Abstract
Meningiomas are the most common primary intracranial tumour in adults1. Patients with symptoms are generally treated with surgery as there are no effective medical therapies. The World Health Organization histopathological grade of the tumour and the extent of resection at surgery (Simpson grade) are associated with the recurrence of disease; however, they do not accurately reflect the clinical behaviour of all meningiomas2. Molecular classifications of meningioma that reliably reflect tumour behaviour and inform on therapies are required. Here we introduce four consensus molecular groups of meningioma by combining DNA somatic copy-number aberrations, DNA somatic point mutations, DNA methylation and messenger RNA abundance in a unified analysis. These molecular groups more accurately predicted clinical outcomes compared with existing classification schemes. Each molecular group showed distinctive and prototypical biology (immunogenic, benign NF2 wild-type, hypermetabolic and proliferative) that informed therapeutic options. Proteogenomic characterization reinforced the robustness of the newly defined molecular groups and uncovered highly abundant and group-specific protein targets that we validated using immunohistochemistry. Single-cell RNA sequencing revealed inter-individual variations in meningioma as well as variations in intrinsic expression programs in neoplastic cells that mirrored the biology of the molecular groups identified.
Similar content being viewed by others
Main
Although previous studies on meningioma have provided important insights into the possibility for molecular data to refine meningioma classification3,4,5,6,7,8, the formal integration of multiple molecular datatypes in a unified analysis has not been performed. Here we assembled a large cohort of meningiomas that were enriched for the uncommon, higher-grade tumours with matched multidimensional molecular and high-quality clinical data. We generated matched DNA somatic copy number, DNA point mutation, DNA methylation, transcriptomic and proteomic data to create a resource—similar to The Cancer Genome Atlas—for meningiomas that we further supplemented with single-cell RNA sequencing data. By integrating multiple datatypes in a unified analysis—as has been achieved for other cancers9,10,11,12—we define a molecular taxonomy for meningiomas that has direct clinical relevance.
Patient samples and clinical data
We used meningioma samples from 121 patients to define molecular groups, and 80 samples from an independent cohort to assess generalizability. Samples were selected on the basis of availability of clinical data as well as the quality and quantity of tissue for analyses. Our cohort reflects the real-life diversity of patients with meningiomas and includes a substantial number of WHO (World Health Organization) grade 2 and 3 meningiomas, which have been understudied to date because of their rarity. We performed whole-exome sequencing for germline polymorphisms, somatic point mutations and somatic copy-number alterations; EPIC array profiling for DNA methylome analysis; and mRNA sequencing for transcriptome analysis on all 121 tumours in the discovery cohort. Whole-cell proteomics was performed on 96 of these tumours (Fig. 1a). DNA methylation was also performed on five healthy meninges samples for methylome comparison. Eight tumours and two healthy meninges samples were profiled by single-nucleus RNA sequencing to examine intratumoral heterogeneity. Grading was confirmed by two independent neuropathologists in accordance with the most recent 2016 WHO classification criteria. All samples were annotated with detailed high-quality clinical data elements that were established a priori (see Methods and Supplementary Table 1).
a, Flow diagram showing the relationship between molecular datasets on tumours in this study: whole-exome sequencing (seq), DNA methylation and mRNA sequencing (RNA-seq) (n = 124), proteomics (n = 96), and single-cell data (n = 8). A total of 121 samples were used for discovery on bulk analyses, with an additional 3 samples assembled specifically for single cell analyses. b, t-distributed stochastic neighbor embedding (t-SNE) reduction of individual platform data with annotated unsupervised cluster assignments for each individual platform. c, Alluvial plot showing the relationships between unsupervised cluster assignments from individual platform analyses using DNA methylation, RNA sequencing and copy-number data. The widths of the nodes and edges are proportional to the number of samples. d, Multiplatform higher-order integration of genetic, epigenetic and transcriptomic data by cluster-of-cluster assignments. Cluster assignments for each independent platform (rows) are shown for each sample (columns). Membership for a given cluster is noted by a black tick. Annotation for clinical factors (WHO grade and the extent of resection (Simpson grade)) is shown above the matrix. e, Kaplan–Meier estimates of recurrence-free survival of patients according to molecular group. f, Brier prediction curves for recurrence-free survival comparing classification by molecular group to WHO grade, DNA methylation cluster assignments by the DKFZ (German Cancer Research Center)3, and cluster assignments by the individual datatypes in this study. The integrated Brier score is shown for each datatype.
Interdependencies of datatypes
To examine relationships between datatypes, we computed the Mutual Information (MI) metric for each gene between all pairwise combinations of datatypes and compared this to a permuted null distribution13. MI values of zero indicate orthogonal information. We found that the distribution of MI values was statistically significantly different between different datatype comparisons (Extended Data Fig. 1a). Moreover, consensus clustering of normalized MI values using genes where MI was significant for at least one datatype pair revealed four different gene clusters, each defined by distinct patterns of dependence between datatypes at different levels of the central dogma. These results show the potential value of formal unsupervised integration of multiple datatypes in meningioma.
Multiplatform integrative analyses
We next sought to combine whole-exome sequencing and copy number, DNA methylation, and mRNA sequencing data using cluster-of-cluster assignments (COCA)9,10,11,12. In this approach, cluster assignments from individual platform analyses are subjected to additional (second-order) clustering to examine the higher-order relationship between samples across molecular features.
Unsupervised sample-wise clustering of gene-level somatic copy-number alterations (CNAs), DNA methylome data and transcriptome data in isolation revealed six stable subgroups for each datatype with clinically relevant and significant differences in outcome (Fig. 1b, Extended Data Fig. 1b, d, f). Cluster assignments across datatypes were neither identical nor orthogonal (Fig. 1c) and cluster associations with outcome were unique for each datatype (Extended Data Fig. 1c, e, g).
COCA analysis combining six copy-number clusters with six DNA methylation and six mRNA abundance clusters converged to reveal four stable molecular groups (MG1–MG4) of meningioma (Fig. 1d, Extended Data Fig. 1h). RNA cluster assignments were strongly associated with MG1, MG3 and MG4, whereas CNA and DNA methylation cluster assignments were most strongly associated with MG2, and the relative importance of these datatypes was confirmed by formal unsupervised integration of two datatypes at a time (Supplementary Table 2). Tumours spanning all WHO grades were represented in each molecular group, with the exception of MG1, which was composed of only WHO grade 1 and 2 tumours. Tumours of higher WHO grade were enriched in MG3 and MG4 (Fisher’s exact test, P = 5.49 × 10−7). Notably, a clear one-to-one relationship between molecular group and WHO grade was not evident (Extended Data Fig. 1i), which prompted us to examine the clinical relevance of these newly defined integrative molecular groups.
Clinical relevance of integrative molecular groups
Although the discovery of the four molecular groups in this study was agnostic to patient outcomes, these groups were characterized by distinct and divergent patterns of recurrence-free survival (Fig. 1f). Overall, patients with MG3 and MG4 tumours had statistically shorter times to recurrence (log-rank test, P = 5 × 10−15), with the most unfavorable outcomes for MG4 tumours. Classification by molecular groups was independently associated with recurrence-free survival as assessed by multivariable Cox regression, even after accounting for known prognostic clinical factors—including WHO grade, extent of surgical resection and receipt of adjuvant radiotherapy (see Supplementary Table 3). Significant differences in recurrence patterns persisted across molecular groups when tumours were analysed separately according to WHO grade (Extended Data Fig. 1j–l). For predicting time to recurrence, classification by molecular group was superior to WHO grade and previously described methylation-based classifications3 as well as classification by cluster assignments from each datatype individually (Fig. 1f). We confirmed the generalizability of molecular-grade classification and outcomes in an independent cohort using mRNA signatures (Extended Data Fig. 2). This framework provides a blueprint for future independent validation and ongoing assessment of generalizability.
Mutational profiles of molecular groups
We next examined the somatic point-mutational profiles of molecular groups. While NF2 was, predictably, the most frequently point-mutated gene, the prevalence of such mutations differed significantly across molecular groups without distinct positional bias (Fig. 2a, Extended Data Fig. 3a). Nearly all MG1 meningiomas had mutations in NF2, whereas such mutations were extremely rare in the MG2 tumours (88% compared with 9%; Fisher’s exact test, P = 5.9 × 10−8). Conversely, the previously described mutations in TRAF7, AKT1, KLF4 and POLR2A were exclusively identified in the MG2 tumours at frequencies of 25%, 13%, 13% and 6%, respectively (Fisher’s exact test, P = 1.20 × 10−8).
a, Oncoprint showing the recurrent somatic mutations identified in tumour samples in this study. New and recurrent mutations in epigenetic regulators and tumour suppressor genes are detected. The colours in the oncoprint represent different types of somatic alterations. The relative proportions of the six different possible base-pair substitutions across all variants in each sample are shown in the bottom of the panel. b, Network of distinguishing gene circuits for each molecular group by mRNA abundance. Nodes represent pathways and edges represent shared genes between pathways. Nodes coloured in red are upregulated pathways whereas those coloured in blue are downregulated pathways in each molecular group. Light purple edges represent the pathways that are targeted by vorinostat. The inset shows a boxplot comparing inferred immune cell infiltrates by ESTIMATE between molecular groups. c, Results of cell viability assay testing the efficacy of vorinostat and 5-azacytidine on patient-derived meningioma cell lines that recapitulate specific molecular groups. Cell lines aligned to MG4 are coloured orange, to MG1 are coloured red, and to MG3 are coloured green. *P < 0.05 (Student’s t-test). Treatment with vorinostat results in a greater than 50% reduction of cell viability in MG4 cell lines only, whereas other general treatments—such as 5-azacytidine—do not show any anti-tumour activity in meningiomas. d, Tumour volumes in intracranial MG4-xenografted mice measured by serial MRI measurements during treatment with vorinostat or control. *P < 0.05 (Student’s t-test). e, Kaplan–Meier overall survival distribution of mice with MG4-intracranial xenografts treated with vorinostat versus control. Statistical significance was assessed using a log-rank test.
We found previously unidentified, statistically significant, recurrent nonsynonymous somatic driver mutations in genes that are involved in chromatin modelling and epigenetic regulation (KDM6A, CHD2), as well as in tumour suppressor genes (PTEN; Supplementary Table 4). Recurrent inactivating mutations in additional chromatin modelling (CREBBP, q = 0.127) and tumour suppressor (FBXW7, q = 0.226; RB1, q = 0.250) genes were also identified as subthreshold hits (Supplementary Table 4). These mutations occurred at frequencies similar to those of other known meningioma driver genes (3–5%, Fig. 2a), and were collectively enriched in the aggressive phenotypes of meningioma, distinguishing MG3 and MG4 tumours from MG1 and MG2 tumours (Fisher’s exact test, P = 0.002). MG4 tumours had significantly greater mutational burden compared to MG1–MG3 tumours (P = 1.6 × 10−3, Kruskal–Wallis test; Extended Data Fig. 3b). The majority of point mutations in meningioma were clonal, with only a small subset seen as late-evolving drivers (Extended Data Fig. 3c–e). The specificity of different mutations for distinct molecular groups was particularly notable given that the generation of molecular groups was independent of point mutations.
Genomic disruptions across molecular groups
We next investigated the pattern of genome-wide CNAs across molecular groups (Extended Data Fig. 4a). MG1 tumours were relatively diploid with the exception of uniform loss of chromosome 22q, which—in combination with concurrent NF2 point mutations—results in biallelic NF2 inactivation. There were two subsets of MG2 tumour: one in which tumours were copy-number neutral but harboured mutations in TRAF7, AKT1, KLF4 or SMO; and the other in which tumours did not harbour mutations but had consistent polysomies of chromosomes 5, 12, 13, 17 and 20. MG3 and MG4 meningiomas were high-aneuploidy tumours with losses in chromosomes 22q (93% and 86%, respectively), 1p (77% and 89%), 6q (30% and 38%), 14 (47% and 35%) and 18 (19% and 38%). MG4 meningiomas also showed gain of chromosome 1q and a loss of chromosome 10, which were uncommon in MG3 meningiomas (34% versus 2% for chromosome 1q, P = 2.9 × 10−4, Fisher’s exact test; and 38% versus 14% for chromosome 10, P = 0.025, Fisher’s exact test). Some MG3 and MG4 tumours containing wild-type NF2 showed silencing of NF2 expression that was not associated with changes in methylation of the NF2 gene (Extended Data Fig. 4b, c). The degree of total genomic disruption, quantified as the percentage of the genome that was altered, was higher in MG3 (median 16.9%) and MG4 (median 19.5%) meningiomas compared with MG1 (median 3.5%) and MG2 (median 9.6%) tumours (P = 5.2 × 10−6, Kruskal–Wallis test). This was further supported by more frequent non-recurrent interchromosomal fusion events in MG3 and MG4 tumours compared to MG1 and MG2 meningiomas (Extended Data Fig. 4d, Supplementary Table 5). Taken together, these data point to an increase in genomic instability in MG3 and MG4 tumours, which have the most unfavorable outcomes.
Gene-expression networks of molecular groups
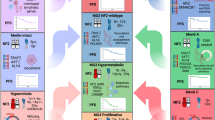
We next investigated the gene-expression pathways associated with each molecular group (Fig. 2b, Extended Data Fig. 5a). MG1 tumours showed greater immune infiltration and enrichment of pathways involved in immune regulation and signalling (Fig. 2b, inset, Extended Data Fig. 5b). By contrast, immune signatures were downregulated in MG4 tumours, and these tumours instead showed enrichment for pathways involved in cell-cycle regulation, as well as several critical and complementary proliferation-associated transcription factor networks (such as MYC, FOXM1, E2F) and protein complexes (for example mTORC1, CDKs, kinesins). MG3 tumours were uniquely enriched for pathways that converged onto the metabolism of several macromolecules. Although we identified two subsets of MG2 tumour by mutations and copy number, the transcriptomes of these subsets were distinctly correlated (Extended Data Fig. 5c, d), and collectively enriched for vascular and angiogenic pathways (Fig. 2b). Consequently, we designated the molecular groups as immunogenic (MG1), benign NF2 wild-type (MG2), hypermetabolic (MG3) and proliferative (MG4). It is notable that the association of molecular groups with outcomes was independent of molecular signatures of proliferation (Extended Data Fig. 5e, Supplementary Table 6).
We next sought to determine whether the distinct expression pathways could be exploited to identify new medical therapies for meningiomas, by mapping drugs approved by the United States Food and Drug Administration (FDA) to target genes in our enrichment network. We found that vorinostat, a histone deacetylase inhibitor, targeted several critical pathways that were specifically upregulated in proliferative (MG4) meningiomas (Fig. 2b). Treatment with vorinostat selectively decreased the viability of cell lines derived from patients with MG4 tumours only, and not cell lines derived from patients with tumours belonging to other molecular groups (Fig. 2c, Extended Data Fig. 6a, b). By contrast, treatment of the same cell lines with a comparable agent, 5-azacytidine, had no effect on cell viability. In mice with intracranial xenografts of patient-derived MG4 cell lines, treatment with vorinostat also attenuated tumour growth (Fig. 2d) and improved survival (Fig. 2e) compared with the control (Extended Data Fig. 6c, d). Overall, these findings suggest that tumours of different molecular groups might differ in their sensitivity to treatment with vorinostat, which warrants further investigation.
Proteogenomic characterization of molecular groups
Using a single-shot liquid chromatography–tandem mass spectrometry approach, we quantified a total of 6,568 unique protein groups in 96 tumours with somatic mutation, epigenome and transcriptome data in our cohort. Enrichment scores of gene sets by mRNA and proteome data were correlated well when comparing samples of similar molecular groups (Extended Data Fig. 7a–c). Functional inference using protein data alone converged on biological networks that were highly similar to those obtained by transcriptome data (Fig. 3a, Extended Data Fig. 7d). Specifically, immunogenic (MG1) tumours were enriched for proteins involved in immunoregulation, whereas hypermetabolic (MG3) meningiomas harboured enrichment of protein pathways converging on nucleotide and lipid metabolism, and proliferative (MG4) meningiomas were enriched for protein gene sets that regulate the cell cycle and cell proliferation.
a, Hierarchical clustering of genes from select pathways identified by gene set enrichment analysis (GSEA). Selected genes have been labelled. Gene annotation to pathway(s) is shown in the side bar. b, Scatter plot of hazard ratios of genes by gene expression (x axis) and protein abundance (y axis). Genes with significant associations with outcome are coloured in red. Selected genes are labelled. Pearson correlation and its associated 95% confidence interval are shown. c, Immunohistochemistry results for group-specific markers. Selected are four representative cases (rows). Images shown for each patient are at the same region of the slide for each antibody. Scale bars, 50 µm. Each case was subjected to unbiased digital quantitation. Below the images of representative stains are the receiver–operating characteristic curves for each antibody with the area under the curve, and 95% confidence interval with the DeLong approach.
We next compared the association of mRNA and protein abundance with outcomes. Overall, the associations of protein and gene abundance with outcome correlated well (Pearson’s ρ = 0.49, 95% confidence interval 0.47–0.50, P < 2.2 × 10−16). Concordance was 213 times more likely (odds ratio = 213.17, 95% confidence interval 113.74–422.26) than non-concordance amongst the 682 genes that were significantly associated with outcome by either mRNA or protein data (Fig. 3b). It is noteworthy that genes associated with poorer outcomes in both datatypes were involved in both the cell cycle (false discovery rate (FDR) = 3.98 × 10−7, hypergeometric test) and metabolism by oxidative phosphorylation (FDR = 2.9 × 10−55, hypergeometric test).
We then identified, using proteomic data, proteins that were highly enriched in each molecular group: S100B for MG1, SCGN for MG2, ACADL for MG3 and MCM2 for MG4 (Supplementary Table 7, see Methods). We validated the enrichment of these proteins in each group by immunohistochemistry in a blinded fashion. Unbiased, digital quantification of each protein marker showed strong concordance between immunohistochemistry and proteomic data, and protein markers were found to discriminate between molecular groups well (Fig. 3c). These results show potential for molecular group classifications to be adopted in conventional neuropathology laboratories, following further independent validation.
Methylation characteristics of molecular groups
We next searched for differences in genome-wide DNA methylation patterns between healthy meninges and meningiomas. We identified two sets of probes that differentiated healthy meninges from meningiomas as a whole (Extended Data Fig. 8a). In one set, probes were fully hypomethylated in healthy meninges and progressively gained methylation across molecular groups, whereas in the other set, probes were fully hypermethylated in healthy meninges, and progressively lost methylation across molecular groups. (Extended Data Fig. 8b). These patterns were similar when examining previously defined regions of the genome that either gain or lose methylation as a function of mitotic age14,15,16 (for example, epigenetic mitotic clocks, Extended Data Fig. 8c), pointing to the possibility that aberrant DNA methylation processes might be associated with the most aggressive molecular groups, although differences in cell type composition could also be a contributing factor.
We then identified transcription factors that were enriched in each molecular group on the basis of hypomethylated enhancer regions within each group (Extended Data Fig. 8d), known transcription-factor binding site motifs and correlations with gene expression17. Hypomethylation at enhancer regions was associated with transcription factors that aligned to the biology of each molecular group that we defined by gene and protein expression (Extended Data Fig. 8e, f).
Single-cell map of meningiomas
To investigate heterogeneity in meningiomas, we performed droplet-based single-nuclear RNA sequencing on eight tumours that were selected to span all molecular groups and WHO grades, as well as two healthy meninges samples for comparison.
In total, 54,393 high-quality and accurately genotyped single nuclei were analysed, and 14 distinct clusters were identified (Fig. 4a–d, Supplementary Figs. 1, 2). Cells were assigned to cell type on the basis of consensus between expression-based clustering (Extended Data Fig. 9a), inference of CNAs (Extended Data Fig. 9b, c) and annotation by canonical markers (Extended Data Fig. 10a). The majority of cells in our data were neoplastic (69%), whereas 14% were immune cells (macrophages and T cells), 10% were fibroblasts and 6% were endothelial cells.
a–d, Side-by-side-by-side-by-side t-distributed stochastic neighbor embedding (t-SNE) plots of 54,393 nuclei from ten samples, coloured by patient of origin (a), cluster number (b), cell type (c) and molecular group of tumour of origin (d). e, Stacked bar plot showing the distribution of cell-type fractions in bulk RNA sequencing data. Samples are grouped according to molecular group. Deconvolution was performed using CIBERSORTx. f, t-SNE plot showing cluster results for each tumour sample using all cells (left) and tumour cells only (right). Clustering was performed by both Seurat and DBSCAN. Cells that are coloured grey by the DBScan algorithm did not meet the parameters for clustering (see Methods). For clustering of all cells, the different colours represent the cluster colour scheme in b. For clustering of neoplastic cells, the different colours represent discrete tumour subpopulations. Annotation of patient sample and molecular group is shown to the left of the t-SNE plots. g, Hierarchical clustering of pairwise similarities between NMF programs on the basis of number of shared genes. Four groups of similar programs (meta-programs, black boxes) were identified. The NMF scores for each gene within modules are plotted below as a heat map. Select genes are labelled. Programs (column) are labelled as in the above heat map.
Non-neoplastic cells from different patients clustered together by cell type, whereas neoplastic cells clustered distinctly by patient, representing the inter-individual variability of meningiomas (Fig. 4a, Extended Data Fig. 10b, Supplementary Table 8). When neoplastic cells were considered in isolation, the variability between cells of different tumours was much larger than the variability within tumours (F = 65,538, P < 2.2 × 10−16, one-way ANOVA), and within the limits of differences in detection rates of genes between cells, the expression of neoplastic cells most closely resembled bulk molecular signatures of their tumour of origin (Extended Data Fig. 10c). Cycling neoplastic cells were enriched in MG3 and MG4 tumours (P = 2.2 × 10−2 and P = 1.49 × 10−2, respectively, mixed-effects) whereas immune cells were enriched in MG1 tumours (P = 1.8 × 10−2, mixed-effects; Extended Data Fig. 10d, e). Indeed, deconvolution of bulk RNA sequencing data using single-cell RNA sequencing signatures confirmed that macrophages were enriched in MG1 tumours, with additional differences in cell composition across molecular groups and healthy meninges (Fig. 4e, Extended Data Fig. 10f).
Heterogeneity by single cell
We first looked for discrete patterns of variation by clustering gene expression profiles of single cells from each sample individually using two independent clustering algorithms (Seurat and DBSCAN). When considering all cells within a sample, MG1–MG3 tumours showed several discrete clusters that were largely explained by the abundance of stromal or immune cell types, whereas MG4 tumours—which were predominantly composed of neoplastic cells—did not show distinct clusters (Fig. 4f). To examine the neoplastic component of each tumour more carefully, we then selected the neoplastic cells of each tumour for additional sub-clustering using the same algorithms. Again, using both algorithms we found that most samples harboured one dominant cluster, and less commonly a second minor cluster of neoplastic cells. Copy-number profiles of neoplastic cells were, in general, similar to those observed by bulk analyses and again did not show substantial variability between cells (Extended Data Fig. 9b, c). These findings were in line with our results from clonality assessment of bulk mutation data (Extended Data Fig. 3c–e), highlighting the relative rarity of subclonal expansion in meningiomas.
We then used non-negative matrix factorization to identify programs that were intrinsically expressed in neoplastic cells and shared between samples. In total, we identified 24 such programs across neoplastic cells of different samples that clustered to four ‘meta-programs’ on the basis of the degree of similarity by shared genes between modules (Fig. 4g, Extended Data Fig. 11a). The meta-programs were highly similar to the biology of the integrative molecular groups that we defined earlier, and the distributions of the activation of these programs across cells of different tumours reflected this (Extended Data Fig. 11b). The most prominent program was related to cell cycle (FDR = 3.13 × 10−32, hypergeometric test), and this program was reflective of discrete patterns of variability in most tumours (Extended Data Fig. 11b, c). Other programs included cellular metabolism (FDR = 7.66 × 10−3, hypergeometric test), inflammatory TNF signalling (FDR = 5.99 × 10−13, hypergeometric test) and a general mesenchymal program (FDR = 2.12 × 10−15, hypergeometric test), which generally showed more continuous patterns of variability (Extended Data Fig. 11c, d). Overall, these programs represent more subtle patterns of variation in meningiomas; however, the similarity of these programs—which are intrinsic to neoplastic cells—to the biology that we defined for the molecular groups introduced in this study points to the importance of these processes in meningioma biology. Indeed, deconvolution and partitioning of our bulk mRNA data using neoplastic and non-neoplastic signatures derived from our single-cell RNA sequencing data showed a high degree of similarity to the molecular groups that we define in this study (Extended Data Fig. 10g).
Conclusions
Here we present a resource for the meningioma community that contains matched multidimensional bulk and single-cell molecular and high-quality clinical data. By integrating multiple datatypes in a unified analysis, we define a molecular taxonomy for meningiomas (Extended Data Fig. 12) that could supersede existing molecular and clinically used classifications and has the potential to inform future iterations of recognized grading schemes.
Methods
Patient samples and clinical annotation
Clinical data was collected for each sample using pre-established common data elements (CDEs) designed for reporting on molecular studies of meningioma. Definitions for CDEs were agreed upon using a systematic process of discovery, internal validation, external validation and distribution. A total of 19 core CDEs (including age, sex, country of care, history of neurofibromatosis, history of malignancy, previous exposure to cranial radiation or chemotherapy, history of multiple meningiomas, timing of surgery, location of tumour, extent of resection at surgery, histopathological grade (WHO) and year of WHO classification system, recurrence status, time to recurrence from index surgery, previous irradiation to meningioma, time to last follow-up) were collected for all samples and an additional 14 supplemental CDEs (including race/ethnicity, hispanic race, diagnosis of meningioma syndrome, tumour size, Simpson grade, performance status at recurrence or last follow-up, second intervention for recurrence, time to second intervention, histopathological subtype of recurrent tumour, vital status, cause of death, time to death) were collected per sample, where possible. Collection of samples and clinical data was carried out in accordance with individual institutional ethics and review board guidelines.
For the present study focusing on integration of multiplatform molecular studies, tissue and blood samples were selected on the basis of sufficient availability of specimens (>500 mg tissue and >1 ml of blood or plasma). In total, 124 fresh-frozen meningioma samples and 5 healthy meninges samples from patients were collected for molecular analyses from the University Health Network Brain Tumour BioBank (Toronto) under the institutional Research Ethics Board. Samples were collected fresh from the patients at the time of surgical resection and immediately snap-frozen in liquid nitrogen and stored at −80 °C. Healthy meninges were collected from patients who underwent neurosurgery for non-oncological disease.
Clinical data was collected as per pre-established consensus definitions as indicated above. In brief, for each case, haematoxylin and eosin (H&E) slides were reviewed by two experienced neuropathologists independently to confirm the diagnosis of meningioma, to grade tumours according to the current 2016 WHO criteria, and to subtype tumours according to recognized histopathological classifications, where appropriate. Given the tendency for local aggressiveness in a subset of meningiomas, tumour recurrence and time to recurrence were the primary outcomes of interest in this study. Recurrence was defined as tumour growth following gross total resection or tumour progression following subtotal resection that resulted in a change in management and the time to recurrence was determined by calculating the duration from the date of surgery to first postoperative imaging documenting tumour recurrence. The extent of resection (Simpson grade) was extracted from the surgeon’s operative report and checked using postoperative magnetic resonance imaging (MRI). Additional clinical information, including—but not limited to—sex, age at surgery, previous treatment, post-operative treatment and tumour location were annotated for each sample.
DNA and RNA processing
DNA and RNA were extracted from adjacent but regionally distinct tissue for each patient. DNA was extracted from tumour and matched normal tissue (whole blood) as well as from healthy meninges samples using the DNeasy Blood and Tissue Kit (Qiagen) and quantified using the Nanodrop 1000 instrument (Thermo Scientific). Total RNA was isolated from tumour samples using the RNeasy Mini Kit (Qiagen) and quantified using the PicoGreen assay. RNA integrity was assessed using the Agilent 2100 Bioanalyzer (RNA; Agilent) and samples with RNA integrity number (RIN) > 7 were selected for further sequencing.
Genome-wide DNA methylation
Illumina Infinium MethylationEPIC BeadChip array (Illumina) was used to obtain genome-wide DNA methylation profiles on 250–500 ng of bisulfite-treated DNA (EZ DNA Methylation Kit, Zymo) per tumour and healthy meninges samples. Raw methylation files (*.idat) were imported, processed and normalized (ssNoob) using minfi18 (v.1.34). Probes that failed to hybridize (detection P value > 0.01) in one or more samples were removed from downstream analyses. Probes that overlapped with known single-nucleotide polymorphisms (SNPs), cross-reactive probes and probes that localized on X and Y chromosomes were also removed for all unsupervised analyses. Differentially methylated probes were identified using a modelling approach based on limma19. When comparing meningiomas to healthy meninges, CpG sites were considered differentially methylated if the absolute mean differences in β value were >0.35 and adjusted P value (FDR-corrected) was <0.05. When comparing each molecular group to healthy meninges, this threshold was adjusted to absolute mean differences of β > 0.1 and adjusted P (FDR-corrected) < 0.05. Probe annotation was performed using the UCSC Genome Browse (hg38 assembly).
Whole-exome sequencing
Exome libraries were prepared using 100 ng DNA from tumour tissue or matched normal DNA. Exome capture was performed using Agilent SureSelect Human Exome Library Preparation V5 or V6 COSIMC + kits and sequenced (pair-ended) on a HiSeq 2500 platform to a median of 191X. Raw sequencing data (fastq files) were aligned to the hg19 reference genome using BWA-MEM v.0.7.1220 with default parameters. PCR duplicate marking, indel realignment and base quality score recalibration were performed using Picard v.1.72 and GATK v.3.6.021. Data quality assessment was performed using CalculateHSMetrics from Picard. Somatic mutations were identified using Mutect V1.1.722 and Strelka v1.0.1323 for tumours with matched peripheral blood controls and Mutect2 V1.1 for tumours without matched peripheral blood controls. All mutations in genes that are recognized drivers in meningiomas (NF2, SMARCB1, TRAF7, AKT1, KLF4, SMO, POLR2A, DMD) were retained for statistical analyses. For the discovery of new, functionally relevant genes, germline variants with GnomAD24 population frequency >0.01% were removed to retain putative somatic mutations. Variants with allele frequency of >10% and a TGL frequency database of variants of <1% were retained to filter out initial passenger events. Genes with at least two somatic protein-altering mutations were selected, and the statistical basis for the filtered mutations was checked using MutSigCV25 for the overall cohort. We used a threshold of FDR <0.1 to consider variants as driver events, as described by the MutSigCV developers25. The functional effects of variants were subsequently annotated using Variant Effect Predictor v.92.026, OncoKB Precision Oncology Knowledge Base27, CancerHotspots.org28 and the dbNSFP database29. Statistically significant variants that were predicted to be actionable/driver alterations, or effects of which were predicted to be pathogenic or likely pathogenic, are reported and shown in Fig. 2a. Tumour mutation burden was calculated as the fraction of total number of protein-altering (nonsynonymous) somatic mutations across the callable exome space (in Mb).
Gene-level copy-number profiling
To assess allele-specific copy-number profiles, we used Sequenza v.2.1.219 for tumour-normal pairs and CNVkit v.0.9.630 for unmatched tumour samples using a pooled reference set of 60 peripheral blood samples from individuals that were unrelated to the study. We used conventional thresholds set by cBioportal31 to classify chromosomal gains and deletions (log2ratio > 0.7 as a high-level gain and log2ratio < −0.7 as a deep deletion). The degree of genomic disruption per sample was computed as the fraction of the genome that was affected by copy-number gains or losses.
RNA sequencing
mRNA libraries were generated using NEB Ultra II directional mRNA library prep kit according to the manufacturer’s protocol. Libraries were sequenced on the Illumina HiSeq 2500 high output flow cell (2 × 126bp), sequenced with 3 samples per lane to obtain approximately 70 million reads per sample. Raw sequencing data (fastq files) were processed and aligned to the human reference genome (GRCh38) using STAR (v.2.6.0a)32. Duplicate reads were removed, and reads were sorted using SamTools (v.1.333). Raw gene expression counts were computed for each sample using featureCounts in the package Rsubread (v.1.5.034) and subsequently normalized by counts per million (CPM) and subjected to TMM (trimmed mean of M) normalization using edgeR (v.3.22.3)35. TMM removes genes with low counts by CPM cutoff to filter out noise. The values for CPM cutoff were determined empirically by identifying the minimum value required to achieve the best normalization across samples. Using only protein-coding genes, the best CPM cutoff was determined to be 1.
Mutual information analysis
The MI metric13 was computed for each gene using all pairwise combinations of molecular data in our study (DNA methylation, CNAs, mRNA abundance, protein abundance). The MI metric measures the amount of information that is known about a gene by one datatype when the paired datatype is already known. Conceptually, MI is related to classic correlations (such as Spearman or Pearson correlations); however, statistical assumptions regarding linearity and ordering are not absolute, making this approach appropriate for the modelling of complex relationships such as those in cancer genomics. MI values of zero indicate completely independent variables, such that knowledge of one variable has no bearing on the knowledge of the other. For each pairwise comparison, data were discretized into 21 bins for each gene, and the MI between two datatypes was defined as MIxy = Hx + Hy − Hxy, where Hx and Hy the marginal entropies of datatypes x and y and Hxy is the joint entropy calculated using the R package Entropy (v.1.2.1). MI was normalized over the mean entropy of the two input vectors. To assess the statistical significance of normalized MI values, permutation testing was performed. Gene-level data were permutated 100,000 times to generate a null MI distribution and P values were calculated as the proportion of null MI values that were greater than or equal to the true observed MI. P values were FDR-adjusted and the significance threshold was set at an FDR of 5%. Consensus clustering36 was performed on those genes for which MI was significant for at least one datatype pair, after subsetting for genes with data available for all four datatypes. The divisive analysis clustering (diana) algorithm was applied to z-scored normalized MI values, using a maxK of 10 with 1,000 resampling repetitions. For methylation data, the Pearson correlation between gene-level RNA abundances and corresponding probe β values was calculated, and the probe with the greatest negative correlation was selected. For genes with annotated probes but without corresponding RNA abundance measures, the probe with the highest variance in β across samples was selected. This was done to achieve a 1:1 gene:probe relationship.
Single-platform clustering analyses
To identify the optimal number of clusters using mRNA data, gene-level somatic copy data and DNA methylation data, we performed consensus clustering using the ConsensusClusterPlus36 R package for each individual datatype separately. Consensus clustering was performed using the top 5,000 most variably expressed genes, 1,000 most variably altered genes and 10,000 most variably methylated CpG sites, as determined by median absolute deviation of logCPM, log2CNV ratios (where CNV is copy-number variation) and β values across all samples for RNA sequencing, gene-level copy number and DNA methylation data, respectively. Clustering was performed using Pearson correlation for the distance metric and Ward linkage algorithm with 1,000 resampling repetitions (ε = 0.8). For each platform, we computed the average silhouette width as well as plots of the cumulative distribution function of the consensus matrix for each k subgroups to identify the optimal k at which the cumulative distribution function reaches an approximate maximum. For gene-level copy number and gene expression we determined the optimal k = 6. For DNA methylation data, both k = 5 and k = 6 provided similar results. Given previous reports of k = 6 methylation subgroups, we selected k = 6 as the optimal number of methylation-based clusters. Samples were then projected into a two-dimension space using t-SNE for cluster assignment and visualization for each individual platform separately. Divergence from expected recurrence-free survival patterns in our samples using a previously established methylation-based cluster classification3 led us to use data-driven methylation cluster groupings for our analyses in this paper. Adjusted Rand indices were calculated on cluster assignments for each pairwise combination of datatypes to determine the degree of cluster overlap.
Cluster-of-cluster assignments
To comprehensively integrate mRNA, copy number and DNA methylation data, we used the COCA algorithm that has been used by the The Cancer Genome Atlas to identify molecular subtypes of systemic cancers9,10,11,12. Cluster assignments from unsupervised t-SNE-based individual platform clustering were first binarized into indicator variables that were combined to construct a matrix of clusters (columns are binarized cluster memberships and rows are samples). This second-order matrix was then subjected to an additional round of consensus clustering to examine the relationship between samples across molecular features. The optimal number of subgroups was selected by computing and maximizing the average silhouette width from k = 2 to k = 10. To examine the relative importance of each datatypes, COCA was repeated with all combinations of two datatypes at a time. Cluster assignments by integration of three versus two datatypes were compared for overlap by computing Adjusted Rand Indices (ARI).
Estimation of the cancer cell fraction
The cancer cell fraction of variant i (CCFi) was calculated as follows:
where ui is a function of the variant allele fraction of variant i (fi), sample purity (ρ), the local copy number of the tumour cells at site i (ntotal,t,i) and the local copy number of the normal cells at site i (ntotal,n,i, assumed to be 2) (ref. 37):
The variant allele fraction of variant i (fi) was directly calculated using the number of reference reads for locus i (rref,i) and the number of alternate reads for locus i (rmut,i).
For each sample, we estimated sample purity (\(\rho \)) as previously described using DNA methylation data38. The local copy number of the tumour cells at site i (ntotal,t,i) was transformed from the segment mean at site i (si):
The mutation multiplicity of variant i (mi) was determined using the following equation:
Finally, if the CCFi was greater than 0.80, then variant i was considered clonal.
Differential gene-expression analysis
Differential gene-expression analysis was computed using gene-wise negative binomial generalized linear models with quasi-likelihood tests (F test, edgeR35 v.3.22.3). Genes were ranked by combining the direction of fold changes (FC) and computed P values using the following formula: sign(log2FC) × −log10(P), where sign(log2FC) determines the direction of the change (upregulated is positive and downregulated is negative) and −log10(P) determines the magnitude of ranking. Gene-set enrichment analysis (GSEA, v.3.0) was performed as previously described, using ranked scores as input to determine whether differentially expressed genes belong to common biological pathways39.
Pathway analysis and network maps
Pathway analyses and network maps were generated as previously described39. Pathways were defined by the gene set file Human_GOBP_AllPathways_no_GO_iea_June_20_2019_symbol.gmt that is maintained and updated regularly by the Bader laboratory (http://download.baderlab.org/EM_Genesets/). GeneSet size was limited to range between 10 and 200, and 2,000 permutations were carried out. The results of the pathway analysis were visualized using the EnrichmentMap App (v.1.2.0) in Cytoscape (v.3.7.2). Network maps were generated for nodes with FDR q value < 0.01, P < 0.0001, and nodes sharing gene overlaps with Jaccard coefficient > 0.25 were connected by a green line (edge). Clusters of related pathways were identified and annotated using a Cytoscape app that uses a Markov Cluster algorithm that connects pathways by shared keywords in the description of each pathway (AutoAnnotate, v.1.2). The resulting groups of pathways are designated as the major pathways in a circle.
FDA drug mapping
In order to discover realistic and new therapeutic agents, we examined whether FDA-approved drugs could be repurposed for the treatment of meningioma by examining for the presence of FDA-approved drug targets in our network analyses. Drugs were selected by the number of target genes in the leading edge of significant GSEA pathways for indicated comparison, then each drug was ranked by the number of genes plus pathways targeted. Finally, the number of significant genes targeted were divided by the total number of target genes of the drug to assess the specificity. This scoring system selected the drugs targeting the greatest number of driving genes in significant biological pathways with high specificity. The resulting list of drugs were grouped by common targets to produce a higher-level summary of the class of drugs with the highest possibility of effective treatment. Individual drugs were visualized on pathway maps using post-analysis function in the Enrichment Map plugin of Cytoscape app.
Gene fusion identification
Interchromosomal and intrachromosomal gene fusion events were detected using FusionCatcher v.1.1.0 with default parameters. FusionCatcher aligns reads to the human reference genome (GRCh38) using Bowtie40 (v.1.2), Bowtie241 (v.2.3), BLAT42 (v.0.35) and STAR BLAT32 (v.2.7). Adjacent and read-through fusions were filtered out from analyses and fusions with Counts_of_common_mapping_reads = 0 were selected to reduce false positive detection of genes with similar sequence homology. A stringent threshold for conservative estimation of fusion events (unique spanning reads ≥25) was used to assess interchromosomal and intrachromosomal fusion events.
Generalization cohort
Large (n > 50), multi-omic meningioma datasets in the literature with matched individual patient outcome data were not available for use as independent validation. Therefore, to confirm the generalizability of the association with integrative molecular groups and their association with outcomes, we assembled an independent cohort of 80 meningioma patient samples with longitudinal outcome data and generated mRNA-sequencing data. Assignment of molecular group for each new sample was performed by a single-sample GSEA (ssGSEA) using the top 50 highly expressed genes for each group in the initial discovery cohort. Cluster assignment was determined by maximal scores from ssGSEA analysis and checked by unsupervised hierarchical clustering of ssGSEA scores. Kaplan–Meier estimates of survival with log-rank tests for association were performed to test the association of molecular groups in the new independent cohort with outcome. The association of molecular groups with outcomes was compared to WHO grade by generation of Brier prediction curves and computation of Brier scores.
The discriminative capacity of gene-expression profiles to distinguish molecular groups overall was quantitated using true gene-expression classifiers (generalized linear model, default alpha and lambda parameters) for each molecular group in the discovery cohort. To do this, we randomly split our cohort into training and test sets, with 90% of the data in the training set and the remaining 10% of the data in the test set. Expression classifiers for each molecular group were trained using the top 50 highly expressed genes for each molecular group, and the performance for each classifier was tested using held-out samples in test cohort by computing the area under the receiver–operative characteristic curve. This process was repeated for a total of 50 iterations of training and testing.
Epigenetic mitotic clock analyses
We used previously described mitotic clocks (epiTOC16, epiTOC215 and solo-WCGW14) that are based on DNA methylation to examine regions of the genome that are either fully methylated or unmethylated in multiple fetal tissues but gain or lose methylation as a function of mitotic age. The epiTOC model calculates a weighted average methylation over 354 CpGs on the 850K array at gene promoters marked by the PRC2 complex that are constitutively unmethylated in fetal tissue and increase in methylation with age and cell division. The epiTOC2 model estimates the mitotic age (adjusted for chronological age of patient) using a weighted subset of 151 CpGs from the epiTOC model that are most likely to change in DNA methylation levels with age. The solo-WCGWs are a set of CpGs at the WCGW motif without flanking CpGs that are hypomethylated in fetal tissues and gain methylation with age and cell division. A total of 6,214 solo-WCGWs that were originally described are found on the EPIC array. Of note, 648 of these are uniformly hypomethylated across multiple fetal tissue types, as previously described, and therefore a weighted average of these 648 CpG sites was used to derive the ‘HypoClock’ score.
Transcription factor analyses
We identified master transcription factors for each molecular group as previously described using ElmerV217. First, differentially methylated distal CpGs at non-promoter (probes further than 2 kb from the transcription start site) sites were identified between each molecular group and every other molecular group independently as well as all other molecular groups as a group. Putative target genes were identified for each differentially methylated CpG by computing the correlation between methylation of the probe and the expression of the closest 10 upstream and 10 downstream genes. Motif occurrences were identified using HOMER within 250-bp region for significantly hypomethylated probes with putative gene targets and enrichment for motifs are calculated by computing the odds ratio (and 95% confidence interval) that each probe in a probe set contains motif occurrences in comparison to a background of all distal probes on the 850K array. Transcription factors were considered enriched if the lower bound of the 95% confidence interval was greater than 1.1. Finally, the mean methylation of all probes in probe-gene pairs that contained a given motif instance within 250 bp were compared to the average expression of a set of 1,639 transcription factors43,44. These were then ranked by degree of anticorrelation using −log10(FDR) in order to identify master regulator transcription factors by transcription factor subfamily.
Shotgun proteomics
Approximately 1–2 mg of fresh frozen meningioma tumours were pulverized using a Covaris cryoPREP Pulverizer. Pulverized tissue was then solubilized in 300 µl of 50% (v/v) 2,2,2-trifluoroethanol in phosphate-buffered saline (pH 7.4) with a 5 min incubation at 95 °C, repeated probe sonication, freeze-thaw cycling, followed by a two-hour heated incubation at 60 °C. Protein lysate (100 µg) was denatured with 5 mM dithiothreitol for 30 min at 60 °C and reduced disulfite bonds were subsequently alkylated with 25 mM iodoacetamide for 30 min at room temperature in the dark. Proteins were digested into peptides with 4 µg of trypsin at 37 °C overnight. Peptides were then desalted and purified using C18-based solid-phase capture. Eluted peptides were lyophilized and solubilized in mass-spectrometry-grade water with 0.1% methanoic acid and peptide concentration was quantified using a NanoDrop Lite spectrophotometer (at 280 nm). For each sample, an Easy1000 nanoLC was used to load 2 µg of peptides onto a 2 cm trap column (Thermo Scientific). The peptides were separated along a four-hour gradient using a 50 cm EasySpray analytical column coupled by electrospray ionization to an Orbitrap Fusion (Thermo Scientific) tribrid mass spectrometer. Peptides were detected using a Top25 data-dependent acquisition method. The acquired data was searched using Maxquant (v.1.6.2.345) against a UniProt complete human protein sequence database (v.2019_04) with an FDR of 1% for peptide spectral matches. Two missed cleavages were permitted along with the fixed carbamidomethyl modification of cysteines, the variable oxidation of methionine and variable acetylation of the protein N terminus. Relative label-free protein quantitation was calculated using MS1-level peak integration along with the matching-between-runs feature enabling a 2 min retention time matching window. Proteins identified with a minimum of two peptides were carried forward for further analysis. Protein groups with log2FC > 2 (that is, 4-fold higher expression or more), and FDR < 0.05 were considered specific for each group.
Validation of proteomic findings by immunohistochemical analyses
To validate the enrichment of group-specific proteins identified by proteomic data, we performed immunohistochemical analyses for S100B, SCGN, ACADL and MCM2 in a cohort of 44 tumours with known molecular group status. Experimentation and analyses were performed blinded to molecular group status. In brief, consecutive 5-µm formalin-fixed paraffin sections were rehydrated and heat-mediated antigen retrieval using sodium citrate buffer (pH 6) was performed. Slides were washed in 3% H2O2 in methanol and blocked in 5% BSA in PBST for 1 h at room temperature followed by overnight incubation at 4 °C with anti-S100B (ThermoFisher, 701340, dilution 1:100), anti-SCGN (Sigma, HPA006641, dilution 1:500) anti-ACADL (Sigma, HPA011990-100UL, dilution 1:200) or anti-MCM2 (Cell Signaling, 12079S, dilution 1:200). The expression signals were developed using DAB Peroxidase Kit and the slides were counterstained with haematoxylin, dehydrated, and coverslipped. Whole-slide images were digitized and obtained using virtual microscopy. Tumour tissue was annotated in each whole slide by an experienced and blinded neuropathologist and subsequently subjected to unbiased quantitative digital pathological assessment using the Multiplex IHC module on HALO software (Indica Labs).
Droplet-based single-nuclear RNA-sequencing
Ten frozen archived tumour specimens and two frozen archived healthy meninges were minced with a sterile scalpel and homogenized using a dounce tissue grinder (size A and B, Sigma Aldrich) in ice cold lysis buffer (0.32 M sucrose, 5 mM CaCl2, 3 mM MgAc2, 20 mM Tris-HCl, 0.1 EDTA, 40 U ml−1 RNase inhibitor and 0.1% Triton X-100 in DEPC-treated water). Homogenized tissue was centrifuged at 500g for 10 min at 4 °C, washed in two rounds using ice cold wash buffer (1× PBS, 12 mM EGTA pH 8.0 and 0.2 U μl−1 RNase inhibitor) and the nucleus pellet was subsequently resuspended in resuspension buffer (1× PBS, 0.04% BSA) prior to filtration using 40 μm Flowmi cell strainer (Sigma Aldrich). Isolated nuclei were stained with DAPI and fluorescence-sorted (BD Influx BRV, Becton Dickinson Biosciences) to retain healthy nuclei. DAPI+ nuclei were washed and resuspended in resuspension buffer. Nuclei were counted and approximately 6,000–8,000 nuclei were loaded onto a 10x Chromium controller using the Chromium Single Cell 3’ Library & Gel Bead v3 (10x Genomics) for each sample. Single nuclei were partitioned into barcoded gel beads in emulsion in the Chromium instrument, followed by cell lysis and reverse transcription of RNA in the droplets. Breaking of the emulsion was followed by cDNA amplification and library construction as per the manufacturer’s recommendations. Samples were sequenced Illumina NovaSeq (10x specific protocol) with a median target sequencing depth of 60,000 reads per nucleus.
snRNA-seq raw data processing, filtering and validation of cells to patients
Raw sequencing data (bcl files) were converted to demultiplexed fastq files (Illumina bcl2fastq, v.2.19.1) and aligned to the human genome reference sequence (hg38). Expression matrix of unique molecular identified counts per gene per nuclei was obtained using CellRanger (10x Genomics). As the first step for validating cells to patients, we looked to confirm that cells had data that covered known SNP regions. To do this, we quantified the number of unique molecular identifiers (UMIs) mapping to a panel of 7.4 million SNPs identified through the 1,000 Genomes Project46 with minor allele frequency > 5% using cellsnp-lite. Two of our samples had highly sparse coverage of known SNP regions and were not reliably genotypable, and were therefore removed from further analyses.
To validate the assignment of cells to patients for samples that had potential overlap in processing, we compared SNPs derived from single-cell RNA sequencing data to SNPs derived from bulk RNA-seq data using demuxlet47. Demuxlet was developed to deconvolute sample identity when multiple samples are pooled by barcoded single-cell sequencing. Variant call format files from bulk RNA sequencing data were generated and compared to variants identified in single-cell data by demuxlet. Only cells with genotypes that aligned to the expected sample were retained for further analyses. Potential doublets were identified using scDblFinder (v.3.13) and removed.
From all remaining cells, we quantified two quality measures for each cell: the number of UMIs detected, and fraction of mitochondrial transcripts. Low-quality cells in which >1.5% of transcripts derived from the mitochondria and cells with low complexity libraries in which less than 1,000 UMIs were detected were removed. After data filtering, a total of 54,393 high-quality single nuclei that were genotyped to 10 samples were retained for analyses.
snRNA-seq clustering of all cells
Library size normalization was performed as previously described using scran, in which hierarchical clustering of cells using Spearman distances subset cells into more groups, and then scaling factors per cell were determined by randomly pooling cells, computing summed library sizes, and comparing to average library size across all cells in each group48,49. Normalized UMI counts were used for clustering by optimizing a shared-nearest-neighbour modularity function with Seurat50. First, principal component analysis was performed using highly variable genes (FDR < 0.001) identified by scran. The number of significant principal components (PC, 10) was determined on the basis of the inflection point of a ‘scree’ plot. Next, a shared-nearest-neighbour graph was built from distances computed in first 10 PC space and clusters were identified by optimizing the modularity function within this space with a resolution set to 0.1. Gene expression and clustering results were visualized using t-SNE of the selected principal components.
Cell type classification
Cells were assigned to different cell types based on consensus by:
(1) Similarity of expression profiles: As neoplastic and stromal/immune compartments are expected to have different expression profiles, we first correlated (Pearson) the expression profile of each cell to every other cell. Unsupervised hierarchical Pearson clustering with Ward linkages on the matrix of correlation values was performed and two major clusters (putative neoplastic and non-neoplastic) of cells were identified.
(2) Copy number profiles. We used inferCNV(v1.1.1)51 to infer CNAs of neoplastic and non-neoplastic cells with snRNA-seq data. Cells from healthy meninges were used as the reference set. Genes were ordered from the human GRCh38 assembly, and a heat map illustrating relative expression intensities of neoplastic nuclei to reference population across the genome was generated for visualization. Almost all neoplastic clusters harboured loss of chromosome 22q that was not observed in non-neoplastic cells that were generally devoid of significant CNA. We further computed a general metric of aneuploidy using inferred CNA data by first scaling CNA to the range of −1 to 1, and then summing the absolute copy number ratios for all genes. The degree of aneuploidy was later used to compare cells of high versus low potency.
(3) Expression of canonical markers: Significantly differentially expressed genes were identified for each cluster using FindAllMarkers in Seurat and these were inspected for canonical immune and stromal cell markers. Enrichment of these markers across clusters was visualized by bubble plots and was indicative of cell-type annotation. Predictions regarding cell cycle phases were made for neoplastic cells on the basis of the expression of a core set of genes, as previously described52.
Correlation of CNA inferred from snRNA-seq data and bulk whole-exome sequencing data
To correlate CNA data from snRNA-seq and bulk whole-exome sequencing data, inferred CNA ratios from snRNA-seq were scaled to values between −1 and 1 such that the two datasets were similarly scaled. Arm-level copy number ratios were then computed from snRNA-seq and bulk CNA data independently, as follows:
Where CNi is the copy number ratio of the ith gene in segment s and Li is the length of the ith gene. Pearson and Spearman correlations were then computed on arm-level CNA ratios from both datatypes.
snRNA-seq clustering of individual samples
To examine heterogeneity within tumours, we clustered cells from each patient independently using two independent approaches (Seurat and DBSCAN). Clustering by Seurat50 was performed as described above, with resolution set to 0.05 to account for the smaller number of cells with single sample analyses.
DBSCAN identifies clusters by identifying dense regions in space, ensuring that the neighbourhood of a radius (ε) has to contain a minimum number of neighbours (minPts). DBSCAN identifies outliers of cells that do not belong to any clusters (considered noise). To cluster cells by DBSCAN we first normalized raw expression levels for each sample as follows:
where CPMi for genes i to n was computed as 106 × UMIi/\(\Sigma \)(UMI1…n). These values were then centred to the average expression of the gene across all cells in the sample to define relative expression of each gene in each cell. Using this data, each sample was subjected to dimensionality reduction by t-SNE (with a perplexity of 30) followed by density clustering using DBSCAN (parameters ε = 1.8 and minPts = 5). Cells that did not meet these parameters were considered unclassifiable and coloured grey in the t-SNEs.
Statistical evaluation of between- and within-patient variation
We used a one-way ANOVA test on the top 10 principal components for all neoplastic cells to compare between-patient variability and within-patient variability as previously described53. The F statistic by ANOVA divides the variability observed in the dataset into between-patient components and within-patient components. F statistics >1 indicate that the between-patient variation is greater than the within-patient variation.
Statistical evaluation of two cell features
To examine whether two features of a cell were associated, we used mixed-effects logistic regression models that are able to account for cell-to-patient dependencies, as previously described54. We specifically used these models to test for the enrichment of immune cells in MG1, the enrichment of cycling cells in MG3 and MG4.
Non-negative matrix factorization to identify intrinsic gene expression programs
To identify the intrinsic expression program, we applied NMF to relative expression levels used for DBSCAN analyses after transforming all negative values to zeros, as previously described54,55,56. Factors k ranged from six to nine and genes were ranked by NMF scores for each expression programs identified. A total of 39 expression programs were identified across eight tumour samples. We then performed hierarchical clustering of programs using the extent of shared genes as a distance metric (using the top 50 genes in each program) to identify meta-signatures that were recurrent across samples. We calculated the Pearson correlation coefficient between NMF scores and the fraction of mitochondrial genes to assess for the relationship of each program with technical confounders. One cluster of programs (25–39) showed higher positive correlation with fraction of mitochondrial genes quantitated. This was confirmed by manual inspection of the genes, which showed several mitochondrial and ribosomal genes that highly score in these programs. These programs were excluded from further analyses as they were favoured to reflect technical artifacts. We then computed activation scores of each NMF program from all cells using AUCell34(v.1.8.0) and compared the distribution of activation scores across tumours.
Deconvolution of bulk RNA-seq data using snRNA-seq signatures
We used CIBERSORTx57(v.1.0) to deconvolute bulk mRNA-seq data from all samples in this study. We first used CIBERSORTX to generate a gene signature matrix for each single-cell cluster from our single-cell RNA sequencing data. Genes with weights greater than 400 were selected for each cluster and used in consensus k-means clustering with 5,000 repeats to partition bulk RNA sequencing data into four groups for comparison with bulk molecular classification.
We then generated a signature matrix for each cell type (macrophage, T cell, endothelial cell, fibroblast, neoplastic) using CIBERSORTx, and then used this to determine cell-type composition of each of our samples with bulk RNA sequencing data using single cell Correction S mode with 100 permutations.
Patient-derived cell lines
Fresh tumour specimens were obtained intraoperatively from five patients from whom informed consent for tissue banking was obtained previously. Cell suspensions were created and maintained as previously reported (PMID 26174772) on ThermoFisher BioLite 100 mm Tissue Culture dishes in DMEM/F12 (Life Technologies, 10565) supplemented with 1 mM non-essential amino acids (Life Technologies, 11140), 100 U ml−1 antibiotic-antimycotic (Life Technologies, 15240) and 10% fetal bovine serum (Life Technologies, 16141) in a humidified atmosphere with 5% CO2. Once confluent, cells were passaged following trypsinization. DNA and RNA were extracted from an aliquot of each cell line. DNA was subjected to bisulfite conversion for DNA methylation profiling. To demonstrate that these cell lines are faithful models of meningiomas, we compared the genome-wide methylome profiles of cell lines to meningiomas from our cohort as well as a published panel of 2,798 tumours from 40 brain tumour types58. We found that all cell lines in this experiment clustered together with human meningioma tumours. In addition, classification of our cell lines using a publicly available DNA methylation-based random-forest model (DKFZ MolecularNeuropathology.org online classifier v.3.1.5) assigned all primary patient-derived cell lines into the meningioma methylation class with high calibrated scores (0.97–0.99). To assign cell lines to molecular groups, we generated mRNA sequencing data from cell lines and performed ssGSEA using the top 50 highly expressed genes for each molecular group from the cohort of tumours in our dataset. Cell lines were assigned to molecular group by maximal ssGSEA scores.
Cell viability assay
Meningioma cells (ranging from 1,500–4,500 cells based on the plating efficiency of each cell line) were plated in technical triplicates in Corning 96-well white-walled plates. Cells were treated with vorinostat (SAHA/MK0683, InvivoChem V0255; diluted to concentrations 100 nM, 500 nM, 1,000 nM, 5,000 nM) or 5-azacytidine (InvivoChem V0404; 10 nM, 50 nM, 100 nM, 500 nM, 1,000 nM) for 10 days. A medium-only control was used for each replicate of each drug treatment, and a DMSO control was used for vorinostat and 5-azacytidine-treated cells. Three separate biological replicates separated by at least one passage of each cell line were completed. After the completion of treatment, CellTitre-Glo luminescent cell viability assay was performed on all samples in accordance with the manufacturer’s instructions (Promega, G7570). Cells were incubated for 10 min with the CellTitre-Glo reagent and luminescence was measured using a 96-well plate reader (GloMax-96 microplate luminometer; Promega). Background luminescence was measured in blank wells with medium without cells and subtracted from experimental values automatically. Statistical analyses of intergroup differences between cell lines at each dose of each respective drug were performed using a two-way ANOVA followed by Tukey’s test.
In vivo patient-derived xenograft
For intracranial xenograft experiments, 1 × 106 MG4 patient-derived cells were injected into the subdural space of NSG mice. Mice were anaesthetized and their craniums were fixed in a stereotaxic frame. An incision was made 3 mm lateral to the midline on the right side of the skull. The bregma was visualized and a burrhole was drilled using an automated 1.5 mm drillbit 3 mm lateral and 1 mm anterior to the bregma. Cells were injected at a depth of 1 mm to the skull surface using a 26-gauge needle and stereotactic Hamilton syringe in 5 μl of media over 3 min. After injection, the syringe was slowly removed over 2 min to limit reflux of cells. The incision was closed with 6-0 absorbable sutures and Vetbond tissue adhesive was applied on top. Mice were treated with either vorinostat (50 mg kg−1 1:1 DMSO:PBS) or vehicle control (1:1 DMSO/PBS at equivalent volume) via intraperitoneal injection daily for 10 days, starting on post-implantation day 7. All mice were imaged at 3–5 days post xenograft implantation using a Bruker 7-Tesla preclinical MRI (STTARR imaging facility, Toronto, Ontario) to confirm intracranial implantation. Additional serial MRI scans were performed every 3 to 7 days based on the availability of our imaging facility to document interval tumour growth. MRI volumetric analysis of tumours was performed by an individual blinded to treatment group using the Horos/OsirixTM open source DICOM reader (GNU Lesser General Public License, v.3 (LGPL-3.0)). Xenograft tumours were segmented on each MRI slice manually and then reconstructed automatically to obtain a volume measurement for each mouse at each radiographic time point. Statistical analyses comparing the mean xenograft volume between the vorinostat-treated and control mice were performed at each time point using a Mann–Whitney U-test, with statistical significance set at P < 0.05. Mice were euthanized when they reached their physiological or experimental endpoint in accordance with our animal care facility and the Canadian Council on Animal Care (CCAC) guidelines. Specifically, the endpoint was reached when mice lost >20% of their starting body weight, demonstrated considerable lethargy and decreased activity, had visible cranial enlargement, or had tumour volumes exceeding 500 mm3 on MRI volumetric measurements. None of the mice in our study exceeded these endpoints without being mandatorily euthanized and no mouse tumours achieved or exceeded the volumetric endpoint.
Survival analyses
For comparison of survival between independent groups, Kaplan–Meier survival plots were generated using the package survminer and log-rank tests were performed to test the null hypothesis of no differences between independent subgroups. Univariable hazard ratios with 95% confidence interval and P values for clinical factors as well as MG1–MG4 were computed by fitting Cox proportional hazards models. Multivariable survival analyses were performed by fitting Cox proportional hazards models that included all factors that were significant on univariable analyses. Prediction error curves were generated to compare the discriminative capacity of Cox proportional hazards models by leave-one-out cross-validation.
Reporting summary
Further information on research design is available in the Nature Research Reporting Summary linked to this paper.
Data availability
Raw sequencing data for all datatypes have been deposited into public repositories. Proteomic data has been deposited to the Mass Spectrometry Interactive Virtual Environment (MassIVE, https://massive.ucsd.edu/; ID MSV000086901). DNA methylation idat files have been deposited to the Gene Expression Omnibus (GEO; GSE180061). Whole-exome sequencing (fastq), bulk mRNA (fastq) and snRNA (fastq) datasets have been deposited to the European Genome Archive (https://www.ebi.ac.uk/ega/) under study ID EGAS00001004982 and dataset IDs EGAD00001007051, EGAD00001007494 and EGAS00001004982. The processed genomic data has been submitted to cBioportal at https://www.cbioportal.org/study/summary?id=mng_utoronto_2021. Source data are provided with this paper.
Code availability
Specific code will be made available upon request to G.Z.
References
Ostrom, Q. T. et al. CBTRUS statistical report: primary brain and other central nervous system tumors diagnosed in the United States in 2012–2016. Neuro. Oncol. 21, v1–v100 (2019).
Goldbrunner, R. et al. EANO guidelines for the diagnosis and treatment of meningiomas. Lancet Oncol. 17, e383–e391 (2016).
Sahm, F. et al. DNA methylation-based classification and grading system for meningioma: a multicentre, retrospective analysis. Lancet Oncol. 18, 682–694 (2017).
Clark, V. E. et al. Genomic analysis of non-NF2 meningiomas reveals mutations in TRAF7, KLF4, AKT1, and SMO. Science 339, 1077–1080 (2013).
Brastianos, P. K. et al. Genomic sequencing of meningiomas identifies oncogenic SMO and AKT1 mutations. Nat. Genet. 45, 285–289 (2013).
Clark, V. E. et al. Recurrent somatic mutations in POLR2A define a distinct subset of meningiomas. Nat. Genet. 48, 1253–1259 (2016).
Harmancl, A. S. et al. Integrated genomic analyses of de novo pathways underlying atypical meningiomas. Nat. Commun. 8, 14433 (2017).
Nassiri, F. et al. DNA methylation profiling to predict recurrence risk in meningioma: development and validation of a nomogram to optimize clinical management. Neuro. Oncol. 21, 901–910 (2019).
Hoadley, K. A. et al. Cell-of-origin patterns dominate the molecular classification of 10,000 tumors from 33 types of cancer. Cell 173, 291–304.e6 (2018).
Koboldt, D. C. et al. Comprehensive molecular portraits of human breast tumours. Nature 490, 61–70 (2012).
Cancer Genome Atlas Research Network. Comprehensive, integrative genomic analysis of diffuse lower-grade gliomas. N. Engl. J. Med. 372, 2481–2498 (2015).
Hoadley, K. A. et al. Multiplatform analysis of 12 cancer types reveals molecular classification within and across tissues of origin. Cell 158, 929–944 (2014).
Reshef, D. N. et al. Detecting novel associations in large data sets. Science 334, 1518–1524 (2011).
Zhou, W. et al. DNA methylation loss in late-replicating domains is linked to mitotic cell division. Nat. Genet. 50, 591–602 (2018).
Teschendorff, A. E. A comparison of epigenetic mitotic-like clocks for cancer risk prediction. Genome Med. 12, 56 (2020).
Yang, Z. et al. Correlation of an epigenetic mitotic clock with cancer risk. Genome Biol. 17, 205 (2016).
Silva, T. C. et al. ELMER v.2: an R/Bioconductor package to reconstruct gene regulatory networks from DNA methylation and transcriptome profiles. Bioinformatics 35, 1974–1977 (2019).
Aryee, M. J. et al. Minfi: a flexible and comprehensive Bioconductor package for the analysis of Infinium DNA methylation microarrays. Bioinformatics 30, 1363–1369 (2014).
Ritchie, M. E. et al. limma powers differential expression analyses for RNA-sequencing and microarray studies. Nucleic Acids Res. 43, e47 (2015).
Li, H. & Durbin, R. Fast and accurate short read alignment with Burrows–Wheeler transform. Bioinformatics 25, 1754–1760 (2009).
McKenna, A. et al. The Genome Analysis Toolkit: A MapReduce framework for analyzing next-generation DNA sequencing data. Genome Res. 20, 1297–1303 (2010).
Cibulskis, K. et al. Sensitive detection of somatic point mutations in impure and heterogeneous cancer samples. Nat. Biotechnol. 31, 213–219 (2013).
Saunders, C. T. et al. Strelka: accurate somatic small-variant calling from sequenced tumor–normal sample pairs. Bioinformatics 28, 1811–1817 (2012).
Lek, M. et al. Analysis of protein-coding genetic variation in 60,706 humans. Nature 536, 285–291 (2016).
Lawrence, M. S. et al. Mutational heterogeneity in cancer and the search for new cancer-associated genes. Nature 499, 214–218 (2013).
McLaren, W. et al. The Ensembl Variant Effect Predictor. Genome Biol. 17, 122 (2016).
Chakravarty, D. et al. OncoKB: a precision oncology knowledge base. JCO Precis. Oncol. 1–16 (2017).
Chang, M. T. et al. Identifying recurrent mutations in cancer reveals widespread lineage diversity and mutational specificity. Nat. Biotechnol. 34, 155–163 (2016).
Dong, C. et al. Comparison and integration of deleteriousness prediction methods for nonsynonymous SNVs in whole exome sequencing studies. Hum. Mol. Genet. 24, 2125–2137 (2015).
Talevich, E., Shain, A. H., Botton, T. & Bastian, B. C. CNVkit: genome-wide copy number detection and visualization from targeted DNA sequencing. PLoS Comput. Biol. 12, e1004873 (2016).
Cerami, E. et al. The cBio Cancer Genomics Portal: an open platform for exploring multidimensional cancer genomics data. Cancer Discov. 2, 401–404 (2012).
Dobin, A. et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics 29, 15–21 (2013).
Li, H. et al. The Sequence Alignment/Map format and SAMtools. Bioinformatics 25, 2078–2079 (2009).
Aibar, S. et al. SCENIC: single-cell regulatory network inference and clustering. Nat. Methods 14, 1083–1086 (2017).
Robinson, M. D., McCarthy, D. J. & Smyth, G. K. edgeR: a Bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics 26, 139–140 (2010).
Wilkerson, M. D. & Hayes, D. N. ConsensusClusterPlus: a class discovery tool with confidence assessments and item tracking. Bioinformatics 26, 1572–1573 (2010).
Dentro, S. C., Wedge, D. C. & Van Loo, P. Principles of reconstructing the subclonal architecture of cancers. Cold Spring Harb. Perspect. Med. 7, a026625 (2017).
Aran, D., Sirota, M. & Butte, A. J. Systematic pan-cancer analysis of tumour purity. Nat. Commun. 6, 8971 (2015).
Reimand, J. et al. Pathway enrichment analysis and visualization of omics data using g:Profiler, GSEA, Cytoscape and EnrichmentMap. Nat. Protocols 14, 482–517 (2019).
Langmead, B., Trapnell, C., Pop, M. & Salzberg, S. L. Ultrafast and memory-efficient alignment of short DNA sequences to the human genome. Genome Biol. 10, R25 (2009).
Langmead, B. & Salzberg, S. Fast gapped-read alignment with Bowtie 2. Nat. Methods 9, 357–359 (2013).
Kent, W. J. BLAT—the BLAST-Like Alignment Tool. Genome Res. 12, 656–664 (2002).
Lambert, S. A. et al. The human transcription factors. Cell 172, 650–665 (2018).
Wingender, E., Schoeps, T., Haubrock, M., Krull, M. & Dönitz, J. TFClass: expanding the classification of human transcription factors to their mammalian orthologs. Nucleic Acids Res. 46, D343–D347 (2018).
Cox, J. & Mann, M. MaxQuant enables high peptide identification rates, individualized p.p.b.-range mass accuracies and proteome-wide protein quantification. Nat. Biotechnol. 26, 1367–1372 (2008).
The 1000 Genomes Project Consortium A global reference for human genetic variation. Nature 526, 68–74 (2015).
Kang, H. M. et al. Multiplexed droplet single-cell RNA-sequencing using natural genetic variation. Nat. Biotechnol. 36, 89–94 (2018).
L. Lun, A. T., Bach, K. & Marioni, J. C. Pooling across cells to normalize single-cell RNA sequencing data with many zero counts. Genome Biol. 17, 75 (2016).
Lun, A. T. L., McCarthy, D. J. & Marioni, J. C. A step-by-step workflow for low-level analysis of single-cell RNA-seq data with Bioconductor. F1000Research 5, 2122 (2016).
Butler, A., Hoffman, P., Smibert, P., Papalexi, E. & Satija, R. Integrating single-cell transcriptomic data across different conditions, technologies, and species. Nat. Biotechnol. 36, 411–420 (2018).
Tickle, T. I., Georgescu, C., Brown, M. & Haas, B. Infer copy number variation from single-cell RNA-seq data. https://doi.org/10.18129/B9.bioc.infercnv (2019).
Tirosh, I. et al. Dissecting the multicellular ecosystem of metastatic melanoma by single-cell RNA-seq. Science 352, 189–196 (2016).
Azizi, E. et al. Single-cell map of diverse immune phenotypes in the breast tumor microenvironment. Cell 174, 1293–1308.e36 (2018).
Jerby-Arnon, L. et al. Opposing immune and genetic mechanisms shape oncogenic programs in synovial sarcoma. Nat. Med. 27, 289–300 (2021).
Kinker, G. S. et al. Pan-cancer single-cell RNA-seq identifies recurring programs of cellular heterogeneity. Nat. Genet. 52, 1208–1218 (2020).
Izar, B. et al. A single-cell landscape of high-grade serous ovarian cancer. Nat. Med. 26, 1271–1279 (2020).
Newman, A. M. et al. Determining cell type abundance and expression from bulk tissues with digital cytometry. Nat. Biotechnol. 37, 773–782 (2019).
Capper, D. et al. DNA methylation-based classification of central nervous system tumours. Nature 555, 469–474 (2018).
Acknowledgements
F.N. is supported by the Canadian Institute of Health Research (CIHR) Vanier Scholarship, AANS/CNS Section on Tumors & NREF Research Fellowship Grant, and Hold’em for Life Oncology Fellowship. G.Z is funded by a CIHR Project Grant Award (173241), a Canadian Cancer Society Innovation Grant (706898) as well as a Quest for Cures Grant (GN-000430) and Clinical Biomarkers Grant (GN-000693) from the Brain Tumour Charity UK. P.C.B. was supported by the NIH/NCI under awards P30CA016042, U24CA248265 and P50CA211015. We also thank N. Dewitt for editing, and the Mary Hunter Meningioma Program and Toronto Western Hospital Foundation.
Author information
Authors and Affiliations
Contributions
F.N., K.A. and G.Z. conceived and designed the study. F.N., S.S., and J.Z.W. collected all biomaterials and clinical data. K.A., A.G. and S. Karimi reviewed the pathological sections. F.N. prepared specimens for whole-exome sequencing, DNA methylation, mRNA sequencing and single-cell RNA sequencing. S. Khan and A.M. carried out proteomic experiments. O.S., S. Karimi and F.N. carried out immunohistochemical experiments and analyses. F.N., J.Z.W. and Q.W. carried out in vitro and in vivo experimentation. F.N., J.L., Y.M., V.P., A.C., R.H.-W., R.I.C., L.Y.L., C.Y.C. and B.M. contributed to the data processing and analyses. F.N., K.A., P.C.B., G.D.B., D.D.d.C., T.K. and G.Z. contributed to data interpretation. F.N. and A.M.W. organized the figures. F.N. and G.Z. wrote the first draft as well as subsequent revisions and the response to reviewers. All authors contributed to the final data interpretation and critical revision of the manuscript and approved the final version of the manuscript. G.Z. supervised all aspects of the study.
Corresponding author
Ethics declarations
Competing interests
D.D.d.C. and A.C. are listed as inventors on patents filed that are unrelated to this project. D.D.d.C. received research funding from Pfizer and Nektar therapeutics that was not related to this project. D.D.d.C is co-founder, shareholder and CSO of Adela, Inc. P.C.B sits on the Scientific Advisory Boards of BioSymetrics Inc. and Intersect Diagnostics Inc. G.T. has served on advisory boards of AbbVie, Bayer and BMS; received consulting fees from AbbVie, Bayer; received speaker fees from Medac and Novocure; received travel grants from Novocure, Medac and BMS; received research grants from Roche Diagnostics and Medac, all not related to this work.
Additional information
Peer review information Nature thanks Itay Tirosh, Roel Verhaak and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Individual datatype classification of meningiomas.
a, Violin plots showing the distribution of the normalized mutual information (MI) for each pairwise comparison of datatype. Median is shown as white dot. The number of total genes and number of genes with statistically significant (FDR < 5%) MI values are shown. Below this is a heatmap showing the consensus clustering of genes where MI was significant for at least one datatype pair. Rows represent a gene for which data exists from all data types. b,d,f, Unsupervised consensus hierarchical clustering of (b), 5,000 genes that show that highest median absolute deviation across expression values, (d), 10,000 CpG sites that show that highest median absolute deviation across β-values, (f), 1,000 genes that show that highest median absolute deviation across copy number ratios. Heatmap of consensus matrices with K = 6 groups (b,d,f) are displayed. Overall, six groups were most stable across all platforms. c,e,g, Kaplan Meier-plot displaying recurrence-free survival (RFS) distributions of unsupervised cluster assignments by (c) mRNA data, (e) DNA methylation data, (g) copy number data. The associations with outcomes are unique for the 6 cluster groups obtained on individual platform analyses. h, Average silhouette widths for unsupervised consensus hierarchical clustering from K = 2 to K = 10. The silhouette score is a measure of stability of number of groups. Higher scores indicate greater stability and robustness. Average silhouette width is highest at K = 4 subgroups. i, Alluvial plot demonstrating associations between WHO grade and integrative molecular groups defined in this study. j-l, Kaplan Meier-plot displaying recurrence-free survival (RFS) distributions of patients stratified and colored by molecular group assignments for WHO grade 1 tumors (j), WHO grade 2 tumors (k), and WHO grade 3 tumors (l).
Extended Data Fig. 2 Generalizability of the association of molecular groups with outcome.
a, Ensemble of Receiver Operating Characteristic (ROC) curves from 50 iterations of trained MG-versus-other models. Overlaid for each model is the mean Area Under the Curve (AUC) and its associated 95% confidence interval for samples in corresponding test sets. b, Heatmap showing results of single-sample Gene-Set Enrichment Analysis (ssGSEA) using mRNA data in an independent cohort of 80 meningioma samples. Each sample in the validation set was assigned a score for molecular groups 1, 2, 3 and 4 using gene-expression based signatures from the discovery cohort. MG designation was determined by highest scores from ssGSEA assignments. Unsupervised hierarchical clustering using scores from MG assignments revealed four distinctive groups of tumors w with 97% of samples having concordant assignment by maximal scores. Samples almost always showed high scores that were distinctive to only a single group, highlighting the robustness of classification in an independent cohort. c, Brier prediction curve for recurrence-free survival comparing molecular group to WHO grade in the generalization cohort. The models tested were those developed on the discovery cohort. Prediction errors are consistently lowest using molecular groups in comparison to the validation cohort. d, Kaplan Meier-plot displaying recurrence-free survival (RFS) distribution of patients stratified and colored by molecular group assignments for generalization set. P value reported is a Log Rank Test. Distributions are highly similar to those obtained in discovery cohort.
Extended Data Fig. 3 Most mutations are clonal in meningioma.
a, Lollipop plots showing the distribution of NF2 mutations by genomic regions within each molecular group. b, Mutational burden (nonsynonymous mutations per megabase) of meningiomas stratified by molecular groups in comparison to other TCGA solid cancers. Every dot represents a sample and horizontal lines are median number of mutations in each cancer type. Mutational burden in each cancer is ordered by percentile rank. Cancer types are ordered on the horizontal axis based on their median numbers of somatic mutations. Mutational burden of MG4 tumors is statistically higher than tumours in MG1-3 (P = 1.6 × 10−3, Kruskal Wallis test). c, Distribution of the number of mutations that are considered clonal per each patient sample (column). A total of 26% of tumors exhibited only clonal point mutations. In the median tumor, 75% of single nucleotide variants were clonal. d, Cancer cell fraction of all variants in each patient sample (columns) ordered as in (c). Variants are colored according to the classification in the legend. e, Cancer cell fraction of recurrent oncogenic driver mutations (columns). Variants are colored according to the classification in the legend.
Extended Data Fig. 4 Genomic disruptions differ among molecular groups.
a, Genome-wide copy-number alterations computed from whole-exome sequencing data. Arrangements of copy number profile are matched to the samples from mutation plot above. Only mutations that are relevant to discussion in text are shown.b, Boxplots showing the mRNA expression of NF2 stratified by molecular group. Each dot is a sample. Samples are colored by NF2 mutation status and shapes are according to NF2 deletion status by CNA. Some MG3 and MG4 meningiomas that are NF2 wildtype show silencing of NF2 expression. c, Boxplots comparing the mean methylation level of NF2 wildtype MG3 and MG4 meningiomas with high versus low NF2 expression using all probes (left), those mapping to the promoter region (middle), and those mapping to the gene body (right). d, Circos plot showing the landscape of interchromosomal gene rearrangements detected using a stringent threshold for conservative estimation of fusion events (unique spanning reads ≥ 25) in each molecular group. Total number of interchromsomal fusion in MG1, MG2, MG3 and MG4 are 2, 7,18, and 23, respectively.
Extended Data Fig. 5 Gene expression profiles of molecular groups.
a, Hierarchical clustering of the expression of genes from select pathways identified in Fig. 2a. Selected genes have been labeled. Redundancy of genes to pathways is shown in the side bar. b, Boxplots showing the results for estimates of immune and stromal infiltration by DNA methylation (LUMP score on left and methylCIBERSORT in middle) and somatic DNA alterations (right, ABSOLUTE score). c, Scatterplots comparing normalized enrichment scores between molecular groups using Gene Set Variation Analysis (GSVA). Each dot is a pathway. Shown at the top of each panel are Pearson correlations and associated 95%CI. MG2 tumors were divided into tumors that are driven by CNA (MG2-CNA) and tumors that are driven by mutations (MG2-Mut). Correlations were highest when comparing MG2 tumors driven by CNA to MG2 tumors driven but mutations (red box). d, Hierarchical clustering of normalized enrichment scores from (c) identifies MG2-CNA and MG2-Mut tumors as one coherent group. e, Boxplots comparing the activation of molecular signatures of proliferation between MGs. Statistical significance is denoted by asterisks.
Extended Data Fig. 6 Molecular characterization of patient derived cell lines.
a, t-distributed Stochastic Neighbor Embedding (tSNE) plot of genome-wide DNA methylation profiles of patient derived cell lines (red), to meningiomas (blue), and 2798 previously published tumors from 40 other brain tumor types58. b, Heatmap showing results of single-sample Gene-Set Enrichment Analysis (ssGSEA) using mRNA data from cell lines. Each cell line was assigned a score for molecular groups 1, 2, 3 and 4 using gene-expression based signatures from the discovery cohort. Molecular group designation was determined by highest scores from ssGSEA assignments. c, Gross morphological images of a representative MG4-xenografted mice. Extra axial tumor is outlined in dashed yellow lines. Compression on adjacent neural structures is evident after partial (middle panel) and complete (right panel) separation of meningioma from brain. d, Serial sections and immunostaining for MCM2 in representative MG4-xenograted mice. Scale bar is 2mm. Small areas of tumor that have invaded the brain can be seen staining for MCM2.
Extended Data Fig. 7 Proteomic and gene expression data converge to similar biology driving each molecular group.
a, Hierarchical clustering of normalized enrichment scores obtained by Gene-Set Variation Analysis (GSVA) using proteomic data (rows) and mRNA data (columns). b, Distribution of correlation of mRNA expression to protein abundance in all samples (grey), MG1 meningiomas (red), MG2 meningiomas (blue), MG3 meningiomas (green) and MG4 meningiomas (orange). Vertical line indicates overall median correlation across all samples (Spearman’s r = 0.279, 95%CI 0.273-0.284). c, Scatterplots comparing normalized enrichment scores by GSVA using gene expression (x-axis) and protein abundance (y-axis) stratified by MG classifications. Each dot represents a pathway. Pathways that are statistically significant and concordant by protein and mRNA data are colored green while those that are discordant are colored green. Pearson correlations and 95% confidence intervals are indicated at the top of each panel. d, Network of activated gene circuits by proteome data in N = 96 samples. Protein groups were ranked for each subtype by degree of differential expression. Gene-set enrichment analysis was performed on the ranked gene lists and enriched pathways are visualized using the EnrichmentMap plugin in Cytoscape App. Nodes represent pathways and edges represent shared genes between pathways. Pathways above horizontal line are up-regulated (red nodes) in each molecular group while pathways below horizontal line are down-regulated (blue nodes) in each molecular group.
Extended Data Fig. 8 Differences in genome-wide methylation across meningioma groups.
a, Hierarchical clustering of highly differentially methylated CpGs (absolute ∆β > 0.35, FDR < 0.05) between all meningiomas and healthy meninges. Annotations of molecular groups are on the right side of heatmap. b, Boxplots showing the distribution of β values for probes in (a) that are hypomethylated in healthy meninges (left) and hypermethylated in healthy meninges (right). Pairwise comparisons in each boxplot are statistically significant (p < 0.05), unless explicitly stated otherwise (ns, not significant). c, Boxplots showing the distribution of using epigenetic mitotic clocks with epiTOC model (left), epiTOC2 model (middle), and HypoClock model (right). Pairwise comparisons in each boxplot are statistically significant (p < 0.05), unless explicitly stated otherwise (ns, not significant). d, Number of unique and overlapping probes that are differentially methylated (absolute ∆β > 0.1, FDR < 0.05) when comparing each molecular group to healthy meninges. e, Scatterplots comparing master regulator transcription factor expression with average β values at sites enriched for the motif of that transcription factor. Samples are colored according to molecular group. Pearson correlation with 95% confidence intervals are reported. Hypomethylation at motifs of immunological-lineage-specific transcription factors such as PU.1, RUNX1/2 and IRF5/8 were enriched in immunogenic (MG1) tumors (P = 1.05 × 10−8, hypergeometric test) and associated with enhancer hypomethylation. Similarly, master regulators of cell proliferation such as MYBL2, LHX4, and FOXM1 were hypomethylated in proliferative (MG4) tumors and associated with increased abundance of these transcription factors (P = 1.24 × 10−3, hypergeometric test).
Extended Data Fig. 9 Meningiomas show low within patient variation of expression and copy number profile.
a, Pairwise correlations of expression profiles of all cells ordered by hierarchical clustering. Each cell is annotated to tumor of origin from Fig. 4a and cluster assignments from Fig. 4b at top and side bars. b, Inferred genome-wide copy number variations of single nuclei of healthy meninges (reference, top panel), immune cells (middle panel), and neoplastic cells (bottom panel). Sample and cluster annotation are shown on the left. The copy number plot of these tumors are homogenous and subclones of cells within tumors with distinct copy number profiles are not common. Annotation to patient of origin and cluster on the left of each heatmap. c, Scatterplots showing the relationship between arm-level CNA inferred by snRNA-seq (x-axis) to matched CNA by bulk whole exome sequencing (y-axis). Two representative samples are shown.
Extended Data Fig. 10 The transcriptome of MGs is shaped by the expression profiles from both neoplastic and non-neoplastic cells.
a, Bubble plot showing the expression of lineage specific markers for distinct cell types. b, Stacked barplot showing the relationship of samples to clusters. Samples are colored by patient of origin as in Fig. 4a. Barplot to the right shows the number of cells within each cluster. c, The top heatmap shows hierarchical clustering results of single cells by molecular group scores. Each cell was scored for the bulk signature of each molecular group and scores were compared to a permuted random gene set. Shown are cells with at least one score with FDR < 0.2. Scores were scaled such that the sum of all scores for each cell is equal to one. Below is a matched heatmap showing the number of genes detected for each MG signature in each cell. In a subset of cells, low scores are associated with low detection rate of genes (yellow and pink boxes). d, Stacked barplot showing the distribution of immune versus non-immune cells across molecular groups (left) and cycling versus non-cycling neoplastic cells across molecular groups (right) to clusters. Samples are colored by molecular group of tumor as in Fig. 4d. e, Barplot showing the total number of cells that are immune versus non-immune (left) and cycling versus non-cycling (right) by MG status of tumor of origin. f, Boxplots comparing the cell type composition of bulk RNA seq samples after deconvolution using single cell RNA-seq signatures. Pairwise comparisons in each boxplot are statistically significant (p < 0.05), unless explicitly stated otherwise (ns, not significant). g. Heatmap showing the expression of marker genes for single cell clusters (determined by CIBERSORTx) in bulk RNA seq data. Each column represents one tumor. Rows are designated marker genes for each cluster. Tumors were partitioned into 4 partitions by consensus k-means clustering with samples and gene sets clustered by hierarchical clustering using Pearson distance metric.
Extended Data Fig. 11 Discrete and continuous patterns of variability can be identified in meningioma.
a, Hierarchical clustering of similarities between NMF programs. Top panel indicates Pearson correlations between number of mitochondrial and ribosomal genes detected with NMF scores for each program. A cluster of programs (dashed lines) showed positive correlation with the expression of mitochondrial and ribosomal genes (confirmed by manual inspection). These programs were considered to be reflective of technical artifacts and not included in subsequent analyses. b, Violin plots showing the distribution of activation scores for NMF programs across MGs. c, Side-by-side tSNEs showing the relationship of discrete clustering results with activation scores of each NMF program. Shown are four representative samples. Activation scores of cell cycle program are closely associated with discrete clusters, whereas scores of metabolism, inflammatory, and mesenchymal program are not associated with discrete clusters. d, Heatmap showing the average expression of genes defining NMF programs (annotated to left) in representative sample CAM_0071. Cells are ranked and ordered according to the activation score of the metabolism program. There is a continuous pattern of gene expression variability in these programs.
Extended Data Fig. 12 Graphical summary of findings.
Shown is a schematic representation that summarizes the major molecular findings and conclusions of our study: unsupervised consensus clustering combining DNA copy number, DNA methylation, and mRNA sequencing data revealed four robust groups of tumors with prototypical biology and distinct clinical outcomes.
Supplementary information
Supplementary Information
This file contains the Supplementary Discussion, Supplementary References and Supplementary Figs. 1, 2.
Supplementary Tables
This file contains Supplementary Tables 1, 2, 3 and 6.
Supplementary Tables
This file contains Supplementary Tables 4, 5, 7 and 8.
Source data
Rights and permissions
About this article
Cite this article
Nassiri, F., Liu, J., Patil, V. et al. A clinically applicable integrative molecular classification of meningiomas. Nature 597, 119–125 (2021). https://doi.org/10.1038/s41586-021-03850-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-021-03850-3
- Springer Nature Limited
This article is cited by
-
Clinical implications of DNA methylation-based integrated classification of histologically defined grade 2 meningiomas
Acta Neuropathologica Communications (2024)
-
Chronic hyperglycemia and intracranial meningiomas
BMC Cancer (2024)
-
Multicenter radio-multiomic analysis for predicting breast cancer outcome and unravelling imaging-biological connection
npj Precision Oncology (2024)
-
MerlinS13 phosphorylation regulates meningioma Wnt signaling and magnetic resonance imaging features
Nature Communications (2024)
-
NF2: An underestimated player in cancer metabolic reprogramming and tumor immunity
npj Precision Oncology (2024)