Abstract
Microbial colonization of the mammalian intestine elicits inflammatory or tolerogenic T cell responses, but the mechanisms controlling these distinct outcomes remain poorly understood, and accumulating evidence indicates that aberrant immunity to intestinal microbiota is causally associated with infectious, inflammatory and malignant diseases1,2,3,4,5,6,7,8. Here we define a critical pathway controlling the fate of inflammatory versus tolerogenic T cells that respond to the microbiota and express the transcription factor RORγt. We profiled all RORγt+ immune cells at single-cell resolution from the intestine-draining lymph nodes of mice and reveal a dominant presence of T regulatory (Treg) cells and lymphoid tissue inducer-like group 3 innate lymphoid cells (ILC3s), which co-localize at interfollicular regions. These ILC3s are distinct from extrathymic AIRE-expressing cells, abundantly express major histocompatibility complex class II, and are necessary and sufficient to promote microbiota-specific RORγt+ Treg cells and prevent their expansion as inflammatory T helper 17 cells. This occurs through ILC3-mediated antigen presentation, αV integrin and competition for interleukin-2. Finally, single-cell analyses suggest that interactions between ILC3s and RORγt+ Treg cells are impaired in inflammatory bowel disease. Our results define a paradigm whereby ILC3s select for antigen-specific RORγt+ Treg cells, and against T helper 17 cells, to establish immune tolerance to the microbiota and intestinal health.
Similar content being viewed by others
Main
The mammalian gastrointestinal tract is continuously colonized with microbiota, opportunistic microorganisms or pathogens, which induce robust responses that are often dominated by immune cells expressing the lineage-specifying transcription factor RORγt1,2,3,4,5,6,7,8,9,10,11,12,13,14,15,16,17. Depending on the adopted fate, these RORγt+ cells orchestrate immunity, inflammation or tolerance in the intestine9,10,11,12,13,14,15,16,17, and substantial alterations in the numbers or function of RORγt+ immune cells occur in multiple chronic human diseases, including inflammatory bowel disease (IBD), human immunodeficiency virus infection and cancer1,2,3,4,5,6,7,8,9,10,11,12,13,14,15,16,17,18. Despite these advances, the full spectrum of cellular heterogeneity among RORγt+ immune cells, the potential for functional interactions among subsets, and the pathways that are necessary to establish immune tolerance in the context of a complex microbiota remain poorly defined.
Defining all RORγt+ cells in the mLN
RORγt+ cell types include T helper 17 (TH17) cells, Treg cells, γδ T cells and ILC3s, as well as a few other potential cell types that were only recently characterized17,19,20. To uncover the full spectrum of cellular heterogeneity and look for pathways impacting tolerance or inflammation in response to microorganisms, we performed single-cell RNA sequencing (scRNA-seq) on all RORγt+ cells from the intestine-draining mesenteric lymph nodes (mLN) of healthy RORγt–eGFP reporter mice (Extended Data Fig. 1a). T cells are dominant in this tissue, and we therefore sequenced an equal ratio of T cells (GFP+TCRβ+) to non-T cells (GFP+TCRβ−) to increase the resolution of rare RORγt+ cell types. This revealed 14 distinctive clusters, each annotated on the basis of select marker genes and visualized by uniform manifold approximation and projection (UMAP; Fig. 1a). Two clusters were excluded from future analyses, including cluster 8, which was identified as a doublet population (Extended Data Fig. 1b), and cluster 7, which was identified as B cells but could not be verified by RORγt reporter or fate-mapping approaches (Extended Data Fig. 1c,d), probably representing a contaminant. The remaining clusters exhibited expression of RORγt and could be divided into T cells versus non-T cells on the basis of expression of Cd3e (Extended Data Fig. 1e,f). T cell subsets were dominated by RORγt+ Treg cells defined by expression of Trac, Cd4 and Foxp3 in clusters 0, 5 and 13, followed by TH17 cells expressing Trac, Cd4 and Il17a in cluster 1, and γδ T cells expressing Trdc in clusters 4 and 9 (Fig. 1b and Extended Data Fig. 1g). Among the non-T cells, nearly all clusters were ILC3s, or ILC3-like cells, on the basis of expression of Id2 and Il7r, including a dominant population of lymphoid tissue inducer (LTi)-like ILC3s that express Ccr6 and Cd4 in cluster 2, T-bet+ ILC3s that express Ncr1 in clusters 3 and 12, and a minor population of group 2 innate lymphoid cell (ILC2)-like cells that express high levels of Gata3 and Il17rb in cluster 10 (Fig. 1c). Subsets of conventional type 2 dendritic cells were suggested to express RORγt in the spleen20, but in a recent report we were unable to identify this population in the intestine21. Further, we could not identify this population in our single-cell analyses of the mLN and found minimal RORγt in conventional type 2 dendritic cells in this tissue as determined by flow cytometry on reporter or fate-mapping mice (Extended Data Fig. 1c,d,h–k). Instead, the remaining two non-T cell clusters (cluster 6 and 11) were defined by expression of Aire (Fig. 1c), most closely resembling extrathymic AIRE-expressing cells (eTACs) that several groups demonstrated are RORγt+, are enriched in lymph nodes during early-life developmental windows, and share transcriptional similarities with ILC3s, dendritic cells and thymic epithelial cells19,22.
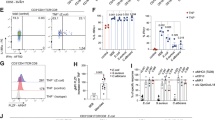
a, UMAP of scRNA-seq data from RORγt+ cells in mLN from healthy RORγt–eGFP reporter mice. TCRβ+GFP+ and TCRβ−GFP+ cells pooled from 3 mice and 12,948 cells were sequenced. b,c, Dot plots showing the mean expression (colour) of the indicated genes in clusters grouped by adaptive lymphocytes (b) and innate lymphocytes (c). The dot size represents the proportion of cells in a cluster with the gene detected. d, A volcano plot of differentially expressed genes between cluster 2 and cluster 6 of the scRNA-seq dataset. FC, fold change. e, Histograms showing the expression levels of the indicated proteins in LTi-like ILC3s (black lines) and RORγt+ eTACs (red shaded lines) from the mLN of RORγt–eGFP reporter mice. Max., maximum. f, Quantitative PCR analysis of Aire expression in sort-purified LTi-like ILC3s and RORγt+ eTACs from mLN, relative to Hprt (n = 3). g,h, Frequencies of ‘fate-mapped’ LTi-like ILC3s and RORγt+ eTACs in the mLN of CD127-Cre (n = 3), CD11c-Cre (n = 3), CLEC9A-Cre (n = 4) (g) and AIRE-Cre fate-mapping mice (n = 4) (h). ND, not detected. The data in f–h are representative of two or three independent experiments, shown as means ± s.e.m.; the statistics shown in f–h were obtained by unpaired Student’s t-test (two-tailed). The statistics in d and Supplementary Table 1 were obtained by the Wilcoxon test as implemented by Seurat; red dots are significantly different.
Given this heterogeneity among innate RORγt+ immune cells, we next sought to understand the phenotype and functions of ILC3s versus RORγt+ eTACs. Differential analysis of LTi-like ILC3s (cluster 2) and the dominant RORγt+ eTACs (cluster 6) revealed distinct gene signatures (Fig. 1d). Further validation revealed that LTi-like ILC3s are defined by high expression levels of CD127, CD25, CD122, LY6A (also known as SCA1) and CXCR6, whereas RORγt+ eTACs are defined by high expression levels of SIGLECG, DPP4 and the integrin α4β7 (Fig. 1e and Extended Data Fig. 2a–c). Notably, LTi-like ILC3s express significantly higher levels of RORγt than eTACs (Fig. 1d,e and Extended Data Fig. 2c). We developed a gating strategy to isolate these cell types and confirmed that expression of Aire is exclusive to RORγt+ eTACs and not observed in ILC3s (Fig. 1f and Extended Data Fig. 3a). These gating strategies do not rely on AIRE and therefore include eTAC subsets that express low levels of this transcription factor. Notably, RORγt+ eTACs did not exhibit expression of previously defined AIRE-dependent tissue-specific antigens, but rather expressed several genes associated with a transient-amplifying AIRE+ thymic epithelial cell23,24 (Supplementary Table 1). A previous study inferred that ILC3s have the potential to convert into RORγt+ eTACs in vitro following RANK–RANKL stimulation19, raising fundamental questions about the in vivo distinction of these two cell types. We determined that most RORγt+ eTACs fate-mapped positive for CD127-Cre, despite lacking CD127 staining, and exhibited limited fate-mapping for CD11c-Cre or CLEC9A-Cre, which is comparable to LTi-like ILC3s (Fig. 1g and Extended Data Fig. 3a,b). In addition, RORγt+ eTACs fate-mapped positive for AIRE-Cre, as do a minor proportion of LTi-like ILC3s in the mLN (Fig. 1h and Extended Data Fig. 3b), suggesting the potential of interconversion among these cell types. Collectively, these data define the full spectrum of innate and adaptive immune cells that express the lineage-specifying transcription factor RORγt in the mLN of healthy mice, revealing a dominance of RORγt+ Treg cells and LTi-like ILC3s, as well as determining phenotypic distinctions between RORγt+ eTACs and LTi-like ILC3s.
RORγt+ Treg cells require MHCII+ ILC3s
We next sought to look for functional interactions between innate and adaptive RORγt+ cells in mLN by examining their spatial proximity in the interfollicular zone, an area we previously found to contain most ILC3s and other RORγt+ T cells25,26. Indeed, ILC3s were rare in B cell follicles and T cell zones, but robustly present in interfollicular zones, which contrasts with more diffuse localization patterns of CD11c+ cells (Extended Data Fig. 4). Markedly, we observed that CD3−RORγt+CD127+ ILC3s were adjacent to CD3+RORγt+FOXP3+ Treg cells in the interfollicular zones, where a substantial portion of total RORγt+ Treg cells were associated with ILC3s (Fig. 2a and Extended Data Fig. 4). CD103+ dendritic cells have the potential to induce the differentiation of RORγt+ Treg cells from naive T cells27,28,29,30; however, both CD103+ and CD103− dendritic cells are redundant in promoting peripheral Treg differentiation31, indicating that other cell types regulate this process. Our previous work demonstrated that LTi-like ILC3s limit microbiota-specific TH17 cells through major histocompatibility complex class II (MHCII) and a process termed intestinal selection26,32. Therefore, we next interrogated whether LTi-like ILC3s impact RORγt+ Treg cells through MHCII and observed that LTi-like ILC3s abundantly express MHCII protein and transcripts in the mLN of healthy mice (Fig. 2b and Extended Data Fig. 5a). Further, MHCII∆ILC3 mice that were generated by crossing H2-Ab1fl/fl mice with Rorccre mice32 exhibited a highly selective deletion of MHCII in ILC3s relative to other immune cells in the draining mLN and large intestine (Fig. 2c and Extended Data Fig. 3c). In mice lacking ILC3-specific MHCII, we identified a marked and significant reduction in the frequency of RORγt+ Treg cells in both mLN and large intestine relative to littermate control mice (Fig. 2d–h). We previously reported that MHCII∆ILC3 mice exhibit spontaneous intestinal inflammation32, and consistent with this, we observe significantly increased total cell numbers of CD4+ T cells and TH17 cells within the large intestine, with a significant decrease in numbers of RORγt+ Treg cells (Extended Data Fig. 5b–e). These results were specific to MHCII+ LTi-like ILC3s, as deletion of MHCII by targeting T cells with CD4-Cre or ILC2s with interleukin-5 (IL-5)-Cre, and deletion of RORγt by targeting T-bet+ ILC3s with Ncr1-Cre, did not affect the frequency of RORγt+ Treg cells or other CD4+ T cell subsets in the large intestine (Extended Data Fig. 5f–k). RORγt+ eTACs also express MHCII as previously shown19, but Rorccre only modestly targeted deletion of MHCII on RORγt+ eTACs, and this is probably due to their significantly lower level of expression of RORγt relative to ILC3s (Figs. 1d,e and 2c and Extended Data Fig. 5a). Further, targeting of RORγt+ eTACs indicates a redundant role for this population in modulating RORγt+ Treg cells as comparable frequencies were present in RorccreAirefl/fl mice, AirecreRorcfl/fl mice and AirecreH2-Ab1fl/fl mice relative to littermate controls (Fig. 2i–k and Extended Data Fig. 5l–o). LTi-like ILC3s were also comparable in AirecreRorcfl/fl mice relative to littermates (Extended Data Fig. 5n), indicating that RORγt+ eTACs are redundant to maintain this population in homeostasis. RORγt+ Treg cells were present at comparable frequencies in Clec9acreH2-Ab1fl/fl mice relative to littermate controls, indicating a redundancy of conventional dendritic cells (Fig. 2l and Extended Data Fig. 5p). Finally, an ex vivo co-culture system revealed that RORγt+ Treg cells increased in frequency and cell number when in the presence of LTi-like ILC3s, as well as exhibiting a reduced BIM level, increased NUR77 staining and minor changes in cell death at this time point (Extended Data Fig. 6a–e), indicating that LTi-like ILC3s are sufficient to support RORγt+ Treg cells.
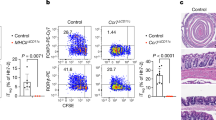
a, Left (images), tile-scanned (left) and magnified (column 2) images of mLN, and serial sections (columns 3–5) showing the same interfollicular area as in the left image stained for ILC3s (CD3−IL-7Rα+RORγt+) and RORγt+ Treg cells (CD3+IL-7Rα−FOXP3+RORγt+); the white asterisk indicates the association between ILC3s and RORγt+ Treg cells. Scale bars, 50 μm (left), 10 μm (column 2) and 2 μm (columns 3–5). Right, quantification of the percentage of RORγt+ Treg cells in association with ILC3s in the interfollicular zone (n = 3). b, Representative plots of the frequency of MHCII on LTi-like ILC3s in mLN. c, Quantification of MHCII expression on major MHCII-expressing cells from mLN and large intestine-lamina propria (LI-LP) of H2-Ab1fl/fl (n = 5 for B cells (B), monocytes (Mo), macrophages (Mf), dendritic cells (DC), conventional type I and 2 dendritic cells (cDC1 and cDC2) and neutrophils (Neu); n = 4 for ILC2, ILC3 and eTACs) and MHCIIΔILC3 mice (n = 4). d–g, Representative flow cytometry plots of the frequency (percentage of CD4+ T cells) (d,f) and quantification (e,g) of RORγt+ Treg cells in mLN (d,e) and LI-LP (f,g) of H2-Ab1fl/fl and MHCIIΔILC3 mice (n = 9). h–l, Quantification of RORγt+ Treg cells (percentage of Treg cells) in mLN of MHCIIΔILC3 mice (n = 5) (h), AirecreH2-Ab1fl/fl mice (n = 4) (i), RorccreAirefl/fl mice (n = 5) (j), AirecreRorcfl/fl mice (n = 5) (k) and Clec9acreH2-Ab1fl/fl mice (n = 4) (l) relative to littermate control mice. The data in d–g are pooled from two independent experiments. The data in h–l are representative of two independent experiments. The data are shown as mean ± s.e.m.; the statistics shown in c were obtained by multiple unpaired t-test (two-tailed); the statistics shown in e,g,h–l were obtained by unpaired Student’s t-test (two-tailed).
ILC3s select microbiota-specific Treg cells
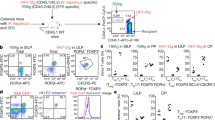
RORγt+ T cells develop in response to antigens derived from microbiota that colonize the mammalian intestine. For example, it has been demonstrated that segmented filamentous bacteria (SFB) promote antigen-specific TH17 cells33, and Helicobacter hepaticus promote antigen-specific RORγt+ Treg cells34 in the intestine of wild-type mice. Therefore, we next examined whether MHCII+ ILC3s regulate both TH17 cells and RORγt+ Treg cells that recognize distinct microbiota-derived antigens in the intestine. To accomplish this, we transferred congenically marked naive T cells from SFB-specific TCR transgenic mice and H. hepaticus-specific TCR transgenic mice into recipients that were colonized with both microorganisms (Extended Data Fig. 7a). Two weeks post transfer, we analysed the donor T cell populations and observed comparable upregulation of CD44 in recipient littermate controls and MHCIIΔILC3 mice (Extended Data Fig. 7b,c). This indicates that ILC3s are not required to prime antigen-specific T cells and is consistent with the absence of ILC3s from T cell zones of lymph nodes26 (Extended Data Fig. 4). SFB-specific T cells differentiated into TH17 cells in littermate control mice, and a significantly expanded TH17 cell population was observed in mice lacking ILC3-specific MHCII (Fig. 3a,b). The impairment of RORγt+ Treg cells is insufficient to drive expansion of effector TH17 cells, such as the SFB-specific T cell populations, as comparable intestinal TH17 cells were observed in Foxp3creRorcfl/fl mice (Extended Data Fig. 7d). By contrast, we observed that intestinal H. hepaticus-specific T cells failed to express FOXP3 in mice lacking ILC3-specific MHCII, and rather adopted a RORγt+ TH17 cell phenotype relative to those transferred to littermate controls (Fig. 3c,d and Extended Data Fig. 3d). In the absence of ILC3-specific MHCII, the H. hepaticus-specific T cells upregulated T-bet and co-produced significantly more interferon-γ (IFNγ) and IL-17A (Fig. 3e–h). Importantly, we observed a significant impairment in the differentiation of H. hepaticus-specific RORγt+ Treg cells when ILC3s were targeted in Il22creH2-Ab1fl/fl mice, and in this context, there was a partial but selective deletion of MHCII on ILC3s in the mLN and no targeting of MHCII+ eTACs (Extended Data Fig. 7e–g). These data demonstrate that ILC3-specific MHCII is necessary to limit the expansion of microbiota-specific TH17 cells as we previously described26,32, while simultaneously enforcing microbiota-specific RORγt+ Treg populations and preventing their ability to expand as pro-inflammatory TH17 cells.
a–d, Flow cytometry plots (a,c) and frequencies (b,d) of RORγt+ Treg cells (RORγt+FOXP3+) and TH17 cells (RORγt+FOXP3−) in Peyer’s patch for SFB-specific (CD45.1−CD90.1+CD4+ T cells; a,b) and in LI-LP for H. hepaticus-specific (CD45.1+CD90.1−CD4+ T cells; c,d) from H2-Ab1fl/fl and MHCII∆ILC3 mice (n = 4). e,f, Representative flow cytometry plots of the frequency (T-bet+ among H. hepaticus-specific RORγt+FOXP3− TH17 cells) (e) and quantification (f) of T-bet+ H. hepaticus-specific TH17 in LI-LP of H2-Ab1fl/fl and MHCII∆ILC3 mice as shown in c (n = 4). g,h, Representative flow cytometry plots of the frequency (IL-17A+ and/or IFNγ+ cells among H. hepaticus-specific RORγt+FOXP3− TH17 cells) (g) and quantification (h) of IFNγ+ and IFNγ+IL-17A+ cells among H. hepaticus-specific TH17 in LI-LP of H2-Ab1fl/fl and MHCII∆ILC3 mice as shown in c (n = 4). i,j, Representative flow cytometry plots of the frequency (i) and quantification (j) of MHCII expression on ILC3s, dendritic cells and eTACs (gated as CD127−CD90−) in mLN and LI-LP of MHCIIneg and MHCIIILC3+ mice (n = 6 or 9). ND, not detected. k,l, Representative flow cytometry plots of the frequency (left and middle) and quantification (right) of RORγt+ Treg cells (RORγt+FOXP3+ among H. hepaticus-specific CD4+ T cells) analysed for transgenic T cells in mLN (k) and LI-LP (l) of MHCIIneg and MHCIIILC3+ mice (n = 6). The data in a–h are representative of two independent experiments. The data in i–l are pooled from two or three independent experiments. The data are shown as mean ± s.e.m.; the statistics were obtained by unpaired Student’s t-test (two-tailed).
We also interrogated whether MHCII+ ILC3s are sufficient to promote microbiota-specific RORγt+ Treg cells by using mice in which MHCII expression is restricted to ILC3s26. Revisiting H2-Ab1-Stopfl/fl mice (MHCIIneg) and those crossed to Rorccre mice (MHCIIILC3+) revealed robust expression of MHCII restricted to ILC3s, but not on conventional dendritic cells in the mLN and large intestine, as well as a lack of MHCII on RORγt+ eTACs in the mLN that is probably due to the limited expression of Rorc (Fig. 3i,j and Extended Data Fig. 7h). We next transferred congenically marked naive T cells from H. hepaticus-specific TCR transgenic mice into recipients that were colonized with H. hepaticus as above. Two weeks post transfer, we analysed the donor T cell populations and observed comparable upregulation of CD44 and downregulation of CD62L in recipient MHCIIneg and MHCIIILC3+ mice (Extended Data Fig. 7i). This suggests that even with the lack of MHCII in this MHCIIneg mouse model, some endogenous priming of naive H. hepaticus-specific T cells remains. However, when analysing the fate of these T cells, it was clear that MHCII expression on ILC3s is sufficient for H. hepaticus-specific T cells to efficiently adopt a RORγt+ Treg fate in both the mLN and large intestine (Fig. 3k,l). These results collectively demonstrate that MHCII+ ILC3s are necessary and sufficient to critically select for the fates of RORγt+ T cells by simultaneously promoting Treg cells and limiting TH17 cells of distinct antigen specificities.
ILC3s support RORγt+ Treg cells through ITGAV
We next mechanistically examined how MHCII+ ILC3s promote RORγt+ Treg cells. Analyses of the remaining RORγt+ Treg population in MHCIIΔILC3 mice revealed a significant increase in the BIM level but comparable levels of Ki-67 (Extended Data Fig. 7j,k), suggesting that there are alterations in cell survival but not proliferation. Defined receptor–ligand analyses in our scRNA-seq data demonstrated that MHCII+ LTi-like ILC3s may interact with RORγt+ Treg cells through cytokine–cytokine receptors or processing of transforming growth factor-β (TGFβ) mediated by the integrin αV (ITGAV; Fig. 4a). Subsets of ILC3s produce IL-2 to support Treg homeostasis in the small intestine35, but ablation of IL-2 in all RORγt+ cells did not impact the presence of RORγt+ Treg cells in the large intestine (Extended Data Fig. 7l). We also previously demonstrated that ILC3-mediated sequestration of IL-2 mechanistically contributed to their ability to limit effector T cell responses26, and analyses of IL-2 binding and CD25 levels revealed that RORγt+ Treg cells were significantly more efficient at competing for this survival cytokine compared with naive or effector T cell populations (Extended Data Fig. 7m). This indicates a contribution of IL-2 competition among innate and adaptive RORγt+ cell types. We next explored integrins that mediate processing of TGFβ, as previous reports identified a critical role for dendritic cell-mediated processing of TGFβ by the integrin αVβ8 in the support of Treg cells36,37,38 and that loss of TGFβR on Treg cells results in upregulation of BIM39. We explored which β-integrins pair with ITGAV on ILC3s and found moderate levels of Itgb1, Itgb3 and Itgb5 expression on ILC3s in the mLN and large intestine, but a lack of Itgb6 and Itgb8 (Fig. 4a,b). Using our ex vivo co-culture system, we found that blockade of ITGAV or Itgb3 significantly abrogated ILC3-mediated support of RORγt+ Treg cells and suppression of TH17 cells (Fig. 4c). This indicates that ILC3s interact with RORγt+ Treg cells in part through αVβ3 integrin, which has previously been linked to processing of latent TGFβ and several other extracellular ligands40. Consistent with this, our observations show that ILC3s in the mLN and intestine stained highly for ITGAV, and that this could be successfully deleted by crossing Itgavfl/fl mice with Rorccre mice, whereas ITGAV remained intact on RORγt+ eTACs in these mice, probably owing to the limited level of expression of Rorc (Fig. 4d and Extended Data Fig. 8a). Consistent with the role we identified for MHCII+ ILC3s, our observations showed a significant reduction in the frequency of RORγt+ Treg cells and a significant increase in the frequency of TH17 cells in both the mLN and large intestine of RorccreItgavfl/fl mice relative to littermate control mice (Fig. 4e). Further, comparable frequencies of RORγt+ Treg cells were present in mLN of Cd4creItgavfl/fl mice relative to littermate control mice, suggesting that ITGAV on ILC3s, but not on CD4+ T cells, was involved in promoting RORγt+ Treg cells (Extended Data Fig. 8b–e). In addition, H. hepaticus-specific T cells failed to robustly differentiate into RORγt+ Treg cells, and rather expanded as TH17 cells in both the mLN and large intestine of RorccreItgavfl/fl mice relative to littermate controls (Fig. 4f and Extended Data Fig. 8f). Taken together, these data indicate that MHCII+ ILC3s select for antigen-specific RORγt+ Treg cells, and that this occurs in part by ILC3 expression of αVβ3 integrin, which can process latent TGFβ and interact with other extracellular ligands that could be important in this process.
a, A dot plot showing selected genes expressed in ILC3 (y axis) and Treg (x axis) cell clusters. b, Quantitative PCR analysis of Itgb1, Itgb3, Itgb5, Itgb6 and Itgb8 expression in sort-purified LTi-like ILC3s from LI-LP (n = 7) and mLN (n = 6), relative to Hprt. ND, not detected. c, Sort-purified RORγt+CD4+ T cells and LTi-like ILC3s from LI-LP and mLN (n = 12, each dot represents samples pooled from 2 mice) were co-cultured for 72 h under the indicated conditions, and the frequencies of RORγt+ Treg cells and TH17 cells were analysed by flow cytometry. d, Quantification of ITGAV on LTi-like ILC3s and eTACs in mLN (n = 5), and quantification of ITGAV on ILC3s in LI-LP, of Itgavfl/fl and RorccreItgavfl/fl mice (n = 5 or 3 per group). e, Representative flow cytometry plots of the frequency (percentage of CD4+ T cells) (left and middle panels) and quantification (right panels) of RORγt+ Treg cells and TH17 cells in mLN (top left and top middle panels) and LI-LP (bottom left and bottom middle panels) of Itgavfl/fl (n = 9) and RorccreItgavfl/fl mice (n = 8). f, Representative flow cytometry plots of the frequency (left and middle panels) and quantification (right panels) of RORγt+ Treg cells (RORγt+FOXP3+ among H. hepaticus-specific CD4+ T cells) and TH17 cells (RORγt+FOXP3− among H. hepaticus-specific CD4+ T cells) in mLN (top left and top middle panels) and LI-LP (bottom left and bottom middle panels) of Itgavfl/fl and RorccreItgavfl/fl mice (n = 4). The data in d,f are representative of two independent experiments. The data in c,e are pooled from two independent experiments. The data are shown as means ± s.e.m.; the statistics shown in c were obtained by one-way analysis of variance with Tukey’s multiple comparisons test; the statistics shown in d–f were obtained by unpaired Student’s t-test (two-tailed).
Altered ILC3–Treg interactions in IBD
IBD is a human disease characterized by chronic inflammation of the gastrointestinal tract with a substantial alteration in the numbers or functionality of ILC3s4,35,41,42,43 and RORγt+ T lymphocytes44,45,46,47,48,49. However, a comprehensive understanding of the interactions between ILC3s and RORγt+ T cells in human health and IBD is lacking. This prompted us to perform scRNA-seq on the innate lymphoid cell and T cell compartments from the inflamed intestine versus adjacent non-inflamed intestine in human IBD (Extended Data Fig. 9a). Seventeen clusters were identified and visualized by UMAP (Fig. 5a). We separated the clusters of immune cells into T cells versus non-T cells as determined by expression of CD3E (Extended Data Fig. 10a). Among non-T cells, cluster 4 was identified as ILC3s by expression of ID2, KIT and RORC (Fig. 5b), and representation of these cell types was reduced in the inflamed tissue relative to adjacent non-inflamed tissue (Extended Data Fig. 10b). This decrease of ILC3s in the inflamed human intestine was associated with a significant decrease in the expression levels of genes associated with ILC3 identity including RORC, RORA, KIT, IL1R1, CCR6 and IL22 and those involved in antigen processing and presentation including CD74, HLA-DRA, CD83, CD81, CTSH and IRF4 (Fig. 5c). Further, this impairment of ILC3s during intestinal inflammation was independently validated with colonic biopsies from a cohort of paediatric patients with Crohn’s disease relative to age-matched controls (Fig. 5d and Extended Data Fig. 9b). Among the T cells, clusters 6 and 8 were identified as Treg cells by expression of CD4 and FOXP3 (Fig. 5e). A direct comparison of differentially expressed genes revealed that cluster 6 represents RORγt+ Treg cells as they exhibited higher levels of expression of RORA, RORC, CCR6 and CXCR6, whereas cluster 8 exhibits features of thymic-derived Treg cells with a higher level of expression of HELIOS (encoded by IKZF2); Fig. 5f and Supplementary Table 2). Representation of RORγt+ Treg cells (cluster 6) was also reduced in the inflamed human intestine relative to matched non-inflamed tissue, and this was associated with a significant decrease in the expression levels of genes associated with RORγt+ Treg identity, function and TCR signalling (Fig. 5g,h). Several of these changes in abundance or transcriptional signatures among ILC3s and Treg cells were found in another published dataset from human IBD samples44 (Extended Data Fig. 10c–e). Further, a significant reduction in the frequency of RORγt+ Treg cells was independently validated in colonic biopsies from a cohort of paediatric patients with Crohn’s disease relative to age-matched controls (Fig. 5i, Extended Data Fig. 9b and Supplementary Table 3). Finally, there was a modest but significant positive correlation between the frequency of ILC3s and the frequency of RORγt+ Treg cells in all analysed intestinal biopsies (Fig. 5j and Extended Data Fig. 10f). Comparable results were validated in a second independent cohort of paediatric patients with Crohn’s disease (Extended Data Fig. 10g–i). These correlative data suggest that ILC3s promote RORγt+ Treg cells to support intestinal health in humans, and these cellular interactions become altered in IBD.
a, UMAP plots of scRNA-seq data for innate lymphoid cells and T lymphocytes from inflamed and non-inflamed intestine in human IBD. b, A dot plot showing the mean expression (colour) of the indicated genes in clusters grouped by low CD3E expression level; the dot size represents the proportion of cells in a cluster with the gene detected. c, A dot plot showing the mean expression (colour) of the indicated genes in cluster 4 as indicated in a in non-inflamed (NI) versus inflamed (Infla) tissue. d, Quantification of ILC3 frequency in a cohort of patients with IBD. Healthy donors, n = 17; patients with IBD, n = 18. e, A dot plot showing the mean expression (colour) of the indicated genes in clusters grouped by high CD3E expression. f, A volcano plot of differentially expressed genes between cluster 6 and cluster 8 as indicated in a. g, A bar graph showing the composition of T lymphocytes as indicated in a. h, A dot plot showing the mean expression (colour) of the indicated genes in cluster 6 in non-inflamed versus inflamed tissue as indicated in a. i, Quantification of the frequency of RORγt+ Treg cells among total Treg cells in a cohort of patients with IBD as indicated in d. j, Correlation between the ILC3 and RORγt+ Treg frequencies as indicated in d,i. The data in d,i are shown as means ± s.e.m.; statistics shown in d,i were performed using Mann–Whitney U-test (unpaired, two-tailed), correlative analyses in j are compared by Pearson’s rank correlation coefficient (R2). The statistics in f and Supplementary Table 2 were obtained by the Wilcoxon test as implemented by Seurat; red dots are significantly different.
Our collective datasets indicate that LTi-like ILC3s critically select for RORγt+ Treg cells and against TH17 cells to orchestrate immunologic tolerance to microbiota and maintain intestinal homeostasis (Extended Data Fig. 10j), which substantially broadens our current understanding of intestinal selection26,32 and mucosal immunology. This selection process probably occurs after initial priming by dendritic cells or other redundant antigen-presenting cells in response to colonization with microorganisms, impacts T cells with distinct antigen specificities, and requires antigen presentation through MHCII on LTi-like ILC3s with contributions from ITGAV and gradients of competition for IL-2. αVβ3 integrin could support processing of latent TGFβ or interact with other extracellular ligands. These are important findings in the emerging paradigm that there is sophisticated cross-regulation between RORγt+ lymphocytes during homeostasis, which is essential to preserve the ability to rapidly respond to intestinal injury, infection or inflammation. We further phenotypically define distinctions between ILC3s and RORγt+ eTACs. A previous study demonstrated that in vitro RANK–RANKL stimulation supports the interconversion potential of these two cell types19, raising the possibility that RORγt+ eTACs contribute to these pathways at distinct developmental windows and during infection or inflammation. However, our results demonstrate that LTi-like ILC3s are non-redundant in selecting for microbiota-specific RORγt+ Treg cells. We also identify a positive correlation between ILC3s and RORγt+ Treg cells in the human intestine, and a fundamental disruption of these cell types in IBD. Intriguingly, the cells of each of these RORγt+ populations express IL-17, IL-23R and the integrin α4β7, and it will be important to consider how targeting these molecules with available biologic therapies impacts these cellular pathways. Taken together, these findings fundamentally advance our understanding of the heterogeneity and functional interactions by which RORγt+ cells coordinate intestinal health and immune tolerance to the microbiota. Our data indicate that disruption of ILC3s in human IBD inherently underlies the dysregulation of microbiota-specific RORγt+ Treg cells and TH17 cells, thus shifting the balance from tolerance to inflammation. Defining strategies to prevent this impairment of ILC3s, or promote their expansion, may hold the key to reinstating tolerance to microbiota in human IBD. Finally, as we have shown50, it is possible that MHCII+ ILC3s could be harnessed to achieve antigen-specific tolerance through a similar induction of Treg cells in many autoimmune and related chronic inflammatory diseases.
Methods
Data reporting
Sample size estimates for animal experiments were determined using power analysis (power = 90% and α = 0.05) based on the mean and standard deviation from our previous studies and/or pilot studies using at least three animals per group. No randomization method was used in animal experiments, because littermate group allocation was performed through animal genotypes; as a result, the investigators were not blinded to allocation during experiments.
Mice
C57BL/6J mice, Thy1.1 transgenic mice, CD45.1 transgenic mice, Rosa26–loxP-flanked STOP yellow fluorescent protein gene (eYFP) mice51, H2-Ab1fl/fl mice52, Red5-Cre (Il5cre) mice53, Foxp3cre mice54, Cd4cre mice55, Itgaxcre (Cd11ccre) mice56, Clec9acre mice57, Il22cre mice58, Rorcfl/fl mice59, Itgavfl/fl mice60, SFB (7B8) TCR transgenic mice33 and H. hepaticus (HH7-2) TCR transgenic mice34 were purchased from Jackson Laboratories. Rorccre mice and Rorc(γt)–GfpTG (RORγt–eGFP) mice61 were provided by Gerard Eberl. Airecre mice62 were provided by Yong Fan. MHCIIΔILC3 mice were generated as previously described32. MHCIIILC3+ mice were generated by crossing Rorccre mice with IAbbStopfl/fl mice63 provided by Terri M. Laufer. Ncr1cre mice64 were provided by Eric Vivier (Inserm). Il7rcre mice65 were provided by David Artis (Weill Cornell Medicine) with permission from H. R. Rodewald. Il2fl/fl mice66 were provided by Kendall Smith (Weill Cornell Medicine). All mice were on a C57BL/6 background and maintained in specific-pathogen-free facilities in Weill Cornell Medicine. Male and female mice were used at 5 to 12 weeks of age. All protocols were approved by the Institutional Animal Care and Use Committee at Weill Cornell Medicine, and all experiments were performed in accordance with its guidelines.
Flow cytometry and cell sorting
Single-cell suspensions were incubated on ice with conjugated antibodies in PBS containing 2% FBS and 1 mM EDTA. Unlabelled anti-CD16/32 (clone 2.4G2, BD Biosciences) was used to block Fc receptors when analysing myeloid cells. Dead cells were excluded with Fixable Aqua Dead Cell Stain (Thermo Fisher Scientific). The staining antibodies for flow cytometry were mainly purchased from Thermo Fisher Scientific, Biolegend or BD Biosciences. The following were used for mouse cell-surface staining: CCR6 (29-2L17), NKp46 (29A1.4), CD3ε (145-2C11), CD4 (RM4-5, GK1.5), CD5 (53-7.3), CD8α (53-6.7), CD11b (M1/70), CD11c (N418), CD19 (1D3), Gr1 (RB6-8C5), LY6G (1A8), CD45R (also known as B220; RA3-6B2), CD45.1 (A20), CD45.2 (104), CD45 (30-F11), CD64 (X54-5/7.1), CD90.1 (OX-7), CD90.2 (30-H12), CD127 (A7R34), CD51 (RMV-7), F4/80 (BM8), FcεR1α (MAR-1), MHCII (M5/114.15.2), NK1.1 (PK136), TCRγδ (GL3), KLRG1 (2F1/KLRG1), CD44 (IM7), CD62L (MEL-14), CD25 (PC61), CXCR6 (SA051D1), SIGLECG (SH1), CD26 (H194-112), NUR77 (12.14), CD172α (P84), XCR1 (ZET), LY6C (HK1.4), CD132 (TUGm2), CD138 (281-2), CD117 (ACK2), CD122 (TM-β1), SCA1 (D7) and the integrin α4β7 (DATK32). The following were used for mouse intracellular staining: FOXP3 (FJK-16S), GATA3 (L50-823), IL-17A (17B7), IFNγ (XMG1.2), Ki-67 (SolA15), BIM (C34C5), RORγt (B2D) and T-bet (4B10). Lineage markers for mouse were as follows: CD3ε, CD5, CD19, B220, GR1, NK1.1, CD11b and CD11c, unless otherwise indicated. The following were used for cell-surface staining of human samples: CD3 (UCHT1), CD11c (3.9), CD14 (TuK4), CD19 (HIB19), CD34 (581), CD4 (SK3), CD45 (HI30), CD25 (BC96), CD94 (DX22), CD117 (104D2), CD123 (6H6), CD127 (A019D5), FcεR1 (AER-37(CRA1)) and NKp44 (44.189). The following were used for intracellular staining of human samples: FOXP3 (PCH101) and HELIOS (22F6). Human RORγt+ Treg cells were gated as CD45+CD3+CD4+FOXP3+HELIOS−RORγt+; human ILC3s were gated as CD45+CD3−CD11c−CD14−CD19−CD34−CD94−CD123−FcεR1−CD127+CD117+.
For intracellular staining, cells were fixed and permeabilized with FOXP3/Transcription Factor Staining Buffer Set following the manufacturer’s instructions (Thermo Fisher Scientific). Briefly, cells were incubated with FOXP3 Fixation/Permeabilization working solution for 30 min at room temperature or overnight at 4 °C, and then stained for intracellular targets by incubating with conjugated antibodies in 1× permeabilization buffer for 30 min at room temperature. For intracellular cytokine staining, cells were first incubated for 4 h in RPMI with 10% FBS, 50 ng ml−1 phorbol 12-myristate 13-acetate, 750 ng ml−1 ionomycin and 10 μg ml−1 brefeldin A, all obtained from Sigma-Aldrich. Antibodies for flow cytometry were purchased from BioLegend, Thermo Fisher Scientific or BD Biosciences. IL-2 binding capacity was assessed using a biotinylated IL-2 fluorokine assay kit (R&D Systems), following the manufacturer’s instructions. Flow cytometry data were collected using an LSR Fortessa (BD Biosciences) and analysed with FlowJo V10 software (Tree Star). Cell sorting was performed with an Aria II (BD Biosciences).
Quantitative PCR
Sort-purified cells were lysed in RLT buffer (Qiagen). RNA was extracted via RNeasy mini kits (Qiagen), as per the manufacturer’s instructions. Reverse transcription of RNA was performed using Superscript reverse transcription according to the protocol provided by the manufacturer (Thermo Fisher Scientific). Real-time PCR was performed on cDNA using SYBR green chemistry (Applied Biosystems). Reactions were run on a real-time PCR system (ABI 7500; Applied Biosystems). Data for samples were normalized to Hprt and are shown as a fold change compared with the controls.
Preparation of single-cell suspensions from intestine or lymph nodes
Small intestine, caecum and colon were removed, opened longitudinally and rinsed with ice-cold PBS. Peyer’s patches were removed from small intestine. Dissected intestinal tissues were cut into pieces of approximately 1.5 cm, and intestinal epithelial cells were dissociated by incubating in HBSS (Sigma-Aldrich) containing 5 mM EDTA (Thermo Fisher Scientific), 1 mM dithiothreitol (Sigma-Aldrich) and 2% FBS with shaking at 200 r.p.m. for 20 min at 37 °C. Dissociation of epithelial cells was performed twice. Samples were vortexed and rinsed with PBS after each step. The epithelial cell fraction was discarded. Remaining tissues were enzymatically digested in RPMI containing 0.4 U ml−1 dispase (Thermo Fisher Scientific), 1 mg ml−1 collagenase III (Worthington), 20 μg ml−1 DNase I (Sigma-Aldrich) and 10% FBS on a shaker for 45 min at 37 °C. Leukocytes were enriched by 40% (for flow cytometry) or 40%/80% (for sorting) Percoll (GE Healthcare) gradient centrifugation. mLN were chopped and incubated in RPMI containing 2% FBS and 2 mg ml−1 collagenase D (Sigma-Aldrich) with shaking at 200 r.p.m. for 20 min at 37 °C, and cells were then dissociated using a Pasteur pipette, and filtered through a 70-μm cell strainer in PBS containing 0.5 mM EDTA and 2% FBS.
Adoptive transfer of microbiota-specific TCR transgenic T cells
Recipient mice (CD45.2+, CD90.2+) were colonized with H. hepaticus (ATCC 51449/Hh3B1) by oral gavage 7 days before T cell transfer as described previously34. In some experiments, antibiotics (1 g l−1 ampicillin, 1 g l−1 colistin and 5 g l−1 streptomycin in drinking water for 3 days) were applied to permit robust colonization of H. hepaticus. SFB is continuously colonizing mice in our facility. Naive CD4+ T cells of H. hepaticus-specific (CD45.1+, CD90.2+) and SFB-specific (CD45.2+, CD90.1+) mice were isolated from spleen and lymph nodes by a naive CD4+ T cell isolation kit following the manufacturer’s instructions (Miltenyi Biotech) or by a FACSAria cell sorter (BD Biosciences) and were gated as CD45+CD3+CD4+CD25−CD44lowCD62Lhi. Recipient mice received 10,000–100,000 cells per mouse of both TCR transgenic CD4+ T cell types retro-orbitally and were analysed 2 weeks after transfer.
Immunofluorescence and image analysis
Tissue sections from experimental mice were cut and stained as described previously67,68. Briefly, 6-μm-thick sections of tissue were cut, fixed in cold acetone at 4 °C for 20 min and then stored at −20 °C before staining. The detection of RORγt in frozen tissue sections using immunofluorescence has been described previously69. Antibodies to the following mouse antigens were used: purified Armenian hamster anti-mouse CD3 (clone 145-2C11, 1:100, Biolegend), CD11c (clone HL3 - isotype, 1:100, BD Pharmingen), rat anti-mouse IL-7Rα eFluor660 (clone A7R34, 1:25, Thermo Fisher), biotin anti-mouse FOXP3 (clone FJK-16s, 1:50, Thermo Fisher) and rat anti-mouse RORγt (clone AFKJS-9, 1:25, Thermo Fisher). Detection of RORγt expression required amplification of the signal as described previously69. Purified RORγt antibodies were detected with donkey anti-rat IgG–FITC (1:150, Jackson ImmunoResearch), then rabbit anti-FITC–AF488 (1:200, Invitrogen) and then donkey anti-rabbit IgG–AF488 (1:200, Invitrogen). Purified anti-CD3 antibodies were detected with DyLight 594 goat anti-Armenian hamster IgG (clone Poly4055, 1:200, Biolegend), and biotinylated FOXP3 antibodies were detected with SA-AF555 (1:500, Invitrogen). Sections were counterstained with 4′,6-diamidino-2-phenylindole (DAPI; Invitrogen) and mounted using ProLong Gold (Invitrogen). Slides were analysed on a Zeiss 780 Zen microscope (Zeiss). High-resolution (×63) images of the interfollicular areas were taken. RORγt+ Treg cells were enumerated using the Zen software (Zeiss), by examining 8 to 12 images of interfollicular zones from at least 3 individual mice. Each RORγt+ Treg in the image was then quantified as being either adjacent or not adjacent to an ILC3 on the basis of observed co-localization of markers on the surface membrane (CD3 for T cell membrane, and IL-7R but not CD3 for ILC3 membrane, in addition to intracellular transcription factors).
Ex vivo ILC3 and T cell culture
A T cell stability assay was conducted as previously reported70 with minor modifications. Sort-purified RORγt+ CD4+ T cells (CD45+RORγtGFP+CD5+CD3+CD4+) and ILC3s (CD45+RORγtGFP+CD3−CD5−CD19−B220−TCRγδ−Gr1−NK1.1−CD11b−CD11c−CD127+CCR6+) were plated in a round-bottom 96-well plate at a 1:1 or 2:1 ratio (104 to 105 cells per well). In some experiments, technical replicates were necessarily used instead of biological replicates when cell numbers were limited. No statistical analyses were made when using technical replicates. Large intestine and mLN were used to collect RORγt+ CD4+ T cells and ILC3s. Cells were incubated in RPMI, supplemented with 10% FBS, 1 mM sodium pyruvate, 10 mM HEPES, 2 mM GlutaMax, 80 μM 2-mercaptoethanol, 100 U ml−1 penicillin, 100 μg ml−1 streptomycin (all from Thermo Fisher Scientific), 10 ng ml−1 recombinant IL-7 (Thermo Fisher Scientific), anti-integrin αV (Biolegend), anti-integrin β1 (BD Biosciences), anti-integrin β3 (BD Biosciences) and anti-integrin αVβ6 (MilliporeSigma) at 10 μg ml−1 at 37 °C and 5% CO2 for 72 h. Cells were analysed by flow cytometry.
scRNA-seq
CD45+TCRβ+GFP+ (T cells) and CD45+TCRβ−GFP+ (non-T cells) were sorted from the mLN of RORγt–eGFP mice and mixed equally at a 1:1 ratio to enrich the non-T cell populations. CD45+CD19−CD14−CD123−FcεRIα−CD34−CD94−CD4−CD127+ innate lymphoid cells and CD45+CD3+ T cells were sorted from resected inflamed tissue from a patient with Crohn’s disease versus adjacent tissue and mixed at a 1:4 ratio. For human IBD samples, 13,072 cells from non-inflamed tissue and 14,104 cells from inflamed tissue were sequenced and reported in the datasets. scRNA-seq libraries were generated using the 10x Genomics Chromium system with 3′ version 3 chemistry. Libraries were sequenced on an Illumina NovaSeq instrument. Reads were processed by Cell Ranger version 3.1.0 (10x) using the mm10 reference genome, resulting in a filtered HDF5 file. scRNA-seq data were further processed and analysed using R version 3.6.3 (R Core Team 2020) and the Seurat package version 3.2.3 (ref. 71). Specifically, Cell Ranger output was imported using the Read10X_h5 function. Seurat objects were created using only genes appearing in at least three cells. Cells were further filtered to exclude those with fewer than 600 genes detected, more than 5,000 genes detected, or more than 10% mitochondrial reads. Read counts were then normalized using the NormalizeData function. The graph representing cells with similar expression patterns was generated with the FindNeighbors function using the 20 largest principal components. Cell clusters were generated using the Louvain algorithm implemented by the FindClusters function with the resolution parameter equal to 0.4. Marker genes for each cluster were determined using the Wilcoxon test on the raw counts, implemented by the function FindAllMarkers, and including only positive marker genes with log fold changes greater than 0.25 and Bonferroni-corrected P values less than 0.01. Cluster names were determined by manual inspection of the lists of cluster marker genes. Dimensionality reduction by UMAP was performed using the RunUMAP function with the 20 largest principal components. All visualizations of scRNA-seq data were generated using the Seurat package as well as ggplot2 version 3.3.3 (ref. 72).
Cell–cell interaction network
Selected genes expressed in ILC3 clusters (2 and 12) as well as selected genes expressed in Treg clusters (0, 5 and 13) were submitted to the STRING database73. The resulting list of known protein–protein interactions was filtered to include only entries with either database annotations (database_annotated) or experimental evidence (experimentally_determined_interaction).
Human intestinal tissue isolation
De-identified surgically resected intestinal tissues from an adult individual with Crohn’s disease, or intestinal biopsies from the colon of paediatric individuals with Crohn’s disease and sex- and age-matched controls without IBD, were obtained from the JRI IBD Live Cell Bank Consortium at Weill Cornell Medicine following protocols approved by the Institutional Review Board (protocol number 1503015958). Informed consent was obtained from all subjects. Biopsies were cryopreserved in 90% FBS and 10% dimethylsulfoxide for future side-by-side comparison. Following thawing, tissues were incubated in 0.5 mg ml−1 collagenase D and 20 mg ml−1 DNase I for 1 h at 37 °C with shaking. After digestion, remaining tissues were further dissociated mechanically by a syringe plugger. Cells were filtered through a 70-mm cell strainer and used directly for staining.
Statistics
P values for datasets were determined by an unpaired two-tailed Student’s t-test with a 95% confidence interval for comparison between two independent groups. For repeated comparison between two groups, a multiple unpaired two-tailed t-test was applied. For datasets that were not normally distributed, a Mann–Whitney U-test was performed. One-way analysis of variance followed by Tukey’s multiple comparison test was used for comparison between more than two groups. Simple linear regression was used to analyse correlation. All statistical analyses were performed with GraphPad Prism version 9 (GraphPad Software Inc.). Investigators were not blinded to group allocation during experiments.
Reporting summary
Further information on research design is available in the Nature Research Reporting Summary linked to this article.
Data availability
All data necessary to understand and evaluate the conclusions of this paper are provided in the manuscript and supplementary materials. scRNA-seq data have been deposited in the Gene Expression Omnibus database under the accession numbers GSE184175 and GSE184291. A reanalysis was performed on publicly available scRNA-seq data with the accession number GSE134809. Source data are provided with this paper.
References
Honda, K. & Littman, D. R. The microbiota in adaptive immune homeostasis and disease. Nature 535, 75–84 (2016).
Maloy, K. J. & Powrie, F. Intestinal homeostasis and its breakdown in inflammatory bowel disease. Nature 474, 298–306 (2011).
Maynard, C. L., Elson, C. O., Hatton, R. D. & Weaver, C. T. Reciprocal interactions of the intestinal microbiota and immune system. Nature 489, 231–241 (2012).
Zhou, L. & Sonnenberg, G. F. Essential immunologic orchestrators of intestinal homeostasis. Sci. Immunol. 3, eaao1605 (2018).
Belkaid, Y. & Hand, T. W. Role of the microbiota in immunity and inflammation. Cell 157, 121–141 (2014).
Hooper, L. V. & Macpherson, A. J. Immune adaptations that maintain homeostasis with the intestinal microbiota. Nat. Rev. Immunol. 10, 159–169 (2010).
Harrison, O. J. & Powrie, F. M. Regulatory T cells and immune tolerance in the intestine. Cold Spring Harb. Perspect. Biol. 5, a018341 (2013).
Janney, A., Powrie, F. & Mann, E. H. Host–microbiota maladaptation in colorectal cancer. Nature 585, 509–517 (2020).
Harrington, L. E. et al. Interleukin 17–producing CD4+ effector T cells develop via a lineage distinct from the T helper type 1 and 2 lineages. Nat. Immunol. 6, 1123–1132 (2005).
Park, H. et al. A distinct lineage of CD4 T cells regulates tissue inflammation by producing interleukin 17. Nat. Immunol. 6, 1133–1141 (2005).
Ohnmacht, C. et al. The microbiota regulates type 2 immunity through RORγt+ T cells. Science 349, 989–993 (2015).
Sefik, E. et al. Individual intestinal symbionts induce a distinct population of RORγ+ regulatory T cells. Science 349, 993–997 (2015).
Buonocore, S. et al. Innate lymphoid cells drive interleukin-23-dependent innate intestinal pathology. Nature 464, 1371–1375 (2010).
Cella, M. et al. A human natural killer cell subset provides an innate source of IL-22 for mucosal immunity. Nature 457, 722–725 (2009).
Sawa, S. et al. Lineage relationship analysis of RORγt+ innate lymphoid cells. Science 330, 665–669 (2010).
Spits, H. et al. Innate lymphoid cells — a proposal for uniform nomenclature. Nat. Rev. Immunol. 13, 145–149 (2013).
Eberl, G. RORγt, a multitask nuclear receptor at mucosal surfaces. Mucosal Immunol. 10, 27–34 (2017).
Goc, J. et al. Dysregulation of ILC3s unleashes progression and immunotherapy resistance in colon cancer. Cell 184, 5015–5030 (2021).
Yamano, T. et al. Aire-expressing ILC3-like cells in the lymph node display potent APC features. J. Exp. Med. 216, 1027–1037 (2019).
Cabeza-Cabrerizo, M., Cardoso, A., Minutti, C. M., Pereira da Costa, M. & Reis e Sousa, C. Dendritic cells revisited. Annu. Rev. Immunol. 39, 131–166 (2021).
Zhou, W. et al. ZBTB46 defines and regulates ILC3s that protect the intestine. Nature 609, 159–165 (2022).
Gardner, J. M. et al. Extrathymic Aire-expressing cells are a distinct bone marrow-derived population that induce functional inactivation of CD4+ T cells. Immunity 39, 560–572 (2013).
Wells, K. L. et al. Combined transient ablation and single-cell RNA-sequencing reveals the development of medullary thymic epithelial cells. eLife 9, e60188 (2020).
Anderson, M. S. et al. Projection of an immunological self shadow within the thymus by the aire protein. Science 298, 1395–1401 (2002).
Mackley, E. C. et al. CCR7-dependent trafficking of RORγ+ ILCs creates a unique microenvironment within mucosal draining lymph nodes. Nat. Commun. 6, 5862 (2015).
Hepworth, M. R. et al. Group 3 innate lymphoid cells mediate intestinal selection of commensal bacteria-specific CD4+ T cells. Science 348, 1031–1035 (2015).
Coombes, J. L. et al. A functionally specialized population of mucosal CD103+ DCs induces Foxp3+ regulatory T cells via a TGF-beta and retinoic acid-dependent mechanism. J. Exp. Med. 204, 1757–1764 (2007).
Farache, J. et al. Luminal bacteria recruit CD103+ dendritic cells into the intestinal epithelium to sample bacterial antigens for presentation. Immunity 38, 581–595 (2013).
Sun, C. M. et al. Small intestine lamina propria dendritic cells promote de novo generation of Foxp3 T reg cells via retinoic acid. J. Exp. Med. 204, 1775–1785 (2007).
Tanoue, T., Atarashi, K. & Honda, K. Development and maintenance of intestinal regulatory T cells. Nat. Rev. Immunol. 16, 295–309 (2016).
Russler-Germain, E. V. et al. Gut Helicobacter presentation by multiple dendritic cell subsets enables context-specific regulatory T cell generation. eLife 10, e54792 (2021).
Hepworth, M. R. et al. Innate lymphoid cells regulate CD4+ T-cell responses to intestinal commensal bacteria. Nature 498, 113–117 (2013).
Yang, Y. et al. Focused specificity of intestinal TH17 cells towards commensal bacterial antigens. Nature 510, 152–156 (2014).
Xu, M. et al. c-MAF-dependent regulatory T cells mediate immunological tolerance to a gut pathobiont. Nature 554, 373–377 (2018).
Zhou, L. et al. Innate lymphoid cells support regulatory T cells in the intestine through interleukin-2. Nature 568, 405–409 (2019).
Belkaid, Y. & Oldenhove, G. Tuning microenvironments: induction of regulatory T cells by dendritic cells. Immunity 29, 362–371 (2008).
Worthington, J. J., Czajkowska, B. I., Melton, A. C. & Travis, M. A. Intestinal dendritic cells specialize to activate transforming growth factor-β and induce Foxp3+ regulatory T cells via integrin αvβ8. Gastroenterology 141, 1802–1812 (2011).
Worthington, J. J. et al. Integrin αvβ8-mediated TGF-β activation by effector regulatory T cells is essential for suppression of T-cell-mediated inflammation. Immunity 42, 903–915 (2015).
Ouyang, W., Beckett, O., Ma, Q. & Li, M. O. Transforming growth factor-beta signaling curbs thymic negative selection promoting regulatory T cell development. Immunity 32, 642–653 (2010).
Sarrazy, V. et al. Integrins αvβ5 and αvβ3 promote latent TGF-β1 activation by human cardiac fibroblast contraction. Cardiovasc. Res. 102, 407–417 (2014).
Bernink, J. H. et al. Human type 1 innate lymphoid cells accumulate in inflamed mucosal tissues. Nat. Immunol. 14, 221–229 (2013).
Teng, F. et al. A circadian clock is essential for homeostasis of group 3 innate lymphoid cells in the gut. Sci. Immunol. 4, eaax1215 (2019).
Vivier, E. et al. Innate lymphoid cells: 10 years on. Cell 174, 1054–1066 (2018).
Martin, J. C. et al. Single-cell analysis of Crohn's disease lesions identifies a pathogenic cellular module associated with resistance to anti-TNF therapy. Cell 178, 1493–1508 (2019).
Hovhannisyan, Z., Treatman, J., Littman, D. R. & Mayer, L. Characterization of interleukin-17-producing regulatory T cells in inflamed intestinal mucosa from patients with inflammatory bowel diseases. Gastroenterology 140, 957–965 (2011).
Jaeger, N. et al. Single-cell analyses of Crohn's disease tissues reveal intestinal intraepithelial T cells heterogeneity and altered subset distributions. Nat. Commun. 12, 1921 (2021).
Boland, B. S. et al. Heterogeneity and clonal relationships of adaptive immune cells in ulcerative colitis revealed by single-cell analyses. Sci. Immunol. 5, eabb4432 (2020).
Smillie, C. S. et al. Intra- and inter-cellular rewiring of the human colon during ulcerative colitis. Cell 178, 714–730 (2019).
Friedrich, M., Pohin, M. & Powrie, F. Cytokine networks in the pathophysiology of inflammatory bowel disease. Immunity 50, 992–1006 (2019).
Grigg, J. B. et al. Antigen-presenting innate lymphoid cells orchestrate neuroinflammation. Nature 600, 707–712 (2021).
Srinivas, S. et al. Cre reporter strains produced by targeted insertion of EYFP and ECFP into the ROSA26 locus. BMC Dev. Biol. 1, 4 (2001).
Hashimoto, K., Joshi, S. K. & Koni, P. A. A conditional null allele of the major histocompatibility IA-beta chain gene. Genesis 32, 152–153 (2002).
Nussbaum, J. C. et al. Type 2 innate lymphoid cells control eosinophil homeostasis. Nature 502, 245–248 (2013).
Rubtsov, Y. P. et al. Regulatory T cell-derived interleukin-10 limits inflammation at environmental interfaces. Immunity 28, 546–558 (2008).
Lee, P. P. et al. A critical role for Dnmt1 and DNA methylation in T cell development, function, and survival. Immunity 15, 763–774 (2001).
Caton, M. L., Smith-Raska, M. R. & Reizis, B. Notch–RBP-J signaling controls the homeostasis of CD8− dendritic cells in the spleen. J. Exp. Med. 204, 1653–1664 (2007).
Schraml, B. U. et al. Genetic tracing via DNGR-1 expression history defines dendritic cells as a hematopoietic lineage. Cell 154, 843–858 (2013).
Ahlfors, H. et al. IL-22 fate reporter reveals origin and control of IL-22 production in homeostasis and infection. J. Immunol. 193, 4602–4613 (2014).
Choi, G. B. et al. The maternal interleukin-17a pathway in mice promotes autism-like phenotypes in offspring. Science 351, 933–939 (2016).
Lacy-Hulbert, A. et al. Ulcerative colitis and autoimmunity induced by loss of myeloid αv integrins. Proc. Natl Acad. Sci. USA 104, 15823–15828 (2007).
Lochner, M. et al. In vivo equilibrium of proinflammatory IL-17+ and regulatory IL-10+ Foxp3+ RORγt+ T cells. J. Exp. Med. 205, 1381–1393 (2008).
Fan, Y. et al. Thymus-specific deletion of insulin induces autoimmune diabetes. EMBO J. 28, 2812–2824 (2009).
Archambault, A. S. et al. Cutting edge: conditional MHC class II expression reveals a limited role for B cell antigen presentation in primary and secondary CD4 T cell responses. J. Immunol. 191, 545–550 (2013).
Narni-Mancinelli, E. et al. Fate mapping analysis of lymphoid cells expressing the NKp46 cell surface receptor. Proc. Natl Acad. Sci. USA 108, 18324–18329 (2011).
Schlenner, S. M. et al. Fate mapping reveals separate origins of T cells and myeloid lineages in the thymus. Immunity 32, 426–436 (2010).
Popmihajlov, Z., Xu, D., Morgan, H., Milligan, Z. & Smith, K. A. Conditional IL-2 gene deletion: consequences for T cell proliferation. Front. Immunol. 3, 102 (2012).
Withers, D. R. et al. Cutting edge: lymphoid tissue inducer cells maintain memory CD4 T cells within secondary lymphoid tissue. J. Immunol. 189, 2094–2098 (2012).
Kim, S. et al. CD117+ CD3− CD56− OX40Lhigh cells express IL-22 and display an LTi phenotype in human secondary lymphoid tissues. Eur. J. Immunol. 41, 1563–1572 (2011).
Dutton, E. E. & Withers, D. R. Identification of murine and human innate lymphoid cells in frozen tissue sections using immunofluorescence. Methods Mol. Biol. 2121, 51–58 (2020).
Yang, B. H. et al. Foxp3+ T cells expressing RORγt represent a stable regulatory T-cell effector lineage with enhanced suppressive capacity during intestinal inflammation. Mucosal Immunol. 9, 444–457 (2016).
Butler, A., Hoffman, P., Smibert, P., Papalexi, E. & Satija, R. Integrating single-cell transcriptomic data across different conditions, technologies, and species. Nat. Biotechnol. 36, 411–420 (2018).
Hadley, W. ggplot2: Elegant Graphics for Data Analysis (Springer, 2016).
von Mering, C. et al. STRING: known and predicted protein–protein associations, integrated and transferred across organisms. Nucleic Acids Res. 33, D433–D437 (2005).
Acknowledgements
We thank members of the Sonnenberg Laboratory for discussions and critical reading of the manuscript. Research in the Sonnenberg Laboratory is supported by the National Institutes of Health (R01AI143842, R01AI123368, R01AI145989, U01AI095608, R21CA249274, R01AI162936 and R01CA274534), an Investigators in the Pathogenesis of Infectious Disease Award from the Burroughs Wellcome Fund, the Meyer Cancer Center Collaborative Research Initiative, the Dalton Family Foundation and Linda and Glenn Greenberg. W. Zhou, J.G., L.Z. and W. Zhang are supported by fellowships from the Crohn’s and Colitis Foundation (831404, 519428, 608975 and 901000, respectively). D.R.W. and F.G. are supported by a Senior Research Fellowship from the Wellcome Trust to D.R.W. (110199/Z/15/Z). J.G.F. is supported by P30-ES002109 and R35CA210088. G.F.S. is a CRI Lloyd J. Old STAR. We thank the Epigenomics Cores of Weill Cornell Medicine and G. Putzel for bioinformatics assistance, J. Conrad for administrative assistance, and S. Mozumder for technical assistance. The JRI IBD Live Cell Bank is supported by the JRI, the Jill Roberts Center for IBD, Cure for IBD, the Rosanne H. Silbermann Foundation, the Sanders Family and Weill Cornell Medicine Division of Pediatric Gastroenterology, Hepatology, and Nutrition.
Author information
Authors and Affiliations
Consortia
Contributions
M.L., H.S. and G.F.S. conceived the project. M.L. performed most experiments and analysed the data. H.S., L.K., F.G., J.G., W. Zhou, L.Z., W. Zhang and J.Z. helped with experiments and data analyses. J.G.F., Z.S., Y.F., T.M.L., G.E. and D.R.W. provided essential tools, scientific advice and expertise. R.E.S. and JRI Live Cell Bank contributed to clinical sample acquisition and processing. M.L. and G.F.S. wrote the manuscript, with input from all authors.
Corresponding author
Ethics declarations
Competing interests
H.S. is currently employed by EA Pharma. The other authors declare no competing interests.
Peer review
Peer review information
Nature thanks the anonymous reviewers for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 scRNA-seq profiling of RORγt+ cells from mouse mLN.
a, Gating strategy to sort GFP+TCRβ+ and GFP+TCRβ− cells (1:1, n = 3) for scRNA-seq. b, Doublet test showing cluster 8 as doublets. c, d, Representative flow cytometry plot of the frequency of RORγt+ cells in CD127+ ILC fractions (CD45+CD3ε−CD5−NK1.1−Ly6G−TCRγ/δ−B220−CD11b−CD11c−KLRG1−CD127+), CD19+ B cell fractions (CD45+CD19+) and CD172a+ cDC2 fractions (CD45+ CD3ε−CD5−NK1.1−Ly6G−TCRγ/δ−B220−CD64−CD11c+MHCII+XCR1−CD172a+) from RORγt-eGFP reporter mice (n = 3) (c) and RorccreRosa26lsl−YFP fate mapped mice (n = 3) (d). e–k, Violin plot showing the expression of Rorc (e), Cd3e (f), Foxp3 (g), Xcr1 (h), Clec9a (i), Clec10a (j), Clec12a (k) among all the identified clusters.
Extended Data Fig. 2 Characterization of RORγt+ eTACs and LTi-like ILC3s in mouse mLN.
a, Violin plot showing the expression of Siglecg and Dpp4 among all the identified clusters of non-T lymphocytes. b, Gating strategy to identify ILC3s and RORγt+ eTACs from mLN of RORγt-eGFP reporter mice (n = 4) for data shown in Fig. 1e, f. c, Quantification of indicated genes expression in LTi-like cells and RORγt+ eTACs shown in Fig. 1e (n = 4). Data in c are representative of three independent experiments. Data are shown as means ± s.e.m., statistics shown in c are obtained by unpaired Student’s t-test (two-tailed).
Extended Data Fig. 3 Gating strategies and fate mapping of RORγt+ eTACs and LTi-like ILC3s in mouse mLN.
a, Gating strategy to identify LTi-like ILC3s and RORγt+ eTACs for fate mapping analyses in Fig. 1g, h. b, Representative flow cytometry plots showing expression of CD127, CD11c, CLEC9A and AIRE among “fate-mapped” LTi-like ILC3s and RORγt+ eTACs in mLN shown in Fig. 1g, h. c, Gating strategy to identify LTi-like ILC3s, RORγt+ eTACs and RORγt+ Tregs of mLN in Fig. 2c–l, Fig. 4e, the same gating strategy applied to the LI-LP. d, Gating strategy to identify H. Hepaticus (Hh)-specific CD4+ T cells in mouse LI-LP.
Extended Data Fig. 4 Immunofluorescence and quantification of different cell types in mLN.
a, Representative flow cytometry plots of frequency (left) and percentage (right) of RORγt+FoxP3+ Tregs and RORγt+FoxP3− TH17 cells among total RORγt+CD4+ T cells in mLN of WT mice (n = 5). b, Tile-scanned (left) and magnified (right) images of mLN stained for expression of IL7Rα (red), CD3 (blue), FOXP3 (cyan), RORγt (green) and DAPI (grey). c–e, Quantification of percentage of RORγt+ Tregs among total Tregs (c), total numbers per mm2 of ILC3s (d) and RORγt+ Tregs (e) in interfollicular zone of mLN of WT-1 (n = 11 areas), WT-2 (n = 8 areas) and WT-3 (n = 12 areas) mice. f, Tile-scanned images and serial sections of mLN stained for expression of IL7Rα (red), CD11c (blue), FOXP3 (cyan), RORγt (green) and DAPI (grey). Left panel is without IL7Rα and RORγt staining, middle panel is without FOXP3 staining, and right panel is a merge with a magnified image. g, Tile-scanned (left) and magnified (right) images of mLN stained for expression of CD3 (red), FOXP3 (cyan), CD11c (green) and DAPI (grey). Scale bars: 50 μm, 20 μm (in magnified images). Data in a, b, f, g are representative of two independent experiments. Data in c–e are representative of two independent experiments containing a total of 5 mice. Data are shown as means ± s.e.m., statistics shown in a are obtained by unpaired Student’s t-test (two-tailed).
Extended Data Fig. 5 MHCII+ ILC3s selectively regulate T cell homeostasis in the gut.
a, UMAP plots of scRNA-seq data showing expression of H2-Ab1, H2-Ab1 and Cd74 are enriched in clusters of LTi-like ILC3s and RORγt+ eTACs across all the identified clusters in mouse mLN. b–e, Cell numbers of CD4+ T cells (b), TH17 cells (c), RORγt−Tregs (d) and RORγt+ Tregs (e) in mLN (upper panel) and LI-LP (lower panel) of H2-Ab1fl/fl and MHCIIΔILC3 mice (n = 9, pooled from two independent experiments). f–k, Large intestine of H2-Ab1fl/fl (n = 6) and Cd4creH2-Ab1fl/fl (MHCIIΔT cell) mice (n = 4) (f, g), H2-Ab1fl/fl (n = 4) and Il5creH2-Ab1fl/fl (MHCIIΔILC2) mice (n = 6) (h, i), Rorcfl/fl (n = 10) and Ncr1creRorcfl/fl mice (n = 9) (j, k) were analyzed. Proportion of MHCII-expressing CD4+ T cells (f), ILC2s (h) and ILC3s (j). Frequency of each subset among CD4+ T cells and RORγt+ Tregs among total Tregs (g, i, k). TH17: Foxp3−RORγt+; Treg: Foxp3+; TH1: Foxp3−RORγ−T-bet+; TH2: Foxp3−RORγt−Gata3+. l, Quantification of RORγt+ Tregs among total CD4+ T cells in LI-LP of Airefl/fl and RorccreAirefl/fl mice (n = 5). m, n, Quantification of eTACs among total CD127−CD90− cells (m) and LTi-like ILC3s among CD45+CCR6+ cells (n) in mLN of Rorcfl/fl and AirecreRorcfl/fl mice (n = 5). o, Quantification of MHCII expression among eTACs (o, left) and LTi-like ILC3s (o, right) in mLN of H2-Ab1fl/fl and AirecreH2-Ab1fl/fl mice (n = 4). p, Quantification of MHCII expression among DCs in mLN of H2-Ab1fl/fl and Clec9acreH2-Ab1fl/fl mice (n = 4). Data are representative of two independent experiments unless otherwise indicated. Data shown as mean ± s.e.m. Statistics in f, h, j, l–p, right of g, i, k are obtained by unpaired Student’s t-test. Statistics shown in left of g, i, k are obtained by multiple unpaired t-test. Statistics are calculated by two-tailed test.
Extended Data Fig. 6 LTi-like ILC3s support RORγt+ Tregs in a co-culture system.
a-c, Sort-purified RORγt+CD4+ T cells and LTi-like ILC3s from LI-LP and mLN (n = 6 or 5 or 3 per group as technical replicates) were co-cultured for 72 h and RORγt+ Tregs were analyzed by flow cytometry. Frequency and cell number of RORγt+ Tregs (a, RORγt+Foxp3+ among CD4+ T cells), MFI of Bim (b) and Nur77 (c) in RORγt+ Tregs. d, e, Dead cells were quantified in RORγt+ Tregs (d) and TH17 cells (e) after co-culture with or without LTi-like ILC3s for 72 h. RORγt+CD4+ T cells and LTi-like ILC3s were sort-purified from mLN and LI-LP and pooled for co-culture assay (n = 12, each dot represents samples pooled from 2 mice). Data in a–c are representative of two independent experiments. Data in d, e are pooled from two independent experiments. Data are shown as mean ± s.e.m., statistics shown in d, e were obtained by unpaired Student’s t-test (two-tailed).
Extended Data Fig. 7 MHCII+ LTi-like ILC3s select for microbiota specific RORγt+ Tregs.
a, H. hepaticus (Hh)-specific and/or SFB-specific CD4+ T cells were transferred to H2-Ab1fl/fl and MHCIIΔILC3 mice colonized with H. hepaticus 2 weeks before experiment as shown in Fig. 3a–h, k, l. b, c, Frequency of CD44+ ratio among SFB-specific (b) or Hh-specific (c) CD4+ T cells were analyzed in Peyer’s patch for SFB (CD45.1−CD90.1+CD4+ T cells) and in LI-LP for Hh-specific (CD45.1+CD90.1−CD4+ T cells) transgenic T cells (n = 4). d, Quantification of RORγt+ Tregs and TH17 cells among total CD4+ T cells in LI-LP of Rorcfl/fl and Foxp3creRorcfl/fl mice (n = 4). e, f, Quantification of MHCII expression on LTi-like ILC3s (n = 9) (e) and eTACs (n = 4) (f) in mLN of H2-Ab1fl/fl and Il22creH2-Ab1fl/fl mice. g, Quantification of RORγt+ Tregs (among Hh-specific CD4+ T cells) and TH17 cells (among Hh-specific CD4+ T cells) were analyzed in LI-LP of H2-Ab1fl/fl and Il22creH2-Ab1fl/fl mice (n = 9). h, Representative flow cytometry plots of the frequency of MHCII expression on ILC3s, DCs in LI-LP of MHCIIneg and MHCIIILC3+ mice (n = 6) as shown in Fig. 3j. i, Frequency of CD44hiCD62Llo ratio among Hh-specific CD4+ T cells were analyzed in mLN and LI-LP for Hh-specific CD4+ T cells as shown in Fig. 3k, l (n = 6). j, k, RORγt+ Tregs in LI-LP of H2-Ab1fl/fl and MHCIIΔILC3 mice (n = 4) were analyzed by flow cytometry. Histogram and MFI of Bim (j). Proportions of Ki-67 positive cells (k). l, Quantification of RORγt+ Tregs among CD4+ T cells in LI-LP of Il2fl/fl and RorccreIl2fl/fl mice (n = 4). m, Quantification of CD25 staining or IL-2 binding in mLN of WT mice (n = 3). Naïve T cells: CD44loCD62Lhi; effector T: CD44hiCD62Llo; TH17 cells: RORγt+FoxP3−; Tregs: FoxP3+; RORγt+Tregs: RORγt+FoxP3+. Data in e, g are pooled from two independent experiments with similar results. Data in b–d, f, i–l are representative of two independent experiments. Data are shown as mean ± s.e.m., statistics shown in m are obtained by one-way ANOVA with Tukey’s multiple comparisons test, statistics shown in b–g, i–l are obtained by unpaired Student’s t-test (two-tailed).
Extended Data Fig. 8 Itgav on LTi-like ILC3s contributes to the selection of microbiota-specific RORγt+ Tregs.
a, Representative flow cytometry plot of the frequency of Itgav expression on LTi-like ILC3s and eTACs in mLN of Itgavfl/fl and RorccreItgavfl/fl mice as shown in Fig. 4d (n = 5). b–d, Quantification of ITGAV on CD4+ T cells (b), LTi-like ILC3s (c) and eTACs (d) in mLN of Itgavfl/fl and Cd4creItgavfl/fl mice (n = 6). e, Quantification of RORγt+ Tregs in mLN of Itgavfl/fl and Cd4creItgavfl/fl mice (n = 6). f, H. Hepaticus (Hh)-specific CD4+ T cells were transferred to Itgavfl/fl and RorccreItgavfl/fl mice colonized with H. hepaticus 14 days before experiment related to Fig. 4f. Data in b–e are representative of two independent experiments. Data are shown as mean ± s.e.m., statistics shown in b–e are obtained by unpaired Student’s t-test (two-tailed).
Extended Data Fig. 9 Gating strategy for ILC3s and T cell subsets in the human intestine.
a, Gating strategy to sort ILCs and T cells from small intestine of the IBD patient for scRNA-seq in Fig. 5. b, Gating strategy to analyse ILC3s and RORγt+ Tregs by flow cytometry as shown in Fig. 5.
Extended Data Fig. 10 ILC3s select microbiota specific Tregs to establish tolerance in the gut.
a, Violin plot of CD3E expression among clusters of scRNA-seq data as shown in Fig. 5a. b, Bar graph showing the composition of non-T lymphocytes as indicated in Fig. 5a in non-inflamed tissue (NI) versus inflamed tissue (Infla). c, Bar graph showing the composition of ILC3 lymphocytes in non-inflamed tissue (NI) versus inflamed tissue (Infla) from human IBD samples as published44. d, e, A dot plot showing the mean expression (colour) of indicated genes in ILC3 cluster (d) and Treg cluster (e) in non-inflamed versus inflamed tissue from human IBD samples as published44. f, Correlation analyses between the RORγt+ Tregs (RORγt+Helios− among CD4+ T cells) and TH17 cells (RORγt+FoxP3− among CD4+ T cells) in the cohort of CD patients as in Fig. 5d, i, j. g, h, Quantification of frequency of ILC3s among CD127+CD117+ subset (g) and RORγt+ Tregs among total Tregs (h) in a second independent cohort of individuals. Healthy donor n = 15, Crohn’s disease (CD) patients n = 15. i, Correlation analyses between the ILC3 (ILC3 among CD127+CD117+ subset) and RORγt+ Tregs (RORγt+ Helios− among FoxP3+ Tregs) in a second independent cohort of human samples as in (g, h). j, LTi-like ILC3s are necessary and sufficient in selecting for the differentiation fate of microbiota specific RORγt+ Tregs, and selecting against TH17 cells, via antigen presentation with contributions from integrin αv and gradients of competition for IL-2. This collectively enforces immunologic tolerance to microbiota and maintains intestinal homeostasis. Data in g, h are shown as means ± s.e.m., statistics shown in g, h are performed using Mann–Whitney U-test (unpaired), correlative analyses in f, i are compared by Pearson’s rank correlation coefficient (R2). Statistics are calculated by two-tailed test.
Supplementary information
Supplementary Table 1
List of genes that are differentially expressed between RORγt+ eTACs and LTi-like ILC3s in the mouse mLN.
Supplementary Table 2
List of genes that are differentially expressed between RORγt+ Treg cells and RORγt- Treg cells in the human intestine.
Supplementary Table 3
Clinical metadata on paediatric patients with IBD and matched healthy controls.
Source data
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Lyu, M., Suzuki, H., Kang, L. et al. ILC3s select microbiota-specific regulatory T cells to establish tolerance in the gut. Nature 610, 744–751 (2022). https://doi.org/10.1038/s41586-022-05141-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-022-05141-x
- Springer Nature Limited
This article is cited by
-
Impaired immune tolerance mediated by reduced Tfr cells in rheumatoid arthritis linked to gut microbiota dysbiosis and altered metabolites
Arthritis Research & Therapy (2024)
-
The dynamic shifts of IL-10-producing Th17 and IL-17-producing Treg in health and disease: a crosstalk between ancient "Yin-Yang" theory and modern immunology
Cell Communication and Signaling (2024)
-
Sequential immunotherapy: towards cures for autoimmunity
Nature Reviews Drug Discovery (2024)
-
Immune microniches shape intestinal Treg function
Nature (2024)
-
The emerging family of RORγt+ antigen-presenting cells
Nature Reviews Immunology (2024)