Abstract
Cytokines were the first modern immunotherapies to produce durable responses in patients with advanced cancer, but they have only modest efficacy and limited tolerability1,2. In an effort to identify alternative cytokine pathways for immunotherapy, we found that components of the interleukin-18 (IL-18) pathway are upregulated on tumour-infiltrating lymphocytes, suggesting that IL-18 therapy could enhance anti-tumour immunity. However, recombinant IL-18 previously did not demonstrate efficacy in clinical trials3. Here we show that IL-18BP, a high-affinity IL-18 decoy receptor, is frequently upregulated in diverse human and mouse tumours and limits the anti-tumour activity of IL-18 in mice. Using directed evolution, we engineered a ‘decoy-resistant’ IL-18 (DR-18) that maintains signalling potential but is impervious to inhibition by IL-18BP. Unlike wild-type IL-18, DR-18 exerted potent anti-tumour effects in mouse tumour models by promoting the development of poly-functional effector CD8+ T cells, decreasing the prevalence of exhausted CD8+ T cells that express the transcriptional regulator of exhaustion TOX, and expanding the pool of stem-like TCF1+ precursor CD8+ T cells. DR-18 also enhanced the activity and maturation of natural killer cells to effectively treat anti-PD-1 resistant tumours that have lost surface expression of major histocompatibility complex class I molecules. These results highlight the potential of the IL-18 pathway for immunotherapeutic intervention and implicate IL-18BP as a major therapeutic barrier.
Similar content being viewed by others
Main
Cytokines are secreted proteins that provide instructive cues to immune cells and are therefore attractive candidates for use in cancer immunotherapy. However, the clinical application of cytokines has been hampered by their biological pleiotropism, which reduces their therapeutic specificity and can cause toxicity2. A major effort in cytokine research is to engineer ‘designer’ cytokines with tailored biological activities4 that will precisely activate anti-tumour immune programs. To identify ways to improve cytokine immunotherapy, we analysed transcriptional datasets to characterize patterns of cytokine and cytokine receptor expression on CD8+ tumour-infiltrating lymphocytes (TILs). IL-18 and the subunits of its receptor (IL-18Rα/Rβ) were enriched in both activated and dysfunctional tumour CD8+ T cells (Extended Data Fig. 1a), suggesting that IL-18 agonism could effectively stimulate anti-tumour responses.
IL-18 is a member of the IL-1 cytokine family and mediates inflammation downstream of the NLRP3 and NLRP1 inflammasomes5. It drives MYD88 signalling through heterodimerization of its receptor subunits IL-18Rα (encoded by IL18R1) and IL-18Rβ (IL18RAP). Originally termed interferon-γ-inducing factor (IGIF), IL-18 has been found to stimulate innate lymphocytes and antigen-experienced, but not naive, T cells6. Therapeutically, recombinant IL-18 (rIL-18) has been reported to synergize with immune checkpoint inhibitors (ICIs)7 and chimeric antigen receptor T (CAR-T) cells in mouse tumour models8. rIL-18 has been administered to patients in clinical trials and found to be safe and well-tolerated9. However, clinical development of rIL-18 has been curtailed by its lack of efficacy3. IL-18 is negatively regulated by a decoy receptor called IL-18 binding protein (IL-18BP), a secreted antagonist that binds IL-18 with extremely high affinity (KD < 1 nM)10. In patients treated with rIL-18, serum concentrations of IL-18BP increased by 10- to 100-fold9,11. Therefore, we hypothesized that IL-18BP produced in the tumour microenvironment (TME) might act as a ‘secreted immune checkpoint’ to limit effective rIL-18 immunotherapy.
IL-18R and IL-18BP are prevalent in the TME
We initially sought to characterize the expression of IL-18 pathway components in mouse tumours. Through immunophenotyping of MC38 and YUMMER1.7 tumours and matched spleens, we found that IL-18Rα was widely expressed on natural killer (NK) cells, but markedly upregulated on CD4+ and CD8+ T cells in the tumour compared to the spleen (Extended Data Fig. 1b–d). Within the T cell compartment, acquisition of IL-18Rα expression was exclusive to antigen-experienced CD44+ T cells (Extended Data Fig. 1e, f). In addition, both Il18bp transcripts and IL-18BP protein were highly expressed in the TME and further increased by treatment with mouse (m) IL-18 in an IFN-γ-dependent fashion (Extended Data Fig. 1g–j).
To determine whether these results translated to human tumours, we analysed IL18BP expression in the Cancer Genome Atlas (TCGA) database. Expression of IL18BP was increased across many tumour types compared to matched normal tissue controls (Extended Data Fig. 2a). Expression of IL18BP strongly correlated with expression of CD3E, CD8A, and PDCD1 (R = 0.59–0.88), indicating that the increase in expression was associated with the presence of activated CD8+ T cells (Extended Data Fig. 2b–d). We confirmed expression of IL-18BP protein in the TME by immunohistochemical staining of tissue microarrays for several tumour types (Extended Data Fig. 2e). Enzyme-linked immunosorbent assay (ELISA) showed that IL-18BP expression was also elevated in the serum of patients with non-small cell lung cancer (NSCLC) and was increased further by anti-PD-1 or anti-PD-L1 treatment (Extended Data Fig. 2f).
To assess the functional effect of IL-18BP on IL-18 therapy, we engrafted MC38 tumours into either wild-type C57BL/6 mice or Il18bp−/− mice and treated them with mIL-18 or vehicle. Whereas mIL-18 had no effect on tumour growth in wild-type mice, it significantly inhibited tumour growth in Il18bp−/− mice (Extended Data Fig. 2g; P < 0.001). Together, these data indicate that IL-18BP expression is common in cancer and that it may act as a soluble immune checkpoint.
Engineering a decoy-resistant IL-18
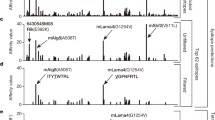
Given the potential limitation of IL-18BP on rIL-18 immunotherapy, we sought to create a decoy-resistant IL-18 variant (DR-18) that retains full signalling capacity through the IL-18 receptor, but is impervious to inhibition by IL-18BP (Fig. 1a). This posed an engineering challenge, as IL-18Rα and IL-18BP bind IL-18 at a highly overlapping interface, and IL-18BP binds IL-18 with an affinity that is more than three orders of magnitude higher than that of IL-18Rα (Extended Data Fig. 3a–c). Although point mutations (E6A and K53A) in human (h) IL-18 have been purported to reduce neutralization by IL-18BP12, we found that proteins with these mutations retained IL-18BP binding without improvements in selectivity towards IL-18Rα (Extended Data Fig. 3d). We therefore used directed evolution with yeast surface display to screen more than 250 million mIL-18 variants that were randomized at 13 receptor contact positions for those that bound to IL-18Rα but not IL-18BP (Fig. 1a, Extended Data Fig. 3e). After five rounds of selection for IL-18Rα and counter-selection against IL-18BP (Extended Data Fig. 3f), we obtained a population that exclusively bound IL-18Rα (Fig. 1b). Sequencing of this population revealed 11 unique sequences, from which we created two ‘consensus sequences’—CS1 and CS2 (Fig. 1c, Extended Data Fig. 3g). We recombinantly expressed these variants and measured their affinities for IL-18Rα and IL-18BP by surface plasmon resonance. All of the selected variants retained IL-18Rα binding, with negligible binding to IL-18BP (KD > 10 μM) (Fig. 1c).
a, Schematic representations of DR-18 (left) and yeast-display directed evolution process (right). b, Representative histogram assessing staining for IL-18Rα (100 nM, left) and IL-18BP (200 nM, right) by flow cytometry of yeast display library after each round of selection. c, The sequences and dissociation constants (KD) of clones summarized for selected DR-18 variants, with differences from wild-type IL-18 indicated for each mutant at the given amino acid position (top). Green shading highlights converging residues to form consensus sequence (bottom). KD measurements are shown for binding of mIL-18 and DR-18 variants to IL-18Rα and IL-18BP measured by SPR. –, not tested; NBD, no binding detected. d, Quantification of intracellular IFN-γ staining in splenic NK cells stimulated with IL-18, CS1 or CS2 in the presence of IL-12. e, Quantification of intracellular IFN-γ staining in splenic NK cells stimulated with IL-18 or CS1 or CS2 in the presence of IL-12 and varying concentrations of IL-18BP. Data are representative of three independent experiments (d, e) with n = 3 mice per condition and presented as mean ± s.e.m.
To evaluate the functionality of DR-18, we stimulated NK cells ex vivo with either mIL-18 or the DR-18 variants CS1 and CS2 and measured production of IFN-γ by the cells. CS1 had equal potency to IL-18 (EC50 = 74 and 54 pM, respectively), whereas CS2 was around 1.5 log more potent (EC50 = 2.4 pM; Fig. 1d). We then measured the decoy-resistance of the DR-18 variants by fixing the concentration of the variants while titrating increasing amounts of IL-18BP. Whereas mIL-18 was sensitive to IL-18BP (IC50 = 3.6 nM), NK cells stimulated with either CS1 or CS2 maintained robust IFN-γ production irrespective of IL-18BP concentration (Fig. 1e). On the basis of these results, we elected to proceed with CS2 (hereafter referred to as DR-18) for subsequent studies.
DR-18 is active against mouse tumours
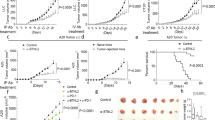
We compared the efficacy of DR-18 and mIL-18 in the treatment of syngeneic mouse colorectal and melanoma tumours (Fig. 2a, see Methods). Whereas treatment with mIL-18 was ineffective, treatment with DR-18 produced strong tumour growth inhibition, enhanced survival, and (in some mice) complete tumour regression (Fig. 2b, c, Extended Data Fig. 4a–g). The efficacy of DR-18 was commensurate or superior to that of anti-PD-1 monotherapy, and the combination of DR-18 and anti-PD-1 produced a synergistic response that resulted in complete tumour regression in most of the treated mice (Fig. 2b, c, Extended Data Fig. 4a, b, e).
a, Wild-type (WT) mice were subcutaneously (s.c.) engrafted with 0.5 × 106 MC38 cells. From day 7 (tumour size 50–100 mm3), mice were treated twice weekly with phosphate-buffered saline (PBS), IL-18 or DR-18 (s.c.), or with anti-PD-1 intraperitoneally (i.p.). For depletion and neutralization studies, mice were injected i.p. with anti-CD4, anti-CD8, anti-NK1.1, or anti-IFN-γ at the indicated time points (red arrows). b, c, Mean tumour growth (b) and Kaplan–Meier survival curves (c) for mice bearing MC38 tumours after treatment. Black arrows denote days of treatment. d, Kaplan–Meier survival curves of mice engrafted with MC38 tumours and treated with DR-18, alone or with indicated depletion/neutralization. e, Kaplan–Meier survival curves of wild-type or Il18r1−/− mice engrafted with MC38 tumours following treatment. f, Kaplan–Meier survival curves of Rag2−/− mice reconstituted with bulk CD3+ T cells from either wild-type or Il18r1−/− donor mice, subsequently engrafted with MC38 tumours, and treated with PBS or DR-18. b–f, Data are representative of three independent experiments, with minimum n = 5 mice per group; b, mean ± s.e.m.
To exclude the possibility that DR-18 activity could be attributable to increased IL-18 receptor affinity (as opposed to its independence from IL-18BP), we compared the efficacy of DR-18 (CS2) to the related variant CS1, which has similar affinity towards IL-18Rα to that of mIL-18. CS1 produced similar efficacy to DR-18, whereas even high doses (1 mg kg−1) of mIL-18 did not elicit anti-tumour responses (Extended Data Fig. 4h). These results indicate that the anti-tumour efficacy of DR-18 is driven by its independence from IL-18BP.
DR-18 acts on T cells to treat tumours
To determine the contribution of particular immune cell populations to the effects of DR-18 on tumours, we performed antibody-mediated depletion studies. In MC38 tumours, the efficacy of DR-18 was abrogated by depletion of CD8+ T cells and partially inhibited by depletion of NK1.1+ cells (Fig. 2a, d). In YUMMER1.7 tumours, effective treatment with DR-18 required CD8+ and CD4+ T cells, but not NK1.1+ cells (Extended Data Fig. 5a). Consistent with these results, DR-18 treatment was ineffective towards tumours engrafted in Rag2−/− mice (Extended Data Fig. 5b). In both models, DR-18 activity was dependent on IFN-γ (Fig. 2d, Extended Data Fig. 5a). Similarly, DR-18 treatment was ineffective in Il18r1−/− mice, confirming that DR-18 activity is mediated through the IL-18 receptor (Fig. 2e). To assess the ability of DR-18 to promote memory responses, mice surviving primary MC38 engraftment following DR-18 treatment were re-challenged with MC38; nearly all the mice (14/15) rejected the second tumour inoculation (Extended Data Fig. 5c).
To determine whether antigen-specific CD8+ T cells are targeted by DR-18 treatment, we adoptively transferred Thy1.1+P14+ CD8+ T cells into mice bearing GP33-expressing B16 melanomas and subsequently treated the mice with either DR-18 or vehicle. Following treatment with DR-18, there was a substantial increase in the frequency, number, and functionality (IFN-γ production) of intratumoral P14+ CD8+ T cells (Extended Data Fig. 5d–g). Furthermore, analysis of the endogenous CD8+ (Thy1.1−) T cell response revealed that the majority of infiltrating CD8+ TILs were CD44+CD39+ (Extended Data Fig. 5h, i), a phenotype that is associated with tumour antigen-specific cells13. This activity could be localized directly to an effect on intratumoral cells, as DR-18 increased the frequency of activated CD8+ TILs even in the presence of FTY720, an inhibitor of T cell egress from lymphoid tissues (Extended Data Fig. 5j).
Within tumours, IL-18Rα is predominantly expressed on intratumoral T and NK cells, although some myeloid cells express it at low levels (Extended Data Fig. 5k). To determine whether T cells were sufficient to mediate the efficacy of DR-18, we adoptively transferred T cells from either wild-type or Il8r1−/− donors into Rag2−/− recipients, which we engrafted with MC38 tumours and treated with DR-18 or vehicle. Whereas transfer of wild-type T cells restored the response of Rag2−/− mice to DR-18 treatment, Il18r1−/− T cells conferred no benefit (Fig. 2f, Extended Data Fig. 5l). Furthermore, depletion of XCR1+ cDC1 cells—which are essential for priming anti-tumour CD8+ T cell responses14—did not affect the efficacy of DR-18 during treatment (Extended Data Fig. 5m). Together, these results indicate that the direct activity of DR-18 on T cells is sufficient to drive anti-tumour responses.
DR-18 remodels the immune TME
To investigate the consequences of DR-18 treatment on the TME, we performed single-cell RNA sequencing (scRNA-seq) of YUMMER1.7 tumours after treatment with DR-18, mIL-18, or vehicle (see Methods). Globally, DR-18 induced striking changes in clusters of lymphocytes, macrophages, granulocytes, and fibroblasts (Extended Data Fig. 6a–d). Consistent with the antibody depletion studies above, Il18r1 and Il18rap were expressed predominantly in T cells and NK cells. By contrast, most Il18 and Il18bp transcripts were expressed in fibroblasts, granulocytes and macrophages (Extended Data Fig. 6e).
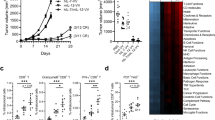
Analysis of Cd3e+ clusters yielded five CD8+ T cell clusters (CD8_1 to CD8_5) and two CD4+ T cell clusters (CD4_1 and CD4_2; Fig. 3a). Cluster CD8_1 was exclusive to and predominant after DR-18 treatment, and was characterized by high levels of effector molecule transcripts (Ifng, Prf1, and Gzmb), cell surface proteins (Il8r1 and Klrg1), and co-stimulatory receptors (Fig. 3b–d, Extended Data Fig. 6f), consistent with an effector (TEFF) phenotype15,16. Conversely, most T cells from tumours treated with mIL-18 or vehicle were found in cluster CD8_2, which exhibited high expression of Tox, Cd101, Cd38, and Cd244 (Fig. 3b–d, Extended Data Fig. 6f), indicative of an exhausted (TEX) phenotype17,18,19. CD8_3 is likely to represent naive-like T (TN) cells, which express lymphoid homing markers and high levels of Tcf7. CD8_4 and CD8_5 showed expression of cell division genes and the transcription factors Tcf7 and Id3, similar to stem-like precursor CD8+ T cells that have been found in patients with chronic viral infection or tumours20,21,22 (Fig. 3d). Overall, the two CD4 clusters were present at lower frequencies and were differentiated by the expression of Foxp3 in cluster CD4_1 (Fig. 3c, Extended Data Fig. 6f).
a, t-SNE projection showing Cd3e-positive cells present in YUMMER1.7 and YUMMER1.7-B2m−/− tumours coloured by cluster (see Methods). b, t-SNE projection showing distribution of Cd3e+ cells coloured by treatment. c, Bar plot showing frequency of T cells within each cluster as a function of treatment. d, Heatmap showing mean expression of curated gene list in CD8+ T cell clusters from c. a–d, Data are from one experiment with n = 3 tumours pooled per condition. e–n, Wild-type mice were subcutaneously engrafted with MC38 tumours and treated twice weekly with PBS, IL-18, anti-PD-1, or DR-18 from day 7 after tumour implantation. Tumours were analysed on day 15 by flow cytometry. e, Numbers of tumour-infiltrating CD4+ T cells, CD8+ T cells, and NK cells in each treatment group. f, g, Representative flow plots (f) and absolute numbers of polyfunctional CD8+ T cells (g) measured by co-staining for intracellular IFN-γ, TNF-α, and granzyme B (GZMB) following ex vivo stimulation with phorbol myristate acetate (PMA) and ionomycin. h, i, Representative flow plots (h) and quantification of intracellular TOX staining (i) in intratumoral CD44+ CD8+ T cells. j, k, Representative flow plots (j) and absolute cell numbers (k) of DN (PD-1−TIM3−), SP (PD-1+TIM3−) or DP (PD-1+TIM3+) intratumoral CD44+CD8+ T cells in each treatment group. l, m, Representative flow plots (l) and absolute numbers (m) of TCF1+ intratumoral SP CD8+ T cells. n, Absolute numbers of IFN-γ+ intratumoral DP CD8+ T cells. e–n, Data are representative of three independent experiments with n = 5 mice per group and presented as the mean ± s.e.m. P values were calculated using two-tailed unpaired Student’s t-tests (e, i, m, n) or two-way ANOVA (g, k).
Treatment with DR-18 also affected intratumoral myeloid populations (Extended Data Fig. 6g–j). Tumours from mice treated with vehicle or mIL-18 primarily contained immunosuppressive macrophages expressing Cx3cr1, Apoe, Mrc1 and Mertk23 and monocytes expressing Ccr2 and Ly6c (Extended Data Fig. 6g, k). By contrast, DR-18-treated tumours predominantly contained cells from cluster Mac_2, which exhibited high expression of Nos2, Arg1, and Cd274, consistent with a proinflammatory IFN-γ-activated signature. We confirmed this phenotype by flow cytometry (Extended Data Fig. 6l, m). Additionally, we observed increased abundance of granulocytes following DR-18 treatment (Extended Data Fig. 6a–c), similar to effects previously observed following effective ICI treatment24.
DR-18 increases polyfunctional TILs
To validate the TIL phenotypes identified using scRNA-seq, we performed immunophenotyping of YUMMER1.7 or MC38 tumours from mice treated with vehicle, mIL-18, or DR-18. We found an increase in the number and proliferative index (Ki67+) of CD4+ T cells, CD8+ T cells, and NK cells in tumours treated with DR-18 (Fig. 3e, Extended Data Fig. 7a, b), including an increase in CD44+ CD8+ T cells that co-expressed CD39 and PD-1 (Extended Data Fig. 7c–e). Intratumoral CD4+ T cells, CD8+ T cells, and NK cells also showed enhanced staining for IFN-γ, granzyme B, and CD107a after treatment with DR-18 (Extended Data Fig. 7f–k). Notably, tumours from DR-18-treated mice showed a substantial increase in the number of polyfunctional CD8+ T cells and NK cells (Fig. 3f, g, Extended Data Fig. 7l, m), a feature of an effective anti-tumour response17. Concordant with their increased polyfunctionality, CD8+ T cells from DR-18-treated tumours expressed lower levels of TOX (Fig. 3h, i).
DR-18 expands intratumoral stem-like T cells
Durable immunotherapeutic responses require a stem-like CD8+ T cell precursor population that expresses TCF1 and PD-120,21,25,26. These stem-like cells differentiate into terminal effector cells, which lose TCF1 expression, acquire TIM3 and have a greater functional capacity, but are limited in their ability to self-renew. In tumours treated with vehicle or mIL-18, the distribution of cells between the PD-1−TIM3− double-negative (DN), PD-1+ single-positive (SP), and PD-1+TIM3+ double-positive (DP) populations was similar. Treatment with DR-18, and to a lesser extent anti-PD-1, shifted the distribution towards the SP and predominantly DP fractions (Fig. 3j, k, Extended Data Fig. 8a, b). Analysis of TCF1 expression revealed that DR-18—but not vehicle, mIL-18, or anti-PD-1— greatly expanded the number of intratumoral TCF1+CD44+ CD8+ T cells, specifically in the SP population (Fig. 3l, m, Extended Data Fig. 8c–g). In the DP population, DR-18 did not affect TCF1 levels but did increase IFN-γ expression, as did anti-PD-1 treatment (Fig. 3n, Extended Data Fig. 8h–i). Notably, both TCF1+ precursor and PD-1+CD44+ CD8+ T cells were expanded in the tumour draining lymph nodes (tdLNs) in mice treated with DR-18 (Extended Data Fig. 8j), suggesting that DR-18 expands a reservoir of stem-like and activated T cells outside the tumour.
DR-18 drives anti-tumour NK responses
Downregulation or loss of MHC class I is a common feature of human cancers and a resistance mechanism to immunotherapy27. Though loss of MHC class I renders tumour cells sensitive to NK-mediated cytotoxicity, NK cells readily become exhausted in the TME28. We investigated whether DR-18 treatment could restore NK cell function in MHC class I-deficient tumours, which are refractory to treatment with ICI (Fig. 4a, Extended Data Fig. 9a–e). In MC38-B2m−/−, YUMMER1.7-B2m−/− and RMA-S (tapasin-deficient) tumours, treatment with DR-18 resulted in tumour regression in 30–75% of mice (Fig. 4a, Extended Data Fig. 9a–e). Antibody depletion studies indicated that NK1.1+ cells were essential for the anti-tumour activity of DR-18 in this setting (Fig. 4a, Extended Data Fig. 9b, d). Analysis of the intratumoral NK1.1+ population after treatment revealed an increase in the number of NK cells, with minimal NKT cells or type 1 innate lymphoid (ILC1) cells present (Extended Data Fig. 9f).
a, Kaplan–Meier survival curve of wild-type mice engrafted s.c. with MC38-B2m−/− tumours and treated twice weekly with PBS, anti-PD-1 plus anti-CTLA-4, IL-18, DR-18, or DR-18 plus anti-CD8 and anti-NK1.1 antibodies. b, c, t-SNE projections showing Ncr1+ cells present in YUMMER1.7 and YUMMER1.7-B2m−/− tumours coloured by unique cluster (b) and by treatment (c). d, Heatmaps showing mean expression of curated genes for each of the NK cell clusters shown in b. e, f, Quantification of surface KLRG1 (e) and CD27/CD11b staining (f) by percentage on intratumoral NK cells from the indicated treatment groups. g–i, Representative flow plots (g), absolute numbers (h) and frequency (i) of intratumoral polyfunctional NK cells measured by co-staining for intracellular IFN-γ and granzyme B following ex vivo stimulation with PMA and ionomycin. Data are representative of three independent experiments with n = 5 mice per group and presented as mean ± s.e.m. P values were calculated using two-tailed unpaired Student’s t-tests (e) and two-way ANOVA (f, h).
scRNA-seq profiling of YUMMER1.7-B2m−/− tumours treated with DR-18 or vehicle identified four clusters of Ncr1+ cells (NK_1 to NK_4; Fig. 4b, Extended Data Fig. 9g, see Methods). Unlike vehicle, DR-18 treatment induced NK_2 cluster cells (Fig. 4b, c), which exhibited increased expression of effector molecule transcripts (Ifng, Prf1, Gzmb), activating transcription factors (Irf8, Zeb2, Cbfb), and a distinct Ly49 repertoire (Fig. 4d). Flow cytometry analysis showed an increase in NK cell maturation with enhanced proliferation and polyfunctionality following DR-18 treatment (Fig. 4e–i, Extended Data Fig. 9h–j). Shedding light on the mechanism of NK-mediated tumour killing elicited by DR-18, administration of neutralizing IFN-γ antibodies completely ablated DR-18 anti-tumour responses, whereas FasL antagonist antibodies or perforin (Prf1) deficiency had no effect (Extended Data Fig. 9k, l).
Human IL-18 engineered for IL-18BP resistance
To assess the feasibility of translating the DR-18 approach to human IL-18, we developed a functional human (h) DR-18 through a parallel directed evolution process. Surface plasmon resonance (SPR) showed that hDR-18 bound strongly to IL-18Rα, but not to IL-18BP, for both human and cynomolgus (Macaca fascicularis) proteins (Extended Data Fig. 10a). Similarly, hDR-18 stimulated NFκB signalling in hIL-18 reporter cells, but was not inhibited by IL-18BP (Extended Data Fig. 10b, c). Finally, hDR-18 elicited IFN-γ production in both human and cynomolgus peripheral blood mononuclear cells (Extended Data Fig. 10d, e).
Discussion
The powerful activity of DR-18 in mouse tumour models highlights the potential of the IL-18 pathway to enhance anti-tumour immunity. DR-18 expands stem-like TCF1+ precursor CD8+ T cells and biases their differentiation towards polyfunctional TEFF cells and away from TOX+ TEX cells. This mechanism appears distinct from the effect of blocking PD-1, which augments the effector function of TEX cells, but does not affect the numbers of stem-like CD8+ T cells, as we found here and was previously described21. DR-18 also promotes NK cell responses against ICI-refractory tumours that have lost MHC class I expression, a major resistance mechanism that is not addressed by currently approved immunotherapies.
The efficacy of DR-18 therapy contrasts with prior inferences about IL-18 biology that have been made on the basis of mouse genetics and pharmacologic studies with rIL-18. IL-18 is generally not required for tumour immunosurveillance29 and some reports have suggested that IL-18 has tumour-promoting roles30,31,32,33. However, pleiotropy is a common feature of cytokines, and factors such as dose, schedule, and site of action can greatly affect their biological activity. For example, low doses of IL-2 can be used to therapeutically expand immunosuppressive Treg cells34, but high doses stimulate CD8+ T cells for tumour immunotherapy35. Thus, our results do not necessarily contradict these previous findings, but rather highlight aspects of IL-18 pleiotropism that can be optimally tuned for cancer immunotherapy.
Dysregulation of IL-18 is associated with numerous autoinflammatory diseases, which are characterized by increases in systemic concentrations of free IL-18 relative to IL-18BP10,. Similarly, a case of biallelic loss of IL18BP was recently associated with fulminant viral hepatitis36. It is therefore conceivable that DR-18 therapy may evoke greater toxicity than was seen previously in clinical trials of rIL-18. Nevertheless, our preclinical studies thus far indicate that DR-18 is efficacious and well-tolerated across a wide range of bioactive doses from 0.001 to 1.0 mg kg−1.
In summary, our results highlight a crucial role of IL-18BP as a secreted immune checkpoint that fundamentally alters the biological effects of IL-18. The ability of DR-18 to act on CD8+ TEFF cells, stem-like TCF1+ CD8+ T cells, and NK cells provides a strong rationale for the clinical development of DR-18 and other IL-18 receptor agonists.
Methods
Patient specimens
Human plasma samples were collected at the University of Navarra in Pamplona, Spain, from two cohorts: cohort 1 included 22 healthy donors and cohort 2 included 52 patients with NSCLC who were treated with anti-PD-1 or anti-PD-L1 monoclonal antibodies. Samples from patients with NSCLC were collected sequentially at the time of first visit to the clinic (baseline) and after starting anti-PD-1 treatment (approximately 2–3 weeks later). Complete baseline and treatment characteristics of the patients are summarized in Supplementary Table 1. The study protocols were approved by the Institutional Review Board at the University of Navarra (approval 11/2010) and all patients provided written informed consent.
Mice
Mice were maintained at Yale University in accordance with the guidelines of the Institutional Animal Care and Use Committee (IACUC). The following mouse strains were used this study: WT C57BL/6 (Charles River, C57BL/6NCrl or Jackson Labs, C57BL/6J 000664), BALB/c (Charles River, BALB/cAnNCrl), Il18bp−/− and Il18r1−/− (Yale, R. Flavell), Rag2−/− (Yale, D. Schatz), XCR1DTR and TCRP14 Transgenic (Yale, A. Iwasaki) and Perforin−/− (Yale, M. Bosenberg). Experiments were conducted using age- and gender-matched mice in accordance with approved institutional protocols.
Cell lines
The following tumour cell lines were used in this study: YUMMER1.7 (Yale, M. Bosenberg)37, MC38 (Yale, M. Bosenberg), RMA-S cell line (Yale, C. Rothlin)28, CT-26 (ATCC, CRL-2638), B16-F10 (ATCC, CRL-6475) and B16-F10GP33 (Yale, A. Iwasaki). The YUMMER1.7 cell line was cultured in DMEM/F12 (Thermo Fisher, 11320-033) including l-glutamine and 2.438 g/l sodium bicarbonate and supplemented with 1 × non-essential amino acids (Thermo Fisher, 11140050), 10% fetal bovine serum (Thermo Fisher, 16140-071) and 1× penicillin and streptomycin (Thermo Fisher, 15140-122). All other cell lines were cultivated in RPMI-1640 (Thermo Fisher, 21870-076) supplemented with 10% fetal bovine serum and 1 × penicillin and streptomycin. All cells were cultured at 37 °C, 5% CO2 and kept at low passage (~3–5 passages) once obtained from vendors or collaborators.
MC38-B2m−/− and YUMMER1.7- B2m−/− cell lines were generated using CRISPR–Cas9 technology. In brief, B2m guides (forward: CACCGAGTATACTCACGCCACCCAC; reverse: AAACGTGGGTGGCGTGAGTATACTC) targeting exon 2 of the B2m gene were cloned into a Cas9-single guide RNA (sgRNA) expression vector (PX458, Addgene, 48138) with a GFP reporter. The plasmid was transiently transfected into each parental line using Lipofectamine 3000 (Fisher Scientific, L3000-001) according the manufacturer’s instructions. Forty-eight hours after transfection, GFP-positive cells were sorted (BD FACS Aria) as single cells into 96-well plates. B2m deletion was confirmed separately in each clone using flow cytometry showing a lack of MHC class I surface staining following 12 h of in vitro incubation with recombinant murine IFN-γ (Peprotech, 315-05) at 100 U/ml. Single cell-derived clones of each line that exhibited similar morphology, in vitro growth characteristics, and in vivo tumour formation characteristics to the parental lines were selected for experiments.
Stable H2B–GFP-expressing YUMMER1.7 and YUMMER1.7-B2m−/− lines were generated and used for scRNA-seq experiments. In brief, lentivirus was produced using pCMV-VSV-G (Addgene, 8454) and psPAX2 (Addgene, 8454) with an H2B–GFP plasmid (Addgene, 25999) in 293T cells. 293T cells were transfected with 10 μg of a 1:2:4 (VSV-G: PAX-2: H2B-GFP) mixture of plasmid DNA in X-treme GENE 9 transfection reagent (Sigma, 6365779001) according to the manufacturer’s instructions in Opti-MEM (Thermo Fisher, 31985088). After overnight incubation, the medium was replaced with normal growth medium (DMEM) and incubated for 48 h, after which time the medium was harvested and filtered through a 0.45-μM filter. Untitred viral supernatant was applied to YUMMER1.7 and YUMMER1.7-B2m−/− lines for 48 h. Single GFP+ tumour cells were sorted into individual wells of 96-well plates by fluorescence-activated cell sorting (FACS) (BD FACS Aria). Single-cell-derived clones of YUMMER1.7 and YUMMER1.7-B2m−/− expressing GFP that exhibited similar morphology, in vitro growth characteristics, and in vivo tumour formation characteristics to the parental lines were selected for experiments.
Cell lines tested negative for mycoplasma contamination using a universal mycoplasma detection kit (ATCC 30-1012K).
ELISA
IL-18BP and IFN-γ ELISAs were performed using Human IL-18 BPa Quantikine ELISA Kit (R&D system, DBP180), mouse IL-18BPd DuoSet ELISA kit (R&D system, DY122-05), mouse IFN-γ ELISA MAX Deluxe kit (Biolegend, 430804), human IFN-γ Quantikine ELISA Kit (R&D system, DIF150), and primate IFN-γ DuoSet ELISA (R&D system, DY961) according to the manufacturers’ instructions.
Immunohistochemistry
Human tumour tissue microarrays (TMAs) were obtained from the Yale Tissue Microarray Facility. IL-18BP immunohistochemical staining of the TMAs was performed by the Yale Dermatopathology laboratory using anti-IL-18BP antibody clone EP1088Y (Abcam) and was previously validated38,39. The melanoma TMA was stained with Azure blue so that melanin (turned green) could be differentiated from the DAB chromagen (brown). All scorable tumour cores were included in this analysis. Melanoma TMA (YTMA-192) contained 282 scorable tumour cores. Breast cancer TMA (YTMA-353) contained 114 scorable tumour cores. Head and neck cancer TMA (YTMA-305) contained 76 scorable tumour cores. Gastric cancer TMA (YTMA-141) contained 62 tumours scorable tumour cores. Ovarian TMA (YTMA-264) contained 226 scorable tumour cores. Where available, cell lines and normal tissue on the TMAs were used as controls. Scoring was performed by a board-certified pathologist (M.W.B.) in a blinded fashion. Cores were scored as negative (0) or positive (1+, 2+, or 3+).
Mouse IL-18BP immunohistochemical staining was performed on Il18bp−/− spleen, wild-type spleen (IL-18 treated) and tumour (MC38) using anti-IL-18BP antibody (clone EP1088Y; Abcam). Tissue was fixed in 4% PFA overnight on ice. Post-fixation samples were embedded in paraffin and sectioned at 5 μm before staining. The number of IL-18BP-positive cells per high-power field was quantified in representative sections from each condition.
mRNA quantification
Whole blood and tumour samples were harvested in Trizol and total RNA was extracted using the RNeasy kit (Qiagen, Q74104) according to the manufacturer’s instructions. The total RNA was reverse transcribed using Oligo(dT) primers and Maxima H Minus Reverse Transcriptase (Thermo Fisher, EP0752). Il18bp expression was assayed by real-time PCR using iQ SYBR Green Supermix (Bio-rad, 1708880) with primers (forward: GAGGGCCACACAAGTCGC; reverse: GCTGGGCCAGAATGATGTGA) on an ABI 7900HT real-time PCR system. Actb (β-actin) was used as a control (forward: AGGTGACAGCATTGCTTCTG; reverse: GCTGCCTCAACACCTCAAC). All values were normalized to expression levels in naive mouse spleen. Measurements were performed in triplicate for each sample.
Protein expression, purification and biotinylation
The mature form of mouse IL-18 (amino acids 37–193) and associated variants were assembled as gene blocks (Integrated DNA Technologies, IDT) and cloned into a pTZ-SH vector for expression of N-terminal SUMO-tagged and C-terminal hexahistidine-tagged proteins in Escherichia coli BL21 (DE3) Rosetta strain (Fisher Scientific, 70954-3). Protein expression was induced with 0.5 mM IPTG at 16 °C for 20 h. The fusion proteins were first purified using an Ni-NTA column (Fisher Scientific, P188223), followed by removal of the SUMO tag with the SUMO protease Ulp1. Next, the protein solution was buffer-exchanged to remove imdizaole and re-applied to a second Ni-NTA column to remove the free SUMO tag. The eluted protein was concentrated and separated from aggregates by gel-filtration (Column SEC70, Bio-rad). Protein from the mono-disperse peak was pooled and loaded on a final Ni-NTA column for endotoxin removal with 0.2% Triton X-114 at 4 °C. Finally, the eluted protein was buffer exchanged into sterile, endotoxin free PBS using a PD-10 column (GE Healthcare) and flash-frozen in liquid nitrogen for long-term storage at −80 °C.
The mouse IL-18Rα ectodomain (amino acids 19–329) and IL-18 binding protein (IL-18BP, amino acids 31–194) were expressed by transient transfection of Expi293 cells (Thermo Fisher). Both sequences were cloned into the pEZT_D_Lux vector with an N-terminal H7 signal peptide and a C-terminal AviTag and hexahistidine tag. Plasmids were transfected into Expi293 cells by ExpiFectamine 293 Transfection Kit (Thermo Fisher, A14524) per the manufacturer’s instructions. Cells were harvested 3–5 days after transfection. Proteins were captured from cell supernatant via Ni-NTA chelating resin and further purified by size exclusion chromatography (Column SEC70, Bio-rad) into a final buffer of HEPES buffered saline (HBS; 10 mM HEPES, pH 7.5, 150 mM NaCl).
For protein biotinylation, proteins were expressed with a C-terminal biotin acceptor tag (AviTag)-GLNDIFEAQKIEWHE. After Ni-NTA chromatography, protein biotinylation was carried out at room temperature for 2 h with soluble BirA ligase enzyme in 0.1 mM bicine (pH 8.3), 10 mM ATP, 10 mM magnesium acetate, and 0.5 mM biotin (Avidity, BIO500). Biotinylated proteins were then purified by gel-filtration as described above. Biotinylation efficiency was assessed using an SDS/PAGE streptavidin-shift assay.
Yeast display of IL-18
The nucleotide sequence of the mature form of mouse IL-18 (amino acids 37–193) was cloned into an N-terminal displayed vector, pYAL, and displayed on the surface of yeast strain Saccharomyces cerevisiae EBY100. Yeast were maintained and expanded in liquid synthetic dextrose medium with casamino acids (SDCAA) at 30 °C and then induced to express IL-18 in liquid synthetic glucose medium with casamino acids (SGCAA) at 20 °C for 24–48 h. The displayed protein level was verified by staining with a C-terminal Myc tag antibody (Cell Signaling Technology, 3739S). Biotinylated IL-18Rα and IL-18BP binding was detected using a fluorescent streptavidin phycoerythrin secondary and quantified by flow cytometry using a Sony SA3800 flow cytometer.
Mouse IL-18 library construction and selection
Thirteen residues in IL-18 which were in contact with both IL-18Rα and IL-18BP were identified by aligning the structure of hIL-18–hIL-18Rα–hIL-18Rβ complex (Protein Data Bank (PDB) ID 3WO4) to the structure of hIL-18: ectromelia virus IL-18BP complex (PDB ID 3F62). A library randomizing these residues was constructed using assembly PCR with the degeneration primers. The PCR products were further amplified with primers containing homology to the pYAL vector and co-electroporated into EBY100 competent yeast together with linearized pYAL vector. The resulting library was later measured to contain 4.0 × 108 transformants.
Transformed yeast were recovered and expanded in SDCAA medium at 30 °C, induced by 1:10 dilution into SGCAA medium and cultured at 20 °C for 24–48 h. The appropriate numbers of induced yeast were used in each round to ensure at least tenfold coverage of the expected diversity, and not less than 108 cells. All selection steps were carried out at 4 °C using PBE buffer (PBS with 0.5% BSA and 2 mM EDTA). For round 1, the yeast library was counter-selected with anti-Cy5/Alexa Fluor 647 microbeads (Miltenyi, 130-091-395) with an LS MACS column (Miltenyi, 130-042-401) to remove non-specific binders. Positive selection was performed by labelling yeast with 1 μM biotinylated IL-18Rα, followed by magnetic selection with Alexa Fluor 647 microbeads and the LS MACS column. For round 2, counter-selection reagent was changed to 1 μM biotinylated IL-18BP while the IL-18Rα concentration was kept at 1 μM. For rounds 3–5, selection was performed by incubating yeast with Alexa Fluor647-conguated IL-18Rα at concentrations of 100 nM (round 3), 100 nM (round 4), or 10 nM (round 5) in the presence of 250 nM pre-formed and biotin-capped IL-18BP:SA-PE tetramers. IL-18 display levels were determined by staining with Alexa Fluor 488-conjugated anti-Myc (Cell Signaling Technologies, 2279S). Yeast were selected by FACS sorting with a Sony SH800 cell sorter by excluding IL-18BP (PE) binders and gating the top 1% of display-normalized IL-18Rα binders. After each round of selection, recovered yeast were expanded in SDCAA medium at 30 °C overnight and later induced at 20 °C by a 1:10 dilution into SGCAA medium for 24–48 h.
Surface plasmon resonance
SPR experiments were conducted using a Biacore T100 and carried out at 25 °C. Interactions were measured using either conventional multiple-cycle programs or a single-cycle kinetics program. Mouse, human, or cynomolgus biotinylated IL-18Rα or IL-18BP were immobilized onto a Biacore biotin capture chip (Series S CAP sensor chip, GE Healthcare) to yield a Rmax of ∼50 RU (IL-18Rα) or ~10 RU (IL-18BP). Measurements were made with half-log dilutions of the IL-18 variants in HBS-P+ buffer (10 mM HEPES pH 7.4, 150 mM NaCl, 0.005% surfactant P20). The surface was regenerated by three 60-s injections of regeneration buffer (3/4 (v/v) 8 M guanidine hydrochloride +1/4 (v/v) 1 M sodium hydroxide). Experiments were performed in multiple channels for duplicate measurements (F2-1 and F4-3). All data were analysed with the Biacore T100 evaluation software version 2.0 with a 1:1 Langmuir binding model.
Isolation of lymphocytes
Spleens were dissociated using a 70-μm cell strainer and red blood cells were lysed using ACK buffer (Thermo Fisher, A1049201). Lymph nodes were dissociated using a 70-μm cell strainer. Tumours were cut into small pieces in the presence of 3 ml RPMI-1640 supplemented with 1% FBS. Collagenase IV (Gibco, 17104019, final concentration 1 mg/ml) and Dnase I (Roche, 10104159001, final concentration 0.2 mg/ml) were added and samples were incubated at 37 °C for tissue digestion. After 30 min of digestion, 6 ml of RPMI-1640 with 10% FBS was added to neutralize protease activity and tumour tissues were forced through 70-μm cell strainers to prepare single-cell suspensions. Cells were then washed twice and resuspended in 1–3 ml of RPMI-1640 with 1% FBS medium for downstream analysis. Cell concentrations were counted using a Beckman Coulter particle counter.
Ex vivo stimulation of splenocytes and PBMCs
For the mouse IL-18/DR-18 stimulation assay, approximately 1 × 106 mouse splenic lymphocytes were stimulated ex vivo with a range of concentrations of mouse IL-18 or DR-18 from 0.00316 to 316 ng/ml. For the mouse IL-18BP inhibition assay, 1 × 106 splenic lymphocytes were stimulated ex vivo with a fixed concentration of IL-18 (1 nM) or DR-18 (0.5 nM) and range of IL-18BP concentrations (0.01 to 100 nM). All stimulations were performed in the presence of mIL-12 (10 ng/ml, Peprotech, 210-12) and protein transport inhibitor (1:200, BD Golgiplug, 554724) for 4 h in RPMI containing 10% fetal bovine serum at 37 °C. Cells cultured with IL-12 alone were used as negative controls, and PMA/ionomycin was used as a positive control. After 4 h, IFN-γ expression was measured on NK cells by flow cytometry, gating on CD3−/NK1.1+. Data were normalized by subtracting the background MFI from the unstimulated control and defining the largest mean value as 100%.
For human and cynomolgus peripheral blood mononuclear cell (PBMC) stimulation assays, approximately 0.6 × 106 human (PPA, 15-00012) or 0.3 × 106 cynomolgus (BioIVT, M3-010-C20M) PBMCs were stimulated ex vivo with gradient of human IL-18 or DR-18 ranging from 0.0316 to 1,000 ng/ml. All stimulations were performed in the presence of hIL-12 (10 ng/ml, Peprotech, 200-12H) for 12 h in RPMI containing 10% fetal bovine serum. IFN-γ production after stimulation was measured using corresponding human and cynomolgus ELISA kits per the manufacturers’ instructions.
Human IL-18 reporter assay
The IL-18 HEK-Blue assay (InvivoGen, hkb-hmil18) was performed according to the manufacturer’s instructions. In brief, 5 × 104 cells were seeded into each well of a 96-well plate and stimulated with 0–100 ng/ml of hIL-18 or DR-18 for 24 h at 37 °C and 5% CO2. Forty microlitres of cell culture supernatant was then taken from each well and mixed with 160 μl QUANTI-Blue medium in a 96-well plate, incubated for 3 h at 37 °C and 5% CO2, and then read in a microplate reader at 655 nm.
Tumour treatment studies
The numbers of tumour cells engrafted were as follows: 0.5 × 106 MC38 cells, 0.5 × 106 YUMMER1.7 cells, 0.5 × 106 MC38-B2m−/− cells, 1 × 106 YUMMER1.7-B2m−/− cells, 1 × 106 RMA-S cells, 0.25 × 106 CT26 cells, 0.25 × 106 B16-F10 cells, and 0.25 × 106 B16-F10GP33 cells. Tumours were engrafted subcutaneously into the flanks of 8–10-week-old age-matched female or male mice. YUMMER1.7 and YUMMER1.7-B2m−/− cells were implanted only into male mice, as this cell line was derived from a male mouse. Treatment was initiated when mean tumour size was between 50 and 100 mm3 (usually at day 7 post engraftment); mice with tumours smaller than 30 mm3 or larger than 150 mm3 were excluded from randomization. Remaining mice were randomized into designated groups to ensure an approximately equal average tumour size. Mice were then treated with the designated test articles by intraperitoneal injection twice weekly for a total of 3–5 doses (as indicated in the text). Pilot dose-finding studies with MC38 tumours indicated that DR-18 treatment resulted in tumour growth inhibition (TGI), tumour regression, and clearance at doses as low as 10 μg/kg and at schedules as infrequent as administration once every two weeks, with 0.32 mg/kg given bi-weekly representing the maximally efficacious regimen. Test articles were diluted in sterile PBS and dosed as follows: anti-PD-1 (RMP1-14, Bio X Cell) 8 mg/kg, anti-CTLA-4 (9H10, Bio X Cell) 8 mg/kg, IL-18 0.32 mg/kg and DR-18 0.32 mg/kg administered twice weekly. Control groups were treated with sterile PBS or isotype control antibodies. Tumour growth was tracked twice weekly by caliper measurement. Tumour volume was calculated using volume = 0.5 × length × width × width. Mice were euthanized when tumours reached endpoints (volume greater than or equal to 1,000 mm3 (MC38, CT26, and B16-F10) or 500 mm3 (YUMMER1.7 and RMA/S), or volume greater than or equal to 500 mm3 (MC38, CT26, and B16-F10) or 250 mm3 (YUMMER1.7 and RMA/S) plus tumour ulceration). Survival analyses reflect this endpoint.
For immune cell depletion (CD4/CD8/NK) and effector molecule neutralization (IFN-γ/FasL) studies, antibodies against CD8a (2.43, Bio X cell or TIB210, Bio X cell), CD4 (GK1.5, Bio X cell), NK1.1 (PK136, Bio X Cell), IFN-γ (R4-6A2, Bio X or XMG1.2, Bio X cell), and FasL (MFL3, Bio X cell) were used. Antibodies were administered by intraperitoneal injection starting on day 6 (one day before therapy initiation) and were continued twice weekly for the duration of the experiment. All depleting antibodies were used at 8 mg/kg per treatment. Lymphocyte depletions were confirmed in peripheral blood lymphocytes by flow cytometry with the following antibodies: CD8a (53-6.7), CD4 (RM4-5) and NKp46 (29A1.4).
For tumour re-challenge studies, mice exhibiting complete tumour regression as a result of DR-18 treatment were re-inoculated subcutaneously with twice the initial dose of MC38 tumour cells (1 × 106) 30 days after the initial tumours were cleared. As a control, naive C57BL/6J mice were challenged at the same time. Tumour growth and survival were monitored twice weekly as stated above for up to 60 days.
For ablation of cDC1 studies, WT and XCR1DTR were injected i.p., with 25 ng diphtheria toxin (DT) (150, List Biological Lab) per gram of body weight on day 6 post tumour engraftment. To maintain DT ablation, mice received 100 ng DT per gram of body weight twice weekly after the initial DT injection. cDC1 depletion was confirmed by flow cytometry.
For FTY720 experiments, FTY720 (S5002, Selleck Chemicals) was reconstituted in water (10 mg/ml) and diluted in PBS. Wild-type mice were treated i.p. with 3 mg/kg starting on day 6 (one day before therapy initiation) and continuing twice weekly together with DR-18 treatment for the duration of the experiment. FTY720 efficacy was confirmed by measuring the reduction in CD3+ T cells in the blood.
All animal experiments were randomized immediately prior to initiation of treatment. The investigators were not blinded to allocation during experiments and outcome assessment.
Adoptive transfer experiments
CD3+ naive T cells were purified from the spleen and lymph nodes of wild-type or Il18r1−/− mice using a mouse pan-naive T cell isolation kit (Stemcell, 19848) by magnetic based bead sorting. Purified T cells were then transferred into Rag2−/− mice (nine million cells per recipient) by retro-orbital intravenous injection. Tumour treatment studies were then conducted as described above.
Immunophenotyping of tumour-engrafted mice
Mice with MC38, MC38-B2m−/−, YUMMER1.7, and YUMMER1.7-B2m−/− tumours for immunophenotyping were engrafted and treated in the same way as for tumour growth studies. Mice were euthanized 24 h after the third dose of saline, anti-PD-1, IL-18, or DR-18 (day 15 after engraftment), or at indicated time point, and spleens, draining lymph nodes, and tumours were harvested for analysis. Tumours were dissociated for analysis as previously described. Cell surface staining of single-cell suspensions from spleens and tumours was performed using fluorophore-conjugated antibodies (BD Biosciences, eBioscience, BioLegend, Tonbo, & R&D Systems; see below). All samples were stained with Zombie Yellow (Biolegend, 423104) to identify dead cells and incubated with Fc receptor blocking antibody (Fisher Scientific, 553142). Intracellular staining was performed by fixing and permeabilizing with the eBioscience Foxp3/Transcription Factor Staining Set (Thermo Fisher, 00-5523-00) or Cytofix/Cytoperm kit (BD Biosciences, 554714). For endogenous cytokine production measurement, single-cell suspensions of splenocytes and tumour cells were incubated with Golgi-plug (1:500, BD Biosciences, 555029) for 4 h at 37 °C. For polyfunctional cytokine production measurement, single-cell suspensions of splenocytes and tumour cells were stimulated with a PMA/ionomycin cocktail (1:200, Biolegend, 423304) and incubated for 4 h at 37 °C. Cells cultured in medium alone (without stimulation) were used as a negative control.
Cell staining was performed using the following fluorophore-conjugated antibodies: CD45 (30-f11), CD8α (53–6.7), CD4 (RM4-5), TCRβ (H57-597), CD3e (17A2), CD19 (ID3), NK1.1 (PK136), IL-18Rα (P3TUNYA), B2m (A16041A), Ki67 (16A8), KLRG1 (2F1), IFN-γ (XMG1.2), GZMB (QA16A02), TNF-α (MP6-XT22), CD107A (1D4B), PD-1 (RMP1-30 and 29F.1A12), TIM3 (RMT3-23), CD44 (IM7), CD62L (MEL-14), CD103 (2E7), CD11b (M1/70), CD27 (LG.3A10), EOMES (Dan11mag), FOXP3 (FJK-16 s), CD200R1 (OX-110), CD49b/DX5 (DX5), F4/80 (BM8.1), CD64 (X54-5/7.1), CD11c (N418), Ly6C (HK1.4), Ly6G (1A8), I-A/I-E (M5/114.15.2), CD90.1 (OX-7), CD39 (24DMS1), NOS2 (CXNFT), TCF1 (C63D9) and TOX (REA473). Fluorescence spectra were acquired using a LSRII flow cytometer and analysed by FlowJo (Version 10). For flow cytometry analysis, naive mouse splenocytes or fluorescent minus one (FMO) staining was used for gating.
scRNA-seq sample preparation
Biological replicates from n = 3 pooled mice were processed for each experimental condition from YUMMER1.7 and YUMMER1.7-B2m−/− tumours. Biological replicates were then pooled together at the single-cell suspension stage with an equivalent number of cells from each replicate. The following populations were sorted and purified: P1: GFP−CD45+CD3+ (T cells), P2: GFP−CD45+CD3− (non-T immune cells), P3: GFP+/−CD45−CD3− (tumour and stromal cells). P1, P2, and P3 for each sample were then mixed back together at a 2:1:1 ratio, respectively. Five thousand cells from each of the mixed sorted samples for each condition were loaded onto the 10x Genomics Chromium System. Library preparation was performed using 10x Genomics reagents according to the manufacturer’s instructions by the Yale Center for Genome Analysis (YCGA) and passed quality control. Libraries were sequenced using an Illumina HiSeq 4000 (one library per lane) at the YCGA.
scRNA-seq analysis
Samples were processed using the Cellranger software suite commands cellranger mkfastq for processing raw call files into fastq files. Cellranger count was used to align reads to a custom mm10 reference modified to include eGFP (marking tumour cells), to filter reads, and to generate a cell-by-gene matrix for each sample library. Libraries were aggregated using cellranger aggr without normalization to generate a single cell-by-gene matrix. Based on Gapdh expression, the top 14,000 cells ranked by nUMI were retained for analysis. The Seurat package for R v.2.3.440 was used to process the matrix and perform downstream analysis. Expression values were log-normalized with a scaling factor of 104, and the 2,509 most variable genes were detected and used for further analysis with the FindVariableGenes function. Values were scaled to number of UMIs and percent mitochondrial genes, and principle component analysis (PCA) was performed on the most variable genes. The FindClusters command was used to perform a shared nearest neighbour (SNN) modularity optimization-based clustering algorithm using a resolution of 1.0, and tSNE dimensional reduction was calculated on the first 50 principle components to visualize data. Clusters consisting of cells with low/null expression of Gapdh and Eno1 (non-cells), or co-expression of cell-type exclusive markers (doublets) such as Cd3e and Cd68 were removed from further analysis by the SubsetData command, and variable genes were re-identified, data were re-scaled and PCA clustering and tSNE were re-run as described. Clusters containing the following cell types were identified using cell type markers: Tumour cells (eGFP), Myeloid cells (Cd68), NK cells (Ncr1), T cells (Cd3e), Neutrophils (Lcn2), and subsets of these groups were identified by markers noted in heatmaps (Extended Data Fig. 6d). Cell type assignments for each cluster were verified by comparing with ImmGen data sets41. T cells, NK cells, and myeloid cells were subsetted and re-analysed separately as described above. Cluster frequencies by library were normalized to the number of cells per library and column plots were generated using ggplot2 v. 3.2.0 (Extended Data Fig. 6c). Gene expression t-SNE plots were plotted using ggplot2 v 3.2.0. For heatmaps, mean scaled expression values of each gene were calculated per cluster and plotted using pheatmap v 1.0.12 with values scaled by row (gene). Cell cycle scoring was performed using the Seurat CellCycleScoring command using mouse gene sets orthologous to previously described human gene sets42.
Analysis of TCGA data
IL18BP expression in individual cancer versus counterpart normal tissues was analysed using TCGA cancer databases. Median and mean values were calculated. Human IL18BP mRNA differentiated expression, correlation with CD3E, CD8A and PDCD1 data for multiple cancers and matched normal tissues were obtained from TCGA and the GTEx projects and analysed by webserver GEPIA2. The original microarray data was normalized by cancer browser (http://xena.ucsc.edu/welcome-to-ucsc-xena/) and then analysed using R. The error bars in figures represent s.e.m.
Statistical analysis
Statistical analyses were conducted using R v.3.4.0 and Prism 8 (GraphPad Software). Pearson’s r was calculated with the R function cor (). Ordinary one-way or two-way ANOVA with Tukey’s multiple comparisons test, or a two-tailed paired or unpaired Student’s t-test (see Figure legends) were used to determine statistical significance (*P < 0.05, **P < 0.01, ***P < 0.001, ****P < 0.0001). The mean and s.e.m. are presented in the Figures. No statistical methods were used to predetermine sample size.
Reporting summary
Further information on research design is available in the Nature Research Reporting Summary linked to this paper.
Data availability
All data generated during this study are available within the paper. The scRNA-seq data were deposited on Gene Expression Omnibus (GSE146609). Source data are provided with this paper.
References
Atkins, M. B., Kunkel, L., Sznol, M. & Rosenberg, S. A. High-dose recombinant interleukin-2 therapy in patients with metastatic melanoma: long-term survival update. Cancer J. Sci. Am. 6 (Suppl. 1), S11–S14 (2000).
Waldmann, T. A. Cytokines in cancer immunotherapy. Cold Spring Harb. Perspect. Biol. 10, a028472 (2018).
Tarhini, A. A. et al. A phase 2, randomized study of SB-485232, rhIL-18, in patients with previously untreated metastatic melanoma. Cancer 115, 859–868 (2009).
Spangler, J. B., Moraga, I., Mendoza, J. L. & Garcia, K. C. Insights into cytokine-receptor interactions from cytokine engineering. Annu. Rev. Immunol. 33, 139–167 (2015).
Mantovani, A., Dinarello, C. A., Molgora, M. & Garlanda, C. Interleukin-1 and related cytokines in the regulation of inflammation and immunity. Immunity 50, 778–795 (2019).
Guo, L., Junttila, I. S. & Paul, W. E. Cytokine-induced cytokine production by conventional and innate lymphoid cells. Trends Immunol. 33, 598–606 (2012).
Ma, Z. et al. Augmentation of immune checkpoint cancer immunotherapy with IL18. Clin. Cancer Res. 22, 2969–2980 (2016).
Hu, B. et al. Augmentation of antitumor immunity by human and mouse CAR T cells secreting IL-18. Cell Rep. 20, 3025–3033 (2017).
Robertson, M. J. et al. Clinical and biological effects of recombinant human interleukin-18 administered by intravenous infusion to patients with advanced cancer. Clin. Cancer Res. 12, 4265–4273 (2006).
Dinarello, C. A., Novick, D., Kim, S. & Kaplanski, G. Interleukin-18 and IL-18 binding protein. Front. Immunol. 4, 289 (2013).
Robertson, M. J. et al. A dose-escalation study of recombinant human interleukin-18 using two different schedules of administration in patients with cancer. Clin. Cancer Res. 14, 3462–3469 (2008).
Kim, S. H. et al. Site-specific mutations in the mature form of human IL-18 with enhanced biological activity and decreased neutralization by IL-18 binding protein. Proc. Natl Acad. Sci. USA 98, 3304–3309 (2001).
Simoni, Y. et al. Bystander CD8+ T cells are abundant and phenotypically distinct in human tumour infiltrates. Nature 557, 575–579 (2018).
Salmon, H. et al. Expansion and activation of CD103+ dendritic cell progenitors at the tumor site enhances tumor responses to therapeutic PD-L1 and BRAF inhibition. Immunity 44, 924–938 (2016).
Apetoh, L. et al. Consensus nomenclature for CD8+ T cell phenotypes in cancer. OncoImmunology 4, e998538 (2015).
Best, J. A. et al. Transcriptional insights into the CD8+ T cell response to infection and memory T cell formation. Nat. Immunol. 14, 404–412 (2013).
Wherry, E. J. & Kurachi, M. Molecular and cellular insights into T cell exhaustion. Nat. Rev. Immunol. 15, 486–499 (2015).
Scott, A. C. et al. TOX is a critical regulator of tumour-specific T cell differentiation. Nature 571, 270–274 (2019).
Seo, H. et al. TOX and TOX2 transcription factors cooperate with NR4A transcription factors to impose CD8+ T cell exhaustion. Proc. Natl Acad. Sci. USA 116, 12410–12415 (2019).
Kurtulus, S. et al. Checkpoint blockade immunotherapy induces dynamic changes in PD-1−CD8+ tumor-infiltrating T cells. Immunity 50, 181–194.e186 (2019).
Miller, B. C. et al. Subsets of exhausted CD8+ T cells differentially mediate tumor control and respond to checkpoint blockade. Nat. Immunol. 20, 326–336 (2019).
Hudson, W. H. et al. Proliferating transitory T cells with an effector-like transcriptional signature emerge from PD-1+ stem-like CD8+ T cells during chronic infection. Immunity 51, 1043–1058.e1044 (2019).
Murray, P. J. et al. Macrophage activation and polarization: nomenclature and experimental guidelines. Immunity 41, 14–20 (2014).
Gubin, M. M. et al. High-dimensional analysis delineates myeloid and lymphoid compartment remodeling during successful immune-checkpoint cancer therapy. Cell 175, 1014–1030.e1019 (2018).
Jeannet, G. et al. Essential role of the Wnt pathway effector Tcf-1 for the establishment of functional CD8 T cell memory. Proc. Natl Acad. Sci. USA 107, 9777–9782 (2010).
Gattinoni, L. et al. Wnt signaling arrests effector T cell differentiation and generates CD8+ memory stem cells. Nat. Med. 15, 808–813 (2009).
Sade-Feldman, M. et al. Resistance to checkpoint blockade therapy through inactivation of antigen presentation. Nat. Commun. 8, 1136 (2017).
Ardolino, M. et al. Cytokine therapy reverses NK cell anergy in MHC-deficient tumors. J. Clin. Invest. 124, 4781–4794 (2014).
Street, S. E., Trapani, J. A., MacGregor, D. & Smyth, M. J. Suppression of lymphoma and epithelial malignancies effected by interferon gamma. J. Exp. Med. 196, 129–134 (2002).
Nakamura, K. et al. Dysregulated IL-18 is a key driver of immunosuppression and a possible therapeutic target in the multiple myeloma microenvironment. Cancer Cell 33, 634–648.e635 (2018).
Terme, M. et al. IL-18 induces PD-1-dependent immunosuppression in cancer. Cancer Res. 71, 5393–5399 (2011).
Vidal-Vanaclocha, F. et al. IL-18 regulates IL-1beta-dependent hepatic melanoma metastasis via vascular cell adhesion molecule-1. Proc. Natl Acad. Sci. USA 97, 734–739 (2000).
Park, C. C. et al. Evidence of IL-18 as a novel angiogenic mediator. J. Immunol. 167, 1644–1653 (2001).
Hirakawa, M. et al. Low-dose IL-2 selectively activates subsets of CD4+ Tregs and NK cells. JCI Insight 1, e89278 (2016).
Rosenberg, S. A. IL-2: the first effective immunotherapy for human cancer. J. Immunol. 192, 5451–5458 (2014).
Belkaya, S. et al. Inherited IL-18BP deficiency in human fulminant viral hepatitis. J. Exp. Med. 216, 1777–1790 (2019).
Wang, J. et al. UV-induced somatic mutations elicit a functional T cell response in the YUMMER1.7 mouse melanoma model. Pigment Cell Melanoma Res. 30, 428–435 (2017).
Carbotti, G. et al. The IL-18 antagonist IL-18-binding protein is produced in the human ovarian cancer microenvironment. Clin. Cancer Res. 19, 4611–4620 (2013).
Lin, L. et al. Interleukin-37 expression and its potential role in oral leukoplakia and oral squamous cell carcinoma. Sci. Rep. 6, 26757 (2016).
Butler, A., Hoffman, P., Smibert, P., Papalexi, E. & Satija, R. Integrating single-cell transcriptomic data across different conditions, technologies, and species. Nat. Biotechnol. 36, 411–420 (2018).
Heng, T. S. & Painter, M. W. The Immunological Genome Project: networks of gene expression in immune cells. Nat. Immunol. 9, 1091–1094 (2008).
Tirosh, I. et al. Dissecting the multicellular ecosystem of metastatic melanoma by single-cell RNA-seq. Science 352, 189–196 (2016).
Singer, M. et al. A distinct gene module for dysfunction uncoupled from activation in tumor-infiltrating T cells. Cell 166, 1500–1511.e1509 (2016).
Acknowledgements
We thank all members of the Ring and Bosenberg labs for advice and technical assistance; E. Folta-Stogniew for assistance with SPR; and A. Iwasaki for mice and reagents. The cartoon in Fig. 1a was created using BioRender (https://biorender.com/). This work was supported by grants from the National Cancer Institute Immuno-Oncology Translation Network (U01CA233096; to A.M.R. and M.W.B.), Gabrielle’s Angel Foundation (to A.M.R.), and the Blavatnik Fund for Innovation at Yale (to A.M.R.). A.M.R. is additionally supported by an NIH Director’s Early Independence Award (DP5OD023088), the Pew-Stewart Scholars Program, and the Robert T. McCluskey Foundation. M.F.S. is supported by a Miguel Servet contract from Instituto de Salud Carlos III, Fondo de Investigacion Sanitaria (Spain). W.D. is supported by a Career Development Award from the Dermatology Foundation. O.-E.W. is supported by a NSF Graduate Research Fellowship (1752134) and by a NIH training grant (T32AI055403). The T100 Biacore instrumentation used was supported by NIH Award S10RR026992-0110.
Author information
Authors and Affiliations
Contributions
T.Z., W.D., O.-E.W., K.P.H., M.W.B., and A.M.R. designed experiments. T.Z., W.D., O.-E.W., K.P.H., C.E.R., and S.F. performed experiments. T.Z., W.D., O.-E.W., M.K.M., K.P.H., J.W., M.W.B., and A.M.R. analysed data. M.F.S. provided samples from patients with NSCLC. R.J. and R.A.F. provided Il18bp−/− and Il18r1−/− mice. T.Z., O.-E.W., W.D., M.W.B., and A.M.R. wrote the paper. M.W.B. and A.M.R. supervised the research. A.M.R. conceived of the project.
Corresponding author
Ethics declarations
Competing interests
A.M.R., T.Z., and S.F. are named inventors on a patent application that describes the DR-18 molecule. A.M.R. is the founder of Simcha Therapeutics, the commercial licensee of DR-18, and holds equity in the immuno-oncology companies Forty-Seven Inc., ALX Oncology, and Medicenna Therapeutics. W.D. and M.W.B. serve as consultants for Eli Lilly. W.D. has research funding from Pfizer for unrelated work.
Additional information
Peer review information Nature thanks Jim Wells, Dietmar Zehn and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Regulation of the IL-18 signalling axis in the TME.
a, Distribution of intratumoral CD8+ T cell transcripts by dysfunction and activation gene scores, adapted from Singer et al.43. Blue points mark a curated list of cytokine and cytokine receptors. Red points mark Il18, Il18r1, Il18rap. b–f, Wild-type C57BL/6 mice were subcutaneously engrafted with the indicated mouse syngeneic tumour. Tumour and spleen were removed and analysed by flow cytometry at day 7 after engraftment. b, c, Representative flow plots (b) and quantification of percentage of IL-18Rα expression (c) on splenic and intratumoral CD4+ T cells (NK1.1−, CD3+, TCRβ+, CD4+), CD8+ T cells (NK1.1−, CD3+, TCRβ+, CD8+), NK cells (CD3−, TCRβ−, NK1.1+, CD200R1−) from wild-type mice engrafted with MC38 cells. d, Percentage IL-18Rα expression on splenic and intratumoral CD4+ T cells, CD8+ T cells, and NK cells from wild-type mice engrafted with YUMMER1.7 cells. e, f, Representative flow plots (e) and quantification of surface CD44 and IL-18Rα expression (f) on CD8+ T cells in spleen and tumour from MC38 tumour-bearing mice. g, Normalized quantification of Il18bp by qPCR in blood and tumour lysate assessed at day 7 after s.c. engraftment of wild-type mice with MC38 tumours, treated with PBS, IL-18 or anti-IFN-γ + IL-18. h, Quantification of plasma IL-18BP protein level by ELISA at day 7 after s.c. engraftment of wild-type mice with MC38 tumours, treated with either PBS or IL-18. i, Representative immunohistochemistical staining for IL-18BP from l18bp−/− mouse spleen, wild-type spleen, MC38 tumour, or MC38 tumour treated with one dose of IL-18, and assessed on day 7. Scale bar, 25 μm. j, Quantification of IL-18BP+ cells per high power field (HPF) in representative sections from each group in i. b–f, i, Data are representative of two independent experiments with n = 5 mice per group. g, h, j, Data are pooled from two independent experiments with n = 5 mice per group. P values were calculated using two-way ANOVA (c, d, f) or two-tailed unpaired Student’s t-tests (g, h, j), and data are presented as mean ± s.e.m.
Extended Data Fig. 2 IL-18BP level is elevated in human cancers and correlated with T cell infiltration.
a, Expression of IL18BP transcripts in normal (blue) or cancer (red) tissues from the TCGA database. CHOL, cholangiocarcinoma; DLBC, diffuse large B cell lymphoma; GBM, glioblastoma multiforme; HSNC, head and neck squamous carcinoma; KIRC, kidney renal clear cell carcinoma; PAAD, pancreatic adenocarcinoma; SKCM, skin cutaneous melanoma; STAD, stomach adenocarcinoma (*P < 0.01). b–d, Correlation of IL18BP expression with T cell markers CD3E (b), CD8A (c) and PDCD1 (d) from the TCGA database for SKCM (n = 558), BRCA (breast adenocarcinoma, n = 1,085), HNSC (n = 44), STAD (n = 221), and OV (ovarian cancer, n = 426). e, Frequency of IL-18BP immunohistochemistry staining levels in human tumour tissue microarrays. Each sample was scored as negative (0) or positive (1+, 2+, or 3+). Representative images are shown for each staining level. f, Quantification of plasma IL-18BP protein level by ELISA for healthy donors (n = 22) and patients with NSCLC (n = 52) at baseline before treatment and at the time of the following CT scan after receiving treatment with anti-PD-(L)1 (n = 52). g, Representative mean tumour growth in wild-type (left) and Il18bp−/− (right) mice engrafted s.c. with MC38 tumours and treated with PBS or IL-18. Data are representative of three independent experiments with n = 5 mice per group. P values were calculated using one-way ANOVA (a, f) or two-way ANOVA (g), and data are presented as mean ± s.e.m.
Extended Data Fig. 3 Generation of decoy-resistant IL-18.
Related to Fig. 1. a, Structural alignment of hIL-18 (green)–hIL-18Rα/Rβ (cyan) complex (PDB ID 3WO4) with hIL-18–vIL-18BP (blue) complex (PDB ID 3F62). b, c, Representative surface plasmon resonance (SPR) sensorgrams of mouse WT IL-18 binding to IL-18Rα (b) or IL-18BP (c). IL-18Rα measurements were conducted using a conventional multiple cycle program, whereas IL-18BP measurements were conducted using a single-cycle program. d, Dose–response curves of IL-18BP protein antagonizing IL-18Rα in complex with indicated IL-18 and mutants (E42A, K89A and E42A/K89A). Experiments were performed in duplicate (n = 2). e, Randomized positions of murine IL-18 to create DR_18, with the corresponding degenerate codon and the potential amino acid at each position. f, Summary of the experimental design for directed evolution and yeast selection process to generate DR-18. Yeast libraries were selected for IL-18Rα binding and counter-selected against IL-18BP using magnetic-activated cell sorting (MACS; rounds 1 and 2) and subsequently FACS (rounds 3–5). Blue text (right) indicates positive selection reagent, and red text (left) shows the counter-selection reagent. g, Structural representation of DR-18 mutation positions in IL-18Rα and IL-1BP binding overlap region. Side chains from a minimized set of mutations up to six consensus residues (1N, 50M, 52K, 55E, 56V and 59L) are displayed as stick models. b–d, Data are representative of two independent experiments and are presented as mean ± s.e.m.
Extended Data Fig. 4 Anti-tumour efficacy of DR-18 in syngeneic tumour models.
Related to Fig. 2. Wild-type mice were engrafted s.c. with 0.5 × 106 MC38 or YUMMER1.7 tumour cells. From day 7 after engraftment, when tumour size reached 50–100 mm3, mice were treated twice weekly with PBS, or 0.32 mg/kg IL-18 or DR-18. a, Representative spider plots depicting MC38 tumour growth in wild-type mice treated with PBS, IL-18, DR-18, anti-PD-1, IL-18 + anti-PD-1, or DR-18 + anti-PD-1. b, e, Representative mean tumour growth (a) and Kaplan–Meier survival curves (e) of wild-type mice engrafted s.c. with YUMMER1.7 cells and treated with PBS, IL-18, DR-18, anti-PD-1, IL-18 + anti-PD-1, or DR-18 + anti-PD-1. Arrows signify days of treatment. c, f, Mean tumour growth (c) and Kaplan–Meier survival curves (f) of wild-type Balb/c mice engrafted s.c. with 0.25 × 106 CT-26 colon carcinoma cells and treated with PBS, anti-PD-1, IL-18, or DR-18. d, g, Mean tumour growth (d) and and Kaplan–Meier survival curves (g) of wild-type mice engrafted s.c. with 0.25 × 106 B16-F10 melanoma cells and treated with PBS, IL-18, or DR-18 from day 7. h, Mean tumour growth of wild-type mice engrafted s.c. with MC38 cells and treated with PBS, IL-18, CS1 or CS2 (DR-18), with the indicated doses. a–d, h, Data are representative of three independent experiments with n = 5 mice per group. e–g, Data are summary of three (e) or two (f–h) independent experiments with n = 5 mice per group. Data are presented as mean ± s.e.m.
Extended Data Fig. 5 DR-18 acts on antigen-specific T cells in the tumour.
Related to Fig. 2. a, Kaplan–Meier survival curves of mice engrafted with YUMMER1.7 tumours and treated with PBS, DR-18, or DR-18 and anti-CD4, anti-CD8, anti-NK1.1, or anti-IFN-γ antibodies. Treatment parameters were the same as for MC38 tumours (Fig. 2a, see Methods). b, Kaplan–Meier survival curves of wild-type or Rag2−/− mice engrafted s.c. with MC38 tumours and treated with PBS or DR-18. c, Kaplan–Meier survival curves of mice re-challenged with 1.0 × 106 MC38 tumour cells after successful DR-18 treatment compared to naive mice engrafted with 1.0 × 106 MC38 cells for the first time. d–i, Thy1.1+ P14 CD8 T cells (2.0 × 106) were adoptively transferred into wild-type mice, which were then engrafted s.c. with 0.25 × 106 B16F10-gp33 tumour cells one day later. Mice were then treated twice weekly with PBS or DR-18 (0.32 mg/kg) once tumours reached 50–100 mm3 for a total of three treatments. d–f, Representative flow plots (d), quantification of frequency (e), and absolute number of antigen-specific tumour-infiltrating P14 CD8+ T cells (f), as defined by Thy1.1+ and Gp33-tetramer. g, Percentage of IFN-γ on intratumoral antigen-specific (Thy1.1+Tetramer+) P14 CD8+ cells after PBS (n = 4) or DR-18 (n = 4) treatment. h, i, Representative flow plots (h) and quantification (i) of CD44 and CD39 expression on endogenous (Thy1.1−Tetramer−) tumour-infiltrating CD8+ T cells. j, Quantification of CD44 and PD-1 expression on tumour-infiltrating CD8+ T cells from wild-type mice engrafted with MC38 cells and treated with PBS or DR-18 in combination with FTY720. k, Representative flow cytometry plots of IL-18Rα expression on CD4+ T cells (CD45+, NK1.1−, CD3+, TCRβ+, CD4+), CD8+ T cells (CD45+, NK1.1−, CD3+, TCRβ+, CD8+), NK cells (CD45+, CD3−, TCRβ−, NK1.1+), B cells (CD45+, CD3−, TCRβ−, NK1.1−, CD19+), monocytes (CD45+, CD64+, CD11b+, Ly6C+), macrophages (CD45+, CD64+, MHCII+, F4/80+), dendritic cells (CD45+, CD64−, CD11c+), and neutrophils (CD45+, CD64−, CD11b+, Ly6G+) from MC38 tumours (n = 5), matched spleens (n = 5), un-engrafted wild-type spleens (n = 3), or Il18r1−/− spleens (n = 3). l, Representative mean tumour growth of MC38 tumours implanted into Rag2−/− mice that were adoptively transferred with T cells isolated from wild-type (n = 5) or Il18r1−/− (n = 8) mice and treated with PBS or DR-18. Black arrows signify day of treatment. m, Representative mean tumour growth of MC38 tumours implanted into wild-type (n = 4) and XCR1DTR (n = 5) mice treated with PBS or DR-18 in combination with DT. Green arrows signify DT treatment regimen. d–j, Data are representative of two independent experiments and presented as mean ± s.e.m. P values were calculated using one-tailed (e, f) or two-tailed (g) unpaired Student’s t-tests or two-way ANOVA (i, j).
Extended Data Fig. 6 Single-cell transcriptomic landscape of the TME in DR-18-treated tumours.
Related to Fig. 3. Wild-type mice were engrafted s.c. with 0.5 × 106 YUMMER1.7 cells. From day 7 after engraftment, mice were treated twice weekly with PBS, IL-18 or DR-18 (s.c.). On day 15, tumours were removed and sorted, and 5,000 cells per condition were used for scRNA-seq analysis. a, t-SNE projection showing all cells present in YUMMER1.7 and YUMMER1.7-B2m−/− tumours, coloured by cluster. Cells from the YUMMER1.7-B2m−/− experiments are shown in light grey and are discussed later. b, t-SNE projection showing distribution of cells coloured by treatment condition (dark grey, PBS; blue, IL-18; magenta, DR-18). c, Percentage of cells in each cluster by treatment. d, t-SNE projection of all cells analysed showing expression of genes supporting cell-type assignments. e, t-SNE projections of all cells analysed showing expression of IL-18 pathway-related genes. f, t-SNE projections showing expression of selected genes in computationally isolated Cd3e-positive T cells. g, t-SNE projections of Cd68+ cells, showing expression of Cx3cr1, Nos2, Arg1, and Mertk. h, t-SNE projection of Cd68+ cells, coloured by graph-based cluster. i, t-SNE projection of myeloid cell distribution as a function of treatment condition. j, Frequency of macrophages within each cluster identified in h. k, Heatmap showing differentially expressed genes in Cd68+ cells by clusters identified in h. l, m, Frequency (l) and absolute numbers (m) of Nos2+ macrophages from MC38 engrafted tumours following PBS, IL-18 or DR-18 treatment. Data are representative of three independent experiments with n = 5 mice per group and presented as mean ± s.e.m. P values calculated using one-way ANOVA.
Extended Data Fig. 7 DR-18 treatment enhances lymphocyte effector function.
Related to Fig. 3. Wild-type mice were engrafted s.c. with 0.5 × 106 MC38 or YUMMER1.7 cells. From day 7 after engraftment mice were treated twice weekly i.p. with PBS, IL-18, or DR-18. Tumours were harvested and analysed on day 15 after engraftment. a, Absolute number of tumour-infiltrating CD4+ T cells, CD8+ T cells and NK cells from YUMMER1.7 tumours in each treatment group. b, Quantification of intracellular Ki-67+ staining by percentage of indicated intratumoral lymphocyte population. c, Absolute number of antigen-experienced (CD44+) CD8+ T cells from MC38 tumours in each treatment group. d, Representative flow plots of PD-1 and CD39 staining on intratumoral CD44+ CD8+ T cells from MC38 tumours in indicated treatment groups. e, Absolute number of PD-1−CD39−, PD-1+CD39− and PD-1+CD39+ CD44+CD8+ T cells from d in each treatment group. f, Quantification by Luminex of IFN-γ protein level from tumour lysate of indicated treatment groups. g, Quantification of intracellular IFN-γ staining by percentage of indicated intratumoral lymphocyte populations from YUMMER1.7 tumours. h, i, Quantification of intracellular IFN-γ staining by percentage in tumour (h) or spleen (i) for each indicated intratumoral lymphocyte population from mice engrafted with MC38 cells. j, k, Quantification of intracellular granzyme B staining by MFI (j) and CD107 staining in intratumoral CD8+ T cells and NK cells (k) from MC38 tumours in the indicated treatment groups. l, m, Absolute number (l) and frequency (m) of polyfunctional NK cells measured by co-staining of intracellular IFN-γ and granzyme B following ex vivo stimulation with PMA and ionomycin. Data are representative of three independent experiments with five mice per group. P values calculated using two-tailed unpaired Student’s t-tests (a), one-way ANOVA (b, f, g, j, k) or two-way ANOVA (c, e, h, i, l) and data presented as mean ± s.e.m.
Extended Data Fig. 8 DR-18 promotes intratumoral TCF1+ precursor CD8+ T cells.
Related to Fig. 3. a, Frequency of surface PD-1 and TIM3 staining on MC38 intratumoral CD44+CD8+ T cells in indicated treatment groups. b, Absolute cell number of DN (PD-1−TIM3−), SP (PD-1+TIM3−) or DP (PD-1+TIM3+) CD8+ T cells from YUMMER1.7 tumours in each treatment group. c, Frequency of intracellular TCF1 in intratumoral CD44+CD8+ T cells in MC38 tumours in each treatment group. d, Absolute number of TCF1+CD44+CD8+ T cells from MC38 tumours in each treatment group. e, Frequency of intracellular TCF1 staining in MC38 intratumoral CD44+CD8+ T cells subsets based on PD-1 and TIM3 surface expression. f, Representative histograms of intracellular TCF1 staining in intratumoral SP PD1+TIM3−CD44+CD8+ T cells from YUMMER1.7 tumours. g, Absolute number of TCF1+ SP PD1+TIM3− CD44+CD8+ T cells from YUMMER1.7 tumours in each treatment group. h, i, Frequency of intracellular IFN-γ in DP (PD-1+TIM3+) CD44+CD8+ T cells from MC38 (h) and YUMMER1.7 (i) tumours in each treatment group. j, Absolute number of TCF1+ cells in SP (PD-1+TIM3−) CD44+CD8+ T cells in draining lymph nodes from MC38 tumours in each treatment group. Data are representative of three independent experiments with five mice per group. P values were calculated using one-way (c, d, g–j) or two-way ANOVA (a, b, e) and data are presented as mean ± s.e.m.
Extended Data Fig. 9 DR-18 is effective towards MHC class I-deficient tumours.
Related to Fig. 4. a, Representative spider plots showing tumour growth in wild-type mice engrafted s.c. with MC38-B2m−/− tumours and treated with PBS, or anti-PD-1 + anti-CTLA-4, IL-18 or DR-18. b, c, Kaplan–Meier survival curves (b) and representative spider plots (c) of wild-type mice engrafted s.c. with YUMMER1.7-B2m−/− tumours and treated twice weekly with PBS, anti-PD-1 + anti-CTLA-4, IL-18, DR-18, or DR-18 + anti-CD8 and anti-NK1.1 antibodies. Depleting antibodies were given on the same schedule as for MC38-B2m−/− (see Methods). d, e, Kaplan–Meier survival curves (d) and spider plots showing tumour growth (e) of wild-type mice engrafted s.c. with 1 × 106 RMA-S tumour cells and treated with PBS, anti-PD-1, IL-18, DR-18 or DR-18 + anti-NK1.1. Spider plot data are representative of two independent experiments and survival curve data are pooled from two independent experiments. f, Absolute number of intratumoral NK cells (NK1.1+CD3−EOMES+DX5+CD200R−), NKT cells (NK1.1+CD3+) and ILC1 cells (NK1.1+CD3−EOMES−DX5−CD200R+) in wild-type mice engrafted s.c. with MC38-B2m−/− tumours in indicated treatment groups. g, t-SNE projection of different clusters in YUMMER1.7-B2m−/− tumours as a function of treatment. YUMMER1.7 tumours are shown in light grey (discussed in Fig. 3, see Methods). h, Representative flow cytometry plots of IFNγ and Ki67 staining on intratumoral NK cells from YUMMER1.7-B2m−/− tumours treated with PBS or DR-18. i, j, Frequency of Ki67+IFNγ+ NK cells from YUMMER1.7-B2m−/− (i) and MC38-B2m−/− (j) tumours treated with PBS or DR-18. k, Representative mean tumour growth in wild-type mice engrafted s.c. with MC38-B2m−/− tumours and treated with PBS, DR-18, or DR-18 + anti-FasL or anti-IFN-γ. l, Representative mean tumour growth of Prf1−/− mice engrafted s.c. with MC38-B2m−/− cells and treated with PBS (n = 3) and DR-18 (n = 5) as indicated. h–l, Data are representative of two independent experiments with five mice per group. P values calculated using two-tailed unpaired Student’s t-test and data presented as mean ± s.e.m.
Extended Data Fig. 10 Ex vivo characterization of a human DR-18 variant.
a, SPR sensorgrams of human DR-18 (hDR-18) binding to human or cynomolgus macaque (M. fascicularis) IL-18Rα and IL-18BP. b, Concentration–response curves of hIL-18 or hDR-18 on HEK Blue IL-18R reporter cells. c, Titration of hIL-18BP on a fixed concentration of hIL-18 (1 ng/ml) or hDR-18 (0.1 ng/ml) on HEK Blue IL-18R reporter cells. d, e, Concentration–response curves of IFN-γ stimulation by hIL-18 or hDR-18 on human PBMCs (d) or cynomolgus macaque PBMCs (e). IFN-γ concentration measured by ELISA. Data are representative of at least two independent experiments with n = 2 for b–e and presented as mean ± s.e.m.
Supplementary information
Source data
Rights and permissions
About this article
Cite this article
Zhou, T., Damsky, W., Weizman, OE. et al. IL-18BP is a secreted immune checkpoint and barrier to IL-18 immunotherapy. Nature 583, 609–614 (2020). https://doi.org/10.1038/s41586-020-2422-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-020-2422-6
- Springer Nature Limited
This article is cited by
-
Single-cell transcriptional atlas of tumor-associated macrophages in breast cancer
Breast Cancer Research (2024)
-
The 4th NextGen therapies of SJIA and MAS, part 4: it is time for IL-18 based trials in systemic juvenile idiopathic arthritis?
Pediatric Rheumatology (2024)
-
Phenome-wide Mendelian randomisation analysis of 378,142 cases reveals risk factors for eight common cancers
Nature Communications (2024)
-
Dictionary of immune responses to cytokines at single-cell resolution
Nature (2024)
-
Biological and clinical roles of IL-18 in inflammatory diseases
Nature Reviews Rheumatology (2024)