Abstract
The cation channel of sperm (CatSper) is essential for sperm motility and fertility1,2. CatSper comprises the pore-forming proteins CATSPER1–4 and multiple auxiliary subunits, including CATSPERβ, γ, δ, ε, ζ, and EFCAB91,3,4,5,6,7,8,9. Here we report the cryo-electron microscopy (cryo-EM) structure of the CatSper complex isolated from mouse sperm. In the extracellular view, CATSPER1–4 conform to the conventional domain-swapped voltage-gated ion channel fold10, following a counterclockwise arrangement. The auxiliary subunits CATSPERβ, γ, δ and ε—each of which contains a single transmembrane segment and a large extracellular domain—constitute a pavilion-like structure that stabilizes the entire complex through interactions with CATSPER4, 1, 3 and 2, respectively. Our EM map reveals several previously uncharacterized components, exemplified by the organic anion transporter SLCO6C1. We name this channel–transporter ultracomplex the CatSpermasome. The assembly and organization details of the CatSpermasome presented here lay the foundation for the development of CatSpermasome-related treatments for male infertility and non-hormonal contraceptives.
Similar content being viewed by others
Main
Fertilization—the union of a sperm and an egg—is a fundamental biological process that seeds a new life. Several key steps during fertilization, including sperm hyperactivation, acrosome reaction, and sperm–egg fusion, are regulated by Ca2+ signalling11,12. The sperm-specific cation channel CatSper, which is primarily localized to the principal piece of mature spermatozoa flagellum, is responsible for multiple Ca2+-dependent physiological responses during fertilization2,13. CatSper-mediated Ca2+ signalling initiates a tyrosine phosphorylation cascade that controls sperm motility14. CatSper also has a central role in sperm chemotaxis15,16. For example, CatSper mediates the Ca2+ influx induced by the steroid sex hormone progesterone in human sperm17,18.
CatSper contains the largest number of subunits of any known ion channel1,3,4,5,6,7,8,9. Among the identified components, CATSPER1–4 share relatively low sequence identities (about 20%) among themselves and with the related voltage-gated ion channels (VGICs). None of the auxiliary subunits of CatSper shares sequence homology with those of VGICs, suggesting that CatSper has a unique assembly mechanism.
Mutations in CatSper impair male fertility19,20,21,22,23,24,25,26. Deletion of CATSPER1, 2, 3, or 4 in mice leads to abnormal sperm motility and male sterility due to the loss of sperm hyperactivation1,27,28,29. In humans, the expression of CATSPER1 is substantially reduced in subfertile individuals with deficient sperm motility30. CATSPER2 mutations are associated with human asthenoteratozoospermia, which cause non-syndromic male infertility22. CatSper has thus been implicated as a potential target for the treatment of male infertility. Conversely, it is also an attractive target for the development of non-hormonal contraceptives31,32.
Despite the physiological importance of CatSper, none of the ten identified subunits has been structurally characterized, to our knowledge. To elucidate the assembly of CatSper, we have isolated the endogenous CatSper complex from mouse sperm and determined its cryo-EM structure. Our results show that CatSper is a channel–transporter ultracomplex, which we name the CatSpermasome.
Cryo-EM analysis of CatSper
To prepare a biochemically functional protein sample, we sought to purify the CatSper complex from its endogenous source. We generated transgenic knock-in mice by inserting a 3×FLAG–GFP tag into the N terminus of CATSPER1 (Extended Data Fig. 1a). We detected GFP fluorescence in the principal piece of the spermatozoa from knock-in but not wild-type mice (Extended Data Fig. 1b, c). CatSper was purified from the testis and epididymis of mature male knock-in mice for cryo-EM analysis (Extended Data Fig. 1d–f). The presence of CatSper components was confirmed by mass spectrometry (MS) analysis (Extended Data Fig. 2). Details of protein purification, data acquisition, and image processing can be found in the Methods.
About 560,000 selected particles yielded an EM reconstruction at an overall resolution of 2.9 Å (Fig. 1, Extended Data Fig. 3, Extended Data Table 1). The characteristic micelle enabled the distinction of a large extracellular region, a transmembrane region surrounded by detergent micelles, and an intracellular region (Fig. 1). The excellent density supported unambiguous subunit assignment for all known components, except for the two small cytosolic subunits CATSPERζ and EFCAB9. After assignment of the known subunits, several extra densities indicated the presence of uncharacterized components (Fig. 1a, b). Among them, TMEM262, hereafter referred to as CATSPERη, and SLCO6C1 were identified from the EM map and MS analyses. TMEM249 is likely to be another component of CatSper. A cytosolic entity and a single transmembrane helix remain unassigned. A detailed summary of model building is presented in the Methods and Extended Data Table 2.
a, The overall cryo-EM map of the CatSpermasome shown in three side views. CATSPER1–4 are coloured yellow, orange, pale green and cyan; CATSPERβ, γ, δ and ε are coloured pink, blue, light blue and green, respectively. The previously uncharacterized subunits SLCO6C1 and CatSperη are coloured slate and purple, respectively. The map that probably belongs to TMEM249 is coloured wheat. The cytosolic region that may belong to CATSPERζ and EFCAB9 is coloured light cyan. The transmembrane map near CatSperη coloured salmon and the cytosolic map coloured grey are not assigned. The detergent micelle is shown in transparency. For visual clarity, a composite map from five reconstruction maps was used (see Methods, Extended Data Fig. 3). b, Top views of the extracellular (left) and transmembrane (right) regions of the CatSpermasome. CATSPER1–4 are highlighted by a dotted circle. The map figures were generated in ChimeraX with a contour level of 0.4. c, Local resolution map of the final overall reconstruction estimated by cryoSPARC and generated in Chimera.
Overall assembly of the CatSpermasome
The resolved complex has dimensions of approximately 220 Å × 170 Å × 240 Å. The central channel domain constituted by CATSPER1–4 adopts the canonical VGIC architecture. The single C-terminal transmembrane helix in each auxiliary subunit (CATSPERβ, γ, δ and ε), attaches to the adjacent voltage-sensing domain (VSD) of CATSPER4, 1, 3 and 2, respectively, and the bulky extracellular domains cover the entire channel domain (Fig. 1a, b, Supplementary Video 1).
The channel domain displays a pseudo-four-fold symmetry, but the overall transmembrane region has no symmetry because of the presence of other transmembrane components (Fig. 1b). On one side, SLCO6C1 is close to CATSPERε and CATSPER2. On the other side, there are three transmembrane components adjacent to CATSPERβ and CATSPER4: CATSPERη, a protein that is probably TMEM249, and an uncharacterized component with a transmembrane helix (Fig. 1b).
The cytosolic region consists of two separate but interacting parts. One entity, composed of characteristic α-helices, is likely to belong to the subcomplex of CATSPERζ and EFCAB9, as the two EF-hand containing lobes of EFCAB9 can be well fitted into the map (Extended Data Fig. 4). The identity of the other part that is sandwiched by the elongated S6 segments of CATSPER2 and CATSPER3 remains unknown owing to limited local resolution.
The heterotetrameric CATSPER1–4 channel
CATSPER1–4, with S1–S4 of each subunit forming the VSD and S5–S6 enclosing the ion-conducting pore, are arranged counterclockwise in the extracellular view with the conventional domain-swapping of the pore domain and VSD from the adjacent protomer (Fig. 2a, Extended Data Fig. 5). Similar to voltage-gated calcium and sodium (Cav and Nav) channels, the selectivity filter is supported by two pore helices, P1 and P2, but the extracellular loops that connect S5 and P1 (L5) and P2 and S6 (L6) are substantially shorter and lack any glycosylation modification (Extended Data Fig. 6a).
a, Left, the overall structure of the heterotetrameric CATSPER1–4 is assembled anticlockwise in the extracellular view. Right, the ion permeation path of the channel calculated by HOLE is illustrated with coloured dots based on the pore radius (red, <1.5 Å; green, 1.5–2.3 Å; purple, >2.3 Å). b, The overall structure of the selectivity filter and pore helices of CATSPER1–4 is similar to that of Cav1.1 (grey). CATSPER1–4 are superimposed on repeats II, I, IV and III of Cav1.1, respectively. c, A closed channel. The corresponding pore radii along the permeation path are shown. Insets, the closed intracellular gate is formed by two hydrophobic rings. SF, selectivity filter. d, The VSDs of CATSPER1–4 (left to right) adopt various conformations. The Cα atoms of the residues in positions R1–R6 are shown as spheres. Positively charged residues, An1 residues, and CTC residues are shown as sticks. Polar or acidic residues that participate in gating charge coordination and non-Arg/Lys residues in positions R1–R6 are shown as lines.
The conformation of the selectivity filter of CATSPER1–4 is similar to that of Cav1.1 (Fig. 2b). The four key residues that guard the entrance of the selectivity filter are Asp536, Asp295, Asp227, and Asp237 from CATSPER1, 2, 3 and 4, respectively. Sequence alignment of the selectivity filter and the pore helices in CATSPER1–4 and Cav1.1 indicates a common Thr-X-Asp/Glu-X-Trp motif (Extended Data Fig. 6b). Densities that belong to cations can be seen in the vestibule of the selectivity filter (Extended Data Fig. 5b).
Along the ion permeation path, the selectivity filter and the intracellular gate represent two constriction sites (Fig. 2c). The intracellular gate, with a radius of less than 1 Å, is sealed by two layers of hydrophobic residues from the S6 segments. Therefore, the current structure represents a closed channel.
Distinct conformations of the four VSDs
CatSper is gated by voltage, albeit with a lower sensitivity than Cav channels33. Sequence alignment of the VSDs of CATSPER1–4 (VSD1–4) indicates they have conserved elements for voltage sensitivity, including the positively charged residues Arg/Lys on the S4 segments, the charge transfer centres (CTCs), and a conserved acidic residue Asp/Glu on S2 (denoted An1 in Extended Data Fig. 6c). The CTC of each VSD includes an acidic residue Glu (An2) on S2, an occluding residue Phe/Tyr (F), and an adjacent conserved acidic residue on S334. Whereas there are usually 4–6 positively charged residues on S4 of Cav channels (R1–R6), the number of positively charged residues on S4 of CATSPER1–4 vary substantially, with seven (R0–R6) in CATSPER1, four (R2–R5) in CATSPER2, and only two each in CATSPER3 (R4–R5) and CATSPER4 (R3–R4) (Fig. 2d). The S4 of VSD4 is mostly relaxed from the 310 helix to the α-helix and the occluding residue on S2 of CATSPER4 is replaced by a Leu. These characteristic VSD features are reminiscent of NALCN (a Na+ leak channel that also has relatively weak voltage sensitivity35).
The four VSDs have different conformations (Fig. 2d). When the four VSDs are superimposed relative to CTC and An1, the S4 segments vary and the heights of the Cα atoms of the corresponding positively charged residues are CATSPER3>CATSPER2>CATSPER4>CATSPER1 (Extended Data Fig. 6d). The R4 residues in CATSPER3 and CATSPER2 are located above the occluding residue on S2, whereas those in CATSPER4 and CATSPER1 are below it (Fig. 2d). Despite this, the exact functional state of each VSD has yet to be defined.
The auxiliary subunits CATSPERβ, γ, δ, ε
All the extracellular segments in CATSPERβ, γ, δ and ε contain multiple modular domains (Extended Data Fig. 7a, b). CATSPERδ lacks an N-terminal domain (NTD) that is conserved in the other three subunits. Following the NTD, all four subunits consist of a seven-bladed β-propeller domain, an Ig-like domain, and a stem domain before the transmembrane helix. CATSPERβ has an extra head domain and CATSPERε has an additional NTD2 domain. All these extracellular domains (ECDs) are mainly composed of β-strands and modified with multiple glycosylation sites and disulfide bonds (Extended Data Fig. 8, Extended Data Table 2).
The ECDs are stabilized on the distal region through extensive interactions (Fig. 3a, Extended Data Fig. 7c, d). The lack of interaction between the stem domains leaves four side openings that allow free passage of ions to the selectivity filter of the channel domain. Despite the large sizes of the four auxiliary subunits, only the stem and transmembrane domains are involved in the association with the channel domain.
a, CATSPERβ, γ, δ and ε interact with CATSPER4, 1, 3 and 2, respectively. CATSPERβ, γ, δ and ε are shown in surface representation and CATSPER1–4 as cartoon. Details from the square boxes are presented in b. b, Close-up views of the interactions between the stem domains of CATSPERδ (left) and CATSPERγ (right) and the adjacent channel subunits. The key residues that mediate the interactions in the interface are shown as sticks. Hydrogen bonds and cation-π interactions are indicated by red dashed and brown dotted lines, respectively.
The transmembrane helices of CATSPERβ, γ, δ and ε interact with the S0 and S2 segments of CATSPER4, 1, 3 and 2, respectively, through extensive van der Waals contracts and limited polar interactions (Fig. 3a, Extended Data Fig. 7e). In addition, the stem domains of CATSPERδ and CATSPERγ contribute to the specificity of the pairwise association between the channel subunits and the auxiliary subunits. For instance, Arg918 from the stem domain of CATSPERγ inserts into the aqueous outer cavity of the VSD1 and forms hydrogen bonds with Glu377 and Glu381 in CATSPER1 (Fig. 3b). The stem domain of CATSPERδ not only forms hydrogen bonds with VSD3, but also interacts closely with the L5 and L6 loops of CATSPER2 through hydrogen bonds and cation-π interactions. By contrast, the stem domains of CATSPERε and CATSPERβ are at a distance from their adjacent VSDs (Fig. 3a, Extended Data Fig. 7e).
Novel components of the CatSpermasome
We assigned SLCO6C1 (close to VSD2) on the basis of the MS results and the characteristic density of the 12-TM major facilitator superfamily (MFS) fold (Figs. 1a, 4a, Extended Data Fig. 2). In support of this assignment, a comparative proteomic study showed a fivefold decrease in SLCO6C1 expression in Catsper1-null mice9. In addition, there was an extra density for the unique extracellular Kazal domain of SLCO6C1 (Fig. 4b, Extended Data Fig. 9a). We identified a predicted glycosylation site (N535) in the Kazal domain, further validating the protein identity (Fig. 4c).
a, Overall structure of the mouse CatSpermasome. SLCO6C1, CATSPERη and their neighbouring subunits are coloured as in Fig. 1. Other subunits are coloured light grey. b, SLCO6C1 interacts with CATSPERε through its C domain and Kazal domain. The overall structure of SLCO6C1 adopts a canonical MFS fold, with the N domain, C domain and extracellular Kazal domain coloured light blue, slate, and cyan, respectively. The sugar moieties of an identified glycosylation site are shown as sticks. c, Interaction between the Kazal domain of SLCO6C1 and the stem domain of CATSPERε. Residues that may be involved in the specific interaction are shown as sticks. Hydrogen bond and cation-π interactions are indicated by red dashed and brown dotted lines, respectively. Inset, electron density map (σ = 4.0) of a segment around the glycosylation site N535 in the Kazal domain. d, CATSPERη interacts with the stem domain of CATSPERε through its extracellular loops and transmembrane regions. e, Close-up views of the interaction interfaces between CATSPERη and CATSPERε in the extracellular region (left) and the transmembrane region (right).
SLCO6C1 is a testis-specific organic anion transporter protein (belonging to the OATP6 family) whose structure and physiological function have not been well studied36. In humans, the close related homologue SLCO6A1 is also predominantly expressed in testis37. SLCO6C1 contains three parts: the N domain, the C domain and the extracellular Kazal domain between TM9 and TM10 of the C domain (Fig. 4b, Extended Data Fig. 9b). The structure of SLCO6C1 adopts an outward-facing conformation, forming an angle of approximately 22° (Extended Data Fig. 9b). SLCO6C1 interacts with CATSPERε through two interfaces. In the transmembrane region, TM9 interacts with the transmembrane domain of CATSPERε, probably through hydrophobic interactions. In the extracellular region, the Kazal domain of SLCO6C1 associates with the stem domain of CATSPERε through potential hydrogen bonds and cation-π interactions (Fig. 4c). The glycosylation site in SLCO6C1 and the residues that are involved in its interactions with CATSPERε are highly conserved among mouse, rat, and human, predicting similar assembly in these species (Extended Data Fig. 9c).
Close to VSD4, extra densities for six transmembrane helices were resolved (Fig. 1b, Extended Data Fig. 10a). The middle three transmembrane helices were assigned to CATSPERη based on high-quality EM density and MS results (Extended Data Figs. 2, 10b). The loop between TM1 and TM2 and the C-terminal loop of CATSPERη constitute a platform that supports the stem domain of CATSPERβ (Fig. 4d, e). In addition, TM2 and TM3 of CATSPERη form extensive hydrophobic interactions with the transmembrane domain of CATSPERβ, further stabilizing the CatSpermasome.
While the single transmembrane density near CATSPERη is too small to be identified, the densities of two transmembrane helices connected to a cytosolic globular domain near CATSPERη probably belong to TMEM249 (Extended Data Figs. 2, 10a). A predicted structure of TMEM249 fits well into the density (Extended Data Fig. 10c). Consistent with this analysis, TMEM249 is also localized to the principal piece of sperm by immunofluorescence detection (Extended Data Fig. 11). TMEM249 contributes to the assembly of the CatSpermasome by interacting with CATSPERη and VSD4 in the transmembrane and cytosolic regions, respectively (Extended Data Fig. 10d).
Perspectives
Our structure offers an implication for the gating mechanism of CatSper. The S6 segments of CATSPER2 and CATSPER3 extend more than 30 Å into the cytosol, held by an unidentified cytosolic component (Fig. 1a, Extended Data Fig. 4a). These interactions may be further stabilized by the attachment of the CATSPERζ–EFCAB9 subcomplex. The cytosolic components may undergo conformational changes upon stimulation (for example by Ca2+ and/or intracellular alkalinization), which in turn trigger the movement of the S6 segments and opening of the channel. Structures of CatSper in the open conformation and other intermediate states are required to examine this potential gating mechanism.
The discovery of the previously unknown components of the CatSpermasome is serendipitous. Although little was known about the physiological functions of these components, the current structure implies that they are important for the assembly and stability of the CatSpermasome.
The identification of an organic anion transporter in the CatSpermasome is particularly notable. SLCO6C1 might mediate the transport of steroid hormones such as dehydroepiandrosterone, as reported in its rat counterpart38. The transport of the substrate through SLCO6C1 is likely to be mediated by an alternating access mechanism that could couple to the opening of CatSper. Although functional channel–transporter complexes have been reported39, their structural observation has been limited to KATP, the complex between the potassium channel Kir6.2 and the ABC transporter module SUR140. However, SUR1 acts only as an ADP sensor of KATP. Whether SLCO6C1 in the CatSpermasome has a regulatory role as a sensor or works through a distinct coupling mechanism remains to be investigated.
In summary, the structure reveals the overall assembly of the CatSpermasome. Future characterization of the physiological and pathological importance of SLCO6C1, CATSPERη and TMEM249, and the identification of the unassigned components of the CatSpermasome, will contribute to the mechanistic understanding of sperm fertility.
Methods
Animals
All animal maintenance and experimental procedures were conducted in accordance with institutional guidelines, and all animal studies were approval by the Institutional Animal Care and Use Committee (IACUC) of Westlake University, Hangzhou, China. Male and female mice (strain: C57BL/6J) of 2–5 months old were used for experiments in this study. Mice were maintained at strict barrier facilities with macroenvironmental temperature and humidity ranges of 20–26 °C and 40–70%, respectively. Mouse rooms had a 12 h light/12 h dark cycle. The housing conditions were closely monitored and controlled. No statistical methods were used to predetermine sample size. The experiments were not randomized and the investigators were not blinded to allocation during experiments and outcome assessment.
Transgenic mice generation and genotyping
The 3×FLAG-EGFP-TEV-CATSPER1 knock-in mice were generated by CRISPR–Cas9 gene editing. Cas9 protein, two gRNAs (gRNA1: CAGTCATGGATCAATCTTCA, gRNA2: AGATTGATCCATGACTGTCT) and a donor vector containing the DNA sequence of 3×FLAG-EGFP-TEV before the first exon of the Catsper1 gene were injected into fertilized eggs. The embryos were transferred to recipient female mice to obtain F0 mice. The genotype of knock-in mice was confirmed by PCR using two pairs of primers (F1: TTTCGTCAACATGGAAGGCTG, R1: TTGAAGTCGATGCCCTTCAG; F2: GGAAGATTCCGAGAGAAGAGTCAG, R2: TTGATGGCTTGGGTCTAAGCTAC) and sequencing (Extended Data Fig. 1).
In vitro fertilization
In vitro fertilization (IVF) experiments were carried out according to standard protocols1,41. Eggs from mature knock-in (KI)/+ or KI/KI female mice (9–12 weeks old) were synchronized with 10 units of PMSG (pregnant mare serum gonadotropin) followed by 10 units of HCG (human chorionic gonadotropin) 60.5 h and 13 h before collection, respectively. Sperm from the epididymis of KI/KI male mice were collected and capacitated in vitro for 1 h before use. Eggs were incubated with the sperm for 4–6 h at 37 °C, and unbound sperm were washed away. After overnight incubation at 37 °C, two-cell-stage or four-cell-stage embryos were collected and transferred to recipient female mice. The genotypes of the offspring mice were confirmed by PCR.
Confocal fluorescence microscopy imaging
For fluorescence imaging of sperm flagellum, fresh mouse sperm taken from the epididymis were immediately washed with 1× PBS (phosphate-buffered saline). After centrifugation at 1,000g for 2 min at room temperature, the sperm cells were resuspended in 1× PBS to an appropriate density and the drops of sample were smeared evenly onto clean glass slides and dried in the air at room temperature for 2 h. For immunofluorescence, the samples were fixed with 4% (v/v) paraformaldehyde in PBS buffer at ambient temperature for 1 h. After fixation, the slides were rinsed and the sperm cells were permeabilized with 2% (v/v) Triton X-100 and 1% (w/v) bovine serum albumin in PBS buffer for 30 min. The slides were incubated with primary antibodies (1:50, Abmart) at 4 °C overnight and then rinsed in PBS buffer (4 × 5 min). The slides were subsequently incubated with goat anti-rabbit IgG AF594 secondary antibody (same as Alexa Fluor 594, 1:500, Abmart) at room temperature for 2 h and then rinsed in PBS buffer (4 × 5 min) to remove the unbound secondary antibodies. Finally, the sperm nuclei were dyed with Hoechest 33342 (Life Technologies Corporation) and the slides were sealed for confocal microscopy imaging.
Images were acquired using a confocal fluorescence microscope (FV3000) at an objection lens magnification of 40.0×. EGFP fluorescence of knock-in mice and the auto-fluorescence signal in the middle piece of both wild-type and knock-in mice were detected in dark-field mode, with 488 nm laser wavelength for excitation and the detection wavelength at 500–540 nm. For immunofluorescence samples, second antibody and Hoechst signals were detected in 570–620 nm and 430–470 nm wavelength ranges, with 561 nm and 405 nm wavelengths for excitation, respectively. All fluorescence signals were merged onto the brightfield image.
Endogenous purification of the mouse CatSpermasome
All purification steps were carried out at 4 °C unless otherwise stated. Testis and epididymis tissue from about 50 KI/+ or KI/KI mice were homogenized in buffer containing 0.3 M sucrose, 25 mM HEPES-Na, pH 7.0, 2 mM EDTA and protease inhibitors including 2 mM phenylmethylsulphonyl fluoride (PMSF), 2.6 μg/ml aprotinin, 1.4 μg/ml pepstatin and 10 μg/ml leupeptin. The homogenate was centrifuged at 200,000g for 1 h. The pellet was solubilized at 4 °C for 2 h in buffer containing 25 mM HEPES-Na, pH 7.0, 400 mM NaCl, 1% (w/v) lauryl maltose neopentyl glycol (LMNG, Anatrace), 0.12% (w/v) cholesteryl hemisuccinate tris salt (CHS, Anatrace), 2 mM EDTA and protease inhibitors. During this process, glutathione S-transferase (GST)-fused GFP nanobodies were added. After centrifugation at 20,000g for 1 h, the supernatant was applied to glutathione sepharose 4B resin (GS4B, GE Healthcare). The protein-loaded resin was washed with 25 mM HEPES-Na, pH 7.0, 400 mM NaCl, 0.015% GDN (Anatrace), 1 mM EDTA and 2 mM PMSF. The target protein complex was eluted with buffer containing 13 mM reduced glutathione, 100 mM HEPES-Na, pH 8.0, 150 mM NaCl, 0.015% GDN and 1 mM EDTA. The eluent was then loaded to anti-FLAG G1 affinity resin (GenScript). The resin was washed with buffer containing 25 mM HEPES-Na, pH 7.0, 400 mM NaCl, 0.015% GDN, 1 mM EDTA and 2 mM PMSF. The protein was then eluted with buffer containing 25 mM HEPES-Na, pH 7.0, 150 mM NaCl, 0.015% GDN, 1 mM EDTA and 250 μg/ml FLAG peptide. The eluent was concentrated and cross-linked using 2 mM bissulfosuccinimidyl suberate (BS3) (Thermo Fisher Scientific) at 4 °C for 1.5 h and then quenched by 25 mM Tris-HCl, pH 8.0. The sample was then submitted to size-exclusion chromatography (Superose 6, 10/30, GE Healthcare) in buffer containing 25 mM HEPES-Na, pH 7.0, 150 mM NaCl, 0.015% GDN and 1 mM EDTA. The fractions containing the CatSpermasome were pooled for electron microscopy analysis or mass spectrometric analysis.
Cryo-EM sample preparation and data acquisition
For cryo-EM sample preparation, aliquots (3.5 μl) of the protein sample were loaded onto glow-discharged lacey carbon Cu grids coated with ultrathin carbon film (Ted Pella Inc., 400 mesh). The grids were blotted for 8 s after waiting for 60 s and immersed in liquid ethane using Vitrobot (Mark IV, Thermo Fisher Scientific) under 100% humidity at 8 °C. The imaging system comprised of a Titan Krios operating at 300 kV, a Gatan K3 Summit detector, and a GIF Quantum energy filter with a 20-eV slit width. Movie stacks were automatically acquired in super-resolution mode (81,000× magnification) using AutoEMation42, with a defocus range from −1.5 μm to −2.5 μm. Each stack was exposed for 2.56 s with 0.08 s per frame, resulting in 32 frames and approximately 50 e−/Å2 of total dose.
Image processing
A flowchart of the data processing process can be found in Extended Data Fig. 3f. Using MotionCor243, 16,648 movie stacks were motion-corrected with twofold binning. Patch-based CTF parameters of the dose-weighted micrographs (1.077 Å per pixel) were determined by cryoSPARC v344, and around five million particles were picked iteratively. 2D classification enriched images of good classes, resulting in a total of 1.2 million particles being selected with a box size of 400 pixels. Following an ab-initio reconstruction for initial map generation and 3D classification, non-uniform refinement jobs45 yielded a reconstruction at an overall resolution of 2.9 Å (Map 1). A local refinement job further improved the map quality of the extracellular domains (ECD) with an ECD mask. To improve the map quality of the cytosolic and peripheral transmembrane regions, we exported the subtracted refined particles by applying four different masks near the transmembrane regions of CATSPERβ, δ and ε, and the cytosolic region, respectively. We then performed multiple 3D classifications without orientation searching using RELION 3.0 (--skip-align)46,47. After two rounds of classification for each region with class number K = 5, the best classes were selected for subsequent non-uniform refinement jobs in cryoSPARC, yielding four reconstructions at overall resolutions of 3.4–3.5 Å with improved local map quality (Map 2–Map 5). The resolution of the masked region in each map was estimated by applying the same mask used for skip align 3D classification. Map resolutions were determined by the gold-standard Fourier shell correlation (FSC) 0.143 criterion using Phenix.mtriage48. All the map figures were generated in ChimeraX49 or Chimera50.
Model building and refinement
Model building was combined with multiple strategies including de novo building, homologue modelling, and structure prediction and docking. The majority of the model was built against Map 1, and Map 2–Map 5 were used for facilitating the modelling and analysis of peripheral transmembrane regions and the cytosolic regions. All these maps as well as a composite map shown in Fig. 1 have been deposited in the Electron Microscopy Data Bank under the same accession code.
For modelling of CATSPER1–4, initial models were first generated by homologue modelling using Cav1.1 (PDB: 6JP5) or Nav1.4 (PDB: 6AGF) as template. The initial models were then fit into the transmembrane channel region of the electron density map in Chimera50. Except for the S3 and S4 segments of CATSPER2, whose map quality is moderate, densities of the channel domains are of high quality and allow accurate side chain assignment. CATSPER1–4 were distinguished by side chain assignments as well as the various lengths of the L5 and L6 loops. The structures were then manually adjusted in Coot51. The ion permeation path of the channel was calculated by HOLE52.
The auxiliary subunits CATSPERβ, γ, ε were built de novo in Coot. The modelling of CATSPERδ was started from a homologue model using CATSPERε as template. The average resolution of the large extracellular region is near 3 Å, which allows unambiguous side chain assignment. Sequence assignment was guided by bulky residues such as Phe, Tyr, Trp and Arg. Multiple glycosylation sites and disulfide bonds were identified in CATSPERβ, γ, δ and ε, which in turn helped to validate the accuracy of the modelling.
For modelling of SLCO6C1 (Uniprot ID: Q8C0X7), a starting AI-facilitated prediction model was generated by tFold (https://drug.ai.tencent.com/). The model was fitted into the density map in Chimera and then manually adjusted in Coot. The bulky residues are mostly consistent with the traceable side chains of the density map. A protruding density that belongs to sugar moieties in the extracellular Kazal domain is close to a predicted glycosylation site (N535), further validating the assignment.
For modelling of CATSPERη (Uniprot ID: D3Z338), a predicted model was generated by tFold. The model of the three-transmembrane-helix bundle can be fitted well into the density near the transmembrane and stem domains of CATSPERβ. Out of the six clearly resolved transmembrane helices in the local region, the density quality of the three helices that belong to CATSPERη is the highest (probably owing to its close contact with the stem domain of CATSPERβ), which allows unambiguously bulky side chain assignment. The model was then manually adjusted in Coot.
After building of CATSPERη, the remaining nearby densities were not continuous, with one single transmembrane helix in one side and another two helices in the other side of CATSPERη, suggesting that they may belong to distinct components. Notably, the densities of the two helices are connected to a cytosolic globular density that contains a discernible β-sheet. We selected all the possible transmembrane protein targets from the MS list and predicted their structures by tFold. Among them, the predicted structure of TMEM249 (Uniprot ID: A0A2R8VHF7), which contains two N-terminal transmembrane helices and a C-terminal cytosolic domain, can be fitted well into the density. Therefore, TMEM249 is highly likely to be the third newly identified component, although the side chains cannot be assigned. The remaining single helix is too small to be confidently identified. TMEM89, TMEM190 and TMEM210, which are testis-specific small proteins with single predicted transmembrane helix, are potential candidates. Currently, a poly-Ala helix was built for this density.
The cytosolic region is flexible with a relatively low resolution of about 6–10 Å. The best map for this region clearly shows two separate parts. One of them is sandwiched by the elongated S6 segments of CATSPER2 and CATSPER3. However, the density quality is not very good and no predicted structures from the MS list fit the map. This density thus remains to be identified. The other part of the cytosolic density, which is mainly composed of α-helices, can be fitted by the predicted structures of EFCAB9 and CATSPERζ by tFold and trRosseta53, respectively. Owing to the limited resolution, reliable side chain assignment of this part is impractical.
The models were refined against the corresponding maps by PHENIX48 in real space (phenix.real_space_refine) with secondary structure and geometry restraints generated by ProSMART54. Overfitting of the overall models was monitored by refining the models in one of the two independent half-maps from the gold-standard refinement approach and testing the refined model against the other map55. Statistics of 3D reconstruction and model refinement, and a detailed summary of the final model can be found in Extended Data Tables 1 and 2, respectively. All structure figures were prepared in PyMOL56.
MS analysis
SDS–PAGE was used to remove the detergent from the protein sample. The SDS–PAGE gel containing the protein sample band was cut and digested by trypsin with prior reduction and alkylation in 50 mM ammonium bicarbonate at 37 °C overnight. The digested products were extracted twice with 1% formic acid in 50% acetonitrile aqueous solution and dried to reduce volume by speedvac.
For liquid chromatography with tandem MS (LC–MS/MS) analysis, the peptides were separated by a 65-min gradient elution at a flow rate of 0.300 μl/min with the Thermo EASY-nLC1200 integrated nano-HPLC system which is directly interfaced with the Thermo Q Exactive HF-X mass spectrometer. The analytical column was a home-made fused silica capillary column (75 μm internal diameter, 150 mm length; Upchurch Scientific) packed with C-18 resin (300 A, 3 μm, Varian, Lexington, MA). Mobile phase A consisted of 0.1% formic acid, and mobile phase B consisted of 100% acetonitrile and 0.1% formic acid. The mass spectrometer was operated in the data-dependent acquisition mode using the Xcalibur 4.1 software and there was a single full-scan mass spectrum in the Orbitrap (400–1,800 m/z, 60,000 resolution) followed by 20 data-dependent MS/MS scans at 30% normalized collision energy. Each mass spectrum was analysed using the Thermo Xcalibur Qual Browser and Proteome Discoverer for the database searching.
Reporting summary
Further information on research design is available in the Nature Research Reporting Summary linked to this paper.
Data availability
The cryo-EM maps of the mouse CatSpermasome and the corresponding atomic coordinate have been deposited in the Electron Microscopy Data Bank and the Protein Data Bank under the accession codes EMD-31076 and 7EEB, respectively. The mass spectrometry data have been deposited in the MassIVE database (https://massive.ucsd.edu/ProteoSAFe/static/massive.jsp) under the accession number MSV0000987325. All data analysed during this study are included in this Article and its Supplementary Information. Any other relevant data are available from the corresponding author upon reasonable request.
References
Ren, D. et al. A sperm ion channel required for sperm motility and male fertility. Nature 413, 603–609 (2001).
Ren, D. & Xia, J. Calcium signaling through CatSper channels in mammalian fertilization. Physiology 25, 165–175 (2010).
Quill, T. A., Ren, D., Clapham, D. E. & Garbers, D. L. A voltage-gated ion channel expressed specifically in spermatozoa. Proc. Natl Acad. Sci. USA 98, 12527–12531 (2001).
Lobley, A., Pierron, V., Reynolds, L., Allen, L. & Michalovich, D. Identification of human and mouse CATSPER3 and CATSPER4 genes: characterisation of a common interaction domain and evidence for expression in testis. Reprod. Biol. Endocrinol. 1, 53 (2003).
Liu, J., Xia, J., Cho, K. H., Clapham, D. E. & Ren, D. CatSperβ, a novel transmembrane protein in the CatSper channel complex. J. Biol. Chem. 282, 18945–18952 (2007).
Wang, H., Liu, J., Cho, K. H. & Ren, D. A novel, single, transmembrane protein CATSPERG is associated with CATSPER1 channel protein. Biol. Reprod. 81, 539–544 (2009).
Chung, J. J., Navarro, B., Krapivinsky, G., Krapivinsky, L. & Clapham, D. E. A novel gene required for male fertility and functional CATSPER channel formation in spermatozoa. Nat. Commun. 2, 153 (2011).
Chung, J. J. et al. CatSperζ regulates the structural continuity of sperm Ca2+ signaling domains and is required for normal fertility. eLife 6, e23082 (2017).
Hwang, J. Y. et al. Dual sensing of physiologic pH and calcium by EFCAB9 regulates sperm motility. Cell 177, 1480–1494 (2019).
Long, S. B., Campbell, E. B. & Mackinnon, R. Voltage sensor of Kv1.2: structural basis of electromechanical coupling. Science 309, 903–908 (2005).
Whitaker, M. Calcium at fertilization and in early development. Physiol. Rev. 86, 25–88 (2006).
Okabe, M. The cell biology of mammalian fertilization. Development 140, 4471–4479 (2013).
Wang, H. F., McGoldrick, L. L. & Chung, J. J. Sperm ion channels and transporters in male fertility and infertility. Nat. Rev. Urol. 18, 46–66 (2021).
Chung, J. J. et al. Structurally distinct Ca2+ signaling domains of sperm flagella orchestrate tyrosine phosphorylation and motility. Cell 157, 808–822 (2014).
Seifert, R. et al. The CatSper channel controls chemosensation in sea urchin sperm. EMBO J. 34, 379–392 (2015).
Yoshida, M. & Yoshida, K. Sperm chemotaxis and regulation of flagellar movement by Ca2+. Mol. Hum. Reprod. 17, 457–465 (2011).
Strunker, T. et al. The CatSper channel mediates progesterone-induced Ca2+ influx in human sperm. Nature 471, 382–386 (2011).
Lishko, P. V., Botchkina, I. L. & Kirichok, Y. Progesterone activates the principal Ca2+ channel of human sperm. Nature 471, 387–391 (2011).
Avenarius, M. R. et al. Human male infertility caused by mutations in the CATSPER1 channel protein. Am. J. Hum. Genet. 84, 505–510 (2009).
Hildebrand, M. S. et al. Genetic male infertility and mutation of CATSPER ion channels. Eur. J. Hum. Genet. 18, 1178–1184 (2010).
Jin, Z. R. et al. Roles of CatSper channels in the pathogenesis of asthenozoospermia and the therapeutic effects of acupuncture-like treatment on asthenozoospermia. Theranostics 11, 2822–2844 (2021).
Avidan, N. et al. CATSPER2, a human autosomal nonsyndromic male infertility gene. Eur. J. Hum. Genet. 11, 497–502 (2003).
Zhang, Y. et al. Sensorineural deafness and male infertility: a contiguous gene deletion syndrome. J. Med. Genet. 44, 233–240 (2007).
Luo, T. et al. A novel copy number variation in CATSPER2 causes idiopathic male infertility with normal semen parameters. Hum. Reprod. 34, 414–423 (2019).
Wang, J. X. et al. Patient with CATSPER3 mutations-related failure of sperm acrosome reaction with successful pregnancy outcome from intracytoplasmic sperm injection (ICSI). Mol. Genet. Genom. Med. 9, e1579 (2020).
Brown, S. G. et al. Homozygous in-frame deletion in CATSPERE in a man producing spermatozoa with loss of CatSper function and compromised fertilizing capacity. Hum. Reprod. 33, 1812–1816 (2018).
Quill, T. A. et al. Hyperactivated sperm motility driven by CATSPER2 is required for fertilization. Proc. Natl Acad. Sci. USA 100, 14869–14874 (2003).
Qi, H. et al. All four CatSper ion channel proteins are required for male fertility and sperm cell hyperactivated motility. Proc. Natl Acad. Sci. USA 104, 1219–1223 (2007).
Jin, J. et al. Catsper3 and Catsper4 are essential for sperm hyperactivated motility and male fertility in the mouse. Biol. Reprod. 77, 37–44 (2007).
Nikpoor, P., Mowla, S. J., Movahedin, M., Ziaee, S. A. M. & Tiraihi, T. CatSper gene expression in postnatal development of mouse testis and in subfertile men with deficient sperm motility. Hum. Reprod. 19, 124–128 (2004).
Long, J. E., Lee, M. S. & Blithe, D. L. Male contraceptive development: update on novel hormonal and nonhormonal methods. Clin. Chem. 65, 153–160 (2019).
Cheng, C. Y. & Mruk, D. D. New frontiers in nonhormonal male contraception. Contraception 82, 476–482 (2010).
Kirichok, Y., Navarro, B. & Clapham, D. E. Whole-cell patch-clamp measurements of spermatozoa reveal an alkaline-activated Ca2+ channel. Nature 439, 737–740 (2006).
Tao, X., Lee, A., Limapichat, W., Dougherty, D. A. & MacKinnon, R. A gating charge transfer center in voltage sensors. Science 328, 67–73 (2010).
Xie, J. F. et al. Structure of the human sodium leak channel NALCN in complex with FAM155A. Nat. Commun. 11, 5831 (2020).
Hagenbuch, B. & Stieger, B. The SLCO (former SLC21) superfamily of transporters. Mol. Aspects Med. 34, 396–412 (2013).
Fietz, D. et al. Membrane transporters for sulfated steroids in the human testis—cellular localization, expression pattern and functional analysis. PLoS One 8, e62638 (2013).
Suzuki, T. et al. Identification and characterization of novel rat and human gonad-specific organic anion transporters. Mol. Endocrinol. 17, 1203–1215 (2003).
Abbott, G. W. Chansporter complexes in cell signaling. FEBS Lett. 591, 2556–2576 (2017).
Li, N. N. et al. Structure of a pancreatic ATP-sensitive potassium channel. Cell 168, 101–110.e10 (2017).
Perez, G. I. et al. Prolongation of ovarian lifespan into advanced chronological age by Bax-deficiency. Nat. Genet. 21, 200–203 (1999).
Lei, J. & Frank, J. Automated acquisition of cryo-electron micrographs for single particle reconstruction on an FEI Tecnai electron microscope. J. Struct. Biol. 150, 69–80 (2005).
Zheng, S. Q. et al. MotionCor2: anisotropic correction of beam-induced motion for improved cryo-electron microscopy. Nat. Methods 14, 331–332 (2017).
Punjani, A., Rubinstein, J. L., Fleet, D. J. & Brubaker, M. A. cryoSPARC: algorithms for rapid unsupervised cryo-EM structure determination. Nat. Methods 14, 290–296 (2017).
Punjani, A., Zhang, H. & Fleet, D. J. Non-uniform refinement: adaptive regularization improves single-particle cryo-EM reconstruction. Nat. Methods 17, 1214–1221 (2020).
Zivanov, J. et al. New tools for automated high-resolution cryo-EM structure determination in RELION-3. eLife 7, e42166 (2018).
Scheres, S. H. W. RELION: implementation of a Bayesian approach to cryo-EM structure determination. J. Struct. Biol. 180, 519–530 (2012).
Adams, P. D. et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr. D Biol. Crystallogr. 66, 213–221 (2010).
Goddard, T. D. et al. UCSF ChimeraX: Meeting modern challenges in visualization and analysis. Protein Sci. 27, 14–25 (2018).
Pettersen, E. F. et al. UCSF Chimera—a visualization system for exploratory research and analysis. J. Comput. Chem. 25, 1605–1612 (2004).
Emsley, P. & Cowtan, K. Coot: model-building tools for molecular graphics. Acta Crystallogr. D Biol. Crystallogr. 60, 2126–2132 (2004).
Smart, O. S., Neduvelil, J. G., Wang, X., Wallace, B. A. & Sansom, M. S. P. HOLE: a program for the analysis of the pore dimensions of ion channel structural models. J. Mol. Graph. Model. 14, 354–360 (1996).
Yang, J. et al. Improved protein structure prediction using predicted interresidue orientations. Proc. Natl Acad. Sci. USA 117, 1496–1503 (2020).
Nicholls, R. A., Fischer, M., McNicholas, S. & Murshudov, G. N. Conformation-independent structural comparison of macromolecules with ProSMART. Acta Crystallogr. D Biol. Crystallogr. 70, 2487–2499 (2014).
Amunts, A. et al. Structure of the yeast mitochondrial large ribosomal subunit. Science 343, 1485–1489 (2014).
DeLano, W. L. The PyMOL Molecular Graphics System http://www.pymol.org (2002).
Acknowledgements
We thank N. Yan and H. Yu for critical reading of the manuscript; M. Jiang and Y. Ru for help with the immunofluorescence experiments; the Cryo-EM Facility and HPC Center of Westlake University for providing cryo-EM and computation support; S. Feng and the Mass Spectrometry & Metabolomics Core Facility of Westlake University for protein sample MS analysis; and J. Bao, D. Wu and the Laboratory Animal Resources Center of Westlake University for help with animal maintenance and IVF experiments. This work was supported by Westlake Laboratory (Westlake Laboratory of Life Sciences and Biomedicine) (W101486022101) and an Institutional Startup Grant from the Westlake Education Foundation (101486021901) to J.W.
Author information
Authors and Affiliations
Contributions
J.W. conceived and supervised the project; S.L. and Y.Z. prepared the protein sample under the guidance of Z.Y. and J.W.; S.L. collected the cryo-EM data and M.K. calculated the cryo-EM map; J.W. built the model; all authors contributed to data analysis; and J.W. wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Nature thanks the anonymous reviewers for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Mouse genotyping, confocal imaging of spermatozoa and endogenous purification of the CatSpermasome from mouse sperm.
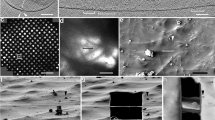
a, The genotype of each mouse was verified by two PCR reactions. Top, schematic of the genotyping procedure. Bottom, a group of representative PCR results. NC, negative control (empty template); WT, wild type; KI/+, heterozygote knock-in; KI/KI, homozygote knock-in. In total, 49 wild-type, 676 KI/+, and 207 KI/KI mice were verified. b, An drawing of a mouse sperm. CatSper was mainly distributed at the principal piece of spermatozoa. c, EGFP fluorescence was detected in the principal piece (red arrows) of knock-in mouse spermatozoa, but not wild-type mouse spermatozoa. Blue arrows, auto-fluorescence signal observed in the middle piece of spermatozoa. Shown here are one of the two images taken for each sample. Scale bars, 30 μm. d, Schematic of CatSpermasome purification. e, The purified protein sample was subjected to gel filtration analysis. The peak fractions of CatSpermasome (arrow) were collected and concentrated for cryo-EM and MS studies. Inset, the cross-linked protein sample was visualized on SDS–PAGE by silver staining. The corresponding protein band (arrow) in a separate gel without staining was cut out for MS analysis. For gel source data, see Supplementary Fig. 1. f, A representative EM micrograph of the CatSpermasome sample stained with uranyl acetate (one micrograph out of five in total for the negative staining sample). Scale bar, 50 nm.
Extended Data Fig. 2 Mass spectrometric analysis of the purified CatSpermasome.
a, MS samples were the same as used for the cryo-EM study. MS detected proteins are shown in order of decreasing confidence. All previously characterized CatSper components are listed (yellow). The top six entries with highest peptide spectrum match (PSM) values are all CATSPER proteins. Most of the contaminating proteins are cytoskeletal proteins. The newly identified components SLCO6C1 and CATSPERη are highlighted in red. TMEM249, which is probably another new component, is shaded in light blue. b, Representative MS spectra for specific peptides of CATSPER1, SLCO6C1, CATSPERη, and TMEM249.
Extended Data Fig. 3 Cryo-EM analysis of mouse CatSpermasome.
a, A representative motion-corrected micrograph of the CatSpermasome cryo-EM sample out of a dataset of 16,648 images. Scale bar, 50 nm. b, Two-dimensional class averages. Box size, 430 Å. c, Gold standard FSC curves for the 3D reconstructions. The curves were calculated with masks for the entire protein (overall map), and for masked regions of corresponding maps. See f for each map. d, Validation of the final structure models. FSC curves of the final refined model versus the summed map that it was refined against (black); of the model versus the first half-map (blue); and of the model versus the second half-map (red). The small difference between the blue and red curves indicates that the refinement of the atomic coordinates did not suffer from overfitting. e, Angular distribution of the particles of the final reconstruction generated by cryoSPARC. f, Flowchart of EM data processing (see ‘Image processing’ in Methods).
Extended Data Fig. 4 Structural features of the cytosolic regions.
a, The cytosolic region consists of two separate but interacting parts. Cytosolic map 1 is in close contact with the S6 segments of CATSPER2 and CATSPER3. A density surrounded in the bottom of cytosolic map 1 (shown in green) is connected to the density of the S6 segment of CATSPER3 and may belong to the carboxyl end of CATSPER3. Owing to limited resolution, the identity of cytosolic map 1 remains to be determined. Cytosolic map 2, however, is likely to be the subcomplex of EFCAB9 and CATSPERζ. The maps were generated in ChimeraX. b, Predicted structures of EFCAB9 by tFold and CATSPERζ by trRosseta. EFCAB9 has two EF-hand motif-containing lobes, which is very similar to calmodulin. CATSPERζ consists mainly of α-helices. c, Docking of the predicted structures of EFCAB9 and CATSPERζ into cytosolic map 2. The two lobes of EFCAB9 are in a compact conformation instead of the extended conformation in the predicted structure. The main body of CATSPERζ (light cyan in b) can be fitted into the remaining density near EFCAB9 in cytosolic map 2. Several fitted α-helices are indicated by arrows.
Extended Data Fig. 5 EM maps of CATSPER1–4.
a, Electron density maps of each segment of the transmembrane helices of CATSPER1–4. The boundaries of each displayed segment are labelled. The densities, shown as blue meshes, are contoured at 3–4σ in PyMOL. b, Electron density map of the selectivity filter and the pore helices. The densities are contoured at 4σ. Two tentatively assigned Na+ ions are shown as purple spheres. c, Electron density maps of detergent-like molecules. These densities may also belong to cholesterol or steroid hormones under physiological conditions. Three GDN molecules are tentatively assigned to these densities. The densities are found in a semi-open cavity formed by the S3, S4 and S4–5 segments of CATSPER1, but not CATSPER2–4, whose corresponding cavities are smaller. d, Structural superimposition of CATSPER1–4 indicates that CATSPER1 has a larger cavity for binding of detergent-like molecules.
Extended Data Fig. 6 Structural details and sequence alignment of the pore domain and VSDs.
a, Overall structure of the pore domain of CATSPER1–4. The critical DDDD residues in the selectivity filter are shown as sticks. Each S6 segment contains a π-helix turn (red arrows). b, Sequence alignment of the selectivity filter and the pore helices among mouse CATSPER1–4, human CATSPER1–4, and rabbit Cav1.1. The invariant Thr and Trp residues are shaded cyan. The DDDD residues in the selectivity filter of CATSPER1–4 and the corresponding residues EEEE in Cav1.1 are highlighted red. c, Sequence alignment of the VSDs among CATSPER1–4. The boundary of each segment is shaded light grey. Positively charged residues on S4 segments are shaded blue and residues corresponding to positions R1–R6 are boxed. An1 and CTC residues on segments S2 and S3 are shaded purple. d, Structural comparison of the VSDs among CATSPER1–4. The four VSDs are superimposed relative to CTC and An1 on S2. For visual clarity, the S1 segments are omitted and only the side chains of aligned residues and R4 residues on the S4 segments are shown.
Extended Data Fig. 7 Structure of the auxiliary subunits CATSPERβ, γ, δ and ε.
a, The overall structures of CATSPERβ, γ, δ and ε share similar domain organizations. The structures are shown in cartoon form and the sugar moieties in the glycosylation sites are shown as sticks. The NTDs, β-propeller domains, Ig-like domains, stem domains and transmembrane domains are coloured green, blue, orange, yellow and salmon, respectively. The head domain in CATSPERβ and the NTD2 domain in CATSPERε are coloured slate and cyan, respectively. b, Domain organization of CATSPERβ, γ, δ and ε. The boundaries for each domain and the identified glycosylation sites are labelled. The disulfide bonds are indicated by orange lines. See Extended Data Table 2 for details. c, Inter-subunit interactions among CATSPERβ, γ, δ and ε, shown in four side views. CATSPERβ is domain coloured and CATSPERγ, δ and ε are coloured as in Fig. 1. The side openings formed by two adjacent subunits are indicated by dotted lines. For visual clarity, the transmembrane helices are omitted. d, Extracellular view of the auxiliary subunits. The top of the channel is sealed by the Ig-like domains of CATSPERβ, γ, δ and ε. Bottom, close-up view of the interactions among the Ig-like domains (box in top image). The residues that mediate the interactions in the interface are shown as sticks. Hydrogen bonds are indicated by red dashed lines. e, Interactions between the transmembrane domains of CATSPERβ, γ, δ and ε and the adjacent VSDs of CATSPER1–4. The residues that contribute to the interface interactions are shown as sticks. Potential hydrogen bonds are indicated by red dashed lines. The stem domains of CATSPERβ and CATSPERε are further from the adjacent channel subunits than those of CATSPERγ and CATSPERδ (double-headed arrows).
Extended Data Fig. 8 Representative EM maps of the auxiliary subunits CATSPERβ, γ, δ and ε.
The selected segments cover almost every domain of CATSPERβ, γ, δ and ε. Representative densities for the glycosylation sites from each subunit are also presented. The densities, shown as blue meshes, are contoured at 4–5σ in PyMOL.
Extended Data Fig. 9 Structural analysis and sequence alignment of SLCO6C1.
a, Side view (left) and central slice view (right) of the electron density map of SLCO6C1 and the adjacent components. The map was generated in ChimeraX. b, SLCO6C1 is captured in an outward-facing conformation. The structure is shown in cylindrical helices cartoon mode and coloured by domain. Right, the N and C domains of SLCO6C1 each contain six transmembrane helices and are pseudo-symmetric, with an r.m.s.d. of ~6 Å when superimposed. c, Sequence alignment among mouse SLCO6C1, rat SLCO6C1 and human SLCO6A1. The glycosylation site (red triangle) and the residues that may be involved in interactions with CATSPERε (yellow triangles) are conserved among species. The uniport IDs for the aligned sequences are: mSLCO6C1: Q3V161; rSLCO6C1: G3V7R7; and hSLCO6A1: Q86UG4.
Extended Data Fig. 10 Structural characterization of CATSPERη and TMEM249.
a, Side view (left) and central slice view (right) of the electron density maps of CATSPERη and the adjacent components. The map was generated in ChimeraX. b, Electron density maps of CATSPERη. The map allows accurate assignment of the side chains of the bulky residues. The densities, shown as blue meshes, are contoured at 4σ in PyMOL. c, Predicted structure of TMEM249 by tFold perfectly fits into the density near CATSPERη with minor adjustments. The structure has the characteristics of two transmembrane helices and a β-sheet (arrows) in the cytosolic domain. d, TMEM249 interacts with CATSPERη and VSD4 in the transmembrane and cytosolic regions, respectively. Unlike CATSPERη, TMEM249 does not interact with the stem domain of CATSPERβ.
Extended Data Fig. 11 Immunofluorescence detection of representative CatSpermasome components in wild-type sperm.
CATSPER1, CATSPER4, CATSPERβ and TMEM249 are mainly distributed in the principal piece of sperm (red fluorescence signal). The sperm head, middle piece and principal piece are indicated by black, blue, and red arrows, respectively. NC, negative control (without primary antibody). Shown here are one of three images taken for each sample. Scale bars, 20 μm.
Supplementary information
Supplementary Figure 1
Uncropped raw data gel from Extended Data Fig. 1 - SDS-PAGE gel for Extended Data Fig. 1e showing the BS3-crosslinked purified mouse CatSpermasome sample by silver staining.
Video 1
: Overall EM map of the mouse CatSpermasome The displayed overall composite map was in the same color scheme as Fig. 1. The movie was generated in ChimeraX
Rights and permissions
About this article
Cite this article
Lin, S., Ke, M., Zhang, Y. et al. Structure of a mammalian sperm cation channel complex. Nature 595, 746–750 (2021). https://doi.org/10.1038/s41586-021-03742-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-021-03742-6
- Springer Nature Limited
This article is cited by
-
Structural biology and molecular pharmacology of voltage-gated ion channels
Nature Reviews Molecular Cell Biology (2024)
-
Sperm-Specific CatSper is Not Conserved in All Vertebrates and May Not be the Only Progesterone-Responsive Ion Channel Present in Sperm
The Journal of Membrane Biology (2024)
-
Control of intracellular pH and bicarbonate by CO2 diffusion into human sperm
Nature Communications (2023)
-
Cryo-EM structures of human organic anion transporting polypeptide OATP1B1
Cell Research (2023)
-
Advances in the study of genetic factors and clinical interventions for fertilization failure
Journal of Assisted Reproduction and Genetics (2023)