Abstract
The enzymes of the mitochondrial electron transport chain are key players of cell metabolism. Despite being active when isolated, in vivo they associate into supercomplexes1, whose precise role is debated. Supercomplexes CIII2CIV1-2 (refs. 2,3), CICIII2 (ref. 4) and CICIII2CIV (respirasome)5,6,7,8,9,10 exist in mammals, but in contrast to CICIII2 and the respirasome, to date the only known eukaryotic structures of CIII2CIV1-2 come from Saccharomyces cerevisiae11,12 and plants13, which have different organization. Here we present the first, to our knowledge, structures of mammalian (mouse and ovine) CIII2CIV and its assembly intermediates, in different conformations. We describe the assembly of CIII2CIV from the CIII2 precursor to the final CIII2CIV conformation, driven by the insertion of the N terminus of the assembly factor SCAF1 (ref. 14) deep into CIII2, while its C terminus is integrated into CIV. Our structures (which include CICIII2 and the respirasome) also confirm that SCAF1 is exclusively required for the assembly of CIII2CIV and has no role in the assembly of the respirasome. We show that CIII2 is asymmetric due to the presence of only one copy of subunit 9, which straddles both monomers and prevents the attachment of a second copy of SCAF1 to CIII2, explaining the presence of one copy of CIV in CIII2CIV in mammals. Finally, we show that CIII2 and CIV gain catalytic advantage when assembled into the supercomplex and propose a role for CIII2CIV in fine tuning the efficiency of electron transfer in the electron transport chain.
Similar content being viewed by others
Main
Complex I (CI), CIII2 and CIV of the electron transport chain (ETC) are embedded in the inner mitochondrial membrane and couple oxidoreduction reactions, shown in Fig. 1a, to pumping of protons across the inner mitochondrial membrane, generating the electrochemical gradient used by ATP synthase15,16. CII contributes to the gradient indirectly by reducing ubiquinone. Supercomplexes CICIII2/respirasome may accept electrons mostly from NADH (via CI), while CIII2CIV can accept, via the pool of quinol, electrons originating from both NADH and succinate (CII). Within supercomplexes from different species, the interaction interface between CI and CIII2 is conserved, while CIV binds in different positions4,6,7,9,10,11,12,13,17. Therefore, the organization of mammalian CIII2CIV cannot be inferred from yeast and plant structures. Another open question is the exact role of supercomplex assembly factor 1 (SCAF1), which is known to be required for assembly of CIII2CIV14 but perhaps not for the respirasome, and is absent in plants and yeast. The functional role of the supercomplexes is also debated5. Scaf1-knockout mice and zebrafish show impairment of exercise performance2, growth and fertility18, pointing towards a role of CIII2CIV in the optimization of cellular metabolism (Supplementary Information).
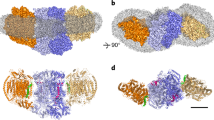
a, Scheme of the respiratory chain with the reactions that each complex catalyses (left). An overview of the existing supercomplexes in mammalian cells is also shown (right). IMS, intermembrane space. b, Cryo-electron microscopy maps of the different CIII2CIV conformations (intermediate locked, assembled locked and mature unlocked, contoured at 2.5, 2.5 and 2.7σ, respectively), coloured by complex as in a. c, Atomic models built into maps shown in b, viewed from the side (top) or from the matrix (bottom). The assembly factor SCAF1 is depicted in blue, while the CIII2 subunits Sub10–Sub11–ISP, not fully resolved in the assembly intermediate, are depicted in dark red. The two CIII protomers are in different shades of yellow.
To answer these questions, we characterized CIII2CIV in its native conformation by solubilizing mitochondrial membranes from ovine and mouse hearts in digitonin, purifying the sample by chromatography in amphipol A8-35 (Extended Data Fig. 1) and performing cryo-electron microscopy analyses. The remaining respirasome and CICIII2 were separated by 3D classification (Extended Data Figs. 2–5) and analysed mainly with respect to CIII2 for comparison. The overall proportion of CIII2CIV (approximately 30%) was lower than CICIII2/respirasome in both species, although relatively higher in mouse than in ovine (Extended Data Fig. 1). Consistent with previous studies1,5,19, in both mouse and ovine samples, CI is found almost exclusively within the supercomplexes, while CIII2 exists also in a free form.
The resulting structures (at approximately 3.1 Å and 3.4 Å resolution for mouse CIII2 and CIV, respectively (Methods)) show that two main conformations of CIII2CIV exist in mammals (Fig. 1b, c, Extended Data Fig. 4b, Supplementary Video 1): one that we refer to as mature unlocked, or simply unlocked, with CIV only peripherally attached to CIII2; and a locked class, with CIV rotated from its unlocked position to tightly contact CIII2 via an extensive interface (Fig. 1b, c). Both conformations are entirely novel compared to yeast CIII2CIV2: in the locked conformation CIV is shifted laterally and is much closer to CIII2; in the unlocked conformation, CIV is almost 180o rotated compared to the yeast structure (Extended Data Fig. 8a). The mammalian supercomplex is also different from plant CIII2CIV13; in plant and yeast, CIV is oriented with COX1 towards CIII2 and COX3 away from it, as opposed to mammals where COX3 faces CIII2 in the unlocked class. The locked state could be further classified to the locked assembled class, containing fully assembled CIII2 and CIV, and the locked intermediate class, containing assembly intermediate of CIII2 (Fig. 1b, c). In the mouse dataset, the unlocked, locked intermediate and locked assembled classes comprised 41%, 21% and 38% of CIII2CIV particles, respectively (Extended Data Fig. 2). In the ovine dataset, the unlocked state dominated with 93% of particles, with only 7% being in the locked state (which was not resolved into subclasses, probably due to the low number of particles; Extended Data Fig. 4a). Comparison of the ovine and mouse structures shows (Extended Data Fig. 4b) that the locked and unlocked conformations are conserved among the mammalian species and since the mouse data yielded higher resolution maps, we used those to build atomic models and refer to them in the rest of this paper, unless otherwise specified. The mouse structures are overall similar to other known mammalian structures, with all the cofactors similarly coordinated and arranged at similar distances (Extended Data Fig. 7f).
Analysis of side-chain density revealed that the C terminus of SCAF1 (residues 56–113) replaces COX7A1 in CIV from all CIII2CIV classes (Extended Data Fig. 7b). The interface between CIII2 and CIV mainly involves non-catalytic subunits and it changes markedly in the two conformations (Figs. 1b, c, 2a, b): in the locked class, subunit 8 (Sub8), Sub6 and Sub7 of CIII2 interact with SCAF1, COX7C and COX8B of CIV, whereas in the unlocked class, the iron-sulfur protein (ISP), the only catalytic subunit involved in the interface, Sub7 and Sub10 of CIII2 interact with COX3, COX5B and COX6A2 of CIV. Sub6 and Sub7 (along with Sub8 and Core1) are involved in the interaction with CI in the respirasome7,8,10, suggesting that a common area of CIII2 interacts either with CI or with CIV in different supercomplexes. Side-chain interactions are shown in Fig. 2a and described in Supplementary Information. Most of these residues are conserved across mammalian species, indicating a conserved interaction between the complexes. Confirming its key role in the assembly of CIII2CIV, SCAF1 links CIV and CIII2 in all classes (Fig. 1c), providing an anchor point and a pivot to guide the locked-to-unlocked transition.
a, Locked class, with interacting subunits in orange (in CIII2) and cyan (in CIV), plus SCAF1 in dark blue. The two side views are depicted. The insets show closed-up views of the interacting sites. Lipids and side chains involved in the interface are shown as sticks. CDL, cardiolipin; 3PE, phosphatidylethanolamine, PC1, phosphatidylcholine. b, Same as a but for the unlocked class. c, Unlocked CIII2CIV model showing the carved density for Sub9 (yellow) and SCAF1 (blue), contoured at 2.5σ. The MPP domains of CIII2 are encircled. d, CIII2 from CICIII2 showing the density of Sub9, contoured at 1.5σ, as in c. The two possible orientations of CICIII2 relative to Sub9 are visualized by the two CI models (in grey) flanking CIII2.
In the intermediate locked class, CIII2 lacks, either entirely or partially, three subunits: ISP, for which only the N-terminal loop is visible in the protomer away from CIV; Sub10, only partially ordered; and Sub11, completely invisible. The subunits lacking or disordered in the intermediate class would occupy the positions that are structurally blocked by CIV in the unlocked class, suggesting that assembly of CIII2 within the supercomplex proceeds unidirectionally via the locked intermediate–locked assembled–unlocked class pathway. Moreover, the CIII2 subunits ISP and Sub10, which are involved in the interface of the unlocked class, are not fully folded in the intermediate class, confirming that the assembly of CIII2CIV starts with the locked class.
Analysis of CIII2 conformational variability within CICIII2, described in Supplementary Information, indicates that CIII2 ‘breathes’ between two conformational states, one ‘open’, close to CIII2 conformation in the intermediate CIII2CIV, and another ‘closed’, reminiscent of the mature CIII2CIV classes. This suggests that, although CIII2 gets compacted upon assembly, it retains a high degree of flexibility, which may be functionally important.
Unexpectedly, the N terminus of SCAF1 is found inside one of the CIII2 matrix processing peptidase (MPP) cavities (Fig 2c, Extended Data Figs. 6, 7). These large cavities on the matrix side of each protomer are formed by subunits Core1 and Core2, which are homologous to mitochondrial MPP and potentially involved in the proteolysis of the precursor of ISP, producing CIII Sub9 (also bound within the MPP cavity) as an N-terminal fragment (78 residues). The activity of MPP has been shown for plant CIII and only tentatively demonstrated for mammalian CIII20. The SCAF1 chain extends from the MPP cavity facing CIV, towards CIV, in all of our classes (Figs. 1b, c, 2c, Extended Data Fig. 6). We could trace the entire SCAF1 from the N terminus inside CIII2 to the C terminus at the end of the transmembrane helix bound to CIV, with a short break for a disordered loop connecting the two complexes. Therefore, SCAF1 forms a physical connection between the two enzymes, its N terminus acting as a ‘hook’ being ‘swallowed’ by the MPP cavity. This unique arrangement ensures that a strong bond is keeping the supercomplex together.
Our observation of the CIII2 assembly intermediate supports the proposal21 that a full CIV docks on an assembly intermediate of CIII before the ISP subunit attaches. In the protomer facing CIV of the intermediate class, SCAF1 occupies the position of the ISP N terminus, thus preventing it from getting attached to the surface of CIII (Extended Data Fig. 6c). Therefore, it is possible that only one copy of an ISP precursor is processed at this stage, resulting in a visible ISP N terminus (residues 1–15) in the opposite CIII protomer and a single Sub9 bound inside MPP. In the assembled locked class, SCAF1 moves approximately 10 Å to its final position and the N terminus of ISP takes its rightful place. Therefore, the assembled locked class features all of the fully folded CIII2 subunits and then, finally, CIV moves to the unlocked conformation, where SCAF1 is no longer involved in the interface but still links the two complexes (Figs. 1b, c, 3a). SCAF1 is present in CIII2 and CIV from all classes, showing that it is not only an assembly factor but also a stable component of the supercomplex.
a, Sequence of the assembly events as observed in our structures (top) as opposed to the assembly of CI-containing supercomplexes (bottom): CIII2 is depicted in yellow, SCAF1 in blue, CIV in cyan and CIII2 subunits Sub10, Sub11 and ISP in red. Subunit COX7A of CIV, present in the respirasome instead of SCAF1, is depicted in light blue. b, Putative cyt c (black) positions in CIII2CIV locked (top) and unlocked (bottom) conformations, obtained by superposition of the crystal structures. The distance between the two cyt c-binding sites in the respective classes is indicated. c, Closed-up view of the putative cyt c position in the locked class: Sub8 of CIII2 (yellow) would clash with cyt c (black). d, Surface charges (calculated with APBS in PyMOL) of the unlocked class and cyt c. On the top, the arrows indicate the binding surfaces of cyt c on CIII and CIV. Cyt c is flipped 180o, to show the charge of its interacting surface. At the bottom, cyt c (black) is superimposed as in b, and the arrows indicate the location of the quinone (Q)-binding sites, showing that both are fully accessible to the membrane.
CIII analysis of CI-containing supercomplexes revealed the absence of any N-terminal SCAF1 density in CICIII2, and we saw that CIII2 from the respirasome also did not feature any such density, neither in the mouse sample, nor in the ovine, where we could distinguish the tight and the loose conformations of the respirasome (Fig. 2d, Extended Data Figs. 3, 5, 6a). This puts an end to the long-standing debate over the involvement of SCAF1 in the formation of the respirasome: the absence of the SCAF1 N terminus inside CIII2 from respirasomes clearly shows that SCAF1 is not required (Supplementary Information). In addition, we have refined CIV from the ovine respirasome to a sufficient resolution (4.1 Å) to show that, in CIV in the heart, the position of SCAF1 is occupied by COX7A1 (Extended Data Figs. 5b, 7b), which is the heart-specific and skeletal muscle-specific homologue of COX7A. By contrast, COX7A2 is ubiquitously expressed and was found in the human kidney-derived structure of the respirasome22.
In contrast to previous X-ray models of CIII2, containing a copy of the Sub9 model in each of the two MPP cavities, our maps revealed that only one copy of Sub9 is present. Unexpectedly, Sub9 straddles both cavities—the N terminus (residues 1–36) shares the cavity with SCAF1, whereas its C terminus is packed into the opposite cavity (Extended Data Fig. 6b). The density for the entire Sub9 is clear and could be resolved only because the attachment of a single copy of CIV makes the entire supercomplex asymmetric for structural analysis (Fig. 2c). This explains why in crystal structures, which lack this asymmetry, the density for Sub9 was weak and fragmented, due to averaging of both orientations23. Sub9 N-terminal and C-terminal parts form continuous β-sheets with the walls of the Core2 subunits. The N terminus of SCAF1 occupies the bulk of the cavity, folding into a two-stranded β-sheet, followed by a short helix and a loop going out of the MPP mouth (Extended Data Figs. 6c, 7a). SCAF1 interacts mostly with Core1, and surface charge analysis indicates that the negatively charged MPP cavity matches the positively charged surface of SCAF1, suggesting that SCAF1 is driven into the cavity mostly by electrostatic interactions.
We found this arrangement of Sub9 not only in CIII2CIV but also in CICIII2 (Fig. 2c, d, Extended Data Fig. 6a, Supplmentary Information), without any SCAF1 density, confirming that CIII2 is always asymmetric due to the presence of only one copy of Sub9 per dimer, either with an inserted SCAF1 N terminus (CIII2CIV) or without (CICIII2 or respirasome).
There is enough space only for one copy of SCAF1 to enter the MPP cavity containing the N terminus of Sub9, while a second SCAF1 would not be able to enter the other MPP cavity due to heavy steric clashes with the C terminus of Sub9. The MPP cavity is fully occupied by Sub9 and SCAF1 throughout all of our CIII2CIV classes, suggesting that there is no rearrangement of the cavity upon insertion of ISP. Thus, the asymmetry of the CIII2 dimer created by Sub9 clearly explains why only CIII2CIV is observed in mammals, as opposed to CIII2CIV2 in S. cerevisiae, where CIII2 lacks Sub9 and MPP activity24. Our data support doubts about the activity of MPP in mammalian CIII2 (Supplementary Information).
Our structures reveal the detailed mechanism of CIII2CIV assembly (Fig. 3a): SCAF1 drives the formation of CIII2CIV by mediating the docking of CIV onto an assembly intermediate of CIII2. The N terminus of SCAF1 enters the MPP cavity of CIII2, while its C-terminal transmembrane helix contacts Sub7 and Sub8 of CIII2, leaving the rest of the helix available for incorporation into CIV. After docking of CIV, CIII2 maturation proceeds with the folding of ISP, Sub10 and Sub11, forming the assembled locked conformation. At this point, CIV shifts to the unlocked class, where two of the newly added subunits of CIII2 (ISP and Sub10) form contacts with CIV. This transition is possibly initiated by the initial activity of CIII2, which requires the enzyme to ‘breathe’ (Supplementary Video 2), dislodging CIV.
The question stemming from our findings is the specific metabolic role of the two assembled conformations: given that the intermediate class lacks the integrated ISP subunit, this conformation must be catalytically inactive. In our mouse data, the locked assembled and the unlocked mature classes seem to be equally represented, whereas in the ovine data, the unlocked class is predominant. The mouse sample represents ‘fresher’ material; it is therefore conceivable that, similarly to the loose respirasome7, the unlocked class accumulates over time.
The supercomplex may provide a kinetic advantage by shortening the distance between CIII2 and CIV, compared to the 3D diffusion of cytochrome (cyt) c between individual complexes randomly distributed in the membrane25,26. Previously discussed protein-formed ‘substrate channels’27,28, which would facilitate direct transfer of quinone or cyt c within supercomplexes, are not observed in either respirasomes4,7 or CIII2CIV11,12,13, including our structures (Fig 1). When bound to CIV (as inferred from Protein Data Bank ID 5IY5), in the locked class, cyt c clashes with Sub8 of CIII (Fig. 3b, c), whereas there are no such clashes in the unlocked class. The distance between the probable cyt c-binding sites on CIII2 (inferred from Protein Data Bank ID 3CX5) and CIV in the unlocked class (8.5 nm; Fig 3b) is comparable to that in yeast (6.1 nm)11 and plant (7 nm) CIII2CIV1-2 (ref. 13). CIII2 cyt c1 and CIV CUA are separated by 6 nm in the locked class and 8.5 nm in the unlocked class (Extended Data Fig. 7f), which is too far for direct electron tunnelling. Therefore, the electron transfer between the complexes can only be mediated by the mobile cyt c.
The locked class is likely to be inactive due to cyt c–Sub8 clash, and only the unlocked class would represent the fully active, thus mature, CIII2CIV. This explains why a transition from the assembled locked to the mature unlocked class is required. In yeast, the positively charged cyt c can slide along a negatively charged patch on the surface of CIII2 and CIV without being released into the bulk solvent, thus further speeding up the reaction by 2D diffusion29. This charge distribution is conserved in our structures (Fig. 3d), suggesting that a similar mechanism may be present. In cyt c-mediated QH2:O2 oxidoreductase activity assays, we found that at saturating cyt c concentrations both CIII2 and CIV are about 1.7 and 1.9 times, respectively, more active within the supercomplex than as individual enzymes (Extended Data Fig. 7e). This is a larger advantage than in yeast29 and could be due to effects of the presence of SCAF1 in both enzymes, which remain to be investigated. At intermediate cyt c concentrations (1 μM), the supercomplex advantage increased to a 2.5-fold higher rate than that catalysed by two mixed individual complexes. Conversely, at very low cyt c concentrations (5–10 nM), there was barely any activity. This is in contrast to yeast where CIII2CIV1-2 was active under such conditions and has been suggested to be using just one cyt c bound per supercomplex29. Together with the large advantage at intermediate cyt c concentrations, this indicates that in mammals, the 2D diffusion mechanism is still operational but probably with two or more cyt c bound per supercomplex29. The reasons for this difference with yeast could be related to the opposite orientation of CIV versus CIII2. In conclusion, our structural and functional data point towards an evolutionarily conserved kinetic advantage of CIII2CIV over the individual complexes.
CIII2CIV would also have some kinetic advantage, in terms of CIII/CIV reactions, over the respirasome where the distance between cyt c sites is about 10 nm and one of the quinone-binding sites in CIII2 is partially shielded by CIV7, while both quinone sites are fully exposed in CIII2CIV (Fig. 3d). This would mean that kinetically it might be advantageous for mitochondria to use or express more CIII2CIV when substrate availability favours CII activity over CI28. Such a kinetic fine tuning of oxidative phosphorylation activity is possible without the proposed existence of separate quinone pools2,28, which is inconsistent with functional experiments4,30. The relative amount of CIII2CIV is probably regulated by the availability of SCAF1: insertion of its N terminus into the MPP cavity of the CIII assembly intermediate would make it destined for CIII2CIV, while SCAF1-free CIII2 can assemble into other supercomplexes.
Methods
Animals
Ovine hearts from which the supercomplexes were purified were obtained from a local abattoir, hence no ethics approval was required. For mouse samples, animal treatments were performed in accordance with the Austrian ‘Tierversuchs-Verordnung 2012 -TVV2012 (522. Verordnung)- Methoden our Totung von Tieren’, corresponding to the Annex IV (Methods of Killing Animals) of the European directive 2010/63/EU. Without any previous procedure, animals were killed by cervical dislocation and hearts were removed (procedure referred to as ‘organ removal’). According to the laws in Austria, no further approval by the Federal Ministry of Austria for Education, Science and Research was needed. The mice were housed within the breeding facility of the IST Austria (Am Campus 1, 3400 Klosterneuburg). The facility has been approved by the Authorities (Austrian Federal Ministry of Science, Research and Economy) under the following licence number: BMWFW-66.018/003-II/3b/2014. The mice were housed in commercially available individually ventilated cages made of polysulfon under standard laboratory conditions (room temperature 21–24 °C; relative humidity 55 ± 10%; photoperiod 12-h light:12-h dark) and supplied with standard diet and water ad libitum.
Sample purification
The sample was kept on ice for the whole procedure. Mitochondria from sheep hearts were purified as previously described31. The same procedure was adapted to the mouse hearts: CD-1 mice were killed by cervical dislocation and their hearts were excised and placed singularly on ice in pre-cooled tubes. The organs were minced with scissors in 1.4 ml buffer A (10 mM Tris-HCl pH 7.8, 250 mM sucrose and 5 mM β-mercaptoethanol) per gram of tissue, then the minced tissue was placed in a glass homogenizer equipped with a Teflon pestle, in 1.6 ml buffer B (10 mM Tris-HCl pH 7.8, 1 mM Tris-succinate pH 7.8, 250 mM sucrose, 5 mM β-mercaptoethanol and 0.2 mM EDTA) per gram of tissue. The sample was homogenized until no solid tissue was visible, then another 1.6 ml buffer B per gram of tissue was added and the sample was homogenized for 30 strokes. The sample was then placed in a tube and the homogenizer was rinsed with another 1.6 ml buffer B per gram of tissue, to collect the leftover tissue. The heavy cell debris was removed by 5 mins of centrifugation at 1,000g, then the supernatant was divided into 2-ml aliquots and the mitochondria were spun down for 10 min at 9,000g. The mitochondrial pellet was washed with 1 ml buffer B per tube, spun down for 2 min at 15,000g and resuspended again in 0.5 ml buffer B per tube, thereby allowing for the pooling of two tubes per washing step in a total volume of 1 ml per tube until the whole sample was pelleted into one tube and washed one last time in 0.5 ml buffer B. The mitochondrial pellet was then treated in the same way for ovine and mouse material: intact mitochondria were lysed by osmotic shock in water using a homogenizer and membranes were washed in buffer MM (20 mM HEPES pH 7.7, 50 mM NaCl, 10% glycerol, 1 mM EDTA, 2 mM DTT, 0.002% PMSF and cOmplete Ultra EDTA-free cocktail), as described in ref. 31. The washed mitochondrial membranes were then solubilized in 6:1 w/w digitonin-to-protein ratio for 1 h on a wheel in buffer M7 (20 mM HEPES pH 7.7, 50 mM NaCl, 10% glycerol, 1 mM EDTA, 2 mM DTT, 0.002% PMSF, cOmplete Ultra EDTA-free cocktail and 50 μg/ml cardiolipin, which was added only to the mouse preparation). After solubilization, the sample was spun down for 45 min at 21,000g and loaded on a MonoQ ion-exchange column, equilibrated in buffer QA (20 mM HEPES pH 7.7, 50 mM NaCl, 10% glycerol, 2 mM EDTA, 1 mM DTT, 0.2% digitonin and 10 μg/ml cardiolipin, which was added only to the mouse preparation). The separation of CIII-containing species from ATPase was achieved by applying buffer QB (20 mM HEPES pH 7.7, 1 M NaCl, 10% glycerol, 2 mM EDTA, 1 mM DTT, 0.2% digitonin and 10 μg/ml cardiolipin, which was added only to the mouse preparation) to obtain a linear gradient elution with NaCl (0.05–0.5 M in 20 ml, corresponding to 20 column volumes). The peak containing CIII (followed by A420 in comparison to A280) was then exchanged into amphipol A-35, at a final 1% (w/v) concentration for 30 min on a rotating wheel. To remove the detergent and dilute the excess salt, the sample was then diluted fivefold in SEC buffer (20 mM HEPES pH 7.7, 200 mM NaCl and 2 mM EDTA) and re-concentrated to 0.5 ml to be loaded on the SEC column. Gel filtration was performed twice sequentially on a Superose 6 10/300 column, to improve the purity of the sample. The final eluted fractions were concentrated to about A280 = 5 (approximately 4 mg/ml) and used for grid preparation. Extended Data Figure 1 shows chromatograms and native electrophoresis for the mouse and ovine preparations. For western blot, antibody 14742-1-AP from Proteintech was used to detect Core2, and PA5-96994 from Invitrogen was used to detect COX7A2L. Both antibodies were used at a 1:1,000 dilution. Goat polyclonal antibody anti-rabbit IgG (horseradish peroxidase conjugated; ab205718, Abcam) was used as a secondary antibody, at a 1:10,000 dilution.
Grid preparation
Quantifoil 0.6/1 300 mesh copper grids were manually coated with an approximately 1-nm layer of continuous carbon and glow discharged just before use for 5 s at about 25 mA for the mouse samples. The ovine samples were frozen on non-coated quantifoil 0.6/1 300 mesh copper grids, glow discharged for 2 min at about 25 mA. Of the sample (at 4 mg/ml for murine and 4.5 mg/ml for ovine), 2.7 μl containing 0.2% CHAPS to reduce protein denaturation at the air–water interface32 was applied to the grids in a humidified chamber (100% humidity at 4°) of a Vitrobot Mark IV, blotted for 5 s with 25 force and plunge-frozen in liquid ethane.
Data collection
The mouse CIII2CIV and CICIII2 dataset was collected on a Titan Krios microscope, whereas the ovine samples and the mouse respirasome were collected on a Glacios microscope, both equipped with a FalconIII camera. Micrographs were acquired in linear mode, at a nominal pixel size of 1.2 Å for Glacios and 1.09 Å for Titan microscopes. The total dose applied to the micrographs was 90 (Krios), 93 (Glacios mouse respirasome) 100 (Glacios ovine respirasome) and 92 (Glacios ovine CIII2CIV) e−/Å2, with 46 (Krios), 60 (Glacios mouse respirasome), 39 (Glacios ovine respirasome) and 69 (Glacios ovine CIII2CIV) frames per micrograph and a defocus range of −0.5 μm to −2 μm. After centring the beam in a hole and collecting a micrograph, the surrounding eight holes were imaged using beam-image-shift33, to improve the throughput. The details about the Krios data collection are listed in Extended Data Table 1.
Data processing
The processing pipelines for each dataset are illustrated in Extended Data Figs. 2–5. The micrographs were mainly processed using Relion/3.134. MotionCor235 was used for motion correction, CTFFIND436 for the initial estimation of CTF parameters, gctf37 for per-particle defocus estimation when continuous carbon grids were used and Topaz38 for accurate picking after preliminary processing. Local resolution was calculated using ResMap 1.1.4. The maps were refined both as overall consensus refinement of the entire supercomplex and by focusing on either CIII2 or CIV for each class, leading to the following gold-standard resolutions for the entire mouse supercomplex, focused CIII2 and focused CIV, respectively: unlocked was 3.3 Å, 3.1 Å and 3.5 Å; locked assembled was 3.2 Å, 3.1 Å and 6.7 Å; locked intermediate was 3.6 Å, 3.4 Å and 6.3 Å; locked combined (pooled intermediate and assembled) focused on CIV was 3.4 Å. For the ovine supercomplex, the corresponding values were 4.9 Å, 4.3 Å and 5.9 Å for unlocked and 9.2 Å (only globally refined) for locked (Extended Data Figs. 2–4). Structures of mouse CIII2CIV were refined into composite maps, formed by a combination of focus-refined maps for each class. To our knowledge, these are the first structures of mouse CIII2 and CIV, which may facilitate structure-based mutagenesis and functional analyses. The dataset-specific processing pipelines are described in the following sections.
Mouse CIII2CIV and CICIII2
For the mouse Krios dataset, after motion correction and CTF estimation, particles were picked with Laplacian of Gaussian (LoG) spatial filter in Relion and subjected to multiple rounds of 3D classification, to obtain good references for Topaz picking. Given the presence of multiple species in the sample, multireference 3D classification was performed at first, with CICIII2 (PDB 6QBX), CIII2CIV (PDB 1NTZ and 5IY5) and CIII2 (PDB 1NTZ) as starting models, filtered to 60 Å. Subsequent classic 3D classification then allowed us to separate the best particles. After Topaz picking with the best CIII2CIV or CICIII2 classes (to obtain two pools of particles, one for each supercomplex), per-particle defoci were estimated and one more round of 3D classification separated the particles of interest from remaining junk and contamination. Subsequently, 3D refinement, CTF refinement and polishing were applied to both the CIII2CIV and the CICIII2 pools. The two pools of particles were then treated differently. CIII2CIV particles were classified without alignment around CIII2 to separate the assembly intermediate from the assembled complex, then the two good classes were classified to identify the three final conformations (intermediate locked, assembled locked and mature unlocked, or simply unlocked). These maps were then globally refined and focus-refined around CIII2 and CIV. For the locked classes, intermediate and assembled were pooled to focus-refine CIV. The CICIII2 particles, instead, were focus-refined around CIII2, then classification without alignment was performed in the Sub9 region, with mask inside the MPP cavities formed by Core1 and Core2 subunits, to separate the two orientations. The particles belonging to each class were then reoriented, to have CIII2 in the centre of the box and oriented along the C2 symmetry axis, with a global refinement. This procedure allowed us to easily flip one of the two classes by 180o and then pool back the particles to focus-refine asymmetric CIII2.
Ovine CIII2CIV
Similar to the mouse Krios dataset, after motion correction and CTF estimation, particles were picked with LoG spatial filter in Relion and subjected to multiple rounds of 3D classification, first with CICIII2, CIII2CIV and CIII2 as starting models and then with CIII2CIV only, either in the locked or in the unlocked conformation. The unlocked class particles were pooled and refined, first globally and then focused on CIII2 and CIV. The final locked class only contained 5,000 particles, therefore it was only globally refined.
Ovine CICIII2CIV
Similar to the mouse Krios dataset, after motion correction and CTF estimation, particles were picked with LoG spatial filter in Relion and subjected to multiple rounds of 3D classification, first with CICIII2, CIII2CIV and CIII2 as starting models and then with CICIII2CIV only, with models filtered to 60 Å. The final pool of respirasome particles was globally refined, then focus-refined on the membrane arm of CI and subsequently classified without searches to separate the different conformations of the supercomplex components. The two best classes, out of 20, corresponding to the tight and loose conformations of the respirasome, were 3D-refined, CTF-refined, polished and focus-refined on CIII2. For the CIV map, instead, the best pool of CIV-containing particles, without distinguishing between tight and loose, were refined around the CI membrane arm and CIV to best resolve the COX7A isoform.
Mouse CICIII2CIV
Similar to the mouse Krios dataset, after motion correction and CTF estimation, particles were picked with LoG spatial filter in Relion. Multiple rounds of 3D classification separated CICIII2CIV from junk and CICIII2 contamination. The final pool of respirasome particles was globally refined, CTF-refined, polished and focus-refined on CIII2.
Model building
Initial models of each chain were obtained using mouse sequences and the online tool PHYRE239. The initial models were then docked on pre-existing CIII2 (PDB 1NTZ and 3L75) and CIV (PDB 5IY5 and 5Z62) models and merged in Coot40. The Sub9 and SCAF1 (chains T and I of CIII2, respectively (Extended Data Table 2)) were manually built in Coot. The mouse CIII2 and CIV structures are overall very similar to other known mammalian structures, such as bovine, with all of the cofactors similarly coordinated. The flexible ISP domains were docked as rigid bodies into their rather weak density and not re-built any further. The density for the loop connecting the N and C terminus of SCAF1 was absent due to flexibility in this area, but the loop was still built as polyalanine to facilitate the visualization of the likely connection.
Model refinement
Model refinement was performed in Phenix using the described protocol4. Initially, the best models for CIII2 and CIV were refined using the highest resolution maps available, that is, CIII2 from CICIII2 and CIV from the locked classes, and adjusted manually in Coot as needed. Then they were applied to the other maps and refined either individually or as supercomplexes. The composite maps for the refinement of CIII2CIV were obtained by first fitting in Chimera the CIII2 and CIV focus-refined maps (from a particular class) into the map from the overall refinement of that particular class (intermediate, assembled locked or unlocked). Focused maps were then carved within 5 Å of the PDB model for each complex and summed up in Chimera (using the ‘vop maximum’ command to avoid adding up densities at the interface between complexes) into a composite map for each class.
Details about the models (chain IDs, composition and completeness) are listed in Extended Data Table 2.
Chimera and PyMOL were used to prepare the figures.
Activity measurements
The activities of CIII2 and CIV were measured spectrophotometrically by following the oxidation of decylubiquinol (DQH2) at 278 nm (ε = 13.55 mM−1 cm−1)41 and of cyt c at 550 nm (ε = 18.5 mM−1 cm−1)42, respectively. Reduced substrates were prepared following established procedures41,43,44,45 with minor modifications. For DQH2, 200 μl of a 10 mM stock of decylubiquinone (SC358659, ChemCruz) was diluted in equal volume of 100 mM potassium phosphate pH 6.0 and reduced with 5 mg of sodium borohydride. The solution was vortexed until it became colourless, then 20 μl of 1.2 M HCl was added and the solution was vortexed again. Addition of sodium borohydride and acid was repeated and followed by spin down of the sample for 10 min at 10,000g. The reduced quinol in the supernatant was then extracted in three cycles using 0.8 ml of hexane and washed twice with 5 ml of 5 M NaCl acidified with 10 mM HCl. The final organic phase was evaporated under nitrogen stream and decylubiquinol was solubilized in acidic ethanol (containing 6 mM HCl). The concentration of reduced quinol was calculated from the absorbance at 278 nm, using an extinction coefficient of 3.1. Reduced cyt c was prepared from a 6 mM stock of oxidized cyt c (C2037-1G, Sigma-Aldrich) in 50 mM potassium phosphate buffer pH 7.0, reduced by addition of 0.2 mg of sodium dithionite. The reducing agent was removed using a PD Minitrap G-25 (GE Healthcare) desalting column. The assays were carried out at 30 oC, under constant stirring, in a UV-transparent quartz cuvette, using a Shimadzu UV-2600 UV-VIS spectrophotometer. The activity was measured with 2–10 nM protein sample in assay buffer, containing 20 mM HEPES pH 7.4, 0.1% LMNG, 0.2% CHAPS, 0.25 mg/ml of 4:1 DOPC:CL lipid mixture, 100 μM DQH2 and 5 μM rotenone (R8875-1G, Sigma-Aldrich), to inhibit any contaminating CI. DQH2 and rotenone were freshly added to each reaction mixture. The samples were incubated for 2–5 min before starting the reaction by addition of cyt c. Given the scarcity of the sample, due to the limiting amount of tissue as starting material (mouse hearts), the measurements were performed as follows: after pre-incubation of the sample in assay buffer, 1 μM oxidized cyt c was added and DQH2 oxidation was followed at 278 nm for 30 s, after which 50 μM oxidized cyt c was added and the reaction was followed for 60 s. At this point, a mixture of 25 μM antimycin A (A8674-50MG, Sigma-Aldrich) and fenamidone (33965-100MG-R, Sigma-Aldrich) was used to inhibit CIII2 and CIV activity was monitored at 550 nm upon addition of 50 μM reduced cyt c for 60 s. Finally, 1 mM KCN (60178-52G, Sigma Aldrich) was used to stop the reaction. A sample without protein was used to subtract the background due to non-enzymatic reactions. Experiments with isolated CIII2 and CIV (mixed in equal amounts) were performed under identical conditions using CIII2 and CIV-containing fractions from mouse supercomplex preparations. Each measurement was repeated three times, except for the 5–10 nM cyt c measurements, which were repeated two times. The amount of protein in each sample was estimated using the Pierce 660 nm Protein Assay Reagent (22660, Thermo Scientific), and for the CIII2CIV supercomplex sample, the contribution from the small amount of CI-containing supercomplexes was estimated using the NADH:FeCy activity, as previously described4, with pure CI at known concentrations as reference.
Reporting summary
Further information on research design is available in the Nature Research Reporting Summary linked to this paper.
Data availability
Structures of the CIII2CIV intermediate, assembled locked, mature unlocked and CIII2 from CICIII2 were deposited in the PDB (access IDs 7O3E, 7O37, 7O3C and 7O3H, respectively) and corresponding cryo-electron microscopy density maps in the EMDB (IDs 12705, 12702, 12703 and 12706, respectively). Any other relevant data and information are available from the corresponding author on request. The following models, deposited in the PDB (https://www.rcsb.org), have been cited throughout the paper: 1NTZ, 3CX5, 3L75, 5IY5, 5J4Z, 5Z62, 6HWH, 6HU9, 6QBX and 7JRP.
References
Schägger, H. & Pfeiffer, K. Supercomplexes in the respiratory chains of yeast and mammalian mitochondria. EMBO J. 19, 1777–1783 (2000).
Calvo, E. et al. Functional role of respiratory supercomplexes in mice: SCAF1 relevance and segmentation of the Qpool. Sci. Adv. 6, eaba7509 (2020).
Lobo-Jarne, T. et al. Human COX7A2L regulates complex III biogenesis and promotes supercomplex organization remodeling without affecting mitochondrial bioenergetics. Cell Rep. 25, 1786-1799.e4 (2018).
Letts, J. A., Fiedorczuk, K., Degliesposti, G., Skehel, M. & Sazanov, L. A. Structures of respiratory supercomplex I+III2 reveal functional and conformational crosstalk. Mol. Cell 75, 1131-1146.e6 (2019).
Lobo-Jarne, T. & Ugalde, C. Respiratory chain supercomplexes: structures, function and biogenesis. Semin. Cell Dev. Biol. 76, 179–190 (2018).
Davies, K. M., Blum, T. B. & Kühlbrandt, W. Conserved in situ arrangement of complex I and III2 in mitochondrial respiratory chain supercomplexes of mammals, yeast, and plants. Proc. Natl Acad. Sci. USA 115, 3024–3029 (2018).
Letts, J. A., Fiedorczuk, K. & Sazanov, L. A. The architecture of respiratory supercomplexes. Nature 537, 644–648 (2016).
Sousa, J. S., Mills, D. J., Vonck, J. & Kühlbrandt, W. Functional asymmetry and electron flow in the bovine respirasome. eLife 5, e21290 (2016).
Gu, J. et al. The architecture of the mammalian respirasome. Nature 537, 639–643 (2016).
Guo, R., Zong, S., Wu, M., Gu, J. & Yang, M. Architecture of human mitochondrial respiratory megacomplex I 2 III 2 IV 2. Cell 170, 1247–1257 (2017).
Hartley, A. M. et al. Structure of yeast cytochrome c oxidase in a supercomplex with cytochrome bc 1. Nat. Struct. Mol. Biol. 26, 78–83 (2019).
Rathore, S. et al. Cryo-EM structure of the yeast respiratory supercomplex. Nat. Struct. Mol. Biol. 26, 50–57 (2019).
Maldonado, M., Guo, F. & Letts, J. A. Atomic structures of respiratory complex III2, complex IV, and supercomplex III2-IV from vascular plants. eLife 10, e62047 (2021).
Cogliati, S. et al. Mechanism of super-assembly of respiratory complexes III and IV. Nature 539, 579–582 (2016).
Spikes, T. E., Montgomery, M. G. & Walker, J. E. Structure of the dimeric ATP synthase from bovine mitochondria. Proc. Natl Acad. Sci. USA 117, 23519–23526 (2020).
Pinke, G., Zhou, L. & Sazanov, L. A. Cryo-EM structure of the entire mammalian F-type ATP synthase. Nat. Struct. Mol. Biol. 27, 1077–1085 (2020).
Wiseman, B. et al. Structure of a functional obligate complex III2IV2 respiratory supercomplex from Mycobacterium smegmatis. Nat. Struct. Mol. Biol. 25, 1128–1136 (2018).
García‐Poyatos, C. et al. Scaf1 promotes respiratory supercomplexes and metabolic efficiency in zebrafish. EMBO Rep. 21, e50287 (2020).
Lopez-Fabuel, I. et al. Complex I assembly into supercomplexes determines differential mitochondrial ROS production in neurons and astrocytes. Proc. Natl Acad. Sci. USA 113, 13063–13068 (2016).
Xia, D. et al. Structural analysis of cytochrome bc1 complexes: implications to the mechanism of function. Biochim. Biophys. Acta 1827, 1278–1294 (2013).
Pérez-Pérez, R. et al. COX7A2L is a mitochondrial complex III binding protein that stabilizes the III2+IV supercomplex without affecting respirasome formation. Cell Rep. 16, 2387–2398 (2016).
Zong, S. et al. Structure of the intact 14-subunit human cytochrome c oxidase. Cell Res. 28, 1026–1034 (2018).
Zong, S. et al. UQCRFS1N assembles mitochondrial respiratory complex-III into an asymmetric 21-subunit dimer. Protein Cell 9, 586–591 (2018).
Hunte, C., Koepke, J., Lange, C., Roßmanith, T. & Michel, H. Structure at 2.3 Å resolution of the cytochrome bc1 complex from the yeast Saccharomyces cerevisiae co-crystallized with an antibody Fv fragment. Structure 8, 669–684 (2000).
Stuchebrukhov, A., Schäfer, J., Berg, J. & Brzezinski, P. Kinetic advantage of forming respiratory supercomplexes. Biochim. Biophys. Acta Bioenerg. 1861, 148193 (2020).
Berndtsson, J. et al. Respiratory supercomplexes enhance electron transport by decreasing cytochrome c diffusion distance. EMBO Rep. 21, e51015 (2020).
Blanchi, C., Genova, M. L., Castelli, G. P. & Lenaz, G. The mitochondrial respiratory chain is partially organized in a supercomplex assembly: kinetic evidence using flux control analysis. J. Biol. Chem. 279, 36562–36569 (2004).
Lapuente-Brun, E. et al. Supercomplex assembly determines electron flux in the mitochondrial electron transport chain. Science 340, 1567–1570 (2013).
Moe, A., Di Trani, J., Rubinstein, J. L. & Brzezinski, P. Cryo-EM structure and kinetics reveal electron transfer by 2D diffusion of cytochrome c in the yeast III–IV respiratory supercomplex. Proc. Natl Acad. Sci. USA 118, e2021157118 (2021).
Fedor, J. G. & Hirst, J. Mitochondrial supercomplexes do not enhance catalysis by quinone channeling. Cell Metab. 28, 525-531.e4 (2018).
Letts, J. A., Degliesposti, G., Fiedorczuk, K., Skehel, M. & Sazanov, L. A. Purification of ovine respiratory complex i results in a highly active and stable preparation. J. Biol. Chem. 291, 24657–24675 (2016).
Kampjut, D., Steiner, J. & Sazanov, L. A. Cryo-EM grid optimisation for membrane proteins. iScience 24, 102139 (2021).
Cheng, A. et al. High resolution single particle cryo-electron microscopy using beam-image shift. J. Struct. Biol. 204, 270–275 (2018).
Zivanov, J., Nakane, T. & Scheres, S. H. W. Estimation of high-order aberrations and anisotropic magnification from cryo-EM data sets in RELION-3.1. IUCrJ 7, 253–267 (2020).
Zheng, S. Q. et al. MotionCor2: anisotropic correction of beam-induced motion for improved cryo-electron microscopy. Nat. Methods 14, 331–332 (2017).
Rohou, A. & Grigorieff, N. CTFFIND4: fast and accurate defocus estimation from electron micrographs. J. Struct. Biol. 192, 216–221 (2015).
Zhang, K. Gctf: real-time CTF determination and correction. J. Struct. Biol. 193, 1–12 (2016).
Bepler, T. et al. Positive-unlabeled convolutional neural networks for particle picking in cryo-electron micrographs. Nat. Methods 16, 1153–1160 (2019).
Kelley, L. A., Mezulis, S., Yates, C. M., Wass, M. N. & Sternberg, M. J. E. The Phyre2 web portal for protein modeling, prediction and analysis. Nat. Protoc. 10, 845–858 (2015).
Emsley, P., Lohkamp, B., Scott, W. G. & Cowtan, K. Features and development of Coot. Acta Crystallogr. D 66, 486–501 (2010).
Sherwood, S. & Hirst, J. Investigation of the mechanism of proton translocation by NADH:ubiquinone oxidoreductase (complex I) from bovine heart mitochondria: does the enzyme operate by a Q-cycle mechanism? Biochem. J. 400, 541–550 (2006).
Kubota, T., Yoshikawa, S. & Matsubara, H. Kinetic mechanism of beef heart ubiquinol: cytochrome c oxidoreductasel. J. Biochem. 111, 91–98 (1992).
Rieske, J. S. [44] Preparation and properties of reduced coenzyme Q-cytochrome c reductase (complex III of the respiratory chain). Methods Enzymol. 10, 239–245 (1967).
Trumpower, B. L. & Edwards, C. A. Purification of a reconstitutively active iron-sulfur protein (oxidation factor) from succinate•cytochrome c reductase complex of bovine heart mitochondria. J. Biol. Chem. 254, 8697–8706 (1979).
[No authors listed] Reduced cytochrome c solution. Cold Spring Harb. Protoc. 2015, pdb.rec084475 (2015).
Acknowledgements
We thank the pre-clinical facility of the IST Austria and A. Venturino for assistance with the animals; and V.-V. Hodirnau for assistance during the Titan Krios data collection, performed at the IST Austria. The data processing was performed at the IST high-performance computing cluster. This project has received funding from the European Union’s Horizon 2020 research and innovation program under the Marie Skłodowska-Curie grant agreement no. 754411.
Author information
Authors and Affiliations
Contributions
I.V. purified the sample, prepared cryo-electron microscopy grids, acquired and processed EM data, built and analysed the atomic models and wrote the initial draft of the manuscript. L.A.S. designed and supervised the project, analysed data and models and revised the manuscript. I.V. and L.A.S. acquired funding.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Nature thanks Werner Kühlbrandt and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Peer reviewer reports are available.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Sample preparation overview.
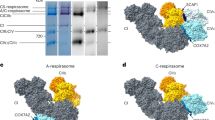
a, Purification of the mouse sample: on top, chromatograms of the MonoQ (left), the first SEC (middle) and the second SEC (right). The pooled fractions are highlighted in blue. At the bottom, native PAGE and western blot of the purification steps: from left to right, Coomassie-stained solubilised material (SM), then Coomassie staining, complex I in-gel activity assay, complex IV in-gel activity assay, western blot for Core2, western blot for SCAF1 of the purification steps. b, Purification of the ovine sample: the same steps as shown in a. For Gel source data, please see Supplementary Fig. 1. The mouse purification was repeated at least 5 times, the ovine preparation was repeated at least 10 times, all attempts of replication were successful, and one representative preparation per sample type is shown in the figure.
Extended Data Fig. 2 Processing pipeline for the mouse CIII2CIV dataset.
Scheme of the processing pipeline, with representative micrograph, pre-processing, classification steps and final maps, coloured by local resolution, with relative angular distribution plot.
Extended Data Fig. 3 Processing pipeline for the mouse CICIII2 data, FSC curves of the mouse maps and representative densities from the cryo-EM maps.
a, Processing pipeline as in Extended Data Fig. 2: the same set of micrographs was used, but the classification focused on CICIII2. b, Gold-standard half-map and model-map FSC curves for the mouse dataset. c–h, Representative densities of CIII2 (c–e) and CIV (f–i) for α-helices -contoured at 2.7σ- (c and f), β-sheets -contoured at 2.7σ- (d and g), cofactors -contoured at 1.9σ- (e and i) and lipids -contoured at 1.8σ- (h, from CIII2).
Extended Data Fig. 4 Processing pipeline and maps of the ovine CIII2CIV dataset.
a, Processing pipeline of the ovine CIII2CIV dataset, as in Extended Data Figs. 2, 3. b, Final maps of the ovine CIII2CIV locked (left, 2.8σ contour) and unlocked (right, 3σ contour for CIII and 6σ for CIV), shown from the side (top) and from the matrix (bottom). The respective mouse models, coloured as in Figs. 1, 3 , were superimposed on CIII2, to show the conservation of the conformations across mammalian species.
Extended Data Fig. 6 MPP cavity in different supercomplexes.
a, Cryo-EM map density (grey mesh) carved around Sub9 (orange cartoon) and SCAF1 (blue) positions in supercomplexes CIII2CIV, CICIII2 and CICIII2CIV. For CIII2CIV and the mouse CICIII2, the unique positions of a single copy each of Sub9 and SCAF1 were used, while for respirasomes, where the limited resolution prevented Sub9-focused classification and so CIII2 adopts a random orientation, two-fold symmetry-related positions of these proteins were also added. Contour levels are 2, 1.9 and 2.1σ for mouse CIII2CIV mature unlocked, assembled locked and intermediate locked respectively; 8σ for ovine CIII2CIV mature unlocked; 2.9σ for mouse CICIII2; 6.7σ for mouse CICIII2CIV; 6.7 and 5.3σ for ovine CICIII2CIV tight and loose respectively. b, Sub9 (orange) integration into the MPP walls: UQCRC1 (Core1) and UQCRC2 (Core2) are shown in grey (Core1 light and Core2 dark), except for the strands of Core2 forming continuous sheets with Sub9, which are shown in yellow. c, SCAF1 (blue), Sub7 (yellow) and ISP (red) arrangement in the intermediate (left) and assembled locked (right) classes. The rest of CIII is depicted in grey. The density for Sub7 is carved in grey, to show the ordering of the loop in the assembled versus intermediate class. In the insets at the bottom, the densities for SCAF1 and ISP are shown in the two classes. 2.9σ contour for CIII2CIV intermediate and 2.3σ for assembled.
Extended Data Fig. 7 SCAF1 in CIII and CIV and functional implications.
a, Surface charge representation (as in Fig. 3) of Core1 and SCAF1 in the unlocked class (full model is grey with a sketched eye indicating the perspective). SCAF1 cartoon is green. A white asterisk indicates the MPP "mouth". b, SCAF1 (cyan) and COX7A1 (blue) models fitted to the locked (left, 2.5σ contour) and unlocked class (right, 2.7σ contour) CIV maps. The arrows in b and c indicate bulky residues used to identify the subunit based on the map density. SCAF1 model, but not COX7A1, fits the density in both classes. c, Ovine SCAF1 (cyan) and COX7A1 (blue) models fitted to CIV map from respirasome (7.3σ contour): contrary to CIII2CIV, COX7A1 model fits the density, unlike SCAF1. d, Sequence alignment of mammalian COX7A isoforms. The SCAF1 portions interacting with CIII2 are indicated above the alignment. The black arrows indicate the positions highlighted in b and c. e, Activity measurements (described in the Methods) of CIII2+CIV (light grey) and supercomplex CIII2CIV (dark grey). The electron transfer rates (in e-/s) are calculated from the CIII2-driven oxidation of DQH2 or from the CIV-driven oxidation of reduced cyt c, as indicated. The bars represent the mean, the dots depict the single measurements. The 5-10 nM ox cyt c represents 2.5-5:1 cyt c/CIII2CIV ratio (two independent experiments). All other measurements come from 3 independent experiments (two values in the CIII2+CIV sample with 50 μM red cyt c overlap, therefore only two dots are visible). f, Cofactor distances within and between CIII2 and CIV, in the unlocked conformation. Cyt c is bound to CIII2 and CIV as in Fig. 3b. The transparent cartoon represents the protein subunits, the cofactors are sticks (hemes) and spheres (FeS clusters and Cu centers), in orange for CIII2, cyan for CIV and black for cyt c. The dashed lines represent the IMM.
Extended Data Fig. 8 Structural differences among supercomplexes and comparison between CIII2 movements in CICIII2 and intermediate vs assembled CIII2CIV.
a, Top left, CIII2CIV and respirasome (ovine, PDB 5j4z) aligned by CIII2: CI of respirasome is grey, CIV of respirasome is blue, CIII2 is yellow and CIV of CIII2CIV is cyan (the colour scheme for mammalian CIII2CIV is consistent throughout the panel). CIII2CIV locked is shown on the left, unlocked on the right. The two possible orientations of CIII2CIV based on CIII2 are shown at the top and at the bottom. Top right, plant CIII2CIV (PDB 7jrp) and mammalian CIII2CIV aligned by CIII2 in the locked (top) and unlocked (bottom) conformations. The plant CIV is blue. Bottom left, yeast CIII2CIV2 (PDB 6hu9) and mammalian CIII2CIV aligned by CIII2 in the locked (left) and unlocked (right) conformations. The yeast CIV is blue. Bottom right, M. smegmatis CIII2CIV2 (PDB 6hwh) and mammalian CIII2CIV aligned by CIII2 in the locked (left) and unlocked (right) conformations. The bacterial CIV is blue. The straight arrows indicate the shift of CIV towards mammalian CIV, and the curved arrows indicate the 90 or 180o flip of CIV around its vertical axis. b, Side (top) and IMS (bottom) views of the extreme frames of CIII2 3DVA mode 0 (open on the left and closed in the middle, 5σ contour). Ovals indicate the position of the ISP Fe-S domains, only visible in the closed state. On the right, overlay of the two extremes, with closed in dark grey and open in light grey. Arrows indicate the “breathing” movement of the TM and IMS domains. c, Same as b, but with CIII2 intermediate (left) and assembled (right) models from CIII2CIV fitted to the densities and coloured as in Figs. 1, 3.
Supplementary information
Supplementary Information
This file contains additional discussion relevant for the main text.
41586_2021_3927_MOESM4_ESM.jpg
Supplementary Fig. 1 Uncropped gels. All the original, uncropped scans of the native gels and western blots shown in Extended Data Fig. 1 are shown.
41586_2021_3927_MOESM5_ESM.mov
Supplementary Video 1 Overview of the structural features of CIII2CIV.The mature unlocked state map and model are shown, followed by a transition through the three different conformations, viewed from the side and from the top. Finally, the cytochrome c binding sites, inferred from the crystal structures cited in the manuscript, are shown. The colour scheme is the same as in the rest of the manuscript.
41586_2021_3927_MOESM6_ESM.mov
Supplementary Video 2 “Breathing” movement of CIII2 in CICIII2. The result of the 3DVA analysis on CIII2 is shown, viewed from the side first, then from the IMS. The fluctuating movements between a closed state, with the two ISP C-terminal domains visible, and an open state, with no density corresponding to ISP, are depicted.
Rights and permissions
About this article
Cite this article
Vercellino, I., Sazanov, L.A. Structure and assembly of the mammalian mitochondrial supercomplex CIII2CIV. Nature 598, 364–367 (2021). https://doi.org/10.1038/s41586-021-03927-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-021-03927-z
- Springer Nature Limited
This article is cited by
-
SCAF1 drives the compositional diversity of mammalian respirasomes
Nature Structural & Molecular Biology (2024)
-
Detailed analysis of Mdivi-1 effects on complex I and respiratory supercomplex assembly
Scientific Reports (2024)
-
High-resolution in situ structures of mammalian respiratory supercomplexes
Nature (2024)
-
Euglena’s atypical respiratory chain adapts to the discoidal cristae and flexible metabolism
Nature Communications (2024)
-
Electron transport system supercomplexes affect reactive-oxygen species production and respiration in both a hibernator (Ictidomys tridecemlineatus) and a nonhibernator (Rattus norvegicus)
Journal of Comparative Physiology B (2024)