Abstract
The liver is the most common site of metastatic disease1. Although this metastatic tropism may reflect the mechanical trapping of circulating tumour cells, liver metastasis is also dependent, at least in part, on the formation of a ‘pro-metastatic’ niche that supports the spread of tumour cells to the liver2,3. The mechanisms that direct the formation of this niche are poorly understood. Here we show that hepatocytes coordinate myeloid cell accumulation and fibrosis within the liver and, in doing so, increase the susceptibility of the liver to metastatic seeding and outgrowth. During early pancreatic tumorigenesis in mice, hepatocytes show activation of signal transducer and activator of transcription 3 (STAT3) signalling and increased production of serum amyloid A1 and A2 (referred to collectively as SAA). Overexpression of SAA by hepatocytes also occurs in patients with pancreatic and colorectal cancers that have metastasized to the liver, and many patients with locally advanced and metastatic disease show increases in circulating SAA. Activation of STAT3 in hepatocytes and the subsequent production of SAA depend on the release of interleukin 6 (IL-6) into the circulation by non-malignant cells. Genetic ablation or blockade of components of IL-6–STAT3–SAA signalling prevents the establishment of a pro-metastatic niche and inhibits liver metastasis. Our data identify an intercellular network underpinned by hepatocytes that forms the basis of a pro-metastatic niche in the liver, and identify new therapeutic targets.
Similar content being viewed by others
Main
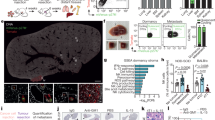
To understand the mechanisms that underlie the formation of a pro-metastatic niche in the liver, we used the LSL-KrasG12D/+;LSL-Trp53R127H/+;Pdx1-cre (KPC) mouse model of pancreatic ductal adenocarcinoma (PDAC)4,5. We looked for features of a pro-metastatic niche in the livers of over-16-week-old tumour-bearing KPC mice and 8- to 10-week-old non-tumour-bearing (NTB) KPC control mice, which lack PDAC but harbour pancreatic intraepithelial neoplasia (PanIN)6. Compared to control mice, the livers of KPC mice contained increased numbers of myeloid cells, accompanied by an increase in the deposition and expression of fibronectin and type I collagen (COL1) (Fig. 1a, Extended Data Fig. 1a–d). Orthotopic implantation of KPC-derived PDAC cells into wild-type mice recapitulated these changes (Extended Data Fig. 1e–i). As shown previously7,8, matrix deposition did not require myeloid cells (Extended Data Fig. 1j–l). These results are consistent with evidence that myeloid cell accumulation and extracellular matrix deposition are key components of a pro-metastatic niche7,8,9,10.
a, Images and quantification of myeloid cells, fibronectin (FN), and COL1 in the liver. Arrows indicate Ly6G+ cells. Numbers in parentheses on plots indicate the number (n) of mice. Data pooled from two experiments. TB, tumour-bearing; NTB, non-tumour-bearing. b, Images of the liver and quantification of PDAC–YFP cells. Control mice (n = 14) and NTB KPC mice (n = 10) were intrasplenically injected with PDAC–YFP cells, and the liver was analysed after 10 days. Data representative of two independent experiments. c, Scatter plot of transcriptome data. FPKM, fragments per kilobase of exon per million mapped fragments (n = 5 for both groups). Scale bars, 50 μm (a) and 1 cm (b). Statistical significance calculated using one-way analysis of variance (ANOVA) with Dunnett’s test (a) and two-tailed Mann–Whitney test (b). Data represented as mean ± s.d.
We next evaluated the susceptibility of the liver to metastatic colonization. Yellow fluorescent protein (YFP)-labelled KPC-derived PDAC cells (PDAC–YFP)6 were injected into control mice and KPC mice. The metastatic burden was threefold higher in KPC mice, and metastatic lesions were detected in the livers of KPC mice at increased frequency and size with enhanced proliferation (shown using Ki-67) (Fig. 1b, Extended Data Fig. 2a, b). Similar findings were observed using a YFP-negative KPC-derived cell line (Extended Data Fig. 2c, d). Orthotopic implantation of PDAC cells also increased the susceptibility of the liver to metastatic colonization, and this finding was independent of the presence of CD4+ and CD8+ T cells (Extended Data Fig. 2e–s).
We next performed mRNA sequencing on RNA isolated from the livers of control and KPC mice. We identified 275 differentially expressed genes (Extended Data Fig. 3a, b, Supplementary Data 1) and found that genes upregulated in KPC mice were associated with immune-related processes (Extended Data Fig. 3c). Notably, genes encoding myeloid chemoattractants, including SAA and members of the S100 family, were upregulated in KPC mice11,12,13 (Fig. 1c, Extended Data Fig. 3d, e). We also found enrichment of immune-related pathways, particularly the IL-6–JAK–STAT3 signalling pathway (Extended Data Fig. 3f, Supplementary Table 1). We validated our results by examining the livers of KPC mice for the presence of phosphorylated STAT3 (pSTAT3). STAT3 was activated in 80–90% of hepatocytes from KPC mice, compared to less than 2% of hepatocytes in control mice (Extended Data Fig. 3g, h). By contrast, we did not detect activation of STAT1 signalling (Extended Data Fig. 3i). Orthotopic implantation of PDAC cells also induced phosphorylation of STAT3 in hepatocytes (Extended Data Fig. 3j, k).
As IL-6 is fundamental to STAT3 signalling in hepatocytes14, we examined the livers of control mice (Il6+/+) and IL-6 knockout mice (Il6−/−) orthotopically injected with PBS or PDAC cells. Tumour-implanted Il6−/− mice displayed a decrease in STAT3 activation, particularly in hepatocytes (Fig. 2a, Extended Data Fig. 4a). This loss in STAT3 activation was accompanied by reductions in myeloid cell accumulation and extracellular matrix deposition without alterations in the morphology and density of liver sinusoids (Fig. 2a and Extended Data Fig. 4a-d). We also observed reduced expression of SAA, other chemoattractants, and extracellular matrix proteins (Fig. 2b, Extended Data Fig. 4e). Genetic ablation of Il6, however, did not alter proliferation, vascular density, or primary tumour growth (Extended Data Fig. 4f, g). Il6−/− mice were also less susceptible than control mice to metastatic colonization, and blockade of the IL-6 receptor (IL-6R) similarly inhibited the formation of a pro-metastatic niche in the liver (Fig. 2c–e, Extended Data Fig. 4h–s). Notably, genetic ablation of Il6 or blockade of IL-6R did not completely inhibit STAT3 signalling, suggesting that IL-6-independent mechanisms may contribute to STAT3 activation.
a, b, n = 5 and 6 for Il6+/+ mice and n = 4 and 5 for Il6−/− mice orthotopically injected with PBS or PDAC cells, respectively. a, Quantification of pSTAT3+ cells, myeloid cells, and fibronectin. b, mRNA levels of Saa1 and Saa2 in the liver. c–e, n = 4 and 5 for Il6+/+ mice and n = 4 for Il6−/− mice orthotopically injected with PBS or PDAC cells, respectively. All groups were intraportally injected with PDAC–YFP cells on day 10. c, d, Images of liver and flow cytometric analysis. e, Quantification of PDAC–YFP cells. Data representative of two independent experiments (a–e). Scale bars, 1 cm. Statistical significance calculated using one-way ANOVA with Dunnett’s test. Data represented as mean ± s.d.
IL-6 promotes the development and progression of PDAC15,16,17,18. To identify the source of IL-6, we orthotopically injected PBS or PDAC cells into Il6+/+ and Il6−/− mice and measured the concentration of IL-6 at distinct anatomic sites (Extended Data Fig. 5a). We detected IL-6 only in tumour-implanted Il6+/+ mice, with the highest concentration of IL-6 found in the primary tumour (Extended Data Fig. 5b, c). Although Il6 mRNA was undetectable in the liver, lung, and malignant cells, we observed Il6 mRNA in host cells adjacent to CK19-expressing PDAC cells (Extended Data Fig. 5d–g). Human primary tumours displayed a similar expression pattern (Extended Data Fig. 5h). Moreover, Il6 mRNA was detected in α-SMA+ stromal cells located adjacent to PanIN and PDAC cells in KPC mice (Extended Data Fig. 5i–k). We also found that primary pancreatic tumour supernatant activated STAT3 signalling in hepatocytes, and this was reduced in the presence of anti-IL-6R antibodies (Extended Data Fig. 6a, b). These results show that IL-6 released by non-malignant cells within the primary tumour is a key mediator of STAT3 signalling in hepatocytes.
To study a role for hepatocytes in directing liver metastasis, we generated mice that lacked Stat3 in hepatocytes (Stat3flox/flox Alb-cre). Compared to control mice (Stat3flox/flox), tumour-implanted Stat3flox/flox Alb-cre mice lacked features of a pro-metastatic niche (Fig. 3a–c, Extended Data Fig. 6c) and failed to produce SAA (Fig. 3d–f). However, deletion of Stat3 in hepatocytes did not affect liver sinusoid density or morphology and did not alter the size, proliferation, or vascular density of the primary tumour (Extended Data Fig. 6d–f). The livers of tumour-implanted Stat3flox/flox Alb-cre mice were also less susceptible to metastatic colonization (Extended Data Fig. 6g–l). In addition to its expression in hepatocytes (Extended Data Fig. 6m), mRNA for SAA was detected in colonic cells19 and in cells present in the periphery of the primary tumour (Extended Data Fig. 6n). However, both cell types maintained comparable levels of SAA mRNA despite deletion of Stat3 in hepatocytes.
a, Study design for b–f (n = 4 for Stat3flox/flox mice injected with PBS or PDAC cells; n = 8 and 7 for Stat3flox/flox Alb-cre mice injected with PBS and PDAC cells, respectively). b, c, Quantification of pSTAT3+ cells, myeloid cells, and fibronectin. d, mRNA levels of Saa1 and Saa2 in the liver. e, Images of Saa1 and Saa2 mRNA in liver cells. Dashed lines and asterisks indicate sinusoids and hepatocytes, respectively. f, Concentration of circulating SAA. Data representative of two independent experiments (a–f). Scale bars, 50 μm. Statistical significance calculated using one-way ANOVA with Dunnett’s test. Data represented as mean ± s.d.
SAA proteins are acute phase reactants20. Consistent with elevated levels of circulating SAA in tumour-implanted mice (Fig. 3f), patients with PDAC displayed elevated levels of circulating SAA (Extended Data Fig. 7a). Overexpression of SAA and pSTAT3 by hepatocytes was also observed in five of seven patients with liver metastases (Fig. 4a, Extended Data Fig. 7b). Notably, high levels of circulating SAA correlated with worse outcomes (Extended Data Fig. 7c). Elevated levels of circulating SAA were also observed in patients with non-small-cell lung carcinoma (NSCLC) with liver metastases, and overexpression of SAA by hepatocytes was detected in the livers of patients with colorectal carcinoma (CRC) (Extended Data Fig. 7d, e). In addition, compared to tumour-implanted control mice (Saa+/+), double-knockout Saa1−/− Saa2−/− mice (hereafter referred to as Saa−/− mice) implanted with PDAC or MC-38 CRC cells failed to show features of a pro-metastatic niche in the liver, though genetic ablation of Saa1 and Saa2 had no effect on primary tumour growth (Fig. 4b–e, Extended Data Fig. 7f–s). SAA was also necessary for IL-6-mediated formation of a pro-metastatic niche and for fibrosis and myeloid cell recruitment in the setting of liver injury (Extended Data Fig. 8).
a, Images of SAA in the livers of healthy donors (top) and patients with PDAC with liver metastases (bottom). Dashed lines and asterisks indicate sinusoids and hepatocytes, respectively. Data representative of one experiment. b, Quantification of pSTAT3+ cells, myeloid cells, and fibronectin (n = 5 for all groups orthotopically injected with PBS or PDAC cells). For c–e, n = 4 and 5 for Saa+/+ mice and n = 5 and 6 for Saa−/− mice orthotopically injected with PBS and PDAC cells, respectively. All groups were intraportally injected with PDAC–YFP cells on day 10. c, d, Images of liver and flow cytometric analysis. e, Quantification of PDAC–YFP cells. Data representative of two independent experiments (b–e). Scale bars, 50 μm (a) and 1 cm (c). Statistical significance calculated using one-way ANOVA with Dunnett’s test. Data represented as mean ± s.d.
Tissue inhibitor of metalloproteinases 1 (TIMP1)7,8 and macrophage migration inhibitory factor (MIF)9,10 have been implicated in the promotion of metastasis. However, expression of these molecules was not affected by IL-6–STAT3–SAA signalling (Extended Data Fig. 9). We next determined whether formation of a pro-metastatic niche in the liver is dependent on the anatomical proximity of the pancreas to the liver. To this end, we looked for features of a pro-metastatic niche in the livers of CD45.1 and CD45.2 mice that were parabiotically joined (Extended Data Fig. 10a). Although only CD45.2 mice were implanted with PDAC cells, both mice displayed myeloid cell accumulation and fibrosis in the liver (Extended Data Fig. 10b–g), suggesting that formation of this niche is not dependent on the anatomical distance between the tumour and the liver. We also investigated whether SAA has a role in establishing a pro-metastatic niche in the lung. Development of PDAC in KPC mice induced accumulation of Ly6G+ myeloid cells and deposition of fibronectin within the lung, but IL-6–STAT3–SAA signalling was not required for the formation of a pro-metastatic niche in the lung (Extended Data Fig. 10h–o).
Our data provide insight into the mechanisms that direct liver metastasis. Although recent studies have suggested a role for tumour-intrinsic factors in driving metastatic spread of cancer7,8,9,10,21,22,23, we provide evidence that inflammatory responses mounted by hepatocytes are critical to liver metastasis. Mechanistically, hepatocytes orchestrate this process through activation of IL-6–STAT3 signalling and the subsequent production of SAA, which alters the immune and fibrotic microenvironment of the liver to establish a pro-metastatic niche (Extended Data Fig. 10p). Our findings suggest that therapies that target hepatocytes might prevent liver metastasis in patients with cancer.
Methods
Mice
CD45.2 (wild type, C57BL/6J), CD45.1 (B6.SJL-Ptprca Pepcb/BoyJ), Il6 knockout (Il6−/−, B6.129S2-Il6tm1Kopf/J), Stat3flox/flox (B6.129S1-Stat3tm1Xyfu/J), and Alb-cre+/+ (B6.Cg-Tg(Alb-cre)21Mgn/J) mice were obtained from the Jackson Laboratory. Stat3flox/flox mice were bred to Alb-cre+/+ mice to generate Stat3flox/+ Alb-cre+/− mice, which were backcrossed onto Stat3flox/flox mice to generate Stat3flox/flox Alb-cre+/− mice. These mice were then bred to each other to create Stat3flox/flox Alb-cre+/+ and Stat3flox/flox Albumin-cre+/− mice (Stat3flox/flox Alb-cre), and Stat3flox/flox Albumin-cre−/− mice (Stat3flox/flox). KrasLSL-G12D/+ Trp53LSL-R172H/+ Pdx1-cre (KPC) mice and Trp53LSL-R172H/+ Pdx1-cre (PC) mice were as previously described4,5. Saa1 and Saa2 double-knockout (Saa−/−) mice were as previously described24 and provided by the University of Kentucky College of Medicine. Saa−/− mice used for experiments had been bred to obtain a 99.9% C57BL/6 background using the Jackson Laboratory Speed Congenic Service24. All transgenic mice were bred and maintained in the animal facility of the University of Pennsylvania. Animal protocols were reviewed and approved by the Institute of Animal Care and Use Committee of the University of Pennsylvania. In general, mice were monitored three times per week for general health and euthanized early based on defined endpoint criteria including tumour diameter ≥1 cm, ascites, lethargy, loss of ≥10% body weight, or other signs of sickness or distress.
Clinical samples
All patient samples were obtained after written informed consent and were de-identified. Studies were conducted in accordance with the 1996 Declaration of Helsinki and approved by institutional review boards of the University of Pennsylvania and the Mayo Clinic. To obtain plasma from healthy donors, patients with PDAC patients, and patients with NSCLC, peripheral whole blood was drawn in EDTA tubes (Fisher Scientific). Within 3 h of collection, blood samples were centrifuged at 1,600g at room temperature for 10 min with the brake off. Next, the plasma was transferred to a 15-ml conical tube without disturbing the cellular layer and centrifuged at 3,000g at room temperature for 10 min with the brake off. This step was repeated with a fresh 15-ml conical tube. The plasma was then stored at –80 °C until analysis. Biopsy results, computed tomography, and/or magnetic resonance imaging records were used to determine sites of metastasis in patients with PDAC or NSCLC whose plasma samples were used for assessment of SAA levels. Liver specimens from healthy donors were obtained by percutaneous liver biopsy, and acquisition of liver specimens from patients with liver metastases was as previously described25. Liver specimens from patients with CRC with liver metastases were obtained from the Cooperative Human Tissue Network (CHTN). Patient characteristics are shown in Supplementary Table 2.
Cell lines
PDA.69 cell line (PDAC cells) was used for intrasplenic and orthotopic injection, and PDA.8572 cell line (PDAC–YFP cells) was used for intrasplenic, intraportal, and retro-orbital injections. These cell lines were derived from PDAC tumours that arose spontaneously in KPC mice, as previously described4,26. The MC-38 cell line, which was used for orthotopic implantation, was purchased from Kerafast. Cell lines were cultured in DMEM (Corning) supplemented with 10% fetal bovine serum (FBS, VWR), 83 μg/ml gentamicin (Thermo Fisher), and 1% GlutaMAX (Thermo Fisher) at 37 °C, 5% CO2. Only cell lines that had been passaged fewer than 10 times were used for experiments, and trypan blue staining was used to ensure that cells with >95% viability were used for studies. Cell lines were tested routinely for Mycoplasma contamination at the Cell Center Services Facility at the University of Pennsylvania. All cell lines used in our studies tested negative for Mycoplasma contamination.
Animal experiments
For all animal studies, mice of similar age and gender were block randomized in an unblinded fashion. Male and female mice aged between 8 to 12 weeks were used unless indicated otherwise. Mice were age- and gender-matched with appropriate control mice for analysis. Sample sizes were estimated based on pilot experiments and were selected to provide sufficient numbers of mice in each group for statistical analysis.
For orthotopic and intrasplenic injections of pancreatic tumour cells, mice were anaesthetized using continuous isoflurane, and their abdomen was sterilized. After administering analgesic agents and assessing the depth of anaesthesia, we performed a laparotomy (5–10 mm) over the left upper quadrant of the abdomen to expose the peritoneal cavity. For orthotopic injection, the pancreas was exteriorized onto a sterile field, and sterile PBS or pancreatic tumour cells (5 × 105 cells suspended in 50 μl of sterile PBS) were injected into the tail of the pancreas via a 30-gauge needle (Covidien). Successful injection was confirmed by the formation of a liquid bleb at the site of injection with minimal fluid leakage. The pancreas was then gently placed back into the peritoneal cavity. For intrasplenic injection, 150 μl sterile PBS was drawn into a syringe and then sterile PBS or pancreatic tumour cells (5 × 105 cells suspended in 100 μl sterile PBS) was gently drawn into the same syringe in an upright position as previously described27. After the spleen was exteriorized onto a sterile field, pancreatic tumour cells were injected into the spleen via a 30-gauge needle. Successful injection was confirmed by whitening of the spleen and splenic blood vessels with minimal leakage of content into the peritoneum. Splenectomy was then performed by ligating splenic vessels with clips (Horizon) then cauterizing them to ensure that there was no haemorrhage. Afterwards, the remaining blood vessels were placed back into the peritoneal cavity. For both procedures, the peritoneum was closed with a 5-0 PDS II violet suture (Ethicon), and the skin was closed using the AutoClip system (Braintree Scientific). Following surgery, mice were given buprenorphine subcutaneously at a dose of 0.05-0.1 mg/kg every 4–6 h for 12 h and then every 6–8 h for 3 additional days. Mice that were orthotopically injected with pancreatic tumour cells were analysed after 20 days, unless indicated otherwise in study designs. Mice that were intrasplenically injected with PDAC cells were analysed after 10 days.
For intraportal injection of pancreatic tumour cells and hydrodynamic injection of expression vectors, mice were anaesthetized using continuous isoflurane, and their abdomen was sterilized. After administration of analgesic agents, median laparotomy (10 mm) was performed, and the incision site was held open using an Agricola retractor (Roboz). After exposure of the peritoneal cavity, the intestines were located and exteriorized onto a sterile field surrounding the incision site to visualize the portal vein. Throughout the procedure, the intestines were kept hydrated with sterile PBS that was pre-warmed to 37 °C. For intraportal injection, sterile PBS or pancreatic tumour cells (5 × 105 cells suspended in 100 μl sterile PBS) were injected into the portal vein via a 30-gauge needle. Successful injection was confirmed by partial blanching of the liver. For hydrodynamic injection, 1 μg of pLIVE expression vectors was suspended in sterile saline corresponding to 8% of mouse body weight as previously described28. Vectors were injected into the portal vein via a 27-gauge needle within 5–8 s. Successful injection was confirmed by complete blanching and swelling of the liver. For both procedures, a sterile gauge was then held over the injection site for 1 min to ensure that no injected contents would leak into the peritoneal cavity. Afterwards, the intestines were placed back into the peritoneal cavity, and the peritoneum and skin were closed with a suture and autoclips, respectively. Following surgery, mice were given buprenorphine subcutaneously as described above. Intraportal injection of pancreatic tumour cells was performed on day 10, and metastatic burden in the liver was evaluated on day 20, unless indicated otherwise in study designs.
For orthotopic implantation of colorectal tumour cells, wild-type mice were first subcutaneously injected with MC-38 (1 × 106 cells suspended in 100 μl of sterile PBS) into the right flank. After 10 days, mice were euthanized, and subcutaneous tumours were collected. Tumours were then cut into small pieces, each 3 × 3 mm in size, and placed in sterile PBS on ice until implantation. Mice were anaesthetized using isoflurane, and their abdomen was sterilized. Following administration of analgesic agents, median laparotomy was performed as described above. Implantation of colorectal tumour tissues into the caecum was then performed as previously described29. After we placed the intestines back into the peritoneal cavity, the peritoneum and skin were closed with a suture, and mice were given buprenorphine as described above. Mice were analysed after 10 days.
For parabiotic joining of mice, female CD45.2 mice were orthotopically injected with sterile PBS or pancreatic tumour cells as described above and co-housed with age-matched female B6 CD45.1 mice. Each parabiotic pair was housed in a separate cage to maximize bonding between partners. After one week, parabiotic partners were anaesthetized using continuous isoflurane, and their flanks were sterilized. After administration of analgesic agents, longitudinal skin flaps from the lower limb to the upper limb were created, and everted skin flaps were sewn using a suture. In addition, the knees and olecranons of parabiotic partners were joined together using a suture for additional stabilization. Following surgery, mice were given buprenorphine subcutaneously at a dose of 0.05-0.1 mg/kg every 4-6 h for 5 days. Parabiotically joined mice were analysed after 20 days.
For administration of antibodies, the abdomen of mice was sterilized, and anti-CD4 antibodies (GK1.5, 0.2 mg), anti-CD8 antibodies (2.43, 0.2 mg), anti-IL-6R antibodies (15A7, 0.2 mg), or rat isotype control antibodies (LTF-2, 0.2 mg) were suspended in 100 μl sterile PBS. Antibodies were subsequently injected into the peritoneum via a 30-gauge needle. All antibodies used in in vivo experiments were obtained from BioXCell. To deplete F4/80+ myeloid cells, clodronate-encapsulated liposomes (Liposoma) were administered by intraperitoneal injection according to the manufacturer’s protocol. For induction of liver injury, mice were intraperitoneally injected with CCl4 (Sigma, 1 ml/kg body weight) dissolved in sunflower seed oil as previously described30. Detailed information on antibodies and reagents used in experiments can be in found in Supplementary Table 3.
Microscopic analysis
For preparation of formalin-fixed paraffin-embedded (FFPE) sections, dissected tissues were fixed in 10% formalin for 24 h at room temperature, washed twice with PBS, and then stored in 70% ethanol solution at 4 °C until they were embedded in paraffin and sectioned at 5 μm. For preparation of cryosections, dissected tissues were embedded in Tissue-tek O.C.T. (Electron Microscopy Sciences) and frozen on dry ice. Frozen tissues were stored at –80 °C until they were sectioned at 7 μm.
Automated immunohistochemistry, immunofluorescence, and RNA in situ hybridization were performed on FFPE sections using a Ventana Discovery Ultra automated slide staining system (Roche). Reagents were obtained from Roche and ACDBio (Supplementary Table 3) and used according to manufacturer’s protocol. Images were acquired using a BX43 upright microscope (Olympus), an Aperio CS2 scanner system (Leica), or an IX83 inverted multicolour fluorescent microscope (Olympus). Manual immunohistochemistry of mouse tissues for SAA was previously described31. For manual multicoloured immunofluorescence staining, O.C.T. liver cryosections were briefly air dried and fixed with 3% formaldehyde at room temperature for 15 min. For intracellular staining, sections were permeabilized with methanol at –20 °C for 10 min immediately after formaldehyde fixation. Sections were then blocked with 10% normal goat serum in PBS containing 0.1% TWEEN 20 for 30 min. For intracellular staining, 0.3% Triton X-100 was added to the blocking solution for permeabilization of cellular and nuclear membranes. Sections were incubated with primary antibodies (Supplementary Table 3) in the blocking solution for 1 h at room temperature or overnight at 4 °C, followed by washing with PBS containing 0.1% TWEEN 20. Sections were then incubated with secondary antibodies (Supplementary Table 3) in the blocking solution for 1 h at room temperature or overnight at 4 °C. After washing, sections were stained with DAPI to visualize nuclei and subsequently with Sudan Black B in 70% ethanol to reduce autofluorescence, as previously described32. Immunofluorescence imaging was performed on an IX83 inverted multicolour fluorescent microscope (Olympus). For quantification of cells and extracellular matrix proteins, five random fields were acquired from each biological sample.
Flow cytometry
Mice were euthanized, and the liver and lung were removed after the blood was drained by severing the portal vein and inferior vena cava. The liver and lung were rinsed thoroughly in PBS before mincing with micro-dissecting scissors into small pieces (<0.5 × 0.5 mm in size) at 4 °C in DMEM containing collagenase (1 mg/ml, Sigma-Aldrich), DNase (150 U/ml, Roche), and Dispase (1 U/ml, Worthington). Tissues were then incubated at 37 °C for 30 min with intermittent agitation, filtered through a 70-μm nylon strainer (Corning), and washed three times with DMEM. Cells were resuspended in ACK lysing buffer (Life Technologies) at room temperature for 15 min to remove red blood cells. After washing three times with DMEM, cells were counted and stained using Aqua dead cell stain kit (Life Technologies) following the manufacturer’s protocol. For characterization of immune cell subsets, cells were washed three times with PBS containing 0.2 mM EDTA with 2% FBS and stained with appropriate antibodies (Supplementary Table 3). For quantification of PDAC–YFP cells, cells were not stained with any antibodies. Lastly, cells were washed three times with PBS containing 0.2 mM EDTA with 2% FBS and examined using a FACS Canto II (BD Biosciences). Collection and analysis of the peripheral blood was as previously described26. FlowJo (FlowJo, LLC, version 10.2) was used to analyse flow cytometric data and generate 2D t-SNE plots.
Detection of IL-6, SAA, and TIMP1
Mice that were orthotopically implanted with PDAC cells were euthanized, and primary tumours were removed and weighed. In addition, blood samples were collected from the portal vein and left ventricle of the heart using a 27-gauge needle. Tumours were rinsed thoroughly in PBS and minced with micro-dissecting scissors into small pieces (<0.5 × 0.5 mm in size) at 4 °C in serum-free DMEM at 1 mg of tissue per 1 μl medium. Tumour suspensions were then centrifuged at 12,470g at 4 °C for 15 min, and tumour supernatant was collected and stored at –80 °C until analysis. A similar procedure was performed to obtain pancreas supernatant from mice that were orthotopically injected with PBS. To collect the serum, blood samples were allowed to clot at room temperature for 30 min. Samples were then centrifuged at 12,470g at 4 °C for 15 min, and the serum was collected and stored at –80 °C until analysis. IL-6 levels in tumour or pancreas supernatant and serum were assessed using a cytometric bead array (BD Biosciences) following the manufacturer’s protocol. Samples were examined using a FACS Canto II (BD Biosciences), and data were analysed using FCAP Array (BD Biosciences, version 3.0). SAA and TIMP1 levels in mouse serum samples were measured using a commercially available enzyme-linked immunosorbent assay kit (Thermo Fisher) following the manufacturer’s protocol. Similarly, SAA levels in plasma samples collected from healthy donors and patients with PDAC as described under ‘Clinical samples’ were measured using a commercially available human enzyme-linked immunosorbent assay kit (Thermo Fisher) following the manufacturer’s protocol.
RNA and quantitative PCR
Mouse organs and cells were stored in TRIzol (Thermo Fisher) at –80 °C until analysis. Samples were thawed on ice and allowed to equilibrate to room temperature before RNA was isolated using a RNeasy Mini kit (Qiagen) following the manufacturer’s protocol. cDNA synthesis was performed as previously described33. Primers for quantitative PCR were designed using the Primer3 online program34, and sequences were analysed using the Nucleotide BLAST (NCBI) to minimize non-specific binding of primers. Primers were synthesized by Integrated DNA Technologies, and their sequences can be found in Supplementary Table 4. Quantitative PCR was performed as previously described33. Gene expression was calculated relative to Actb (β-actin) using the ∆Ct formula, and fold change in gene expression was calculated relative to the average gene expression of control groups using the ∆∆Ct formula. Genes with Ct greater than or equal to 30 were considered not detected.
QuantSeq 3′ mRNA sequencing and data analysis
RNA was isolated from the livers of control mice and NTB KPC mice as described above and submitted to the Genomics Facility at the Wistar Institute. After the quality of RNA was assessed using a 2100 Bioanalyzer (Agilent), samples were prepared using a QuantSeq 3′ mRNA-Seq library prep kit FWD for Illumina (Lexogen) following the manufacturer’s protocol and analysed on a NextSeq 500 sequencing system (Illumina). FASTQ files were uploaded to the BaseSpace Suite (Illumina) and aligned using its RNA-Seq Alignment application (version 1.0.0), in which STAR was selected to align sequences with maximum mismatches set to 14 as recommended by Lexogen. Output files were analysed using Cufflinks Assembly & DE application (version 2.1.0) in the BaseSpace Suite to determine differentially expressed genes, which were used to generate an expression heatmap and a FPKM scatter plot. In addition, these genes were analysed using ClueGO (version 2.3.3)35 and CluePedia (version 1.3.3)36, which are applications of Cytoscape software (version 3.5.1)37. Functional grouping of biological processes was performed on the basis of kappa score. Gene Ontology data38,39 downloaded on 23 January 2018 were used for analysis. Gene set enrichment analysis (version 3.0)40 was used to determine biological processes that were differentially enriched in experimental groups.
In vitro studies
To isolate primary hepatocytes for in vitro studies, mice were anaesthetized using continuous isoflurane, and their abdomen was sterilized. After administering analgesic agents and assessing the depth of anaesthesia, we performed a laparotomy (10–15 mm) along the midline of the abdomen to expose the peritoneal cavity. The intestines were then located and exteriorized to visualize the inferior vena cava and portal vein. The inferior vena cava was cannulated via a 24 gauge Insyte Autoguard cathether (BD), and the liver was perfused using 50 ml liver perfusion medium (Thermo Fisher) at a flow rate of 8–9 ml/min using a peristaltic pump. At the start of perfusion, the portal vein was severed to drain the blood from the liver. Successful perfusion was confirmed by blanching of the liver, which was subsequently perfused using 50 ml liver digest medium (Thermo Fisher) at the same flow rate. Both liver perfusion medium and liver digest medium were pre-warmed to 42 °C in a water bath. After perfusion, the liver was carefully transferred to a Petri dish containing William’s E medium (Sigma) supplemented with 10% FBS, 83 μg/ml gentamicin, and 1% GlutaMAX.
To dissociate hepatocytes from the liver, cell scrapers were used to create small cuts (5 mm) on the surface of the liver, and the tissue was gently shaken. Dissociated cells were then filtered through a 100-μm nylon strainer (Corning) and centrifuged at 50g at 4 °C for 5 min. After the supernatant was discarded, cells were resuspended in a solution consisting of isotonic Percoll (Sigma) and supplemented William’s E medium (2:3 ratio). Cells were then centrifuged at 50g at 4 °C for 10 min to obtain a pellet enriched in hepatocytes. The supernatant was discarded, and hepatocytes were resuspended in supplemented William’s E medium. Cell viability and number were determined using trypan blue staining, and 5 × 104 hepatocytes were seeded in each well of a 48-well plate pre-coated with collagen. Hepatocytes were incubated in supplemented William’s E medium for 4 h at 37 °C, 5% CO2 to allow attachment to the plate. The medium was then switched to HepatoZYME-SFM (Thermo Fisher) supplemented with 83 μg/ml gentamicin and 1% GlutaMAX. Medium was replenished every 24 h for the next 48–72 h.
For hepatocyte activation assays, hepatocytes were incubated in supplemented HepatoZYME-SFM mixed with (i) serum-free DMEM, (ii) primary pancreatic tumour supernatant, or (iii) serum-free DMEM containing 250 ng/ml IL-6 (Peprotech) for 30 min at 37 °C, 5% CO2. All mixtures were made in a 1:1 ratio, and each condition was run in triplicate. For the in vitro IL-6R blockade experiment, hepatocytes were pre-incubated with 5 μg/ml anti-IL-6R antibodies for 2 h before being stimulated with tumour supernatant. After stimulation, medium was carefully removed, and formaldehyde and methanol were used to fix and permeabilize hepatocytes, respectively, as described above. Hepatocytes were then stained for pSTAT3 (Supplementary Table 3), and their nuclei stained with DAPI. Immunofluorescence imaging was performed on an IX83 inverted multicolour fluorescent microscope (Olympus).
Statistical analysis
Statistical significance was calculated using Prism (GraphPad Software, version 7) unless indicated otherwise. Multiple comparisons testing was performed using one-way ANOVA with Dunnett’s test. Paired group comparisons test was carried out using two-tailed Wilcoxon matched-pairs signed rank test. Unpaired group comparisons test was performed using two-tailed unpaired Student’s t test or two-tailed Mann–Whitney test. Comparison of Kaplan–Meier overall survival curves was performed using log-rank (Mantel-Cox) test. P values less than 0.05 were treated as significant. The experiments were not randomized and the investigators were not blinded to allocation during experiments and outcome assessment, unless stated otherwise.
Reporting summary
Further information on research design is available in the Nature Research Reporting Summary linked to this paper.
Data availability
QuantSeq 3′ mRNA sequencing data have been deposited in the Gene Expression Omnibus (GEO) under accession number GSE109480. Source Data are provided for all figures and extended data figures. All data are available from the corresponding author upon reasonable request.
References
Hidalgo, M. Pancreatic cancer. N. Engl. J. Med. 362, 1605–1617 (2010).
Becker, A. et al. Extracellular vesicles in cancer: cell-to-cell mediators of metastasis. Cancer Cell 30, 836–848 (2016).
Sceneay, J., Smyth, M. J. & Möller, A. The pre-metastatic niche: finding common ground. Cancer Metastasis Rev. 32, 449–464 (2013).
Hingorani, S. R. et al. Trp53R 172H and Kras G12D cooperate to promote chromosomal instability and widely metastatic pancreatic ductal adenocarcinoma in mice. Cancer Cell 7, 469–483 (2005).
Lee, J. W., Komar, C. A., Bengsch, F., Graham, K. & Beatty, G. L. Genetically engineered mouse models of pancreatic cancer: the KPC model (LSL-Kras G12D/+ ;LSL-Trp53 R172H/+ ;Pdx-1-Cre), its variants, and their application in immuno-oncology drug discovery. Curr. Protoc. Pharmacol. 73, 14.39.1–14.39.20 (2016).
Rhim, A. D. et al. EMT and dissemination precede pancreatic tumor formation. Cell 148, 349–361 (2012).
Grünwald, B. et al. Pancreatic premalignant lesions secrete tissue inhibitor of metalloproteinases-1, which activates hepatic stellate cells via CD63 signaling to create a premetastatic niche in the liver. Gastroenterology 151, 1011–1024.e7 (2016).
Seubert, B. et al. Tissue inhibitor of metalloproteinases (TIMP)-1 creates a premetastatic niche in the liver through SDF-1/CXCR4-dependent neutrophil recruitment in mice. Hepatology 61, 238–248 (2015).
Hoshino, A. et al. Tumour exosome integrins determine organotropic metastasis. Nature 527, 329–335 (2015).
Costa-Silva, B. et al. Pancreatic cancer exosomes initiate pre-metastatic niche formation in the liver. Nat. Cell Biol. 17, 816–826 (2015).
Kowanetz, M. et al. Granulocyte-colony stimulating factor promotes lung metastasis through mobilization of Ly6G+Ly6C+ granulocytes. Proc. Natl Acad. Sci. USA 107, 21248–21255 (2010).
Hiratsuka, S. et al. The S100A8-serum amyloid A3-TLR4 paracrine cascade establishes a pre-metastatic phase. Nat. Cell Biol. 10, 1349–1355 (2008).
Hiratsuka, S., Watanabe, A., Aburatani, H. & Maru, Y. Tumour-mediated upregulation of chemoattractants and recruitment of myeloid cells predetermines lung metastasis. Nat. Cell Biol. 8, 1369–1375 (2006).
Schmidt-Arras, D. & Rose-John, S. IL-6 pathway in the liver: From physiopathology to therapy. J. Hepatol. 64, 1403–1415 (2016).
Long, K. B. et al. IL6 receptor blockade enhances chemotherapy efficacy in pancreatic ductal adenocarcinoma. Mol. Cancer Ther. 16, 1898–1908 (2017).
Lesina, M. et al. Stat3/Socs3 activation by IL-6 transsignaling promotes progression of pancreatic intraepithelial neoplasia and development of pancreatic cancer. Cancer Cell 19, 456–469 (2011).
Fukuda, A. et al. Stat3 and MMP7 contribute to pancreatic ductal adenocarcinoma initiation and progression. Cancer Cell 19, 441–455 (2011).
Corcoran, R. B. et al. STAT3 plays a critical role in KRAS-induced pancreatic tumorigenesis. Cancer Res. 71, 5020–5029 (2011).
Eckhardt, E. R. et al. Intestinal epithelial serum amyloid A modulates bacterial growth in vitro and pro-inflammatory responses in mouse experimental colitis. BMC Gastroenterol. 10, 133 (2010).
Urieli-Shoval, S., Linke, R. P. & Matzner, Y. Expression and function of serum amyloid A, a major acute-phase protein, in normal and disease states. Curr. Opin. Hematol. 7, 64–69 (2000).
Notta, F. et al. A renewed model of pancreatic cancer evolution based on genomic rearrangement patterns. Nature 538, 378–382 (2016).
Yachida, S. et al. Distant metastasis occurs late during the genetic evolution of pancreatic cancer. Nature 467, 1114–1117 (2010).
Reichert, M. et al. Regulation of epithelial plasticity determines metastatic organotropism in pancreatic cancer. Dev. Cell 45, 696–711.e8 (2018).
de Beer, M. C. et al. Impact of serum amyloid A on high density lipoprotein composition and levels. J. Lipid Res. 51, 3117–3125 (2010).
Liang, W. S. et al. Genome-wide characterization of pancreatic adenocarcinoma patients using next generation sequencing. PLoS One 7, e43192 (2012).
Long, K. B. et al. IFNγ and CCL2 cooperate to redirect tumor-infiltrating monocytes to degrade fibrosis and enhance chemotherapy efficacy in pancreatic carcinoma. Cancer Discov. 6, 400–413 (2016).
Soares, K. C. et al. A preclinical murine model of hepatic metastases. J. Vis. Exp. 91, 51677 (2014).
Ma, Y., Gao, M., Sun, H. & Liu, D. Interleukin-6 gene transfer reverses body weight gain and fatty liver in obese mice. Biochim. Biophys. Acta 1852, 1001–1011 (2015).
Tseng, W., Leong, X. & Engleman, E. Orthotopic mouse model of colorectal cancer. J. Vis. Exp. 10, 484 (2007).
Klaas, M. et al. The alterations in the extracellular matrix composition guide the repair of damaged liver tissue. Sci. Rep. 6, 27398 (2016).
Webb, N. R. et al. Deficiency of endogenous acute-phase serum amyloid A protects apoE−/− mice from angiotensin II-induced abdominal aortic aneurysm formation. Arterioscler. Thromb. Vasc. Biol. 35, 1156–1165 (2015).
Sun, Y. et al. Sudan black B reduces autofluorescence in murine renal tissue. Arch. Pathol. Lab. Med. 135, 1335–1342 (2011).
Kalbasi, A. et al. Tumor-derived CCL2 mediates resistance to radiotherapy in pancreatic ductal adenocarcinoma. Clin. Cancer Res. 23, 137–148 (2017).
Untergasser, A. et al. Primer3—new capabilities and interfaces. Nucleic Acid Res. 40, e115 (2012).
Bindea, G. et al. ClueGO: a Cytoscape plug-in to decipher functionally grouped gene ontology and pathway annotation networks. Bioinformatics 25, 1091–1093 (2009).
Bindea, G., Galon, J. & Mlecnik, B. CluePedia Cytoscape plugin: pathway insights using integrated experimental and in silico data. Bioinformatics 29, 661–663 (2013).
Shannon, P. et al. Cytoscape: a software environment for integrated models of biomolecular interaction networks. Genome Res. 13, 2498–2504 (2003).
The Gene Ontology Consortium. Expansion of the Gene Ontology knowledgebase and resources. Nucleic Acids Res. 45, D331–D338 (2017).
Ashburner, M. et al. Gene ontology: tool for the unification of biology. Nat. Genet. 25, 25–29 (2000).
Subramanian, A. et al. Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc. Natl Acad. Sci. USA 102, 15545–15550 (2005).
Acknowledgements
We thank members of the Zaret laboratory for assistance with isolation and culture of primary hepatocytes and members of the Genomics Facility (Wistar Institute) and the Molecular Pathology and Imaging Core (University of Pennsylvania) for technical support; E. J. Wherry, D. J. Powell, J. R. Conejo-Garcia, I. E. Brodsky, and E. L. Stone for discussions and advice; and the Mayo Clinic Arizona for provision of liver tissue sections collected from patients with pancreatic cancer. This work was supported by National Institutes of Health grants F30 CA196106 (J.W.L.), T32 HL007439 (J.W.L.), T32 CA009140 (M.L.S.), R01 CA197916 (G.L.B.), R01 HL134731 (N.R.W. and F.C.d.B.), the University of Pennsylvania Molecular Pathology and Imaging Core of the Center for Molecular Studies in Digestive and Liver Diseases grant P30 DK050306, the 2015 Pancreatic Cancer Action Network-AACR Career Development Award 15-20-25-BEAT supported by an anonymous foundation (G.L.B.), the 2017 Stand Up to Cancer (SU2C) Innovative Research Grant SU2C-AACR-IRG 13-17 (G.L.B.), a Research Support Grant from the University of Kentucky Office of the Vice President for Research (N.R.W.), the American Surgical Association Foundation Fellowship (P.M.P.), the University of Pennsylvania Pancreatic Cancer Research Center (E.L.C.), and the Abramson Cancer Center Translational Centers of Excellence (E.L.C.).
Reviewer information
Nature thanks Tim Greten, Anirban Maitra, Robert Schwabe and the other anonymous reviewer(s) for their contribution to the peer review of this work.
Author information
Authors and Affiliations
Contributions
Experiments and data analysis were performed by J.W.L., M.L.S., P.M.P., S.K.T., C.A.K., D.D., W.L.G., X.H., A.J., M.C.d.B, F.C.d.B, N.R.W., and G.L.B; generation of mouse pancreatic tumour cell lines by W.L.G. and G.L.B.; immunofluorescence and immunohistochemistry by J.W.L., M.L.S., J.H.L., D.D., W.L.G., and A.J.; RNA in situ hybridization by J.W.L. and X.H.; tumour cell culture by J.W.L., M.L.S., and C.A.K.; animal studies by J.W.L., M.L.S., S.K.T., C.A.K., K.G., and W.L.G.; flow cytometry by J.W.L., M.L.S., and S.K.T.; QuantSeq 3′ mRNA sequencing and data analysis by J.W.L.; quantitative PCR by J.W.L., J.H.L., and X.H.; cytokine bead array by J.W.L., M.L.S., and C.A.K.; primary hepatocyte studies by J.W.L.; enzyme-linked immunosorbent assays by J.W.L. and M.C.d.B.; hydrodynamic injection studies by J.W.L, D.L., and M.G.; parabiotic joining by J.W.L. and P.M.P. M.C.d.B, F.C.d.B., and N.R.W. provided Saa−/− mice; D.X. and A.S. provided liver specimens from healthy donors; M.J.B. and R.K.R. provided liver specimens from patients with PDAC; T.A.B., A.L.C., K.S.M., J.C.T., S.S.Y., M.H.O., C.A., and E.L.C. provided plasma samples; M.G. established the hydrodynamic injection procedure; D.L. designed and prepared the IL-6 expression vector ; P.M.P., M.G., D.L., E.L.C., M.C.d.B., F.C.d.B., and N.R.W. provided experimental advice; J.W.L. and G.L.B designed the study; and J.W.L., M.L.S., and G.L.B. prepared and wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Primary PDAC development induces myeloid cell accumulation and fibrosis within the liver.
a, Gating strategy for identification of F4/80+, Ly6G+, CD3+, and CD19+ cells. Representative images from flow cytometric analysis of cells isolated from the liver of a tumour-bearing KPC mouse are shown. b, Quantification of immune cells in the liver by flow cytometry. Numbers in parentheses indicate the number (n) of mice. Data pooled from four experiments. c, Representative Sirius red staining of the liver (n = 5 for all groups) viewed using standard light microscopy (top) and polarized light (bottom). d, mRNA levels of Fn1, Col1a1, and Des in the liver (n = 6 for all groups). Data pooled from two experiments (c, d). e–i, Wild-type mice were orthotopically injected with PBS or PDAC cells and analysed on day 20. e, Quantification of immune cells in the liver by flow cytometry (n = 5 for both groups). f, t-distributed stochastic neighbour embedding (t-SNE) 2D plots of immune cells analysed in e. g, Quantification of myeloid cells in the liver (n = 7 for PBS and n = 6 for PDAC). h, i, Images and quantification of fibronectin and COL1 in the liver (n = 6 for both groups). Data representative of at least two independent experiments (e–i). j, Study design for k and l (n = 4 for mice injected with PBS; n = 4 and 3 for mice injected with PDAC cells and then intraperitoneally injected with PBS and clodronate-encapsulated liposomes (CEL), respectively). k, Images and quantification of F4/80+ cells in the liver. l, Quantification of COL1 and fibronectin in the liver. Data representative of one experiment (j–l). Scale bars, 100 μm (c) and 50 μm (other panels). Statistical significance calculated using two-tailed unpaired Student’s t-test (d), one-way ANOVA with Dunnett’s test (k, l), and two-tailed Mann–Whitney test (other panels). NS, not significant. Data represented as mean ± s.d., except b and e, which are shown as box plots (centre line, median; box limits, upper and lower quartiles; whiskers, maximum and minimum values).
Extended Data Fig. 2 Primary PDAC development enhances liver susceptibility to metastatic colonization.
a, b, Control mice (n = 14) and NTB KPC mice (n = 10) were intrasplenically injected with PDAC–YFP cells, and the liver was analysed after 10 days. a, Images of the liver showing metastatic lesions (yellow, CK19) and Ki-67 (purple). Scale bars, 4 mm (left) and 200 μm (right). b, Quantification of lesions (left) and Ki-67+ tumour cells (right). Data pooled from three experiments (a, b). c, d, n = 9 and 13 for control mice and n = 6 and 13 for NTB KPC mice intrasplenically injected with PBS and PDAC (PDA.69), respectively. c, Images of the liver (top) and metastatic lesions in the liver (stained with PDX1). Scale bars, 1 cm (top) and 50 μm (bottom). d, Liver weights and mRNA levels of Pdx1 in the liver relative to Gapdh. Data pooled from five experiments (c, d). e, Study design for f and g. Wild-type mice were injected with PBS or PDAC cells and then injected with PDAC–YFP cells on day 10. The liver was removed 2 h (f, n = 4 for both groups) or 24 h (g, n = 5 for both groups) after intraportal injection. f, g, Images and quantification of tumour cells in the liver. Scale bars, 50 μm. Data representative of one experiment (e–g). h, Study design for i–l (n = 4 for both groups). i, Images of the liver and flow cytometric analysis. Scale bars, 1 cm. j, Quantification of PDAC–YFP cells. k, l, Images of the liver showing metastatic lesions (yellow, CK19) and Ki-67 (purple). Scale bars, 4 mm (k) and 200 μm (l). Data representative of at least three independent experiments (h–l). m, Study design for n and o (n = 5 for both groups). n, Gating strategy for identification of T cell subsets. Images from flow cytometric analysis of cells isolated from the liver of a wild-type mouse are shown. EM, effector memory. o, Quantification of CD4+ T cell subsets (top) and CD8+ T cell subsets (bottom) in the liver. Data representative of two independent experiments (m–o). p–s, Wild-type mice were orthotopically injected with PBS or PDAC cells (n = 4 for both groups). One group received anti-CD4 and anti-CD8 antibodies on days 8 and 13. Both groups were intraportally injected with PDAC–YFP cells on day 10. p, Flow cytometric analysis of peripheral blood and quantification of CD4+ and CD8+ T cells. q, Images of the liver, flow cytometric analysis, and quantification of PDAC–YFP cells. Scale bars, 1 cm. r, Images of the liver showing metastatic lesions (yellow, CK19), Ki-67 (purple, top), and CD4+ cells (purple, bottom) and CD8+ cells (brown, bottom). Scale bars, 200 μm (top) and 50 μm (bottom). s, Quantification of lesions and Ki-67+ tumour cells (top) and CD4+ cells and CD8+ cells (bottom). Data representative of one experiment (p–s). Statistical significance calculated using one-way ANOVA with Dunnett’s test (d) or two-tailed Mann–Whitney test (other panels). NS, not significant. Data represented as mean ± s.d.
Extended Data Fig. 3 Primary PDAC development induces expression of myeloid chemoattractants and activates STAT3 signalling in the liver.
a, Study design for b–e (n = 5 for both groups for b–d and n = 10 and 9 for control mice and NTB KPC mice, respectively, for e). b, Heat map showing differentially expressed genes in the liver. c, Enriched biological processes in the liver of NTB KPC mice. Left, significance; right, number of genes in each group. d, FPKM values for chemoattractant genes in the liver. e, mRNA levels of chemoattractant genes in the liver. Data representative of one experiment (a–e). f, Enrichment of IL-6–JAK–STAT3 signalling genes in the liver (n = 5 for control mice and NTB KPC mice). FDR, false discovery rate; NES, normalized enrichment score. Data representative of one experiment. g, Left, images showing hepatocytes (stained for albumin) and pSTAT3. Right, percentage of hepatocytes that were pSTAT3+ in control mice (n = 4), NTB KPC mice (n = 5), and TB KPC mice (n = 5). h, Left, images of F4/80+ cells and pSTAT3. Right, percentage of F4/80+ cells that were pSTAT3+ in control mice (n = 9), NTB KPC mice (n = 8), and TB KPC mice (n = 5). i, Images of F4/80+ cells and pSTAT1 in the liver of control mice (n = 9), NTB KPC mice (n = 8), and TB KPC mice (n = 5). Data pooled from two experiments (g–i). j, k, Wild-type mice were orthotopically injected with PBS (n = 7) or PDAC cells (n = 6). Left, images of hepatocytes (j), F4/80+ cells (k), and pSTAT3. Right, percentage of hepatocytes (j) and F4/80+ cells (k) that are pSTAT3+. Scale bars, 50 μm. Statistical significance calculated using ClueGO35 (c), two-tailed unpaired Student’s t-test (d), GSEA40 (f), one-way ANOVA with Dunnett’s test (g, h), and two-tailed Mann–Whitney test (other panels). NS, not significant. Data represented as mean ± s.d., except d, e, which are shown as box plots (centre line, median; box limits, upper and lower quartiles; whiskers, max and min values).
Extended Data Fig. 4 IL-6 promotes the formation of a pro-metastatic niche in the liver.
a–g, n = 5 and 6 for Il6+/+ mice and n = 4 and 5 for Il6−/− mice orthotopically injected with PBS and PDAC cells, respectively. a, Images of pSTAT3+ cells, myeloid cells, and fibronectin. Arrows indicate Ly6G+ cells. b, c, Images and quantification of COL1. d, Images of sinusoids (brown, stained with CD31) in the liver. e, mRNA levels of Lcn2, S100a8, S100a9, Ccl6, Cxcl1, Fn1, Col1a1, and Des in the liver. f, Images of pancreas and primary tumour stained with CD31 (brown), CK19 (yellow), and Ki-67 (purple). g, Quantification of the weight of pancreas or primary tumour (left), number of Ki-67+ tumour cells (middle), and vascular area (right). Data representative of one (b–d, Fn1, Col1a1, and Des in e–g) or two independent experiments (a, all other genes in e). h, Study design for i–k (n = 5 for all groups). All groups were injected with PDAC–YFP cells on day 10. i, j, Images of the liver showing metastatic lesions (yellow, CK19) and Ki-67 (purple). Scale bars, 4 mm (i) and 200 μm (j). k, Quantification of lesions (left) and Ki-67+ tumour cells (right). Data representative of one experiment (h–k). l, Study design for m–o (n = 4 and 5 for mice injected with PBS and treated with isotype control or anti-IL-6R antibodies, respectively; n = 7 and 8 for mice injected with PDAC cells and treated with isotype control or anti-IL-6R antibodies, respectively, unless indicated otherwise below). m, n, Images and quantification of pSTAT3+ cells, myeloid cells, and fibronectin. For fibronectin, n = 7 for mice injected with PDAC cells and treated with anti-IL-6R antibodies. Arrows indicate Ly6G+ cells. o, mRNA levels of Saa1 and Saa2. p, Study design for q–s (n = 7 and 8 for mice injected with PBS and treated with isotype control and anti-IL-6R antibodies, respectively; n = 7 for mice injected with PDAC cells and treated with isotype control and anti-IL-6R antibodies). All groups were injected with PDAC–YFP cells on day 10. q, r, Images of the liver and flow cytometric analysis. Scale bars, 1 cm. s, Quantification of PDAC–YFP cells. Data representative of two independent experiments (l–s). Scale bars, 50 μm unless indicated otherwise. Statistical significance calculated using one-way ANOVA with Dunnett’s test. NS, not significant. Data represented as mean ± s.d.
Extended Data Fig. 5 Non-malignant cells are the predominant source of IL-6.
a, Study design for b–e (n = 5 and 6 for Il6+/+ mice injected with PBS and PDAC cells, respectively; n = 4 and 5 for Il6−/− mice injected with PBS and PDAC cells, respectively). SN, supernatant. b, Concentration of IL-6 in pancreas supernatant and serum collected from indicated sites in Il6+/+ mice injected with PBS or PDAC cells. c, Concentration of IL-6 in pancreatic tumour supernatant and serum collected from indicated sites in Il6−/− mice injected with PBS or PDAC cells. Solid lines indicate data points from individual mice, and dashed lines indicate the lower limit of detection (b, c). d, Images of CK19 and Il6 mRNA in the pancreas and primary tumour. e, Images of Il6 mRNA in the liver and lung of Il6+/+ mice injected with PDAC cells. Data representative of two independent experiments (a–e). f, Study design for g and h (n = 4 for Il6+/+ mice injected with PDAC cells and n = 5 for human samples). g, Images of α-SMA, CD31, CK19, and Il6 mRNA in perivascular cells (top), stromal cells (bottom, left), endothelial cells (bottom, middle) and malignant cells (bottom, right) present within the mouse primary tumour. h, Images of α-SMA (yellow), CD31 (yellow), CK19 (yellow), and IL6 mRNA (brown) in perivascular cells (top, left), stromal cells (top, right), endothelial cells (bottom, left) and malignant cells (bottom, right) present within the human primary tumour. Data representative of one experiment (f–h). i, Study design for j and k (n = 5). j, k, Representative images of α-SMA, CK19, and Il6 mRNA detected in PanIN (j) and invasive PDAC (k). Data representative of one experiment (i–k). Scale bars, 50 μm. Statistical significance calculated using two-tailed Wilcoxon test. NS, not significant. Dagger, blood vessel; double dagger, PanIN lesion. Data from individual mice shown in b, c.
Extended Data Fig. 6 STAT3 signalling in hepatocytes promotes the formation of a pro-metastatic niche in the liver.
a, b, Representative images and quantification of pSTAT3+ hepatocytes (n = 3 technical replicates per condition) treated with or without IL-6 (a) or anti-IL-6R (b). Arrows indicate pSTAT3+ hepatocytes. SN, pancreatic tumour supernatant. Data representative of two independent experiments. For c–f and n, n = 4 for Stat3flox/flox mice and n = 8 and 7 for Stat3flox/flox Alb-cre mice orthotopically injected with PBS or PDAC cells, respectively. c, mRNA levels of Fn1 in the liver. d, Images of sinusoids (brown, stained for CD31) in the liver. e, Images of pancreas and primary tumour stained for CD31 (brown), CK19 (yellow), and Ki-67 (purple). f, Quantification of the weight of pancreas or primary tumour (left), number of Ki-67+ tumour cells (middle), and vascular area (right). Data representative of one experiment (c–f). g, Study design for h–l (n = 4 and 5 for Stat3flox/flox mice and Stat3flox/flox Alb-cre mice, respectively). All groups were injected with PDAC–YFP cells on day 10. h, Flow cytometric analysis. i, Quantification of PDAC–YFP cells. j, k, Images of the liver showing metastatic lesions (yellow, CK19) and Ki-67 (purple). Scale bars, 4 mm (j) and 200 μm (k). l, Quantification of lesions (left) and Ki-67+ tumour cells (right). Data representative of one experiment (g–l). m, Images of SAA detected by immunohistochemistry (brown, n = 5 for wild-type mice orthotopically injected with PBS or PDAC cells). Dashed lines and asterisks indicate sinusoids and hepatocytes, respectively. n, Images of Saa1 and Saa2 mRNA (brown) detected by RNA in situ hybridization. Data representative of one experiment (m, n). Scale bars, 50 μm unless indicated otherwise. Statistical significance calculated using one-way ANOVA with Dunnett’s test. NS, not significant. Data represented as mean ± s.d.
Extended Data Fig. 7 SAA promotes the formation of a pro-metastatic niche in the liver.
a, Concentration of circulating SAA in healthy donors (n = 69), patients with locally advanced PDAC (n = 28), and patients with liver metastases (n = 43). Data represented as a box plot (centre line, median; box limits, upper and lower quartiles; whiskers, max and min values). b, Images of SAA (yellow) and pSTAT3 (purple) in the liver of healthy donors and patients with PDAC with liver metastases. Dashed lines and asterisks indicate sinusoids and hepatocytes, respectively. c, Kaplan–Meier survival curve for patients with PDAC with liver metastases who had low (<250 μg ml–1, black, n = 21) or high (>250 μg ml–1, red, n = 22) levels of circulating SAA. d, Concentration of circulating SAA in patients with locally advanced NSCLC (n = 8) and patients with NSCLC with liver metastases (n = 13). Data shown as a box plot (centre line, median; box limits, upper and lower quartiles; whiskers, max and min values). e, Images of SAA (brown) in the liver of CRC patients with liver metastases. Dashed lines and asterisks indicate sinusoids and hepatocytes, respectively. Data representative of one experiment (a–e). f, g, n = 5 and 4 for Saa+/+ mice and n = 4 and 5 for Saa−/− mice orthotopically injected with PBS or CRC cells (MC-38), respectively. f, Quantification of myeloid cells, fibronectin, and COL1. g, Images of fibronectin (left) and COL1 (right). Data representative of one experiment (f, g). h–o, n = 5 for all groups unless indicated otherwise. h, Images of pSTAT3+ cells, myeloid cells, and fibronectin. i, j, Images and quantification of COL1. k, Images of sinusoids (brown, CD31) in the liver. l, mRNA levels of Lcn2, S100a8, S100a9, Ccl6, Cxcl1, Fn1, Col1a1, and Des in the liver. m, mRNA levels of Saa1 and Saa2. n, Images of pancreas and primary tumour stained for CD31 (brown), CK19 (yellow), and Ki-67 (purple). o, Quantification of the weight of pancreas or primary tumour (left), number of Ki-67+ tumour cells (middle), and vascular area (right). For weight, n = 4 for Saa+/+ mice injected with PDAC cells. Data representative of one (i–k, Fn1, Col1a1, and Des in l, n, o) or two independent experiments (h, all other genes in l, m). p, Study design for q–s (n = 8 and 5 for Saa+/+ mice and n = 5 and 7 for Saa−/− mice injected with PBS and PDAC cells, respectively). All groups were injected with PDAC–YFP cells on day 10. q, r, Images of the liver showing metastatic lesions (yellow, CK19) and Ki-67 (purple). Scale bars, 4 mm (q) and 200 μm (r). s, Quantification of lesions (left) and Ki-67+ tumour cells (right). Data representative of one experiment (p–s). Scale bars, 50 μm unless indicated otherwise. Statistical significance calculated using two-sided Mann–Whitey test (a, d), Mantel–Cox test (c), and one-way ANOVA with Dunnett’s test (other panels). NS, not significant. Data represented as mean ± s.d. unless indicated otherwise.
Extended Data Fig. 8 SAA is a downstream mediator of IL-6 signalling that drives myeloid cell accumulation and fibrosis within the liver.
a, Study design for b–d (n = 5 for all groups, except n = 4 for Saa−/− mice). b, Concentration of IL-6 in the serum collected from control mice (left), Il6−/− mice (middle), and Saa−/− mice (right) on indicated days. c, d, Images and quantification of pSTAT3+ cells, myeloid cells, fibronectin, and COL1. Arrows indicate Ly6G+ cells. Data representative of one experiment (a–d). e, Study design for f–h (n = 5 for mice injected with pLIVE-vector; n = 6 for mice injected with pLIVE-IL-6). f, Concentration of IL-6 in the serum. g, Images of the liver showing metastatic lesions (yellow, CK19) and Ki-67 (purple). h, Quantification of lesions (left) and Ki-67+ tumour cells (right). Data representative of one experiment (e–h). i, Study design for j (n = 5 for all groups, except n = 4 for Saa−/− mice injected with CCl4). j, Images and quantification of myeloid cells, fibronectin, and COL1. Arrows indicate Ly6G+ cells. Data representative of one experiment (i, j). Dashed lines indicate the lower limit of detection (b, f). Scale bars, 50 μm. Statistical significance calculated using two-tailed Mann–Whitney test (b, f, h) or one-way ANOVA with Dunnett’s test (other panels). NS, not significant. Data represented as mean + s.d. or mean ± s.d.
Extended Data Fig. 9 IL-6–STAT3–SAA signalling axis does not affect expression of MIF and TIMP1.
a, Study design for b–e (n = 5 and 4 for mice injected with PBS and PDAC cells). b, c, Images of CK19 (yellow) and TIMP1 (purple) in the pancreas, primary tumour, and liver. Data representative of at least three independent experiments (a–c). d, FPKM values of genes in the liver of control mice (n = 5) and NTB KPC mice (n = 5) obtained from QuantSeq 3′ mRNA sequencing. Data represented as box plots (centre line, median; box limits, upper and lower quartiles; whiskers, max and min values). e, mRNA levels of Timp1 in the liver of control mice (n = 6), NTB KPC mice (n = 7), and tumour-bearing KPC mice (n = 6) relative to Actb. Data representative of one experiment (d, e). f, j, n, Study designs for g–i, k–m and o–q, respectively. For f, n = 5 for Il6+/+ mice injected with PBS and n = 5 or 6 for Il6+/+ mice injected with PDAC cells. n = 4 and 5 for Il6−/− mice injected with PBS and PDAC cells, respectively. For j, n = 5 for all groups, except n = 4 or 5 for Saa+/+ injected with PDAC cells. For n, n = 4 for Stat3flox/flox mice and n = 8 and 7 for Stat3flox/flox Alb-cre mice injected with PBS and PDAC cells, respectively. g, k, o, mRNA levels of Mif and Timp1 in the indicated organs relative to Actb. h, l, p, Images of CK19 (yellow) and TIMP1 (purple) in the pancreas and primary tumour. i, m, q, Concentration of TIMP1 in the serum. Data representative of one experiment (f–q). Scale bars, 50 μm. Statistical significance calculated using one-way ANOVA with Dunnett’s test (i, m, q) or two-tailed Mann–Whitney test (other panels). NS, not significant. ND, not detected. Data represented as mean ± s.d. unless indicated otherwise.
Extended Data Fig. 10 Primary PDAC development induces a systemic response that promotes the formation of a pro-metastatic niche in the liver.
a, Study design for b–g (n = 4 and 8 for CD45.2 mice injected with PBS (group 1) and PDAC cells (group 2), respectively). b, Assessment of chimaerism in parabiotically joined mice. Flow cytometric analysis of peripheral blood gated on CD45.1+ and CD45.2+ cells as a percentage of CD45+ cells. c, Dorsal (top) and ventral (bottom) views of parabiotically joined mice. Dagger, site of laparotomy for orthotopic injection; double dagger, skin suture for parabiotic joining. d, Images and quantification of myeloid cells and fibronectin. Arrows indicate Ly6G+ cells. e, mRNA levels of Saa. f, Concentration of IL-6 in pancreas supernatant and serum. Solid lines indicate data points from individual mice. Dashed lines indicate the lower limit of detection. g, Concentration of circulating SAA. Data representative of one (f, g) or two independent experiments (a–e). h, i, n = 5 for all groups. h, Images of myeloid cells and fibronectin in the lung of control mice, NTB KPC mice, and tumour-bearing KPC mice. i, Quantification of myeloid cells and fibronectin. Data representative of one experiment (h, i). j, Study design for k–m. k, n = 4 and 5 for Il6+/+ mice and n = 5 and 4 for Il6−/− mice injected with PBS and PDAC cells, respectively. For l, n = 5 for all groups. For m, n = 4 for Stat3flox/flox mice and n = 8 and 7 for Stat3flox/flox Alb-cre mice injected with PBS and PDAC cells, respectively. k–m, Quantification of myeloid cells and fibronectin. Data representative of one experiment (j–m). n, Study design for o (n = 4 for all groups of mice). All groups of mice were injected with PDAC–YFP cells on day 10. o, Quantification of PDAC–YFP cells. Data representative of one experiment (n, o). p, Conceptual model of generation of pro-metastatic niche in the liver. Scale bars, 50 μm. Statistical significance calculated using two-sided Mann–Whitney test (d, e, g), two-sided Wilcoxon test (f), and one-way ANOVA with Dunnett’s test (other panels). NS, not significant. Data represented as mean ± s.d., except d, e, g, which are shown as box plots (centre line, median; box limits, upper and lower quartiles; whiskers, max and min values). Data from individual mice are shown in f.
Supplementary information
Supplementary Information
This file contains Supplementary Data 1 and Supplementary Tables 1-4.
Source data
Rights and permissions
About this article
Cite this article
Lee, J.W., Stone, M.L., Porrett, P.M. et al. Hepatocytes direct the formation of a pro-metastatic niche in the liver. Nature 567, 249–252 (2019). https://doi.org/10.1038/s41586-019-1004-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-019-1004-y
- Springer Nature Limited
This article is cited by
-
The double-edged effects of IL-6 in liver regeneration, aging, inflammation, and diseases
Experimental Hematology & Oncology (2024)
-
Hepatocytes coordinate immune evasion in cancer via release of serum amyloid A proteins
Nature Immunology (2024)
-
Cell competition in primary and metastatic colorectal cancer
Oncogenesis (2024)
-
From biology to the clinic — exploring liver metastasis in prostate cancer
Nature Reviews Urology (2024)
-
Serum amyloid A promotes glycolysis of neutrophils during PD-1 blockade resistance in hepatocellular carcinoma
Nature Communications (2024)