Abstract
Heart failure with preserved ejection fraction (HFpEF) is a common syndrome with high morbidity and mortality for which there are no evidence-based therapies. Here we report that concomitant metabolic and hypertensive stress in mice—elicited by a combination of high-fat diet and inhibition of constitutive nitric oxide synthase using Nω-nitro-l-arginine methyl ester (l-NAME)—recapitulates the numerous systemic and cardiovascular features of HFpEF in humans. Expression of one of the unfolded protein response effectors, the spliced form of X-box-binding protein 1 (XBP1s), was reduced in the myocardium of our rodent model and in humans with HFpEF. Mechanistically, the decrease in XBP1s resulted from increased activity of inducible nitric oxide synthase (iNOS) and S-nitrosylation of the endonuclease inositol-requiring protein 1α (IRE1α), culminating in defective XBP1 splicing. Pharmacological or genetic suppression of iNOS, or cardiomyocyte-restricted overexpression of XBP1s, each ameliorated the HFpEF phenotype. We report that iNOS-driven dysregulation of the IRE1α–XBP1 pathway is a crucial mechanism of cardiomyocyte dysfunction in HFpEF.
Similar content being viewed by others
Main
HFpEF is a growing public health problem, accounting for approximately half of hospital admissions of individuals with heart failure1. The complex clinical phenotype that characterizes HFpEF stems from the presence of multiple comorbidities, including obesity, hypertension and diabetes2. Clinical data suggest that systemic inflammation and an imbalance in the levels of nitric oxide (NO) are crucial for the development of HFpEF3. However, definitive experimental evidence supporting these concepts has not been described owing to limitations in preclinical models that fail to recapitulate the full range of features found in HFpEF. As a result, fundamental pathophysiological mechanisms remain unclear, and there are no evidence-based clinical therapies available for individuals with HFpEF2,4,5.
A ‘two-hit’ mouse model of HFpEF
Because patients with HFpEF very often have the comorbidities of hypertension, obesity and metabolic dysfunction2, we formulated a two-hit hypothesis, one in which the coincidence of metabolic stress (obesity and metabolic syndrome) and mechanical stress (hypertension induced by constitutive NO synthases suppression) is a major mechanism that underlies HFpEF pathophysiology. To test this, male C57BL/6N wild-type mice were divided into four treatment groups and exposed to one of the following regimens: a high-fat diet (HFD) (60% kilocalories from fat (lard)); l-NAME (0.5 g l−1 in drinking water); a combination of both treatments (two-hit model; HFD + l-NAME); or standard (chow) diet for 5 or 15 weeks (Fig. 1a). As expected, HFD induced body weight increases and glucose intolerance (Extended Data Fig. 1a–c), whereas l-NAME treatment raised both systolic and diastolic blood pressure (Extended Data Fig. 1d, e). Longitudinal echocardiographic evaluation revealed persistent preservation of the left ventricular ejection fraction (LVEF) in all groups (for at least one year in the HFD + l-NAME cohort; Fig. 1b, c, Extended Data Fig. 1f and Extended Data Table 1), coupled with significant alterations in left ventricular global longitudinal strain, uniquely in mice exposed to the combination of HFD and l-NAME at both 5- and 15-week time points (Fig. 1d and Extended Data Fig. 1g). Whereas varying degrees of diastolic dysfunction were observed in the different experimental groups, animals concomitantly exposed to both HFD and l-NAME manifested signs of increased left ventricular filling pressure as measured both by noninvasive Doppler imaging (Fig. 1e, f, Extended Data Fig. 1h, i and Extended Data Table 1) and invasive (Extended Data Table 1) analyses. Consistent with the documented elevated filling pressures, mice exposed to HFD + l-NAME uniquely exhibited a robust increase in lung weight (Fig. 1g and Extended Data Fig. 1j), indicative of pulmonary congestion and a preclinical surrogate for heart failure. Cardiac hypertrophy (Fig. 1h and Extended Data Fig. 1k) and cardiomyocyte hypertrophy (Extended Data Fig. 2a, b), as well as cardiac fibrosis (Extended Data Fig. 2a, c) and a reduction in myocardial capillary density (Extended Data Fig. 2a, d), were observed in mice treated with l-NAME or the combination of HFD + l-NAME. Chronic l-NAME administration also increased aortic stiffness, measured by pulse-wave velocity (Extended Data Fig. 2e), and impaired endothelial function in the coronary arteries, reducing coronary flow reserve (Extended Data Fig. 2f, g). Whereas excess weight affected exercise performance (Fig. 1i and Extended Data Fig. 1l) after 15 weeks of combination treatment, HFD + l-NAME-fed mice displayed a significant reduction in running distance compared to the other cohorts (Fig. 1i). Importantly, exercise intolerance in HFD + l-NAME-fed mice occurred in the absence of major histopathological (Extended Data Fig. 3a), molecular (Extended Data Fig. 3b) or strength (Extended Data Fig. 3c–f) abnormalities in skeletal muscle, which suggests that—during the period of observation described here—the reduction in exercise tolerance in our HFpEF model is not due to a deficit in skeletal muscle function. Whereas some overlapping features were observed in l-NAME- and HFD-treated groups, the presence of the greatest hypertrophic response, worsened diastolic function and pulmonary congestion—which was observed exclusively in the HFD + l-NAME-fed mice—lend additional support to the notion that this is a bona fide model of HFpEF. Because abnormalities in cardiomyocyte contraction and relaxation are common features in HFpEF6,7, adult mouse ventricular myocytes were collected after five weeks of dietary manipulation, revealing significant reductions in contraction velocity and impaired relaxation only in the HFD + l-NAME-treated cohort (Extended Data Fig. 4a–f).
a, Experimental design. C57BL/6N mice were maintained on different dietary regimens and analysed at 5 and 15 weeks. b, Representative left ventricular M-mode echocardiographic tracings. Images are representative of 15 independent mice. c, Percentage of LVEF. n = 15 mice per group. d, Left ventricular global longitudinal strain (GLS). n = 10 mice per group. e, Representative pulsed-wave Doppler (top) and tissue Doppler (bottom) tracings. Images are representative of 15 independent mice. f, Ratio between mitral E wave and E′ wave (E/E′). n = 15 mice per group. g, Ratio between wet and dry lung weight (LW). n = 15 mice per group. h, Ratio of heart weight to tibia length (HW/TL). n = 15 mice per group. i, Running distance during exercise exhaustion test. n = 8 mice per group. Data are mean ± s.e.m. c, d, f–i, One-way analysis of variance (ANOVA) followed by Sidak’s multiple-comparisons test. Numbers above square brackets show significant P values.
The IRE1α–XBP1 signalling pathway is inactive in HFpEF
Evidence thus far suggested that concomitant exposure to HFD + l-NAME elicits unique functional and structural changes in the heart that mirror the clinical features of human HFpEF. Taking advantage of the recent observation that accumulation of misfolded proteins occurs in clinical HFpEF8, we evaluated the unfolded protein response (UPR)—an evolutionarily conserved adaptive response capable of mitigating stress in conditions that disrupt protein quality control9. The IRE1α–XBP1 axis is the evolutionarily most conserved branch of the UPR, one that has recently been identified as a regulator of cardiomyocyte stress responsiveness10. The role of the IRE1α–XBP1 signalling pathway in HFpEF is unknown.
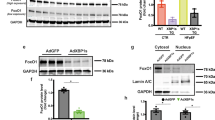
Analysis of transcript levels of multiple UPR markers in hearts from all four treatment groups at five weeks revealed activation of UPR sensors and effectors in mice treated with either HFD or l-NAME (Fig. 2a). In hearts from mice exposed to a combination of HFD + l-NAME, we detected no changes in transcript (Fig. 2a) and protein (Fig. 2b) levels of activating transcription factor 6 (ATF6) and a mild activation of the protein kinase RNA-like ER kinase (PERK) UPR branches (Fig. 2b). Notably, we detected a significant reduction below baseline in the transcript levels of binding immunoglobulin protein (Bip (also known as Hspa5)), CCAAT/enhancer-binding protein (C/EBP) homologous protein (CHOP (encoded by Ddit3)) and the spliced form of Xbp1 (Xpb1s) uniquely in hearts (Fig. 2a, c) and cardiomyocytes (Fig. 2d) from HFD + l-NAME-treated mice. Reduced levels of Xpb1s in hearts from mice exposed to HFD + l-NAME were associated with a reduction in IRE1α phosphorylation (Fig. 2e, f). These observations were replicated in left ventricular samples from a previously validated rat model of HFpEF: ZSF1-obese rats (Extended Data Fig. 4g–o).
a, mRNA levels of Xbp1s, Bip, Atf6, Atf4 and Chop in left ventricles of mice from different experimental groups. n = 5 mice per group. b, Immunoblot images of PERK, ATF6 and GAPDH proteins in left ventricular samples of mice fed a chow or HFD + l-NAME diet. n = 3 mice per group. c, Electrophoretic analysis of spliced (s) and unspliced (u) Xbp1 transcripts in left ventricular samples of mice fed a chow or HFD + l-NAME diet. Tunicamycin-treated NRVMs were used as positive control. TUN, tunicamycin. Images are representative of three independently performed experiments with similar results. d, mRNA levels of Xbp1s, Bip and Chop in adult mouse ventricular myocytes (AMVMs) from mice fed a chow or HFD + l-NAME diet. n = 4 mice per chow group; n = 5 mice per HFD + l-NAME group. Adult mouse ventricular myocytes were isolated from individual mice. e, Immunoblots of phosphorylated (p-)IRE1α, IRE1α and GAPDH from left ventricles of mice fed a chow or HFD + l-NAME diet. n = 3 mice per group. f, Densitometric analysis of the ratio of p-IRE1α/IRE1α protein bands. n = 3 mice per group. g, mRNA levels of XBP1s and XBP1u in human myocardial biopsies from non-failing controls, and patients with HFpEF or HFrEF. Control n = 15; HFpEF n = 13 (XBP1s) and n = 14 (XBP1u); HFrEF n = 15 patients. h, Immunoblot images of p-IRE1α and IRE1α in human myocardial biopsies from controls, patients with HFpEF or HFrEF. n = 8 subjects per group. i, Densitometric analysis ratio between p-IRE1α and IRE1α protein bands. n = 8 patients per group. j, Ratio between mitral E wave and A wave (E/A) of control and Xbp1s transgenic (Tg) mice fed a chow or HFD + l-NAME diet over time. k, Ratio between mitral E wave and E′ wave of control and Xbp1s transgenic mice fed a chow or HFD + l-NAME diet over time. n = 5 mice per chow control and chow transgenic groups; n = 7 mice per HFD + l-NAME control and HFD + l-NAME transgenic groups. Each mouse was analysed at all three time points. Dox, doxycycline. l, Running distance during exercise exhaustion test. m, Ratio between wet and dry lung weight at the end of the study. n = 5 mice per chow control and chow transgenic groups; n = 7 mice per HFD + l-NAME control and HFD + l-NAME transgenic groups. Data are mean ± s.e.m. a, g, i, One-way ANOVA followed by Sidak’s multiple-comparisons test. d, f, Two-tailed unpaired Student’s t-test. j–m, Two-way ANOVA followed by Sidak’s multiple-comparisons test. Numbers above square brackets show significant P values. For gel source data, see Supplementary Fig. 1.
In contrast to our findings in HFpEF rodents, we detected robust activation of this UPR pathway and increased Xbp1 splicing in a preclinical model of pressure-overload-induced heart failure with reduced ejection fraction (HFrEF) induced by severe transverse aortic constriction11 (Extended Data Fig. 5a–i). In addition, in a model of milder pressure overload (by transverse aortic constriction) before emergence of systolic dysfunction (three weeks), we observed similar levels of diastolic dysfunction and pulmonary congestion as in HFD + l-NAME-fed mice (Extended Data Fig. 6a–f), although without a reduction in the level of Xbp1s myocardial transcripts (Extended Data Fig. 6g) or reduction in IRE1α phosphorylation (Extended Data Fig. 6h), which supports the notion that in a transverse aortic constriction model the HFpEF phenotype is time-delimited, in contrast to observations in HFD + l-NAME mice (Extended Data Table 1) and in patients with HFpEF1.
To test for human relevance, we turned to endomyocardial biopsies of control and failing human hearts (Extended Data Table 2). Consistent with our observations in both ZSF1-obese rats and HFD + l-NAME-treated mice, the levels of XBP1s transcript were also reduced uniquely in human hearts from patients with HFpEF, whereas the unspliced form of XBP1 (XBP1u) did not change among hearts from controls, or patients with HFpEF or HFrEF (Fig. 2g). Accordingly, IRE1α phosphorylation levels were also downregulated exclusively in human HFpEF myocardium (Fig. 2h, i). Taken together, these data confirm that defective IRE1α splicing activity and XBP1s downregulation occur in both experimental and clinical HFpEF but not in HFrEF, which suggests distinct roles for this transcription factor in the two different types of heart failure.
Because our findings reveal that downregulation of Xbp1s correlated with the development of HFpEF, we used a cardiomyocyte-specific, doxycycline-responsive Tet-off Xbp1s transgenic mouse line10 to test for a causal role of XBP1s in HFpEF pathogenesis. Five weeks of exposure of transgenic Xbp1s and littermate control animals to the HFD + l-NAME regimen (Extended Data Fig. 6i) was sufficient to induce a cardiac phenotype characterized by diastolic dysfunction with preserved LVEF (Fig. 2j, k and Extended Data Fig. 6j). Notably, overexpression of Xbp1s for two weeks (Extended Data Fig. 6k) ameliorated diastolic dysfunction (Fig. 2j, k), improved exercise tolerance (Fig. 2l), reduced lung congestion (Fig. 2m) and decreased the expression of genes associated with heart failure (Extended Data Fig. 6l) without affecting LVEF (Extended Data Fig. 6j) or cardiac hypertrophy (Extended Data Fig. 6m). Taken together, these data demonstrate that overexpression of Xbp1s in cardiomyocytes is sufficient to partially ameliorate the diastolic dysfunction and signs of heart failure observed in HFpEF mice, which suggests a model in which reduced abundance of XBP1s in cardiomyocytes is a driving force in HFpEF pathogenesis.
iNOS-dependent S-nitrosylation of IRE1α in HFpEF
As previously noted, the current paradigm of HFpEF pathophysiology holds that comorbidities drive cardiac remodelling in HFpEF through microvascular endothelial inflammation12. A systemic pro-inflammatory state reduces endothelial NOS (eNOS) activity in coronary endothelial cells, limiting NO bioavailability for cardiomyocytes. In our model, l-NAME—which we use as a driver of endothelial dysfunction-based hypertension—is a substantially more potent inhibitor of eNOS and neuronal NOS (nNOS) compared to iNOS13. Notably, l-NAME-induced events are associated with upregulation of iNOS in the cardiovascular system14. Similarly, iNOS is upregulated in rodents exposed to HFD15. These observations suggest that the inflammatory state elicited by multiple comorbidities observed in clinical HFpEF might be replicated by the combination of HFD + l-NAME, pointing to iNOS as a common disease feature.
The combination of HFD and l-NAME elicited a systemic proinflammatory state (Extended Data Fig. 7a, b) coupled with increased levels of the Nos2 (which encodes iNOS) transcript (Fig. 3a) and protein (Extended Data Fig. 7c) in the myocardium relative to either treatment alone, and in the absence of changes in Nos3 (which encodes eNOS) transcripts (Extended Data Fig. 7d) or eNOS and nNOS proteins (Extended Data Fig. 7e). Changes in Nos2 levels were mirrored at the cardiomyocyte level (Fig. 3b). The level of Nos2 transcripts was increased in left ventricular samples from ZSF1-obese rats (Fig. 3c) and, importantly, NOS2 transcript levels were increased exclusively in human HFpEF hearts (Fig. 3d), which confirms that there is increased abundance of iNOS in the myocardium of both experimental and clinical HFpEF. Elevated iNOS activity and nitrosative stress promote S-nitrosylation of cysteine residues within multiple proteins and can perturb their function16. Accordingly, we observed a mild increase in total protein nitrosylation in hearts of HFD + l-NAME-fed mice (Extended Data Fig. 7f, g).
a, mRNA levels of Nos2 in left ventricles of mice from different experimental groups. n = 4 mice per chow group; n = 6 mice per HFD, l-NAME and HFD + l-NAME groups. b, mRNA levels of Nos2 in adult mouse ventricular myocytes isolated from chow-fed or HFD + l-NAME mice. n = 5 mice per group. Adult mouse ventricular myocytes were isolated from individual mice. c, mRNA levels of Nos2 in left ventricles of Wistar–Kyoto (WKY) and ZSF1-obese rats. n = 5 rats per group. d, mRNA levels of NOS2 in human myocardial biopsies from control (non-failing) patients or patients with HFpEF or HFrEF. n = 11 patients per control group, n = 11 patients per HFpEF group and n = 10 patients per HFrEF group. e, Immunoblots of S-nitrosylated IRE1α (SNO-IRE1α), IRE1α and GAPDH in left ventricular samples of wild-type and Nos2 knockout (Nos2 KO) mice after five weeks of chow or HFD + l-NAME diet. Blots are representative of four independently performed experiments with similar results. −Asc, without ascorbate; +Asc, with ascorbate; GSNO, S-nitrosoglutathione. f, Densitometric analysis of SNO-IRE1α with ascorbate protein bands in each group. n = 4 mice per group. g, Xbp1s mRNA level of NRVMs transduced with increasing multiplicities of infection of α-galactosidase adenovirus (AdLacZ) or AdiNOS for 24 h. n = 4 biologically independent experiments. h, Immunoblots of SNO-IRE1α and IRE1α in NRVMs transduced with wild-type IRE1α adenovirus (AdIRE1α WT) or AdIRE1α with mutations in two target nitrosylation sites (AdIRE1α M1+M2) and AdLacZ or AdiNOS for 24 h. Blots are representative of three independently performed experiments with similar results. i, Densitometric analysis of the ratio of SNO-IRE1α to IRE1α protein band intensities. n = 3 biologically independent experiments. j, Xbp1s mRNA level of NRVMs transduced with AdIRE1α WT or AdIRE1α M1+M2 and AdLacZ or AdiNOS for 24 h. n = 5 biologically independent experiments. Data are mean ± s.e.m. a, d, g, i, j, One-way ANOVA followed by Sidak’s multiple-comparisons test. b, c, Two-tailed unpaired Student’s t-test. f, Two-way ANOVA followed by Sidak’s multiple-comparisons test. Numbers above square brackets show significant P values. For gel source data, see Supplementary Fig. 1.
The observations of increased iNOS coupled with reduced levels of XBP1s in the myocardium of both experimental and human HFpEF raised the possibility of a connection between iNOS activity and XBP1s production. Liver from morbidly obese mice is marked by iNOS-dependent S-nitrosylation of IRE1α that inhibits its endonuclease activity, resulting in reduced XBP1 splicing15. Notably, in hearts of HFD + l-NAME-treated mice, we observed aberrant S-nitrosylation of IRE1α (Fig. 3e, f). To determine whether this mechanism occurs in cardiomyocytes, neonatal rat ventricular myocytes (NRVMs) were infected with an adenovirus driving iNOS expression (AdiNOS) (Extended Data Fig. 8a, b). Transduction of NRVMs with AdiNOS significantly increased NO production in culture medium collected from NRVMs (Extended Data Fig. 8c) without affecting cardiomyocyte viability (Extended Data Fig. 8d) and—importantly—resulted in a biphasic response in the levels of Xbp1s transcript, with an initial increase and subsequent decrease below basal levels (Fig. 3g). iNOS overexpression in cardiomyocytes was also sufficient to drive increases in global protein nitrosylation (Extended Data Fig. 8e) and IRE1α S-nitrosylation (Fig. 3h, i) in association with reduction in Xbp1s transcript levels (Fig. 3j). iNOS-dependent inhibition of IRE1α splicing activity was further corroborated by the observation of reduced IRE1α phosphorylation (Extended Data Fig. 8f) and blunted Xbp1s upregulation (Extended Data Fig. 8g) in cardiomyocytes treated with the endoplasmic reticulum stressor tunicamycin.
To test directly for an effect of nitrosylation on IRE1α endoribonuclease activity, we used an adenovirus that contained ERN1 (which encodes IRE1α) with mutations in two target nitrosylation sites within the RNase domain15. Notably, this mutant had significantly reduced iNOS-dependent IRE1α S-nitrosylation (Fig. 3h, i), coupled with a lack of nitrosylation-dependent suppression of XBP1 splicing (Fig. 3j). Collectively, these results establish that iNOS governs the activity of IRE1α by increasing S-nitrosylation of IRE1α, leading to a reduction in XBP1s generation in cardiomyocytes.
Restoration of IRE1α–XBP1 axis ameliorates HFpEF
To confirm that iNOS-dependent suppression of the IRE1α–XBP1 axis contributes to the pathophysiology of HFpEF, Nos2 knockout mice (Extended Data Fig. 9a) were challenged with our HFpEF protocol (Extended Data Fig. 9b). Despite the absence of differences in LVEF (Extended Data Fig. 9c), body weight (Extended Data Fig. 9d), blood pressure (Extended Data Fig. 9e, f), glucose tolerance (Extended Data Fig. 9g, h) and cardiac hypertrophy (Extended Data Fig. 9i), significant attenuation of diastolic dysfunction (Fig. 4a, b), improvement in exercise tolerance (Fig. 4c) and reduced lung congestion (Fig. 4d) were observed in Nos2 knockout mice subjected to the HFD + l-NAME regimen. Under HFpEF conditions, iNOS gene ablation restored XBP1 splicing (Fig. 4e) and Xbp1s transcript levels (Fig. 4f), as well as IRE1α phosphorylation (Fig. 4g, h). Notably, depletion of iNOS diminished global myocardial protein nitrosylation (Extended Data Fig. 7f, g), significantly reducing S-nitrosylation of IRE1α (Fig. 3e, f). These data confirm the crucial role of iNOS in the pathogenesis of HFpEF and in the regulation of the IRE1α–XBP1 axis in the heart.
a, Ratio between mitral E wave and A wave. b, Ratio between mitral E wave and E′ wave. c, Running distance during exercise exhaustion test. d, Ratio between wet and dry lung weight of wild-type and Nos2 knockout mice after five weeks of chow or HFD + l-NAME diet. a, b, n = 10 mice per group; c, d, n = 5 mice per group. e, Electrophoretic analysis of spliced and unspliced Xbp1 transcripts in left ventricular samples of wild-type and Nos2 knockout mice after five weeks of chow or HFD + l-NAME diet. Tunicamycin-treated NRVMs were used as positive control. Blots are representative of three independently performed experiments with similar results. f, g, mRNA levels of Xbp1s (f; n = 5 mice per group) and immunoblots of p-IRE1α, IRE1α and GAPDH (g) in left ventricular samples of wild-type and Nos2 knockout mice after five weeks of chow or HFD + l-NAME diet. n = 4 mice per group. h, Densitometric analysis of the ratio of p-IRE1α/IRE1α. n = 4 mice per group. Data are mean ± s.e.m. a–d, f, h, Two-way ANOVA followed by Sidak’s multiple-comparisons test. Numbers above square brackets show significant P values. For gel source data, see Supplementary Fig. 1.
To test whether pharmacological inhibition of iNOS ameliorates cardiac relaxation and exercise intolerance in HFpEF, we exposed mice fed a HFD + l-NAME diet for five weeks to l-N6-(1-iminoethyl)lysine (l-NIL), a specific inhibitor of iNOS17 (Extended Data Fig. 10a). After three days of l-NIL treatment, the iNOS inhibitor decreased NO metabolites in urine of mice exposed to HFD + l-NAME compared with mice treated with vehicle (Extended Data Fig. 10b). In the absence of significant changes in the percentage of LVEF (Extended Data Fig. 10c) and blood pressure (Extended Data Fig. 10d, e), l-NIL administration partially improved ventricular relaxation (Extended Data Fig. 10f, g) and exercise performance (Extended Data Fig. 10h) in HFD + l-NAME-treated mice. Notably, these observations in HFD + l-NAME-fed mice treated with l-NIL occurred in the absence of concomitant restoration of Xbp1s transcript levels (Extended Data Fig. 10i). In contrast to Nos2 deletion—in which long-term reduction in the activity of iNOS prevents the aberrant S-nitrosylation of IRE1α, restoring XBP1s levels in the myocardium—the amelioration of left ventricular diastolic properties and exercise tolerance induced by short-term iNOS inhibition appears to be independent of myocardial XBP1s levels, suggesting additional beneficial mechanisms of iNOS inhibition on cardiac relaxation.
Discussion
HFpEF is a lethal disorder for which there are no effective clinical therapies. Given the limitations of current preclinical models, we developed a mouse model of HFpEF that recapitulates the vast majority of the clinical features of HFpEF. The interventions that we used to replicate the clinical phenotype are based on the (limited) pathophysiological observations that are available from the human condition; alterations in NO pathway(s) have been repeatedly observed in human HFpEF hearts, and our model takes advantage of this fact to manipulate NO biology from the start. By contrast, only limited evidence is available to implicate the renin–angiotensin–aldosterone system in HFpEF, which may explain why preclinical models based on manipulation of the renin–angiotensin–aldosterone system elicited only a limited number of HFpEF features, and therapies that target this system have failed in clinical trials. Using multiple approaches in preclinical models of HFpEF and human myocardial samples from patients with heart failure, we uncovered evidence that implicates systemic inflammation, iNOS activation, nitrosative stress and suppression of the IRE1α–XBP1 axis of the UPR in disease pathophysiology. Indeed, our findings suggest that metabolic inflammation and its master mediator, iNOS, are critical elements of the pathophysiology of HFpEF.
Current treatments of HFpEF that have focused on increasing NO bioavailability have reported neutral18,19,20 or negative21,22 results. Our findings provide a biological explanation for the failure of NO-inducing approaches as therapeutic strategies. Therefore, turning attention to strategies that focus on reducing the activation of iNOS and other pro-inflammatory mediators, as well as to molecular mediators of nitrosative stress (such as components of the iNOS–S-nitrosylase complex)23, may hold promise for the treatment of HFpEF.
Working from the realities of clinical HFpEF, including multi-organ comorbidities, we developed a mouse model of HFpEF that recapitulates the myriad features of HFpEF. Further, we show that iNOS-dependent suppression of the IRE1α–XBP1 axis is a pathophysiological driver of this syndrome.
Methods
Experimental animals
All experiments involving animals conformed to the Guide for the Care and Use of Laboratory Animals published by the US National Institutes of Health (NIH Publication eighth edition, update 2011) and were approved by the Institutional Animal Care and Use Committee of the University of Texas Southwestern Medical Center. The studies were in compliance with all ethical regulations. C57BL/6N mice were used for wild-type studies. Tetracycline-responsive elements (TRE)-XBP1s mice were crossed with mice containing the tetracycline transactivator (tTA) transcription factor driven by the α-myosin heavy chain promoter (αMHC-tTA) to generate mice with cardiomyocyte-specific inducible overexpression of XBP1s (XBP1s transgenic) as previously described10. Nos2 knockout mice (Nos2−/−, B6.129P2-Nos2tm1Lau/J) were purchased from Jackson Laboratory to establish an in-house colony. ZSF1-obese (ZSF1-LeprfaLeprcp/Crl, strain 378) and WKY rats were obtained from Charles River Laboratories. Male adult (8- to 12-week-old) mice were used in the experiments. Analyses in rats were carried out when the animals reached 20 weeks of age. Mice and rats were maintained on a 12-h light/dark cycle from 06:00 to 18:00 and had unrestricted access to food (2916, Teklad for chow groups and D12492, Research Diet for the HFD groups) and water. l-NAME (0.5 g l−1, Sigma-Aldrich) was supplied in the drinking water for the indicated periods of time, after adjusting the pH to 7.4. l-NIL (Cayman Chemicals) was administered intraperitoneally at a dose of 80 mg kg−1 body weight twice a day for 3 days. Transverse aortic constriction (TAC) or severe TAC (sTAC) was surgically induced as previously described24.
Conventional echocardiography and Doppler imaging
Transthoracic echocardiography was performed using a VisualSonics Vevo 2100 system equipped with MS400 transducer (Visual Sonics). LVEF and other indices of systolic function were obtained from short-axis M-mode scans at the midventricular level, as indicated by the presence of papillary muscles, in conscious, gently restrained mice. Apical four-chamber views were obtained in anaesthetized mice for diastolic function measurements using pulsed-wave and tissue Doppler imaging at the level of the mitral valve. Anaesthesia was induced by 5% isoflurane and confirmed by lack of response to firm pressure on one of the hindpaws. During echocardiogram acquisition—under body-temperature-controlled conditions—isofluorane was reduced to 1.0–1.5% and adjusted to maintain a heart rate in the range of 415–460 beats per min. Parameters collected include: heart rate, left ventricular end-diastolic diameter, left ventricular end-systolic diameter, end-diastolic interventricular septal wall thickness, left ventricular end-diastolic posterior wall, left ventricular fractional shortening, LVEF, peak Doppler blood inflow velocity across the mitral valve during early diastole, peak Doppler blood inflow velocity across the mitral valve during late diastole, isovolumic relaxation time, peak tissue Doppler of myocardial relaxation velocity at the mitral valve annulus during early diastole and early filling deceleration time. At the end of the procedures all mice recovered from anaesthesia without difficulties. All parameters were measured at least three times, and means are presented.
Speckle-tracking echocardiography and strain analysis
B-mode traces acquired from the parasternal long-axis view were used to calculate global strain in longitudinal dimensions using VevoStrain software (Visual Sonics) and a speckle-tracking algorithm. Velocity and displacement were also calculated in both the long and short axes. Values generated by strain analysis in the longitudinal dimension are negative, indicative of fibre shortening. B-mode images were selected based on quality (high frame rates) and on the ability to visualize both the endocardial and epicardial left ventricular wall borders. Borders of the endocardium and epicardium were traced, and semi-automated strain analysis was performed. Average peak global strain values were obtained from six independent anatomical segments of the left ventricle.
Coronary flow reserve
Coronary flow velocity was measured in cine traces of the left proximal coronary artery using pulsed-wave Doppler at baseline and under hyperaemic conditions induced by inhalation of 1.5% and 3.0% isoflurane, respectively. Coronary flow reserve is expressed as the ratio of peak blood flow velocity during hyperaemia and peak blood flow velocity at baseline.
Tail-cuff blood pressure recordings
Systolic blood pressure was measured noninvasively in conscious mice using the tail-cuff method and a CODA instrument (Kent Scientific). Mice were placed in individual holders on a temperature-controlled platform (37 °C) and recordings were performed under steady-state conditions. Before testing, all mice were trained to become accustomed to short-term restraint. Blood pressure was recorded for at least four consecutive days and readings were averaged from at least eight measurements per session.
Pulse-wave velocity
Pulse-wave velocity (PWV) was measured noninvasively over the entire aorta. Pulsed Doppler images were obtained at the ascending aorta (immediately cranial to the aortic valve) and the abdominal aorta (immediately cranial to the iliac bifurcation). Subsequently, the distance between the two locations was approximated by an external body tape measurement. PWV was calculated as the ratio of ΔX and ΔT (PWV = ΔX/ΔT), in which ΔX is the distance between the two anatomical locations at which the Doppler signals were recorded and ΔT is the electrocardiography-based transit time between averaged waveforms at both locations.
Pressure–volume analysis
An apical approach was used to obtain invasive pressure–volume measurements from anaesthetized (pentobarbital sodium, 40–50 mg kg−1) mice using a mouse 1.4-F catheter with pressure and conductance sensors (SPR 839, Millar Instruments) as previously described25. The inferior vena cava was located and occluded during a pause in ventilation to acquire load-independent indices. Data were analysed using PV Laboratory Chart software (version 8).
Exercise exhaustion test
After three days of acclimatization to treadmill exercise, an exhaustion test was performed in the experimental groups of mice. Mice ran uphill (20°) on the treadmill (Columbus Instruments) starting at a warm-up speed of 5 m min−1 for 4 min after which speed was increased to 14 m min−1 for 2 min. Every subsequent 2 min, the speed was increased by 2 m min−1 until the mouse was exhausted. Exhaustion was defined as the inability of the mouse to return to running within 10 s of direct contact with an electric-stimulus grid. Running time was measured and running distance calculated.
Grip-strength test
Muscle strength was assessed using a grip-strength test performed by the Neuro-Models Core Facility at UT Southwestern Medical Center. Mice from different experimental groups were horizontally drawn along a straight line (to avoid the effect of body-weight differences) and allowed to grasp a pull-bar assembly connected to the grip-strength meter (Columbus Instruments). Mice were then gently pulled by the tail away from the sensor until the grasp was broken, and the peak force (g) was recorded. The test was considered complete when six trials per limb pair (five forelimb and five hindlimb) were recorded. All measurements were performed by the operator in a blinded fashion.
Intraperitoneal glucose-tolerance test
Intraperitoneal glucose-tolerance tests were performed by injection of glucose (2 g kg−1 in saline) after 6-h fasting. Tail blood glucose levels (mg dl−1) were measured with a glucometer before (0 min) and at 15, 30, 45, 60 and 120 min after glucose administration.
Histology
Hearts and skeletal muscles (soleus and gastrocnemius/plantaris) were collected and either flash-frozen in embedding medium, which contained a 3:1 mixture of Tissue Freezing Medium (Triangle Biomedical Sciences) and gum tragacanth (Sigma-Aldrich), or fixed in 4% paraformaldehyde overnight and processed for routine paraffin histology (5-μm sections stained with haematoxylin and eosin, Masson’s trichrome or picrosirius red). Frozen sections were cut on a cryotome and stained for ATPase activity as previously described26. Wheat germ agglutinin (WGA) staining and lectin staining were used to measure the cross-sectional area of cardiomyocytes and to quantify capillary density, respectively. After deparaffinization and antigen retrieval with hot citrate buffer (Antigen Retrival Citra, BioGenex), slides were incubated with WGA conjugated to Alexa Fluor 488 (50 mg ml−1, 1 h, room temperature) or with biotin-conjugated lectin from Griffonia simplicifolia (overnight, 4 °C). Cardiomyocyte size, fibrosis area and capillary numbers were visualized with a Leica DM2000 upright photomicroscope and quantified using ImageJ software version 2.0 (three microscopic fields per heart). For immunofluorescence studies, cardiac sections were permeabilized with 0.3% Triton X-100 in phosphate-buffered saline (PBS), blocked with 5% goat serum in PBS and then incubated with KDEL (ADI-SPA-827-D, Enzo Life Sciences) antibody 1:200 in 3% bovine serum albumin (BSA) in PBS overnight at 4 °C. After washing, sections were incubated with the anti-mouse secondary antibody conjugated to Alexa Fluor 488 (1:500 in 3% BSA in PBS, 1 h at room temperature). Sections were then mounted using anti-fade reagent supplemented with DAPI (Invitrogen) and images were obtained using a Zeiss LSM 880 upright confocal fluorescence microscope.
Isolated muscle preparation and stimulation
Muscle preparation was accomplished as previously described27. In brief, soleus muscles were isolated surgically from mice fed chow or HFD + l-NAME and mounted on Grass FT03.C force transducers connected to a Powerlab 8/SP data acquisition unit (AD Instruments), bathed in physiological salt solution at 37 °C and perfused continuously with 95% O2 and 5% CO2. After calibration, muscles were adjusted to their initial lengths at a point in which the passive force is 0.5g and were stimulated with two platinum wire electrodes to establish the optimal length. Muscles were then stimulated at 150 Hz in 2-s bursts until the maximal tetanic contraction was achieved. Stress (mN mm−2) was calculated to normalize contraction responses to tissue cross-sectional areas. For evaluation of relaxation, muscles were stimulated at 150 Hz for 40 s. Values were expressed as the percentage of the force during the protocol relative to peak force.
Cardiomyocyte isolation and treatment
AMVMs were isolated from littermates of each experimental group. In brief, hearts were isolated and immediately digested through retrograde perfusion with perfusion buffer (113 mM NaCl, 4.7 mM KCl, 0.6 mM KH2PO4, 0.6 mM Na2HPO4, 1.2 mM MgSO4, 10 mM HEPES, 12 mM NaHCO3, 10 mM KHC03, 30 mM taurine, 10 mM BDM and 5.5 mM glucose) for 1 min followed by buffer containing liberase TM (0.025 mg ml−1, Roche) and trypsin (0.025%) for 14 min. The left ventricle was dissected and minced in perfusion medium supplemented with 10% dialysed fetal bovine serum. After filtration, AMVMs were allowed to sediment by gravity and non-cardiomyocytes were discarded. Calcium reintroduction was performed stepwise from 0 to 1.8 mM in six steps. NRVMs were isolated from 1–2-day-old Sprague Dawley rat pups as previously described10. Cell preparations typically contained greater than 95% cardiomyocytes. When appropriate, tunicamycin from Streptomyces sp. (Sigma-Aldrich, 1 μg ml−1 for 24 h) was used.
Sarcomere length measurements in paced cardiomyocytes
Sarcomere shortening and relaxation were measured in freshly isolated AMVMs using the integrated IonOptix LLC contractility/photometry system. In brief, AMVMs were electrically stimulated at 0.5 Hz using a field stimulator and changes in sarcomere length were recorded. Basal and peak sarcomere length, maximum departure/return velocities and time to peak were measured. All measurements were performed at room temperature.
Adenovirus production, purification and transduction
Mouse Nos2 cDNA was obtained from Addgene (19295) and subcloned into an adenovirus expression vector using Adeno-X Adenoviral System 3 (CMV promoter and Zsgreen1 reporter, Takara) following the manufacturer’s instructions. Adeno-X 293 cell line was purchased from Clontech (Adeno-X 293 Cell Line, 632271). The cell line was authenticated by the vendor and mycoplasma detection tested negative. Adenoviruses were obtained from culture supernatant and cell lysates and the viral titre was determined. NRVMs were transduced with increasing multiplicities of infection (MOIs) of virus and cells were collected 24 h after infection. An AdLacZ construct contained in the kit was used as control at the same MOI. Adenoviruses carrying wild-type human ERN1 (which encodes IRE1α) and ERN1 with mutations in two target nitrosylation sites in the RNase domain (ERN1 M1+M2) were donated by G. S. Hotamisligil and L. Yang.
RNA isolation and qPCR
Total RNA was extracted from mouse and rat hearts, AMVMs or NRVMs using TRIzol reagent or Quick-RNA MicroPrep kit (Zymo Reaserch). A total of 500 ng RNA was used for reverse transcription using iScript reagent (Bio-Rad). qPCR reactions were performed in triplicate with SYBR master mix (Bio-Rad). For human hearts, total RNA was extracted using the miRNeasy Mini Kit (Qiagen) then reverse-transcribed into cDNA using the High Capacity cDNA Reverse Transcription Kit (Applied Biosystems). Real-time PCR was performed in duplicate using TaqMan Gene Expression Assay probes with specific primers for target sequences. The 2−ΔΔCt relative quantification method, using 18S or GAPDH for normalization, was used to estimate the amount of target mRNA in samples, and fold ratios were calculated relative to mRNA expressions levels from control samples. The following PCR primers sequences were used, including forward, reverse sequences for each gene, respectively:
Xbp1s mouse/rat GGTCTGCTGAGTCCGCAGCAGG, GAAAGGGAGGCTGGTAAGGAAC; Bip mouse CATGGTTCTCACTAAAATGAAAGG, GCTGGTACAGTAACAACTG; Atf6 mouse AGAAAGCCCGCATTCTCCAG, ACTCCCAGAATTCCTACTGATGC; Atf4 mouse ATGGCCGGCTATGGATGAT, CGAAGTCAAACTCTTTCAGATCCATT; Chop mouse CTGCCTTTCACCTTGGAGAC, CGTTTCCTGGGGATGAGATA; Nos2 mouse GTTCTCAGCCCAACAATACAAGA, GTGGACGGGTCGATGTCAC; Nos3 mouse TCAGCCATCACAGTGTTCCC, ATAGCCCGCATAGCGTATCAG; Nppa mouse CTGGGACCCCTCCGATAGAT, TTCGGTACCGGAAGCTGTTG; Nppb mouse TTTGGGCTGTAACGCACTGAA, TGTGGCAAGTTTGTGCTCCA; Myhc isoform 1 mouse CCTTGGCACCAATGTCCCGGCTC, GAAGCGCAATGCAGAGTCGGTG; Myhc isoform 2A mouse ATGAGCTCCGACGCCGAG, TCTGTTAGCATGAACTGGTAGGCG; Myhc isoform 2X mouse AAGGAGCAGGACACCAGCGCCCA, ATCTCTTTGGTCACTTTCCTGCT; Tnf mouse ATCGGTCCCCAAAGGGATGA, GGTGGTTTGCTACGACGTG; Il1b mouse TAACTGCACCCACTTCCCAG, AGGCTTGGCAACCCAAGTAA; Il6 mouse CTCTGCAAGAGACTTCCATCCA, GACAGGTCTGTTGGGAGTGG; 18S mouse/rat AAACGGCTACCACATCCAAG, CCTCCAATGGATCCTCGTTA; XBP1s human CTGAGTCCGCAGCAGGTG, GTCCAGAATGCCCAACAGGA; XBP1u human CGGAAGCCAAGGGGAATGAA, TGTTCTGGAGGGGTGACAAC; NOS2 human TCCCGAAGTTCTCAAGGCAC, TTCTTCACTGTGGGGCTTGC; GAPDH human TCAACGACCACTTTGTCAAGCTCA, GCTGGTGGTCCAGGGGTCTTACT.
Electrophoretic analysis of XBP1 splicing
XBP1 splicing was assessed as previously described28. In brief, cDNAs from mouse and rat hearts were subjected to semiquantitative PCR using the following primers (5′-CCTTGTGGTTGAGAACCAGG-3′ and 5′-CTAGAGGCTTGGTGTATAC-3′) to amplify a 451-base pair (bp) Xbp1u fragment or a 425-bp Xbp1s fragment, respectively. PCR products were then digested using PstI restriction enzyme, which only cleaves the intron included in the unspliced Xbp1 cDNA to yield a 154-bp and a 297-bp fragment, leaving the spliced Xbp1 cDNA intact. The 18S transcript was also amplified as internal control and the PstI-digested products were analysed by 2% agarose gel electrophoresis.
Immunoblot analysis
Protein extracts from frozen mouse, rat and human hearts, AMVMs and NRVMs were prepared by lysis in ice-cold modified RIPA buffer (150 mM NaCl, 50 mM Tris HCL pH 7.4, 1% Triton X-100, 0.5% sodium deoxycholate, 0.1% SDS, 5 mM EDTA and 2 mM EDTA) containing protease and phosphatase inhibitors. Proteins were separated by sodium dodecyl sulfate polyacrylamide gel electrophoresis (SDS–PAGE) on 4–20% gradient gels (Bio-Rad) and transferred to nitrocellulose membranes. An Odyssey scanner (LI-COR version 3.0) was used as detection system. For human myocardial samples, protein lysates were prepared using cell lysis buffer (Cell Signaling Technology) with the addition of 1 mM phenylmethylsulfonyl fluoride. Lysates were prepared using the Retsch Mixer Mill. Capillary-based immunoassay was performed using the Wes-Simple Western method with the anti-rabbit detection module (ProteinSimple). Protein expression was measured by chemiluminescence and quantified as area under the curve using the Compass for Simple Western program (ProteinSimple). Proteins were detected with the following primary antibodies: p-IRE1α Ser724 (NB100-2323, Novus Biological); IRE1α (3294, Cell Signaling Technology), GRP78 (3177, Cell Signaling Technology); GRP94 (2104, Cell Signaling Technology); PERK (3192, Cell Signaling Technology); ATF6 (24169-1-AP, Proteintech); iNOS (13120, Cell Signaling Technology); nNOS (4234, Cell Signaling Technology); eNOS (9572, Cell Signaling Technology); and GAPDH (10R-G109a, Fitzgerald).
Cytokine/chemokine analyses using antibody array membranes
RayBio C-Series mouse cytokine antibody array C1 (AAM-CYT-1-8) was used according to the manufacturer’s instructions. In brief, the membranes were placed in the plastic tray provided in the kit, and 200 μl of plasma from mice fed with chow or HFD + l-NAME was incubated overnight in the designated well at 4 °C. Subsequently, the samples were aspirated, the membranes were washed with wash buffer and then incubated with 1 ml of biotinylated antibody cocktail overnight at 4 °C. After washing, membranes were incubated with HRP–streptavidin overnight at 4 °C. Detection buffer was used to develop a chemiluminescent signal and the LI-COR instrument was used to detect signal intensity.
Measurement of nitrite and nitrate
The amounts of nitrite and nitrate, the breakdown products of nitric oxide, were measured in urine samples or cell medium using the nitrate/nitrite colorimetric assay kit (Cayman Chemicals) according to the manufacturer’s instructions.
Lactate dehydrogenase assay
To evaluate cell survival, lactate dehydrogenase (LDH) release was quantified using the CytoTox96 cytotoxicity kit (Promega) according to the manufacturer’s instructions. Each experiments was performed in three biological replicates each time in triplicate. LDH release was calculated as follows: (medium LDH)/(medium LDH + intracellular LDH).
Determination of S-nitrosylation
Hearts were excised, snap-frozen in liquid nitrogen and stored at −80 °C until homogenization. Cardiac tissue or NRVM samples were homogenized in lysis buffer (PBS supplemented with 10% glycerol, 1 mM EDTA, 1 mM EGTA, 1% Triton X-100, protease/phosphatase inhibitors and 0.1 mM neocuproine). Lysates were spun at 10,000g and supernatants were used for protein determination using the BCA method (Thermo Fisher Scientific). Free cysteines were blocked at room temperature in HENS buffer (100 mM HEPES (pH 7.0), 1 mM EDTA, 0.1 mM neocuproine and 2.5% SDS) using 100 mM N-ethylmaleimide (NEM). Proteins were acetone-precipitated and pellets were washed using methanol. The protein pellet was resuspended in modified HENS buffer (100 mM HEPES, 1% SDS, 1 mM EDTA and 0.1 mM neocuproine) pH 8.0 and labelled using 40 mM ascorbic acid and 0.4 mM Tandem Mass Tags (Thermo Fisher Scientific). A negative control without ascorbic acid and a positive control with 1 mM GSNO were included. Immunoprecipitation was carried out using 500 μg labelled protein, 5 μl IRE1α antibody and 25 μl Dynabeads protein G in immunoprecipitation buffer (overnight, 4 °C). Beads were magnetically separated and washed five times with immunoprecipitation buffer and then eluted using a 2× Laemmli buffer. For some experiments, S-nitrosylated proteins were separated using resin-assisted capture protocols. Total lysates (2 mg) were blocked twice with 100 mM NEM in HENS buffer pH 7.0 at 55 °C for 30 min to ensure complete cysteine blockage. Proteins were acetone-precipitated and washed with methanol to remove unreacted NEM. Then, samples were incubated with freshly prepared 50 mM ascorbate and 55 mg thiopropyl-sepharose and rotated end-over-end in the dark for 4 h. The bound SNO proteins were washed with modified HENS (1% SDS) buffer five times; SNO proteins were then eluted with Laemmli sample buffer supplemented with 5% β-mercaptoethanol and analysed by SDS–PAGE and western blot.
Human myocardial tissue
Human myocardial tissue samples were collected under protocols approved by Institutional Review Board (IRB) at Johns Hopkins University and consent for biopsy procedures or use of explanted tissues prospectively obtained in all cases. Patients with HFpEF were referred for cardiac catheterization and right ventricular endomyocardial biopsy owing to clinical suspicion of infiltrative cardiomyopathy. HFrEF samples were obtained from the right ventricular septum of explanted dilated failing hearts before orthotopic heart transplantation. Control samples were obtained from the right ventricular septum of nonfailing, explanted, unused donor hearts. Control and HFrEF samples were provided by collaborators at the Perelman School of Medicine at the University of Pennsylvania through an IRB-approved protocol.
Statistical analysis
Data are mean ± s.e.m. Differences were analysed by two-tailed unpaired Student’s t-test for experiments with two groups and one-way or two-way ANOVA plus Sidak’s post hoc test for multiple comparisons as appropriate in experiments including ≥3 groups. A value of P < 0.05 was considered statistically significant. All experiments were performed with at least three biological replicates. Statistical analyses were conducted using GraphPad Prism software 8.0 or SPSS Statistics version 19 (SPSS) for the comparison of categorical variables in Extended Data Table 2. No statistical analysis was used to predetermine sample sizes; estimates were made based on our previous experience, experimental approaches, availability and feasibility required to obtain statistically significant results. Experimental mice were randomly assigned to each experimental or control group. Investigators were blinded to the genotypes of the individual animals during the experiments and outcome assessments.
Reporting summary
Further information on research design is available in the Nature Research Reporting Summary linked to this paper.
Data availability
The authors declare that the data supporting the findings of this study are available within the paper and its Supplementary Information. Each data point corresponding to figures that describe the results from in vivo or in vitro model studies are provided as separate Source Data for Figs. 1c, d, f–i, 2a, d, f, g, i–m, 3a–d, f, g, i, j, 4a–d, f, h and Extended Data Figs. 1a–l, 2b–f, 3b–e, g, 4a–l, n, p, 5b–g, 6b–g, j–m, 7b, d, g, 8b–d, g, 9c–i, 10b–i. Other source data related to the study are available from the corresponding author upon reasonable request.
References
Dunlay, S. M., Roger, V. L. & Redfield, M. M. Epidemiology of heart failure with preserved ejection fraction. Nat. Rev. Cardiol. 14, 591–602 (2017).
Shah, S. J. et al. Phenotype-specific treatment of heart failure with preserved ejection fraction: a multiorgan roadmap. Circulation 134, 73–90 (2016).
Chirinos, J. A. & Zamani, P. The nitrate–nitrite–NO pathway and its implications for heart failure and preserved ejection fraction. Curr. Heart Fail. Rep. 13, 47–59 (2016).
Butler, J., Braunwald, E. & Gheorghiade, M. Recognizing worsening chronic heart failure as an entity and an end point in clinical trials. J. Am. Med. Assoc. 312, 789–790 (2014).
Roh, J., Houstis, N. & Rosenzweig, A. Why don’t we have proven treatments for HFpEF? Circ. Res. 120, 1243–1245 (2017).
Primessnig, U. et al. Novel pathomechanisms of cardiomyocyte dysfunction in a model of heart failure with preserved ejection fraction. Eur. J. Heart Fail. 18, 987–997 (2016).
Methawasin, M. et al. Experimentally increasing the compliance of titin through RNA binding motif-20 (RBM20) inhibition improves diastolic function in a mouse model of heart failure with preserved ejection fraction. Circulation 134, 1085–1099 (2016).
González-López, E. et al. Wild-type transthyretin amyloidosis as a cause of heart failure with preserved ejection fraction. Eur. Heart J. 36, 2585–2594 (2015).
Wang, Z. V. & Hill, J. A. Protein quality control and metabolism: bidirectional control in the heart. Cell Metab. 21, 215–226 (2015).
Wang, Z. V. et al. Spliced X-box binding protein 1 couples the unfolded protein response to hexosamine biosynthetic pathway. Cell 156, 1179–1192 (2014).
Rothermel, B. A. et al. Differential activation of stress-response signaling in load-induced cardiac hypertrophy and failure. Physiol. Genomics 23, 18–27 (2005).
Paulus, W. J. & Tschöpe, C. A novel paradigm for heart failure with preserved ejection fraction: comorbidities drive myocardial dysfunction and remodeling through coronary microvascular endothelial inflammation. J. Am. Coll. Cardiol. 62, 263–271 (2013).
Furfine, E. S., Harmon, M. F., Paith, J. E. & Garvey, E. P. Selective inhibition of constitutive nitric oxide synthase by l-N G-nitroarginine. Biochemistry 32, 8512–8517 (1993).
Kopincová, J., Púzserová, A. & Bernátová, I. l-NAME in the cardiovascular system — nitric oxide synthase activator? Pharmacol. Rep. 64, 511–520 (2012).
Yang, L. et al. S-Nitrosylation links obesity-associated inflammation to endoplasmic reticulum dysfunction. Science 349, 500–506 (2015).
Gould, N., Doulias, P. T., Tenopoulou, M., Raju, K. & Ischiropoulos, H. Regulation of protein function and signaling by reversible cysteine S-nitrosylation. J. Biol. Chem. 288, 26473–26479 (2013).
Moore, W. M. et al. l-N 6-(1-Iminoethyl)lysine: a selective inhibitor of inducible nitric oxide synthase. J. Med. Chem. 37, 3886–3888 (1994).
Borlaug, B. A., Koepp, K. E. & Melenovsky, V. Sodium nitrite improves exercise hemodynamics and ventricular performance in heart failure with preserved ejection fraction. J. Am. Coll. Cardiol. 66, 1672–1682 (2015).
Sharma, K. & Kass, D. A. Heart failure with preserved ejection fraction: mechanisms, clinical features, and therapies. Circ. Res. 115, 79–96 (2014).
Redfield, M. M. et al. isosorbide mononitrate in heart failure with preserved ejection fraction. N. Engl. J. Med. 373, 2314–2324 (2015).
Zamani, P. et al. Isosorbide dinitrate, with or without hydralazine, does not reduce wave reflections, left ventricular hypertrophy, or myocardial fibrosis in patients with heart failure with preserved ejection fraction. J. Am. Heart Assoc. 6, e004262 (2017).
Oeser, C. Heart failure: nitrates reduce activity levels in HFpEF. Nat. Rev. Cardiol. 13, 2–3 (2016).
Jia, J. et al. Target-selective protein S-nitrosylation by sequence motif recognition. Cell 159, 623–634 (2014).
Zhu, H. et al. Cardiac autophagy is a maladaptive response to hemodynamic stress. J. Clin. Invest. 117, 1782–1793 (2007).
Kong, Y. et al. Suppression of class I and II histone deacetylases blunts pressure-overload cardiac hypertrophy. Circulation 113, 2579–2588 (2006).
Oh, M. et al. Calcineurin is necessary for the maintenance but not embryonic development of slow muscle fibers. Mol. Cell. Biol. 25, 6629–6638 (2005).
Zhi, G. et al. Myosin light chain kinase and myosin phosphorylation effect frequency-dependent potentiation of skeletal muscle contraction. Proc. Natl Acad. Sci. USA 102, 17519–17524 (2005).
Wang, J. M. et al. IRE1α prevents hepatic steatosis by processing and promoting the degradation of select microRNAs. Sci. Signal. 11, eaao4617 (2018).
Acknowledgements
This work was supported by grants from NIH: HL-120732 (J.A.H.), HL-128215 (J.A.H.), HL-126012 (J.A.H.), F32HL136151 (K.M.F.), F32HL142244 (D.T.), 2T32HL007227-41 (V.S.H.), HL-135827 (D.A.K.), HL-119012 (D.A.K.), R01-HL102478 (P.P.A.M.), the American Heart Association (AHA): 16POST30680016 (F.A.), 16PRE29660003 (S.Y.K.), 14SFRN20510023 (J.A.H.), 14SFRN20670003 (J.A.H.) and 16SFRN28620000 (K.S. and D.A.K.), AHA and the Theodore and Beulah Beasley Foundation 18POST34060230 (G.G.S.), University Federico II of Naples and Compagnia di San Paolo STAR program (G.G.S.), Fondation Leducq TransAtlantic Network of Excellence11CVD04 (J.A.H.), Cancer Prevention and Research Institute of Texas RP110486P3 (J.A.H.), Senator Paul D. Wellstone Muscular Dystrophy Cooperative Research Center U54-HD087351 (P.P.A.M.) and by Fondo Nacional de Desarrollo Científico y Tecnológico, FONDAP 15130011 (S.L.). We thank G. S. Hotamisligil and L. Yang for providing ERN1 wild-type and ERN1 M1+M2-expressing adenoviruses. We thank A. Rosenberg for his help with the ProteinSimple capillary immunoassay analysis (partly supported by The BioTechne Grant Foundation).
Author information
Authors and Affiliations
Contributions
G.G.S., F.A., D.T., K.M.F., E.V., S.Y.K. and T.M.H. performed the experiments. G.G.S. designed the experiments, performed the analyses and wrote the manuscript. F.A. designed the nitrosylation experiments, performed the analysis, isolated the adult mouse ventricular myocytes and performed the contractility studies. X.L. isolated the NRVMs. N.J. managed mouse colonies. H.I.M. performed the PV loop experiments and mouse surgeries. Z.V.W. provided the XBP1s transgenic mice and XBP1s adenovirus. P.P.A.M. and J.H. performed the skeletal muscle experiments, D.I.L., V.S.H., K.S. and D.A.K. provided human data. S.L. and T.G.G. contributed to the experimental design and manuscript preparation. J.A.H. conceived the project and contributed to manuscript preparation.
Corresponding author
Ethics declarations
Competing interests
G.G.S., T.G.G. and J.A.H. are co-inventors on a patent application (PCT/US/2017/037019) that was filed in June 2017 (provisional application filed in June 2016). The patent relates to the diet used for modeling HFpEF.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Systemic and cardiac phenotype of mice after 5 or 15 weeks of different dietary regimens.
a, Body weight of mice from different experimental groups after 5 or 15 weeks of diet (n = 10 mice per group for each time point). b, Intraperitoneal glucose-tolerance test (ipGTT) after 5 or 15 weeks of diet (5 weeks, n = 10 mice per group; 15 weeks, n = 5 mice per group). c, Area under the curve of the ipGTT experiment at 5 and 15 weeks (5 weeks, n = 10 mice per group; 15 weeks, n = 5 mice per group). d, e, Systolic blood pressure (SBP) (d) and diastolic blood pressure (DBP) (e) of different experimental groups after 5 or 15 weeks of treatment (n = 10 mice per group for each time point). f, Percentage of LVEF. g, Left ventricular global longitudinal strain. h, Ratio between mitral E wave and A wave. i, Ratio between mitral E wave and E′ wave. j, Ratio between wet and dry lung weight. k, Ratio between heart weight and tibia length. l, Runninng distance during exercise exhaustion test of mice after five weeks of diet. f, h–l, n = 10 mice per group. g, n = 5 mice per group. Data are mean ± s.e.m. a, c–l, One-way ANOVA followed by Sidak’s multiple-comparisons test; numbers above square brackets show significant P values. b, Two-way ANOVA followed by Sidak’s multiple-comparisons test; P values are as follows: 5 weeks 15 min, ***P = 0.0003 chow versus HFD, ***P = 0.0004 chow versus HFD + l-NAME; 30 min ***P = 0.0008 chow versus HFD, **P = 0.006 chow versus HFD + l-NAME; 45 min *P = 0.010 chow versus HFD, ***P = 0.0008 chow versus HFD + l-NAME; 60 min *P = 0.049 chow versus HFD, **P = 0.0096 chow versus HFD + l-NAME; 5 weeks 15 min **P = 0.008 chow versus HFD, ****P < 0.0001 chow versus HFD + l-NAME; 30 min **P = 0.005 chow versus HFD, ****P < 0.0001 chow versus HFD + l-NAME; 45 min **P = 0.009 chow versus HFD, ****P < 0.0001 chow versus HFD + l-NAME; 60 min *P = 0.028 chow versus HFD, **P = 0.0020 chow versus HFD + l-NAME.
Extended Data Fig. 2 Heart morphology and vascular characterization of mice after five weeks of different dietary regimens.
a, Representative images of haematoxylin and eosin (H&E), WGA, Masson’s trichrome (MT) and lectin staining in transversal sections of left ventricle of mice of different experimental groups. Images are representative of four independently performed experiments with similar results. Scale bars, 500 μm (haematoxylin and eosin) and 50 μm (WGA, Masson’s trichrome and lectin). b, WGA quantification of cardiomyocyte cross-sectional area (n = 4 mice per group). c, Percentage of fibrosis area in Masson’s trichrome-stained transversal sections (n = 4 mice per group). d, Myocardial capillary density (n = 4 mice per group). e, Aortic PWV of mice of different experimental groups (n = 5 mice per group). f, Representative pulsed-wave Doppler tracings of coronary flow in mice fed with chow (top) or HFD + l-NAME (bottom) under basal conditions (left; 1.5% isofluorane (Iso)) and after hyperaemic stimulus (right; 3% isofluorane). Images are representative of five independent mice. g, Coronary flow reserve (CFR) quantification (n = 5 mice per group). Data are mean ± s.e.m. b–d, e, g, One-way ANOVA followed by Sidak’s multiple-comparisons test. Numbers above square brackets show significant P values.
Extended Data Fig. 3 Histological and functional analyses of skeletal muscle in mice after five weeks of different dietary regimens.
a, Representative images of haematoxylin and eosin, WGA, metachromatic ATPase (ATPase), Masson’s trichrome and picrosirius red (PSR) staining of soleus and gastrocnemius/plantaris (G/P) muscles from mice fed with chow or HFD + l-NAME. Images are representative of three independently performed experiments with similar results. Scale bars, 50 μm. b, mRNA level of Myhc isoforms (MyHC-1, MyHC-2A or MyHC-2X) of soleus and gastrocnemius/plantaris muscles from mice fed with chow or HFD + l-NAME (n = 5 mice per group). c, Relaxation curve of isolated soleus muscles from mice fed with chow or HFD + l-NAME (n = 5 mice per chow group; n = 6 mice per HFD + l-NAME group). d, e, In vivo forelimb (d) and hindlimb (e) grip force measurements of mice from different experimental groups (n = 8 mice per chow group; n = 4 mice per HFD group; n = 3 mice per l-NAME group; n = 6 mice per HFD + l-NAME group). f, Maximal tetanic stresses in soleus muscles from chow and HFD + l-NAME mice (n = 5 mice per chow group; n = 6 mice per HFD + l-NAME group). Data are mean ± s.e.m. b, c, f, Two-tailed unpaired Student’s t-test (chow versus HFD + l-NAME soleus; chow versus HFD + l-NAME gastrocnemius/plantaris). d, e, One-way ANOVA followed by Sidak’s multiple-comparisons test. Numbers above square brackets show significant P values.
Extended Data Fig. 4 AMVM contractility after 5 weeks of different dietary regimens and functional characterization and analysis of the IRE1α–XBP1 axis in ZSF1-obese 20-week-old rats.
a, Baseline sarcomere length. n = 4 mice and n = 35 cells per chow group; n = 3 mice and n = 30 cells per HFD group; n = 3 mice and n = 30 cells per l-NAME group; n = 3 mice and n = 31 cells per HFD + l-NAME group. b, Time to peak. n = 4 mice and n = 36 cells per chow group; n = 3 mice and n = 30 cells per HFD group; n = 3 mice and n = 30 cells per l-NAME group; n = 3 mice and n = 31 cells per HFD + l-NAME group. c, Maximum return velocity. n = 4 mice and n = 36 cells per chow group; n = 3 mice and n = 29 cells per HFD group; n = 3 mice and n = 30 cells per l-NAME group; n = 3 mice and n = 31 cells per HFD + l-NAME group. d, Change (Δ) in sarcomere length compared to baseline. n = 4 mice and n = 36 cells per chow group; n = 3 mice and n = 30 cells per HFD group; n = 3 mice and n = 30 cells per l-NAME group; n = 3 mice and n = 31 cells per HFD + l-NAME group. e, Maximum departure velocity. n = 4 mice and n = 35 cells per chow group; n = 3 mice and n = 30 cells per HFD group; n = 3 mice and n = 29 cells per l-NAME group; n = 3 mice and n = 28 cells per HFD + l-NAME group. f, Representative tracings of cardiomyocyte contraction–relaxation during pacing. Each trace depicts one cell representative of the average for each experimental group. g–k, Measurements in 20-week old WKY and ZSF1-obese rats. g, Body weight. h, Percentage LVEF. i, Ratio between mitral E wave and A wave. j, Ratio between mitral E wave and E′ wave. k, Ratio between heart weight and tibia length. l, Lung weight to tibia length ratio. g–i, l, n = 5 rats per group; k, n = 5 rats per WKY group and n = 4 rats per ZSF1-obese group. m, Electrophoretic analysis of spliced and unspliced Xbp1 transcripts in left ventricular samples from WKY and ZSF1-obese rats. Tunicamycin-treated NRVMs were used as positive control. n = 3 rats per group. n, Xbp1s mRNA levels in the left ventricles of WKY and ZSF1-obese rats. n = 5 rats per group. o, Immunoblots of p-IRE1α, IRE1α and GAPDH from left ventricular samples of WKY and ZSF1-obese rats. n = 5 rats per group. p, Densitometric analysis of the ratio of p-IRE1α to IRE1α bands. n = 5 rats per group. Data are mean ± s.e.m. a–e, One-way ANOVA followed by Sidak’s multiple-comparisons test. g–l, n, p, Two-tailed unpaired Student’s t-test. Numbers above square brackets show significant P values. For gel source data, see Supplementary Fig. 1.
Extended Data Fig. 5 Functional characterization and UPR activation in mice after one week of sTAC.
a, Experimental design. C57BL/6N male mice were exposed to sham (control) or sTAC surgery and were evaluated after 1 week. b, Percentage of LVEF. c, Ratio between mitral E wave and A wave. d, Ratio between mitral E wave and E′ wave. e, Ratio between wet and dry lung weight. f, Ratio between heart weight and tibia length of sham control and sTAC mice. n = 5 mice per group. g, Xbp1s mRNA level in left ventricles of sham and sTAC mice. n = 4 mice in sham group; n = 5 mice in sTAC group. h, Immunoblots of GRP94, GRP78 and GAPDH in left ventricular samples of sham and sTAC mice. Data are representative of three independently performed experiments with similar results. Arrow indicates the regulated band. i, KDEL (Lys-Asp-Glu-Leu) sequence immunofluorescence staining in left ventricular sections of sham and sTAC mice. Hoechst, nuclei. Scale bars, 50 μm. Images are representative of three independently performed experiments with similar results. Data are mean ± s.e.m. b–g, Two-tailed unpaired Student’s t-test. Numbers above square brackets show significant P values. For gel source data, see Supplementary Fig. 1.
Extended Data Fig. 6 Functional characterization and analysis of the IRE1α–XBP1 axis in mice after TAC and phenotype of cardiomyocyte-restricted Xbp1s transgenic mice.
a, Experimental design. C57BL/6N mice were exposed to sham (control) or TAC surgery and followed up to five weeks. b–f, Analyses of different experimental groups of mice after sham or before (pre-TAC) and after TAC for one, three or five weeks. b, Ratio between mitral E wave and A wave. c, Ratio between mitral E wave and E′ wave. d, Percentage of LVEF. e, Ratio between heart weight and tibia length. f, Ratio between lung weight and body weight. b–d, n = 10 mice per group for pre-TAC, TAC 1 week and TAC 3 weeks groups; n = 5 mice for the TAC 5 weeks group; e, f, n = 10 mice per group for TAC 1 week and TAC 3 weeks groups; n = 5 mice per group for sham and TAC 5 week groups. g, Xbp1s mRNA levels in the left ventricles of sham-operated mice and mice with TAC for three weeks. n = 5 mice per group. h, Immunoblots of p-IRE1α, IRE1α and GAPDH from left ventricular samples of sham-operated mice and mice with TAC for three weeks. n = 3 mice per group. i, Experimental design. Control (CTR) and Xbp1s transgenic mice (TG) were fed with chow or HFD + l-NAME (green triangle). After five weeks, echocardiographic assessment was performed and doxycycline (Doxy) was removed from the drinking water to induce transgene expression (grey triangle). Two weeks after transgene induction (blue triangle), mice were subjected to functional analysis and tissue collection. j, Percentage of LVEF of different experimental cohorts over time. n = 5 mice per chow control and chow transgenic groups; n = 7 mice per HFD + l-NAME control and HFD + l-NAME transgenic groups. Each mouse was analysed at all three time points. k, Xbp1s mRNA level in left ventricles of control and Xbp1s transgenic mice fed with chow or HFD + l-NAME for seven weeks. n = 3 mice per group. l, Nppa and Nppb left ventricular mRNA levels. m, Ratio between heart weight and tibia length at the end of the study. n = 5 mice per chow control and chow transgenic groups; n = 7 mice per HFD + l-NAME control and HFD + l-NAME transgenic groups. Data are mean ± s.e.m. b–f, One-way ANOVA followed by Sidak’s multiple-comparisons test. g, Two-tailed unpaired Student’s t-test. j–m, Two-way ANOVA followed by Sidak’s multiple-comparisons test. Numbers above square brackets show significant P values. For gel source data, see Supplementary Fig. 1.
Extended Data Fig. 7 Myocardial nitrosative stress and inflammatory markers in mice after five weeks of different dietary regimens.
a, Top, cytokine/chemokine antibody array in plasma samples from chow and HFD + l-NAME-treated mice with visual estimation of differently abundant cytokines. Membranes are representative of two independently performed experiments with similar results. Bottom, list of cytokines and chemokines represented by the antibody array membrane. b, Tnf, Il1b and Il6 mRNA levels in left ventricles of mice fed with chow or HFD + l-NAME. n = 5 mice per group. c, iNOS and GAPDH in left ventricular samples of mice fed with chow or HFD + l-NAME. NRVMs infected with mouse AdiNOS (MOI = 100) were used as positive controls for iNOS bands. Arrows indicate the regulated bands. n = 3 mice per group. d, Nos3 (eNOS) mRNA levels in left ventricles from different experimental groups of mice. n = 4 mice per chow group; n = 6 mice per group for HFD, l-NAME and HFD + l-NAME groups. e, Immunoblots of nNOS, eNOS and GAPDH of different experimental groups of mice (n = 3 mice per group). f, Immunoblots of nitrosylated cysteines (Cys-SNO) and GAPDH in left ventricular samples of wild-type and Nos2 knockout (iNOS KO) mice after five weeks of chow or HFD + l-NAME diet. W/B, without blocking. n = 3 mice per group. g, Densitometric analysis of Cys-SNO:GAPDH ratios. n = 3 mice per group. Data are mean ± s.e.m. b, Two-tailed unpaired Student’s t-test. d One-way ANOVA followed by Sidak’s multiple-comparisons test. g, Two-way ANOVA followed by Sidak’s multiple-comparisons test. Numbers above square brackets show significant P values. For gel source data, see Supplementary Fig. 1.
Extended Data Fig. 8 iNOS overexpression in cardiomyocytes reduces IRE1α activation and XBP1s levels without affecting cardiomyocyte viability.
a, Immunoblots of iNOS and GAPDH proteins of NRVMs infected with increasing MOIs of AdLacZ or AdiNOS for 24 h. Blots are representative of three independently performed experiments with similar results. b, Nos2 (iNOS) mRNA levels in NRVMs transduced with increasing MOIs of AdLacZ or AdiNOS for 24 h. n = 4 biologically independent experiments. c, Nitrite/nitrate concentration in the medium from NRVMs transduced with increasing MOIs of AdLacZ or AdiNOS for 24 h. n = 4 biologically independent experiments. d, LDH release in NRVMs transduced with increasing MOIs of AdLacZ or AdiNOS for 24 h. n = 3 biologically independent experiments. e, Immunoblots of Cys-SNO, iNOS and GAPDH in NRVMs transduced with AdLacZ or AdiNOS for 24 h (MOI = 100). Blots are representative of three independently performed experiments with similar results. f, Immunoblots of p-IRE1α, IRE1α, iNOS and GAPDH in NRVMs transduced with increasing MOIs of AdLacZ or AdiNOS in the presence or absence of tunicamycin for 24 h. Blots are representative of three independently performed experiments with similar results. g, Xbp1s mRNA level of NRVMs transduced with MOI of 100 of AdLacZ or AdiNOS in the presence or absence of tunicamycin for 24 h. n = 3 biologically independent experiments. Data are mean ± s.e.m. b–d, g, One-way ANOVA followed by Sidak’s multiple-comparisons test. Numbers above square brackets show significant P values. For gel source data, see Supplementary Fig. 1.
Extended Data Fig. 9 Phenotype of Nos2 knockout mice after five weeks of different dietary regimens.
a, DNA genotyping of wild-type and Nos2 knockout mice. This signature was used for genotyping. b, Experimental design. Wild-type and Nos2 knockout mice were fed a chow or HFD + l-NAME diet for five weeks. Subsequently, mice were subjected to functional analysis and tissues were collected. c, Percentage of LVEF. d, Body weight.e, Systolic blood pressure. f, Diastolic blood pressure. g, Intraperitoneal glucose-tolerance test of different experimental groups of mice. c, n = 10 mice per group; d–g, n = 5 mice per group. h, Area under the curve of the intraperitoneal glucose-tolerance tests. n = 5 mice per group. i, Ratio between heart weight and tibia length of different experimental groups of mice. n = 5 mice per group. Data are mean ± s.e.m. c–i, Two-way ANOVA followed by Sidak’s multiple-comparisons test. c–f, h, i, Numbers above square brackets show significant P values. g, 15 min ***P = 0.0002 chow wild type versus HFD + l-NAME wild type, ****P < 0.0001 chow wild type versus HFD + l-NAME Nos2 knockout; 30 min ***P = 0.0002 chow wild type versus HFD + l-NAME wild type, ****P < 0.0001 chow wild type versus HFD + l-NAME Nos2 knockout; 45 min ****P < 0.0001 chow wild type versus HFD + l-NAME wild type, ***P = 0.0002 chow wild type versus HFD + l-NAME Nos2 knockout; 60 min ****P < 0.0001 chow wild type versus HFD + l-NAME wild type, ****P < 0.0001 chow wild type versus HFD + l-NAME Nos2 knockout; 120 min **P = 0.010 chow wild type versus HFD + l-NAME wild type, **P = 0.007 chow wild type versus HFD + l-NAME Nos2 knockout. For gel source data, see Supplementary Fig. 1.
Extended Data Fig. 10 Functional characterization and myocardial levels of XBP1s in mice treated with an iNOS inhibitor.
a, Experimental design. C57BL/6N mice were fed a chow or HFD + l-NAME diet (brown triangle) for five weeks and subsequently injected intraperitoneally (i.p.) with l-NIL at the dose of 80 mg kg−1 body weight or vehicle twice a day for three days (blue triangles). After that point, mice were subjected to functional analysis and tissues were collected (red triangle). b, Urinary nitrite/nitrate concentration in mice fed a HFD + l-NAME diet treated with vehicle or l-NIL. n = 5 mice per group. c, Systolic blood pressure. d, Diastolic blood pressure. e, Percentage of LVEF. f, Ratio between mitral E wave and A wave. g, Ratio between mitral E wave and E′ wave. h, Running distance during exercise exhaustion test. i, Xbp1s mRNA levels in left ventricles of mice from different experimental groups. b–h, n = 5 mice group; i, n = 3 per group. Data are mean ± s.e.m. b, Two-tailed unpaired Student’s t-test. c–i, Two-way ANOVA followed by Sidak’s multiple-comparisons test. Numbers above square brackets show significant P values.
Supplementary information
Supplementary Figures
This file contains uncropped gel data scans.
Source data
Rights and permissions
About this article
Cite this article
Schiattarella, G.G., Altamirano, F., Tong, D. et al. Nitrosative stress drives heart failure with preserved ejection fraction. Nature 568, 351–356 (2019). https://doi.org/10.1038/s41586-019-1100-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-019-1100-z
- Springer Nature Limited
This article is cited by
-
Inhibition of ferroptosis reverses heart failure with preserved ejection fraction in mice
Journal of Translational Medicine (2024)
-
Redox proteomics of PANC-1 cells reveals the significance of HIF-1 signaling protein oxidation in pancreatic ductal adenocarcinoma pathogenesis
Journal of Translational Medicine (2024)
-
Empagliflozin protects against heart failure with preserved ejection fraction partly by inhibiting the senescence-associated STAT1–STING axis
Cardiovascular Diabetology (2024)
-
Linking immune modulation to cardiac fibrosis
Nature Cardiovascular Research (2024)
-
Protein kinase N promotes cardiac fibrosis in heart failure by fibroblast-to-myofibroblast conversion
Nature Communications (2024)