Abstract
The human liver is an essential multifunctional organ. The incidence of liver diseases is rising and there are limited treatment options. However, the cellular composition of the liver remains poorly understood. Here we performed single-cell RNA sequencing of about 10,000 cells from normal liver tissue from nine human donors to construct a human liver cell atlas. Our analysis identified previously unknown subtypes of endothelial cells, Kupffer cells, and hepatocytes, with transcriptome-wide zonation of some of these populations. We show that the EPCAM+ population is heterogeneous, comprising hepatocyte-biased and cholangiocyte populations as well as a TROP2int progenitor population with strong potential to form bipotent liver organoids. As a proof-of-principle, we used our atlas to unravel the phenotypic changes that occur in hepatocellular carcinoma cells and in human hepatocytes and liver endothelial cells engrafted into a mouse liver. Our human liver cell atlas provides a powerful resource to enable the discovery of previously unknown cell types in normal and diseased livers.
Similar content being viewed by others
Main
The liver serves as a central metabolic coordinator with a wide array of essential functions, including the regulation of glucose and lipid metabolism, protein synthesis, and bile synthesis. Furthermore, the liver is a visceral organ that is capable of remarkable natural regeneration after tissue loss1. However, the prevalence of liver diseases and mortality associated with them have risen markedly within recent decades2. The liver cellular landscape has barely been explored at single-cell resolution, which limits our molecular understanding of liver function and disease biology. The recent emergence of sensitive single-cell RNA sequencing (scRNA-seq) methods3 allows us to investigate cell types in healthy and diseased tissue.
To characterize the human liver at single-cell resolution, we developed a robust pipeline for scRNA-seq of cryopreserved and freshly isolated samples of patient-derived human liver and assembled an atlas consisting of 10,372 cells from nine donors. We performed in-depth analysis of all liver cell types with a focus on epithelial liver cell progenitors.
scRNA-seq of the human liver
We used mCEL-Seq24 for scRNA-seq of non-diseased liver tissue from nine patients who underwent liver resections for colorectal cancer metastasis or cholangiocarcinoma without history of chronic liver disease (Fig. 1a, see Methods). We sorted and sequenced viable cells both in an unbiased fashion and by enriching specific cell populations on the basis of cell surface marker expression (Extended Data Fig. 1, see Methods). Because fresh liver tissue material is scarce and difficult to preserve, and biobanks with cryopreserved liver samples represent rich resources, we generated scRNA-seq data from cryopreserved cells as well as single-cell suspensions from freshly prepared liver samples (see Methods). We then used RaceID3 for the identification of cell types4,5 (see Methods).
a, Outline of the protocol used for scRNA-seq of human liver cells. Samples from liver resections were digested to prepare single-cell suspensions. Cells were sorted into 384-well plates and processed according to the mCEL-Seq2 protocol. b, t-SNE map of single-cell transcriptomes from normal liver tissue from nine donors highlighting the main liver cell compartments. ‘Other’ denotes various small populations comprising 22 red blood cells and 46 cells that cannot be unambiguously annotated. ‘Other endothelial cells’ cannot be unambiguously classified as LSECs or MaVECs. c, t-SNE map of single-cell transcriptomes highlighting RaceID3 clusters, which reveals subtype heterogeneity in all major cell populations of the human liver. Numbers denote clusters. d, Heat map showing the expression of established marker genes for each cell compartment. Colour bars indicate patient, major cell type, and RaceID3 cluster. Scale bar, log2-transformed normalized expression. b, c, n = 10,372 cells.
Cells from different patients, isolated from freshly prepared or cryopreserved single-cell suspensions, co-clustered (Extended Data Fig. 1). Furthermore, fresh and cryopreserved cells from the same patient did not have markedly different gene signatures (Extended Data Fig. 1e–h). However, there were compositional differences both between fresh and cryopreserved samples derived from the same patient and among different fresh (or cryopreserved) samples. We attribute these differences to variability in cell viability and cell type composition across samples.
As scRNA-seq of randomly sampled populations yielded almost exclusively hepatocytes and immune cells (Extended Data Fig. 1i), we applied additional sorting strategies to enrich for endothelial cells (Extended Data Fig. 1a–c) and EPCAM+ cells (see below).
Our atlas comprises all the main liver cell types defined by the expression of marker genes, including hepatocytes, EPCAM+ bile duct cells (cholangiocytes), CLEC4G+ liver sinusoidal endothelial cells (LSECs), CD34+PECAMhigh macrovascular endothelial cells (MaVECs), hepatic stellate cells and myofibroblasts, Kupffer cells, and immune cells (Fig. 1b–d, Supplementary Table 1). To facilitate interactive exploration of our human liver cell atlas, we created a web interface: http://human-liver-cell-atlas.ie-freiburg.mpg.de/.
Zonation of human liver cell types
Hepatocytes are spatially heterogeneous and zonated along the portal–central axis of the liver lobule6,7,8. According to metabolic sub-specialization, the liver lobule has been divided into the periportal zone surrounding the portal triad (portal vein, hepatic artery and bile duct), the central zone nearest to the central vein, and the remaining mid zone6,7,8. Whereas previous observations have suggested that non-parenchymal cells such as LSECs and Kupffer cells have specialized subtypes6, it has been hard to demonstrate heterogeneity of these cell types, and most studies have been carried out in rodents.
We were able to directly compare the signatures of MaVECs and LSECs, and identified several previously unknown subpopulations (Extended Data Fig. 2, Supplementary Note 1).
scRNA-seq has been highly informative on hepatocyte zonation in mouse9, and the first single-cell analysis of human hepatocyte and endothelial cell zonation at limited resolution was done recently10. To infer continuous transcriptome-wide zonation, we reasoned that the major axis of variability for a cell type could reflect gene expression changes associated with zonation. Hence, we ordered LSECs and hepatocytes by diffusion pseudo-time (dpt)11, here interpreted as pseudo-space, along this axis and applied self-organizing maps (SOMs) to infer co-expression modules (Fig. 2, see Methods).
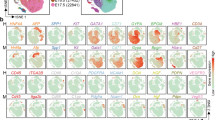
a, Diffusion maps (left) and SOMs (middle) of single-cell transcriptome-derived zonation profiles for hepatocytes (n = 2,534 cells). DPT indicates diffusion-pseudotime and is interpreted here as a spatial zonation coordinate. Right, zonation profiles of GLUL (central), APOE (midzonal), CYP1A2 and CYP2E1 (central/midzonal) and ALB and PCK1 (periportal); bottom left, immunostaining for GLUL, APOE, CYP1A2, and CYP2E1 from the Human Protein Atlas31. See Extended Data Fig. 3g for additional images. b, Diffusion maps (left) and SOMs (middle) of single-cell transcriptome-derived zonation profiles for endothelial cells (n = 1,361 cells). Right, zonation profiles of BTNL9 and ANPEP (periportal), LYVE1 and FCN3 (midzonal), and ICAM1, FCN3 and ENG (central); bottom left, immunostaining for ICAM1 and ANPEP from the Human Protein Atlas. P, portal tracts; C, central. Colour bars at the bottom of SOMs show RaceID3 cluster as in Fig. 1. The y axis of the zonation profiles indicates normalized expression.
We first validated our strategy by recovering the previously characterized zonation of mouse hepatocytes9 (Extended Data Fig. 3a–d). For our human hepatocytes, this approach recovered zonated expression patterns of landmark genes: for example, ALB and PCK1 (periportal module 1), CYP1A2 and CYP2E1 (central/midzonal modules 34 and 24, respectively), and GLUL (central module 33)7,9 (Fig. 2a, Extended Data Fig. 3e–g, Supplementary Tables 2, 3). In total, 1,384 out of 3,395 expressed genes (41%) included in the hepatocyte analysis exhibited significant zonation (Benjamini–Hochberg corrected ANOVA, P < 0.01). Pathway enrichment analysis revealed that periportal hepatocyte modules are enriched in genes involved in biological oxidation, consistent with an oxygen gradient that peaks in the periportal zone6,7,8, and in the glycogen synthesis pathway (Extended Data Fig. 3h). In accordance with its zonation in mouse hepatocytes, the urea cycle enzyme CPS1 peaks in periportal hepatocytes (Extended Data Fig. 3g). Midzonal hepatocyte modules are enriched in, for example, metabolism of xenobiotics by cytochrome P450. Immunostainings for selected genes validate the predicted zonation at the protein level (Fig. 2a).
LYVE1 and CD14 have been identified as markers that distinguish midzonal and central LSECs from periportal LSECs12. Analysis of LSEC zonation showed that 806 out of 1,198 expressed genes (67%) exhibited significant zonation (Benjamini–Hochberg corrected ANOVA, P < 0.01) (Fig. 2b, Extended Data Fig. 3i, Supplementary Tables 4, 5). Central and midzonal endothelial cells (modules 1 and 3) exhibited peaked expression of LYVE1 and FCN3, which encodes a ficolin protein that can switch on the lectin pathway of complement activation. Notably, pathway enrichment analysis of the central and midzonal endothelial modules recovered pathways, such as binding and uptake of ligands by scavenger receptors, that are shared with midzonal hepatocytes (Extended Data Fig. 3j). Together with a more detailed gene expression analysis (Supplementary Note 2) this observation suggests that genes and functions are co-zonated across hepatocytes and endothelial cells.
Finally, a comparison between mouse9,13 and human cells revealed only limited evolutionary conservation of gene expression zonation (Supplementary Note 3, Extended Data Fig. 3k, l, Supplementary Tables 6, 7), reflecting widespread evolutionary changes.
Human liver immune cell populations
A detailed analysis of the CD163+VSIG4+ Kupffer cell compartment revealed subpopulations with distinct gene expression signatures (Supplementary Note 4, Extended Data Fig. 4), in agreement with a recent study10. Moreover, we detected shared gene expression and pathways between Kupffer cell subsets and endothelial cells (Supplementary Note 4, Extended Data Fig. 4), providing further evidence that different cell types show functional co-operation.
We identified an MS4A1+CD37+ subset of B cells, which corresponds to circulating B cells with upregulated MHC class II components, and a liver-resident MZB1+ subset of B cells that expresses DERL3, SSR4 and IGHG4 (Extended Data Fig. 5).
Finally, we recovered a population of CD56+ (also known as NCAM1+) natural killer (NK) cells (cluster 5), as well as CD56– (cluster 3) and CD56+ (cluster 1) CD8A+ NKT cells, which expressed different combinations of chemokine ligands, granzymes, and killer cell lectin-like receptor genes (Extended Data Fig. 6). In clusters 12 and 18, a number of heat-shock genes are upregulated. These observations demonstrate an unexpected variety of immune cell subtypes in the human liver.
Putative bipotent epithelial progenitors
Liver regeneration after tissue damage involves the replication of several types of liver cells, including hepatocytes and cholangiocytes. Furthermore, different types of liver damage lead to specific mechanisms of liver regeneration14,15. However, the existence of a population of naive adult stem cells in the human liver and its contribution to turnover and regeneration remains controversial. Rare EPCAM+ cells have been termed hepatic stem cells16; these can form dense round colonies when cultured and are bipotent progenitors of hepatoblasts, which differentiate into cholangiocytes or hepatocytes both in vitro and in vivo16,17.
To search for genuine liver progenitor cells, we sorted and sequenced single EPCAM+ cells from adult human livers. We identified biliary and potential liver progenitor cell surface marker genes that correlated with EPCAM or TROP1 expression; these included TACSTD2 (also known as TROP2), FGFR2, TM4SF4 and CLDN1. Immunohistochemistry confirmed the expression of predicted markers such as ANXA4 and the transcriptional co-activator WWTR1 (Extended Data Fig. 7a).
A focused analysis revealed that the EPCAM+ compartment is transcriptionally heterogeneous and consists of an ASGR1+ hepatocyte-biased population, KRT19highCFTRhighALBlow cholangiocyte populations, and a remaining population of putative naive progenitor cells (Fig. 3a, Extended Data Fig. 7b, c). The EPCAM+ population exhibits only stochastic expression of the proliferation markers MKI67 and PCNA and is negative for the hepatoblast marker AFP (Extended Data Fig. 7d). Hence, the transcriptional heterogeneity of this population is unlikely to arise as a result of proliferation, and the observed subtypes reside in the normal human liver.
a, Expression t-SNE maps of ASGR1 and CFTR for the EPCAM+ compartment only. The colour bar indicates log2 normalized expression. b, StemID218 analysis of the EPCAM+ compartment. Shown are links with StemID2 P < 0.05. Node colour denotes transcriptome entropy. c, FateID analysis of the EPCAM+ compartment highlights populations that are preferentially biased towards hepatocyte progenitors and cholangiocytes, respectively, and reveals similar bias towards both lineages in the central population (clusters 1, 2, 5, 6 and 7). Colour bar indicates lineage probability. d, Expression heat map of selected hepatocyte marker genes (HP, ASGR1), mature cholangiocyte genes (KRT19, CFTR, CXCL8, MMP7), additional progenitor markers (grey), and all genes upregulated in the central population (clusters 1, 2, 5, 6 and 7) within the EPCAM+ compartment (Benjamini–Hochberg corrected P < 0.01; fold change >1.33; see Methods). Four compartments are indicated, resolving the predicted fate bias (Extended Data Fig. 8). e, Correlation of nearest-neighbour-imputed (k = 5) expression (using RaceID3) of TACSTD2 and hepatocyte bias predicted by FateID. Red line, loess regression. R, Spearman’s rank correlation. a–e, n = 1,087 cells. f, Immunostaining for TROP2 from the Human Protein Atlas (n = 3 biologically independent samples). Arrow, bile duct; arrowhead, bile ductule. g, Immunofluorescence labelling of EPCAM and KRT19. EPCAM+KRT19low/– (solid arrow) and EPCAM+KRT19+ (broken arrow) cells are indicated. Nuclei are stained with DAPI. Images are maximum z-stack projections of 6 μm. Scale bar, 10 μm. (n = 3 independent experiments).
To explore the relatedness of these subpopulations, we reanalysed the EPCAM+ population with RaceID3 and used StemID2 for lineage reconstruction4,18 (Fig. 3b, see Methods). This analysis showed that the population in the centre of the t-distributed stochastic neighbour embedding (t-SNE) map (clusters 1, 2, 5, 6, 7) bifurcates into hepatocyte progenitors and cholangiocytes. To provide further evidence for continuous differentiation trajectories connecting naive EPCAM+ progenitors to cholangiocytes and mature hepatocytes, we performed StemID2 and diffusion map analyses on the combined population of mature hepatocytes and EPCAM+ cells (Extended Data Fig. 8a–c).
To better understand the emergence of fate bias towards the two lineages, we used FateID to infer lineage probabilities in each cell4. Consistently, FateID inferred similar probabilities that the central population would differentiate towards hepatocytes and cholangiocytes (Fig. 3c). The fate bias predictions are supported by a differential gene expression analysis revealing upregulation of common genes that encode several signalling pathway components (HES1, SFRP5, FGFR2, FGFR3) in the central population (Fig. 3d), and gradual upregulation of distinct gene sets towards the hepatocyte-biased and cholangiocyte populations (Extended Data Fig. 8e). The expression of TROP2 was negatively correlated with hepatocyte fate bias, exhibiting a gradient that ranged from high expression in mature cholangiocytes to very low expression in the hepatocyte-biased population (Fig. 3e, Extended Data Fig. 7c). Immunostaining for TROP2 in normal human liver tissue showed specific expression in cells of the bile ducts and bile ductules (Fig. 3f). Notably, TROP2 expression has been found in amplifying oval cells in injured mouse livers19.
The central TROP2int population is in itself heterogeneous and contains a MUC6high population (cluster 7) (Extended Data Fig. 7c). MUC6 is highly expressed by pancreatic progenitors and multi-potent bile duct tree stem cells20, which have been proposed to be the origin of the EPCAM+ hepatic stem cells. The TROP2high cholangiocyte clusters comprise a CXCL8+ population (cluster 8) and an MMP7+ population (clusters 4 and 13) (Extended Data Figs. 7c, 8e, f), whereas TROP2low clusters show upregulation of hepatocyte markers such as ALB, HP, HNF4A and ASGR1 (Fig. 3d, Extended Data Figs. 7c, 8e, f).
The central TROP2int population that was stratified as bipotent on the basis of FateID-predicted bias expresses genes that encode early developmental transcription factors such as HES1, which is essential for tubular bile duct formation21, and PROX1, an early specification marker for the developing liver in the mammalian foregut endoderm that is required for hepatocyte proliferation and migration during development22 (Fig. 3d). Furthermore, this population showed lower expression of hepatocyte genes such as HNF4A, HP and ALB and of cholangiocyte genes such as KRT19 and CFTR compared to the hepatocyte-biased and mature cholangiocyte populations, respectively (Fig. 3d, Extended Data Figs. 7c, 8f). We speculate that we enriched for the TROP2int KRT19low/– immature population during cell isolation, as mature bile duct cells require a harsher digestion for their isolation, which can negatively affect other liver cell types. Thus, the actual fraction of KRT19high cells in the tissue is presumably higher. We validated the existence of EPCAM+KRT19low/– cells in addition to EPCAM+KRT19high/+ cells in situ by immunofluorescence (Fig. 3g, Extended Data Fig. 7e).
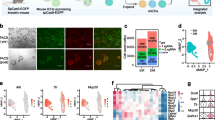
Consistent with our scRNA-seq data, flow cytometry profiles of EPCAM and TROP2 displayed a gradient of TROP2 expression in EPCAM+ cells, and EPCAM expression correlated with TROP2 expression (Fig. 4a). Moreover, forward and side-scatter profiles of EPCAM+ cells indicated that the compartment is heterogeneous and consists of populations with different sizes and morphologies (Fig. 4a). On the basis of the distribution of TROP2 expression, we compartmentalized EPCAM+ cells into three compartments: TROP2low/–, TROP2int, and TROP2high (Fig. 4a). To confirm that the TROP2int population harbours the progenitor population, we attempted to culture bipotent organoids23 from each compartment. In agreement with our prediction, TROP2int cells exhibited the highest organoid-forming capacity, whereas TROP2low/– cells did not form organoids, and TROP2high cells gave rise to much smaller organoids at a strongly reduced frequency compared to TROP2int cells (Fig. 4b). Single-cell culture of TROP2int cells demonstrated the organoid-forming capacity of individual cells from this gate, providing evidence for bipotency at the clonal level (Fig. 4c). As expected, scRNA-seq of the input populations for organoid culture from each compartment showed a marked enrichment of the respective compartment in the original EPCAM+ data (Fig. 4d, e, Extended Data Fig. 8g, h). Notably, flow cytometry profiles of EPCAM and TROP2 for organoid cells grown from the TROP2int compartment recovered TROP2low/–, TROP2int and TROP2high populations in the organoids (Fig. 4f).
a, Fluorescence-activated cell sorting (FACS) plots for EPCAM+ cells showing expression (left) and forward and side scatter (right) of EPCAM and TROP2 (n = 6 independent experiments). The gates for the three compartments are shown. b, Top, organoid culturing of cells from the TROP2low/–, TROP2int and TROP2high compartments (n = 3 independent experiments). Bottom, number of organoids (left), organoid frequency relative to the TROP2int compartments (centre), and size of organoids (right); n = 3 patients, 100 seeded cells each. Scale bar, 400 μm. c, Organoid frequency in single-cell cultures of TROP2int cells (n = 3 independent experiments, 96 cells each). Owing to the small number of cells we were unable to purify single cells from the other gates. b, c, Mean ± s.d. d, Symbol t-SNE map showing organoid cells, original EPCAM+ data (from Fig. 3) and cells sorted from the gates in a. e, Expression t-SNE maps for SERPINA1, KRT19 and CXCL8. Colour bar indicates log2 normalized expression. f, FACS plot of expression of EPCAM and TROP2 in organoid cells grown from the TROP2int compartment, 17 days after initial culture (n = 3 independent experiments). g, Expression heat map of genes that were differentially expressed between patient and organoid cells (Benjamini–Hochberg corrected P < 0.05 (see Methods), mean expression >0.7, log2 fold change >2). d, e, g, n = 2,870 cells.
To elucidate the cell type composition of the organoids in depth, we performed scRNA-seq. Co-analysis of organoid cells and EPCAM+ cells sequenced directly from patients demonstrated marked transcriptome differences (Fig. 4e). Although EPCAM and CD24 were expressed in cells from both organoids and patients, organoid cells showed lower expression of various genes such as AQP1 and the WNT signalling modulator SFRP5, and higher expression of others, such as the proliferation marker MKI67+, reflected by differential enrichment of the corresponding pathways (Fig. 4g, Extended Data Fig. 8i–k). We observed several subpopulations within the organoids, including a non-dividing hepatocyte-biased SERPINA1high population and a non-dividing KRT19high cholangiocyte-biased population, consistent with the signature of the EPCAM+ cells recovered from the patients (Fig. 4e). This further supports the claim that the TROP2int compartment harbours a bipotent progenitor population, which can give rise to hepatocyte and cholangiocyte populations.
In contrast to patient cells, organoid cells showed strong downregulation of ALB but expressed AGR2 and other mucin family genes such as MUC5AC and MUC5B, which are normally expressed, for example, in intestinal cells and gastrointestinal cancers24,25 (Fig. 4g, Extended Data Fig. 8j). These observations reflect that organoid cells express genes that are expressed in other systems, acquire a more proliferative state, and appear to upregulate stem cell-related pathways such as WNT signalling.
In light of these functional validation experiments, the observed gene signature of TROP2int cells, and the in situ location of these cells, our data strongly suggest that the putative liver progenitor population can be defined as a subpopulation of bile duct cells.
Perturbed cell states in liver cancer
Hepatocellular carcinoma (HCC) is the most common type of primary liver cancer26. To demonstrate the value of our atlas as a reference for comparisons with diseased liver cells, we sequenced CD45+ and CD45– cells from HCC tissue from three patients (Extended Data Fig. 9a, b, see Methods).
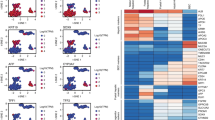
We recovered several cell types from the tumours, including cancer cells, endothelial cells, Kupffer cells, NKT cells and NK cells (Fig. 5a, Extended Data Fig. 9c) and compared them to the normal liver cell atlas. Differential gene expression analysis and immunohistochemistry revealed that cancer cells lose the expression of cytochrome P450 genes such as CYP2E1 and CYP2C8 and the periportally zonated gene CPS1 (Fig. 5b, Extended Data Fig. 9d, e) as well as the metabolic signature of normal hepatocytes (Fig. 5c). They show increased expression of AKR1B10, a known biomarker of HCC with potential involvement in hepatocellular carcinogenesis27 (Extended Data Fig. 9d). Moreover, immunohistochemistry confirmed that IL32, a pro-inflammatory TNFα-inducing cytokine, is highly upregulated in cancer cells (Fig. 5b). Overall, cancer cells show upregulation of WNT and Hedgehog signalling pathways, highlighting similarities between EPCAM+ normal liver progenitors and the observed cancer cell population (Fig. 5c).
a, Symbol t-SNE map highlighting normal liver cells and cells from HCC. n = 11,654 cells, n = 3 patients with HCC. b, Immunostaining for IL32 and CYP2E1 in normal liver and HCC tissue. c, GSEA for genes that were differentially expressed between cancer cells from HCC and normal hepatocytes (n = 15,442 genes). d, GSEA for genes that were differentially expressed between normal endothelial cells and endothelial cells from HCC (n = 15,442 genes). c, d, Benjamini–Hochberg corrected P < 0.01; NES, normalized enrichment score; see Methods. e, Immunostaining for CLEC4G and PECAM1 in normal liver tissue and HCC tissue. All staining images are from the Human Protein Atlas31.
Endothelial cells from tumours show upregulation of, for example, extracellular matrix organization genes such as COL4A2 and SPARC (Fig. 5d, Extended Data Fig. 9f). Strikingly, they do not express LSEC marker genes such as CLEC4G but do express MaVEC marker genes such as PECAM1, AQP1 and CD34 (Fig. 5e, Extended Data Fig. 9f, g). Moreover, HCC LSECs show increased expression of PLVAP, which makes them less permeable and could potentially restrict the access of lymphocytes and soluble antigens28 to the tumour (Supplementary Note 5, Extended Data Fig. 9f, g).
We conclude that the comparison of scRNA-seq data between the cell populations of HCC and the liver cell atlas allows the inference of perturbed gene expression signatures, biomarkers and modulated functions across cell types.
A human liver chimaeric mouse model
Mice harbouring patient-derived xenografted liver cells are a powerful tool for studying human liver cells and diseases in vivo29. To correctly interpret such experiments, it is crucial to understand the differences between cells taken directly from the human liver and human cells that have been transplanted into the mouse liver.
To address this issue, we transplanted human liver cells from patient-derived hepatocyte and non-parenchymal cell fractions into FRG-NOD (Fah−/−Rag2−/−Il2rg−/− non-obese diabetic) mice30 (HMouse); after engraftment, we sorted single human cells in an unbiased fashion and on the basis of hepatocyte and endothelial cell markers for scRNA-seq (Fig. 6a, Extended Data Fig. 10a). We then compared engrafted cells to our reference atlas and observed that we had successfully transplanted both human hepatocytes and endothelial cells (Fig. 6b, Extended Data Fig. 10b, c), which had maintained their fundamental gene signatures, such as expression of ALB or PCK1 and CLEC4G, PECAM1 or CD34, respectively (Extended Data Fig. 10b–f). Nevertheless, many genes were differentially expressed in engrafted cells compared to non-engrafted human liver cells; for example, AKR1B10, which was also expressed by cancer cells from HCC, was expressed in engrafted cells but not non-engrafted cells (Fig. 6c, Extended Data Fig. 10g). Gene set enrichment analysis (GSEA) of differentially expressed genes revealed that HMouse hepatocytes and endothelial cells showed downregulation of pathways such as haemostasis, and upregulation of WNT and Hedgehog signalling as well as cell cycle genes (Fig. 6d), akin to what we observed in HCC cells and cells from liver organoids.
a, Outline of the transplantation of human liver cells (hepatocytes and non-parenchymal cells) into the FRG-NOD mouse and the two strategies for sorting engrafted human cells from the mouse liver. b, Symbol t-SNE map highlighting normal liver cells and cells from the humanized mouse model. The main engrafted cell types (hepatocytes and endothelial cells) are circled. c, Expression t-SNE maps of AKR1B10 and CXCL1/CXCL2. Colour bar indicates log2 normalized expression. n = 10,683 cells. d, GSEA of genes that were differentially expressed between hepatocytes and endothelial cells from humanized mouse (HMouse) and patients (Human). n = 13,614 genes; Benjamini–Hochberg corrected P < 0.01; see Methods.
Discussion
We have established a human liver cell atlas, revealing heterogeneity within major liver cell populations and the existence of an epithelial progenitor in the adult human liver.
Our atlas reveals transcriptome-wide zonation of hepatocytes and endothelial cells, and suggests that different liver cell types may cooperate to carry out essential functions. Although we could validate predicted zonation profiles with antibody staining, it will be essential to perform more large scale in situ gene expression analysis.
The EPCAM+TROP2int population is a strong candidate for potential involvement in homeostatic turnover, liver regeneration, disease pathogenesis and tumour formation. Although our in silico analysis and in vitro organoid culture experiments provide evidence that this population is bipotent, its lineage potential remains to be demonstrated in vivo.
As demonstrated by our HCC analysis, the atlas provides a key reference for the investigation of liver diseases and will contribute to the development of urgently needed human liver models, including organoids and humanized liver chimaeric mouse models.
Methods
Human liver samples
Human liver tissue samples were obtained from patients who had undergone liver resections between 2014 and 2018 at the Center for Digestive and Liver Disease (Pôle Hépato-digestif) at the Strasbourg University Hospitals, University of Strasbourg, France. For the human liver cell atlas, samples were acquired from patients without chronic liver disease (defined as liver damage lasting over a period of at least six months), genetic haemochromatosis with homozygote C282Y mutation, active alcohol consumption (>20 g/d in women and >30 g/d in men), active infectious disease, pregnancy or any contraindication for liver resection. All patients provided written informed consent. The protocols followed the ethical principles of the declaration of Helsinki and were approved by the local Ethics Committee of the University of Strasbourg Hospitals and by the French Ministry of Education and Research (CPP 10-17, Ministère de l'Education Nationale, de l’Enseignement Supérieur et de la Recherche; approval number DC-2016-2616). Data protection was performed according to EU legislation regarding privacy and confidentiality during personal data collection and processing (Directive 95/46/EC of the European Parliament and of the Council of the 24 October 1995). Samples (BP1) and tissue blocks were obtained from Biopredic International.
Tissue dissociation and preparation of single-cell suspensions
Human liver specimens obtained from resections were perfused for 15 min with calcium-free 4-(2-hydroxyethyl)-1-piperazine ethanesulfonic acid buffer containing 0.5 mM ethylene glycol tetraacetic acid (Fluka) followed by perfusion with 4-(2-hydroxyethyl)-1-piperazine ethanesulfonic acid containing 0.5 mg/ml collagenase (Sigma-Aldrich) and 0.075% CaCl2 at 37 °C for 15 min as previously described32. Then the cells were washed with phosphate-buffered saline (PBS) and nonviable cells were removed by Percoll (Sigma-Aldrich) gradient centrifugation. Part of the isolated cells was further separated into primary human hepatocytes (PHHs) and non-parenchymal cells (NPCs) by an additional centrifugation step at 50g for 5 min at 4 °C. The isolated cells were frozen in liquid nitrogen using the CryoStor CS10 solution (Sigma-Aldrich). Human HCC tissues were dissociated using the gentleMACS Dissociator (Miltenyi Biotec) according to the manufacturer’s protocol.
Transplantation of human cells into Fah −/− /Rag2 −/− /Il2rg −/− mice
Fah−/−/Rag2−/−/Il2rg−/− non-obese diabetic (FRG-NOD) breeding mice were kept at the Inserm Unit 1110 SPF animal facility and maintained with 16 mg/l of 2-(2-nitro-4-trifluoro-methyl-benzoyl)-1,3 cyclohexanedione (NTBC; Swedish Orphan Biovitrum) in drinking water. Six-week-old male and female mice were intravenously injected with 1.5 × 109 plaque-forming units (pfu) of an adenoviral vector encoding the secreted form of the human urokinase-like plasminogen activator (Ad-uPA)33. Forty-eight hours later, 106 PHHs and 2 × 105 NPCs from the same liver donor, isolated as previously described, were injected intra-splenically via a 27-gauge needle. For the procedure, mice were kept under gaseous isoflurane anaesthesia and received a subcutaneous injection of buprenorphine (0.1 mg/kg). After transplantation, the NTBC was gradually decreased and completely withdrawn in 7 days. The success of the transplantation was evaluated 2 months after the procedure by dosing human albumin in mouse serum as previously described34. This procedure was approved by the local ethics committee and authorized by the French ministry of higher education and research (authorization number #4485-20l603lll5352125 v3). All procedures are consistent with the guidelines set by the Panel on Euthanasia (AVMA) and the NIH Guide for the Care and Use of Laboratory Animals as well as the Declaration of Helsinki in its latest version, and the Convention of the Council of Europe on Human Rights and Biomedicine. The animal research was performed within the regulations and conventions protecting animals used for research purposes (Directive 86/609/EEC), as well as with European and national laws regarding work with genetically modified organs. The animal facility at the University of Strasbourg, Inserm U1110 has been approved by the regional government (Préfecture) and granted authorization number E67-482-7, 2017/08/24.
Mouse liver cell isolation
The anaesthetized animal was restrained and the skin sprayed with 70% ethanol. The liver and other inner organs were revealed by cutting through the skin and peritoneum. A 24G needle was carefully inserted into the inferior vena cava and secured with a clamp, and chelating solution (0.05 M HEPES pH 7.2, 10 mM EGTA in HBSS without CaCl2 and MgCl2) was run at a low speed (1.5–2 ml/min). The portal vein was then cut and perfusion speed was increased to a flow rate of 7 ml/min. After that, the diaphragm was cut and the anterior vena cava clamped. The chelating perfusion was run for 7 min and then switched to collagenase solution (0.05 M HEPES pH 7.2, 4.7 mM CaCl2, 20 µg/ml Liberase, Sigma LIBTM-RO) at a flow rate of 7 ml/min for 7 min. The liver was then removed and passed through a 70-µm cell strainer with 10 ml ice-cold PBS without CaCl2 and MgCl2. The resulting single-cell suspension was centrifuged at 300g for 5 min at 4 °C and washed twice with ice-cold PBS.
FACS
Liver cells were sorted from mixed, hepatocyte, and non-parenchymal cell fractions on an Aria Fusion I using a 100-μm nozzle. Cells from the HCC samples were not fractionated and were sorted directly after tissue digestion. Zombie Green (Biolegend) was used as a viability dye. Cells were stained with human-specific antibodies against CD45 (Biolegend, cat. no. 304023), PECAM1 (Biolegend, cat. no. 303111), CD34 (Biolegend, cat. no. 343609), CLEC4G (R&D systems, cat. no. FAB2947A), ASGR1 (BD Biosciences, cat. no. 563655), EPCAM (R&D systems, cat. no. FAB960R), and TROP2 (Biolegend, cat. no. 363803). Organoids were stained with antibodies against EPCAM and TROP2. For the humanized mouse samples, cells were stained either with antibodies against ASGR1 and PECAM1 or with human HLA-ABC (BD Biosciences, cat. no. 740407) and mouse H2-Kb (BD Biosciences, cat. no. 553570). Viable cells were sorted in an unbiased fashion or from specific populations based on the expression of markers into the wells of 384-well plates containing lysis buffer.
Single-cell RNA amplification and library preparation
Single-cell RNA sequencing was performed according to the mCEL-Seq2 protocol4,35. Viable liver cells were sorted into 384-well plates containing 240 nl primer mix and 1.2 μl PCR encapsulation barrier, Vapour-Lock (QIAGEN) or mineral oil (Sigma-Aldrich). Sorted plates were centrifuged at 2,200g for a few minutes at 4 °C, snap-frozen in liquid nitrogen and stored at −80 °C until they were processed. We used 160 nl reverse transcription reaction mix and 2.2 μl second-strand reaction mix to convert RNA into cDNA. cDNA from 96 cells was pooled together before clean up and in vitro transcription, generating four libraries from one 384-well plate. We used 0.8 μl AMPure/RNAClean XP beads (Beckman Coulter) per 1 μl sample during all purification steps including library cleanup. Other steps were performed as described in the protocol4,35. Libraries were sequenced on an Illumina HiSeq 2500 and 3000 sequencing system (paired-end multiplexing run, high output mode) at a depth of ~150,000–200,000 reads per cell.
Quantification of transcript abundance
Paired-end reads were aligned to the transcriptome using bwa (version 0.6.2-r126) with default parameters36. The transcriptome contained all gene models based on the human whole genome ENCODE V24 release. All isoforms of the same gene were merged to a single gene locus. Subsequently, gene loci with >75% sequence overlap were merged. The right mate of each read pair was mapped to the ensemble of all gene loci and to the set of 92 ERCC spike-ins in the sense direction. Reads mapping to multiple loci were discarded. The left read contains the barcode information: the first six bases corresponded to the unique molecular identifier (UMI) followed by six bases representing the cell-specific barcode. The remainder of the left read contains a polyT stretch. The left read was not used for quantification. For each cell barcode, the number of UMIs per transcript was counted and aggregated across all transcripts derived from the same gene locus. The number of observed UMIs was converted into transcript counts using binomial statistics37.
Single-cell RNA sequencing data analysis
Overall, 10,372 cells passed the quality control threshold of >1,000 transcripts (Poisson-corrected UMIs37) for the normal human liver cell atlas. For cells from the organoids, 1,052 cells passed the quality control thresholds. For cells from HCC, 1,282 cells passed the quality control threshold of >1,000 transcripts. For cells from the humanized mouse, 311 cells passed the quality control threshold of >1,000 transcripts. All the datasets were analysed using RaceID34. For normalization, the total transcript counts in each cell were normalized to 1 and multiplied by the minimum total transcript count across all cells that passed the quality control threshold (>1,000 transcripts per cell). Prior to normalization, cells expressing >2% of KCNQ1OT1 transcripts, a previously identified marker of low quality cells18, were removed from the analysis. Moreover, transcripts correlating to KCNQ1OT1 with a Pearson’s correlation coefficient of >0.4 were also removed. RaceID3 was run with the following parameters: mintotal = 1000, minexpr = 2, minnumber = 10, outminc = 2, cln = 15.
Diffusion pseudo-time analysis and self-organizing maps
Diffusion pseudotime (dpt) analysis11 was implemented and diffusion maps generated using the destiny R package. The number of nearest neighbours, k, was set to 100. SOMs were generated using the FateID package on the basis of the ordering computed by dpt as input. Only genes with >2 counts after size normalization in at least a single cell were included for the SOM analysis. In brief, smooth zonation profiles were derived by applying local regression on normalized transcript counts after ordering cells by dpt. Next, a one-dimensional SOM with 200 nodes was computed on these profiles after z-transformation. Neighbouring nodes were merged if the Pearson’s correlation coefficient of the average profiles of these nodes exceeded 0.85. The remaining aggregated nodes represent the gene modules shown in the SOM figures.
P values for the significance of zonation were derived by binning dpt-ordered profiles into three equally sized bins to perform ANOVA. The resulting P values were corrected for multiple testing using the Benjamini–Hochberg method. Increasing the number of bins produced similar results.
Conservation of zonation between human and mouse
Expression data from Halpern et al.9 (GEO accession code GSE84498) were used for analysing the evolutionary conservation of hepatocyte zonation between human and mouse. The transcript count data were analysed using RaceID3 to determine cell types, with parameter mintotal = 1,000 and cln = 6. A subgroup of clusters was identified as hepatocytes on the basis of marker gene expression and used for dpt and SOM analysis, as was done for the human data. To obtain a similar number of genes, only genes with at least 1.5 counts after size normalization in at least a single cell were included. To identify orthologues between human and mouse for the references used in this study and by Halpern et al.9 as provided by the authors, we first identified pairs of orthologues based on identical gene identifiers upon capitalization of all letters. We further computed mutual blastn (run with default) best hits. The final list comprises 16,670 pairs of orthologues.
Conservation of zonation was assessed using Pearson’s correlation of zonated expression profiles after binning the human data into nine equally sized bins, akin to the nine zones derived in Halpern et al.9. Conservation of zonation of endothelial cells was evaluated based on published mouse data from Halpern et al.13 using classification into four spatially stratified populations. To calculate Pearson’s correlation coefficient between human and mouse endothelial cells, a diffusion-pseudotime analysis was performed for all human cells mapping to endothelial cell clusters and these profiles were discretized into four equally sized bins.
Lineage analysis of the EPCAM + compartment
For a separate analysis of the EPCAM+ population, all cells from clusters 4, 7, 24 and 39 were extracted and reanalysed using RaceID34 with the parameters mintotal = 1000 and minexpr = 2, minnumber = 10 outminc = 2, and default parameters otherwise. StemID24 was run on these clusters with cthr = 10, nmode = TRUE and knn = 3. FateID4 was run on the filtered and feature-selected expression matrix from RaceID3, with target clusters inferred by FateID using ASGR1 plus ALB and CXCL8 plus MMP7 as markers for hepatocyte and cholangiocyte lineage target clusters, respectively. Using KRT19 and CFTR as mature cholangiocyte markers yields highly similar results.
Differential gene expression analysis
Differential gene expression analysis between cells and clusters was performed using the diffexpnb function from the RaceID package. First, negative binomial distributions reflecting the gene expression variability within each subgroup were inferred on the basis of the background model for the expected transcript count variability computed by RaceID3. Using these distributions, a P value for the observed difference in transcript counts between the two subgroups was calculated and corrected for multiple testing using the Benjamini–Hochberg method as described38.
Pathway enrichment analysis and gene set enrichment analysis
Symbol gene IDs were first converted to Entrez gene IDs using the clusterProfiler39 package. Pathway enrichment analysis and GSEA40,41 were implemented using the ReactomePA42 package. Pathway enrichment analysis was done on genes taken from the different modules in the SOMs. GSEA was done using the differentially expressed genes inferred by the diffexpnb function from the RaceID package.
Validation of protein expression using the Human Protein Atlas
Immunostaining images were collected from the Human Protein Atlas31 (https://www.proteinatlas.org).
Immunofluorescence
Human liver tissue was fixed overnight in 3.7% formaldehyde (Fig. 3g) or cryosectioned and fixed in 2.5% paraformaldehyde for 20 min (Extended Data Fig. 7e). The tissue was embedded in OCT and stored at –80 °C. The tissue was cryosectioned into 7-μm sections. The tissue was washed twice for 5 min in 0.025% Triton 1× PBS. The tissue was then blocked in 10% FBS with 1% BSA in 1× PBS for 2 h at room temperature. The dilution used for the anti-human KRT19 (HPA002465, Sigma, Fig. 3g; MA5-12663, Invitrogen, Extended Data Fig. 7e) and EPCAM (SAB4200704, Sigma, Fig. 3g; PA5-19832, Invitrogen, Extended Data Fig. 7e) antibodies was 1:100 in 100 μl 1× PBS with 1% BSA. The antibodies were incubated overnight at 4 °C in the dark. The tissue was washed twice with 0.025% Triton 1× PBS and then incubated with secondary antibodies donkey anti-rabbit IgG-AF488 ((A21206, Thermo Fisher Scientific), Fig. 3g) and goat anti-mouse IgG-AF568 ((A11019, Thermo Fisher Scientific), Fig. 3g) or sheep anti-mouse IgG-AF488 ((515-545-062, Jackson ImmunoResearch), Extended Data Fig. 7e) at 1:200 dilution and donkey anti-rabbit IgG-RRX ((711-295-152, Jackson ImmunoResearch), Extended Data Fig. 7e) at 1:100 dilution in 1× PBS with 1% BSA for 1 h at room temperature. The tissue was then washed twice with 0.025% Triton 1× PBS. DAPI Fluoromount-G (Southern Biotech) was added to the tissue and a coverslip placed on top. Imaging was done using a Zeiss confocal microscope LSM780 (Fig. 3g) or ZEISS Axio Vert.A1 (Extended Data Fig. 7e). Images were taken at 63× magnification.
Organoid culturing
Organoid culturing was done as previously described43. The cell populations from the EPCAM+ compartment were sorted on an Aria Fusion I using a 100-μm nozzle into tubes containing culture medium supplemented with 10 μM ROCK inhibitor (Y-27632) (Sigma-Aldrich). After sorting, cells were centrifuged in order to remove the medium and then resuspended in 25 μl Matrigel. Droplets of the Matrigel solution containing the cells were added to the wells of a 24-well suspension plate and incubated for 5–10 min at 37 °C until the Matrigel solidified. Droplets were overlaid with 250 μl liver isolation medium and then incubated at 37 °C, 5% CO2. After 3–4 days, the liver isolation medium was replaced with liver expansion medium. For the single-cell culture, from each patient, single cells from the TROP2int gate were sorted into the wells of a non-tissue-culture-treated 96-well plate containing medium with 5% Matrigel. Organoids were passaged 14 days after isolation and then passaged multiple times 5–7 days after splitting. For FACS, single-cell suspensions were prepared from the organoids by mechanical dissociation followed by TrypLE (Life Technologies) digestion as previously described43. Organoid cells were sequenced 5 days after splitting and 17 days after initially sorting the cells for the culture.
Step-by-step protocol
A detailed protocol for scRNA-seq of cryopreserved human liver cells is available at Protocol Exchange44.
Reporting summary
Further information on research design is available in the Nature Research Reporting Summary linked to this paper.
Data availability
Data generated during this study have been deposited in the Gene Expression Omnibus (GEO) with the accession code GSE124395. The human liver cell atlas can be interactively explored at http://human-liver-cell-atlas.ie-freiburg.mpg.de/.
Change history
09 September 2019
In this Article, the issue paginated PDF only included the paginated pages, this has now been updated to include the online-only sections again. The HTML was unaffected.
References
Michalopoulos, G. K. & DeFrances, M. C. Liver regeneration. Science 276, 60–66 (1997).
Ryerson, A. B. et al. Annual report to the nation on the status of cancer, 1975–2012, featuring the increasing incidence of liver cancer. Cancer 122, 1312–1337 (2016).
Grün, D. & van Oudenaarden, A. Design and analysis of single-cell sequencing experiments. Cell 163, 799–810 (2015).
Herman, J. S., Sagar & Grün, D. FateID infers cell fate bias in multipotent progenitors from single-cell RNA-seq data. Nat. Methods 15, 379–386 (2018).
Grün, D. et al. Single-cell messenger RNA sequencing reveals rare intestinal cell types. Nature 525, 251–255 (2015).
Jungermann, K. & Kietzmann, T. Zonation of parenchymal and nonparenchymal metabolism in liver. Annu. Rev. Nutr. 16, 179–203 (1996).
Gebhardt, R. Metabolic zonation of the liver: regulation and implications for liver function. Pharmacol. Ther. 53, 275–354 (1992).
Kietzmann, T. Metabolic zonation of the liver: the oxygen gradient revisited. Redox Biol. 11, 622–630 (2017).
Halpern, K. B. et al. Single-cell spatial reconstruction reveals global division of labour in the mammalian liver. Nature 542, 352–356 (2017).
MacParland, S. A. et al. Single cell RNA sequencing of human liver reveals distinct intrahepatic macrophage populations. Nat. Commun. 9, 4383 (2018).
Haghverdi, L., Büttner, M., Wolf, F. A., Buettner, F. & Theis, F. J. Diffusion pseudotime robustly reconstructs lineage branching. Nat. Methods 13, 845–848 (2016).
Strauss, O., Phillips, A., Ruggiero, K., Bartlett, A. & Dunbar, P. R. Immunofluorescence identifies distinct subsets of endothelial cells in the human liver. Sci. Rep. 7, 44356 (2017).
Halpern, K. B. et al. Paired-cell sequencing enables spatial gene expression mapping of liver endothelial cells. Nat. Biotechnol. 36, 962–970 (2018).
Raven, A. et al. Cholangiocytes act as facultative liver stem cells during impaired hepatocyte regeneration. Nature 547, 350–354 (2017).
Michalopoulos, G. K., Barua, L. & Bowen, W. C. Transdifferentiation of rat hepatocytes into biliary cells after bile duct ligation and toxic biliary injury. Hepatology 41, 535–544 (2005).
Schmelzer, E. et al. Human hepatic stem cells from fetal and postnatal donors. J. Exp. Med. 204, 1973–1987 (2007).
Turner, R. et al. Human hepatic stem cell and maturational liver lineage biology. Hepatology 53, 1035–1045 (2011).
Grün, D. et al. De novo prediction of stem cell identity using single-cell transcriptome data. Cell Stem Cell 19, 266–277 (2016).
Okabe, M. et al. Potential hepatic stem cells reside in EpCAM+ cells of normal and injured mouse liver. Development 136, 1951–1960 (2009).
Cardinale, V. et al. Multipotent stem/progenitor cells in human biliary tree give rise to hepatocytes, cholangiocytes, and pancreatic islets. Hepatology 54, 2159–2172 (2011).
Kodama, Y. et al. Hes1 is required for the development of intrahepatic bile ducts. Gastroenterology 124, A123 (2003).
Sosa-Pineda, B., Wigle, J. T. & Oliver, G. Hepatocyte migration during liver development requires Prox1. Nat. Genet. 25, 254–255 (2000).
Huch, M. et al. Long-term culture of genome-stable bipotent stem cells from adult human liver. Cell 160, 299–312 (2015).
Betge, J. et al. MUC1, MUC2, MUC5AC, and MUC6 in colorectal cancer: expression profiles and clinical significance. Virchows Arch. 469, 255–265 (2016).
Park, S.-W. et al. The protein disulfide isomerase AGR2 is essential for production of intestinal mucus. Proc. Natl Acad. Sci. USA 106, 6950–6955 (2009).
Forner, A., Reig, M. & Bruix, J. Hepatocellular carcinoma. Lancet 391, 1301–1314 (2018).
Matkowskyj, K. A. et al. Aldoketoreductase family 1B10 (AKR1B10) as a biomarker to distinguish hepatocellular carcinoma from benign liver lesions. Hum. Pathol. 45, 834–843 (2014).
Rantakari, P. et al. The endothelial protein PLVAP in lymphatics controls the entry of lymphocytes and antigens into lymph nodes. Nat. Immunol. 16, 386–396 (2015).
Grompe, M. & Strom, S. Mice with human livers. Gastroenterology 145, 1209–1214 (2013).
Azuma, H. et al. Robust expansion of human hepatocytes in Fah–/–/Rag2–/–/Il2rg–/– mice. Nat. Biotechnol. 25, 903–910 (2007).
Uhlén, M. et al. Proteomics. Tissue-based map of the human proteome. Science 347, 1260419 (2015).
Krieger, S. E. et al. Inhibition of hepatitis C virus infection by anti-claudin-1 antibodies is mediated by neutralization of E2-CD81-claudin-1 associations. Hepatology 51, 1144–1157 (2010).
Lieber, A., Peeters, M. J., Gown, A., Perkins, J. & Kay, M. A. A modified urokinase plasminogen activator induces liver regeneration without bleeding. Hum. Gene Ther. 6, 1029–1037 (1995).
Mailly, L. et al. Clearance of persistent hepatitis C virus infection in humanized mice using a claudin-1-targeting monoclonal antibody. Nat. Biotechnol. 33, 549–554 (2015).
Hashimshony, T. et al. CEL-Seq2: sensitive highly-multiplexed single-cell RNA-seq. Genome Biol. 17, 77 (2016).
Li, H. & Durbin, R. Fast and accurate long-read alignment with Burrows-Wheeler transform. Bioinformatics 26, 589–595 (2010).
Grün, D., Kester, L. & van Oudenaarden, A. Validation of noise models for single-cell transcriptomics. Nat. Methods 11, 637–640 (2014).
Anders, S. & Huber, W. Differential expression analysis for sequence count data. Genome Biol. 11, R106 (2010).
Yu, G., Wang, L.-G., Han, Y. & He, Q.-Y. clusterProfiler: an R package for comparing biological themes among gene clusters. OMICS 16, 284–287 (2012).
Subramanian, A. et al. Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc. Natl Acad. Sci. USA 102, 15545–15550 (2005).
Mootha, V. K. et al. PGC-1α-responsive genes involved in oxidative phosphorylation are coordinately downregulated in human diabetes. Nat. Genet. 34, 267–273 (2003).
Yu, G. & He, Q. Y. ReactomePA: an R/Bioconductor package for reactome pathway analysis and visualization. Mol. Biosyst. 12, 477–479 (2016).
Broutier, L. et al. Culture and establishment of self-renewing human and mouse adult liver and pancreas 3D organoids and their genetic manipulation. Nat. Protocols 11, 1724–1743 (2016).
Aizarani, N. et al. Protocol for single-cell RNA-sequencing of cryopreserved human liver cells. Protoc. Exch. https://doi.org/10.21203/rs.2.9620/v1 (2019).
Acknowledgements
This study was supported by the Max Planck Society, the German Research Foundation (DFG) (SPP1937 GR4980/1-1, GR4980/3-1, and GRK2344 MeInBio), the DFG under Germany’s Excellence Strategy (CIBSS – EXC-2189 – Project ID 390939984), and the Behrens-Weise-Foundation (all to D.G.); and by ARC, Paris and Institut Hospital-Universitaire, Strasbourg (TheraHCC, TheraHCC2.0 IHUARC IHU201301187 and IHUARC2019 IHU201901299 to T.F.B.), the European Union (ERC-AdG-2014-671231-HEPCIR, EU H2020-667273-HEPCAR and ERC-PoC-2016-PRELICAN to T.F.B), Agence nationale de recherches sur le sida et les hépatites virales (ANRS, ECTZ35076) and the Foundation of the University of Strasbourg. This work was done under the framework of the LABEX ANR-10-LABX-0028_HEPSYS and Inserm Plan Cancer (Plan Cancer 2014-2019, Action 13.1, appel à projets 2018) and benefits from funding from the state managed by the French National Research Agency as part of the Investments for the future. Institut Universitaire France (IUF) to T.F.B. We thank S. Hobitz, K. Schuldes (MPI-IE FACS facility) and U. Bönisch (MPI-IE Deep Sequencing facility); the CRB (Centre de Ressources Biologiques-Biological Resource Centre of the Strasbourg University Hospitals) for the management of regulatory requirements of patient-derived liver tissue; C. Fauvelle and L. Heydmann for their contributions to the initial single-cell isolations; F. Juehling, F. H. T. Duong and C. Schuster for helpful discussions; the patients for providing informed consent to participate in the study; and the nurses, technicians and medical doctors of the hepatobiliary surgery and pathology services of Strasbourg University Hospitals for their support. This publication is part of the Human Cell Atlas (https://www.humancellatlas.org/publications).
Peer review information
Nature thanks Meritxell Huch, Shalev Itzkovitz and the other anonymous reviewer(s) for their contribution to the peer review of this work.
Author information
Authors and Affiliations
Contributions
T.F.B. and D.G. conceived the study. N.A. designed, optimized and performed cell sorting, scRNA-seq, organoid culture and immunofluorescence, provided validation using the Human Protein Atlas, and performed computational analysis and interpretation of the data. A.S. managed the supply of patient material, isolated single cells from patient tissue, performed animal experiments and carried out immunofluorescence. S. contributed to scRNA-seq analyses and performed scRNA-seq experiments. L.M. performed animal experiments. J.S.H. created the web interface. S.D. isolated single cells from patient tissues. P.P. performed liver resections and provided patient liver samples. T.F.B. established the liver tissue supply pipeline and supervised the animal experiments. D.G. analysed and interpreted the data and supervised experiments and analysis by N.A., S. and J.S.H. D.G., N.A. and T.F.B. coordinated and led the study. N.A. and D.G. wrote the manuscript with input from S., A.S. and T.F.B.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 scRNA-seq analysis of normal liver resection specimens from nine adult patients.
a, FACS plot for CD45 and ASGR1 staining from a mixed fraction (hepatocyte and non-parenchymal cells). b, FACS plot for PECAM1 and CD34 staining from a mixed fraction. c, FACS plot for CLEC4G staining from a mixed fraction. a–c, n = 6 independent experiments. d, t-SNE map showing the IDs of the nine patients from whom the cells were sequenced. Cells were sequenced from freshly prepared single-cell suspensions for patients 301, 304, 325 and BP1, and from cryopreserved single-cell suspensions for patients 301, 304, 308, 309, 310, 311, 315 and 325. Cells were sorted and sequenced mainly in an unbiased fashion from non-parenchymal cell, hepatocyte and mixed fractions for patients 301 and 304. Non-parenchymal and mixed fractions were used to sort specific populations on the basis of markers. CD45– and CD45+ cells were sorted from all patients. CLEC4G+ LSECs were sorted by FACS from patients 308, 310, 315 and 325. EPCAM+ cells were sorted by FACS from patients 308, 309, 310, 311, 315 and 325. e, t-SNE map highlighting data for fresh and cryopreserved cells from patients 301, 304 and 325. Although minor shifts in frequencies within cell populations are visible, transcriptomes of fresh and cryopreserved cells co-cluster. Differential gene expression analysis of fresh versus cryopreserved cells, for example, for endothelial cells of patient 325 in cluster 10 (f), did not reveal any differentially expressed genes. d, e, n = 10,372 cells. f, Bar plot showing the number of differentially expressed genes (Benjamini–Hochberg corrected P < 0.01; see Methods) between fresh and cryopreserved cells within each cluster for patient 325 (top; n = 2,248 cells) and patients 304 (n = 959 cells) and 301 (n = 1,329 cells) (bottom). Only clusters with more than five cells from fresh and cryopreserved samples were included for the computation. g, Scatter plot of mean normalized expression across fresh and cryopreserved cells from patient 325 in endothelial cells of cluster 10 (no differentially expressed genes, left; n = 101 cells) and cluster 11 (maximal number of differentially expressed genes across all clusters, right; n = 272 cells). Red dots indicate differentially expressed genes (Benjamini–Hochberg corrected P < 0.01; see Methods). Diagonal (solid black line) and log2 fold changes of 4 (broken black lines) are indicated. Almost all differentially expressed genes for cluster 11 exhibit log2 fold changes of less than 4. h, Bar plot showing the fraction of sorted cells which passed quality filtering (see Methods) after scRNA-seq. Error bars are derived from the sampling error assuming binomial counting statistics. F, fresh samples; C, cryopreserved samples. i, t-SNE map highlighting cells sequenced from mixed plates representing unbiased samples for patients 301 and 304. Without any enrichment strategy, hepatocytes and immune cells strongly dominate and endothelial cells and EPCAM+ cells are rarely sequenced. j, Table of patient information. CCM, colon cancer metastasis; ICC, intrahepatic cholangiocarcinoma; LR, liver resection.
Extended Data Fig. 2 The endothelial cell compartment is a heterogeneous mixture of subpopulations.
a, Expression heat map of genes upregulated in endothelial cell clusters (Benjamini–Hochberg corrected P < 0.01; n = 1,830 cells; see Methods). For each cluster the top ten upregulated genes were extracted and expression of the joint set is shown in the heat map across all endothelial cell clusters. Genes were ordered by hierarchical clustering. b, Expression t-SNE maps for the LSEC and MaVEC marker genes PECAM1, CLEC4G, CD34, CLEC4M and FLT1. c, Expression t-SNE maps for VWF, AQP1, CCL21, TFF3, UNC5B and IGFBP5. d, Expression t-SNE maps for CPE and CLU. e, Expression t-SNE map for H19. b–e, Colour bars indicate log2 normalized expression. n = 10,372 cells. f, Immunostaining of CD34, CLEC4G, PECAM1 and AQP1 in normal liver tissue from the Human Protein Atlas. The portal area for AQP1 is enlarged to show positive staining of both bile duct cells and portal MaVECs (black arrows).
Extended Data Fig. 3 Evolutionary conservation of zonation profiles.
a, Diffusion maps highlighting inferred dpt and Alb expression (left), and a self-organizing map for mouse hepatocyte single-cell RNA-seq data9 (right; see Methods). See Fig. 2 for details. b, Heat map showing the spatial hepatocyte zonation profiles (nine zones) inferred by Halpern et al.9 using the same ordering of genes as in a. c, Pearson’s correlation coefficient of zonation profiles inferred by Halpern et al.9 and our dpt approach after discretizing dpt-inferred zonation profiles into nine equally sized bins. We found that 1,347 out of 1,684 genes (80%) above the expression cutoff exhibited a positive correlation between the two methods. d, Pearson’s correlation coefficient as a function of average normalized expression. Negative correlations are enriched at low expression, and Pearson’s correlation of zonation profiles positively correlates with expression (Spearman’s R = 0.25; n = 1,684 genes). e, t-SNE map of single-cell transcriptomes highlighting the clusters generated by RaceID3, run separately on hepatocytes (clusters 11, 14, and 17 in Fig. 1c). The map reveals a major group of hepatocyte clusters and a number of small clusters that co-express T cell-related genes, B cell-related genes or progenitor genes. f, t-SNE maps highlighting the expression of ALB, the immune cell marker gene PTPRC, the B cell marker gene IGKC, and the progenitor marker gene EPCAM. The colour bar indicates log2 normalized expression. Co-expression of hepatocyte and immune cell markers could either indicate the presence of doublets or be due to spillover of highly expressed genes such as ALB between cells during library preparation. For the zonation analysis (Fig. 2), only cells in clusters 3, 7, 19, 4, 2, 9, 8 and 11 from the map in e were included. e, f, n = 3,040 cells. g, Immunostaining for the periportal markers CPS1, PCK1, MTHFS, and GATM from the Human Protein Atlas31. The zonation module containing each gene in the SOM (Fig. 2a) is indicated in parentheses. P, portal tracts; C, central veins. h, Pathways enriched for genes in hepatocyte central/mid modules 24 and 33 (top; n = 659 genes) and periportal modules 1 and 3 (bottom; n = 422 genes) (compare with Fig. 2a). i, Immunostaining of the central marker ENG from the Human Protein Atlas31. The zonation module in the SOM (Fig. 2b) is indicated in parentheses. j, Pathways enriched for genes in endothelial central/mid modules 1 and 3 (top; n = 422 genes) and periportal module 20 (bottom; n = 73 genes) (compare with Fig. 2b). h, j, P values in the pathway enrichment analysis were calculated using a hypergeometric model and adjusted using the Benjamini–Hochberg method (see Methods). k, Pearson’s correlation coefficient of hepatocyte zonation profiles of orthologue pairs of human and mouse genes. Mouse data are from Halpern et al.9 (n = 967 genes) l, Pearson’s correlation coefficient of endothelial cell (including MVECs and LSECs) zonation profiles of orthologue pairs of human and mouse genes (n = 977 genes). Mouse data are from Halpern et al.13. See Methods for details.
Extended Data Fig. 4 The human liver contains different Kupffer cell populations.
a, Expression t-SNE maps of marker genes for Kupffer cell subtypes. The colour bar indicates log2 normalized expression (n = 10,372 cells). b, Major pathways upregulated in the CD1C+ antigen-presenting (n = 12 genes) and LILRB5+ metabolic/immunoregulatory (n = 35 genes) Kupffer cell subsets as revealed by Reactome pathway analysis. The number of genes in each pathway is shown on the x axis. P values were calculated using a hypergeometric model and adjusted using the Benjamini–Hochberg method. c, Expression heat map of genes upregulated in Kupffer cell clusters (Benjamini–Hochberg corrected P < 0.01, see Methods). For each cluster, the top ten upregulated genes were extracted and expression of the joint set is shown in the heat map across all Kupffer cell clusters. Genes were ordered by hierarchical clustering.
Extended Data Fig. 5 The human liver contains different B cell populations.
Expression t-SNE maps of the markers for the B cell subtypes. The colour bars indicate log2 normalized expression (n = 10,372 cells).
Extended Data Fig. 6 Heterogeneity of NK and NKT cells in the human liver.
a–c, Expression t-SNE maps of inferred markers of cluster 5 (a), cluster 1 (b) and cluster 3 (c). Cluster 5 comprises mainly CD56+CD8A– NK cells, some of which show upregulated CCL4. Cluster 1 comprises CD56–CD8A+ NKT cells, which show upregulated CCL5. Cluster 3 consists of both CD56+ and CD56–CD8A+ NKT cells. Clusters 1 and 3 express T cell receptor components exemplified by CD3D co-receptor expression. d, Differential expression of killer cell lectin-like receptor genes across the different populations shown in a–c. e, Differential expression of granzyme genes across the different populations shown in a–c. Colour bars indicate log2 normalized expression. a–e, n = 10,372 cells.
Extended Data Fig. 7 scRNA-seq identifies marker genes expressed by EPCAM+ cells.
a, Expression t-SNE maps (left) for EPCAM, CD24, FGFR2, TACSTD2, CLDN1, TM4SF4, WWTR1 and ANXA4 (n = 10,372 cells) and immunohistochemistry from the Human Protein Atlas (right) for CLDN1, TM4SF4, WWTR1, and ANXA4. b, Expression t-SNE maps for ASGR1 and CFTR (n = 10,372 cells). c, t-SNE maps showing expression of KRT19, ALB, TACSTD2 and MUC6 in the EPCAM+ compartment (n = 1,087 cells). a–c, Colour bars indicate log2 normalized expression. d, Expression heat map of proliferation marker genes (MKI67, PCNA), AFP, and identified markers of the EPCAM+ compartment. Genes highlighted in red correspond to newly identified markers of the EPCAM+ compartment. The heat map comprises all clusters to show the specificity of the markers for the progenitor compartment. The expression analysis confirms the absence of proliferating and AFP+ cells. e, Immunofluorescence labelling of EPCAM and KRT19 on human liver tissue. EPCAM+KRT19low/– cells (solid arrow) in the canals of Hering (asterisk) and EPCAM+KRT19+ cells (broken arrow) in the bile duct (arrowhead) are indicated. Nuclei are stained with DAPI. Scale bar, 10 μm (n = 3 independent experiments).
Extended Data Fig. 8 The EPCAM+ compartment segregates into different major subpopulations.
a, Separate RaceID3 and StemID2 analyses of the EPCAM+ and hepatocyte populations reveal a lineage tree connecting EPCAM+ cells to mature hepatocytes via an EPCAM+ hepatocyte progenitor cluster (part of the EPCAM+ population in Fig. 3b). Shown are links with StemID2 P < 0.05. The node colour highlights transcriptome entropy. b, Two-dimensional diffusion map representation of the population shown in a, highlighting expression of the hepatocyte marker ALB (left), EPCAM (centre), and the mature cholangiocyte marker CFTR (right). The maps suggest continuous transitions from the EPCAM+ compartment towards hepatocytes and mature cholangiocytes. c, Expression t-SNE map of EPCAM (top) and the hepatocyte marker ASGR1 (bottom) for the population shown in a. Colour bars indicate log2 normalized expression. b, c, n = 3,877 cells. d, Expression heat map of de novo identified markers of the EPCAM+ compartment, highlighting the expression distribution within clusters of this population only (Fig. 3). e, Expression heat map of all genes that were differentially expressed in the more mature clusters, belonging to the groups denoted as ‘hepatocyte fate’ and ‘cholangiocyte fate’. For each of these clusters, the top ten upregulated genes (Benjamini–Hochberg corrected P < 0.01) were selected, and the joint set of these genes is shown in the figure. f, Expression t-SNE maps of CXCL8, MMP7 and HP. Colour bars indicate log2 normalized expression. d–f, n = 1,087 cells. g, Normalized expression counts of ALB, KRT19 and TACSTD2 in cells sequenced from the gates in Fig. 4a (n = 293 cells). Centre line, mean; boxes, interquartile range; whiskers, 5% and 95% quantiles; data points, outliers. h, t-SNE map of RaceID3 clusters for organoid cells and EPCAM+ cells from patients (Fig. 3), including cells sorted from the gates in a. i, Expression t-SNE maps of EPCAM, CD24 and AQP1 in organoid cells and EPCAM+ cells from patients. j, Expression t-SNE maps of SFRP5, ALB, AGR2 and MKI67. Colour bars indicate log2 normalized expression. h–j, n = 2,870 cells. k, GSEA of genes that are differentially expressed between organoid and EPCAM+ liver cells from patients (Benjamini–Hochberg corrected P < 0.01, n = 11,610 genes; see Methods).
Extended Data Fig. 9 Cell types from patient-derived HCC exhibit perturbed gene expression signatures.
a, FACS plot of CD45 and ASGR1 staining on cells from HCC samples (n = 3 independent experiments). b, Symbol t-SNE map showing the IDs of HCC patients (n = 11,654 cells). c, t-SNE map showing RaceID3 clusters for normal liver cells co-analysed with cells from HCC tissues (n = 3 patients). d, Expression heat map of genes that are differentially expressed between cancer cells from HCC and normal hepatocytes (Benjamini–Hochberg corrected P < 0.05 and log2 fold change >1.6; n = 256 cells; see Methods). Genes highlighted in red correspond to differentially expressed genes validated by immunohistochemistry. e, Immunostaining of CPS1 and CYP2C8 in normal liver and HCC tissues from the Human Protein Atlas. f, Expression heat map of genes that are differentially expressed between endothelial cells from HCC and normal endothelial cells from MaVEC and LSEC clusters. Benjamini–Hochberg corrected P < 0.05; log2 fold change >1.5; n = 1,936 cells; see Methods). Genes highlighted in red correspond to differentially expressed genes validated by immunohistochemistry. g, Immunostaining of CD34, LAMB1, AQP1 and PLVAP in normal liver and HCC tissues from the Human Protein Atlas. h, Heat map of genes that are differentially expressed between normal and HCC-resident NK and NKT cells (Benjamini–Hochberg corrected P < 0.05; n = 2,754 cells; see Methods). i, Heat map of genes that are differentially expressed between normal and HCC-resident Kupffer cells (Benjamini–Hochberg corrected P < 0.05; n = 991 cells; see Methods). j, GSEA of genes that are differentially expressed between normal and HCC-resident NK and NKT cells (n = 15,442 genes). k, GSEA of genes that are differentially expressed between normal and HCC-resident Kupffer cells (n = 15,442 genes).
Extended Data Fig. 10 Transplanted human liver cells in a humanized mouse model exhibit a distinct gene signature compared to cells within the human liver.
a, t-SNE map of RaceID3 clusters of liver cells from patients co-analysed with cells from the humanized mouse liver model. b, Expression t-SNE maps of the hepatocyte marker gene ALB. c, Expression t-SNE maps of the endothelial marker CLEC4G. d, Expression t-SNE maps of HP, PCK1 and CCND1. e, Expression t-SNE maps of the liver endothelial cell zonated genes LYVE1, FCN3 and CD14. f, Expression t-SNE maps of PECAM1, CD34 and AQP1. a–f, Colour bars indicate log2 normalized expression. n = 10,683 cells. g, Heat maps of genes that are differentially expressed between hepatocytes (n = 3,175 cells) and endothelial cells (n = 1,710 cells) from patients (human hepatocytes and human endothelial cells) and from the humanized mouse model (HMouse hepatocytes and HMouse endothelial cells). Benjamini–Hochberg corrected P < 0.05; see Methods.
Supplementary information
Supplementary Information
This file contains Supplementary Notes 1-5 and Supplementary References.
41586_2019_1373_MOESM3_ESM.xls
Supplementary Table 1 Gene expression signatures of all clusters. Official gene symbol, mean expression across all genes not contained within a cluster (mean.ncl), mean expression across all genes contained within a cluster (mean.cl), fold change between cluster and non-cluster cells (fc), differential expression p-value (pv, see Methods), and Benjamini-Hochberg corrected p-value are given. Only genes with P<0.01 are shown for each cluster. (n= 10,372 cells).
41586_2019_1373_MOESM4_ESM.xls
Supplementary Table 2 Human hepatocyte zonation. Official gene symbol, SOM module and Benjamini-Hochberg corrected ANOVA p-value of zonated genes in human hepatocytes (n=3 bins; 2,534 cells; Methods).
41586_2019_1373_MOESM5_ESM.zip
Supplementary Table 3 Human hepatocyte zonation data. Official gene symbol, SOM module and loess-smoothed expression z-scores (see Methods) of DPT-ordered hepatocytes. These data were used to create the zonation map in Fig. 2a.
41586_2019_1373_MOESM6_ESM.xls
Supplementary Table 4 Human endothelial cell zonation. Official gene symbol, SOM module and Benjamini-Hochberg corrected ANOVA p-value of zonated genes in human liver endothelial cells (n=3 bins; 1,361 cells; Methods).
41586_2019_1373_MOESM7_ESM.zip
Supplementary Table 5 Human endothelial cell zonation data. Official gene symbol, SOM module and loess-smoothed expression z-scores (see Methods) of DPT-ordered endothelial cells. These data were used to create the zonation map in Fig. 2b.
41586_2019_1373_MOESM8_ESM.xls
Supplementary Table 6 Conservation of hepatocyte zonation between mouse and human. Official gene symbol, SOM module and Benjamini-Hochberg corrected ANOVA p-value of zonated genes in human and mouse hepatocytes (Methods), and Pearson’s correlation coefficient between mouse and human zonation profiles (n=9 bins).
41586_2019_1373_MOESM9_ESM.xls
Supplementary Table 7 Conservation of liver endothelial cell zonation between mouse and human. Official gene symbol, SOM module and Benjamini-Hochberg corrected ANOVA p-value of zonated genes in human liver endothelial cells (Methods) and Pearson’s correlation coefficient between mouse and human zonation profiles. Mouse zonation p-values are taken from Halpern et al. 13 (n=4 bins).
Rights and permissions
About this article
Cite this article
Aizarani, N., Saviano, A., Sagar et al. A human liver cell atlas reveals heterogeneity and epithelial progenitors. Nature 572, 199–204 (2019). https://doi.org/10.1038/s41586-019-1373-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-019-1373-2
- Springer Nature Limited
This article is cited by
-
Role of liver sinusoidal endothelial cell in metabolic dysfunction-associated fatty liver disease
Cell Communication and Signaling (2024)
-
Spatial multiomics reveals a subpopulation of fibroblasts associated with cancer stemness in human hepatocellular carcinoma
Genome Medicine (2024)
-
Integrated meta-omics reveals the regulatory landscape involved in lipid metabolism between pig breeds
Microbiome (2024)
-
Spatial transcriptomics reveals tumor-derived SPP1 induces fibroblast chemotaxis and activation in the hepatocellular carcinoma microenvironment
Journal of Translational Medicine (2024)
-
The roles of tissue resident macrophages in health and cancer
Experimental Hematology & Oncology (2024)