Abstract
A non-enveloped virus requires a membrane lesion to deliver its genome into a target cell1. For rotaviruses, membrane perforation is a principal function of the viral outer-layer protein, VP42,3. Here we describe the use of electron cryomicroscopy to determine how VP4 performs this function and show that when activated by cleavage to VP8* and VP5*, VP4 can rearrange on the virion surface from an ‘upright’ to a ‘reversed’ conformation. The reversed structure projects a previously buried ‘foot’ domain outwards into the membrane of the host cell to which the virion has attached. Electron cryotomograms of virus particles entering cells are consistent with this picture. Using a disulfide mutant of VP4, we have also stabilized a probable intermediate in the transition between the two conformations. Our results define molecular mechanisms for the first steps of the penetration of rotaviruses into the membranes of target cells and suggest similarities with mechanisms postulated for other viruses.
Similar content being viewed by others
Main
Rotaviruses and other non-enveloped, mammalian, double-strand RNA (dsRNA) viruses have evolved a mechanism to deliver a large (around 700 Å diameter) macromolecular complex into the cytosol of a target cell. These viruses package segmented genomes into icosahedral capsids with several protein layers2,3 (Fig. 1a). The role of the outer layer of an infectious rotavirus particle (a triple-layer particle, or TLP) is to insert the double-layer particle (DLP) that it surrounds into the cell to which the virus has attached. The DLP does not dissociate further: the polymerase and capping enzymes contained within it instead transcribe the 11 genome segments, cap the mRNA products and extrude the completed transcripts into the cytosol2 (Fig. 1b).
a, Overall structure of a rotavirus TLP. b, Schematic of the virus entry pathway derived from live-cell imaging experiments8,16,20. c, Domain organization of the VP8* and VP5* proteins in the upright (left) and reversed (right) conformations. The domains are labelled underneath the primary structure. The residue numbers indicate termini and domain boundaries. α, N-terminal α-helical segment of VP8*. The regions in grey are not detected (disordered) in the reversed-conformation structure. d, Atomic models of upright (left) and reversed (right) conformations of VP5* (and VP8*) on the virion surface, from our cryo-EM reconstructions of wild-type recoated RRV TLPs. VP5*, red, orange and salmon; VP6, green; VP7, yellow; VP8*, magenta. Some VP6 and VP7 subunits are omitted for clarity.
Previous studies have defined the functions of the two TLP outer-layer proteins, VP4 and VP7, in the delivery of the DLP4,5,6,7,8,9. Infection requires the cleavage of VP4—both in cell culture and in vivo—into an N-terminal fragment, VP8*, and a C-terminal fragment, VP5*. VP8* attaches the virion to its cellular receptor (in many cases, a glycolipid)10,11,12,13,14, whereas VP5* perforates the lipid bilayer of the vesicular membrane that surrounds the entering particle after its initial uptake by the target cell (Fig. 1b). The function of VP7, a Ca2+-stabilized trimer15, is to anchor VP4 onto the virion surface and to respond to loss of Ca2+ at an early stage of infection by dissociating and releasing VP8* and VP5* from the DLP16.
On a virion, the ‘spike’ protein VP4 is a trimer with an unusual, asymmetric conformation, both before and after its activation by cleavage of each subunit into VP8* and VP5* fragments6,17. For one of the three VP8* fragments, cleavage releases all but a short, N-terminal segment of VP8* (denoted VP8* α; Fig. 1c). The two intact VP8* fragments associate with the bean-shaped β-barrel domains of the VP5* fragments from which they have been separated by cleavage, forming a two-fold symmetric projection; the β-barrel domain of the third VP5* supports this projection, holding it away from the particle surface like a diagonal cantilever6. The VP8* lectin domains at the tip of the spike cover hydrophobic loops at one end of the two projecting VP5* β barrels with which they associate; the opposite ends of the projecting VP5* barrels similarly cover the hydrophobic loops of the third VP5* β barrel. The C-terminal regions of all three VP5* chains associate with three-fold symmetry into the ‘foot’ of the spike, clamped onto the particle surface6 by VP7.
Recombinant VP4 is monomeric; successive treatment with chymotrypsin and trypsin yields a trimeric product, designated VP5CT, from which VP8* has dissociated completely and from which most residues of the foot have been removed18. An α-helical coiled coil at the centre of VP5CT projects in the same direction as the hydrophobic loops on the β barrels—a chain reversal with respect to the spike on a TLP. We have previously proposed that the transition from the asymmetric, ‘upright’ spike conformation seen on virions to the symmetric, ‘reversed’ conformation seen in VP5CT leads to perforation of a target membrane when coupled with the interaction of the β-barrel hydrophobic loops with the lipid bilayer18. Mutations that reduce the hydrophobicity of the VP5* loops compromise infectivity and reduce the association of VP5* with liposomes in vitro5.
We describe here results from electron cryomicroscopy (cryo-EM), showing that transition to the reversed conformation can occur spontaneously on the surface of the TLP, without the dissociation of VP7 (Fig. 1d), and that association of the hydrophobic loops with a target membrane causes residues of the foot region to enter the lipid bilayer of the membrane. We infer that the formation of an initial membrane lesion by interaction of the foot regions from one or more VP5* trimers with the bilayer is a decisive, early event in the penetration of rotaviruses and the delivery of the DLPs into a host cell.
Reversed VP5* on rotavirus particles
We determined the three-dimensional structures of rhesus rotavirus (RRV) (Supplementary Data 1–3) native TLPs and wild-type recoated TLPs (rcTLPs), prepared as described in Methods (Extended Data Fig. 1). We initially imposed icosahedral symmetry and then classified subparticles corresponding to VP4 positions (Extended Data Fig. 2). Classification of rcTLPs (without further alignment) yielded three distinct classes (Extended Data Figs. 3, 4, Extended Data Table 1): one corresponding to the asymmetric upright spike (Fig. 1d, left), a second corresponding to a three-fold symmetric reversed structure (Fig. 1d, right) and a third corresponding to empty vertices. Classification of spikes from native TLPs, which were treated in the same way as rcTLPs, yielded classes that were identical to those from rcTLPs but with a different distribution of particles among the classes (Extended Data Fig. 5). The three-fold symmetric structure resembled the reversed conformation adopted by VP5CT in solution18 but not previously seen on rotavirus particles. When we omitted the incubation at 37 °C, the TLPs had only upright VP4 spikes and some empty positions at which VP4 was not present.
Inspection of the reversed-conformation map showed that the VP5CT coordinates18 matched closely with the reconstructed density. In VP5CT, three β-barrel domains surround a central, three-chain coiled coil formed by polypeptide-chain segments C-terminal to the β barrels (Fig. 1d, right). Hydrophobic loops at one end of the β barrel project in the same direction as the distal end of the coiled coil. VP5CT, which is produced by proteolytic cleavage of recombinant VP4, terminates at the end of the coiled coil; about 250 additional amino acid residues on intact VP4 form the foot that anchors VP4 in the upright spike conformation onto the DLP (Fig. 1d, left). The VP5* in our rcTLPs was intact (Extended Data Fig. 1b), but the region beneath the VP5CT-like density—normally occupied by the foot—was completely empty in the reversed reconstructions (Fig. 1d, right). Loss of the foot from this cavity indicates that, as expected from the direction of the coiled coil, the final approximately 250 amino acid residues of the VP5* polypeptide chain project outwards. We detected little or no density C-terminal to the coiled coil in the reversed class map, and therefore concluded that the foot residues must be substantially disordered. We have previously described the VP5CT conformation as a ‘folded-back’ structure, and suggested that a conformational transition of the spike might involve folding back of the β-barrel domain towards the still-anchored foot8,18. In the reversed structure that we have now visualized, formation of the coiled coil has instead projected the foot-domain residues outwards, while retaining the orientation of the β barrels (Fig. 1d, Extended Data Fig. 6a).
After the transition to the reversed conformation, VP5* does not dissociate despite the loss of interactions between the foot and the surrounding VP6 and VP7 trimers. The inward-facing end of each of the three VP5* β-barrels contacts a set of loops in the adjacent VP7 through a network of largely polar side chain interactions (Fig. 2a, right, Extended Data Fig. 7). VP8* remains associated with the TLP, even after the transition19 (Extended Data Fig. 8a). On a TLP, the foot domain of VP5* includes—along its three-fold axis—the initial approximately 16 residues of the VP4 polypeptide chain, two of which then connect outwards to the VP8* lectin domains at the apex of the spike, while the third terminates at residue 27 (Fig. 2b, left). As the foot unfolds and projects outwards along the three-fold axis of the spike, the three VP8* N-terminal segments must therefore remain trapped in the foot cavity, with the connections to the two lectin domains emerging through a channel at the VP5*–VP7 interface (Fig. 2b, right). These emerging connections could, in principle, also participate in the VP5*–VP7 contacts.
a, Top views of the upright (left), intermediate (middle) and reversed (right) conformations. VP5*, red, orange and salmon; VP7 chains interacting with VP5*, yellow. Magnified views of the interfaces are shown in Extended Data Fig. 7. Gaps between VP5* and VP7 shell molecules that allow for the exit of VP5* foot domains are marked by blue dashed boxes. Scale bar, 50 Å. b, Side views of the upright (left), intermediate (middle) and reversed (right) conformations. VP8*, magenta. In the reversed structure (right), VP8* is shown schematically: no density corresponding to VP8* was observed, but biochemical data reveals its presence. Scale bar, 50 Å.
An early cryo-EM study of the structure of rotaviruses found that brief incubation of RRV at pH 11, followed by a return to pH 8, caused an apparent collapse of the spikes into three symmetrically disposed protuberances19. We repeated the protocol from this previous study with both native TLPs and wild-type VP4 rcTLPs, and found that exposure to pH 11 induced nearly complete transition to the reversed conformation we describe above (Extended Data Fig. 8b). Therefore, we suggest that the protuberances described in the previous study were the three β-barrel domains as seen at low resolution (Supplementary Discussion).
Trapping a potential intermediate
The transition of VP5* from an upright (Fig. 1d, left) to a reversed (Fig. 1d, right) conformation might plausibly proceed in two steps: the rearrangement of the β-barrel domains into a three-fold cluster, followed by an eversion along the three-fold axis of the remainder of VP5* in which formation of the coiled coil thrusts the unfolded foot domain outwards (Fig. 2b). We screened a series of double cysteine mutants designed to produce disulfide links within the foot and therefore stabilize its folded structure. We obtained the most robust recoating from a variant with cysteines at positions 567 and 590 in RRV VP4 (Fig. 3), denoted VP4(S567C/A590C). Recoating DLPs with this mutant protein (along with wild-type VP7) yielded TLPs with an infectivity 1,600-fold lower than that of controls recoated with wild-type VP4 (Fig. 3a).
a, Particle (P) to focus-forming unit (FFU) ratios for TLPs, wild-type (WT) rcTLPs and rcTLPs containing the mutant protein VP4 (S567C/A590C), determined as described in Methods. n = 3 biologically independent experiments. Data are mean ± s.d. One-way ANOVA with Tukey post-hoc test. ****P < 0.0001, NS, not significant. For source data, see Supplementary Data 4. b, Structure of the foot-locked intermediate obtained from cryo-EM analysis of rcTLPs containing VP4(S567C/A590C). VP5*, red, orange and salmon; VP6, green; VP7, yellow; VP8*, magenta. c, Left, structural comparison between the VP5* foot domains of the upright conformation of wild-type rcTLPs (red) and the intermediate conformation of rcTLPs containing VP4 (S567C/A590C) (grey). Right, structural comparison between the VP5* β-barrel domains of the reversed conformation of wild-type rcTLPs (red) and the intermediate conformation of rcTLPs containing VP4 (S567C/A590C) (grey). d, Comparison of relative amounts of VP5* determined by western blotting (144 ng of each substrate), as described in Methods. Experiment repeated independently twice with similar results. For western blot source images, see Supplementary Fig. 1.
Cryo-EM analysis of rcTLPs bearing VP4(S567C/A590C) spikes yielded, after initial icosahedral reconstruction, approximately 216,000 VP4 positions (60 per virion particle). Classification of the VP4 positions showed the mature spike conformation (same as Fig. 1d, left), empty positions without VP4, and a new class in which VP4 contained three outward projecting β-barrel domains but no central, three-chain coiled coil (Fig. 3b). Instead, the foot domain of VP5* was firmly in place, and its structure—at the resolution we obtained—was the same as that of a VP5* foot domain on a wild-type rcTLP (Figs. 3b, c). The anchored VP8* N-terminal segments were present, but we detected no clearly interpretable density beyond residue 16, where the three polypeptide chains encountered the inward-facing ends of the VP5* β barrels (Fig. 2b).
When rcTLPs containing VP4(S567C/A590C) were treated briefly at pH 11 and then reneutralized, we found that most (69%) of the spikes were in the intermediate conformation (Extended Data Fig. 8c). The remaining 31% had converted to reversed trimers, probably because of inefficient formation of the disulfide bond.
Spike rearrangement during cell entry
A previous electron cryotomography (cryo-ET) study of RRV TLPs entering BSC-1 cells at the rim—where they were thin enough for imaging—showed that the spacing at the virion–membrane interface between the membrane and the outer surface of the VP7 layer could be either ‘loose’ (around 120 Å) or ‘tight’ (around 60 Å), and that projections representing VP4 spanned both spacings8. The spikes spanning the wider gap had approximately the length expected for the upright conformation8. To extend the analysis, we recorded more than 100 new tomographic datasets, extracted individual sub-tomograms of nearly fully engulfed particles, and calculated icosahedral averages for each (Fig. 4a, Extended Data Fig. 9). The loose engulfment corresponds to the spike contacts expected for initial lectin contacts with glycolipid, although the resolution of the icosahedral average is not sufficient to show the contacts directly. The tight engulfment corresponds closely to our expectation from Fig. 1d (right): the volume occupied by the foot in the unrearranged structure is empty, and the gap between the surface of VP7 and the membrane would place the VP5* hydrophobic loops at the surface of the bilayer, with the extruded foot apparently embedded in the membrane. We infer that the transformation that we have detected on the surface of virions in vitro also occurs during cell entry, as a key step in the delivery of the DLP into the cytosol.
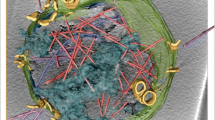
a, Cryo-ET reconstructions of RRV entering at the thin edge of BSC-1 cells. Tomographic analysis is shown for two viral particles that are representative of a ‘loose’ (left) or a ‘tight’ (right) virion–membrane (M) interface. Middle, top, slices of tomographic reconstructions (low-pass-filtered and contrast-enhanced; scale bars, 50 nm); middle, bottom, 3D tomographic reconstructions with icosahedral symmetry imposed. In the magnified views of the icosahedrally averaged tomographic reconstructions, density is partially cut and shown in solid grey; protein subunits of positioned atomic models are shown in ribbon representation and coloured as in Fig. 1. b, Model of membrane interaction and the rearrangement of VP5* and VP8*; see Discussion for details.
Discussion
A transition of VP4 from an upright to a reversed conformation requires the following steps (Fig. 4b): the separation of the VP8* lectin domains from the tips of the two-fold clustered, VP5* β-barrel domains; an outward flip of the third VP5* β barrel and formation of a β-barrel trimer; and the unfolding and outward projection of the foot domain by zippering of the central, three-strand, α-helical coiled coil. Formation of the observed β-barrel trimer requires that the lectin domains dissociate from the tips of the β barrels, but they remain tethered to the particle by the long N-terminal segment of VP8* (VP8* α and VP8* linker, Fig. 4b). Even after the foot has unfolded, the two complete VP8* fragments do not dissociate19 (Fig. 2b, right, Extended Data Fig. 8a).
In its reversed conformation, VP5* remains bound to the virion through contacts with three of the six VP7 subunits that surround it (Fig. 2a, right, Extended Data Figs. 6a, 7); the same contacts are present in the structure of the ‘foot-locked’ mutant VP4(S567C/A590C). A smooth transition from foot-locked to reversed VP5* is possible, because adjacent β barrels have no direct contact in the former structure, and unfolding and outward projection of the foot can occur without dissociation (Extended Data Fig. 6b). The foot-locked structure is therefore a plausible representation of an intermediate in the overall conformational change (Supplementary Discussion).
The subtomogram averages in Fig. 4a show that the molecular rearrangements seen in vitro correspond to events that occur during virus entry into a host cell. The absence of density in the foot cavity supports our proposed function for the conformational change. The very low infectivity of virus particles recoated with foot-locked VP4 indicates that complete rearrangement is necessary for virion penetration.
Figure 4b suggests how the findings reported here might map onto the sequence of events that occur during cell entry, as derived from live-cell imaging experiments8,16,20. Steps in this sequence are membrane interaction, invagination, engulfment, Ca2+ loss (Supplementary Discussion), VP4 and VP7 loss, and DLP release (Fig. 1b). Membrane deformation requires more free energy than is yielded by simple attachment; interaction of the VP5* hydrophobic loops with the membrane bilayer might be necessary for the observed ‘wrapping’ (Fig. 4a, centre). The interaction of the hydrophobic loops might occur initially through the two β-barrel domains that project in the spike conformation. Spontaneous fluctuations could generate a local dissociation of the VP8* lectin domains (arrows in Fig. 4b, left), enabling the adjacent membrane to capture the hydrophobic loops of the two β-barrel domains that project in the upright conformation. Transition to a state that resembles the foot-locked (intermediate) structure could then ensue (Fig. 4b, centre), followed by unfurling of the foot, formation of the coiled coil, and interaction of the foot with the target membrane (Fig. 4b, right).
During entry into a host cell, at least two distinct regions of VP5* interact with the lipid bilayer of the membrane: the hydrophobic loops that are exposed by dissociation of the VP8* lectin domain, and the foot that is translocated outward by the conformational change described here. Determining how multiple local perturbations, each introduced by insertion of three foot regions into the target membrane, promote longer-range disruption (‘perforation’; Fig. 1b) and DLP release will require high-resolution visualization of the inserted foot regions.
Like other non-enveloped animal viruses with dsDNA or dsRNA genomes, rotaviruses initiate infection of a host cell by delivering a modified, but still assembled, particle into the cytosol. The structures described here define—more closely than has been achieved so far for other dsRNA or dsDNA viruses—how rotaviruses deliver a very large, macromolecular payload. The molecular description of the rotavirus delivery mechanism, and those of other viruses that insert a modified virion or subviral particle into the cytosol, could inform efforts to design delivery vehicles for biological molecules that have intracellular targets.
Methods
No statistical methods were used to predetermine sample size. The experiments were not randomized and the investigators were not blinded to allocation during experiments and outcome assessment.
Cells, plasmids and constructs
MA104 cells (American Type Culture Collection, ATCC) were cultured in M199 media (Invitrogen) supplemented with 25 mM HEPES and 10% HI-FBS (Invitrogen). BSC-1 cells (ATCC) were cultured in DMEM (Invitrogen) supplemented with 10% HI-FBS (Invitrogen). For VP4 and VP7 expression, full-length genomic sequences from rhesus rotavirus (G3 serotype, NCBI: txid444185) were amplified by PCR and cloned into pFastbac (Thermo Fisher Scientific) expression vectors. Mutations were introduced by quick-change mutagenesis in DH10α cells (Thermo Fisher Scientific). Purified plasmid constructs were transfected into DH10-Bac cells (Thermo Fisher Scientific). Purified bacmids were transfected into Sf9 cells (ATCC) grown in Sf-900 II SFM media supplemented with 1% penicillin-streptomycin.
Production and purification of RRV TLPs and DLPs
MA104 cells were grown to confluency in 850 cm2 roller bottles, followed by exchange of medium to M199 supplemented with 1 μg ml−1 trypsin (Gibco). Cells were infected with trypsin-activated RRV particles (G3 serotype) at a multiplicity of infection of 0.1 and incubated for 24 h. Media and cell debris were collected and frozen for storage at −80 °C. Virus in thawed supernatants was concentrated by ultracentrifugation (225,500g, 30 min), added to thawed cell debris pellets (which contain most of the virus), and extracted by sonication with Freon-113 as described21,22. The aqueous phases of three successive rounds of freon extraction were combined, and TLPs were separated from DLPs by caesium chloride gradient centrifugation. TLPs were dialysed twice against 2 l of 20 mM Tris-HCl pH 8.0, 100 mM NaCl, 1 mM CaCl2 (TNC) and DLPs twice against 2 l of 20 mM HEPES, pH 7.4, 100 mM NaCl (HN). The concentration of the purified virus was determined by SDS–PAGE densitometry of VP6 bands against standards of DLP stocks ranging from 0.1 to 1.0 mg ml−1.
Expression and purification of VP4 and VP7
Recombinant VP4 and VP7 were expressed in Sf9 insect cells, using the Bac-to-Bac expression system as described previously23,24. In brief, insect cells (2 million cells per ml) were infected with baculovirus inoculum passaged three times in Sf9 cells. For VP4, cells were collected 72 h after infection and resuspended in 100 ml lysis buffer (75 mM Tris-HCl pH 8.0, 100 mM NaCl, 5 mM EDTA, 7.5% glycerol). After freeze thawing, cells were lysed in a Dounce homogenizer and the homogenate cleared by centrifugation (70,409g, 30 min). VP4 was purified from the cleared supernatant by ammonium sulfate precipitation followed by hydrophobic interaction chromatography (Phenyl FF column) and anion exchange chromatography (Q Sepharose column)25. Final purification was performed on a size exclusion chromatography column (S200) equilibrated in HNE (20 mM HEPES pH 7.3, 100 mM NaCl, 1 mM EDTA). Pure VP4 was concentrated to 5 mg ml−1, flash-frozen with liquid nitrogen, and stored at −80 °C. For VP7, cells were removed from media 72 h after infection and the supernatant was loaded onto a Concavalin A resin. After elution with α-methyl-mannose, protein was adsorbed to a resin with immobilized M159 antibody26 (5 mg per ml of Protein G Sepharose resin) and eluted with HNE15. Fractions containing VP7 were pooled and concentrated to 3 mg ml−1, flash-frozen with liquid nitrogen, and stored at −80 °C.
Recoating of RRV DLPs
Recoating DLPs with VP4 and VP7 to form rcTLPs was carried out following previously described protocols16. In brief, 1 M sodium acetate (pH 5.2) was added to a volume of DLP resulting in a final concentration of 100 mM sodium acetate. VP4 was added to a final concentration of 0.9 mg ml−1 (around 33-fold excess) and the mixture was incubated at room temperature for 1 h. VP7 was then added in 2.3-fold excess along with a final addition of sodium acetate and CaCl2 to maintain their respective concentrations of 100 mM and 1 mM. The mixture was incubated at room temperature for 1 h, and recoated particles were separated from excess components by ultracentrifugation (112,000g, 30 min). Pellets were resuspended in 150 μl of TNC, additional TNC was added to a final volume of 1 ml, and centrifugation was repeated. Final pellets were resuspended in 50 μl of TNC.
Infectivity assay of TLPs and rcTLPs
Titres and particles to focus-forming unit (FFU) ratios for TLP and rcTLPs were determined by infectious focus assays as previously described16, and specific infectivities were derived from concentration measurements on the basis of densitometry of a western blot using a primary antibody specific for VP6 (antibody 2B4, Santa Cruz Biotechnology, 1:4,000 dilution). We determined the relative amounts of VP5* by western blotting of 144 ng of purified particles with antibody HS227 (1:5,000 dilution) in combination with horseradish peroxidase (HRP)-coupled goat anti-mouse IgG (1:5,000 dilution).
pH shift experiments with TLPs and rcTLPs
RRV particles were incubated with 5 μg ml−1 trypsin for 30 min at 37 °C and the reaction was quenched by the addition of 1 mM phenylmethylsulfonyl fluoride (PMSF). pH-induced conformational change was achieved by adding a 0.83 M N-cyclohexyl-3-aminopropanesulfonic acid (CAPS) buffer at pH 11.0 to a final concentration of 100 mM. As a control, we used 100 mM of Tris-HCl pH 8.0. Samples were incubated for 30 min at room temperature and neutralized to pH 8.0 by addition of 1 ml of TNC (20 mM Tris-HCl, 100 mM NaCl, 1 mM CaCl2). Viral particles were pelleted in a TLS 55 rotor (50,000 rpm, 1 h, 4 °C). Supernatants were removed and the pellets were resuspended in 50–80 μl TNC.
VP8* fractionation and detection by western blotting
We digested 12 μg of TLPs with 5 μg ml−1 trypsin in a total volume of 12 μl for 30 min at 37 °C. Trypsin was quenched by adding 0.6 μl of aprotinin and 1.2 μl of PMSF to final concentrations of 2 μg ml−1 and 1 mM, respectively. We shifted the pH by adding 1.2 μl of 1 M CAPS buffer at pH 11.0 (or 1.2 μl of 1 M TRIS pH 8.0 for the control samples), incubated the samples for 30 min at room temperature, and re-neutralized them by adding 1 ml TNC (20 mM Tris-HCl pH 8.0, 100 mM NaCl, 1 mM CaCl2) or 1 ml TN2E (20 mM Tris-HCl pH 8.0, 100 mM NaCl, 2 mM EDTA). We monitored the pH shift and re-neutralization using pH strips. Samples were then incubated for 30 min at 37 °C to promote uncoating of TLPs. TLPs and uncoated DLPs were then pelleted in a TLS 55 rotor (50,000 rpm, 1 h, 4 °C), resuspended in 1 ml buffer and pelleted again (50,000 rpm, 1 h, 4 °C). We carefully removed the supernatants and resuspended the pellets in 50 μl either TNC or TN2E. We analysed 10 μl of each sample by SDS–PAGE and silver staining, and also by SDS–PAGE and western blotting with the VP8*-specific antibody HS127 (1:500 dilution of mouse HS1 ascitic fluid and 1:5,000 dilution of HRP-coupled goat anti-mouse IgG).
Sample preparation for single-particle cryo-EM
RRV rcTLPs or TLPs (2.5 μl) at concentrations between 1 and 2 mg ml−1 were treated with 5 μg ml−1 trypsin at 37 °C for 30 min, after which trypsin was inactivated with PMSF at a final concentration of 1 mM. The trypsin-treated particles were then incubated with 2.5 μl TNC for a further 30 min at 37 °C, followed by application of 4 μl to a C-flat holey carbon grid (Electron Microscopy Science), blotting and freezing with a CP3 plunge freezer (Gatan). The efficiency of the spike transition at pH 8.0 from upright to reversed was noticeably higher for rcTLPs than for authentic TLPs. We therefore used the former to generate samples for electron microscopy of particles that had been incubated at pH 8.0. Because expression and purification of recombinant VP4 and VP7 involve quite different conditions than does virion propagation, we have not tried to track down the source of the difference. Previous structural studies show no substantial structural differences, and specific infectivity of the rcTLPs is sometimes even higher than that of authentic virions.
Single-particle cryo-EM and data processing
Vitrified grids were imaged with a Tecnai Polara F30 microscope (FEI/Thermo Fisher Scientific) operated at 300 kV and images were recorded with a Gatan K2 direct electron detector. Movies of rotavirus particles were taken with an exposure time of 10 s and 0.2 s per frame (50 frames) using a physical pixel size of 1.231 Å in counting mode. Movies were motion-corrected with MotionCor2 (5 × 5 patch alignment)28. We used Gautomatch for viral particle picking with template projections obtained with EMAN229 from a previous reconstruction30. For automatic particle picking, the angular sampling was 3° and the low-pass filter cutoff was 40 Å. We determined the contrast transfer function (CTF) parameters using Gctf31 from total-summed images. We fitted defocus and astigmatism parameters at individual particle coordinates and used relion_preprocess32 for particle extraction. We aligned the rotavirus particle images and calculated reconstructions with cisTEM (refine3d v.1.01, reconstruct3d version 1.02)33 as described previously30. We imposed icosahedral symmetry in setting I2 for alignment and reconstruction and used a spherical shell mask for the reference with an inner radius of 222 Å, and an outer radius of 403 Å (corresponding to the VP2, VP6 and VP7 protein layers of the TLP). After the initial alignment, we used relion_motion_refine and relion_ctf_refine to obtain a polished particle stack and parameters for beam-tilt correction, respectively34, and carried out a second round of alignment with cisTEM. The Fourier shell correlation (FSC)-based resolution estimates for density within the spherical shell were 3.1 Å for the wild-type rcTLP reconstruction and 3.9 Å for the rcTLP VP4(S567C/A590C) reconstruction (Extended Data Table 1).
Local reconstruction of VP4 structures
We used subparticle image analysis for local reconstructions and classification of VP4 structures (Extended Data Fig. 2). Similar to what we previously described for structure determination of the VP1 rotavirus polymerase30,35, we used signal subtraction of non-VP4 proteins (except a single VP7 trimer) with relion_project. We extracted subparticle images (corresponding to the location of VP4 proteins) on the basis of the icosahedral alignment from the original polished and signal-subtracted particle stacks using IMOD36. We used the programs refine3d (v.1.01) and reconstruct3D (v.1.02) from cisTEM for subparticle classification. We kept the subparticle alignment parameters constant during iterative classification (40–80 cycles); the high-resolution limit for classification was 8 Å; and we 3D-masked the references with a single mask encompassing the volume of all VP4 conformations and a single adjacent VP7 trimer. Per-particle weighting in the reconstruction step was essentially turned off by setting BSC (the parameter in cisTEM, which converts the difference between a particle score and the score average into a B factor33) to 0.0. After classification, we calculated final maps for each class from non-signal-subtracted particle stacks and applied standard postprocessing steps. Because classification reduced the number of particles per class compared to the icosahedral reconstruction (with 60-fold averaging), the final resolution of the local reconstructions was slightly lower, ranging from 3.3 to 4.3 Å resolution (Extended Data Table 1). FSC plots and local-resolution analyses of the local reconstructions are shown in Extended Data Fig. 3. Density for the VP8* lectin and the dimeric VP5* β-barrel domains was poorly resolved in the upright spike conformation local reconstruction because of flexibility at the base of the dimeric spike projection (Extended Data Fig. 3a). We therefore improved this region of the map using alignment by classification. In this approach, we iteratively classified without alignment (6 classes, 5 Å high-resolution limit for classification, 80 cycles, 3D mask encompassing the VP8* lectin and VP5* β-barrel domains), superimposed the maps of the classes, and updated particle alignment parameters according to the 3D map alignment. The resulting 4 Å-resolution map enabled us to model the connection between the VP8* N terminus (located in the foot) and the peripheral VP8* lectin domains.
Structure modelling, refinement and validation
Our atomic models consist of the trimetric VP4 spike protein (VP5* and VP8*) surrounded by six trimers of VP6 and VP7 each (some of the VP6 and VP7 subunits are related by icosahedral symmetry of the virion). We initially placed subunits obtained from published structures with Protein Data Bank (PDB) IDs 4V7Q6 and 1SLQ (in the case of the VP5* reversed conformation)18 into our local reconstruction maps. We manually adjusted and completed the models using the programs O37 and Coot38, and used phenix.real_space_refine39 for structure refinement. We used standard stereochemical and B-factor restraints, as well as Ramachandran, rotamer, and secondary structure restraints. Residues included in the models are summarized in Extended Data Table 1. We validated the models with MolProbity40. Model statistics are in Extended Data Table 1. The refined structures show expected FSC values with the cryo-EM maps (Extended Data Fig. 3). Magnified views of density map sections are shown in Extended Data Fig. 4a. To locate structural changes between the upright and reversed structures, we calculated per-residue Cα distances after subunit-wise superposition of VP4, VP7 and VP6 subunits from the upright and reversed conformation structures (Extended Data Fig. 4b).
Sample preparation for cryo-ET
BSC-1 cells, grown on gold grids and incubated with rotavirus TLPs, were plunge-frozen for cryo-ET as previously described8. In brief, Quantifoil gold grids (EMS Q250-AR2) with a holey carbon film (R2/2, 200 mesh) were glow-discharged at −40 mA for 30 s, coated with 0.1% poly-l-lysine hydrobromide (Sigma P1524-25MG), and sterilized with 70% ethanol for 10 min. BSC-1 cells were cultured on these grids in DMEM with 10% FBS and 1x Glutamax at 37 °C and 5% CO2 in a 35-mm-diameter glass bottom dish (MatTek, P35G-1.5-14-C) for 24 h before inoculation. Grids were then washed by aspirating the medium from the dish and replacing it with MEM α (Gibco 41061-029) buffer at 37 °C. This process was repeated (with a total amount of about 2 ml buffer) until the buffer remained clear. Grids were then transferred to glass light microscope slides covered with Parafilm. Just before inoculation, the virus was activated using trypsin stock solution (50 μg ml−1 in TNC (pH 8.0)) at 37 °C for 30 min. After diluting the activated virus 1:5 or 1:10 in MEM α, 5 μl of the solution was immediately added to the cells at 37 °C for inoculation. The grids were then incubated at 37 °C for a period of between 5 and 45 min, before removing the buffer by blotting the grid from the edge with filter paper while simultaneously adding 5 μl of concentrated, BSA-coated 10-nm colloidal gold (Sigma-Aldrich)41. Grids were then blotted from the front with filter paper for 7–8 s and plunge-frozen in liquid ethane using a manual plunge-freezing device. The frozen grids were stored in liquid nitrogen.
cryo-ET and data processing
Vitrified grids were imaged using a Tecnai F30 transmission electron microscope (FEI/Thermo Fischer Scientific) equipped with a field emission gun operating at an accelerating voltage of 300 kV, GIF2000 post column energy filter (Gatan) operated in zero-loss mode with a 20 eV slit width, and a 2k × 2k charge-coupled device camera (Gatan). The cells were screened in low-dose mode for areas of interest—that is, areas that contained virus particles interacting with the edge of BSC-1 cells that were thin enough for cryo-ET imaging (up to 300-nm thick). In each area of interest, a single-axis tilt series was recorded in low-dose mode using the microscope control software SerialEM42. The sample was tilted over a range of about −65° to +65° with 1.5–2.5° increments. The total accumulated electron dose per tilt series was limited to around 100 e− Å−2. The defocus was −6 μm to −8 μm and the pixel size was 1.077 nm or 0.986 nm, respectively. The software IMOD43 was used for fiducial alignment of the tilt series images and tomogram reconstruction using weighted back projection. We analysed virions from 90 raw cryo-tomograms by extracting virus-particle subtomograms with a box size of 150 × 150 × 150 pixels. We contrast-inverted and normalized the extracted viral particle subtomograms, and 3D-aligned them to a low-pass filtered reference using e2spt_classaverage.py and e2spt_extract_align.py from EMAN2 (v.2.2)44. At this stage, subtomographic viral densities were oriented in a standard I4 icosahedral symmetry setting, which enabled us either to calculate an icosahedral density average for each particle by simply applying icosahedral symmetry, or to extract sub-subtomograms at individual spikes positions, or to average selected subtomograms (individual viruses) or sub-subtomograms (individual VP4 trimer positions) (Fig. 4a, Extended Data Fig. 9).
Figure preparation
We prepared the figures using PyMOL (The PyMOL Molecular Graphics System v.2.1, Schrödinger, LLC), POV-Ray (https://www.povray.org), and matplotlib45. We retrieved rotavirus sequences from GenBank46 with BioPython47, aligned them with MAFFT48, and used ESPript49 to display the multiple sequence alignments of VP4 (Supplementary Data 1), VP7 (Supplementary Data 2) and VP6 (Supplementary Data 3). Serotypes and genotypes were compiled from published studies50,51,52,53.
Reporting summary
Further information on research design is available in the Nature Research Reporting Summary linked to this paper.
Data availability
The maps of the cryo-EM reconstructions have been deposited in the Electron Microscopy Data Bank (EMDB) (accession numbers: EMD-21955, upright conformation; EMD-21956, intermediate conformation; EMD-21957, reversed conformation), and the refined coordinates in the Protein Data Bank (PDB) (PDB IDs: 6WXE, upright conformation; 6WXF, intermediate conformation; 6WXG, reversed conformation). We obtained previously published rotavirus structures from the PDB (PDB IDs: 4V7Q and 1SLQ). The rotavirus protein sequences used to prepare the sequence alignments (shown in Supplementary Data 1–3) were retrieved from GenBank46. The accession numbers for VP4 are: RRV, P12473; SA11, P12976; WA, P11193; S2, AQT31697; DS1, P11196; ST3, P11200; AU-1, P39033; 116E, Q09113; 69M, P26451; NCDV, P17465; UK, P12474; 993/83, Q08010; Gottfried, P23045; OSU, P11114; CH3, Q86184; L338, Q98636; EW, Q83450. The accession numbers for VP7 are: RRV, P12473; SA11, P12976; WA, P11193; S2, AQT31697; DS1, P11196; ST3, P11200; AU-1, P39033; 116E, Q09113; 69M, P26451; NCDV, P17465; UK, P12474; 993/83, Q08010; Gottfried, P23045; OSU, P11114; CH3, Q86184; L338, Q98636; EW, Q83450. The accession numbers for VP6 are: RRV, P12473; SA11, P12976; WA, P11193; S2, AQT31697; DS1, P11196; ST3, P11200; AU-1, P39033; 116E, Q09113; 69M, P26451; NCDV, P17465; UK, P12474; 993/83, Q08010; Gottfried, P23045; OSU, P11114; CH3, Q86184; L338, Q98636; EW, Q83450. All data are available from the corresponding author upon reasonable request.
Code availability
The software programs used to generate and analyse the data of this study are publicly available. Custom-written C shell and Python scripts used to run the programs are available from the corresponding author on reasonable request.
References
Harrison, S. C. in Fields Virology 6th edn (eds Knipe, D. M. & Howley, P. M.) 52–86 (Lippincott Williams and Wilkins, 2013).
Estes, M. K. & Greenberg, H. in Fields Virology 6th edn (eds Knipe, D. M. & Howley, P. M.) 1347–1401 (Lippincott Williams and Wilkins, 2013).
Trask, S. D., Ogden, K. M. & Patton, J. T. Interactions among capsid proteins orchestrate rotavirus particle functions. Curr. Opin. Virol. 2, 373–379 (2012).
Tihova, M., Dryden, K. A., Bellamy, A. R., Greenberg, H. B. & Yeager, M. Localization of membrane permeabilization and receptor binding sites on the VP4 hemagglutinin of rotavirus: implications for cell entry. J. Mol. Biol. 314, 985–992 (2001).
Kim, I. S., Trask, S. D., Babyonyshev, M., Dormitzer, P. R. & Harrison, S. C. Effect of mutations in VP5 hydrophobic loops on rotavirus cell entry. J. Virol. 84, 6200–6207 (2010).
Settembre, E. C., Chen, J. Z., Dormitzer, P. R., Grigorieff, N. & Harrison, S. C. Atomic model of an infectious rotavirus particle. EMBO J. 30, 408–416 (2011).
Aoki, S. T. et al. Structure of rotavirus outer-layer protein VP7 bound with a neutralizing Fab. Science 324, 1444–1447 (2009).
Abdelhakim, A. H. et al. Structural correlates of rotavirus cell entry. PLoS Pathog. 10, e1004355 (2014).
Trask, S. D., Kim, I. S., Harrison, S. C. & Dormitzer, P. R. A rotavirus spike protein conformational intermediate binds lipid bilayers. J. Virol. 84, 1764–1770 (2010).
Dormitzer, P. R. et al. Specificity and affinity of sialic acid binding by the rhesus rotavirus VP8* core. J. Virol. 76, 10512–10517 (2002).
Delorme, C. et al. Glycosphingolipid binding specificities of rotavirus: identification of a sialic acid-binding epitope. J. Virol. 75, 2276–2287 (2001).
Martínez, M. A., López, S., Arias, C. F. & Isa, P. Gangliosides have a functional role during rotavirus cell entry. J. Virol. 87, 1115–1122 (2013).
Ramani, S. et al. The VP8* domain of neonatal rotavirus strain G10P[11] binds to type II precursor glycans. J. Virol. 87, 7255–7264 (2013).
Hu, L. et al. Cell attachment protein VP8* of a human rotavirus specifically interacts with A-type histo-blood group antigen. Nature 485, 256–259 (2012).
Dormitzer, P. R., Greenberg, H. B. & Harrison, S. C. Purified recombinant rotavirus VP7 forms soluble, calcium-dependent trimers. Virology 277, 420–428 (2000).
Salgado, E. N., Garcia Rodriguez, B., Narayanaswamy, N., Krishnan, Y. & Harrison, S. C. Visualization of calcium ion loss from rotavirus during cell entry. J. Virol. 92, e01327-18 (2018).
Rodríguez, J. M. et al. New insights into rotavirus entry machinery: stabilization of rotavirus spike conformation is independent of trypsin cleavage. PLoS Pathog. 10, e1004157 (2014).
Dormitzer, P. R., Nason, E. B., Prasad, B. V. & Harrison, S. C. Structural rearrangements in the membrane penetration protein of a non-enveloped virus. Nature 430, 1053–1058 (2004).
Pesavento, J. B., Crawford, S. E., Roberts, E., Estes, M. K. & Prasad, B. V. pH-induced conformational change of the rotavirus VP4 spike: implications for cell entry and antibody neutralization. J. Virol. 79, 8572–8580 (2005).
Salgado, E. N., Upadhyayula, S. & Harrison, S. C. Single-particle detection of transcription following rotavirus entry. J. Virol. 91, e00651-17 (2017).
Smith, R. E., Zweerink, H. J. & Joklik, W. K. Polypeptide components of virions, top component and cores of reovirus type 3. Virology 39, 791–810 (1969).
Street, J. E., Croxson, M. C., Chadderton, W. F. & Bellamy, A. R. Sequence diversity of human rotavirus strains investigated by northern blot hybridization analysis. J. Virol. 43, 369–378 (1982).
Fiore, L. et al. Antigenicity, immunogenicity and passive protection induced by immunization of mice with baculovirus-expressed VP7 protein from rhesus rotavirus. J. Gen. Virol. 76, 1981–1988 (1995).
Mackow, E. R., Barnett, J. W., Chan, H. & Greenberg, H. B. The rhesus rotavirus outer capsid protein VP4 functions as a hemagglutinin and is antigenically conserved when expressed by a baculovirus recombinant. J. Virol. 63, 1661–1668 (1989).
Trask, S. D. & Dormitzer, P. R. Assembly of highly infectious rotavirus particles recoated with recombinant outer capsid proteins. J. Virol. 80, 11293–11304 (2006).
Greenberg, H. B. et al. Production and preliminary characterization of monoclonal antibodies directed at two surface proteins of rhesus rotavirus. J. Virol. 47, 267–275 (1983).
Padilla-Noriega, L. et al. Serologic analysis of human rotavirus serotypes P1A and P2 by using monoclonal antibodies. J. Clin. Microbiol. 31, 622–628 (1993).
Zheng, S. Q. et al. MotionCor2: anisotropic correction of beam-induced motion for improved cryo-electron microscopy. Nat. Methods 14, 331–332 (2017).
Bell, J. M., Chen, M., Baldwin, P. R. & Ludtke, S. J. High resolution single particle refinement in EMAN2.1. Methods 100, 25–34 (2016).
Jenni, S. et al. In situ structure of rotavirus VP1 RNA-dependent RNA polymerase. J. Mol. Biol. 431, 3124–3138 (2019).
Zhang, K. Gctf: Real-time CTF determination and correction. J. Struct. Biol. 193, 1–12 (2016).
Scheres, S. H. RELION: implementation of a Bayesian approach to cryo-EM structure determination. J. Struct. Biol. 180, 519–530 (2012).
Grant, T., Rohou, A. & Grigorieff, N. cisTEM, user-friendly software for single-particle image processing. eLife 7, e35383 (2018).
Scheres, S. H. Beam-induced motion correction for sub-megadalton cryo-EM particles. eLife 3, e03665 (2014).
Ding, K. et al. In situ structures of rotavirus polymerase in action and mechanism of mRNA transcription and release. Nat. Commun. 10, 2216 (2019).
Mastronarde, D. N. & Held, S. R. Automated tilt series alignment and tomographic reconstruction in IMOD. J. Struct. Biol. 197, 102–113 (2017).
Jones, T. A., Zou, J. Y., Cowan, S. W. & Kjeldgaard, M. Improved methods for building protein models in electron density maps and the location of errors in these models. Acta Crystallogr. A 47, 110–119 (1991).
Emsley, P. & Cowtan, K. Coot: model-building tools for molecular graphics. Acta Crystallogr. D 60, 2126–2132 (2004).
Afonine, P. V. et al. Real-space refinement in PHENIX for cryo-EM and crystallography. Acta Crystallogr. D 74, 531–544 (2018).
Chen, V. B. et al. MolProbity: all-atom structure validation for macromolecular crystallography. Acta Crystallogr. D 66, 12–21 (2010).
Iancu, C. V. et al. Electron cryotomography sample preparation using the Vitrobot. Nat. Protoc. 1, 2813–2819 (2006).
Mastronarde, D. N. Automated electron microscope tomography using robust prediction of specimen movements. J. Struct. Biol. 152, 36–51 (2005).
Kremer, J. R., Mastronarde, D. N. & McIntosh, J. R. Computer visualization of three-dimensional image data using IMOD. J. Struct. Biol. 116, 71–76 (1996).
Galaz-Montoya, J. G., Flanagan, J., Schmid, M. F. & Ludtke, S. J. Single particle tomography in EMAN2. J. Struct. Biol. 190, 279–290 (2015).
Hunter, J. D. Matplotlib: A 2D graphics environment. Comput. Sci. Eng. 9, 90–95 (2007).
Benson, D. A. et al. GenBank. Nucleic Acids Res. 46, D41–D47 (2018).
Cock, P. J. et al. Biopython: freely available Python tools for computational molecular biology and bioinformatics. Bioinformatics 25, 1422–1423 (2009).
Katoh, K., Misawa, K., Kuma, K. & Miyata, T. MAFFT: a novel method for rapid multiple sequence alignment based on fast Fourier transform. Nucleic Acids Res. 30, 3059–3066 (2002).
Gouet, P., Courcelle, E., Stuart, D. I. & Métoz, F. ESPript: analysis of multiple sequence alignments in PostScript. Bioinformatics 15, 305–308 (1999).
Gorziglia, M., Larralde, G., Kapikian, A. Z. & Chanock, R. M. Antigenic relationships among human rotaviruses as determined by outer capsid protein VP4. Proc. Natl Acad. Sci. USA 87, 7155–7159 (1990).
Martella, V. et al. Molecular analysis of the VP7, VP4, VP6, NSP4, and NSP5/6 genes of a buffalo rotavirus strain: identification of the rare P[3] rhesus rotavirus-like VP4 gene allele. J. Clin. Microbiol. 41, 5665–5675 (2003).
Matthijnssens, J. et al. Full genome-based classification of rotaviruses reveals a common origin between human Wa-like and porcine rotavirus strains and human DS-1-like and bovine rotavirus strains. J. Virol. 82, 3204–3219 (2008).
Patton, J. T. Rotavirus diversity and evolution in the post-vaccine world. Discov. Med. 13, 85–97 (2012).
Afonine, P. V. phenix.mtriage: a tool for analysis and validation of cryo-EM 3D reconstructions. Comput. Crystallogr. Newsl. 8, 25 (2017).
Kucukelbir, A., Sigworth, F. J. & Tagare, H. D. Quantifying the local resolution of cryo-EM density maps. Nat. Methods 11, 63–65 (2014).
Acknowledgements
We thank Z. Li, S. Sterling, R. Walsh and S. Rawson for assistance and guidance at the Harvard Medical School Cryo-EM Center for Structural Biology and the Harvard Medical School Molecular Electron Microscopy Suite; C. Xu for assistance at the Brandeis University cryo-EM facility; and H. B. Greenberg for the gift of the HS1 and HS2 antibodies. The work was supported by National Institutes of Health grant CA-13202 (to S.C.H.). S.C.H. is an Investigator in the Howard Hughes Medical Institute.
Author information
Authors and Affiliations
Contributions
T.H., E.N.S., D.N., S.J. and S.C.H. designed the experiments; T.H., R.T., E.N.S. and C.B. conducted the experiments and recorded data; T.H., R.T., D.S. and S.J. analysed the data; T.H. and S.J. determined structures and built models; T.H., S.J. and S.C.H. wrote the paper; and T.H., C.B., D.S., D.N., S.J. and S.C.H. revised and edited the paper.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Nature thanks David Bhella, John Patton and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Peer reviewer reports are available.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Sample preparation and cryo-EM data collection.
a, Schematic protocol for recoating of double-layer particles (DLPs) with recombinant VP4 and VP7. b, Time course of the digestion of wild-type recoated triple-layer particles (wt rcTLPs) with 5 μg ml−1 trypsin at 37 °C. Samples were analysed after the shown incubation times by SDS–PAGE. The experiment was repeated independently twice with similar results. For gel source data, see Supplementary Fig. 1. c, Representative micrograph (aligned and summed movie frames) of wt rcTLPs recorded with a Polara F30 electron microscope equipped with a K2 summit detector (magnification, 40,650). Scale bar, 100 nm. d, Power spectrum of the micrograph shown in c.
Extended Data Fig. 2 Data processing for local reconstructions.
Full workflow for local reconstructions of rotavirus spike proteins.
Extended Data Fig. 3 Resolution analysis of the local cryo-EM reconstructions.
Left, Fourier shell correlation (FSC) curves for the reconstructions and refined models calculated with phenix.mtriage54. Correlations for the two half maps are shown as solid blue lines after applying a mask encompassing the models. Correlations between the refined model and final map are shown as red solid lines. Nominal resolution estimates at conventional FSC values are indicated by arrows. Right, local resolution of the reconstructions calculated with ResMap55. a, Upright conformation. Dashed lines (left) are the FSC analysis for the reconstruction of the distal VP5*/VP8* dimeric density that was obtained through alignment by classification (see Methods). The images on the right show the local resolution of this reconstruction. b, Intermediate conformation. c, Reversed conformation. For source data of the FSC plots, see Supplementary Data 5.
Extended Data Fig. 4 Cryo-EM density and structure comparison.
a, Magnified views of representative regions of the cryo-EM density maps obtained by local reconstruction. Density is shown as grey mesh; polypeptide-chain backbone as ribbon; side-chain atoms as sticks (carbon, main color; nitrogen, blue; oxygen, red; sulfur, orange). b, Per-residue Cα distances after subunit-wise superposition of VP4, VP7 and VP6 subunits from the upright and reversed conformation structures.
Extended Data Fig. 5 Comparison of penetration protein conformations on rcTLPs and native TLPs.
Relative subparticle amounts, and corresponding cryo-EM reconstructions obtained from rcTLPs (top row) and native TLPs (bottom row). Local spike reconstructions of rcTLPs were obtained from two cryo-EM samples prepared from two independent recoating reactions. Local spike reconstructions of native TLPs were obtained from one cryo-EM sample.
Extended Data Fig. 6 Rearrangements of the VP8* and VP5* penetration proteins during transition from upright to reversed conformation on the virion surface.
a, Distinct domains of the VP8* and VP5* spike proteins are coloured separately to illustrate their conformational change during transition from upright (top row) to reversed (bottom row) conformation. Domains that were not observed in our cryo-EM maps because of flexible attachment are drawn schematically. VP8*, magenta; VP5*, red, orange and salmon. b, Formation of the trimeric coiled coil and extrusion of the foot domains. Top row, magnified views of the VP5* foot domain exit sites as observed in the intermediate conformation structure. A partially cut surface representation is shown. VP5*, red, orange and salmon; VP7, yellow. The last modelled residues of the VP5* β-barrel domains—482 (chain 1), 480 (chain 2) and 481 (chain 3)—are located on the outside. The connections to the first modelled VP5* foot domain residue—494 (chain 1), 494 (chain 2) and 498 (chain 3)—are indicated by dashed lines (fuzzy density in the cryo-EM map). Arrows indicate a suggested mechanism for foot-domain reversal, involving zipping-up of the trimeric coiled coil and unfolding and extrusion of foot-domain residues. Bottom row, proposed transition from the intermediate structure (left) to the reversed structure by zipping up of the coiled coil and unfolding and extrusion of the foot domains.
Extended Data Fig. 7 Molecular details of the VP5*–VP7 interfaces for upright and reversed conformations.
a, Upright conformation; b, reversed conformation. In each part of the figure, the left-hand image is a view from outside the TLP, with circles in black and blue corresponding to coloured outlines of the detailed, side-view panels on the right. VP5*, salmon; VP7, yellow. The following VP5* interface residues are conserved (Supplementary Data 1): N268, N376, R467, S469. The VP5* β-barrel N terminus (residues 248–250) is not strictly conserved, but the interaction is based on main-chain hydrogen bonds and can probably be maintained for different side chains as well. The following VP7 interface residues are conserved (Supplementary Data 2): S201, T210, L172–Y175.
Extended Data Fig. 8 Inducing penetration protein reversal at alkaline pH and VP8* association with TLPs.
a, Analysis of rotavirus particles without (lanes 1 and 2) and with (lanes 3 and 4) high pH-induced conformational change of VP8* or VP5* and without (lanes 1 and 3) and with (lanes 2 and 4) EDTA-induced uncoating. Pelleted fractions were analysed by SDS–PAGE and silver staining (left) and by western blotting with the VP8*-specific antibody HS127 (right) (see Methods). The experiment was repeated independently three times with similar results. For gel and western blot source data, see Supplementary Fig. 1. b, Relative VP4 subparticle amounts, and corresponding cryo-EM reconstructions obtained from wild-type rcTLPs. c, Relative VP4 subparticle amounts, and corresponding cryo-EM reconstructions obtained from rcTLPs containing VP4(S567C/A590C). Recoating reactions for all samples were carried out at the same time and with the same VP7 and DLP stock solutions. All cryo-EM samples were prepared in the same blotting session.
Extended Data Fig. 9 Cryo-ET analysis of RRVs entering BSC-1 cells.
a, Sections of the tomogram from which the reconstructions (icosahedral average of single virion subtomograms) in Fig. 4a were obtained. Left, virus with loose membrane contact indicated by an arrow. Right, virus with tight (close) membrane contact indicated by an arrow. Images were low-pass filtered and contrast enhanced for display. Scale bar, 100 nm. b, Tomographic slices of manually selected viruses (not including particle in a) from several tomograms with tight membrane contacts (yellow arrowheads). Images were low-pass filtered and contrast-enhanced for display. Those particles were selected to detect VP4 reversed conformations in additional viruses (other than particle in Fig. 4a). c, Sub-subtomogram classification of VP4 positions extracted from the RRV particles shown in b yielded a class with the upright conformation (left) and a class with the reversed conformation (right). VP4 positions were extracted from 26 selected particles chosen to have an easily identified region of close membrane contact. The particle shown in Fig. 4a was excluded from this selection. Note that the reconstructions shown in Fig. 4a are from single particles with imposed icosahedral symmetry, whereas we show here classified averaged individual volumes extracted at VP4 positions.
Supplementary information
Supplementary Information
This file contains the Supplementary Discussion.
Supplementary Figure
Supplementary Figure 1: This file shows the Western blot and gel source data for Fig. 3d, Extended Data Fig. 1b, and Extended Data Fig. 8a.
Supplementary Data
Supplementary Data 1: Rotavirus VP4 multiple sequence alignment from GenBank accession identifiers: RRV, P12473; SA11, P12976; WA, P11193; S2, AQT31697; DS1, P11196; ST3, P11200; AU-1, P39033; 116E, Q09113; 69M, P26451; NCDV, P17465; UK, P12474; 993/83, Q08010; Gottfried, P23045; OSU, P11114; CH3, Q86184; L338, Q98636; EW, Q83450. Modeled residues are shown as solid bars.
Supplementary Data
Supplementary Data 2: Rotavirus VP7 multiple sequence alignment from GenBank accession identifiers: RRV, P12473; SA11, P12976; WA, P11193; S2, AQT31697; DS1, P11196; ST3, P11200; AU-1, P39033; 116E, Q09113; 69M, P26451; NCDV, P17465; UK, P12474; 993/83, Q08010; Gottfried, P23045; OSU, P11114; CH3, Q86184; L338, Q98636; EW, Q83450. Modeled residues are shown as solid bars.
Supplementary Data
Supplementary Data 3: Rotavirus VP6 multiple sequence alignment from GenBank accession identifiers: RRV, P12473; SA11, P12976; WA, P11193; S2, AQT31697; DS1, P11196; ST3, P11200; AU-1, P39033; 116E, Q09113; 69M, P26451; NCDV, P17465; UK, P12474; 993/83, Q08010; Gottfried, P23045; OSU, P11114; CH3, Q86184; L338, Q98636; EW, Q83450. Modeled residues are shown as solid bars.
Supplementary Data
Supplementary Data 4: This file contains the source data for the bar plot shown in Fig. 3a.
Supplementary Data
Supplementary Data 5: This file contains the source data for the Fourier shell correlation (FSC) plots shown in Extended Data Fig. 3.
Rights and permissions
About this article
Cite this article
Herrmann, T., Torres, R., Salgado, E.N. et al. Functional refolding of the penetration protein on a non-enveloped virus. Nature 590, 666–670 (2021). https://doi.org/10.1038/s41586-020-03124-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-020-03124-4
- Springer Nature Limited
This article is cited by
-
Cryo-EM structures of Banna virus in multiple states reveal stepwise detachment of viral spikes
Nature Communications (2024)
-
mRNA-based VP8* nanoparticle vaccines against rotavirus are highly immunogenic in rodents
npj Vaccines (2023)
-
Visualizing molecular interactions that determine assembly of a bullet-shaped vesicular stomatitis virus particle
Nature Communications (2022)
-
Novel fold of rotavirus glycan-binding domain predicted by AlphaFold2 and determined by X-ray crystallography
Communications Biology (2022)
-
Multiple conformations of trimeric spikes visualized on a non-enveloped virus
Nature Communications (2022)