding deletion using the following primers: pF: 5′-CCACTTCGCAAACGTATTTTCTT-3′ and pR: 5′-AGCTCCGGCTGCCGCATTGAA-3′. We used primers targeting gpdh1 as genotyping control primers: pF: 5′-GTCACCTGAAGGCAAGTCAG-3′ and pR: 5′- CTTGCCATACTTCTTGTCCGT-3′.
Cytopathic cell-rounding and cell-enlargement assays
The cytopathic effect of pTc in mammalian cells was analysed using a standard cell-rounding assay. In brief, cells were exposed to pTc for 24 h, and phase-contrast images of cells were recorded (Olympus Ix51, ×20 objective). The numbers of normal and round-shaped cells were manually counted from at least 3 pictures (30–150 cells per picture) from 3 repeats. The percentage of round-shaped cells was plotted and analysed using GraphPad Prism7.
For insect cells, the cytopathic effect of pTc was analysed based on the enlargement phenotype. Cells were exposed to pTc for 4 days, and phase-contrast images of cells were recorded (Olympus Ix51, ×20 objective). The numbers of normal and enlarged cells were manually counted from at least 3 pictures (30–500 cells per picture) from 3 repeats. The percentage of enlarged cells was plotted and analysed using GraphPad Prism7.
Whole-mount staining of fly tissues
Fly tissues were dissected in Schneider’s medium, fixed in 4% PFA for 30 min and blocked in PBST (0.3% Triton X-100) with 5% goat serum for 1 h. Tissues were then incubated with primary antibodies (1:200) in PBST (0.1% Triton X-100) with 5% goat serum at 4 °C overnight. After washing with PBST six times, tissues were incubated with secondary antibodies (1:500) for 1 h at room temperature, then washed with PBST six additional times. Samples were mounted in antifade mounting buffer (H-1000, Vector). Images were recorded with a Zeiss LSM880 upright confocal system and processed in ImageJ bundled with 64-bit Java 1.8.0_172.
Actin immunostaining in primary haemocytes
In brief, third-stage larvae were washed with PBS three times, and then dissected in Schneider’s medium plus 10% FBS, which enabled the release of haemocytes from the larval body cavity. Corpses and debris were carefully removed, and then the Schneider’s medium containing haemocytes was seeded into a glass-bottom 96-well plate (P96-1.5H-N, Cellvis). After haemocytes attached to the plate, the medium was replaced with Schneider’s medium plus 10% FBS containing different concentrations of pTc for 18 h in a 25 °C insect cell incubator. Cells were then fixed with 4% PFA and stained with Alexa Fluor 555-conjugated phalloidin (8953S, Cell Signaling) for 20 min. Red fluorescence images of cells were recorded (Zeiss LSM880, ×20 objective and ×63 oil objective). The numbers of normal cells and cells with bright actin-clustering were manually counted manually from at least three pictures from three repeats for each concentration of pTc. The percentage of cells with bright actin-clustering was analysed using GraphPad Prism7.
Phagocytosis assay
Primary haemocytes from larvae were collected and seeded into glass-bottom 96-well plates. After haemocytes attached to the plate, the medium was replaced with Schneider’s medium plus 10% FBS containing 0, 25 or 50 pM pTc for 18 h. The medium was then changed into serum-free Schneider’s medium with 40 μg ml–1 E. coli (K-12 strain) bio-particles (pHrodo Red E. coli BioParticles P35361, Invitrogen) for 30 min.
After incubation, the cell monolayer was washed three times with serum containing medium, with the final wash containing Hoechst dye. Cells were then directly imaged to detect pHrodo Red E. coli BioParticles, or haemocytes were fixed with 4% PFA for 15 min and stained with Alexa Fluor-488-conjugated phalloidin (A12379, Thermo Fisher Scientific) for 20 min to reveal F-actin. Fluorescence images of cells were recorded (Zeiss LSM880, ×20 objective and ×63 oil objective). The integrated fluorescence intensity was quantified in ImageJ.
Bacterial infection in flies
Y. entomophaga was recovered from a frozen stock and incubated inBHI medium (53286, Millipore) at 28 °C overnight. When the OD600 reached 1.2, the bacteria were collected and diluted 10-fold for infection. P. luminescens subsp. akhurstii was recovered from a frozen stock and incubated in BHI medium at 30 °C overnight. When the OD600 reached 1.2, bacteria were collected and concentrated 17-fold for infection. D. melanogaster (3–5 days old) with a sex ratio of 1:1 were collected and anaesthetized by CO2, pricked with a 0.1-mm stainless steel insect pin (26002-10, Austerlitz) and transferred to a 24 °C incubator for survival tracking.
Colony forming units were quantified in independent batches ofinfections. Flies were raised in a 16 °C fly incubator (3990FL, Thermo Fisher) before infection. After approximately 70% of the flies were dead in the control-microbe group, all flies were collected and homogenized. For the microbe-treatment groups, 7 groups of 14 flies were collected and quantified for both KO and control in P. luminescens subsp. akhurstii infections, and 8 groups of 10 flies were collected and quantified for Y. entomophaga infections. For the PBS groups, 6 groupsof 14 flies were collected and quantified in P. luminescens subsp. akhurstii infection, and 6 groups of 10 flies were collected and quantified in Y. entomophaga infection. In brief, the homogenates were serially diluted and then deposited in 5 μl spots on NBTA agar plates. Plates were incubated for 19–24 h at 30 °C (before Drosophila gut microbiota colonies appear). For colony forming unit estimation, dilutions were chosen in which approximately 5–30 colonies were present in the microbe group. Under the chosen dilution, we found zero colonies in both PBS-groups.
Isolation of Vsg glycan
Approximately 300 μg of purified Vsg-ECD and sVsg-ECD was suspended in 0.1 M NaOH solution before addition of NaBH4 to a final concentration of 55 mg ml–1, and then the samples were incubated at 45 °C for 45–55 h to release the oligosaccharides. The reaction was terminated through the drop-wise addition of pure acetic acid, and the samples were then loaded onto a Amberchrom 50WX8 column with a mesh size of 200–400 (217514, Sigma-Aldrich), which was equilibrated with 5% acetic acid. The column was then washed with 5% acetic acid. Flow through and wash fractions were collected, pooled and lyophilized. Residual salts in the preparation were removed by co-evaporation with 3× 1 ml of 9:1 methanol/acetic acid solution under a stream of nitrogen. Subsequent dry samples were purified on a 200 mg C18 Sep-Pak column (WAT054945, Waters) by washing with 3 ml of 5% acetic acid. The flow through and wash fractions were combined and lyophilized for further analysis.
MALDI-TOF analysis of Vsg glycans
β-eliminated glycans were permethylated, lyophilized and then redissolved in 10 μl of 75% methanol, from which 1 μl was mixed with 1 μl 2,5-dihydroxybenzoic acid (5 mg ml–1 in 50% acetonitrile with 0.1% trifluoroacetic acid) and spotted on a MALDI polished steel target plate (Bruker Daltonics). Mass spectrometry data from each spot was then acquired on a Bruker UltraFlex II MALDI-TOF mass spectrometer (Bruker Daltonics). Reflective positive mode was used, and data were recorded between m/z 300 and m/z 4000. For each mass spectrometer O-glycan profile, an aggregation of 20,000 laser shots or more were collected for data extraction. Mass signals of a signal/noise ratio of at least 4 were considered. Once acquired, the data were manually analysed, and possible O-glycan compositions were annotated based on previously reported insect core structures44.
Reporting summary
Further information on research design is available in the Nature Research Reporting Summary linked to this article.
Data availability
All data generated or analysed during this study are included in this published article and its supplementary data. All biological materials are available upon request from the corresponding authors. Source data are provided with this paper.
References
Lacey, L. A. et al. Insect pathogens as biological control agents: back to the future. J. Invertebr. Pathol. 132, 1–41 (2015).
Labaude, S. & Griffin, C. T. Transmission success of entomopathogenic nematodes used in pest control. Insects 9, 72 (2018).
Bowen, D. et al. Insecticidal toxins from the bacterium Photorhabdus luminescens. Science 280, 2129–2132 (1998).
Roderer, D. & Raunser, S. Tc toxin complexes: assembly, membrane permeation, and protein translocation. Annu. Rev. Microbiol. 73, 247–265 (2019).
Waterfield, N. R., Bowen, D. J., Fetherston, J. D., Perry, R. D. & ffrench-Constant, R. H. The tc genes of Photorhabdus: a growing family. Trends Microbiol. 9, 185–191 (2001).
Waterfield, N. R., Ciche, T. & Clarke, D. Photorhabdus and a host of hosts. Annu. Rev. Microbiol. 63, 557–574 (2009).
Clarke, D. J. Photorhabdus: a tale of contrasting interactions. Microbiology (Reading) 166, 335–348 (2020).
Leidreiter, F. et al. Common architecture of Tc toxins from human and insect pathogenic bacteria. Sci. Adv. 5, eaax6497 (2019).
Hares, M. C. et al. The Yersinia pseudotuberculosis and Yersinia pestis toxin complex is active against cultured mammalian cells. Microbiology (Reading) 154, 3503–3517 (2008).
Landsberg, M. J. et al. 3D structure of the Yersinia entomophaga toxin complex and implications for insecticidal activity. Proc. Natl Acad. Sci. USA 108, 20544–20549 (2011).
Song, N. et al. Genome-wide dissection reveals diverse pathogenic roles of bacterial Tc toxins. PLoS Pathog. 17, e1009102 (2021).
Waterfield, N., Hares, M., Yang, G., Dowling, A. & ffrench-Constant, R. Potentiation and cellular phenotypes of the insecticidal Toxin complexes of Photorhabdus bacteria. Cell. Microbiol. 7, 373–382 (2005).
Gatsogiannis, C. et al. A syringe-like injection mechanism in Photorhabdus luminescens toxins. Nature 495, 520–523 (2013).
Meusch, D. et al. Mechanism of Tc toxin action revealed in molecular detail. Nature 508, 61–65 (2014).
Piper, S. J. et al. Cryo-EM structures of the pore-forming A subunit from the Yersinia entomophaga ABC toxin. Nat. Commun. 10, 1952 (2019).
Busby, J. N., Panjikar, S., Landsberg, M. J., Hurst, M. R. & Lott, J. S. The BC component of ABC toxins is an RHS-repeat-containing protein encapsulation device. Nature 501, 547–550 (2013).
Gatsogiannis, C. et al. Tc toxin activation requires unfolding and refolding of a β-propeller. Nature 563, 209–213 (2018).
Roderer, D., Hofnagel, O., Benz, R. & Raunser, S. Structure of a Tc holotoxin pore provides insights into the translocation mechanism. Proc. Natl Acad. Sci. USA 116, 23083–23090 (2019).
Gatsogiannis, C. et al. Membrane insertion of a Tc toxin in near-atomic detail. Nat. Struct. Mol. Biol. 23, 884–890 (2016).
Lang, A. E. et al. Photorhabdus luminescens toxins ADP-ribosylate actin and RhoA to force actin clustering. Science 327, 1139–1142 (2010).
Roderer, D. et al. Glycan-dependent cell adhesion mechanism of Tc toxins. Nat. Commun. 11, 2694 (2020).
Ng'ang'a, P. N. et al. Involvement of N-glycans in binding of Photorhabdus luminescens Tc toxin. Cell. Microbiol. 23, e13326 (2021).
Song, N. et al. N-glycans and sulfated glycosaminoglycans contribute to the action of diverse Tc toxins on mammalian cells. PLoS Pathog. 17, e1009244 (2021).
Shalem, O. et al. Genome-scale CRISPR–Cas9 knockout screening in human cells. Science 343, 84–87 (2014).
Tao, L. et al. Frizzled proteins are colonic epithelial receptors for C. difficile toxin B. Nature 538, 350–355 (2016).
Zhou, Y. et al. High-throughput screening of a CRISPR–Cas9 library for functional genomics in human cells. Nature 509, 487–491 (2014).
Viswanatha, R., Li, Z., Hu, Y. & Perrimon, N. Pooled genome-wide CRISPR screening for basal and context-specific fitness gene essentiality in Drosophila cells. eLife 7, e36333 (2018).
Hacker, U., Lin, X. & Perrimon, N. The Drosophila sugarless gene modulates Wingless signaling and encodes an enzyme involved in polysaccharide biosynthesis. Development 124, 3565–3573 (1997).
Kaiser, E. et al. Membrane translocation of binary actin-ADP-ribosylating toxins from Clostridium difficile and Clostridium perfringens is facilitated by cyclophilin A and Hsp90. Infect. Immun. 79, 3913–3921 (2011).
Zhou, G. Q. et al. The Drosophila ortholog of the endolysosomal membrane protein, endolyn, regulates cell proliferation. J. Cell. Biochem. 99, 1380–1396 (2006).
Watt, S. M., Buhring, H. J., Simmons, P. J. & Zannettino, A. W. C. The stem cell revolution: on the role of CD164 as a human stem cell marker. NPJ Regen. Med. 6, 33 (2021).
Buszczak, M. et al. The carnegie protein trap library: a versatile tool for Drosophila developmental studies. Genetics 175, 1505–1531 (2007).
Airoldi, S. J., McLean, P. F., Shimada, Y. & Cooley, L. Intercellular protein movement in syncytial Drosophila follicle cells. J. Cell Sci. 124, 4077–4086 (2011).
Hoffmann, J. A. & Reichhart, J. M. Drosophila innate immunity: an evolutionary perspective. Nat. Immunol. 3, 121–126 (2002).
Kounatidis, I. & Ligoxygakis, P. Drosophila as a model system to unravel the layers of innate immunity to infection. Open Biol. 2, 120075 (2012).
Boman, H. G., Nilsson, I. & Rasmuson, B. Inducible antibacterial defence system in Drosophila. Nature 237, 232–235 (1972).
Bakkers, M. J. G. et al. CD164 is a host factor for lymphocytic choriomeningitis virus entry. Proc. Natl Acad. Sci. USA 119, e2119676119 (2022).
Liu, J. et al. Genome-wide knockout screen identifies human sialomucin CD164 as an essential entry factor for lymphocytic choriomeningitis virus. mBio 13, e0020522 (2022).
Daborn, P. J., Waterfield, N., Blight, M. A. & ffrench-Constant, R. H. Measuring virulence factor expression by the pathogenic bacterium Photorhabdus luminescens in culture and during insect infection. J. Bacteriol. 183, 5834–5839 (2001).
Silva, C. P. et al. Bacterial infection of a model insect: Photorhabdus luminescens and Manduca sexta. Cell. Microbiol. 4, 329–339 (2002).
Li, W. et al. MAGeCK enables robust identification of essential genes from genome-scale CRISPR–Cas9 knockout screens. Genome Biol. 15, 554 (2014).
Viswanatha, R. et al. Bioinformatic and cell-based tools for pooled CRISPR knockout screening in mosquitos. Nat. Commun. 12, 6825 (2021).
Jones, S. A. & Hurst, M. R. Purification of the Yersinia entomophaga Yen-TC toxin complex using size exclusion chromatography. Methods Mol. Biol. 1477, 39–48 (2016).
Kurz, S. et al. Targeted release and fractionation reveal glucuronylated and sulphated N- and O-glycans in larvae of dipteran insects. J. Proteomics 126, 172–188 (2015).
Acknowledgements
We thank K. Vogel-Bachmayr for technical assistance, and R. Stubbendieck for suggestions on the phylogenetic tree of Vsg proteins. This study was partially supported by grants from National Institutes of Health (NIH) (R01NS080833, R01NS117626, R01AI132387 and R01AI139087 to M.D.; R01AI170835 to M.D. and N.P.), Intelligence Advanced Research Projects Activity (IARPA) (grant number W911NF-17-2-0089 to M.D.), NIH NIGMS P41 GM132087 (to N.P.) and from the Max Planck Society (to S.R.). M.D. acknowledges support from the NIH-funded Harvard Digestive Disease Center (P30DK034854) and the Boston Children’s Hospital Intellectual and Developmental Disabilities Research Center (P30HD18655). M.D. holds the Investigator in the Pathogenesis of Infectious Disease award from the Burroughs Wellcome Fund. N.P. is an investigator of the Howard Hughes Medical Institute.
Author information
Authors and Affiliations
Contributions
Y.X., R.V., D.R., S.T., S.R. and M.D. initiated the project. Y.X. and R.V. designed and carried out CRISPR–Cas9 screens. Y.X. designed and carried out the majority of experiments. R.V. conducted the RRA analysis, carried out validation assays using dsRNA approaches, and generated vsg KO cells, the UAS-avsg fly line and all the flow cytometry experiments. O.S. and D.R. prepared, purified and modified Tc toxins. O.S. carried out de-glycosylation of Vsg and BLI analysis. H.Z. generated the vsg KO fly lines. C.A. and C.V. carried out glycan analysis by mass spectrometry. S.R., N.P. and M.D. supervised the project. Y.X., R.V. and M.D. wrote the manuscript with input from all co-authors.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no conflicts of interest.
Peer review
Peer review information
Nature thanks Klaus Aktories, Holger Barth, Richard ffrench-Constant and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Characterizing pTc toxicity in mammalian and insect cells.
a–b, U2OS, 5637, and HeLa cells were exposed to pTc for 24 h. Representative images were from one of three independent experiments and shown in (a). The percentage of rounded cells and CR50 values was quantified and plotted in (b). c–d, S2 cells were treated with the indicated dose of pTc for 4 days. Representative images were from one of three independent experiments and shown in (c). The percentage of enlarged cells and CE50 values was quantified and plotted in (d). Data were analyzed from the total number of images indicated in the Source data from three experiments and shown as mean ± SD or the best-fit value ± 95% CI. Scale bar = 50 μm.
Extended Data Fig. 2 Top-ranked genes identified in genome-wide CRISPR screens.
a, List of most prevalent 38 sgRNAs following the last round of selection with pTc or mTc. b, Prevalence of all sgRNAs targeting Vsg, Mop, FIG4, or VAC14 across three rounds of pTc or mTc screens were quantified and plotted according to their corresponding gRNA abundances.
Extended Data Fig. 3 RNAi knock-down validation of top hits from CRISPR screens.
a, Schematic of the experimental steps. Briefly, S2 cells were bathed in dsRNAs targeting the top hits from pTc CRISPR screen # 1 round 1 and allowed to recover for 3 days. Next, cells were treated with 100 pM pTc for 2 days, re-seeded and grown for an additional 7 days. DNA was stained by Hoechst dye and cells were analyzed by flow cytometry. b, Comparison of untreated cells (red) versus those treated with 100 pM pTc (blue) revealed an increase in DNA content after pTc treatment (left). Comparison of pTc treatment following bathing in vsg dsRNA (green) versus LacZ control dsRNA (gold) showed that vsg dsRNA pre-bathing abrogated pTc-induced increase in DNA content. Gate representing DNA content of 80–90% of untreated cells was set. There was no pre-gating on cell sizes as pTc treatment causes changes in cell sizes and shapes. c, The indicated top hits from pTc CRISPR screens were knocked down in S2 cells using dsRNA and then exposed to pTc as described in (a). Percentage of cells with normal Hoechst levels were plotted. d, Comparison of untreated cells (No Toxin) versus those treated with 100 pM pTc revealed increase in cell size (quantified using the forward scatter/side scatter profile, SSC-A versus FSC-A). Comparison of pTc treatment following bathing in vsg dsRNA (DRSC25227 and DRSC10219) versus LacZ control dsRNA showed that vsg dsRNA pre-bathing abrogated pTc-induced cell size enlargement. e, The indicated top hits from pTc CRISPR screens were knocked down in S2 cells using dsRNA and then exposed to pTc as described in (a). The percentage of enlarged cells was plotted, which gave similar results to the DNA-content measurement in (c).
Extended Data Fig. 4 Characterization of vsg KO S2 cells.
a–c, S2 cells expressing one of the three indicated sgRNAs before or after treatment with 130 pM pTc for 2 weeks were harvested, andthe sequence at the gRNA targeting sites analyzed by deep sequencing. The frequency of deletions of the indicated size were plotted (insertions represented less than 1% and were ignored). Non-frame-shift mutations (potentially retaining Vsg receptor activity, red) versus frameshift mutations (blue) are shown. d, vsg KO2, vsg KO3, and the control S2 cells expressing scrambled sgRNA were exposed to two Tc toxins (Xn-XptA1, 3.5 nM; PI-TcdA4, 3 nM) for 4 days. Both contain the same TcdB2-TccC3 as pTc but utilize distinct A subunits (XptA1 from X. nematophila and TcdA4 from P. luminescens, respectively). Both toxins induced cell enlargement on vsg KO cells and showed no difference on potency on control cells versus vsg KO cells. Data were analyzed from the total number of images indicated in the bar graphs from three experiments and shown as mean ± SD.
Extended Data Fig. 5 TcdA1 binds to Vsg-ECD and Vsg-ECD can reduce binding of TcdA1 to cells.
a, S2 cells stably expressing GFP-Vsg (GFP was inserted after Vsg signal peptide, SP-GFP-Vsg cells) were mixed with the control S2 cells. Cells were incubated with Alexa Fluor-647-conjugated TcdA1 (pentamer, 25 nM) at room temperature for 1 h and then subjected to flow cytometry analysis. S2 cells that express GFP-Vsg showed elevated binding of TcdA1 compared with the control S2 cells. b, Vsg-ECD-Fc produced in S2 cells shown in SDS-PAGE gels with Coomassie blue staining, which showed an apparent molecular weight of ~ 130 kDa, much larger than its predicted molecular weight, indicating that the protein is heavily glycosylated. Representative images were from one of three independent experiments. c, Biotinylated TcdA1 interacted with purified Fc-tagged Vsg-ECD, but not the control IgG1-Fc. For gel source data, see Supplementary Fig. 1. Representative images were from one of three independent experiments. d, BLI analysis showing that TcdA1 did not bind to empty probes, and Morganella morganii TcdA4 (MmTcdA4) from mTc and a Tc toxin (YenTc) from Yersinia entomophaga showed no binding to immobilized Vsg-ECD-Fc. e, Pre-incubation of 15 nM TcdA1 pentamer with 1 μM Vsg-ECD for 2 h in 4 °C reduced TcdA1 binding to U2OS cells transfected with Vsg. Representative images were from one of three independent experiments. Scale bar = 100 μm.
Extended Data Fig. 6 Alignment of Vsg, Vsg orthologs, and an artificial receptor.
a, The protein sequences of the indicated Vsg orthologs were aligned in Mega-X through the CLUSTWS method (Drosophila melanogaster: NP_729535.1; Aedes aegypti: XP_021708011.1; Anopheles gambiae: XP_001688320.1; Tribolium castaneum: XP_972460.1; Homo sapiens: NP_006007.2; Galleria mellonella: XP_026748881.1; Spodoptera frugiperda: KAF9794377.1). b, Percentages of T/S residues within the PTS region in Vsg (residues 41–134 in D. melanogaster Vsg, between two arrows) and the corresponding region in other Vsg orthologs and the artificial receptor were counted and listed.
Extended Data Fig. 7 Expression of Vsg orthologs in U2OS cells and rescue of vsg KO S2 cells with Vsg and aVsg.
a, GFP fused Vsg, gVsg, sVsg, TMEM123, or CD164 were transfected into U2OS cells (GFP was inserted after the signal peptide of Vsgs). Cells were fixed and immunostaining was carried out using an anti-GFP antibody without permeabilization of cell membranes. Representative images from one of three independent experiments were shown. Scale bar = 20 μm. b, vsg KO2 S2 cells were transiently transfected with plasmids encoding blue fluorescent protein (BFP) or BFP plus Vsg or agVsg. Cells were then incubated with 25 pM pTc for 20 h, fixed, and stained with Alexa Fluor-647-conjugated phalloidin. BFP marked transfected cells. The percentages of transfected cells with toxin-induced actin clustering phenotype were plotted. Representative images from one of three independent experiments were shown. Scale bar = 50 μm. c, vsg KO2 S2 cells (expressing GFP from the Vsg gRNA plasmid) were transiently transfected with BFP or BFP + agVsg plasmids, then treated with 30 pM pTc for 3 days. Cells were recorded directly under confocal. The percentages of transfected cells to become enlarged were plotted. Representative images from one of three independent experiments were shown. Scale bar = 50 μm. Data were analyzed from the total number of images indicated in the bar graphs from three experiments and shown as mean ± SD.
Extended Data Fig. 8 O-glycosylation contributes to pTc binding and mass spectrometry analysis of O-glycans on Vsg and sVsg.
a, Vsg-ECD treated with O-glycosidase and PNGase F showed reduced molecular weight. b, Rounding of U2OS cells transiently expressing Vsg-GFP fusion proteins and Vsg-GFP with two N-glycan site mutations (N40T and N121S) via transient transfection. Cells were exposed to pTc for 24 h. The percentage of rounded GFP-positive cells was quantified and plotted. Data were analyzed from the total number of images indicated in the bar graphs from three experiments and shown as mean ± SD. c, Characterization of TcdA1 binding to equal amounts of immobilized Vsg-ECD, Vsg-ECD treated with PNGase F, and Vsg-ECD treated with O-glycosidase, using the BLI assay. d, Vsg-ECD and sVsg-ECD were purified as Fc-tagged proteins in S2 cells. β-eliminated glycans were permethylated and analyzed using mass spectrometry. Mass peaks were manually assigned to the indicated O-glycan moieties. e, The O-glycosylaton sites in Vsg-ECD and sVsg-ECD were predicted using NetOGlyc (http://www.cbs.dtu.dk/services/NetOGlyc/) and marked in red.
Extended Data Fig. 9 Vsg expression in D. melanogaster tissues.
a, Schematic illustration of the GFP-Vsg gene fusion in fly strain 50812. The protein-trap constructs P{PTT-GA} carry an Avic\GFP vital fluorescent protein-trap marker. GFP is inserted into the first intron of vsg, resulting in a fusion of GFP between residues 24 and 25 of Vsg. b, GFP-Vsg (green) was detected in larval ovary tissue and co-localized with phalloidin labeled ring canal structures (red). It was also detected in ring canals across the cell membrane marked by Hts staining (red) in larval lymph gland, eye disks and brain tissues. Hoechst dye marks the nuclei (blue). Scale bar = 10 μm. c, GFP-Vsg was detected in haemocytes and fat body cells. Hoechst dye marks the nuclei (blue). The edge of a fat body cell is marked. Scale bar = 10 μm. Representative images were from one of three independent experiments.
Extended Data Fig. 10 Haemocytes from vsg KO fly are resistant to pTc.
a–b, Haemocytes from the control strain (nos-Cas9;attP2) or vsg KO1 D. melanogaster were exposed to the indicated concentrations of pTc for 18 h. Cells were fixed and stained with phalloidin (red) and Hoechst (blue). Representative images (a), and quantification of the percentage of cells with actin clustering phenotype were shown in (b). Scale bar = 10 μm. c–d, Haemocytes from a vsg KO fly line (Hml-Gal4,UAS-EGFP; KO2) and a rescue line that expresses aVsg in haemocytes (Hml-Gal4,UAS-EGFP/UAS-avsg; KO2) were exposed to 100 pM pTc for 18h. Cells were fixed and stained with phalloidin (red). Representative images (c), and quantification of the percentage of cells with actin clustering were shown in (d). Scale bar = 20 μm. e–f, Haemocytes from Control or vsg KO2 D. melanogaster were treated with pTc (25 pM, 18 h) and then co-incubated with fluorescently labelled E. coli bioparticles (red) for 30 min. Cells were then fixed and stained with Alexa Fluor-488-conjugated phalloidin (green) (e). E. coli bioparticles per cell were counted and plotted in (f). Scale bar = 10 μm. Data were analyzed from the total number of images indicated in the bar graphs from three experiments and shown as mean ± SD.
Supplementary information
Supplementary Information
This file contains Supplementary Figs. 1 and 2 and Supplementary Data 1.
Rights and permissions
Springer Nature or its licensor holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Xu, Y., Viswanatha, R., Sitsel, O. et al. CRISPR screens in Drosophila cells identify Vsg as a Tc toxin receptor. Nature 610, 349–355 (2022). https://doi.org/10.1038/s41586-022-05250-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-022-05250-7
- Springer Nature Limited
This article is cited by
-
Stepwise assembly and release of Tc toxins from Yersinia entomophaga
Nature Microbiology (2024)
-
Yersinia entomophaga Tc toxin is released by T10SS-dependent lysis of specialized cell subpopulations
Nature Microbiology (2024)
-
Engineered targeting OIP5 sensitizes bladder cancer to chemotherapy resistance via TRIP12-PPP1CB-YBX1 axis
Oncogene (2024)
-
Integrating genome-wide CRISPR screens and in silico drug profiling for targeted antidote development
Nature Protocols (2024)