Abstract
The initiation of an intestinal tumour is a probabilistic process that depends on the competition between mutant and normal epithelial stem cells in crypts1. Intestinal stem cells are closely associated with a diverse but poorly characterized network of mesenchymal cell types2,3. However, whether the physiological mesenchymal microenvironment of mutant stem cells affects tumour initiation remains unknown. Here we provide in vivo evidence that the mesenchymal niche controls tumour initiation in trans. By characterizing the heterogeneity of the intestinal mesenchyme using single-cell RNA-sequencing analysis, we identified a population of rare pericryptal Ptgs2-expressing fibroblasts that constitutively process arachidonic acid into highly labile prostaglandin E2 (PGE2). Specific ablation of Ptgs2 in fibroblasts was sufficient to prevent tumour initiation in two different models of sporadic, autochthonous tumorigenesis. Mechanistically, single-cell RNA-sequencing analyses of a mesenchymal niche model showed that fibroblast-derived PGE2 drives the expansion οf a population of Sca-1+ reserve-like stem cells. These express a strong regenerative/tumorigenic program, driven by the Hippo pathway effector Yap. In vivo, Yap is indispensable for Sca-1+ cell expansion and early tumour initiation and displays a nuclear localization in both mouse and human adenomas. Using organoid experiments, we identified a molecular mechanism whereby PGE2 promotes Yap dephosphorylation, nuclear translocation and transcriptional activity by signalling through the receptor Ptger4. Epithelial-specific ablation of Ptger4 misdirected the regenerative reprogramming of stem cells and prevented Sca-1+ cell expansion and sporadic tumour initiation in mutant mice, thereby demonstrating the robust paracrine control of tumour-initiating stem cells by PGE2–Ptger4. Analyses of patient-derived organoids established that PGE2–PTGER4 also regulates stem-cell function in humans. Our study demonstrates that initiation of colorectal cancer is orchestrated by the mesenchymal niche and reveals a mechanism by which rare pericryptal Ptgs2-expressing fibroblasts exert paracrine control over tumour-initiating stem cells via the druggable PGE2–Ptger4–Yap signalling axis.
Similar content being viewed by others
Main
Mesenchymal cells are localized in tight association with stem cells in crypts, separated from them by a layer of extracellular matrix less than a 1 μm in thickness (Extended Data Fig. 1a, b). To investigate the heterogeneity of intestinal mesenchymal cells and identify specific pathways that could control stem-cell dynamics, we performed single-cell RNA-sequencing (RNA-seq) analysis in the mouse intestinal mesenchyme. By sequencing 3,179 non-epithelial, non-immune cells we identified all major mesenchymal cell types of the lamina propria, including vascular and lymphatic endothelial cells, pericytes, smooth muscle cells and glial cells (Fig. 1a). Our analyses also revealed the existence of four different fibroblast populations—which we designate F1 to F4—all characterized by expression of Pdgfra (Fig. 1b); F1 and F2 cells are Pdgfralow, whereas F3 and F4 cells are Pdgfrahigh (Fig. 1b). A similar division of fibroblasts into PDGFRA high and low populations was found in a single-cell dataset3 of the human colonic mesenchyme (Extended Data Fig. 1h). Confocal and two-photon imaging in PdgfraeGFP/+-knockin mice4 confirmed the presence of Pdgfrahigh and Pdgfralow fibroblasts and revealed a distinct localization of these two populations in the adult intestine (Extended Data Fig. 2a, b). A dichotomous and compartmentalized expression of Pdgfra was observed in the intestinal mesenchyme throughout postnatal development and on embryonic day 15, when Pdgfrahigh cells are associated with early villus formation, whereas Pdgfralow cells occupy the rest of the mesenchyme (Extended Data Fig. 2c). Among Pdgfralow cells, F1 fibroblasts comprise two subsets (F1a and F1b), both marked by expression of Cd34 and Has1 (Fig. 1b). Population F2 expresses Cd34 and Fgfr2 and comprises four subsets (F2a–F2d) occupying diverse niches in the intestine, as shown in Fgfr2mCherry-knockin mice5 (Fig. 1b, Extended Data Fig. 2d). Among Pdgfrahigh cells, F3 cells express the Cajal-cell marker Ano2 whereas F4 cells express Acta2, consistent with a myofibroblast phenotype (Fig. 1b). To understand the potential functions of these uncharacterized populations we performed pathway analyses. We observed a robust enrichment of arachidonic acid metabolism genes in F3 (Cajal) cells and in the small population of F2d fibroblasts (Extended Data Fig. 1g, Supplementary Table 1), a pathway strongly associated with colorectal cancer6. Arachidonic acid is processed by cyclooxygenases to prostanoids, highly bioactive lipid mediators with a very short half-life and an autocrine or paracrine function in tissues6. In humans, pharmacological inhibition of cyclooxygenase-2 (Cox-2, also known as Ptgs2) prevents both hereditary and sporadic forms of colorectal cancer through an unknown cellular mechanism, but adverse cardiovascular effects currently impede its clinical application7. By fractionating normal human colonic tissues we found that expression of the Cox-2-encoding gene PTGS2 is nearly undetectable in the epithelium but occurs predominantly in stromal cells; the same pattern as observed in the mouse intestine (Extended Data Fig. 1c, d). Our single-cell analyses showed that in the steady-state, mouse intestine Ptgs2 is predominantly expressed in F3 (Cajal) cells and in the PdgfralowFgfr2+Vcam1high F2d fibroblasts (cluster 13) (Fig. 1c, Extended Data Fig. 1f). In humans, PTGS2 is mainly expressed in PDGFRAlowFGFR2+VCAM1+ fibroblasts (cluster 8) and, to a lesser extent, in other fibroblast populations (Extended Data Fig. 1h). Immunostaining of Cox-2 protein in the mouse intestine verified the presence of a Cox-2-expressing fibroblast population in the muscular layer, a location consistent with that of Cajal cells, and a second rare Cox-2-expressing fibroblast population located around part of the crypts, in close proximity to the stem-cell zone, suggestive of the F2d fibroblast cluster (Fig. 1d). We named these cells rare pericryptal Ptgs2-expressing fibroblasts (RPPFs). In agreement with their pericryptal location, RPPFs are marked by expression of the laminin subunit A1 (encoded by Lama1), a basement membrane protein detected specifically at the mesenchymal–epithelial interface, and also by expression of R-spondin 1 (encoded by Rspo1), a stem-cell niche factor detected mainly at the crypt base (Fig. 1b, Extended Data Fig. 2e, f).
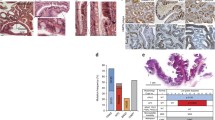
a–c, Single-cell RNA-seq of 3,179 mesenchymal cells from the normal mouse colon. a, t-distributed stochastic neighbour embedding (t-SNE) plot with clustering results. F, fibroblasts. MF, myofibroblasts. b, Violin plots showing the entire range of mesenchymal marker gene expression levels per single cell in each cluster. Endoth, endothelial; lymph, lymphatic; pericyt, pericytes; SMC, smooth muscle cells. c, Ptgs2 expression levels per single cell visualized by t-SNE plot. d, Immunostaining for Cox-2 (green) and vimentin (white) in the normal mouse ileum and colon. Cox-2-expressing fibroblasts are located under the crypt (C) epithelium (white arrowheads) and in the muscularis propria (MP). Results are representative of three independent experiments. Scale bars, 20 μm.
Given the localization of RPPFs within the stem-cell microenvironment, we aimed to understand their role as a potential mesenchymal niche of tumour-initiating stem cells. We used fibroblast-specific Col6-cre mice, which target a substantial fraction of Pdgfra+ intestinal fibroblasts, including fibroblasts surrounding the crypts and Cox-2-expressing fibroblasts, as shown by lineage tracing, flow cytometry in reporter mice and quantitative PCR with reverse transcription (RT–qPCR) analyses in Col6-Cre+ cells separated by fluorescence-activated cell sorting (FACS) (Extended Data Fig. 3a–c). Specific ablation of Cox-2 in Col6-cre+ fibroblasts in Col6-cre-Ptgs2f/f (Ptgs2ΔFibr) mice was efficient and led to a significant reduction of Ptgs2 expression levels in the whole tissue (Extended Data Fig. 3d, e), thereby confirming that fibroblasts are a predominant source of Cox-2 in the steady-state intestine.
To address the role of RPPFs in the mesenchymal niche in early tumour initiation, we used the ApcMin/+ mouse model in which autochthonous intestinal tumours are spontaneously formed by stem cells losing heterozygosity8. This model is highly relevant to human cancer, since somatic or germline mutations in the APC gene, a negative regulator of Wnt–β-catenin signalling, drive sporadic or hereditary forms, respectively, of intestinal neoplasia. Intestinal stem cell-specific Apc ablation is sufficient to drive tumorigenesis8. Notably, although adenoma formation in Apc-mutant mice is known to be Cox-2-dependent, the pathway by which this is mediated remains unknown9,10. Thus, we generated ApcMin/+Ptgs2ΔFibr mice and studied adenoma formation. We found that specific ablation of Ptgs2 in fibroblasts led to a strong reduction in the number of microadenomas formed in the small intestine at the early stage of 5 weeks (Fig. 2a) and significantly fewer macroscopic tumours formed at 5.5 months (Fig. 2b), along with a milder splenomegaly and a significantly prolonged survival (Extended Data Fig. 3f, g). We observed no difference in tumour size in ApcMin/+Ptgs2ΔFibr mice (Extended Data Fig. 3h), which showed that mesenchymal Cox-2 is necessary for tumour initiation but not for tumour growth. To understand how critical Cox-2 expression in the mesenchymal niche is for tumour initiation, compared with other cellular sources of prostanoids, we examined whether selective Cox-2 expression in fibroblasts is sufficient to drive tumour initiation in Apc-mutant mice. For this purpose, we genetically engineered mice in which a loxP-stop-loxP cassette was knocked into the Ptgs2 gene, thereby preventing its expression (Ptgs2OFF) unless excised by Cre-mediated recombination, which reactivates the gene (Ptgs2ON) (Extended Data Fig. 3i). By crossing these with Col6-cre mice, we generated mice in which Ptgs2 is expressed exclusively in fibroblasts (Ptgs2FibrON). We found that control ApcMin/+Ptgs2OFF mice developed only a few intestinal tumours, consistently with the phenotype of Ptgs2-knockout mice in this model9. By contrast, ApcMin/+Ptgs2FibrON mice—in which Ptgs2 is expressed exclusively in fibroblasts—displayed robust tumorigenesis and developed, on average, 30 adenomas per mouse in the small intestine (Fig. 2c). Thus, Cox-2 in fibroblasts is both necessary and sufficient for tumour initiation in Apc-mutant mice. To further establish the role of Cox-2-expressing fibroblasts in controlling tumorigenesis, we used a model of sporadic colonic tumorigenesis, which is driven by random mutations caused by repeated injections of azoxymethane, a potent mutagenic agent. We found that Ptgs2ΔFibr mice displayed a significantly lower incidence of dysplasia and adenoma formation in the colon at 28 weeks of age (Fig. 2d), along with a reduced number—but not reduced size—of adenomas and dysplastic foci per mouse (Extended Data Fig. 3j). These results from two different models demonstrated in vivo that fibroblasts utilize the Cox-2 pathway to provide a tumour initiation-conductive microenvironment for mutated stem cells in the intestine. Thus, we show that resident fibroblasts physiologically control tumorigenesis in trans.
a, Number of β-catenin+ microadenomas in two sections of the small intestine (SI) of 5-week-old ApcMin/+Ptgs2f/f (n = 6) and ApcMin/+Ptgs2ΔFibr (n = 6) mice. Two-tailed t-test. b, Number of macroscopic adenomas in 5.5-month-old ApcMin/+Ptgs2f/f (n = 16) and ApcMin/+Ptgs2ΔFibr (n = 23) mice. Two-tailed Mann–Whitney test. c, Number of macroscopic adenomas in 5.5-month-old ApcMin/+Ptgs2OFF (n = 7) mice in which Ptgs2 expression is blocked, and ApcMin/+Ptgs2FibrON (n = 11) littermates in which Ptgs2 is exclusively expressed in fibroblasts (Extended Data Fig. 3i). Two-tailed Mann–Whitney test (duodenum), t-test (jejunum and colon), Welch’s t-test (ileum and total small intestine). d, Incidence of dysplasia and microadenoma development in the colon of Ptgs2f/f (n = 30) and Ptgs2ΔFibr (n = 24) mice treated with 10 weekly injections of azoxymethane (AOM). Two-sided Fisher’s exact test. e, HPLC–MS/MS analysis of prostanoids in the ileum of littermate Ptgs2f/f (n = 9) and Ptgs2ΔFibr (n = 7) mice. The relative intensity to the respective internal standard is shown. Two-tailed t-test (PGE2 and PGF2α) and Mann–Whitney test (PGI2 and PGD2). Data are mean ± s.e.m. NS, non-significant; *P < 0.05; **P < 0.01; ***P < 0.001; ****P < 0.0001.
To identify which Cox-2-mediated prostanoids are secreted by fibroblasts in the crypt microenvironment in vivo and drive tumour initiation, we performed lipidomic analyses by liquid chromatography–tandem mass spectrometry (HPLC–MS/MS) in the intestine of Ptgs2ΔFibr mice. We identified a significant reduction in the relative abundance of PGE2 and prostaglandin I2 (PGI2) in the whole tissue (Fig. 2e), consistent with the expression of the respective synthases, Ptges and Ptgis, in fibroblasts (Extended Data Fig. 1g), and a trend for decreased prostaglandin D2 (PGD2) and prostaglandin F2α (PGF2α) abundance. Out of these prostaglandins, PGE2 promotes sporadic tumour formation11,12 but its cellular source, its cellular target and the receptor through which it acts have remained unknown. These are important considerations for therapeutic targeting in light of the adverse effects of Cox-2 inhibition7. Since the estimated half-life of PGE2 in vivo is less than 15 s (ref. 13), we hypothesized that PGE2 secreted by RPPFs may diffuse through the thin (less than 1 μm) basal lamina matrix and reach the neighbouring mutant stem cells at a concentration sufficient to signal and drive tumour initiation.
To study the effect of PGE2 on intestinal stem-cell function we cultured crypts in organoid growth media (OGM) supplemented daily with 16,16-dimethyl PGE2 (dmPGE2), a derivative of PGE2 with a prolonged half-life. Notably, PGE2 prevented the formation of budding organoids, leading instead to the development of spheroid-like structures which lack the typical crypt-villus architecture (Extended Data Fig. 4a). This morphology is associated with poor differentiation and increased stemness14. We functionally assessed stem-cell activity and found that PGE2-driven spheroids contain more stem cells with full organoid-forming capacity (Extended Data Fig. 4b). PGE2 signals through four receptors, EP1–EP4 (encoded by Ptger1–Ptger4, respectively), all of which are expressed in the mouse intestine (Extended Data Fig. 5a). We found that EP4 is the major PGE2 receptor expressed in the mouse intestinal epithelium, in stem and progenitor cells and in the normal human colon (Extended Data Fig. 5b–d). We then generated intestinal epithelial cell-specific Villin-cre-Ptger4f/f mice (Ptger4ΔIEC) and found that unlike control crypts, crypts from these mice were able to form budding organoids even when dmPGE2 was added to the OGM (Fig. 3a). On the basis of these results, we focused on the specific role of epithelial Ptger4 in tumour initiation.
a, Indicative 3D cultures of crypts isolated from Ptger4f/f and Ptger4ΔIEC mice grown with OGM or OGM supplemented with 0.1 μM dmPGE2. Scale bar, 100 μm. b, Normal crypts were cultured as 3D organoids in OGM (n = 2) or in co-cultures with primary mouse intestinal fibroblasts (n = 3). The absolute numbers of organoids or spheroids per 3D culture are shown. Results are representative of five independent experiments. Scale bar, 200 μm. c, Crypts isolated from Ptger4f/f and Ptger4ΔIEC mice were grown in co-cultures with wild-type (WT) primary mouse intestinal fibroblasts (n = 3 per genotype). The percentage of organoids and spheroids grown per 3D culture is shown. Results are representative of two independent experiments. Scale bar, 100 μm. In b, c, data are mean ± s.e.m.; two-way ANOVA, ****P < 0.0001. d–l, Single-cell RNA-seq of intestinal crypt–fibroblast co-cultures grown in OGM with 10 μM Ptger4 inhibitor (Ptger4-OFF) or DMSO (Ptger4-ON). Analyses are shown for 1,585 epithelial cells. d, t-SNE plot indicating epithelial cells from co-cultures with Ptger4-ON or Ptger4-OFF. e, t-SNE plot with clustering results. f, Proportion of each epithelial cluster among total epithelial cells in co-cultures with Ptger4-ON or Ptger4-OFF. g–l, Violin plots showing the entire range of metagene expression levels per single cell per cluster for the signatures/transcriptional programs of stem cells15 (g), tuft cells15 (h), RSCs16 (i), β-catenin (j), Yap (k) and early (non-tumour) Apcmin/+ tumorigenesis (l).
To model the mesenchymal niche of intestinal stem cells, we developed a 3D organotypic co-culture comprised of primary intestinal fibroblasts and fresh crypts growing in OGM. When co-cultured with fibroblasts, the majority of crypts developed into spheroids rather than organoids (Fig. 3b). Addition of Ptger4 inhibitors or co-culture with Ptger4ΔIEC crypts was sufficient to restore the growth of budding organoids (Fig. 3c, Extended Data Fig. 4c). Thus, organotypic cultures show that fibroblasts control stem-cell function via paracrine PGE2–Ptger4 signalling.
To understand the specific effects of fibroblast-derived PGE2 on stem-cell function and differentiation we performed single-cell RNA-seq in crypt–fibroblast co-cultures in which Ptger4 signalling was either active (Ptger4-ON) or inhibited (Ptger4-OFF) (Fig. 3d). We analysed 2,192 cells out of which 1,585 were epithelial (Extended Data Fig. 4d). By clustering and aligning these cells with signatures of known intestinal epithelial populations15 (Extended Data Fig. 4e), we observed a markedly different cellular composition between Ptger4-ON and Ptger4-OFF co-cultures (Fig. 3d, e). First, fibroblast-derived PGE2–Ptger4 signalling prevented the terminal differentiation of enterocytes but not that of goblet or Paneth cells (Fig. 3e, f). Second, it induced a substantial expansion of a non-cycling population displaying an intermediate transcriptional profile between stem and tuft cells (cluster 3) (Fig. 3f–h, Extended Data Fig. 4f). These cells express specific marker genes such as Ly6a (which encodes Sca-1), Clu, Msln and Il1rn (Extended Data Fig. 6b). Clu is a marker of revival stem cells16, a quiescent population that functions as reserve stem cells (RSCs)24 and is induced upon irradiation damage in the intestinal epithelium. We found that the overall transcriptional program of cluster 3 cells is highly similar to that of RSCs (Fig. 3i, Extended Data Fig. 4g). Furthermore, Ptger4 is expressed in RSCs and is strongly induced following irradiation damage in the regenerative intestinal epithelium (Extended Data Fig. 5e).
To understand the molecular link between PGE2–Ptger4 and the RSC phenotype, we first tested whether PGE2 activates the β-catenin pathway as reported in wound-associated epithelial cells17. However, we found a strong downregulation of the β-catenin transcriptional program in RSCs compared with cycling stem cells and no overall difference in this pathway between the Ptger4-ON and Ptger4-OFF conditions (Fig. 3j, Extended Data Fig. 4h, k). By contrast, we observed that Ptger4-ON spheroids overexpressed a set of genes reported to be targets of Yap18 as well as overall overexpression of a Yap transcriptional program18 (Extended Data Figs. 4k, 6a, b). This effect was validated in independent PGE2-driven spheroid cultures and confirmed genetically to be mediated by Ptger4 (Extended Data Fig. 6c–e). Yap is a transcriptional effector of Hippo signalling, which is involved in stemness, organ size control, tissue homeostasis, regeneration and cancer19 and is key for RSC function16. Yap is also indispensable for tumorigenesis driven by Apc-deficient stem cells18,20. Indeed, we found that a signature of early Apcmin/+ tumorigenesis correlated with the Yap program (Extended Data Fig. 6f) and both were predominantly expressed in RSCs (Fig. 3k, l, Extended Data Fig. 4i, j).
To directly examine whether Yap mediates fibroblast-driven expansion of RSCs, we isolated crypts from intestinal epithelial cell-specific Villin-cre-Yap1f/f mice (Yap1ΔIEC) and co-cultured them with fibroblasts in OGM. Although Yap1ΔIEC crypts require epiregulin supplementation to grow18, fibroblasts supported their growth in the absence of exogenous epiregulin (Extended Data Fig. 7a). Notably, unlike Yap1f/f crypts, Yap1ΔIEC crypts did not form spheroids in these co-cultures, but instead retained their crypt morphology (Extended Data Fig. 7a). Flow cytometry analyses for the RSC marker Sca-1 showed a robust expansion of Sca-1+ cells in co-cultures of fibroblasts with Yap1f/f crypts which was prevented in co-cultures with Yap1ΔIEC crypts (Extended Data Fig. 7b). Collectively, these analyses revealed that fibroblast-derived PGE2 drives the expansion of an RSC population with a regenerative/tumorigenic program via Ptger4 and Yap.
Next, we examined how PGE2 activates a Yap transcriptional program in crypts. In contrast to an earlier study using a cancer cell line21, stimulation of intestinal organoids with PGE2 did not induce Yap1 expression at the RNA or at the protein level (Extended Data Fig. 6g, h). In addition, day 3 PGE2-driven spheroids showed no difference in total Yap protein expression (Extended Data Fig. 6g). Since G-protein-coupled receptor (GPCR) signalling has been suggested to either activate or inhibit the Hippo–Yap signalling pathway22, and Ptger4 is a GPCR, we hypothesized that PGE2 may activate Yap in the intestinal epithelium by inhibiting its regulation by the Hippo pathway. Indeed, when we stimulated wild-type organoids with dmPGE2, we observed Yap dephosphorylation at Ser127 within 30–60 min (Extended Data Fig. 6i), suggestive of inhibition of Hippo activity and activation of Yap. This effect was mediated by Ptger4 (Fig. 4a). Furthermore, stimulation of wild-type organoids with dmPGE2 led to nuclear translocation of Yap within 30–60 min (Fig. 4b) and transcriptional activation of Yap target genes (Extended Data Fig. 6h), which was prevented by a Ptger4 inhibitor (Fig. 4c). PGE2-stimulation experiments with Yap1ΔIEC organoids or wild-type organoids treated with verteporfin (an inhibitor of the Yap–Tead interaction23) demonstrated that the activation of these genes is mediated by Yap (Fig. 4d, Extended Data Fig. 6j). These results establish that PGE2–Ptger4 signalling inhibits Hippo activity and, consequently, leads to Yap nuclear translocation and induction of a Yap–Tead-dependent gene-expression program in the intestinal crypt.
a, b, Wild-type organoids pretreated with or without 10 μM Ptger4 inhibitor were stimulated with 0.1 μM dmPGE2.Western blots for phosphorylated Yap Ser127 (pYap) and total Yap in cytoplasmic lysates (a), Yap and TBP in nuclear lysates (b). Results are representative of two experiments. c, d, Relative expression of Yap target genes18. c, Wild-type organoids treated with 10 μM Ptger4 inhibitor and 0.1 μM dmPGE2 for 13 h. n = 3 to 4 cultures per condition. One-way ANOVA. d, Yap1ΔIEC and Yap1f/f organoids treated with 0.1 μM dmPGE2 for 13 h. Three cultures per genotype per condition. Two-tailed t-test. e, f, Ptger4f/f (n = 3) and Ptger4ΔIEC (n = 3) mice received 14 Gy of abdominal irradiation. On day 3, the ileum was analysed by haematoxylin and eosin staining (e) and immunostaining for lysozyme (f). Results are representative of three independent experiments. Scale bars, 50 μm. g, Immunostaining for Yap in the small intestine of five-week-old wild-type and ApcMin/+ mice. Nuclear Yap is evaluated on the basis of colocalization with DAPI. Results are indicative of at least eight microadenomas. h, Immunostaining for Sca-1 in the small intestine of five-week-old wild-type and ApcMin/+ mice. Results are representative of two independent experiments. i, k, Quantification of areas of the small intestine with expansion of Sca-1+ epithelial cells in ApcMin/+Yap1f/f (n = 3) and ApcMin/+Yap1ΔIEC (n = 5) mice (i), and ApcMin/+Ptger4f/f (n = 7), ApcMin/+Ptger4ΔIEC (n = 5) mice (k). Two-tailed t-test. j, l, Number of BrdU+ microadenomas per small-intestinal section of ApcMin/+Yap1f/f (n = 4) and ApcMin/+Yap1ΔIEC (n = 5) mice (j), and ApcMin/+Ptger4f/f (n = 5) and ApcMin/+Ptger4ΔIEC (n = 7) mice (l). Two-tailed Mann–Whitney test (j); two-tailed t-test (l). m, Number of macroscopic adenomas in 5.5-month-old ApcMin/+Ptger4f/f (n = 9) and ApcMin/+Ptger4ΔIEC (n = 8) mice. Two-tailed t-test (duodenum and colon); Welch’s t-test (jejunum and total small intestine); Mann–Whitney test (ileum). Data are mean ± s.e.m.
Epithelial ablation of Ptger4 in Ptger4ΔIEC mice is efficient but does not affect stem-cell function and epithelial lineage differentiation in the steady-state intestine, as assessed by single-cell RNA-seq, 5-bromo-2′-deoxyuridine (BrdU)-incorporation experiments and immunostaining for population markers (Extended Data Fig. 8a–f, i). Lineage tracing of Ptger4-deficient Lgr5+ stem cells confirmed that Ptger4 is dispensable for the function of normal stem cells at the steady state (Extended Data Fig. 8h). Similar results were obtained in Ptgs2ΔFibr mice (Extended Data Fig. 3k). To assess the role of PGE2–Ptger4–Yap in stem-cell reprogramming in vivo, we employed an abdominal-irradiation-induced injury model in the context of which slow-cycling RSC populations are mobilized and mediate the epithelial regenerative response24. In the absence of Yap, although no phenotype is observed at the steady state, the epithelial response to irradiation is perturbed, causing increased Paneth cell differentiation18. We found that exposure of Ptger4ΔIEC mice to 14 Gy of abdominal irradiation led to a pronounced expansion of Paneth cells three days after irradiation (Fig. 4e, f), thereby phenocopying the effect of Yap deficiency. These results functionally validate the critical function of Ptger4 for RSC mobilization in vivo.
In early tumour initiation, we found that Yap displays an increased nuclear localization in microadenomas of five-week-old ApcMin/+ mice (Fig. 4g). Furthermore, we observed that Sca-1, a Yap target gene and RSC marker, is detected in the mesenchyme but not in the epithelium in the steady-state in wild-type mice. By contrast, however, in the intestine of five-week-old ApcMin/+ mice, we observed areas of the epithelium in which Sca-1+ epithelial cells were detected as local expansions, whereas Sca-1+ epithelial cells were more widespread in microadenomas (Fig. 4h). Similar expansion of Sca-1+ cells has been described in regenerative contexts such as the response to irradiation and to helminth infection25. Given these data, we examined the role of Yap in the expansion of Sca-1+ cells and in tumour initiation. In five-week-old ApcMin/+Yap1ΔIEC mice, we found a markedly decreased number of Sca-1+ areas in the epithelium compared with littermate ApcMin/+Yap1f/f controls, as well as an almost completely abrogated formation of microadenomas (Fig. 4i, j). These results show that in early tumour initiation, Yap translocates to the nucleus and drives the expansion of Sca-1+ cells and the formation of microadenomas. Nuclear localization of Yap and epithelial Sca-1 expression were also observed in developed tumours in five-month-old ApcMin/+ mice (Extended Data Fig. 9a, b). Moreover, mice treated with repeated azoxymethane injections displayed nuclear localization of Yap in tumours and overexpression of the Yap target gene Clu (Extended Data Fig. 9c, d).
To address whether fibroblast-derived PGE2 activates the Yap program and drives tumour initiation via epithelial Ptger4, we generated ApcMin/+Ptger4ΔIEC mice. Given the crucial role of Yap in tumour initiation, we first examined whether Ptger4 mediates the activation of Yap target genes in these mice. We found that at five weeks of age, ApcMin/+ mice displayed an increased expression of Yap target genes, but not of Yap1 itself, compared with normal controls; however, in ApcMin/+Ptger4ΔIEC littermates, this upregulation of the same Yap targets was abrogated (Extended Data Fig. 9e). Most notably, five-week-old ApcMin/+Ptger4ΔIEC mice displayed an attenuated expansion of Sca-1+ cells in the epithelium compared with their ApcMin/+Ptger4f/f littermates (Fig. 4k). Moreover, ApcMin/+Ptger4ΔIEC mice developed fewer microadenomas at five weeks of age (Fig. 4l). They also displayed a strong reduction in the number of macroscopic tumours formed at 5.5 months (Fig. 4m), a significantly alleviated splenomegaly (Extended Data Fig. 9f) and—consistent with a role of Ptger4 in tumour initiation rather than tumour growth—no difference in the size of the tumours formed (Extended Data Fig. 9g). These results provide definitive genetic evidence that epithelial Ptger4 is the receptor that mediates the tumorigenic effect of PGE2 and explain the role of RPPFs as paracrine drivers of tumour initiation.
On the basis of these results, we addressed the role of PGE2–PTGER4 in the human intestinal stem-cell niche. By isolating crypts from normal parts of the colon from three patients and culturing them in OGM we found that PGE2 drives the formation of spheroid-like structures. This effect was fully prevented by treatment with a PTGER4 inhibitor, thus confirming that PGE2–PTGER4 also controls stem-cell function in the human colonic crypt (Extended Data Fig. 10a). Furthermore, we performed immunostaining for YAP in tissues from 16 patients, including individuals with sporadic adenomas and adenocarcinomas, familial adenomatous polyposis, Lynch syndrome and cancer associated with inflammatory bowel disease (Supplementary Table 3). We observed that YAP displayed a nuclear localization in tumours but not in the neighbouring normal areas of the tissue in all these samples (Extended Data Fig. 10b), supporting its role in tumorigenesis. Of note, both PTGER4 and YAP1 genetic loci were recently identified to be genetically associated with colorectal cancer risk in genome-wide associations studies26, further underlining the relevance of this pathway to human disease.
The results of this study show that PGE2-secreting RPPFs provide a micro-niche favouring the activation of the pro-tumorigenic Yap program in neighbouring stem cells, thereby driving tumorigenesis in the presence of mutations (Extended Data Fig. 10c). This work establishes in vivo that the formation of intestinal tumours requires the paracrine interaction of mutated stem cells with their native mesenchymal microenvironment.
Methods
No statistical methods were used to predetermine sample size. The experiments were not randomized. The investigators were not blinded to allocation during experiments and outcome assessment unless otherwise stated in the Methods.
Mice
Ptgs2f/f mice27 crossed with Col6-cre mice28, Rosa26mT/mG mice29 and ApcMin/+ mice30 were bred in the animal facilities of the BSRC ‘Alexander Fleming’ under specific pathogen-free conditions. Ptger4f/f mice31 and Yap1f/f mice32 crossed with Villin-cre mice33and ApcMin/+ mice30, wild-type mice used for organoid experiments, Lgr5-eGFP-IRES-creERT2 mice34, PdgfraeGFP/+ mice4, as well as Col6-cre mice28 crossed with Rosa26tdTomato/+ mice (Ai14)35 and Ptgs2 Lox-Stop-Lox-knockin mice (Ptgs2LSL; the generation of this mouse strain is described below) were bred in the facilities of the Yale Animal Resources Center. All these mice were maintained on a C57BL/6J genetic background. Fgfr2-T2A-H2B-mCherry mice5 were maintained in the facilities of the Icahn School of Medicine at Mount Sinai on the 129S4 genetic background. Mice were housed in standard cages, on a 12:12 h day:night cycle and were fed a standard rodent chow. Mice were used for experiments at 8–12 weeks of age unless otherwise indicated. For all experiments, littermate, co-housed and sex-matched mice were used. Both male and female mice were used for experiments. No mice were excluded from the analyses performed. End points used for mice developing tumours were changes in activity or mobility, abnormal posture, decreased food and/or water intake and decreased body temperature. Experiments in BSRC ‘Alexander Fleming’ were approved by the Institutional Committee of Protocol Evaluation in conjunction with the Veterinary Service Management of the Hellenic Republic Prefecture of Attika according to all current European and national legislation. All animal experimentation at Yale was performed in compliance with Yale Institutional Animal Care and Use Committee protocols.
Generation of Ptgs2 Lox-stop-Lox-knockin mice
Ptgs2 flox-stop-knockin mice were generated by the University of California, Davis Mouse Biology Program services. JMB8 (C57BL/6N) embryonic stem (ES) cells were targeted with a vector containing a diphtheria toxin A (PGK-DTA) cassette, a 4 kb 5′ arm of homology, two loxP sites within intron 3 of the Ptgs2 gene flanking a STOP cassette sequence (derived from Addgene plasmid 11584), and a frt-flanked PKG-neomycin cassette and a 5.1 kb 3′ arm of homology. The PKG-neomycin element enabled positive selection in ES cells, while the DTA element enabled negative selection in ES cells. Mice bearing the targeted lox-stop-frt-PGK-neomycin-frt-lox Ptgs2 allele in the germline were crossed with the B6N(B6J)-Tg(CAG-Flpo)1Afst/Mmucd transgenic mouse (Mutant Mouse Resource and Research Center MMRC_036512-UCD) and the PGK-neomycin cassette was removed and mice bearing a lox-stop-frt-lox Ptgs2 allele (Ptgs2LSL) were obtained (Extended Data Fig. 3i).
Human study participants
Fresh human colon tissue was obtained from the Yale Pathology Archives on the basis of Yale Human Investigation Committee protocols no. 0304025173, which allows retrieval of tissue from surgical pathology that was consented or has been approved for use with waiver of consent. The data were analysed anonymously from preexisting patient databases and are thus exempt from consent by the human studies committee. Patient characteristics (sex, age, diagnosis) are described in the Supplementary Table 3. All tissue segments were obtained from the uninvolved surgical margins of colon resections. The specific part of the colon resected is indicated in the Supplementary Table 3. Ischaemic time of all samples ranged from 1 h to 3 h. All collected samples were kept on ice-cold RPMI medium before processing.
Formalin-fixed paraffin-embedded colorectal tumour tissue was obtained from the Yale Pathology Archives. The data were analysed anonymously from preexisting patient databases and hence exempt from consent by the human studies committee. Patient characteristics (sex, age, diagnosis) are described in the Supplementary Table 3.
Isolation of human intestinal epithelial cells and stromal cells
For the isolation of intestinal epithelial and stromal cells the tissue was cut into 0.5 cm pieces and incubated five times in HBSS containing 0.5 mM EDTA and 1 mM DTT for 15 min, at 4 °C on a rocker. Epithelial cells were released by vigorous shaking and passed through a 70-μm strainer, washed and used for RNA isolation. For stromal cell isolation the tissue pieces were incubated in DMEM containing 10% FBS, 300 U ml−1 Collagenase XI (Sigma, C7657), 0.1 mg ml−1 Dispase II (Sigma, D4693) and 50 U ml−1 DNase II Type V (Sigma, D8764) for 1 h, at 37 °C, 200 rpm. Cells released after vigorous shaking were passed through a 70-μm strainer, treated with ammonium-chloride-potassium red-blood-cell-lysing buffer, washed with 2% sorbitol and then used for RNA isolation.
Human colonic organoid culture
For the isolation of human colonic crypts the tissue was cut into 0.5-cm pieces and incubated six times in PBS containing 5 mM EDTA and 1 mM DTT for 10 min, at 4 °C on a rocker. Epithelial cells and whole crypts were released by vigorous shaking. The fractions enriched for crypts were further processed. Crypts were washed by centrifugation at 100g, 50g and 30g and then used for organoid development in domes made by Matrigel (Corning, 356231) and IntestiCult Organoid Growth Medium (Human) (Stem Cell Technologies, 06010) according to the manufacturer’s guidelines. When indicated, 16,16-dimethyl PGE2 (Cayman, 14750) dissolved in ethanol was added daily at a final concentration of 0.1 μM. Ethanol was used as a vehicle control for the untreated organoids. The ONO-AE3-208 Ptger4 (EP4) inhibitor (Cayman, 14522) dissolved in DMSO was added at a final concentration of 10 μM 1 h before stimulation and DMSO was used as a vehicle control.
Isolation of mouse intestinal epithelial cells and mesenchymal cells
The intestine was dissected, flushed, opened longitudinally and then cut into 1 cm pieces. The tissues were incubated in HBSS containing 1 mM EDTA, 1 mM DTT, 0.2% FBS, 4-5 times, 10 min each, at 37 °C, 200 rpm. Epithelial cells were released by vigorous shaking, passed through a 70 μm strainer, washed and immediately lysed for RNA isolation. After epithelial cell removal, the remaining stromal part of the intestine was lysed for RNA isolation. For Drop-seq analysis or for FACS-sorting of mesenchymal cells the tissues were processed as above and then incubated in DMEM 10% FBS containing Collagenase XI (300 units/ml, Sigma, C7657), Dispase II (0.1 mg/ml, Sigma, D4693) and DNase II Type V (50 units/ml, Sigma, D8764) for 1 h, at 37 °C, 200 rpm. Cells released after vigorous shaking were passed through a 70 μm strainer and washed with 2% sorbitol. Such cell preparations were directly processed by Drop-seq or by flow cytometry as described below.
Mouse intestinal organoid culture and fibroblast/crypt organotypic co-culture
Crypts were isolated from the last three fourths of the small intestine. The intestine was flushed, cut longitudinally and the villi were scraped off with a glass coverslip. The tissue was then cut into 0.5 cm pieces which were incubated in PBS containing 5 mM EDTA, 0.2% FBS for 30 min at 4 °C on a rocker. Crypts were released by vigorous shaking and were passed through a 70 μm strainer. Six fractions were obtained after vigorous shaking and the ones enriched for crypts were further processed. Crypts were washed by centrifugation at 200g, 100g and 50g and then used for organoid development in domes made by Matrigel (Corning, 356231) and IntestiCult Organoid Growth Medium (Stem Cell Technologies, 06005) according to manufacturer’s guidelines. When indicated, dmPGE2 (Cayman, 14750) dissolved in ethanol was added daily at a final concentration of 0.1 μM. Ethanol was used as a vehicle control for the untreated organoids. Crypts isolated from Yap1ΔIEC mice were cultured in IntestiCult Organoid Growth Medium (Stem Cell Technologies, 06005) supplemented with 0.5 μg ml−1 recombinant mouse epiregulin (RnD 1068-EP-050) or co-cultured with intestinal fibroblasts in OGM without epiregulin.
For the assessment of stem-cell activity, organoids or spheroids were dissociated into single cells by incubation at 37 °C in 0.25% trypsin-EDTA solution (Gibco, 25200056) diluted 1:1 with DMEM without serum. Numbers of live cells were counted after staining with trypan blue. In each experiment, the same number of live single cells per condition (n = 3,000–11,000) were cultured in domes made by Matrigel (Corning, 356231) and OGM.
Intestinal organoids were stimulated with dmPGE2 at a final concentration of 0.1 μM. Ethanol was used as a vehicle control. The ONO-AE3-208 Ptger4 (EP4) inhibitor (Cayman, 14522) dissolved in DMSO was added at a final concentration of 10 μM 1 h before stimulation. Verteporfin (Cayman, 17334) dissolved in DMSO was added at a final concentration of 1 μM 1 h before stimulation. DMSO was used as a vehicle control for ONO-AE3-208 and Verteporfin.
Fibroblasts were isolated from the small intestine of mice. The intestine was dissected, flushed, opened longitudinally and then cut into 1-cm pieces. The tissues were incubated in HBSS containing 1 mM EDTA, 1 mM DTT and 0.2% FBS 4–5 times for 10 min each, at 37 °C, 200 rpm. Epithelial cells were released by vigorous shaking. Then, the tissues were incubated in DMEM 10% FBS containing Collagenase XI (300 U ml−1, Sigma, C7657), Dispase II (0.1 mg ml−1, Sigma, D4693) and DNase II Type V (50 U ml−1, Sigma, D8764) for 1 h, at 37 °C, 200 rpm. Cells released after vigorous shaking were passed through a 70-μm strainer, washed and cultured in DMEM with 10% FBS. For co-culture experiments 2 × 104 fibroblasts were seeded in 48-well plates overnight. Freshly isolated crypts (n = 500) were suspended in 1:1 Matrigel (Corning, 356231) and OGM and added as an overlay on the fibroblasts. Crypts and fibroblasts were co-cultured with OGM. When indicated, the ONO-AE3-208 Ptger4 (EP4) inhibitor dissolved in DMSO was added to the co-cultures every second day at a final concentration of 10 μM. DMSO was used as a vehicle control for the untreated co-cultures.
Quantitative real-time PCR
RNA was isolated with the TRIzol reagent (Thermo Fisher, 15596026) followed by DNase I treatment (Roche, 04716728001) or with the QIAGEN RNA isolation RNeasy plus Mini Kit (QIAGEN, 74134) according to the manufacturer’s instructions. Reverse transcription was performed with the Maxima H Minus Reverse Transcriptase (Thermo Fisher, EP0751). RT–qPCR analyses were performed using iTaq Fast SYBR Green Supermix (Bio-Rad, 1725100) and a CFX96 Touch Real-Time PCR Detection System (Bio-Rad). Data were acquired and analysed with the CFX Manager software (Bio-Rad). Gene expression relative to a control sample was calculated with the RelQuant software (Bio-Rad Laboratories) by normalizing to B2m expression. Where indicated, relative expression (RE) to B2m was calculated as RE = 2−ΔCt. Primers used for human were B2M-F: ATGAGTATGCCTGCCGTGTG, B2M-R: CCAAATGCGGCATCTTCAAAC, PTGS2-F: TGTTGAAAAGTAGTTCTGGG, PTGS2-R: AAGCAGGCTAATACTGATAGG. Primers for mouse were B2m-F: TTCTGGTGCTTGTCTCACTGA, B2m-R: CAGTATGTTCGGCTTCCCATTC, Ptgs2: QT00165347 (QIAGEN) and Ptgs2-F: TCCAACCTCTCCTACTACACCAG, Ptgs2-R: GGGTCAGGGATGAACTCTCTC, Ptger1-F: AAGTTTTGGATTCACTTCCC, Ptger1-R: GAAGGTGTTGAGATTCTTGG, Ptger2-F: CTTGCCTTTCACAATCTTTG, Ptger2-R: ACCCAAGGGTCAATTATAGAG, Ptger3-F: CGCCGCTATTGATAATGATG, Ptger3-R: TTCTTAGCAGCAGATAAACC, Ptger4-F: GTGCGGAGATCCAGATGGTC, Ptger4-R: TCACCACGTTTGGCTGATATAAC, Ly6a-F: GAAAGAGCTCAGGGACTGGAGTGTT, Ly6a-R: TTAGGAGGGCAGATGGGTAAGCAA, Clu-F: GCTGCTGATCTGGGACAATG, Clu-R: ACCTACTCCCTTGAGTGGACA, Il1rn-F: GCTCATTGCTGGGTACTTACAA, Il1rn-R: CCAGACTTGGCACAAGACAGG, Cxcl16-F: CCTTGTCTCTTGCGTTCTTCC, Cxcl16-R: TCCAAAGTACCCTGCGGTATC, Msln-F: CTTAGTCTTGGGTGGATA, Msln-R: TCTTCTGTCTTACAGCCA, Yap1-F: GATGTCTCAGGAATTGAGAAC and Yap1-R: CTGTATCCATTTCATCCACAC.
Western blot
Total protein was extracted with RIPA lysis buffer. Nuclear and cytoplasmic fractions were extracted with NE-PER Nuclear and Cytoplasmic Extraction Reagents (Thermo Fisher, 78833). Protease inhibitors (Thermo Fisher, 87786) and phosphatase inhibitors (Thermo Fisher, 78420) were added. Antibodies against pYAP (Ser127) (D9W2I) (Cell Signaling, 13008), YAP (D8H1X) (Cell Signaling, 14074) and TBP (D5C9H) (Cell Signaling, 44059) were used at a 1:1,000 dilution in 5% BSA, overnight. Antibodies against β-actin (clone C4, Santa Cruz sc-47778) were used at a 1:2,000 dilution in 5% BSA for 2 h at room temperature.
Immunofluorescence and imaging
Two-photon microscopy was performed with a LaVision TriM Scope II (LaVision Biotec) microscope equipped with a Chameleon Vision II (Coherent) two-photon laser in the In Vivo Imaging Facility of Yale School of Medicine. Fresh whole-mount specimens of the small intestine and the colon of a PdgfraeGFP/+ mouse4 were prepared and analysed immediately after mice were euthanized.
Confocal imaging was performed with a Nikon-Ti microscope combined with UltraVox spinning disk (PerkinElmer) and data were analysed by using the Volocity software (PerkinElmer). Rosa26–tdTomato, Pdgfra–eGFP and Fgfr2–mCherry were detected by direct fluorescence. The tissues were dissected, fixed in 4% paraformaldehyde for 4 h at 4 °C, followed by incubation in 30% sucrose/PBS overnight at 4 °C. Tissue samples were frozen in OCT (Tissue-Tek) on dry ice and kept at −80 °C until sectioning. Sections 10 μm thick were prepared with a cryostat (Leica). After washing with PBS, sections were mounted with Fluoroshield histology medium containing DAPI (Sigma, F6057). For Lgr5–eGFP and vimentin immunostaining, the terminal ileum of a Lgr5-eGFP-IRES-creERT2 mouse34, was dissected, frozen and processed as above. The staining was performed with an Alexa Fuor 488-conjugated rabbit polyclonal GFP antibody at 1:200 (Thermo Fisher, A-21311) and an Alexa Fuor 647-conjugated rabbit monoclonal vimentin antibody at 1:200 (Cell Signaling, 9856), overnight at 4 °C. Sca-1 immunostaining was performed in frozen tissue sections processed as above and stained with a rat monoclonal Alexa Fuor 647 antibody (E13-161.7, Biolegend 122517, 1:400) overnight at 4 °C. The number of Sca-1+ areas was quantified in sections of small intestine prepared with the Swiss-roll technique and an EVOS FL Auto 2 Imaging System (Thermo). The evaluation was blinded to mouse genotype.
Formalin-fixed paraffin-embedded tissue sections were deparaffinised, washed and antigen retrieval was performed by microwave heating in citrate buffer. For Cox-2 immunostaining an anti-COX-2 rabbit polyclonal primary antibody was used (Cayman, 160126) at a 1:150 dilution, overnight at 4 °C with an anti-rabbit Alexa Fuor 488 secondary antibody at a 1:1,000 dilution, for 2 h at room temperature. Immunostaining for epithelial lineage markers was performed with conjugated antibodies against lysozyme (FITC-conjugated rabbit polyclonal, DAKO EC 3.2.1.17, 1:100) and Dclk1 (Alexa Fuor 647 rabbit monoclonal, EPR6085, Abcam ab202755, 1:400) and primary antibodies against chromogranin-A (rabbit polyclonal, Abcam ab15160, 1:300) and Olfm4 (rabbit monoclonal, D6Y5A, Cell Signaling 39141, 1:300) followed by an anti-rabbit Alexa Fuor 488 secondary antibody as above. Immunostaining for Yap was performed with a rabbit monoclonal primary antibody (D8H1X, Cell Signaling 14074, 1:50, overnight at 4 °C), a goat anti-rabbit biotinylated IgG secondary antibody (Vector, 1:750) and Streptavidin–Alexa Fuor 488 (1:800). Colocalization of Yap–Alexa Fuor 488 and DAPI was detected with the colocalization mode of Volocity software (PerkinElmer). Immunostaining for laminin A1 was performed with a rat monoclonal antibody (AL-4, R&D MAB4656, 1:50) overnight at 4 °C and an anti-rat Alexa Fuor 594 secondary antibody at a 1:1,500 dilution, for 1 h at room temperature. The numbers of positive cells for each epithelial marker per well-oriented crypt or crypt-villus unit were quantified in a blinded fashion.
BrdU
Administration of BrdU (Sigma) was performed intraperitoneally at a dose of 100 μg per g of body weight 2 h before mice were euthanized. BrdU immunohistochemistry was performed in sections of formalin-fixed paraffin-embedded tissues with the BrdU In-Situ Detection Kit (BD Pharmingen). The sections were counterstained with haematoxylin and analysed with a Leica DMI6000B microscope equipped with the Leica Application Suite LAS v.2.7 software. The number of BrdU+ cells per well-oriented crypt was quantified in a blinded fashion.
Alkaline phosphatase
Alkaline phosphatase activity was detected in deparaffinized sections with the Vector Red Alkaline Phosphatase Substrate Kit (Vector, SK-5100) in 200 mM Tris-HCl, pH 8.5 according to the manufacturer’s instructions. The sections were counterstained with haematoxylin and mounted with DPX or with Cytoseal Xyl (Thermo).
In situ hybridization
In situ hybridization was performed using the C Multiplex Fluorescent Detection Kit v.2 (ACD Bio) according to the manufacturer’s instructions. The colons of eight-week-old wild-type mice were excised, rolled up and immediately frozen in liquid nitrogen before embedding in Tissue-Tek OCT compound (Sakura Finetek). Sections with a thickness of 15 μm were prepared for RNAscope analysis using a mouse Rspo1 probe (ACD Bio 401991), a mouse Ppib positive control (ACD Bio 313911) and the bacterial DapB probe as a negative control (ACD Bio 310043). DAPI was used as a nuclear counterstain.
Evaluation of tumorigenesis in Apc Min/+ mice
Early tumorigenesis in ApcMin/+ mice was examined at the age of five weeks. The entire small intestine was collected and fixed in 10% neutral-buffered formalin solution as a Swiss roll. H&E-stained paraffin sections were examined with a Nikon Eclipse E800 microscope or a Leica DMI6000 B microscope. The number of microadenomas was quantified in sections stained for β-catenin (Fig. 2) or BrdU (Fig. 4). At the age of 5.5 months, tumorigenesis in ApcMin/+ mice was evaluated in the small intestine and the colon. The small intestine was partitioned into three parts of equal length (duodenum, jejunum and ileum). The tissues were opened longitudinally and the number of macroscopic tumours was quantified. The opened small intestine was rolled, fixed in formalin and H&E-stained paraffin sections were obtained. Pictures of all adenomas detected per section were obtained and their maximal diameter was measured by using the ImageJ software or the Leica Application Suite LAS v.2.7. All analyses were blinded to mouse genotype.
Azoxymethane-induced colon tumorigenesis
Mice were injected intraperitoneally with 10 mg kg−1 of azoxymethane (Sigma, A5486) once per week for 10 weeks starting at the age of 6 weeks. Mice were euthanized at the age of 28 weeks. Dysplasia development and adenoma formation were evaluated in H&E-stained paraffin sections of the colon. RNA was extracted from formalin-fixed, paraffin-embedded tissues with the RecoverAll Total Nucleic Acid Isolation Kit (Thermo Fisher, AM1975).
Prostanoid analysis by HPLC–MS/MS
Prostanoids were extracted with acetone followed by liquid/liquid extraction as previously described36, with some modifications. In brief, 10–50 mg of the ileum was homogenized in 500 μl PBS spiked with 100 μM butylated hydroxytoluene on ice. PGE2-d4 (Cayman, no. 314010) and PGD2-d4 (Cayman, no. 312010) were used as internal controls in each sample from the beginning of the extraction procedure at a final concentration of 10 ng ml−1. The samples were deproteinized with acetone. After mixing for 4 min and centrifugation at 2,000g for 10 min at 4 °C, the samples were transferred to clean 15-ml glass tubes, mixed with 800 μl hexane for 30 s and centrifuged for 10 min at 2,000g at 4 °C. The lower phase was acidified to pH 3.5 with formic acid and then mixed with chloroform. After mixing for 30 s and centrifugation for 10 min at 2,000g at 4 °C, the lower chloroform phase was evaporated to dryness under a stream of nitrogen and redissolved in 50 μl of methanol. HPLC–MS/MS analysis was performed using a modification of a method previously described37. From each sample a volume of 5 μl was injected into a Gemini 5 μm C18 110 Å, 100 × 2 mm HPLC column (Phenomenex, 00D-4435-B0) coupled with an Agilent 6490 QQQ Triple Quadrupole mass spectrometer with electrospray ionization in negative mode (Yale West Campus Analytical Core). The flow rate was 0.2 ml min−1 and the column was maintained at ambient temperature. The analysis was performed using an acetonitrile-based gradient system mixing two solvents: solvent A was acetonitrile/water/glacial acetic acid, 45/55/0.02 (v/v/v); solvent B was acetonitrile/water/glacial acetic acid, 90/10/0.02 (v/v/v). The analytes were separated using the following gradient: 0.0–8.0 min, 0% solvent B; 8.0–8.1 min, 0 to 50% solvent B; 8.0–12.0 min 50% solvent B; 12.0–12.1 min, 50 to 70% solvent B; 12.1–20.0 min 70% solvent B; 20.0–20.1 min, 70 to 0% solvent B; 20.1–30.0 min 0% solvent B. The capillary voltage was set at 3,500 V, source temperature at 120 °C, desolvation temperature at 360 °C and cone voltage at 35 V. The detection of prostanoids was based on the multiple reaction monitoring (MRM) method. The transition of precursor masses to specific fragments was monitored using a collision energy of 25–30 eV. PGE2 and PGD2 which have a similar MRM mass transition (m/z 351 → 271) and PGE2-d4/PGD2-d4 which also have similar MRM mass transition (m/z 355 → 275) were distinguished on the basis of their different elution time from the HPLC column. The MRM mass transition for PGF2α was m/z 353 → 193 and for PGI2 was m/z 351 → 215. The data were analysed with the Agilent MassHunter Workstation software, v.B.07.00. For each mass transition the area under the curve was normalized with that of the corresponding internal labelled control and a relative abundance was calculated. The relative abundances calculated were normalized based on the weight of the tissue sample. PGD2-d4 was used as a control for PGD2 and its metabolites. PGE2-d4 was used as a control for the rest of the prostanoids.
Single-cell RNA sequencing and data analysis
Single-cell RNA-seq was performed with the Drop-seq protocol as described previously38,39 with minor modifications. Drop-seq analysis of mesenchymal/lamina propria cells isolated from the middle and distal colon of wild type mice was performed in two biological replicates. For each biological replicate, the colons of n = 2 mice were pooled. The vast majority of intestinal epithelial cells were depleted by EDTA treatment as described above. N = 5 Drop-seq collections were processed in total, two from the first biological replicate and three from the second (Extended Data Fig. 1e). Drop-seq analysis of Ptger4-OFF and Ptger4-ON crypt/fibroblast co-cultures was performed on day 4 of the protocol in one pool of six Ptger4-ON co-cultures and one pool of six Ptger4-OFF co-cultures with one Drop-seq collection per pool. For Drop-seq analysis of crypts, epithelial cells isolated from the small intestine of Ptger4f/f and Ptger4ΔIEC mice, tissues from n = 2 mice per genotype were independently processed as biological replicates. A total of three Drop-seq collections were processed for each genotype, two from the first biological replicate and one from the second (Extended Data Fig. 8c).
The cells were diluted to a concentration of 100 cells per μl and 1-ml aliquots were used as input for each collection of the Drop-seq protocol38,39. The beads were purchased from ChemGenes (no. Macosko201110) and the polydimethylsiloxane co-flow microfluidic droplet generation device was generated by Nanoshift. Samples were processed for cDNA amplification within ∼15 min of collection. Populations of 5,000 beads (∼150 cells) were separately amplified for 15 cycles of PCR and pairs of PCR products were co-purified by the addition of 0.6× AMPure XP beads (Agencourt). Libraries were prepared and tagmented by Nextera XT using 1,000 pg of cDNA input, the custom primer P5_TSO_Hybrid38 and Nextera XT primers N701-N705 (Illumina). Libraries from intestinal mesenchymal cells and crypt epithelial cells were sequenced on the Illumina HiSeq platform (paired end, 2 × 150 bp) and libraries from crypt–fibroblast co-cultures on the Illumina NextSeq 500 platform (paired end, read 1 20 bp; read 2 60 bp), using a Read1CustomSeqB38 primer for read 1.
Single-cell RNA-seq data were processed as described38 to generate a digital expression matrix with transcript count data. This matrix was filtered retaining cells with more than 1,000 transcripts and less than 10% mitochondria transcripts. We then log transformed each entry of the matrix by computing log(TPM/100 + 1), where TPM is transcripts per million (meaning that the sum of all gene levels is equal 1,000,000). After normalization, we used adaptively thresholded low rank approximation (ALRA)40 to impute the matrix and fill in the technical dropped-out values. Subsequently, to visualize the cell subpopulations in two dimensions, we applied principal component analysis followed by t-SNE41, a nonlinear dimension reduction method, to the log-transformed data. DBSCAN42 and graph-based clustering (Seurat, Satija lab) were then used to generate clusters that were overlaid on the t-SNE coordinates to investigate cell subpopulations. Marker genes for each cluster of cells were identified using the Wilcoxon test with Seurat. Pathway enrichment analysis was performed by GSVA43 and P values were calculated with the moderated t-test implemented in the Limma R package. For the adjusted P values the false discovery rate (Benjamini–Hochberg) correction method was used.
In the fibroblast–crypt co-culture, single-cell RNA-seq experiment, epithelial cells were distinguished from fibroblasts and selected on the basis of unbiased clustering and known marker genes as shown in the Extended Data Fig. 4d. Cell-cycle analysis was adapted from Seurat. First, a score was calculated for each cell on the basis of the expression of G2M and S phase markers. Then, a discrete classification of cell cycle was assigned to each cell by comparing its G2M and S scores. Cells expressing neither were classified into the G1 group because they are less likely to be cycling.
A metagene score was assigned on the basis of publicly available bulk and single-cell RNA-seq datasets. For each of these datasets we selected significantly differentially expressed genes and constructed a metagene defined as weighted average of the log-transformed expression of these differentially expressed genes with weights equal to the log fold ratio of these genes in the respective dataset. More specifically, if we assume we have a metagene M that contains m genes: {gene1, gene2, …, genem} and each gene i has log fold change (FCi) in the data we use for the signature of interest, and each gene i has an expression value of xgene i in a given cell in our dataset, then the score for M in this specific cell is calculated as:
Each cell from our single-cell dataset was characterized by a score associated with each of the metagenes. The extent of differential behaviour between distributions of the total cell populations of two different conditions (Ptger4-ON vs Ptger4-OFF cells) was assessed for each metagene using the Kolmogorov–Smirnov test. We built metagenes for stem cells, enterocytes, Paneth cells, goblet cells, enteroendocrine cells and tuft cells by using lists of population-specific genes based upon plate single-cell RNA-seq data from the mouse intestinal epithelium15. We also built the following metagenes: (1) a β-catenin program metagene based on bulk RNA-seq data from organoids bearing a murine stabilized mutant Ctnnb1stab transgene and normal organoids (GSE93947), (2) A Yap program metagene based on bulk RNA-seq data from Yap-overexpressing and normal crypts isolated from doxycycline-treated and untreated YapTg inducible transgenic mice respectively (GSE66567)18, (3) An early ApcMin/+ tumorigenesis program metagene based on microarray gene-expression data from the nonpolypotic sections of terminal ileum from ApcMin/+ and wild-type mice (GSE49970).
Single-cell RNA-seq data of the healthy human colonic mesenchyme3 were obtained from GSE114374. Single-cell RNA-seq data of regenerating intestinal crypts16 were obtained from GSE117783. These datasets were processed by ALRA and the subsequent steps as above.
Abdominal irradiation of mice
For abdominal X-ray irradiation an X-RAD 320 Biological Irradiator (Precision X-ray) was used (Research Irradiator Facilities, Department of Therapeutic Radiology,Yale School of Medicine). Mice were anaesthetized and irradiated individually. 15 mm-thick lead was used for head, limb and tail shielding. Mice were treated at a distance of 50 cm from the radiation source with 320 kV, 12.5 mA X-rays, using a filter consisting of 2.0 mm Al. The mouse abdomen was centred between a 55 mm × 65 mm target window outlined by an adjustable collimator. The dose rate was measured with an ionization chamber by members of the Radiation Safety Division at Yale University. The abdominal dose rate was 235 cGy min−1. Average dose to shielded areas was 3.54 cGy min−1.
Flow cytometry and sorting
Freshly isolated stromal cells from Col6-cre-Rosa26tdTomato/+ mice were stained with monoclonal antibodies against CD45 (Biolegend) and used for FACS sorting. FACS sorting was performed at the Yale Flow Cytometry Facility with a BD FACSAria II sorter equipped with FACSDiva 7 software. Freshly isolated stromal cells from Col6-cre-Rosa26mT/mGPtgs2f/f mice were sorted on the basis of their GFP and tdTomato fluorescent protein expression with a BD FACSAria III sorter (BD) equipped with FACSDiva software at the Flow Cytometry Facility of BSRC Fleming. Single-cell suspensions from organoid cultures and co-cultures were obtained as described above, stained with monoclonal antibodies against Cd24 (Clone M1/69, Biolegend) and Sca-1 (Clone D7, Biolegend) and analysed at the Yale Flow Cytometry Facility with a BD LSRII cytometer equipped with FACSDiva software. Data analysis was performed with the FlowJo software.
Statistical analysis
Statistical analyses were performed with GraphPad Prism 7.01. Normality was tested with the Shapiro–Wilk W test. For W < 0.05, differences in means were tested for statistical significance with two-tailed Mann–Whitney test or Kruskal–Wallis test. For W > 0.05, variances were compared by F test and if similar (F test, P > 0.05), unpaired two-tailed Student’s t-test or one-way ANOVA was applied, otherwise (F test, P < 0.05) unpaired two-tailed Welch’s t-test was applied. For paired comparisons, statistical significance was tested with paired t-test for W > 0.05 or with Wilcoxon matched-pairs signed-rank test for W < 0.05. P values <0.05 were considered as statistically significant. Survival curves were compared by the log-rank test using GraphPad Prism 7.01.
Reporting summary
Further information on research design is available in the Nature Research Reporting Summary linked to this paper.
Data availability
All data that support the findings of this study are available within the paper and its Supplementary Information files. All Drop-seq data that support the findings of this study have been deposited in the Gene Expression Omnibus (GEO) repository with the accession code GSE142431.
Code availability
The code used for single-cell RNA-seq data analysis is available in GitHub (https://github.com/KlugerLab/Scripts_Roulis_et_al_2020).
References
Vermeulen, L. & Snippert, H. J. Stem cell dynamics in homeostasis and cancer of the intestine. Nat. Rev. Cancer 14, 468–480 (2014).
Powell, D. W., Pinchuk, I. V., Saada, J. I., Chen, X. & Mifflin, R. C. Mesenchymal cells of the intestinal lamina propria. Annu. Rev. Physiol. 73, 213–237 (2011).
Kinchen, J. et al. Structural remodeling of the human colonic mesenchyme in inflammatory bowel disease. Cell 175, 372–386 (2018).
Hamilton, T. G., Klinghoffer, R. A., Corrin, P. D. & Soriano, P. Evolutionary divergence of platelet-derived growth factor alpha receptor signaling mechanisms. Mol. Cell. Biol. 23, 4013–4025 (2003).
Molotkov, A., Mazot, P., Brewer, J. R., Cinalli, R. M. & Soriano, P. Distinct requirements for FGFR1 and FGFR2 in primitive endoderm development and exit from pluripotency. Dev. Cell. 41, 511–526 (2017).
Smyth, E. M., Grosser, T., Wang, M., Yu, Y. & FitzGerald, G. A. Prostanoids in health and disease. J. Lipid Res. 50 (Suppl), S423–S428 (2009).
Wang, D. & DuBois, R. N. The role of anti-inflammatory drugs in colorectal cancer. Annu. Rev. Med. 64, 131–144 (2013).
Barker, N. et al. Crypt stem cells as the cells-of-origin of intestinal cancer. Nature 457, 608–611 (2009).
Chulada, P. C. et al. Genetic disruption of Ptgs-1, as well as Ptgs-2, reduces intestinal tumorigenesis in Min mice. Cancer Res. 60, 4705–4708 (2000).
Cherukuri, D. P. et al. Targeted Cox2 gene deletion in intestinal epithelial cells decreases tumorigenesis in female, but not male, Apc Min/+ mice. Mol. Oncol. 8, 169–177 (2014).
Xia, D., Wang, D., Kim, S. H., Katoh, H. & DuBois, R. N. Prostaglandin E2 promotes intestinal tumor growth via DNA methylation. Nat. Med. 18, 224–226 (2012).
Wang, D., Fu, L., Sun, H., Guo, L. & DuBois, R. N. Prostaglandin E2 promotes colorectal cancer stem cell expansion and metastasis in mice. Gastroenterology 149, 1884–1895 (2015).
Bygdeman, M. Pharmacokinetics of prostaglandins. Best Pract. Res. Clin. Obstet. Gynaecol. 17, 707–716 (2003).
Mustata, R. C. et al. Identification of Lgr5-independent spheroid-generating progenitors of the mouse fetal intestinal epithelium. Cell Rep. 5, 421–432 (2013).
Haber, A. L. et al. A single-cell survey of the small intestinal epithelium. Nature 551, 333–339 (2017).
Ayyaz, A. et al. Single-cell transcriptomes of the regenerating intestine reveal a revival stem cell. Nature 569, 121–125 (2019).
Miyoshi, H. et al. Prostaglandin E2 promotes intestinal repair through an adaptive cellular response of the epithelium. EMBO J. 36, 5–24 (2017).
Gregorieff, A., Liu, Y., Inanlou, M. R., Khomchuk, Y. & Wrana, J. L. Yap-dependent reprogramming of Lgr5+ stem cells drives intestinal regeneration and cancer. Nature 526, 715–718 (2015).
Hong, A. W., Meng, Z. & Guan, K. L. The Hippo pathway in intestinal regeneration and disease. Nat. Rev. Gastroenterol. Hepatol. 13, 324–337 (2016).
Cai, J., Maitra, A., Anders, R. A., Taketo, M. M. & Pan, D. β-Catenin destruction complex-independent regulation of Hippo–YAP signaling by APC in intestinal tumorigenesis. Genes Dev. 29, 1493–1506 (2015).
Kim, H. B. et al. Prostaglandin E2 activates YAP and a positive-signaling loop to promote colon regeneration after colitis but also carcinogenesis in mice. Gastroenterology 152, 616–630 (2017).
Yu, F. X. et al. Regulation of the Hippo–YAP pathway by G-protein-coupled receptor signaling. Cell 150, 780–791 (2012).
Liu-Chittenden, Y. et al. Genetic and pharmacological disruption of the TEAD–YAP complex suppresses the oncogenic activity of YAP. Genes Dev. 26, 1300–1305 (2012).
Mills, J. C. & Sansom, O. J. Reserve stem cells: differentiated cells reprogram to fuel repair, metaplasia, and neoplasia in the adult gastrointestinal tract. Sci. Signal. 8, re8 (2015).
Nusse, Y. M. et al. Parasitic helminths induce fetal-like reversion in the intestinal stem cell niche. Nature 559, 109–113 (2018).
Huyghe, J. R. et al. Discovery of common and rare genetic risk variants for colorectal cancer. Nat. Genet. 51, 76–87 (2019).
Ishikawa, T. O. & Herschman, H. R. Conditional knockout mouse for tissue-specific disruption of the cyclooxygenase-2 (Cox-2) gene. Genesis 44, 143–149 (2006).
Armaka, M. et al. Mesenchymal cell targeting by TNF as a common pathogenic principle in chronic inflammatory joint and intestinal diseases. J. Exp. Med. 205, 331–337 (2008).
Muzumdar, M. D., Tasic, B., Miyamichi, K., Li, L. & Luo, L. A global double-fluorescent Cre reporter mouse. Genesis 45, 593–605 (2007).
Moser, A. R., Pitot, H. C. & Dove, W. F. A dominant mutation that predisposes to multiple intestinal neoplasia in the mouse. Science 247, 322–324 (1990).
Schneider, A. et al. Generation of a conditional allele of the mouse prostaglandin EP4 receptor. Genesis 40, 7–14 (2004).
Zhang, N. et al. The Merlin/NF2 tumor suppressor functions through the YAP oncoprotein to regulate tissue homeostasis in mammals. Dev. Cell 19, 27–38 (2010).
Madison, B. B. et al. Cis elements of the villin gene control expression in restricted domains of the vertical (crypt) and horizontal (duodenum, cecum) axes of the intestine. J. Biol. Chem. 277, 33275–33283 (2002).
Barker, N. et al. Identification of stem cells in small intestine and colon by marker gene Lgr5. Nature 449, 1003–1007 (2007).
Madisen, L. et al. A robust and high-throughput Cre reporting and characterization system for the whole mouse brain. Nat. Neurosci. 13, 133–140 (2010).
Roulis, M. et al. Intestinal myofibroblast-specific Tpl2–Cox-2–PGE2 pathway links innate sensing to epithelial homeostasis. Proc. Natl Acad. Sci. USA 111, E4658–E4667 (2014).
Masoodi, M. & Nicolaou, A. Lipidomic analysis of twenty-seven prostanoids and isoprostanes by liquid chromatography/electrospray tandem mass spectrometry. Rapid Commun. Mass Spectrom. 20, 3023–3029 (2006).
Macosko, E. Z. et al. Highly parallel genome-wide expression profiling of individual cells using nanoliter droplets. Cell 161, 1202–1214 (2015).
Shekhar, K. et al. Comprehensive classification of retinal bipolar neurons by single-cell transcriptomics. Cell 166, 1308–1323 (2016).
Linderman, G. C., Zhao, J. & Kluger, Y. Zero-preserving imputation of scRNA-seq data using low-rank approximation. Preprint at https://www.bioRxiv.org/content/10.1101/397588v1 (2018).
van der Maaten, L. & Hinton, G. Viualizing data using t-SNE. J. Mach. Learn. Res. 9, 2579–2605 (2008).
Ester, M., Kriegel, H.-P., Sander, J. & Xu, X. A density-based algorithm for discovering clusters in large spatial databases with noise. KDD 96, 226–231 (1996).
Hänzelmann, S., Castelo, R. & Guinney, J. GSVA: gene set variation analysis for microarray and RNA-seq data. BMC Bioinformatics 14, 7 (2013).
Acknowledgements
We thank C. Lieber, J. Alderman and E. Hughes-Picard for administrative assistance; the Yale Pathology Tissue Service–Tissue Procurement and Distribution Facility for providing human tissue samples; D. Gonzalez for assistance in two-photon imaging; M. Graham for assistance in electron microscopy; M. Samiotaki and T. Wu for assistance in mass spectrometry; and Flavell laboratory members R. Jackson and W. Bailis for discussions. M.R. is supported by a Crohn’s and Colitis Foundation Career Development Award (510777); M.S., L.-S.F., M.S.K. and M.B. were supported by an Austrian Marshall Plan Foundation Master’s Fellowship. This work was supported in part by ERC project MCs-inTEST (340217) (G.K.), the National Natural Science Foundation of China (31930035, 91942311) (B.S.), the Blavatnik Family Foundation and the Howard Hughes Medical Institute (R.A.F.).
Author information
Authors and Affiliations
Contributions
M.R. conceived and designed the study and wrote the manuscript. M.R. designed, performed and analysed experiments in Figs. 1–4 and Extended Data Figs. 1–10, assisted by M.S., L.-S.F., M.S.K., M.B., H.N.B. and J.R.B.; A.K. performed experiments in Fig. 2 and Extended Data Figs. 3, 9, assisted by V.K., N.C. and A.H. P.B. implemented Drop-seq. J.Z., R.Q. and Y.K. analysed Drop-seq data. E.K. and V.A. implemented HPLC–MS/MS analyses. X.Z. and B.S. performed in situ hybridization, H.R.H. and J.J. contributed Ptgs2f/f and Ptgs2LSL mice, R.M.B. contributed Ptger4f/f mice and D.J. contributed human FFPE tissues. G.K. and R.A.F. supervised all research, participated in the interpretation of results and edited the manuscript.
Corresponding authors
Ethics declarations
Competing interests
R.A.F. is a scientific advisor to GlaxoSmithKline and a shareholder and consultant for Zai Lab. All other authors declare no competing interests.
Additional information
Peer review information Nature thanks Garret A. FitzGerald, Dominic Grün, Kun-Liang Guan, Don W. Powell, Omer Yilmaz and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Mesenchymal Ptgs2 expression in the microenvironment of crypt stem cells.
a, Transmission electron microscopy photograph of the base of a mouse ileum crypt. P, Paneth cell; S, columnar basal stem cell; F, fibroblast. Scale bar, 5 μm. Indicative of independent observations in two experiments. b, Immunostaining for Lgr5–eGFP and Vimentin in the ileum of an Lgr5-eGFP-IRES-creERT2 mouse. Scale bar, 20 μm. Indicative of independent observations in one experiment. c, PTGS2 relative gene expression (RE) in intestinal epithelial cells (IECs) and stromal cells isolated from healthy human colonic tissues (n = 6 individuals). Statistical comparison performed with two-tailed Wilcoxon matched-pairs signed-rank test. d, Ptgs2 gene expression in IECs and stromal cells isolated from the ileum and the colon of wild-type mice (n = 4). Statistical significance was determined by two-tailed paired t-test. e, Biological replicates of the Drop-seq experiment shown in Fig. 1a visualized on the respective t-SNE plot depicting n = 3,179 mesenchymal cells. Mesenchymal cells were independently isolated from two groups of wild-type mice (biological replicates 1 and 2). From each of these isolations up to three independent Drop-seq samples were collected (A to C) for a total of five samples. f, All Ptgs2-expressing single cells (n = 1,136) detected in the experiment shown in Fig. 1a, c were analysed separately and re-clustered. Cluster annotations are visualized on a t-SNE plot. Violin plots display the entire distribution of gene expression levels per single cell in each cluster for key mesenchymal marker genes. F, fibroblasts. g, Schematic representation of the arachidonic acid metabolism pathway. For each mesenchymal cluster shown in Fig. 1a, violin plots display the entire distribution of gene expression levels per single cell for six genes involved in the metabolism of arachidonic acid to prostanoids. Data from n = 3,179 single mesenchymal cells are shown. h, Analysis of single-cell RNA-seq data (GSE11434) from the healthy human colonic mesenchyme3. Clustering results for n = 4,348 cells and cluster annotations are visualized on a t-SNE plot. The annotations of stromal populations are matched with the ones reported by Kinchen et al.3 on the basis of the respective markers. Expression levels of PTGS2 per single cell are visualized on a t-SNE plot. The entire range of gene expression levels per single cell for PTGS1, PTGS2 and key mesenchymal marker genes is displayed in violin plots. Data are mean ± s.e.m.; ns, non-significant; *P < 0.05; **P < 0.01.
Extended Data Fig. 2 Location of major fibroblast populations in the mouse intestine.
a, Detection of Pdgfra-expressing mesenchymal cells in the intestine of adult Pdgfra-H2B-eGPF-knockin mice4. Two distinct populations of Pdgfrahigh and Pdgfralow mesenchymal cells were detected in fixed tissue sections by direct eGFP fluorescence (green) and confocal microscopy. Nuclei are stained with DAPI (blue). Pdgfrahigh cells are located under the epithelium along the crypt–villus axis and in the muscularis propria. They form clusters at the tips of villi and the apical part of the colonic mucosa. Pdgfralow cells are located in the inner part of the villi, the pericryptal area and the submucosa. Filled arrows indicate pericryptal Pdgfralow cells. Open arrows indicate subepithelial Pdgfrahigh cells. M, mucosa; V, villus; SM, submucosa; MP, muscularis propria. Scale bars, 20 μm. Data are representative of six independent experiments. b, Detection of Pdgfrahigh and Pdgfralow fibroblasts in the fresh, intact intestine of adult Pdgfra-H2B-eGPF-knockin mice4 by two-photon microscopy. The cells were detected by direct eGFP fluorescence (green). Pdgfrahigh cells are predominant in the muscularis propria, whereas Pdgfralow cells are predominant in the submucosa. Both populations are present in the mucosa. Data are representative of independent observations from one experiment. Scale bars, 100 μm. c, Detection of Pdgfrahigh and Pdgfralow fibroblasts in the intestine of Pdgfra-H2B-eGPF-knockin embryos on embryonic day 15 (E15.0) and in early postnatal development. E15.0: clusters of Pdgfrahigh cells in early villi are indicated by white arrows. Pdgfralow mesenchymal cells occupy the rest of the mesenchyme (asterisks). P0: Pdgfrahigh cells are observed in the villi (V) and Pdgfralow cells are observed both in the villi and in the rest of the mesenchyme (asterisks). P15: Pdgfralow cells surround an early crypt (C) and Pdgfrahigh cells are located at the edges of the crypt (open white arrows). Pdgfralow cells occupy the inner mesenchyme (asterisks). Data are representative of independent observations from one experiment per developmental stage. Scale bars, 20 μm. d, The location of Fgfr2-expressing mesenchymal cells was determined in the intestine of an Fgfr2-T2A-H2B-mCherry-knockin mouse5, by detecting direct mCherry fluorescence (red) in the nucleus (blue, DAPI). The arrows indicate pericryptal Fgfr2+ fibroblasts. Data are representative of independent observations from one experiment. Scale bars, 20 μm. e, Immunostaining for laminin A1 (encoded by Lama1), the epithelial marker E-cadherin and the mesenchymal marker vimentin in the normal mouse intestines shows that laminin A1 is detected specifically at the mesenchymal–epithelial interface. Data are representative of two independent experiments. Scale bars, 5 μm. f, In-situ hybridization analysis showing the location of Rspo1-expressing cells in the normal mouse colon. The position of Rspo1-expressing cells along the crypt axis was quantified in 40 × 80 μm2 sub-epithelial areas at the base, middle and top sections of n = 9 crypts. Unpaired two-tailed Student’s t-test. Mean ± s.e.m. **P < 0.01.
Extended Data Fig. 3 Mice with fibroblast-specific ablation or fibroblast-restricted expression of Cox-2.
a, Immunofluorescence of ileum and colon sections from Col6-cre-Rosa26tdTomato/+ mice (scale bar, 20 μm) and of a small intestinal tumour section from an ApcMin/+-Col6-cre-Rosa26tdTomato/+ mouse (scale bar, 150 μm). Data are representative of two experiments. b, Efficiency of Col6-cre-mediated recombination of a lox-stop-lox tdTomato reporter in Pdgfrahigh and Pdgfralow Cd45− cells determined by flow cytometry in intestinal mesenchymal and lamina propria cells isolated from the small intestine and the colon of Col6-cre-Rosa26tdTomato/+PdgfraeGFP/+ mice in one experiment. c, Ptgs2 relative gene expression (RE) in whole tissue, isolated IECs, FACS-sorted Col6Cre+ fibroblasts (CD45−tdTomato+) and Col6Cre− mesenchymal cells (CD45−tdTomato−) from the small intestine of Col6-cre-Rosa26tdTomato/+ mice (n = 3, pooled). Representative of two experiments. d, Efficiency of Col6-cre-mediated Ptgs2 gene ablation in Col6-Cre+ mesenchymal cells determined by RT–qPCR analysis of Ptgs2 expression in FACS-sorted Col6-cre+ fibroblasts (eGFP+) from the small intestine of Col6-cre-Rosa26mT/mGPtgs2f/+ (n = 3) and Col6-cre-Rosa26mT/mGPtgs2f/f (n = 3) mice. Unpaired two-tailed Welch's t-test. e, Expression of the Ptgs2 gene in whole tissue ileum of littermate Ptgs2f/f and Ptgs2ΔFibr mice (n = 7 each). Two-tailed t-test. f, Spleen weight of 5.5-month-old ApcMin/+Ptgs2f/f (n = 8) and ApcMin/+Ptgs2ΔFibr (n = 6) mice. Average spleen weight of (n = 6) normal littermates (Ptgs2f/f) is displayed for comparison. Two-tailed t-test. g, Survival analysis of ApcMin/+Ptgs2f/f (n = 12) and ApcMin/+Ptgs2ΔFibr (n = 12) mice. A two-tailed P = 0.00009687 was calculated by log-rank test. h, Size of 274 adenomas from 5.5-month-old ApcMin/+Ptgs2f/f (n = 16) and ApcMin/+Ptgs2ΔFibr (n = 18) mice. The whiskers extend from minimum to maximum and the box extends from the 25th to 75th percentiles with the median indicated. Two-tailed Mann–Whitney test. i, Generation of knockin mice bearing a lox-stop-lox cassette insertion in intron-3 of the Ptgs2 gene which prevents its expression (Ptgs2OFF). Col6-cre-mediated excision of the lox-stop-lox cassette reactivates Ptgs2 expression specifically in fibroblasts (Ptgs2FibrON). The orange box depicts an frt site remaining from the flp-mediated removal of an frt-flanked PGK-neomycin selection cassette (see Methods). j, Ptgs2f/f (n = 30) and Ptgs2ΔFibr (n = 24) mice were subjected to 10 weekly intraperitoneal injections with 10 mg kg−1 azoxymethane as displayed. Quantification of the number of dysplastic foci and microadenomas per mouse and quantification of tumour size is shown. Statistical significance was tested by two-tailed Mann–Whitney test. k, Quantification of intestinal epithelial populations in the ileum of littermate Ptgs2f/f and Ptgs2ΔFibr mice (n = 3–5 per genotype). Immunostaining was performed for markers of Paneth cells (lysozyme), tuft cells (Dclk1), enteroendocrine cells (chromogranin A) and stem cells (Olfm4). Goblet cells were identified by periodic acid Schiff (PAS) staining and enterocytes were identified by detecting alkaline phosphatase enzymatic activity. Incorporation and immunohistochemical detection of BrdU was used to determine the numbers of cycling cells. Data for each mouse represent mean number of positive cells per crypt or crypt–villus unit as indicated. N = 400–822 crypts and/or villi were evaluated per staining. Statistical comparisons were performed with two-tailed unpaired t-test except for Olfm4+ cells for which unpaired t-test with Welch’s correction was applied. Scale bars, 50 μm. All data represent mean ± s.e.m. unless otherwise indicated. ns, non-significant; *P < 0.05, **P < 0.01, ***P < 0.001.
Extended Data Fig. 4 PGE2-driven spheroids contain more functional stem cells.
a, Crypts isolated from the small intestine of wild-type mice were grown into organoids by 3D culture with OGM or OGM that was supplemented daily with 0.1 μM 16,16-dimethyl PGE2 (dmPGE2). Indicative images and quantification of the absolute numbers of organoids and spheroids grown per 3D structure are shown. n = 6 3D cultures were evaluated per condition. Scale bar, 100 μm. b, Assessment of stem-cell activity in organoids or PGE2-driven spheroids grown as in a by dissociation into single cells and 3D culture in OGM. Growth of crypts and organoids from the same initial number of cells was quantified on day 14. The results are indicative of five independent experiments starting from independent crypt isolations. c, Normal crypts were grown into organoids with OGM in a 3D co-culture with primary mouse intestinal fibroblasts with or without 10 μM ONO-AE3-208 (Ptger4/EP4 inhibitor). Indicative images and quantification of the absolute numbers of organoids and spheroids grown per 3D structure are shown. n = 6 3D co-cultures were evaluated per condition. Scale bar, 200 μm. d, Separation of n = 2,192 fibroblasts and epithelial cells in single-cell RNA-seq data from fibroblast–crypt organotypic cultures on the basis of the expression of key marker genes. Expression of intestinal epithelial marker genes (Epcam, Atp1b1 and Krt8) and fibroblast marker genes (Sparc, Col1a1 and Col3a1) in single cells from Ptger4-ON and Ptger4–OFF fibroblast–crypt co-cultures is shown projected onto t-SNE plots. e–j, Single-cell data from Ptger4-ON and Ptger4–OFF fibroblast–crypt co-cultures as shown in Fig. 3d, visualized on the respective t-SNE plot depicting n = 1,585 epithelial cells. e, Expression of epithelial population-specific signatures (metagenes) per single epithelial cell. Population signatures were calculated on the basis of single-cell profiling of the mouse intestinal epithelium15. f, Cell cycle analysis of single epithelial cells projected onto the t-SNE plot. g–j, Expression levels of metagenes for the signatures or transcriptional programs of RSC16 (g), β-catenin (h), Yap (i) and early (non-tumour) ApcMin/+ tumorigenesis (j) per single epithelial cell projected onto t-SNE plots. k, Data from n = 1,585 single epithelial cells, visualized in violin plots for each co-culture condition (Ptger4-ON or Ptger4-OFF). The entire range of metagene expression levels per single epithelial cell for the signatures or transcriptional programs of RSCs, β-catenin, Yap and early ApcMin/+ tumorigenesis is displayed. In a, c, two-way ANOVA. Data are mean ± s.e.m. ****P < 0.0001.
Extended Data Fig. 5 Expression of PGE2 receptors in mouse and human tissues.
a, RT–qPCR analysis for Ptger1, Ptger2, Ptger3 and Ptger4 genes across 12 mouse tissues. Expression relative to B2m is displayed as 2−ΔCt. Data represent one experiment. MLN, mesenteric lymph nodes. b, RT–qPCR analysis for Ptger1, Ptger2, Ptger3 and Ptger4 genes in isolated IECs and matched stromal fractions from the small intestine (ileum) and the colon of wild-type mice (n = 4). Statistical comparisons were performed by two-tailed paired t-test. c, Expression levels of Ptger1, Ptger2, Ptger3 and Ptger4 genes determined by RNA-seq in FACS-sorted intestinal epithelial cell populations in 2 or 3 biological replicates and displayed as FPKM (fragments per kilobase of transcript per million mapped reads). Data retrieved from the GSE83394 GEO dataset. d, Expression levels of the human PTGER1, PTGER2, PTGER3 and PTGER4 genes in matched normal colon and tumour tissues from colorectal cancer patients (n = 41), determined by RNA-seq and displayed as FPKM. Data retrieved from The Cancer Genome Atlas for colon adenocarcinoma (TCGA–COAD dataset). Statistical comparisons were performed by two-tailed Wilcoxon matched-pairs signed-rank test. e, Analysis of single-cell RNA-seq data16 (GSE117783) from crypts isolated from the small intestine of normal mice (blue) and mice treated with 12 Gy irradiation (red). n = 6,644 single cells are visualized on t-SNE plots based on the experimental condition (normal, n = 2,882; irradiated, n = 3,762) and the clustering results. Violin plots represent the entire distribution of Ptger4 expression levels per single cell in each cluster and in each condition. The annotations of epithelial populations are matched with the ones reported by Ayyaz et al.16 on the basis of the respective markers. b–d, Mean ± s.e.m.; ns, non-significant; *P < 0.05; **P < 0.01.
Extended Data Fig. 6 PGE2–Ptger4 drive the induction of Yap target genes in intestinal organoids.
a, Volcano plot displaying the results of differential gene-expression analysis performed in single epithelial cells from Ptger4-ON and Ptger4–OFF fibroblast–crypt co-cultures (n = 1,585). Yap1 and Yap target genes18 are indicated. Moderated t-test with false-discovery rate (Benjamini–Hochberg) correction. b, Expression levels of the genes indicated in single epithelial cells from Ptger4-ON and Ptger4–OFF fibroblast–crypt co-cultures (n = 1,585), projected onto t-SNE plots. c, Experimental setup for data shown in d, e. Crypts were grown into organoids or spheroids by 3D culture in OGM or OGM that was supplemented daily with 0.1 μM dmPGE2 for 7 days. Gene expression levels were measured by RT–qPCR on day 7. d, Relative expression of Yap target genes (Ly6a, Clu, Il1rn, Msln and Cxcl16) in day 7 organoids and PGE2-driven spheroids developed from wild-type crypts. N = 3 3D cultures per condition. Two-tailed Welch’s t-test. e, Relative expression of Yap target genes in day 7 organoids and PGE2-driven spheroids developed from crypts isolated from Ptger4f/f and Ptger4ΔIEC mice. n = 3 3D cultures per genotype and condition. One-way ANOVA. f, Correlation between the expression levels of metagenes of a Yap transcriptional program and an early (non-tumour) ApcMin/+ tumorigenesis transcriptional program in single epithelial cells (n = 1,585) from the Ptger4-ON and Ptger4–OFF fibroblast–crypt co-cultures of Fig. 3. g, Small intestinal crypts were grown into organoids or spheroids with OGM or OGM that was supplemented daily with 0.1 μM dmPGE2. Western blot analysis for Yap1 and β-actin was performed in total lysates from untreated organoids, organoids treated with 0.1 μM dmPGE2 for 16 h and untreated spheroids. Data from one organoid and three independent spheroid cultures. h, Relative expression of the Yap1 gene and Yap target genes in wild-type organoid cultures treated with 0.1 μM dmPGE2 for 13 h, as determined by RT–qPCR. n = 3–5 cultures per condition. Statistical comparisons were performed with unpaired two-tailed t-test. For Ly6a, Welch’s correction was applied. i, Western blot analysis for Ser127 pYap and total Yap performed in total lysates from wild-type organoids stimulated with 0.1 μM dmPGE2 for the indicated time-points. Indicative of five independent experiments. j, Relative expression of Yap target genes in wild-type organoids treated with 1 μM verteporfin and 0.1 μM dmPGE2 for 13 h. n = 3–4 cultures per condition. Statistical comparisons were performed with unpaired two-tailed t-test, two-tailed Welch’s t-test or Mann–Whitney test on the basis of the criteria described in Methods. All data are mean ± s.e.m. *P < 0.05, **P < 0.01, ***P < 0.001, ****P < 0.0001.
Extended Data Fig. 7 Genetic ablation of Yap prevents spheroid formation and Sca-1+ stem-cell expansion in fibroblast–crypt organotypic co-cultures.
a, Crypts isolated from the small intestines of Yap1f/f and Yap1ΔIEC mice were grown into organoids by 3D culture with OGM supplemented with 0.5 mg ml−1 recombinant mouse epiregulin (Ereg) as previously described18, or in a co-culture with wild-type primary mouse intestinal fibroblasts with OGM without Ereg supplementation. Indicative images and quantification of the percentages of crypts, organoids and spheroids grown per 3D structure are shown. n = 2 cultures per condition. Data are representative of two independent experiments. Scale bars, 100 μm. Two-way ANOVA. Data represent mean ± s.e.m. ****P < 0.0001. b, Intestinal crypts isolated from the small intestines of Yap1f/f and Yap1ΔIEC mice were co-cultured with wild-type primary mouse intestinal fibroblasts. On day 4, these co-cultures and control Yap1f/f organoid cultures were processed into single-cell suspensions and analysed by flow cytometry for Sca-1 expression in Cd24+ epithelial cells. n = 2 cultures per condition. Scale bars, 100 μm. FSC, forward scatter. Unpaired two-tailed t-test. Mean ± s.e.m. ***P < 0.001.
Extended Data Fig. 8 Ptger4 ablation does not affect epithelial lineage differentiation and stem-cell function.
a, Ptger4 gene expression in crypts isolated from the ileum of littermate Ptger4f/f and Ptger4ΔIEC mice (n = 3 mice per genotype) and in organoids grown from these crypts (n = 3 cultures per genotype) determined by RT–qPCR analysis. Two-tailed unpaired t-test. b, Single-cell RNA-seq (Drop-seq) was performed in crypt epithelial cells isolated from littermate Ptger4f/f and Ptger4ΔIEC mice. Data for 2,439 single epithelial cells are shown in a t-SNE plot. c, Biological replicates visualized on a t-SNE plot. Crypt epithelial cells were independently isolated from two groups of mice per genotype (biological replicates 1 and 2). From the first biological replicate, two independent Drop-seq samples were collected (A and B) for a total number of three samples per genotype. d, Clustering and cluster assignments of 2,439 single epithelial cells displayed on a t-SNE plot. e, Proportion of each epithelial cluster among total crypt epithelial cells in Ptger4f/f and Ptger4ΔIEC mice. f, Violin plots showing the entire range of expression levels for a metagene of the β-catenin transcriptional program in n = 2,439 single epithelial cells from Ptger4f/f and Ptger4ΔIEC mice. g, Analysis of all Ptger4-expressing single cells detected (n = 478). Re-clustering results of Ptger4-expressing single cells with cluster annotations are visualized on a t-SNE plot. The expression levels of key marker genes for these clusters are visualized on t-SNE plots. h, Lineage tracing of Ptger4 heterozygous (Ptger4-HET) and Ptger4-knockout (Ptger4-KO) Lgr5+ stem cells. The small intestines of Lgr5-creERT2-Rosa26tdTomato/+Ptger4f/+ (Ptger4-HET) and Lgr5-creERT2-Rosa26tdTomato/+Ptger4f/f (Ptger4-KO) mice were examined for direct tdTomato fluorescence 5 days after a single injection of 2 mg tamoxifen per mouse. The results shown are representative of independent observations from one experiment. Scale bars, 70 μm. i, Quantification of intestinal epithelial populations in the ileum of littermate Ptger4f/f (n = 5) and Ptger4ΔIEC (n = 5) mice. Immunostaining was performed for markers of Paneth cells (lysozyme), tuft cells (Dclk1), enteroendocrine cells (chromogranin A) and stem cells (Olfm4). Scale bars, 20 μm. Goblet cells were identified by PAS staining and enterocytes were detected by alkaline phosphatase enzymatic activity. Scale bars, 50 μm. Incorporation and immunohistochemical detection of BrdU was used to determine the numbers of cycling cells. Scale bars, 50 μm. Data for each mouse represent mean number of positive cells per crypt or crypt–villus unit as indicated. n = 217–565 crypts and/or villi were evaluated per staining. Statistical comparisons were performed with two-tailed unpaired t-test except for PAS+ cells, for which unpaired Welch's t-test was applied. Mean ± s.e.m.; ns, non-significant; **P < 0.01.
Extended Data Fig. 9 Nuclear localization of Yap and activation of Yap target genes in ApcMin/+ and azoxymethane-induced tumorigenesis.
a, Immunostaining for Yap in the small intestine of five-month-old ApcMin/+ and wild-type littermate control mice. Nuclear localization of Yap is displayed on the basis of colocalization with DAPI. Normal (N) and tumour (T) areas of the ApcMin/+ intestine are indicated. Scale bars, 70 μm. Data are indicative of at least ten different tumour areas. b, Immunostaining for Sca-1 and the epithelial marker E-cadherin in normal and tumour areas of the small intestine of five-month-old ApcMin/+ mice. Indicative of two independent experiments. c, Immunostaining for Yap in the colon of wild-type mice subjected to 10 weekly intraperitoneal injections with 10 mg kg−1 azoxymethane as indicated and in untreated controls. Nuclear localization of Yap is displayed on the basis of colocalization with DAPI. Scale bars, 20 μm. Data indicative of three mice analysed. d, Relative expression of the Yap target gene Clu in normal and tumour areas of the colon of wild-type mice (n = 8) subjected to 10 weekly intraperitoneal injections with 10 mg kg−1 azoxymethane as shown in c. Two-tailed Mann–Whitney test. e, Relative expression of Yap1 and Yap target genes in the small intestine of 5-week-old Ptger4f/f (n = 3), ApcMin/+Ptger4f/f (n = 6) and ApcMin/+Ptger4ΔIEC (n = 8) mice. Statistical comparisons were performed with two-tailed t-test for Yap1 and Il1rn and with two-tailed Mann–Whitney test for Ly6a. f, Spleen weight of 5.5-month-old ApcMin/+Ptger4f/f (n = 8) and ApcMin/+Ptger4ΔIEC (n = 7) mice. Average spleen weight of (n = 2) normal littermates (Ptger4f/f) is displayed for comparison. Two-tailed t-test. g, Size of 72 adenomas from 5.5-month-old ApcMin/+Ptger4f/f (n = 6) and ApcMin/+Ptger4ΔIEC (n = 4) mice. The whiskers extend from minimum to maximum and the box extends from the 25th to 75th percentiles with the median indicated. Two-tailed Mann–Whitney test. Mean ± s.e.m.; ns, non-significant; *P < 0.05; **P < 0.01.
Extended Data Fig. 10 PGE2–PTGER4 controls stem-cell function in human colonic crypts and YAP displays a nuclear localization in human colorectal tumours.
a, Human colonic crypts were grown into organoids by 3D culture with OGM or OGM supplemented daily with 0.1 μM dmPGE2, with or without 10 μM ONO-AE3-208 (PTGER4–EP4 inhibitor). Images indicative of three independent experiments with crypts isolated from three patients are shown. Scale bar, 100 μm. b, Immunostaining for YAP in sections of human colorectal adenomas or adenocarcinomas and neighbouring normal tissue areas. Nuclear localization of Yap is displayed on the basis of colocalization with DAPI. Clearly defined normal (N) and tumour (T) areas are indicated wherever applicable. Images shown are representative of specimens obtained and analysed from n = 16 patients with the types of colorectal tumours indicated. Patient characteristics and the type of colorectal tumour per individual are described in the Supplementary Table 3. c, Schematic representation of the mechanism proposed in the present study. TISC, tumour-initiating stem cell.
Supplementary information
Supplementary Figure 1
This file contains the uncropped western blots.
Supplementary Table 1
Pathway analysis in fibroblast clusters. Top 10 KEGG pathways found to be enriched in each fibroblast population of the mouse colon mesenchyme by Gene Set Variation Analysis.
Supplementary Table 2
Quality control metrics of single cell RNA-seq experiments. Quality control metrics for the Drop-seq experiments performed 1) in mouse colonic mesenchymal cells, 2) in intestinal fibroblast/crypt co-cultures and 3) in primary crypt epithelial cells from Ptger4f/f and Ptger4ΔIEC mice.
Supplementary Table 3
Characteristics of human subjects. Characteristics of human subjects/patients from whom fresh normal tissues or Formalin-Fixed Paraffin Embedded tumor tissues were obtained.
Source data
Rights and permissions
About this article
Cite this article
Roulis, M., Kaklamanos, A., Schernthanner, M. et al. Paracrine orchestration of intestinal tumorigenesis by a mesenchymal niche. Nature 580, 524–529 (2020). https://doi.org/10.1038/s41586-020-2166-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-020-2166-3
- Springer Nature Limited
This article is cited by
-
Colorectal carcinoma peritoneal metastases-derived organoids: results and perspective of a model for tailoring hyperthermic intraperitoneal chemotherapy from bench-to-bedside
Journal of Experimental & Clinical Cancer Research (2024)
-
Beyond genetics: driving cancer with the tumour microenvironment behind the wheel
Nature Reviews Cancer (2024)
-
Tumor initiation and early tumorigenesis: molecular mechanisms and interventional targets
Signal Transduction and Targeted Therapy (2024)
-
Towards Full Thickness Small Intestinal Models: Incorporation of Stromal Cells
Tissue Engineering and Regenerative Medicine (2024)
-
The cyclooxygenase-expressing mesenchyme resists intestinal epithelial injury by paracrine signaling
Cell Regeneration (2023)