Abstract
Pattern-triggered immunity (PTI) and effector-triggered immunity (ETI) in plants enable them to respond to pathogens by activating the production of defence metabolites that orchestrate immune responses1,2,3,4. How the production of defence metabolites is promoted by immune receptors and coordinated with broad-spectrum resistance remains elusive. Here we identify the deubiquitinase PICI1 as an immunity hub for PTI and ETI in rice (Oryza sativa). PICI1 deubiquitinates and stabilizes methionine synthetases to activate methionine-mediated immunity principally through biosynthesis of the phytohormone ethylene. PICI1 is targeted for degradation by blast fungal effectors, including AvrPi9, to dampen PTI. Nucleotide-binding domain, leucine-rich-repeat-containing receptors (NLRs) in the plant immune system, such as PigmR, protect PICI1 from effector-mediated degradation to reboot the methionine–ethylene cascade. Natural variation in the PICI1 gene contributes to divergence in basal blast resistance between the rice subspecies indica and japonica. Thus, NLRs govern an arms race with effectors, using a competitive mode that hinges on a critical defence metabolic pathway to synchronize PTI with ETI and ensure broad-spectrum resistance.
Similar content being viewed by others
Main
Plants have evolved a two-tiered immune system that consists of PTI and ETI1. PTI is controlled by pattern recognition receptors (PRRs) on the plasma membrane that recognize pathogen-associated molecular patterns (PAMPs)2. Pathogens deliver effectors to attenuate plant PTI3. Plants then evolved intracellular NLR receptors to recognize pathogen effector perturbations and activate ETI4, which have been widely adopted in crop breeding for disease resistance5. PTI and ETI have recently been proposed to be mutually potentiating6,7, and they trigger similar biosynthesis of a diverse set of defence-related secondary metabolites and phytohormones8. Methionine (Met) is the precursor of ethylene9, and pathogen effectors hijack the ethylene pathway to favour colonization10,11. Although primary metabolites have been proposed to be involved in plant defence, their immune function and the mechanisms by which NLRs guard and activate their biosynthesis are unclear. How NLRs dominate the arms race against pathogen effectors to orchestrate effective immune responses has long been a puzzle. Magnaporthe oryzae causes blast, the most destructive rice disease12. M. oryzae secretes effectors into the host cell13,14,15,16,17,18,19,20. Several NLRs that confers broad-spectrum blast resistance have been isolated21,22,23, but how these NLRs act remains largely unknown.
PICI1 functions in ETI
The rice NLR PigmR confers resistance to all fungal blast races tested21, suggesting that it potentially integrates PTI and ETI to ensure broad-spectrum resistance. To identify potential signalling regulators, we performed a combined analysis of PigmR-interacting proteins using a yeast-two hybrid (Y2H) screen (referred to as ETI-related proteins) and chitin-induced proteins by quantitative proteomics (referred to as PTI-related proteins), which yielded three overlapping candidates (Extended Data Fig. 1a, Supplementary Table 1). Among these PigmR-interacting and chitin-induced proteins (PICIs), PICI1 contains a PPPDE domain that is conserved in eukaryotes including humans24, and which is predicted to be a new class of deubiquitinatase (Extended Data Fig. 1b). PigmR specifically interacted with PICI1 through its coiled-coil domain in the Y2H screen and in a split luciferase complementation (SLC) assay (Fig. 1a, Extended Data Fig. 1c–h). PICI1–GFP co-localized with PigmR (Extended Data Fig. 1i–l) and co-immunoprecipitated with PigmR–7Myc–6His (Fig. 1b). We generated transgenic plants in a wild-type near-isogenic Pigm line (NIL-Pigm)21 (Extended Data Fig. 2a, b), and found that knockout of PICI1 (PICI1-KO) significantly decreased resistance to avirulent TH12 compared to NIL-Pigm and PICI1-OE (PICI1 overexpression) (Fig. 1c, Extended Data Fig. 2c). PICI1 also interacted with the coiled-coil domains of Pi9 and Pizt23—which confer similar blast resistance to PigmR21—but did not interact with Pish, which confers limited race resistance21 (Extended Data Fig. 2d–h). PICI1-KO in Pizt and Pi9 lines, but not in Pish, decreased resistance to avirulent strains (Extended Data Fig. 2i–q), suggesting that PICI1 is required for full ETI by directly interacting with specific NLRs. We next transiently expressed the blast effector AvrPizt in Pizt-containing rice cells20, which is expected to activate only ETI in the absence of PAMPs. We observed that cell death mediated by the Pizt–AvrPizt interaction was decreased in PICI1-KO (Extended Data Fig. 2r, s), which suggests that PICI1 has a bona fide role in NLR-mediated ETI. Together, these data show that PICI1 has a general role in ETI.
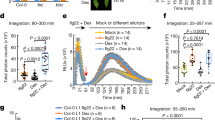
a, PigmR interacts with PICI1 in a Y2H assay. EV, empty vector; SD (−Leu−Trp) and SD (−Leu−Trp−His), SD medium lacking essential amino acids; 3AT, 3-aminotriazole. Another chitin-induced protein Os07g16130 served as a negative control. b, Co-IP of PigmR–7Myc–6His and PICI1–GFP. GFP served as a negative control. c, NIL-Pigm, PICI1-KO and PICI1-OE/NIL-Pigm lines, 5 days post-inoculation (dpi) with punch inoculation (avirulent isolate TH12). NIPB served as a susceptible control. d, Chitin-induced ROS burst. Data are mean ± s.d. (n = 9, biologically independent samples). e, PICI1 was induced after chitin treatment. Leaf sheaths were collected at different times of chitin incubation, with water as a control. Relative PICI1 abundance is indicated with PICI1-KO as a negative control. f, Basal blast resistance of wild-type and transgenic lines of PICI1-KO and PICI1-OE in NIL-Pigm, 5 dpi with the virulent strain TM21. Avirulent TH12 served as a resistant control. For c, f, data were analysed by two-tailed Student’s t-test. Fungal growth (mean ± s.d.; n = 3, biologically independent samples), lesion lengths (mean ± s.d.; n = number of biologically independent samples in the graph). Scale bars, 1 cm. Asterisks represent significant differences (*P < 0.05, **P < 0.01). Exact P values are provided in Supplementary Table 4. Similar results were obtained from three (a, c–f) or two (b) independent experiments.
PICI1 functions in PTI
Next, we investigated the potential role of PICI1 in PTI. We found that the chitin-induced production of reactive oxygen species (ROS) and expression of pathogen-related genes were suppressed in PICI1-KO but enhanced in PICI1-OE (Fig. 1d, Extended Data Fig. 3a). Moreover, chitin induced the expression of PICI1 mRNA and the accumulation of PICI1 protein (Fig. 1e, Extended Data Fig. 3b), similar to PTI triggered by TM21 (a TH12 mutant that is virulent to PigmR) (Extended Data Fig. 3c–e). Consistently, PICI1-KO enhanced susceptibility whereas PICI1-OE increased resistance to TM21 (Fig. 1f). Similar results were also observed with susceptible Nipponbare (NIPB) plants treated with chitin or M. oryzae (Extended Data Fig. 3f–i). The PRR OsCERK1 is essential for chitin signalling25. PICI1-OE/OsCERK1-KO decreased—whereas the PICI1-KO/OsCERK1-KO double mutant did not affect—susceptibility to TH12 compared to OsCERK1-KO (Extended Data Fig. 3j–n), suggesting that PICI1 acts downstream of OsCERK1-mediated PTI.
Furthermore, ROS production and pathogen-related gene expression induced by the bacterial elicitor flg22 were suppressed in PICI1-KO but enhanced in PICI1-OE (Extended Data Fig. 3o, p). PICI1 was also induced by flg22 and by infection with bacterial Xanthomonas oryzae pv. oryzae (Xoo) (Extended Data Fig. 3q, r). PICI1-KO enhanced susceptibility, but PICI1-OE increased resistance, to Xoo (Extended Data Fig. 3s). Thus, PICI1 has a critical role in rice PTI. Notably, PICI1-KO in NIPB (hereafter, PICI1-KO/NIPB) compromised PTI but not Pish-mediated ETI (Extended Data Figs. 2q, 3i), suggesting that PICI1-mediated PTI could be decoupled from ETI.
PICI1 is a plant deubiquitinase
We next determined that recombinant PICI1 and the PPPDE domain of PICI1 (PICI1-PPPDE) cleaved the ubiquitin-AMC substrate (Fig. 2a), towards K48- and K63-linked but not linear ubiquitin (Extended Data Fig. 4a), suggesting that PICI1 is a deubiquitinase. Cys115 in the catalytic dyad, a well-recognized residue in human isopeptide cleavage26, is critical for the activity of PICI1 (Fig. 2a, Extended Data Fig. 4b). Moreover, chitin-induced expression of pathogen-related genes was compromised in protoplasts expressing PICI1(C115S), suggesting its essential role for immune activation (Extended Data Fig. 4c).
a, Deubiquitinase activity was observed for His–PICI1 and His–PICI1-PPPDE, but not His–PICI1(C115S). His alone produced from empty vector (pCOLD) served as a negative control. AU, arbitrary units. Lower-case letters indicate statistical significance (P < 0.05). One-way ANOVA with Tukey’s test. Data are mean ± s.d. (n = 3, biologically independent experiments). Exact P values are provided in Supplementary Table 4. b, PICI1-KO/NIL-Pigm showed an increase in ubiquitin conjugates. Total protein extracts from seedlings before (top) or after (bottom) chitin treatment were analysed using anti-ubiquitin (Ub), anti-K48-linked (K48) or anti-K63-linked (K63) ubiquitin chain antibody. WT, NIL-Pigm. c, Co-IP of OsMETS1–GFP and PICI1–Flag. PICI2 served as a negative control. d, Degradation of OsMETS1–Flag in the presence of PICI1 in N. benthamiana. MG132 was infiltrated 18 h before sampling. The mCherry tag was expressed as an internal control. The RNA expression of mCherry and OsMETS1 was determined by semi-quantitative PCR. e, OsMETS1–Flag was K48-ubiquitin-linked. OsMETS1–Flag and HA–ubiquitin (HA-tagged wild-type, K48O or K63O ubiquitin) were co-expressed with PICI1–cLuc in N. benthamiana, and OsMETS1–Flag was immunoprecipitated and detected. f, Ubiquitinated OsMETS1–GFP in rice, as revealed by immunoprecipitation and detection. g, Deubiquitination of OsMETS1(K196R) by PICI1 but not PICI1(C115S) in rice protoplasts. h, Reduced levels of OsMETS in PICI1-KO compared with the wild type and PICI1-OE in NIPB and NIL-Pigm. i, Increased ubiquitination of endogenous OsMETS in PICI1-KO/NIL-Pigm compared with the wild type. Short and long indicate short exposure and long exposure, respectively. Protein abundance was quantified using ImageJ and is indicated under the lanes (d, h). Actin served as a loading control (b, e, f, h, i). Experiments were repeated three (a, b, d–g) or two (c, h, i) times with similar results.
We next observed an increased abundance of ubiquitin-conjugated proteins—particularly K48-linked ubiquitin, which is critical for protein turnover27—in PICI1-KO compared with the wild type, which was further enhanced after treatment with chitin (Fig. 2b). The decreased resistance was associated with increased ubiquitination in PICI1-KO, which was abolished after infection with TM21 (Extended Data Fig. 4d). Thus, PICI1-mediated deubiquitination is likely to be involved in PigmR-mediated immunity. PICI1 is therefore a deubiquitinase that was previously unknown, to our knowledge, and which does not affect plant growth (Extended Data Fig. 4e).
PICI1 stabilizes methionine synthase
To identify direct substrates of PICI1, we performed PICI1–GFP immunoprecipitation followed by mass spectroscopy (IP–MS) and then intersected potential PICI1-associated proteins with the chitin-induced ubiquitome28, which yielded 20 ubiquitin-modified candidates, including a Met synthase (LOC_Os12g42876) that is highly conserved in monocots and dicots (hereafter named OsMETS1) (Extended Data Fig. 4f–h, Supplementary Table 1). We confirmed the PICI1–OsMETS1 interaction (Fig. 2c, Extended Data Fig. 4i–k), which showed overlapping subcellular localization (Extended Data Fig. 4l–p). These data strongly suggest that OsMETS1 is a substrate of PICI1.
We measured the accumulation of OsMETS1–Flag in the presence of the 26S proteasome inhibitor MG132 or PICI1-cLuc (Fig. 2d, Extended Data Fig. 4q). Consistently, OsMETS1–Flag degraded faster in PICI1-KO/NIL-Pigm than in NIL-Pigm and PICI1-OE/NIL-Pigm (Extended Data Fig. 4r). These results suggest that PICI1 stabilizes OsMETS1 in planta. IP–MS analysis of ubiquitinated OsMETS1–GFP revealed four lysine sites that are probably deubiquitinated during infection (Extended Data Fig. 4s, t). Each of the K-R (lysine-arginine substitution) mutants showed enhanced stability compared to wild-type OsMETS1, with K28R being the most stable (Extended Data Fig. 4u, v). Consistently, global ubiquitome profiling revealed decreased ubiquitination of OsMETS1 K28 after PAMP treatment28. The chitin-induced expression of pathogen-related genes was substantially increased when individual K-R mutants were expressed (Extended Data Fig. 5a). Moreover, the quadruple mutation further increased OsMETS1 stability (Extended data Fig. 5b, c), implying that the other three lysine residues function additively with K28 to regulate the stability of OsMETS1.
PICI1 deubiquitinates OsMETS
We next co-expressed PICI1–cLuc and OsMETS1–Flag with wild-type, K48O or K63O ubiquitin (the latter indicating ubiquitin with only K48 or K63, respectively). PICI1–cLuc reduced the ubiquitinated OsMETS1–Flag generated with wild-type and K48O ubiquitin, but not K63O ubiquitin (Fig. 2e). Furthermore, OsMETS1–GFP was indeed modulated by K48 ubiquitination (Fig. 2f). Notably, PICI1, but not PICI1(C115S), delayed the degradation of OsMETS1–Flag (Extended Data Fig. 5d), and deubiquitinated OsMETS1(K196R) (Fig. 2g). We used OsMETS1(K196R), as wild-type OsMETS1 is barely detected in rice protoplasts (Extended Data Fig. 4v). An in vitro deubiquitination assay further verified the direct deubiquitination of OsMETS1 by PICI1 (Extended Data Fig. 5e). Collectively, our results suggest that PICI1 deubiquitinates OsMETS1 through cleaving K48-linked ubiquitin.
OsMETS1 shares high similarity with LOC_Os12g42884 (hereafter named OsMETS2) (Extended Data Fig. 5f, g). OsMETS2 also interacted with PICI1, and underwent 26S-proteasome-dependent degradation that was delayed by His–PICI1 (Extended Data Fig. 5h–l), suggesting their functional redundancy. We further confirmed the OsMETS–PICI1 interaction in rice (Extended Data Fig. 5m), and examined endogenous OsMETS (OsMETS1 and OsMETS2) stability using an anti-OsMETS antibody (Fig. 2h, Extended Data Fig. 5n–q). The levels of OsMETS protein, but not mRNA, were lower in PICI1-KO than in the wild type and PICI1-OE (Fig. 2h, Extended Data Fig. 5r), consistent with the accelerated degradation of OsMETS in PICI1-KO (Extended Data Fig. 5n). Moreover, OsMETS was indeed modulated by K48 ubiquitination, which was further enhanced in PICI1-KO (Fig. 2i, Extended Data Fig. 5s), confirming that PICI1 cleaves OsMETS K48-linked ubiquitin.
OsMETS functions in PTI and ETI
We first determined that treatment with chitin or flg22 resulted in the accumulation of OsMETS, as was observed for NIL-Pigm inoculated with TM21 or Xoo at the early stage of infection (Fig. 3a, Extended Data Fig. 6a–c). OsMETS also accumulated after TH12 but not TM21infection in NIL-Pigm (Fig. 3b, Extended Data Fig. 6d), implying its ETI function. Of note, TM21-induced accumulation of OsMETS was greatly dampened in PICI1-KO, but prolonged in PICI1-OE (Extended Data Fig. 6e). Increased deubiquitination of OsMETS1 was also observed in PTI and ETI (Extended Data Fig. 6f, g). The data suggest that PTI and ETI induce the accumulation of OsMETS through PICI1. Furthermore, OsMETS1-KO and OsMETS2-KO increased, whereas OsMETS1-OE and OsMETS2-OE decreased, susceptibility to TM21, compared with NIL-Pigm (Fig. 3c, Extended Data Figs. 5p, q, 6h–l), suggesting that OsMETS functions in PTI. Slight susceptibility to TH12 was observed in OsMETS1-KO and OsMETS2-KO (Extended Data Fig. 6m, n), but this was greatly enhanced when RNA interference (RNAi) was used (OsMETS1- and OsMETS2-RNAi) (Fig. 3d, Extended Data Fig. 6o, p), suggesting the redundant roles of OsMETS1 and OsMETS2 in PigmR-mediated ETI. OsMETS-RNAi lines exhibited a dwarf phenotype (Extended Data Fig. 6q, r), with a reduced Met content (Extended Data Fig. 6s). We could not generate OsMETS1 and OsMETS2 double-knockout plants, probably owing to the essential role of Met in growth.
a, OsMETS accumulation was induced in the early stages of TM21 infection. Water served as a control. b, Protein accumulation of OsMETS in NIL-Pigm after inoculation with TH12. Relative OsMETS abundance was indicated with actin as a loading control (a, b). hpi, hours post-infection. c, Basal blast resistance of NIL-Pigm, OsMETS1-KO/NIL-Pigm and OsMETS1 overexpression (OsMETS1-GFP/NIL-Pigm) lines, 5 dpi using spraying inoculation with TM21. d, PigmR-mediated blast resistance, 5 dpi using punch inoculation with TH12. e, Increased basal blast resistance after Met application. NIL-Pigm seedlings were dipped into different concentrations of Met or water from the roots for 48 h, and inoculated at 7 dpi with TM21. Lower-case letters indicate statistical significance (P < 0.05). One-way ANOVA with Tukey’s test. Data are mean ± s.d. (lesion areas, n = 18, biologically independent samples; fungal growth, n = 3, biologically independent samples). f, Ethylene content in NIL-Pigm after treatment with 25 mM Met or water for 96 h. nl gfw−1 h−1 indicates the amount of ethylene per gram fresh weight seedling per hour. Two-tailed Student’s t-test (mean ± s.d.; n = 3, biologically independent samples). g, Effects of AVG and ACC on basal blast resistance in NIL-Pigm. Seedlings were pre-treated with 10 μM AVG, or 20 μM ACC or water for 1 day, then 5 dpi with spraying inoculation (TM21). h, Effect of AVG on PigmR-mediated resistance in NIL-Pigm, punch-inoculated with TH12 for 5 dpi. For c, d, g, h, data were analysed by two-tailed Student’s t-test. Fungal growth (mean ± s.d.; n = 3, biologically independent samples), lesion lengths or areas (mean ± s.d.; n = number of biologically independent samples in the graphs). Scale bars, 1 cm (c–e, g, h). Asterisks indicate significant differences (*P < 0.05, **P < 0.01) (c, d, f–h). Exact P values are provided in Supplementary Table 4 (c–h). Experiments were repeated three times with similar results.
We next tested Met-mediated immunity, and observed dose-dependent Met-induced resistance with the induction of pathogen-related genes (Fig. 3e, Extended Data Fig. 6t–v). In addition, Met rescued OsMETS1-KO susceptibility, and even enhanced resistance, similar to OsMETS overexpression (Extended Data Fig. 6w). Likewise, PICI1-KO decreased Met content, and Met supplementation restored its resistance (Extended Data Fig. 6x, y). In addition, fungal growth was inhibited by 10 and 25 mM but not by 5 mM Met (Extended Data Fig. 7a)—which are much higher concentrations than that seen in rice—suggesting Met-mediated immune activation. Together, these results show that OsMETS-mediated biosynthesis of Met has an important role in blast resistance.
OsMETS functions through ethylene
Met is the precursor of ethylene functioning in plant immunity9. Indeed, treatment with Met increased the production of ethylene (Fig. 3f). Of note, PTI-induced ethylene production was significantly decreased in OsMETS1-KO and PICI1-KO, whereas it was increased in OsMETS1-OE and PICI1-OE (Extended Data Fig. 7b, c). Treatment with ethylene led to the downregulation of OsMETS1 and OsMETS2 (Extended Data Fig. 7d–f), suggesting negative feedback regulation. Pre-treatment with ACC (1-aminocyclopropane-1-carboxylic acid, an ethylene precursor) decreased, whereas AVG (aminoethoxyvinylglycine, an ethylene biosynthesis inhibitor) increased, blast susceptibility (Fig. 3g, Extended Data Fig. 7g, h). In addition, ACC rescued OsMETS-RNAi and PICI1-KO resistance (Extended Data Fig. 7i, j), indicating that ethylene is involved in PICI1–OsMETS-mediated PTI. Notably, pre-treatment with AVG resulted in compromised resistance of NIL-Pigm to TH12, suggesting that ethylene is also critical to PigmR-mediated ETI (Fig. 3h). OsACS2, a key ethylene biosynthesis enzyme, was activated during PTI and ETI, with increased ethylene production (Extended Data Fig. 7k–m). Consistently, OsACS2-KO decreased basal blast resistance (Extended Data Fig. 7o). However, PigmR-mediated ETI was not affected in OsACS2-KO (Extended Data Fig. 7p), suggesting that ETI disturbance may need more deprivation of ethylene owing to the functional redundancy of OsACS proteins29. Therefore, ethylene is actively involved in blast resistance. We further observed that AVG treatment could partially dampen the Met-induced resistance (Extended Data Fig. 7q), suggesting that ethylene biosynthesis is a major function of Met in immune activation.
PigmR protects PICI1 from effectors
To elucidate the mechanism of PICI1 function in PigmR-mediated resistance, we analysed the levels of PICI1 during blast infection. PICI1 protein, but not mRNA, increased in NIL-Pigm inoculated with avirulent TH12, whereas PICI1 gradually declined in NIPB and NIL-Pigm inoculated with virulent strains (Extended Data Fig. 7r, s). The results strongly suggest that fungal effectors are involved in PICI1 turnover. We therefore screened 10 known blast effectors for direct interactions with PICI1, and found that PICI1 bound to 5 of these effectors—AvrPizt20, AvrPii19, AvrPia19, AvrPWL217 and AvrPi918—with a higher affinity than to AvPikC16 and AvrPikD16, whereas AvrBAS413, AvrSlp114, AvrPita15 or MBP tag were not pulled down with PICI1 (Extended Data Fig. 8a). Furthermore, an SLC assay suggested that PICI1 interacted with AvrPi9, AvrPWL2 and AvrPii with a similar or higher binding affinity to the known AvrPii–OsNADP–ME230, but with a lower affinity to that of AvrPikD-HMA31,32 in planta (Extended Data Fig. 8b–d). Moreover, these effectors interacted specifically with PICI1 but not PICI2 (Extended Data Fig. 8e–g), suggesting that PICI1 is a unique immune hub that is co-opted by various effectors. AvrPizt and AvrPia may not interact with PICI1 in planta, probably owing to altered subcellular localization, conformation or modification.
We then generated Avr-expressing transgenic plants and found that AvrPi9—but not AvrPizt, AvrPWL2 or AvrPii—reduced the endogenous levels of PICI1 (Extended Data Fig. 8h). In addition, AvrPi9, but not AvrPizt, co-localized with PICI1 (Fig. 4a, Extended Data Fig. 8i, j). Co-immunoprecipitation and bimolecular fluorescence complementation (BiFC) further specified the AvrPi9–PICI1 interaction (Fig. 4b, Extended Data Fig. 8k). Notably, ectopic expression of AvrPi9 led to enhanced susceptibility with decreased expression of pathogen-related genes (Extended Data Fig. 8l–n), indicating that AvrPi9 suppresses rice PTI. We also found that AvrPi9–mCherry—but not mCherry, AvrPizt–mCherry or AvrPWL2–mCherry—promoted the degradation of PICI1–GFP (Fig. 4c, Extended Data Fig. 8o). Consistently, the endogenous PICI1 exhibited faster degradation in AvrPi9-containing versus AvrPWL2-containing plants (Fig. 4d), which was not affected by MG132 (Fig. 4d, Extended Data Fig. 8p), suggesting non-26S proteasome degradation.
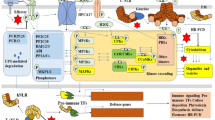
a, Co-localization of PICI1–YFP and AvrPi9–mCherry in rice protoplasts. Scale bar, 5 µm. b, PICI1–GFP but not PICI2–GFP interacts with AvrPi9–mCherry in co-IP. c, Cell-free assay shows AvrPi9-mediated degradation of PICI1–GFP. PICI1–GFP/NIL-Pigm extracts were incubated with AvrPi9–mCherry expressed in N. benthamiana. mCherry and AvrPWL2–mCherry served as negative controls. d, AvrPi9 promotes endogenous PICI1 degradation independently of the 26S proteasome in a rice cell-free assay. AvrPWL2-Flag/NIL-Pigm served as a control. e, Reduced levels of PICI1 in AvrPi9-Flag/NIPB and AvrPi9-Flag/NIL-Pigm compared with the wild type. f, PICI1–GFP degradation is delayed in infected NIL-Pigm. PICI1-GFP/NIL-Pigm extracts inoculated with TH12 or water (36 hpi) were incubated with AvrPi9–mCherry expressed in N. benthamiana. g, Percentages of ProPICI1jap and ProPICI1ind haplotypes in subspecies japonica and indica. h, Nucleotide diversity (π) of PICI1 and flanking regions in indica, japonica and wild rice (O. rufipogon). i, Basal blast resistance of Huajingxian74 and CSSL plants to the virulent isolate YN2, 5 dpi. Scale bar, 1 cm. Two-tailed Student’s t-test. Fungal growth (mean ± s.d.; n = 3, biologically independent samples), lesion lengths (mean ± s.d.; n = numbers of biologically independent samples in the graph). Asterisks indicate significant differences (**P < 0.01). Exact P values are provided in Supplementary Table 4. j, Proposed model for the NLRs–PICI1–OsMETS–ethylene (ET) immune cascade. PICI1 is targeted by both NLRs and Avr effectors, and induced by PAMPs to initiate PTI. Activated NLRs protect PICI1 from effector-mediated degradation to reboot the Met–ethylene biosynthesis, resulting in robust defence through integrating PTI and ETI. Protein abundance was quantified using ImageJ and is indicated under the lanes (c–f). Ponceau S staining (c, d, f) or actin (e) served as loading controls. Experiments were repeated twice (b, e) or three times (a, c, d, f, i) with similar results.
Of note, the AvrPi9-mediated degradation of PICI1–GFP was delayed in PICI1–GFP/NIL-Pigm compared to PICI1–GFP/NIPB (Extended Data Fig. 8p), suggesting that PigmR protects PICI1 from degradation by AvrPi9. To test this, we first determined that PICI1 decreased to a greater extent after overexpression of AvrPi9 in NIPB versus NIL-Pigm (Fig. 4e). Infection with TH12 (without AvrPi9) or TH12AvrPi9 (containing AvrPi9) increased PICI1 in NIL-Pigm but decreased PICI1 in NIPB at a faster rate with TH12AvrPi9 versus TH12 inoculation (Extended Data Fig. 8q–s), suggesting that PICI1 is hijacked by AvrPi9 and other as-yet-unidentified effectors, but protected by PigmR. AvrPi9 is the Avr effector of Pi9, and PigmR is neither activated nor induced in an AvrPi9-dependent manner (Extended Data Fig. 9a–c), suggesting that immune-active PigmR interferes with the AvrPi9–PICI1 interaction to inhibit the degradation of PICI1. Indeed, the AvrPi9-triggered degradation of PICI1–GFP was greatly delayed by avirulent infection (Fig. 4f), accompanied by enhanced PigmR–PICI1 interaction (Extended Data Fig. 9d). AvrPi9–PICI1 interaction was reduced in the presence of PigmR or the coiled-coil domain of PigmR (PigmR-CC) (Extended Data Fig. 9e–h). PigmR-CC also protects PICI1 from AvrPWL2 and AvrPii binding (Extended Data Fig. 9i–l), suggesting that this competitive mechanism might be widely used by PigmR.
We further found that PICI1 and OsMETS accumulated in Pizt and Pi9 plants during avirulent strain infection (Extended Data Fig. 9m, n), and that this PICI1 induction was absent in Pizt-KO/ZH11 (Extended Data Fig. 9o). Pizt-CC can also protect PICI1 from AvrPi9, AvrPWL2 and AvrPii binding in planta (Extended Data Fig. 9p). These results suggest that PICI1 is likely to be protected in general by NLRs.
PICI1 alleles confer immune divergence
Notably, using NIPB as the reference in the 3,000 Rice Genome Project33, 27 single-nucleotide polymorphisms (SNPs) were identified in its promoter, nearly all of which are within indica germplasm (hereafter named ProPICI1ind), together with some aus, admix and aro accessions (Extended Data Fig. 10a, Supplementary Table 2); by contrast, the majority of japonica varieties are the ProPICI1NIPB haplotype (hereafter named ProPICI1jap) (Extended Data Fig. 10b, Supplementary Table 2). Among the 27 SNPs, 13 SNPs are well correlated with indica–japonica differentiation, which was verified in 96 global rice collections34 (Fig. 4g, Supplementary Table 2), implying that PICI1 is differentially expressed between indica and japonica. The decreased nucleotide diversity (π) of the PICI1 promoter in the indica and japonica subspecies compared with that in wild rice (Oryza rufipogon) (Fig. 4h) suggests a selection during domestication. Moreover, fixation index (FST) analysis revealed a greater genetic distance between indica and japonica (Extended Data Fig. 10c), suggesting divergent indica–japonica domestication.
We found that ProPICI1jap showed higher transcriptional activity than ProPICI1ind after treatment with chitin (Extended Data Fig. 10d), consistent with higher fungal induction of PICI1 in japonica than in indica (Extended Data Fig. 10e). Furthermore, a chromosome segment substitution line (CSSL) containing the PICI1jap allele in Huajingxian74 (ProPICI1ind) exhibited an increase in PICI1 induction compared with Huajingxian74 (Extended Data Fig. 10f, g, Supplementary Table 2), indicating that the SNPs are likely to regulate PICI1 expression. In addition, we detected higher basal defence in the CSSL (Fig. 4i, Extended Data Fig. 10f, h). These data are consistent with and may explain previous findings that japonica has higher levels of Met35 and higher field and basal blast resistance than indica36. Collectively, our results suggest that PICI1 contributes to subspecies divergence in basal defence, thus providing a potential breeding target for broad-spectrum blast resistance together with NLRs in rice.
Discussion
We have thus revealed here an NLR–effector arms race that controls the biosynthesis of primary metabolites and immune activation in rice. Rice NLRs including PigmR and blast effectors including AvrPi9 both target PICI1, which stabilizes OsMETS to promote Met–ethylene biosynthesis (Fig. 4j). Therefore, we propose that NLRs counteract effectors to ensure efficient immunity with a ‘competitive’ mode. It would be worth investigating whether the same mechanism is adopted in other pathosystems. The PICI1–OsMETS–ethylene cascade is used in both PTI and ETI, and this ETI–PTI integration contributes to broad-spectrum blast resistance mediated by PigmR and other similar NLRs. However, we cannot exclude roles for other Met derivatives or Met-related processes in blast resistance.
PICI1 is probably hijacked to suppress host immunity by multiple pathogen effectors, similarly to Arabidopsis RIN437,38. Indeed, we observed specific acetylation of K160 in PICI1 after infection (Extended Data Fig. 10i, j, Supplementary Table 1), which promoted the degradation of PICI1—as a PICI1 acetyl-lysine mimic mutant was unstable—and thus weakened interactions with OsMETS1 and PigmR (Extended Data Fig. 10k). How PICI1 is subjected to differential modifications by diverse effectors needs to be further elucidated. However, we suggest that the NLR–PICI1 surveillance mechanism could provide a strategy for breeding disease-resistant crops.
Methods
Plant materials and growth conditions
The japonica variety Nipponbare (NIPB) and near-isogenic Pigm line (NIL-Pigm), OsCERK1-KO/NIPB, Pizt-KO/ZH11 and PigmR-7Myc-6His/NIPB used in this study were collected or generated in our previous study21,22,39. Huajingxian74 and the CSSL were provided by G. Zhang. The following transgenic lines were developed in this study: PICI1-KO/NIPB, PICI1-KO/NIL-Pigm, PICI1-OE(pUBI::PICI1-GFP)/NIPB, PICI1-Flag(pUBI::PICI1-Flag)/NIPB, PICI2-Flag(pUBI::PICI2-Flag)/NIPB, PICI1-OE(pUBI::PICI1-GFP)/NIL-Pigm, OsMETS1-KO/NIL-Pigm, OsMETS2-KO/NIL-Pigm, OsMETS1-OE(pUBI::OsMETS1-GFP)/NIL-Pigm, OsMETS2-OE(pUBI::OsMETS2-GFP)/NIL-Pigm, OsMETS-RNAi/NIL-Pigm, OsACS2-KO/NIL-Pigm, OsCERK1-KO/PICI1-KO/NIPB, OsCERK1-KO/PICI1-OE/NIPB, PICI1-KO/ZH11, PICI1-KO/Ky-Pi9, AvrPi9-Flag/NIL-Pigm, AvrPi9-Flag/NIPB, AvrPWL2-Flag/NIL-Pigm, AvrPii-Flag/NIL-Pigm and AvrPizt-Flag/NIL-Pigm. Rice plants were grown in the experimental fields in Shanghai for the summer season, or Lingshui, Hainan for the winter season under natural field conditions for plant inoculation (punch injection) and seed production. For seedling spraying inoculation, two-week-old seedlings were grown at the greenhouse at 26 °C, 14-h day–10-h night. Nicotiana benthamiana, used for transient expression, protein–protein interaction and degradation analysis, was grown at the greenhouse at 22 °C under long-day conditions (16-h day–8-h night).
Development of transgenic rice plants
For CRISPR–Cas9 constructs, the designed target sequences of PICI1, OsMETS1, OsMETS2 and OsACS2 were synthesized and CRISPR–Cas9 constructs were generated following the protocol reported40. To generate constructs for PICI1, PICI2, OsMETS1/2, AvrPi9, AvrPWL2, AvrPii and AvrPizt fusion overexpression, the corresponding coding sequences were inserted into PUN1301-pUBI-GFP or PUN1301-pUBI-Flag vectors. To generate the OsMETS RNAi construct, a conserved 540-bp coding sequence fragment of OsMETS was selected and inserted as inverted repeats into the RNAi vector PTCK303. The CRISPR–Cas9, RNAi and overexpression plasmids were introduced into NIL-Pigm, ZH11, Ky-Pi9 or NIPB via Agrobacterium (EHA105)-mediated transformation to generate more than 20 independent transgenic lines for each construct, further selected by PCR-based sequencing or western blot. All primer sequences used for cloning can be found in Supplementary Table 3.
Y2H screen and assay
The Y2H screen was conducted to identify the PigmR-interacting candidates as previously described41. In brief, the truncated N-terminal part (CC domain) of PigmR was fused to the GAL4 DNA-binding domain in pDEST32 as the bait to screen the rice cDNA library in pDEST22 according to the manufacturer’s instructions (Invitrogen), with yeast strain AH109. Positive clones were screened and sequenced.
For the Y2H assay, the target coding sequences were cloned into pDEST22(AD) or pDEST32(BD) vectors. Different construct combinations were co-transformed into yeast strain AH109 and clones were grown on selective medium (lacking Trp, Leu and His) containing proper 3-aminotriazole to test the protein–protein interaction. Images were taken 3 days after incubation at 30 °C.
Rice blast inoculation
Rice blast inoculation was performed as usual21,41. In brief, M. oryzae spores were collected in sterile water containing 0.05% Tween-20 and the spore concentration was adjusted to approximately 1 × 105 spores per ml for spraying, punch injection or punch inoculation. For spray inoculation, two-week-old seedlings were sprayed with spore suspensions in a dew growth chamber. Leaves of tillering plants grown in the field were punch-injection-inoculated or punch-inoculated as previously described41. At 5–7 dpi, lesions were evaluated by calculating lesion areas and lengths using the software ImageJ or a ruler. Relative fungal growth was measured by DNA-based quantitative PCR (qPCR) using the threshold cycle value (CT) of M. oryzae 28S rDNA against the CT of rice genomic ACTIN1 DNA.
SLC, BiFC and subcellular localization
The SLC and BiFC assays were described previously41. For SLC assays, the tested coding sequences were cloned into pCAMBIA-35S-nLuc or pCAMBIA-35S-cLuc, and the resulting plasmids were transformed into Agrobacterium (strain GV3101), cultured overnight in LB medium, collected and suspended in infiltration buffer (10 mM MgCl2, 10 mM methylester sulfonate, 150 μM acetosyringone, pH 5.6), and incubated for 2–3 h at 30 °C before infiltration. The suspensions were then infiltrated into 5-week-old N. benthamiana leaves in different combinations. After two days of growth, luciferase substrate (Promega) was sprayed onto the surface of the leaves and the luciferase signals were imaged using a Tanon-5200 imaging system (Tannon). For luciferase activity measurement, leaves were taken and ground into fine powder in liquid nitrogen, homogenized in the lysis buffer following the manufacturer’s instructions (Promega). The extraction solution was then incubated with luciferin substrate in a 96-well plate for 10–15 min, and luminescence was captured with the Varioskan Flash (Thermo Fisher Scientific) plate reader. For the BiFC assay in rice protoplasts, the PICI1, PigmR-CC, Pish-CC, PICI2 and AvrPi9 coding sequences were inserted into PSAT1-nVenus or PSAT1-cCFP, respectively. Different construct combinations were co-expressed in rice protoplasts prepared from seedling sheaths. Fluorescence images were recorded on a confocal microscope (Olympus Fluoview FV1000) at 16 h after transformation.
For subcellular localization of proteins in rice protoplasts, the coding sequences of PICI1, OsMETS1, PigmR, AvrPi9 and AvrPizt were inserted into PA7-35S-YFP or pAN583-35S-mCherry, respectively. The fusion constructs were transformed alone or co-transformed into rice protoplasts. The NLS sequence was fused in-frame with RFP into PA7-35S-RFP as a nuclear marker. Fluorescence was detected using a confocal microscope (Olympus Fluoview FV1000) after incubation in the dark for 16–20 h at 26 °C. To visualize the subcellular localization of PICI1–GFP or OsMETS1–GFP in transgenic rice root, the root tips were stained with 10 μM DAPI for 5 min before capturing the images using a confocal microscope (Olympus Fluoview FV1000).
ROS measurement
For measurement of ROS bursts in rice cells after chitin treatment, the luminol chemiluminescence assay was conducted as described previously with some modifications42. In brief, leaf sheaths from 10-day-old rice plants cultivated in 1/2 MS medium were cut (approximately 3-mm strips) and preincubated overnight in sterile distilled water to recover from wounding stress. The materials were then treated with 1 × 10−6 M chitin [octa-N-acetylchitooctaose (GlcNAc)8], 1 × 10−6 M flg22 or water as a control in reaction buffer containing 20 μM luminol (Wako) and 10 μg ml−1 horseradish peroxidase (Sigma). Immediately after the treatment, luminescence was monitored continuously at 10-s intervals for 25 min with a Glomax 20/20 luminometer (Promega) or 1-min intervals for 40 min with a Varioskan Flash multireader (Thermo Fisher Scientific).
Cell-free degradation assay
For the cell-free protein degradation assay, total proteins were extracted from rice seedlings or N. benthamiana leaves with the extraction buffer (50 mM Tris-MES, pH 8.0, 0.5 M sucrose, 1 mM MgCl2, 10 mM EDTA, 5 mM dithiothreitol, 1 mM phenylmethylsulfonyl fluoride and 1× protease inhibitor cocktail). Samples were incubated at 25 °C with or without 50 μM MG132 and taken at the indicated intervals for the western blotting. All protein immunodetection experiments were performed independently three times with similar results.
Protein extraction, immunoblotting and co-immunoprecipitation
The PICI1 and OsMETS polyclonal antibodies were custom-developed by ABclonal Technology. The PICI1 (100–246 aa) and OsMETS (410–766 aa) proteins were expressed, purified and used as antigens to raise polyclonal antibodies in rabbit.
To extract total proteins from plants, 0.2 g of fresh leaves or rice seedling sheaths were ground into fine powder in liquid nitrogen, homogenized in the protein extraction buffer (150 mM Tris-HCl, pH 7.5, 6 M urea, 2% SDS, and 5% β-mercaptoethanol), boiled for 5 min and then centrifuged at 16,000g for 10 min at 4 °C to remove debris. Supernatants were collected with SDS loading buffer for protein gel blot. For total protein extraction from yeast, a post-alkaline extraction method was performed. In brief, yeast cells are collected, resuspended in 100 μl 0.2 M NaOH, incubated for 5 min at room temperature, then pelleted and resuspended in 50 μl SDS loading buffer, followed by boiling for 5 min, and the resulting supernatants were separated using SDS–PAGE.
The co-immunoprecipitation (co-IP) procedure was described previously41. Protein extracts from N. benthamiana or rice leaves were prepared in the IP extraction buffer (50 mM Tris-HCl, pH 7.5, 150 mM NaCl, 1 mM EDTA, 10% glycerol, 1% Triton X-100, 1 mM PMSF and 1× protease inhibitor cocktail). Different combinations of supernatants were incubated with anti-Myc beads, anti-Flag beads or anti-GFP beads for 2 h at 4 °C and then washed four times with extraction buffer. The bound proteins were eluted from the affinity beads by boiling for 5 min in SDS loading buffer and analysed by western blot.
Quantification of protein accumulation was performed using ImageJ software. Antibodies against the following proteins were used: Myc (1:2,000), GFP (1:2,000), PICI1 (1:1,000), His (1:1,000), MBP (1:1,000), ubiquitin (1:1,000), K48-linkage ubiquitin (1:1,000), K63-linkage ubiquitin (1:1,000), Ac-K-103 (1:1,000), Flag (1:2,000), GAL4-AD (1:1,000), GAL4-BD (1:1,000), mCherry (1:1,000), nLuc (1:1,000), S-tag (1:1,000), HA (1:1,000), OsMETS (1:1,000), actin (1:2,000), goat anti-rabbit IgG secondary antibody (1:10,000) and goat anti-mouse IgG secondary antibody (1:10,000).
Recombinant protein preparation and pull-down assay
To construct His-fusion plasmids for the production of recombinant proteins, the coding sequences were PCR-amplified and inserted into pCOLD (His–PICI1, His–PICI1-PPPDE, His–PICI1(C115S), His–OsMETS1, His–OsMETS1-2, His–OsMETS1(K28R), His–OsMETS1(K196R), His–OsMETS1(K458R) and His–OsMETS1(K740R)). All protein site mutations were generated by mutagenesis kit following the manufacturer’s instructions. The recombinant proteins were produced in the Escherichia coli strain Rosetta (DE3) and induced with 0.5 mM isopropyl β-d-thiogalactoside (IPTG) for 20 h at 18 °C. Bacterial cells were collected and affinity-purified with Ni-NTA agarose (QIAGEN) following the manufacturer’s instructions. To identify the recombinant protein purity and concentration, Coomassie brilliant blue R-250 staining with quantified BSA as a standard was performed. For pull-down assays, bait proteins were captured with corresponding resin (Amylose Resin for MBP-tagged Avr proteins or glutathione sepharose 4B for GST-tagged PICI1), washed three times, then incubated with prey proteins at 4 °C for 2 h in binding buffer (50 mM Tris-HCl, pH 7.5, 100 mM NaCl, 10% (v/v) glycerol). The beads were then washed twice or three times with the washing buffer (50 mM Tris-HCl, pH 7.5, 100 mM NaCl, 10% (v/v) glycerol, 0.1% (v/v) Triton X-100) to remove non-specifically bound proteins, and the precipitates were released by boiling in SDS sample buffer at 100 °C for 5 min and detected by immunoblotting using corresponding antibodies. All PCR primers are listed in Supplementary Table 3.
Deubiquitination assay
Ubiquitin 7-amido-4-methylcoumarin (ubiquitin-AMC, Enzo) was used as a substrate to monitor the enzymatic activity of ubiquitin C-terminal hydrolase (UCH), which releases the fluorogenic AMC component by cleaving the bond between the C-terminus of ubiquitin and AMC. In brief, 1-μg aliquots of recombinant His–PICI1, His–PICI1-PPPDE or His–PICI1(C115S) were incubated with 250 nM ubiquitin-AMC in reaction buffer (50 mM HEPES, pH 7.8, 0.5 mM EDTA and 1 mM dithiothreitol) in a total volume of 100 μl for 1 h. The fluorescence of an AMC moiety was monitored by spectrofluorometry using a Varioskan Flash (Thermo Fisher Scientific) at 380 nm excitation/460 nm emission. For K48-, K63- or linear linked tetra-ubiquitin cleavage assay, recombinant His–PICI1 or His–PICI1-PPPDE was added to 20 μl reaction buffer containing 2.5 μg of K48-, K63- or linear tetra-ubiquitin and held for 1 h at 37 °C. The reactions were stopped by adding SDS loading buffer, followed by separation on SDS–PAGE. Recombinant His alone (pCOLD) was used as a negative control.
RNA analysis
Total RNAs were extracted using TRIzol reagent (Invitrogen) from rice tissues or N. benthamiana leaves. For quantitative PCR with reverse transcription (qRT–PCR) or semi-quantitative PCR, total RNA (around 1 μg) was converted into cDNA using ReverTra Ace qPCR RT Master Mix with the gDNA Remover kit according to the manufacturer’s instructions (TOYOBO). The qRT–PCR reaction was performed using a CFX96 Real-time PCR Instrument (Bio-Rad) with SYBR Premix Ex Taq (Takara) following the manufacturer’s instructions. Rice ACTIN1 served as an internal control to normalize expression levels and the 2-△△CT method was used to calculate the relative expression levels with three biological repeats. All of the qPCR primers are listed in Supplementary Table 3.
Promoter activity analysis
A dual-Luc assay was conducted to measure the promoter activityin rice protoplasts, with RENILLA (REN) as an internal control. The approximately 2-kb promoters of PICI1ind and PICI1jap were isolated and cloned into the vector pGreenII 0800-Luc to generate ProPICI1ind::Luc and ProPICI1jap::Luc constructs. The luciferase activity was calculated by normalizing the REN expression after treatment with 1 × 10−6 M octa-N-acetylchitooctaose (GlcNAc)8.
Measurement of Met and ethylene
For Met quantification, approximately 0.1 g of leaves from two-week or one-month-old plants were collected, ground in liquid nitrogen, and resuspended in 1 ml of precooled methanol/acetonitrile/water (2:2:1, v/v/v) buffer, which were then mixed and homogenized by ultrasonication for 30 min. The extract was incubated at −20 °C for 1 h and centrifuged at 4 °C (12,000 rpm for 30 min). The filtered supernatant was collected and analysed on the QTRAP 6500 (AB SCIEX). A Met standard curve is generated to quantify the Met content in each sample.
Ethylene content was determined following a previously reported protocol11. In brief, leaves of the same position were detached and cut into 3-cm pieces, and then 3 pieces were put into a 10-ml glass vial with 1 ml distilled water, sealed with a gas-proof septum and left in a growth chamber at 28 °C for 48 h. The gas sample was withdrawn from the air space of each bottle using a gas-tight syringe (Hamilton), and injected into a gas chromatograph (Agilent 6890N) equipped with an activated alumina column and flame ionization detectors for ethylene determination. An ethylene standard curve is generated to quantify ethylene emission in each sample.
LC–MS/MS analysis
To identify PICI1-interacting proteins, total proteins were extracted from leaves of two-week-old seedlings (PICI1-GFP/NIL-Pigm) after incubation with strain TH12 for 36 h with the IP extraction buffer mentioned above. Non-transgenic NIL-Pigm served as a negative control. The PICI1–GFP protein was enriched by immunoprecipitation using anti-GFP antibody, eluted with 0.1 M glycine (pH = 2.5) and the supernatant was used for the mass spectrometric (MS) analysis.
To identify OsMETS1 ubiquitination sites, total proteins were extracted from leaves of two-week-old seedlings (OsMETS1-GFP/NIL-Pigm) after they were inoculated with TH12 or water as a control for 36 h with IP extraction buffer. The OsMETS1–GFP protein was enriched by immunoprecipitation using anti-GFP antibody, eluted by SDS loading buffer, separated by SDS–PAGE and after staining with Coomassie brilliant blue R-250, the OsMETS1–GFP protein bands were cut out for the mass spectrometric analysis. To identify PICI1 protein modification sites, total proteins were extracted from leaves of two-week-old seedlings (PICI1-GFP/NIL-Pigm) after they were inoculated with TM21 or water as a control for 36 h with IP extraction buffer. The PICI1–GFP protein was enriched and analysed as mentioned above.
For tandem mass tag (TMT)-based quantitative proteomic analysis, rice leaf sheaths were treated with 1 × 10−6 M octa-N-acetylchitooctaose (GlcNAc)8 or water as a control for 10 min. Each group was processed with two independent biological replicates. The concentration of extracted proteins was determined using a bicinchoninic acid (BCA) kit and the proper amount of protein was digested with trypsin using the filter-aided proteome preparation method and then the resulting peptide was processed according to the manufacturer’s protocol (Thermo Fisher Scientific). The labelled peptides were fractionated using high pH reverse-phase high pressure liquid chromatography (HPLC), followed by tandem mass spectrometry (MS/MS) in Q Exactive (Thermo Fisher Scientific).
The LC–MS/MS analysis was processed by Applied Protein Technology (Shanghai) as previously described28. The entire Rice Annotation Project (RAP) database (http://rapdb.dna.affrc.go.jp/) was searched using Mascot (v.2.3.02, Matrix Science).
Domestication analysis
Nucleotide diversity of PICI1 and flanking regions was calculated using a sliding window approach. The distribution of nucleotide diversity is plotted in sliding windows of 1 kb with a 100-bp step size. Genetic distance (FST) between the japonica accessions, indica accessions and rufipogon accessions for the PICI1 and flanking regions are calculated using a sliding window approach with a 2.5-kb window size and 50-bp step size. The genomic sequences of 27 japonica accessions, 19 indica accessions and 13 rufipogon accessions were acquired from the RicePanGenome (http://db.ncgr.ac.cn/RicePanGenome/).
Accession numbers
The sequence data for PICI1, OsMETS1 and OsMETS2 can be found in the Rice Annotation Project (RAP) database (http://rapdb.dna.affrc.go.jp/) and have been deposited in GenBank: MT920667, MT920668 and MT920669.
Quantification and statistical analysis
No statistical methods were used to predetermine sample size. The experiments were not randomized and the investigators were not blinded to allocation during experiments and outcome assessment. Quantification analyses on lesion areas and protein abundance were conducted by ImageJ software. All values are presented with mean ± s.d. as indicated. Data points are plotted onto the graphs, and the number of samples is indicated in the corresponding figure legends. All statistical analyses were performed by one-way ANOVA followed by Tukey’s test with GraphPad software or by two-sided Student’s t-test with Microsoft Excel software. Detailed information about statistical analysis values for all experiments is provided in Supplementary Table 4.
Reporting summary
Further information on research design is available in the Nature Research Reporting Summary linked to this paper.
Data availability
All data are available within this Article and its Supplementary Information. Original gel blots are shown in Supplementary Fig. 1. Original data points in graphs are shown in the Source Data files. Statistical analyses of this study are provided in Supplementary Table 4. The sequences of PICI1, OsMETS1 and OsMETS2 have been deposited and made publicly available in GenBank with accession codes MT920667, MT920668 and MT920669, respectively. Protein structure models of Protein Data Bank (PDB) ID 2WP7 and PDB ID 1U1U were obtained from the PDB. Source data are provided with this paper.
References
Jones, J. D. & Dangl, J. L. The plant immune system. Nature 444, 323–329 (2006).
Couto, D. & Zipfel, C. Regulation of pattern recognition receptor signalling in plants. Nat. Rev. Immunol. 16, 537–552 (2016).
Xin, X.-F. Kvitko, B. & He, S. Y. Pseudomonas syringae: what it takes to be a pathogen. Nat. Rev. Microbiol. 16, 316–328 (2018).
Kourelis, J. & van der Hoorn, R. A. L. Defended to the nines: 25 years of resistance gene cloning identifies nine mechanisms for R protein function. Plant Cell 30, 285–299 (2018).
Li,W., Deng, Y., Ning, Y., He, Z. & Wang, G. L. Exploiting broad-spectrum disease resistance in crops: from molecular dissection to breeding. Annu. Rev. Plant Biol. 71, 575–603 (2020).
Ngou, B. P. M., Ahn, H.-K., Ding, P. & Jones, J. D. G. Mutual potentiation of plant immunity by cell-surface and intracellular receptors. Nature 592, 110–115 (2021).
Yuan, M. et al. Pattern-recognition receptors are required for NLR-mediated plant immunity. Nature 592, 105–109 (2021).
Bürger, M. & Chory, J. Stressed out about hormones: how plants orchestrate immunity. Cell Host Microbe 26, 163–172 (2019).
Broekaert, W. F., Delauré Sl, M. F. C., Bolle, D., De Bolle, Mf, B. P. A. & Cammue, B. P. The role of ethylene in host-pathogen interactions. Annu. Rev. Phytopathol. 44, 393–416 (2006).
Washington, E. J. et al. Pseudomonas syringae type III effector HopAF1 suppresses plant immunity by targeting methionine recycling to block ethylene induction. Proc. Natl Acad. Sci. USA 113, E3577–E3586 (2016).
Zhao, S. et al. A viral protein promotes host SAMS1 activity and ethylene production for the benefit of virus infection. eLife 6, e27529 (2017).
Dean, R. et al. The top 10 fungal pathogens in molecular plant pathology. Mol. Plant Pathol. 13, 414–430 (2012).
Mosquera, G., Giraldo, M. C., Khang, C. H., Coughlan, S. & Valent, B. Interaction transcriptome analysis identifies Magnaporthe oryzae BAS1–4 as biotrophy-associated secreted proteins in rice blast disease. Plant Cell 21, 1273–1290 (2009).
Mentlak, T. A. et al. Effector-mediated suppression of chitin-triggered immunity by Magnaporthe oryzae is necessary for rice blast disease. Plant Cell 24, 322–335 (2012).
Orbach, M. J., Farrall, L., Sweigard, J. A., Chumley, F. G. & Valent, B. A telomeric avirulence gene determines efficacy for the rice blast resistance gene Pi-ta. Plant Cell 12, 2019–2032 (2000).
Kanzaki, H. et al. Arms race co-evolution of Magnaporthe oryzae AVR-Pik and rice Pik genes driven by their physical interactions. Plant J. 72, 894–907 (2012).
Sweigard, J. A. et al. Identification, cloning, and characterization of PWL2, a gene for host species-specificity in the rice balst fungus. Plant Cell 7, 1221–1233 (1995).
Wu, J. et al. Comparative genomics identifies the Magnaporthe oryzae avirulence effector AvrPi9 that triggers Pi9-mediated blast resistance in rice. New Phytol. 206, 1463–1475 (2015).
Yoshida, K. et al. Association genetics reveals three novel avirulence genes from the rice blast fungal pathogen Magnaporthe oryzae. Plant Cell 21, 1573–1591 (2009).
Li, W. et al. The Magnaporthe oryzae avirulence gene AvrPiz-t encodes a predicted secreted protein that triggers the immunity in rice mediated by the blast resistance gene Piz-t. Mol. Plant Microbe Interact. 22, 411–420 (2009).
Deng, Y. et al. Epigenetic regulation of antagonistic receptors confers rice blast resistance with yield balance. Science 355, 962–965 (2017).
Xie, Z. et al. A nucleotide-binding site-leucine-rich repeat receptor pair confers broad-spectrum disease resistance through physical association in rice. Phil. Trans. R. Soc. B 374, 20180308 (2019).
Zhou, B. et al. The eight amino-acid differences within three leucine-rich repeats between Pi2 and Piz-t resistance proteins determine the resistance specificity to Magnaporthe grisea. Mol. Plant Microbe Interact. 19, 1216–1228 (2006).
Xie, X. et al. PPPDE1 promotes hepatocellular carcinoma development by negatively regulate p53 and apoptosis. Apoptosis 24, 135–144 (2019).
Shimizu, T. et al. Two LysM receptor molecules, CEBiP and OsCERK1, cooperatively regulate chitin elicitor signaling in rice. Plant J. 64, 204–214 (2010).
Suh, H. Y. et al. Crystal structure of DeSI-1, a novel deSUMOylase belonging to a putative isopeptidase superfamily. Proteins 80, 2099–2104 (2012).
Komander, D. & Rape, M. The ubiquitin code. Annu. Rev. Biochem. 81, 203–229 (2012).
Chen, X. L. et al. Proteomic analysis of ubiquitinated proteins in rice (Oryza sativa) after treatment with pathogen-associated molecular pattern (PAMP) elicitors. Front. Plant Sci. 9, 1064 (2018).
Iwai, T., Miyasaka, A., Seo, S. & Ohashi, Y. Contribution of ethylene biosynthesis for resistance to blast fungus infection in young rice plants. Plant Physiol. 142, 1202–1215 (2006).
Singh, R. et al. Magnaporthe oryzae effector AVR-Pii helps to establish compatibility by inhibition of the rice NADP-malic enzyme resulting in disruption of oxidative burst and host innate immunity. Mol. Cells 39, 426–438 (2016).
De la Concepcion, J. C. et al. Polymorphic residues in rice NLRs expand binding and response to effectors of the blast pathogen. Nat. Plants 4, 576–585 (2018).
Maqbool, A. et al. Structural basis of pathogen recognition by an integrated HMA domain in a plant NLR immune receptor. eLife 4, 08709 (2015).
Wang, W. et al. Genomic variation in 3,010 diverse accessions of Asian cultivated rice. Nature 557, 43–49 (2018).
Huang, X. et al. Genome-wide association study of flowering time and grain yield traits in a worldwide collection of rice germplasm. Nat. Genet. 44, 32–39 (2011).
Hu, C. et al. Metabolic variation between japonica and indica rice cultivars as revealed by non-targeted metabolomics. Sci. Rep. 4, 5067 (2014).
Liao, J. et al. Pathogen effectors and plant immunity determine specialization of the blast fungus to rice subspecies. eLife 5, e19377 (2016).
Axtell, M. J. & Staskawicz, B. J. Initiation of RPS2-specified disease resistance in Arabidopsis is coupled to the AvrRpt2-directed elimination of RIN4. Cell 112, 369–377 (2003).
Mackey, D., Holt, B. F. III, Wiig, A. & Dangl, J. L. RIN4 interacts with Pseudomonas syringae type III effector molecules and is required for RPM1-mediated resistance in Arabidopsis. Cell 108, 743–754 (2002).
He, J. et al. A LysM receptor heteromer mediates perception of arbuscular mycorrhizal symbiotic signal in rice. Mol. Plant 12, 1561–1576 (2019).
Ma, X. et al. A robust CRISPR/Cas9 system for convenient, high-efficiency multiplex genome editing in monocot and dicot Plants. Mol. Plant 8, 1274–1284 (2015).
Zhai, K. et al. RRM transcription factors interact with NLRs and regulate broad-spectrum blast resistance in rice. Mol. Cell 74, 996–1009 (2019).
Park, C.-H. et al. The Magnaporthe oryzae effector AvrPiz-t targets the RING E3 ubiquitin ligase APIP6 to suppress pathogen-associated molecular pattern-triggered immunity in rice. Plant Cell 24, 4748–4762 (2012).
Acknowledgements
We thank J. Zhou and J. Li for critical reading and discussion; X. Wei and G. Zhang for providing the rice germplasm and CSSL; J. Zhang for help with ethylene measurement; S. Wang for help with Met measurement; B. Liu, C. Wang and Z. Gu for help with bioinformatic analysis; and X. Fu for sharing HA-Ub plasmids. This work was supported by grants from the National Natural Science Foundation of China (32088102, 31720103913 and U20A2021), the Chinese Academy of Sciences (XDB27040201 and XDA24010304), the National Key Research and Development Program of China (2016YFD0100600), the Science and Technology Commission of Shanghai Municipality (19391900300), the Project of Special Funding for Crop Science Discipline Development of Yangzhou University (yzuxk202006) and the National Key Laboratory of Plant Molecular Genetics. K.Z. was supported by the National Postdoctoral Program for Innovative Talents (BX20190346) and by the Shanghai Postdoctoral Excellence Program.
Author information
Authors and Affiliations
Contributions
K.Z. and Z.H. conceived and designed the experiments. K.Z. and D.L. performed most experiments, including the Y2H, SLC, BiFC, co-IP, subcellular localization, cell-free degradation assay, deubiquitination assay, ROS detection and western blots. K.Z., H.L. and Y.W. performed Met and ethylene measurement. K.Z., H.L. and F.J. performed RNA analysis. K.Z., D.L., B.Y. and X.G. performed pathoassays. D.L. and H.L. performed agronomic traits analysis. K.Z., J.L., Z.L., L.H., X.W. and J.-Y.L. generated material used in this study. J.M. performed the domestication analysis. H.G. and B.H. supervised Y.W. and J.M., respectively. L.Z., E.W., Y.D., C.-K.W., H.G. and B.H. provided theoretical contributions to the project. K.Z., D.L. and Z.H. analysed the data and wrote the paper.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Nature thanks Nicholas Talbot and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Screening of PICIs and PigmR–PICI1 interaction analysis.
a, The workflow for identifying the proteins involved in both PTI and ETI. Tandem Mass Tag (TMT)-based proteomic analysis was performed to investigate the global changes of rice protein profiling with chitin treatment, the blue circle denotes the chitin-induced 413 candidate proteins. Total 11 PigmR-CC interacting proteins were identified through yeast-two-hybrid screen and shown with the pink circle. The overlap between the blue and pink circles indicates 3 PICI candidates. b, Protein alignment of PICI1 and its homologs in A. thaliana, S. cerevisiae, C. elegans, M. musculus and H. sapiens (NP_194296.1, NP_594707.2, NP_741592.1, NP_077244.1 and NP_057160.2, respectively) with MegAlign software. The conserved PPPDE region is underlined in red and the conserved site Cys115 residue is marked with red arrow. c, Immunodetection of protein expression in yeast. pDEST22 (EV), pDEST22-PICI1 (1) and pDEST22-Os07g16130 (2) detected using anti-GAL4-AD antibody. pDEST32-PigmR (BD-PigmR), pDEST32-PigmR-CC (BD-PigmR-CC), pDEST32-PigmR-NBS (BD-NBS) and pDEST32-PigmR-LRR (BD-LRR) detected using anti-GAL4-BD antibody. The asterisks indicate the target proteins. d, Protein alignment of PICI1 and its homologs in rice. e, PigmR and PigmR-CC specifically interact with PICI1 but not its homologs in Y2H assay. f, Immunodetection of protein levels in yeast. Upper panel (AD), pDEST22 (lane 1), pDEST22-PICI1 (lane 2), -Os02g43840 (lane 3), -Os03g01130 (lane 4), -Os03g10200 (lane 5), -Os04g46290 (lane 6), -Os06g01780 (lane 7), -Os06g08360 (lane 8), -Os06g36490 (lane 9), detected using anti-GAL4-AD antibody; middle panel (BD), pDEST32-PigmR-CC, detected using anti-GAL4-BD antibody. The asterisks indicate the target proteins. g, SLC assay of PigmR-PICI1 interaction in N. benthamiana. PIBP1 and Os07g16130 served as positive and negative control, respectively. Fluorescence signal intensity is indicated. h, Western blot confirming expression of proteins in N. benthamiana. Note that cLuc-PICI1 was detected using anti-PICI1 antibody. Short, long = short or long exposure. i, PICI1-YFP was co-expressed with various known organelle markers as indicated in rice protoplasts. mCherry = cytoplasmic marker, NLS-RFP = nucleus marker, ST-mCherrry = trans-Golgi cisternae marker, SYP61-mCherry = trans-Golgi network/early endosome (TGN/EE) marker, ER-mCherry = endoplasmic reticulum marker, OsRac1-mCherrry = cell member marker, Chl Auto = Chloroplast autofluorescence. j, Western blot confirming expression of PICI1-YFP without free YFP in rice protoplast. k, Localization of PICI1-GFP in representative root cells of stable transgenic plants. DAPI staining indicates the nucleus. l, Bimolecular fluorescence complementation (BiFC) verifies the PigmR-CC/PICI1 interaction in rice protoplasts. Pish-CC served as a negative control. Ponceau S staining (c, f) or Actin (h, j) served as loading control. Scale bars, 5 µm (i, l) or 10 μm (k). Experiments were repeated twice (c, f, h) or three times with similar results (e, g, i–l).
Extended Data Fig. 2 PICI1 is involved in PigmR-, Pizt- and Pi9- but not Pish-mediated blast resistance.
a, i, l, o, Schematic of two independent PICI1 knockout lines in NIL-Pigm (a), ZH11 (Pizt-containing) (i), Ky-Pi9 (Pi9-containing) (l) and NIPB (Pish-containing) (o), respectively. b, j, m, p, Protein analysis of PICI1 in the corresponding transgenic plants using an anti-PICI1 antibody. The asterisk indicates the endogenous PICI1 and the band around 63 kDa indicates the PICI1-GFP fusion protein (b, p). c, Blast resistance of NIL-Pigm, PICI1-KO/NIL-Pigm and PICI1-OE/NIL-Pigm lines at 7dpi with punch injection inoculation (TH12). NIPB served as a susceptible control. d, Y2H assay of interactions between PICI1 and the CC domains of PigmR, Pi9, Pizt and Pish. EV, empty vector. e, Immunodetection of protein levels in yeast. Upper panel (AD), pDEST22-PICI1, detected using anti-GAL4-AD antibody; middle panel (BD), PigmR-CC, Pi9-CC, Pizt-CC and Pish-CC, detected using anti-GAL4-BD antibody. f, SLC confirmation of the PICI1-Pizt-CC and PICI1/Pi9-CC interactions in N. benthamiana. PigmR-CC and Pish-CC served as positive and negative control, respectively. Fluorescence signal intensity is indicated. g, Relative luciferase activity of protein-protein interactions was measured. One-way ANOVA with Tukey’s test (mean ± s.d.; n = 5, biologically independent samples). Different letters indicate significant difference at P < 0.05. h, Western blot analysis confirming the expression level of each protein in N. benthamiana. k, n, q, Punch inoculation of PICI1-KO transgenic plants in ZH11 (k), Ky-Pi9 (n) and NIPB (q). Blast resistance of WT and two representative transgenic lines, 5 dpi with avirulent strain YN2 (k, q) or Guy11 (n). TH12 served as a susceptible control (q). r, Transient assay of cell death in rice protoplast. Relative luciferase activity was measured after transformation with avrPizt-mCherry in rice protoplasts derived from ZH11 or PICI1-KO/ZH11. Empty vector (EV) was used as a negative control. Two-tailed Student’s t-test (mean ± s.d.; n = 9, biologically independent samples). s, Western blot analysis confirming expression of proteins in rice protoplasts. Actin (b, h, j, m, p, s) or Ponceau S staining (e) served as loading control. The asterisks indicate the target proteins. For c, k, n, q, data were analysed by two-tailed Student’s t-test. Fungal growth (mean ± s.d.; n = 3, biologically independent samples), lesion lengths or areas (mean ± s.d.; n = numbers of biologically independent samples in the graphs). Scale bars, 1 cm. Asterisks represent significant difference (*P < 0.05, **P < 0.01) (c, k, n, q, r). Exact P values are provided in Supplementary Table 4 (c, g, k, n, q, r). Experiments were repeated twice (b, e, h, j, m, p) or three times with similar results (c, d, f, g, k, n, q, r, s).
Extended Data Fig. 3 PICI1 is involved in chitin- and flg22-induced PTI responses.
a, g, p, Induction of the PTI-related defence genes OsPAL1 and KS4 at 1 h after chitin (a, g) or flg22 (p) incubation in WT, PICI1-KO and PICI1-OE lines in NIL-Pigm (a, p) or NIPB (g) background. Two-tailed Student’s t-test (mean ± s.d.; n = 3, biologically independent samples). b, c, Induction of PICI1 in NIL-Pigm upon chitin/flg22 treatment (b) and during M. oryzaeTM21 infection (c) with water as control. d, h, PICI1 was induced upon M. oryzae TM21 infection (d) and chitin treatment (h) with water as control. The asterisk indicates endogenous PICI1 (d). The relative PICI1 protein abundance was indicated. e, Induction of PTI-related defence genes KS4 and OsPAL1 in NIL-Pigm leaves during M. oryzae TM21 infection. f, o, Chitin- (f) or flg22- (o) induced ROS burst in WT, PICI1-KO and PICI1-OE lines in NIPB (f) or NIL-Pigm (o) background. Data are mean ± s.d. (n = 9, biologically independent samples). i, Disease resistance of PICI1 transgenic plants in NIPB. j, l, Schematic of two independent PICI1/OsCERK1 double knockout (j) and PICI1-OE/OsCERK1-KO (l) lines in NIPB. k, m, Protein levels of PICI1 (k) or PICI1-GFP (m) in PICI1/OsCERK1-KO (k) or PICI1-OE/OsCERK1-KO (m). n, Punch inoculation of OsCERK1-KO, PICI1/OsCERK1-KO and PICI1-OE/OsCERK1-KO transgenic lines. q, r, PICI1 was induced upon flg22 treatment (q) and bacterial Xoo (strain, PXO99A) infection (r) with water as control. s–u, Disease resistance to bacterial Xoo in the PICI1 transgenic lines in NIL-Pigm. Triangles represent the end of bacterial infection sites. For i, n, t, u, data were analysed by two-tailed Student’s t-test. Lesion lengths (mean ± s.d.; n = numbers of biologically independent samples in the graphs), fungal or bacterial growth (mean ± s.d.; n = 3, biologically independent samples). For b, c, e, data are mean ± s.d. (n = 3, biologically independent samples). The rice ACTIN1 served as an internal control (a-c, e, g, p). Actin was detected as a loading control (d, h, k, m, q, r). Blast resistance of representative transgenic lines, 5 dpi with virulent strain TH12 (i, n). Scale bars, 1 cm (i, n) or 5 cm (s). Asterisks represent significant difference (*P < 0.05, **P < 0.01) (a, g, i, n, p, t, u). Experiments were repeated twice (k, m) or three times with similar results (a–i, n, o–u).
Extended Data Fig. 4 PICI1 functions as a deubiquitinase, and interacts with and stabilizes OsMETS1.
a, His-PICI1 and His-PICI1-PPPDE displayed cleavage activity toward K48- and K63-linked but not linear ubiquitin in vitro. His alone served as a control. b, Homology model of PICI1-PPPDE domain (cyan) aligned with Desi-1 (PDB ID code 2WP7) (blue) using PyMOL. The putative catalytic dyad residues of H41 and C115 in PICI1 are shown in yellow and the catalytic residues of Desi-1 H38 and C108 are shown in magenta. The boxed section is magnified for close view. c, Induction of KS4, PR10 and PR5 in PICI1-YFP or PICI1C115S-YFP overexpression protoplast after chitin treatment (1 hpi) with the empty vector control. Lowercase letters indicate statistical significance (P < 0.05). One-way ANOVA with Tukey’s test. Data are mean ± s.d. (n = 3, biologically independent samples). Exact P values are provided in Supplementary Table 4. d, Ubiquitin conjugation in total protein extracts from 2-week-old NIL-Pigm and PICI1-KO/NIL-Pigm, after spraying TH12 or TM21 with water as control for 36 h. e, Morphological phenotype of mature plants of wild-type and PICI1-KO/NIL-Pigm. No obvious change in morphology was observed in the PICI1-KO plants. Scale bar, 10 cm. f, Venn diagram showing the number of total ubiquitinated proteins (blue), PICI1 interacting candidates identified in IP–MS (red), PTI-related ubiquitinated proteins (green) and overlapping proteins. g, Protein alignment of OsMETS1 and its homologs in sorghum, soybean, maize, wheat and Arabidopsis (XP_021301657.1, XP_003542326.1, PWZ52049.1, TraesCS4D02G012900.2 and XP_002871787.1) with MegAlign software. h, Ethylene biosynthesis pathway in plants. S-AdoMet, S-adenosylmethionine; Hcys, homocysteine; THF, tetrahydrofolate; SAH, S-adenosylhomocysteine; Ado, adenosine; ACS, 1-aminocyclopropane-1-carboxylate synthase; ACC, 1-aminocyclopropane-1-carboxylate. i, A SLC assay of OsMETS1 and PICI1 interaction in N. benthamiana. PICI2 served as a negative control. Fluorescence signal intensity is indicated. j, Western blot confirming protein expression in N. benthamiana. k, GST-PICI1 pulls down His-OsMETS1 in vitro. l, Subcellular localization of OsMETS1-GFP in OsMETS1-GFP/NIL-Pigm root. DAPI staining indicates the nucleus. Scale bars, 10 μm. m, Immunodetection of OsMETS1-GFP in transgenic lines, with NIL-Pigm as a negative control. n, Subcellular localization of OsMETS1 and co-localization of PICI1-YFP/OsMETS1-mCherry in rice protoplasts. Scale bars, 5 µm. o, Western blot confirming expression of OsMETS1-mCherry without free mCherry in rice protoplast. mCherry (lane 1), OsMETS1-mCherry (lane 2). p, Western blot confirming expression of OsMETS1-YFP without free YFP in rice protoplast. YFP (lane 1), OsMETS1-YFP (lane 2). q, Degradation of OsMETS1-Flag was proteasome-dependent and delayed by PICI1 in cell-free system. r, PICI1 stabilizes OsMETS1-Flag in rice plants. The lysates from WT, PICI1-KO and PICI1-OE plants were co-incubated with OsMETS1-Flag in the presence or absence of MG132. s, Peptide coverage and ubiquitination sites identified in OsMETS1 by tandem mass spectrometry after M. oryzae (TH12) or water treatment for 36 h. The various peptides identified by mass spectrometry are shown in blue; the lysine acceptor sites detected in red are numbered below with positions; the four specific deubiquitination sites are denoted with asterisks. t, Structure model for OsMETS1. Homology model of the OsMETS1 based on the crystal structure of Arabidopsis Met synthase (PDB ID code 1U1U) and analysed by PyMOL. The four lysines identified as deubiquitination sites were shown in blue. The other lysine acceptor sites were shown in red. u, Cell-free degradation assays show the proteasome-dependent degradation of His-OsMETS1 and delayed degradation of His-OsMETS1 mutant variants. Purified recombinant His-OsMETS1 and its mutants were incubated with protein extracts from non-infected NIPB. His alone served as a negative control. v, The OsMETS1 mutants showed higher protein accumulation compared to wild-type OsMETS1 in protoplasts. The YFP tag protein was expressed as an internal control. The transcript levels were determined by semi-quantitative PCR. Actin was detected as a loading control (d, j, m, o, p). Ponceau S staining served as loading control (q, r, u). The protein abundance was quantified using ImageJ and indicated under lanes (q, r, u). Experiments were repeated twice (j, m, o, p) or three times with similar results (a, c, d, i, k, l, n, q, r, u, v).
Extended Data Fig. 5 PICI1 directly deubiquitinates and stabilizes OsMETS.
a, Induction of PR5 and PR1 in OsMETS1, OsMETS1K28R, OsMETS1K196R, OsMETS1K458R and OsMETS1K740R overexpression protoplasts after chitin treatment (1 hpi). Lowercase letters indicate statistical significance (P < 0.05). One-way ANOVA with Tukey’s test. Data are mean ± s.d. (n = 3, biologically independent samples). Exact P values are provided in Supplementary Table 4. b, Cell-free degradation assays show the delayed degradation of His-OsMETS14KR mutant variant, compared to His-OsMETS1K28R. Purified recombinant fusion proteins were incubated with non-infected NIPB extracts. c, The OsMETS14KR mutant showed higher protein accumulation compared to wild-type OsMETS1 and OsMETS1K28R in protoplasts. The YFP tag protein was expressed as an internal control. The transcript levels were determined by semi-quantitative PCR. d, Stability of the OsMETS1-Flag depends on His-PICI1 but not the His-PICI1C115S in cell-free degradation assay. e, Deubiquitination analysis of ubiquitin-modified OsMETS1-GFP, immunoprecipitated from OsMETS1-GFP/NIL-Pigm plants, by incubating with recombinant His-PICI1 or His-PICI1C115S respectively. f, Tail-to-head organization of OsMETS1 and OsMETS2 genes in rice. g, Protein alignment of OsMETS1 and OsMETS2 with MegAlign software. h, Co-IP assay of PICI1 with OsMETS2 in rice. PICI2 served as a negative control. i, A SLC assay showed that both OsMETS1 and OsMETS2 interacted with PICI1 in N. benthamiana. PICI2 was used as a negative control. Fluorescence signal intensity is indicated. j, Western blot confirming protein expression in N. benthamiana. k, Cell-free degradation assay showed the proteasome-dependent degradation of OsMETS2-Flag in N. benthamiana. l, Cell-free degradation assay showed the delayed degradation of OsMETS2 in presence of recombinant His-PICI1. m, Co-IP assay of PICI1 with endogenous OsMETS in rice. The GFP negative control and PICI1-GFP were constitutively expressed in transgenic rice plants. anti-OsMETS, recognizing both OsMETS1 and OsMETS2. n, Cell-free degradation assay showed that endogenous OsMETS degradation was delayed by PICI1. o, Immunodetection of the recombinant His-OsMETS1 and His-OsMETS2 using the anti-OsMETS antibody. His alone served as a negative control. p, A schematic diagram of two independent OsMETS1-KO lines in NIL-Pigm background. q, Protein levels of OsMETS in NIL-Pigm, OsMETS1-GFP/NIL-Pigm and OsMETS1-KO/NIL-Pigm plants using the anti-OsMETS antibody. The band around 75 kDa indicates the endogenous OsMETS. r, Relative transcript levels of OsMETS in the wild-type, PICI1-KO and PICI1-OE lines in NIL-Pigm and NIPB background. Data are mean ± s.d. (n = 3, biologically independent samples). s, Analysis of endogenous OsMETS ubiquitination. OsMETS was immunoprecipitated from wild-type NIL-Pigm with anti-OsMETS or anti-IgG as negative control. Short, long = Short or long exposure. The protein abundance was quantified using ImageJ and indicated under lanes (b, d, k, l, n, q). Ponceau S staining (b, d, k, l, n, o) or Actin (e, j, q, s) was used as a control for equal loading. The rice ACTIN1 served as an internal control (a, r). Experiments were repeated twice (h, j, m, n, o, s) or three times (a–e, i, k, l, q, r) with similar results.
Extended Data Fig. 6 OsMETS functions in rice PTI and ETI.
a, b, c, d, OsMETS is induced upon treatment of chitin (a) or flg22 (b), and infection of Xoo (PXO99A) (c) or M. oryzae (TM21) (d). e, Protein accumulation of OsMETS in NIL-Pigm, PICI1-KO/NIL-Pigm and PICI1-GFP/NIL-Pigm in early infection stage (TM21). f, g, Decreased ubiquitination of OsMETS1-GFP in PTI (f) and ETI (g). The protein extracts from OsMETS1-GFP/NIL-Pigm plants incubated with chitin for 1h (f) or infected with TH12 for 36 h (g), with water as control, was analysed. NIL-Pigm served as a negative control. h, A schematic diagram of two independent OsMETS2-KO lines in NIL-Pigm. i, Decreased protein levels of OsMETS in OsMETS2-KO/NIL-Pigm compared with NIL-Pigm. j, l, Basal blast resistance of NIL-Pigm, OsMETS2-KO/NIL-Pigm (j) and OsMETS2-GFP/NIL-Pigm (l) at 5 dpi with punch inoculation (TM21). k, Immunodetection of the OsMETS2-GFP in the two independent transgenic lines, with NIL-Pigm as a negative control. m, n, Disease resistance of WT, OsMETS1-KO (m), OsMETS1-OE (m) and OsMETS2-KO (n) were shown, 5 dpi with TH12, NIPB served as a susceptible control. o, Relative transcript levels of OsMETS1, OsMETS2 and OsMETS in NIL-Pigm and two OsMETS-RNAi lines. p, Protein levels of OsMETS in NIL-Pigm and OsMETS-RNAi lines. q, Morphological phenotype of NIL-Pigm and OsMETS-RNAi/NIL-Pigm lines at mature stage in the paddy field. r, OsMETS-RNAi/NIL-Pigm plants significantly reduced plant height (upper panel), tiller number (middle panel), grain yield per plant (lower panel). s, Relative Met contents in WT and OsMETS-RNAi/NIL-Pigm. t, Increased Met accumulation in NIL-Pigm seedling shoots after root incubation with 25 mM Met for 48 h. u, Increased blast resistance after Met treatment in NIPB. v, Induction of the defence gene KS4 at 48 h after root dipping with 25 mM Met solution or water in NIL-Pigm. w, y, Basal blast resistance of Met-treated OsMETS1-KO/NIL-Pigm at 5 dpi (w) or PICI1-KO/NIL-Pigm at 7 dpi (y). Met solution (10 mM) or water was supplied from the roots for 48 h and plants were then inoculated with TM21. x, Relative Met contents in WT and PICI1-KO/NIL-Pigm lines. For j, l, m, n, data were analysed by two-tailed Student’s t-test. Lesion lengths (mean ± s.d.; n = numbers of biologically independent samples in the graphs), fungal growth (mean ± s.d.; n = 3, biologically independent samples). For u, w, y, one-way ANOVA with Tukey’s test. Lesion areas (mean ± s.d.; n = numbers of biologically independent samples in the graphs), fungal growth (mean ± s.d.; n = 3, biologically independent samples). Lowercase letters indicate statistical significance (P < 0.05). For r, s, t, x, data were analysed by two-tailed Student’s t-test (mean ± s.d.; n = numbers of biologically independent samples in the graphs). In o, v, two-tailed Student’s t-test (mean ± s.d.; n = 3, biologically independent samples). Asterisks indicate significant differences (*P < 0.05, **P < 0.01) (j, l-o, r–t, v, x). Exact P values are provided in Supplementary Table 4 (j, l–o, r–y). Scale bars, 1 cm (j, l–n, u, w, y), 10 cm (q). Relative protein abundance was indicated below (a–e, p). Actin was detected as a loading control (a–g, i, k, p). The rice ACTIN1 served as an internal control (o, v). Experiments were independently repeated twice (i, k, p, r) or three times (a–g, j, l–o, s–y) with similar results.
Extended Data Fig. 7 PICI1–OsMETS module confers blast resistance through regulating Met–ethylene biosynthesis.
a, High Met concentrations (10 and 25 mM) inhibit blast fungal growth on medium. The M. oryzae isolate TH12 was inoculated on the complete medium (CM) with Met supplement. The colony diameters were measured. One-way ANOVA with Tukey’s test (mean ± s.d.; n = 5, biologically independent samples). b, c, Ethylene contents in WT and transgenic lines of OsMETS1 (OsMETS1-KO and OsMETS1-OE) (b) and PICI1 (PICI1-KO and PICI1-OE) (c) infected by TM21 (2-week-old, 36 hpi). Two-tailed Student’s t-test (mean ± s.d.; n = 3, biologically independent experiments). d, Induction of ERF63 and ERF73 by ethylene treatment in NIL-Pigm. Two-week-old seedlings were placed into sealed containers with 10 or 100 ppm (μl/L) ethylene, or air as control for 3 h to analyse gene expression. e, f, Suppression of OsMETS1 (e) and OsMETS2 (f) expression by ethylene treatment (100 ppm) in NIL-Pigm. g, Relative expression of ERF63 at 1 d treatment with 10 μM AVG or 20 μM ACC or water. ERF63 was chosen as a marker gene for AVG and ACC responses. Two-tailed Student’s t-test (mean ± s.d.; n = 3, biologically independent samples). h, Effects of AVG and ACC on basal blast resistance in NIPB. Two-week-old seedlings were pre-treated with 10 μM AVG, or 20 μM ACC or water for 1 d and then spray-inoculated with rice blast (TH12), 5 dpi. i, j, Basal blast resistance of ACC-treated PICI1-KO/NIL-Pigm (i) and OsMETS-RNAi/NIL-Pigm (j) plants. ACC solution (20 μM) or water was supplied from the leaves and inoculated with TM21, 5 dpi. k, l, Induction of OsACS2 in NIL-Pigm leaves at early stage by TM21 (k, PTI) or late stage by TH12 (l, ETI) infection. m, Enhanced ethylene production in NIL-Pigm after rice blast infection (TH12 or TM21), with water as a control for 36 h. One-way ANOVA with Tukey’s test (mean ± s.d.; n = 3, biologically independent experiments). n, A schematic diagram of OsACS2-KO lines in NIL-Pigm. o, Basal blast resistance in NIL-Pigm and OsACS2-KO/NIL-Pigm, at 5 dpi with punch inoculation (TM21). p, Disease resistance of wild type and OsACS2-KO lines were shown, 5 dpi with TH12, NIPB served as a susceptible control. q, Met enhances rice blast resistance via ethylene biosynthesis. A Met solution (10 mM) or water was supplied from NIL-Pigm roots for 48 h and pre-treated with 10 μM AVG for 36 h before spray inoculation with rice blast (TM21). One-way ANOVA with Tukey’s test. Lesion areas (mean ± s.d.; n = 5, biologically independent samples), fungal growth (mean ± s.d.; n = 3, biologically independent samples). r, Protein levels of PICI1 in NIL-Pigm (left panel) and NIPB (right panel) plants after inoculation with M. oryzae at the indicated time points. Actin was detected as a loading control. The protein abundance was quantified using ImageJ and indicated under lanes. s, RNA expression of PICI1 was not significantly affected in late stage by blast infection (TH12 and TM21) in NIL-Pigm. For d, e, f, k, l, s, data are mean ± s.d. (n = 3, biologically independent samples). For h, i, j, o, p, data were analysed by two-tailed Student’s t-test. Lesion lengths or areas (mean ± s.d.; n = numbers of biologically independent samples in the graphs), fungal growth (mean ± s.d.; n = 3, biologically independent samples). Lowercase letters indicate statistical significance (P < 0.05) (a, m, q). Asterisks indicate significant differences (*P < 0.05, **P < 0.01) (b, c, g–j, o, p). Exact P values are provided in Supplementary Table 4 (a–c, g-j, m, o–q). The rice ACTIN1 served as an internal control (d–g, k, l, s). Scale bars, 1 cm (h–j, o–q). Experiments were repeated twice (d–g) three times with similar results (a–c, h–j, k–m, o-q, r, s).
Extended Data Fig. 8 PigmR protects PICI1 from AvrPi9-mediated degradation.
a, His-PICI1 could be pulled down by seven effectors (AvrPWL2, AvrPi9, AvrPikC, AvrPia, AvrPizt, AvrPii and AvrPikD fused with MBP) in vitro. MBP alone served as a negative control. b, SLC verification of the interactions between PICI1 and the three blast fungal effectors, AvrPWL2, AvrPi9 and AvrPii in N. benthamiana. Note that AvrPikD/Pik-HMA and AvrPii/OsNADP-ME2 served as positive controls. c, Relative luciferase activity of protein-protein interactions was measured. Data are mean ± s.d. (n = 6, biologically independent samples). d, Western blot confirming protein expression in N. benthamiana. Note that 1-12 represented PICI1/AvrBAS4, PICI1/AvrPWL2, PICI1/AvrPi9, PICI1/AvrPita, PICI1/AvrSlp1, PICI1/AvrPikC, PICI1/AvrPia, PICI1/AvrPizt, PICI1/AvrPii, OsNADP-ME2/PICI1, PICI1/AvrPikD and Pik-HMA/AvrPikD, respectively. e–g, SLC assays showed that PICI1, but not PICI2, specifically interacted with AvrPWL2 (e), AvrPi9 (f) and AvrPii (g) in N. benthamiana. Fluorescence signal intensity is indicated. Protein expression in N. benthamiana was detected by western blot (e-g, down panel). h, Protein levels of endogenous PICI1 in AvrPi9-Flag (AvrPi9-OE), AvrPWL2-Flag (AvrPWL2-OE), AvrPizt-Flag (AvrPizt-OE), and AvrPii-Flag (AvrPii-OE), compared with NIL-Pigm. The protein abundance was quantified and indicated under lanes. i, Subcellular localization of AvrPi9 and co-localization of PICI1-YFP/AvrPizt-mCherry in rice protoplasts. j, Western blot analysis confirming expression of AvrPi9-mCherry or AvrPi9-YFP without free mCherry or YFP in rice protoplasts. k, BiFC confirms the PICI1/AvrPi9 interaction in rice protoplasts. PICI2 served as a negative control. l, Basal blast resistance of WT and AvrPi9-OE lines, 5 dpi with TM21. Scale bars, 1 cm. Data were analysed by two-tailed Student’s t-test. Lesion lengths (mean ± s.d.; n = numbers of biologically independent samples in the graphs), fungal growth (mean ± s.d.; n = 3, biologically independent samples). m, n, Induction of the PTI-related defence genes OsPAL1 (m) and KS4 (n) at 1 h after chitin incubation in WT and AvrPi9-OE. Two-tailed Student’s t-test (mean ± s.d.; n = 3, biologically independent samples). The rice ACTIN1 served as an internal control. o, Cell-free degradation shows AvrPi9FL-mediated PICI1-GFP degradation. Protein extracts were prepared from two-week-old PICI1-GFP/NIL-Pigm seedlings and then incubated with AvrPi9FL-mCherry or AvrPizt-mCherry expressed in N. benthamiana. p, Cell-free degradation shows the AvrPi9FL-mediated PICI1-GFP degradation in NIPB (left panel) and NIL-Pigm (right panel) with or without MG132. Protein extracts were prepared from PICI1-GFP/NIPB and PICI1-GFP/NIL-Pigm seedlings and then incubated with mCherry (upper panels) or AvrPi9-mCherry (lower panels) expressed in N. benthamiana. FL, full length (o, p), and we used the Avrs without signal peptide to make the constructs in the experiments unless otherwise indicated. q, Immunodetection of two independent transformants of TH12AvrPi9 using an anti-S-tag antibody, with its parent TH12 as a negative control. Full length of AvrPi9 tagged by S-tag was transformed into TH12 by Agrobacterium-mediated transformation. r, Pathogenicity test of the transformants with TH12 as a virulent control. s, Protein levels of PICI1 in NIPB (left panel) and NIL-Pigm (right panel) after inoculation with TH12 (upper panel) or TH12AvrPi9 (lower panel). The protein abundance was quantified using ImageJ and indicated under lanes. Ponceau S staining (o–q) or Actin (d–h, j, s) served as loading control. Scale bars, 5 µm (i, k). Asterisks represent significant difference (*P < 0.05, **P < 0.01) (l-n). Exact P values are provided in Supplementary Table 4 (l–n). Experiments were repeated twice (a, d, h, j, q, s) or three times (b, c, e–g, i, k-p, r) with similar results.
Extended Data Fig. 9 PigmR and Pizt protect PICI1 from Avrs binding in a competitive manner.
a, PICI1-interacting effectors including AvrPi9, AvrPWL2 and AvrPii can not activate PigmR-mediated cell death. Relative luciferase activity was measured after transformation with different effectors in NIPB or NIL-Pigm. Empty vector (EV) and MoCDIP1 were used as negative and positive control, respectively. One-way ANOVA with Tukey’s test (mean ± s.d.; n = 3, biologically independent experiments). Different letters indicate significant difference at P < 0.05. b, Western blot analysis confirming expression of proteins in rice protoplasts. Note that 1-5 and 6-10 represented MoCDIP1, EV, AvrPi9, AvrPWL2 and AvrPii in NIPB and NIL-Pigm, respectively. The asterisks indicate the target proteins. c, Increased protein accumulation of PigmR in PigmR-7Myc-His/NIPB after inoculation with TH12 or TH12AvrPi9 with water as a control at the indicated time points. d, PICI1-PigmR interaction is enhanced by M. oryzae infection. Proteins were prepared from transgenic plants PICI1-GFP/NIL-Pigm and PigmR-7Myc-His/NIPB after inoculation with TH12 or water as a control (36 hpi). e, A SLC assay shows that the PigmR disrupts PICI1-AvrPi9 interaction in N. benthamiana. Relative luciferase activity of protein-protein interactions was measured (lower panel). Two-tailed Student’s t-test (mean ± s.d.; n = 6, biologically independent samples). The asterisk represents significant difference (**P < 0.01). f, Western blot confirming protein expression in N. benthamiana. Note that left and right represent PICI1/AvrPi9/GFP and PICI1/AvrPi9/PigmR combination, respectively. g, PigmR-CC outcompetes AvrPi9 for PICI1 binding in vitro. h, PigmR-CC outcompetes AvrPi9 for PICI1 binding in rice protoplasts. i, j, SLC assays show that the PigmR-CC also disrupts PICI1/AvrPWL2 (i) and PICI1/AvrPii (j) interactions in N. benthamiana. Protein expression in N. benthamiana was detected by western blot (lower panel) (i, j). k, l, PigmR-CC outcompetes AvrPii (k) or AvrPWL2 (l) for PICI1 binding in rice protoplasts. The asterisks indicate the target proteins (l). m, n, Protein accumulation of PICI1 and OsMETS in Ky-Pi9 (m) and ZH11 (n) plants after inoculation with avirulent strain Guy11 (m) or YN2 (n) at the indicated time points. o, Protein accumulation of PICI1 in ZH11 and Pizt-KO/ZH11 plants after inoculation with blast strain CH131 (avirulent to Pizt) at the indicated time points. p, SLC assays show that the Pizt-CC also disrupts PICI1/AvrPi9 (left panel), PICI1/AvrPWL2 (middle panel) and PICI1/AvrPii (right panel) interactions in N. benthamiana. Protein expression in N. benthamiana was detected by western blot (lower panel). Actin (b, c, f, i, j, m–o) or Ponceau S staining (p) used as loading control. The protein abundance was quantified using ImageJ and indicated under lanes (c, m-o). Fluorescence signal intensity is indicated (e, i, j, p). Exact P values are provided in Supplementary Table 4 (a, e). Experiments were repeated twice (b, d, f, g, h, k-p) or three times (a, c, e, i, j) with similar results.
Extended Data Fig. 10 PICI1 jap allele shows higher PICI1 expression and increased basal defence, and acetylation of PICI1 promotes its degradation.
a, Twenty-seven SNPs in the PICI1 promoter distribute in different rice varieties based on the 3,000 Rice Genome database33. SNP = 0 indicates the same promoter sequence of PICI1 with NIPB, which was used as the referee. SNP > 0 indicated at least one base change. b, Distribution of the 27 SNPs in japonica and indica rice based on the 3,000 Rice Genome Project database. c, FST of PICI1 and flanking regions between different rice groups. d, Induction of ProPICI1ind-LUC and ProPICI1jap-LUC in rice protoplasts by chitin. LUC activity was measured by normalizing to REN signal. e, japonica rice varieties showed a general higher induction of PICI1 than indica varieties by M. oryzae cocktail at 3 hpi. In box plots, the centre line represents the median, box edges delimit lower and upper quartiles and whiskers show the highest and lowest data points. Two-tailed Student’s t-test (mean ± s.d.; n = numbers of biologically independent samples in the graphs). f, Pathotype test of different M. oryzae strains in two-week-old seedlings of Huajingxian74 using spraying inoculation, indicating YN2 and 99-30-1 virulent toward Huajingxian74. g, Induction of PICI1 at indicated times by M. oryzae infection in Huajingxian74 and CSSL. h, Induction of the defence gene KS4 after blast inoculation in Huajingxian74 and CSSL plants. i, Pathotype test of M. oryzae isolate and Guy11 (avirulent) and TM21 (virulent) toward Pi9-containing rice plant (Ky-Pi9) with punch inoculation. j, Increased acetylation of PICI1 after blast infection. The protein extracts from PICI1-GFP/NIL-Pigm plants challenged with M. oryzae (TM21) or water for 36 h were immunoprecipitated with Ac-K (Acetylated-lysine) or GFP antibody and analysed using anti-GFP or anti-Ac-K antibody, respectively. k, A SLC assay shows the reduced interaction of PICI1K160Q/PigmR-CC and PICI1K160Q/OsMETS1 in comparison with PICI1 in N. benthamiana. The protein and transcript levels of cLuc-PICI1 and cLuc-PICI1K160Q were determined by immunoblot and semi-quantitative PCR. Actin was detected as a loading control (j, k). The rice ACTIN1 served as an internal control (e, g, h). Two-tailed Student’s t-test (mean ± s.d.; n = 3, biologically independent samples) (d, g, h). Asterisks indicate significant differences (*P < 0.05, **P < 0.01) (d, e, g, h). Exact P values are provided in Supplementary Table 4 (d, e, g, h). Scale bars, 1 cm (f, i). Experiments were repeated twice (e, g, h) or three times (d, f, i-k) with similar results.
Supplementary information
Supplementary Figure 1
Uncropped blots and gel images.
Supplementary Table 1
MS/MS and statistical analysis.
Supplementary Table 2
Natural variation analysis for PICI1.
Supplementary Table 3
Primers used in this study.
Supplementary Table 4
Statistical summary—a summary of all statistical analysis is provided. Source data for all of the statistical analysis can be found in the Source Data files.
Source data
Rights and permissions
About this article
Cite this article
Zhai, K., Liang, D., Li, H. et al. NLRs guard metabolism to coordinate pattern- and effector-triggered immunity. Nature 601, 245–251 (2022). https://doi.org/10.1038/s41586-021-04219-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-021-04219-2
- Springer Nature Limited
This article is cited by
-
Early molecular events in the interaction between Magnaporthe oryzae and rice
Phytopathology Research (2024)
-
Deciphering Aphanomyces euteiches-pea-biocontrol bacterium interactions through untargeted metabolomics
Scientific Reports (2024)
-
Allelic variation in rice blast resistance: a pathway to sustainable disease management
Molecular Biology Reports (2024)
-
A genome-wide association study of panicle blast resistance to Magnaporthe oryzae in rice
Molecular Breeding (2024)
-
The OsSGS3-tasiRNA-OsARF3 module orchestrates abiotic-biotic stress response trade-off in rice
Nature Communications (2023)