Abstract
The development of neural circuits relies on axon projections establishing diverse, yet well-defined, connections between areas of the nervous system. Each projection is formed by growth cones—subcellular specializations at the tips of growing axons, encompassing sets of molecules that control projection-specific growth, guidance, and target selection1. To investigate the set of molecules within native growth cones that form specific connections, here we developed growth cone sorting and subcellular RNA–proteome mapping, an approach that identifies and quantifies local transcriptomes and proteomes from labelled growth cones of single projections in vivo. Using this approach on the developing callosal projection of the mouse cerebral cortex, we mapped molecular enrichments in trans-hemispheric growth cones relative to their parent cell bodies, producing paired subcellular proteomes and transcriptomes from single neuron subtypes directly from the brain. These data provide generalizable proof-of-principle for this approach, and reveal molecular specializations of the growth cone, including accumulations of the growth-regulating kinase mTOR2, together with mRNAs that contain mTOR-dependent motifs3,4. These findings illuminate the relationships between subcellular distributions of RNA and protein in developing projection neurons, and provide a systems-level approach for the discovery of subtype- and stage-specific molecular substrates of circuit wiring, miswiring, and the potential for regeneration.
Similar content being viewed by others
Main
Neurons are cells with exceptional structure, characterized by large intracellular distances punctuated with molecular specializations dedicated to local functions. Key subcellular specializations in the establishment of nascent circuitry are growth cones (GCs) at the tips of extending axons1. Although the in vivo molecular diversity of neuron subtypes is now well appreciated5, the in vivo projection-specific molecular diversity of growth cones remains unknown and experimentally inaccessible with current approaches. To enable quantitative and systems-level subcellular readouts from distinct subtype-specific growth cones in the brain, we developed an experimental approach of growth cone sorting and ‘subcellular RNA–proteome mapping’.
To purify GC subtypes from the brain, we fluorescently label neuron populations in vivo, from which we isolate total GCs by subcellular fractionation6,7 (Fig. 1a). Isolated GCs display intact membranes, are enriched in GC marker proteins, and retain encapsulated sequencing-quality RNA (Extended Data Fig. 1). To specifically purify fluorescently labelled GC subtypes from bulk-isolated GCs of the brain, we modified and optimized a fluorescence-activated cell sorter with custom optics and fluidics to sort and collect fluorescent GCs directly (see Methods).
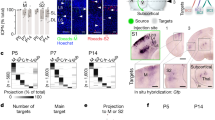
a, Schematic demonstrating two-colour small-particle sorting to separate GFP from RFP GCs. (1) Brains from a GFP and an RFP mouse were homogenized together. (2) Subcellular fractionation yields a GC fraction that contains red and green GCs. (3) GCs were sorted to collect pure green GCs from the mix. b, Small particle sorter plot of forward- and side-scatter of GC sample (blue) overlaid on size-standard beads (grey) for size comparison. Isolated GCs are sub-micrometre particles with a size range centred on 0.5 μm. c, Sorter plot of mixed GFP and RFP GC suspension (as schematized in a), showing separation of GCs into red and green populations. Collection gate to isolate pure green GCs is indicated. d, Collected GCs gated from c were re-analysed revealing GFP-specific GCs. Sorting enables specific purification of gated GCs. n = 3 independent biological replicates.
To monitor and verify our ability to purify individual fluorescent GCs, we used mouse lines that ubiquitously express a variant of red or green fluorescent protein (RFP or GFP, respectively). GFP- and RFP-labelled brains were fractionated together, so that isolated GCs were either red or green, but not both. We loaded isolated GCs into the modified sorter, calibrated gating based on size-standard beads, and determined the optimal conditions for the separation of individual single-colour GCs. To verify collection, we gated on fluorescence to collect green and exclude red GCs. By re-analysing the collected sample, we determined that we indeed isolated pure green GCs (Fig. 1), establishing the conditions and feasibility for the use of this approach to isolate pure subpopulations of labelled circuit-specific GCs from the brain.
We present here the first, to our knowledge, application of circuit-specific GC sorting on the trans-hemispheric projection formed by callosal projection neurons5,8. We specifically labelled upper-layer cortical neurons by in utero electroporation9 in one hemisphere with a plasmid that expressed GFP. Three days after birth (postnatal day (P) 3), we purified trans-hemispheric GCs using GC sorting on the contralateral hemisphere, in which only growing callosal axons that have crossed the midline are fluorescent. Protein was extracted from sorted GCs, and analysed by mass spectrometry to reveal the GC sub-proteome, which we symbolize as {P}GC, of growing trans-hemispheric axons (Fig. 2 and Supplementary Table 1).
As anticipated, a large constituent of {P}GC comprises a nexus of cytoskeletal, membrane and signalling proteins, typically associated with GC functions in axon guidance1. Confirming readout specificity, GFP itself—the marker for selection—was also detected by mass spectrometry in sorted GCs. Beyond these, {P}GC displays a rich range of prominent functional complexes dedicated to anabolism and growth, as well as catabolism and turnover. Notably, {P}GC includes a distinct set of RNA-binding proteins with known dual roles in nuclear spliceosomes as well as cytosolic ribonucleoprotein particles, raising the possibility of novel RNA-binding complexes that regulate GC-localized RNA. The chaperonin complex, specialized in the folding of nascent actin and tubulin polypeptides into functional proteins10, is also robustly present in {P}GC. Together, these findings support local GC synthesis and turnover of proteins in vivo, including cytoskeletal elements, as suggested by in vitro studies11,12.
As a first biological investigation using the new ability to acquire stage- and circuit-specific subcellular molecular data from the brain, we asked to what extent the local transcriptome matches the local proteome in developing neuron projections in vivo. Pioneering studies detected mRNA transcripts in distal processes of cultured neurons11,13,14,15, giving rise to the idea of local translation producing local sub-proteomes in different parts of the neuron16. To examine this in vivo, and to determine the extent to which this happens across gene groups, we combined newly developed subtype-specific GC sorting with subtype-specific neuron cell body sorting17,18,19. Performing these approaches in parallel, we obtained paired, quantitative, internally normalized sub-transcriptome and sub-proteome measurements from diametric GC and cell body (soma) compartments from the developing trans-hemispheric projection of upper-layer cortical neurons in vivo.
We labelled layer II/III neurons of the mouse sensorimotor cortex with membrane-RFP and nuclear-GFP by in utero electroporation. At P3, electroporated hemispheres were triturated into a soma suspension, and sorted for green fluorescence to collect labelled neuron somata. Contralateral hemispheres were fractionated to extract GCs, and GC-sorted for red fluorescence to collect corresponding trans-hemispheric axon GCs. We extracted RNA and protein from sorted GCs and their parent somata, and performed RNA sequencing (RNA-seq) and mass spectrometry to obtain paired measurements from the sub-transcriptomes {R}GC and {R}soma, and the sub-proteomes {P}GC and {P}soma (Fig. 3). This workflow yielded a high-confidence dataset of 955 genes with quantified GC-to-soma ratios for RNAs and proteins (λR and λP, respectively). These values enabled us to map genes based on paired subcellular mRNA and corresponding protein distributions within the developing projection. This revealed pronounced divergence across gene groups, with distinct clusters emerging based on paired distributions (Extended Data Figs. 2–6, Supplementary Tables 2–7 and Supplementary Discussion).
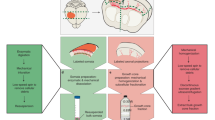
a, Selective labelling of upper layer callosal projection neurons with nuclear-GFP (nuc-GFP; green) and membrane-RFP (mem-RFP; red) by in utero electroporation. Nascent callosal projection at P3 displays ipsilateral somata with green nuclei, and trans-hemispheric axons with red GCs. Scale bar, 600 μm and 100 μm (insets). b, Schematic of subcellular RNA–proteome mapping workflow: neuron labelling; postnatal harvesting; cell dissociation of ipsilateral hemispheres for sorting of GFP+ cell bodies; subcellular fractionation of contralateral hemispheres to isolate GC fraction for RFP+ GC sorting. c, FACS plots with gates for GFP+ somata and RFP+ GCs used to collect trans-hemispheric GCs and their parent cell bodies; RNA-seq and mass spectrometry yield paired measurements of sub-transcriptomes {R}GC and {R}soma, and sub-proteomes {P}GC and {P}soma, from which GC-over-soma ratios of mRNA (λR) and protein (λP) were calculated for each gene. n = 6 independent biological replicates, 3–6 litters each.
A notable pattern in the clustering of λR values emerged from RNA mapping. Although most transcripts are depleted from GCs, we observed that transcripts containing a non-canonical sequence, known as the 5′ terminal oligopyrimidine (TOP) motif3, display marked and consistent GC enrichment. Of the 83 TOP transcripts detected, most displayed trends of GC enrichment, with about half enriched with statistical significance, whereas no TOP transcripts were significantly depleted from GCs (Fig. 4). These results were further validated by quantitative PCR (qPCR) of select TOP and non-TOP reference transcripts, and TOP transcripts were directly visualized in GCs using single-molecule RNA in situ hybridization (Fig. 4c and Extended Data Fig. 7). Collectively, TOP transcripts account for approximately 80% of all significantly GC-enriched transcripts in the mapping dataset (Supplementary Table 4).
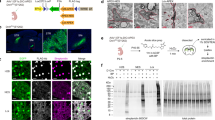
a, Transcript classes schematized according to dependence on mTOR for translation. b, Volcano plot of GC–soma RNA mapping, coloured according to classes in a. λR values plotted for each transcript versus P value (on logarithmic scales). Transcripts visualized in c are labelled. Significance thresholds set to a 0.05 permutation-based false discovery rate (FDR). n = 6 independent biological replicates, 3–6 litters each. c, Single-molecule RNA in situ hybridization (red) for Rack1 (non-ribosomal TOP), Rplp0 (ribosomal TOP), and control transcript Ppib (soma-mapped, canonical) on cultured callosal projection neurons labelled with membrane-GFP (green). Nuclei were stained with DAPI (blue). Scale bar, 10 μm. See Extended Data Fig. 7 for full set. A total of 92, 103 and 84 GCs were imaged for Rack1, Rplp0 and Ppib probes, respectively. n = 4 biological replicates from independent in utero electroporations.
Fewer than 100 genes produce transcripts with bona fide TOP motifs20. However, they collectively produce 5–20% of the total proteome of a cell21,22. TOP transcripts encode the proteins of the translation machinery itself, most notably the protein subunits of ribosomes and translation initiation factors3. As such, their expression is tightly coupled to cellular growth. The TOP motif itself functions as the ON/OFF switch for translation, and is under the direct control of mTOR3,23,24, a hub kinase that integrates growth-factor signalling with the availability of nutrients, energy, and oxygen25. Although 99.8% of all transcripts respond to mTOR signalling with only modest (approximately 20%) changes in translation, TOP translation is fully mTOR-dependent in an all-or-none manner. By contrast, there is a small group of non-canonical transcripts that contain internal ribosome entry sites, or lack poly(A) tails, making their translation entirely mTOR-independent24. Interestingly, this group is the diametric opposite of the TOP group in our dataset, comprising the most extreme outliers of soma enrichment (Fig. 4a, b). These data reveal notable subcellular polarization of the transcriptome within growing projection neurons based on mTOR dependence for translation.
Given the degree of TOP mRNA enrichment in GCs, together with their strict dependence on mTOR for expression, we investigated whether developing projection neurons localize endogenous mTOR to their axon GCs, as previously suggested by overexpression26. We identify mTOR and the mTOR-binding proteins LARP1 and raptor as specifically enriched in GCs at levels comparable to the cardinal GC marker GAP43. LARP1 directly binds and regulates the TOP motif4,27, whereas raptor is a key subunit of mTOR complex 1 (mTORC1)23. We directly visualized these proteins in callosal projection neuron GCs, and observed that mTOR, LARP1, and mTORC1 components specifically accumulate in dense local foci in the ‘palms’ and ‘cuffs’ of axon GCs (Fig. 5 and Extended Data Figs. 8, 9). This pattern is distinct from the granular immunolabelling throughout the neuron of the mTORC2 marker RICTOR and the lysosome marker LAMP1. These data collectively indicate the existence of distinct subcellular foci of mTORC1, LARP1, and their target TOP mRNAs in the GCs of developing axon projections.
a, Western blots of mTOR pathway proteins and controls in homogenate (input) and GC fraction (fr.) preparations. The GC marker GAP43 is a positive control for enrichment; the Golgi marker GM130 is a negative control. b, Quantification of GC enrichment expressed as band intensity ratios of GC fraction over input from blots in a, normalized to corresponding GAP43 ratio (marked by line). mTOR, LARP1, and raptor (mTORC1 marker) display high GC enrichment comparable to GAP43. TSC1, RICTOR (mTORC2 marker), and LAMP1 (lysosome marker) are present in GCs at comparable levels to actin and tubulin. Data are mean ± s.e.m. from n ≥ 3 independent fractionation experiments from distinct litters (n = 7 for mTOR; n = 5 for LARP1, raptor and RICTOR; n = 3 for LAMP1 and TSC1; n = 8 for TUBB3; n = 6 for ACTB and GM130). c, GCs from cultured callosal projection neurons immunolabelled for endogenous mTOR pathway proteins (red in overlays, heat-mapped in underlying panels). mTOR, LARP1, TSC1, and raptor appear in dense local foci within GCs. RICTOR and LAMP1 appear in fine granules distinct from GC foci. Bar (bottom right) indicates the heat-map colour range, and the 10-μm scale. See Extended Data Fig. 8 for full set. In total, 83 GCs were imaged for mTOR, 47 for LARP1, 42 for LAMP1, 49 for TSC1, 26 for raptor, and 30 for RICTOR, from a minimum n = 3 biological replicates from independent in utero electroporations.
The presence of local mTOR foci in callosal projection neuron axon GCs prompted us to examine whether mTOR signalling is necessary for the formation of the trans-hemispheric projection itself during development. We investigated this in vivo using two genetic strategies. In one approach, we electroporated callosal projection neurons with a dominant-negative subunit of PI3 kinase28 (PI3K-DN), a crucial growth-factor signalling pathway that activates mTOR. Compared to matched GFP-only controls, PI3K-DN causes pronounced perturbation in neuron migration, as reported previously29, as well as marked loss of callosal axon extension. In a parallel approach, we acutely knocked out mTOR by Cre electroporation in floxed-mTOR mice (mTOR-KO). mTOR-KO does not significantly affect migration, but instead, specifically prevents the extension of axons across the corpus callosum (Fig. 6 and Extended Data Fig. 10), collectively confirming that mTOR signalling is necessary for trans-hemispheric axon growth.
a, b, Callosal projection neuron migration (a) and callosal axon growth (b) were examined at P3 in brains electroporated with GFP (control), dominant-negative PI3K (PI3K-DN), or Cre in mice with homozygous floxed-mTOR alleles for conditional knockout (mTOR-KO) in electroporated neurons. Inhibition of PI3K signalling hindered the migration of somata (quantified as the percentage of somata en route versus those at destination layers II/III) (a), and stalled callosal axon growth (quantified as normalized GFP intensity in serial bins from ipsilateral-to-contralateral callosum) (b). Deletion of mTOR specifically resulted in failure of callosal axon growth. Scale bars, 100 μm. Data are mean ± s.e.m. from n = 4 mice from different litters for each condition.
Together, these findings place mRNAs of the translation machinery, along with their obligate regulator, mTOR, at the leading edge of growing long-range axon projections. Without mTOR signalling, these projections fail to form. We propose a developmental interpretation for these observations, in which the supply of cellular translation machinery is coupled to axon extension through local signalling. This subcellular organization might be a transient feature of the axon-extension phase of projection neuron development, physically positioning mTOR at sites of most intense cellular growth. It is intriguing to speculate that mTOR foci in GCs might enable sensing of target-derived growth signals locally to globally coordinate transitions of the developmental program of a neuron driven by target-derived signals. It will be interesting to examine this non-standard developmental model directly, along with its implications for axon growth, and possibly regeneration30 (see Supplementary Discussion). Finally, this new line of experimentation using subtype-specific GC sorting and quantitative subcellular RNA–proteome mapping provides a generalizable approach that enables molecular investigations that compare subtype- and stage-specific GCs, GCs from mutant, regenerative, non-regenerative, or reprogrammed neurons to discover molecular specificities behind circuit development, miswiring, and possibly regeneration.
Methods
Animals
Animal experimental protocols were approved by the Harvard University Institutional Animal Care and Use Committee, and complied with all relevant ethical regulations regarding animal research. Experiments using wild-type mice were performed on outbred strain CD1 mouse pups of both sexes (Charles River Laboratories). Mice ubiquitously expressing GFP (Fig. 1) are from the transgenic strain B6 ACTb-EGFP (JAX stock 003291) expressing GFP under the CAG promoter31. Mice ubiquitously expressing RFP (Fig. 1) correspond to a knock-in strain ubiquitously expressing the RFP variant tdTomato, under the CAG promoter from the ROSA26 locus. We created this strain by breeding Ai9 strain32 (JAX stock 007909) females with Vasa-Cre strain (JAX stock 006954) males expressing Cre in embryonic germ cells33 (leading to the removal of a floxed-stop cassette from the original conditional tdTomato knock-in allele) and cross-breeding the red progeny. Both GFP and RFP mouse lines were back bred into the FVB background (JAX strain FVB/NJ) by selecting for fluorescence for over seven generations. For conditional mTOR-knockout experiments (Fig. 6 and Extended Data Fig. 10), we used floxed-mTOR mice (JAX stock 011009) homozygous for mTOR alleles harbouring loxP sites flanking exons 1–5 of the Mtor gene34. No statistical methods were used to predetermine sample size. All animals analysed were P3 or younger, thus no sex determination was attempted. Analyses are thought to include animals of both sexes at approximately equal proportions. Investigators were not blinded to allocation during experiments and outcome assessment unless stated otherwise.
DNA constructs
The following plasmid DNA expression constructs were used for in utero electroporations: membrane-GFP (used in Figs. 2, 4 and 5 and Extended Data Figs. 7–9) is a GFP–GPI fusion construct provided by A.-Katerina Hadjantonakis35; membrane-RFP and nuclear-GFP (used in Fig. 3) corresponds to a 2A bi-cistronic gene encoding myristoylated-tdTomato and Histone2B-GFP from plasmid pCAG-TAG (Addgene plasmid 26771), provided by S. Srinivas4. Expression plasmid pCAG-GFP36 (gift from C. Cepko, Addgene plasmid 11150) was used for GFP control electroporations (used in Fig. 6 and Extended Data Fig. 10). The PI3K-DN plasmid was produced using the following cloning procedures: (1) PCR amplification of the open reading frame of the PI3K regulatory subunit 1 (Pik3r1, accession number: NM_001077495) from mouse brain cDNA; (2) site-directed mutagenesis using QuikChange (Agilent) to delete 36 internal residues (Q478–K513) corresponding to the catalytic subunit-binding site, thus producing the dominant-negative construct PI3K-DN, as previously reported28; and (3) PI3K-DN was subcloned into pCAG-GFP in-frame with GFP, downstream of a 2A element to create a bicistronic expression vector producing GFP and PI3K-DN. Plasmid expressing Cre and GFP for electroporations into floxed-mTOR animals was produced equivalently by subcloning Cre downstream of GFP-2A in pCAG-GFP.
In utero electroporation
Electroporations were performed in utero on embryonic day 15, as previously described37, resulting in specific expression of plasmid DNA in cortical layer II/III neurons, including inter-hemispheric projection neurons (upper layer callosal projection neurons). Dense DNA solution was injected into one lateral ventricle using a pulled glass micropipette. Five current pulses of 35 V were applied, targeting nascent sensorimotor areas of the cortical plate. After term birth, electroporated mouse pups were screened for unilateral cortical fluorescence using a fluorescence stereoscope.
GC fractionation
GC fractions were obtained using modifications of methods described previously6,7. In brief, forebrains of P3 mouse pups were rapidly chilled and homogenized in 0.32 M sucrose buffer supplemented with 4 mM HEPES, HALT protease and phosphatase inhibitors (Thermo), and 1 U ml−1 RNase inhibitors (Promega), with 13 strokes at 900 r.p.m. in a glass-Teflon potter. Postnuclear (input) homogenates were obtained as supernatants after centrifugation at 1,700g for 15 min. Inputs were layered onto 0.83 M sucrose and a 2.5 M sucrose cushion, and spun cooled in a fixed vertical rotor (VTi50, Beckman) at 250,000g for 50 min. The GC fraction was extracted from the 0.32–0.83-M interface.
GC protection assays
To investigate the integrity of GCs in GC fractions, and the specific encapsulation of GC protein and RNA by continuous GC membrane, we applied two parallel hydrolytic enzyme ‘protection assay’ approaches. We performed protection assays with RNase (RNase One, Promega) to test for GC RNA protection, and with protease (trypsin) to test for GC protein protection. Test samples were incubated with either 0.025% trypsin or 30 U ml−1 RNase at 4 °C for 90 min with constant rotation. In parallel with test samples, positive control samples contained 0.3% Triton X-100 detergent in addition to enzyme to disrupt GC membrane integrity and allow RNase or protease access to RNA and protein both outside and inside GCs (Extended Data Fig. 1). Negative control samples contained detergent but no hydrolytic enzyme. Enzyme concentrations, conditions and durations were titrated for complete degradation of all protein and RNA in the Triton X-100-containing control samples, without observable loss in negative control samples.
RNA and proteins remaining after treatment and incubation were measured using capillary electrophoresis (2100 Bioanalyzer, Agilent) or standard western blot, respectively (Extended Data Fig. 1). RNA and protein signals that survived treatment in test samples represent molecules protected inside intact GCs.
GC and soma sorting and collection
GC sorting
To isolate labelled GCs, we took a subcellular fluorescence-based sorting approach, similar in concept to the previously developed FASS approach for synaptosomes38. We customized a Special Order Research Program (SORP) FACSAriaII (BD Instruments) fluorescence-activated sorter for small particle detection. Forward- and side-scatter were detected using a photomultiplier tube (PMT) and a 300 mW 488 nm laser with reduced beam height (6 ± 3 μm) and custom lens assembly with noise-reducing filter and pico-motor focus. Scatter measurements were based on signal peak height and plotted in log mode. Before loading GCs, bleach followed by filtered water was run through the fluidics for at least 20 min. GCs were loaded onto a cooled sample platform and sorted through a cuvette flow cell with a 70-μm nozzle running at 70 psi with PBS as sheath fluid fed through a 0.1-μm filter. Bulk GC fractions in sucrose were diluted 3–6-fold in PBS immediately before loading into the sorter. Comparing the forward- and side-scatter profile of particles in the GC fraction to sub-micrometre polystyrene size-standard beads (BD), most GCs ranged from 0.3 to 0.8 μm in diameter (Fig. 1b), consistent with previous electron microscopic analysis of a heterogeneous bulk GC fraction6. Appropriate bead sizes were used to calibrate drop delays and collection parameters.
Soma sorting
We isolated fluorescent somata of layer II/III inter-hemispheric projection neurons from electroporated hemispheres using established approaches17,19,39. The electroporated areas of the cortical plate were micro-dissected to remove meninges and the ventricular zone (VZ) under a fluorescence stereoscope (Nikon). The micro-dissected pieces of cortex containing fluorescent postmitotic neuron cell bodies were dissociated for sorting as previously described17,18,39,40,41. We used a different customized SORP FACSAriaII equipped with a 100 mW 488 nm laser with large beam height, and an 85 μm nozzle at 45 psi, to collect GFP-labelled neuronal somata (Fig. 3c).
Sorted GC–soma collection for downstream RNA-seq and mass spectrometry
We collected GCs directly into guanidinium-based buffer RLT (Qiagen) containing 2-mercaptoethanol. For each biological replicate, we used an average of 6 electroporated brains, from which we collected on average 2,000,000 fluorescent GCs, and 200,000 fluorescent parent cell bodies. Collection run times averaged 12 h for GCs, and 1.5 h for somata. In addition to electroporation, we tested and successfully sorted fluorescent GCs using alternative methods of fluorescent labelling, including conditional mouse lines, and lipophilic dyes. We extracted both RNA and protein from each sorted sample using a commercial column-based kit according to the manufacturer’s protocol (AllPrep DNA/RNA/Protein kit, Qiagen). Total GC and total soma RNA samples were eluted into 14 μl H2O, and frozen until use for cDNA library preparation and RNA-seq. Total protein from GC and soma samples was precipitated, and pellets were frozen until processing for mass spectrometry.
Mass spectrometry
Protein samples were subjected to on-pellet processing and liquid chromatography–tandem mass spectrometry (LC–MS/MS) on an LTQ Orbitrap Elite (Thermo Fischer) supplied by a NanoAcquity UPLC pump (Waters) for label-free quantitative mass spectrometry. Sample order was randomized between replicate experiments. Specifically, protein pellets were resuspended in 8 M urea, reduced at 56 °C with TCEP, alkylated with iodoacetamide, and digested for 4 h with trypsin. Remaining pellets were sonicated in 80% acetonitrile, and digested in trypsin overnight. Tryptic peptides were separated on a 100-µm microcapillary trapping column packed with 5 cm C18 Reprosil resin of 5 µm particles with 100 Å pores, followed by a 20 cm analytical column of Reprosil resin of 1.8 µm particles with 200 Å pores (Dr. Maisch GmbH). Separation was achieved with a 5–27% acetonitrile gradient in 0.1% formic acid over 90 min at 200 nl min−1. Peptides were ionized by electrospray with 1.8 kV on a custom-made electrode junction sprayed from fused silica pico tips (New Objective). The LTQ Orbitrap Elite was operated in data-dependent mode. Mass spectrometry survey scan was performed in the 395–1,800 m/z range at a resolution of 6 × 104, followed by selection of the 20 most intense ions (TOP20) for collision induced dissociation (CID)-MS2 analysis in the ion trap, using a precursor isolation window width of 2 m/z, an AGC setting of 10,000, and maximum ion accumulation of 200 ms. Singly charged ion species were not subjected to CID fragmentation. Normalized collision energy was set to 35 V and an activation time of 10 ms. Ions in a 10 p.p.m. m/z window around ions selected for MS2 were excluded from further selection for fragmentation for 60 s. The same TOP20 ions were subjected to high collision energy dissociation (HCD)-MS2 analysis in the Orbitrap. The fragment ion isolation width was set to 0.7 m/z, AGC was set to 50,000, the maximum ion time was 200 ms, normalized collision energy was set to 27 V and 1 ms activation time for each HCD-MS2 scan. We analysed output data using MaxQuant and Perseus software (see ‘RNA–proteome mapping data analysis’).
RNA-seq
cDNA libraries from sorted GC and soma samples were prepared from equal masses of RNA using random hexamer primers depleted of rRNA sequences (Ovation Single Cell RNA-Seq System, Nugen). Sample order was randomized between replicate experiments. We prepared libraries according to the manufacturer’s protocol, using six amplification cycles to avoid over-amplification and signal saturation. GC and soma libraries were barcoded and sequenced together on an Illumina Hiseq 2500 sequencer, generating 100-bp paired-end reads. RNA-seq reads were processed using bcbio-nextgen v0.9.5, aligning to GRCm38 with the STAR aligner42 and quantifying counts per gene with Sailfish43 using the Ensembl annotation. We examined a variety of quality control metrics to detect poorly performing samples, using a combination of Qualimap44 and FastqQC (Babraham Bioinformatics).
RNA–proteome mapping data analysis
Replicates and quality control
When the amount of material permitted, we subdivided RNA and protein from a single biological replicate to perform technical replicate RNA-seq and mass spectrometry runs (duplicates or triplicates, depending on available material). Technical replicates included loading partial amounts (halves or thirds) to assess quantitative linearity of output measurements. Technical replicates confirmed low variance between instrument runs and quantitative linearity of readouts. Readouts from technical replicates were merged into a combined data set as a single biological replicate in subsequent statistical analyses.
Biological replicates were subjected to principal component analysis, and cross-correlation analysis. We did not further consider replicates that did not pass quality control tests or were consistent outliers, displayed low complexity, or exhibited identifiable library artefacts. Of the six biological replicates performed for each compartment, five GC RNA-seq, five soma RNA-seq, four GC mass-spec, and six soma mass spectrometry replicates passed quality control (Extended Data Fig. 2). These readouts were considered further for quantification and subcellular RNA–proteome mapping. Individual values of each biological replicate for each gene can be accessed on the Harvard Dataverse repository.
Label-free quantitative mass spectrometry analysis
Mass spectrometry readouts were analysed using the MaxQuant software package following the MaxLFQ method for label-free quantitative (LFQ) proteomics45. Mass spectra were assigned to corresponding peptides and proteins with a 7 p.p.m. peptide tolerance, and peptide-to-spectrum-match FDR of 0.05, and protein matching with a minimum of one unique peptide and FDR of 0.01 using the Andromeda search engine against version 83 of the Ensembl annotation. Proteins that were detected in the same compartment in at least two biological replicates were considered bona fide constituents of the sub-proteomes {P}GC and {P}soma (Supplementary Tables 2 and 3).
For quantification of GC-to-soma ratios λP, LFQ intensities for the proteins in {P}GC and {P}soma were extracted from MaxQuant to the Perseus Platform46 for matrix processing and statistical analysis. Raw LFQ values were normalized across biological replicates of the same compartment using quartile alignment and width adjustment of distributions, so that distribution peaks align at 1. Values were log2 transformed, and imputation was used to assign baseline non-zero values to represent lack of detection. Imputed values were randomly assigned from a distribution simulating baseline detection noise with a distribution peak downshifted by 2 standard deviations and a width of 0.25 standard deviations of the distribution of detected values. This provided non-zero values for ratiometric determination of λP (GC mean normalized LFQ intensity over soma mean normalized LFQ intensity; Supplementary Table 2). Volcano plots of GC–soma represent λP values on the x axis, and two-tailed t-test values across biological replicates on the y axis. Significance thresholds were set to a 0.05 permutation-based FDR.
RNA-seq quantitative analysis and proteome matching
RNA-seq data were internally filtered for transcripts that were detected in at least three of the five GC, or three of the five soma replicate datasets to produce bona fide sub-transcriptomes {R}GC and {R}soma. Sailfish43 transcripts per million (TPM) counts for each gene were matched to the proteome dataset through Ensembl gene IDs, yielding 955 genes with complete RNA–proteome mapping data (Supplementary Table 4). Sailfish raw counts were analysed with Perseus similar to LFQ protein data. Values were quartile aligned with peaks at 1, log2 transformed, and missing values were imputed from the noise, as above. Ratiometric determination of GC enrichment λR (GC mean normalized TPM counts over soma mean normalized TPM counts), and GC–soma mapping volcano plots were performed as with proteins above.
Gene annotation
RNA and protein sequences were aligned and paired using version 83 of the Ensembl annotation. Uniprot and RefSeq entries were matched to Ensembl gene ID using the Synergizer service47. Gene Ontology (GO) annotation was assigned based on the UniProt database. Ad hoc gene groups corresponding to cell compartments were annotated based on the COMPARTMENTS subcellular localization database48. Protein interactions were annotated based on version 10.0 of the STRING database49.
RNA–protein annotation enrichment
1D and 2D annotation enrichment analysis was performed as described50 using the combined dataset of RNA-seq and mass spectrometry biological replicates as input. Annotations considered were GO terms from three categories (‘molecular process’, ‘biological function’ and ‘cellular component’), as well as gene groups as listed in Supplementary Table 7. Significance was determined using a two-sided Benjamini–Hochberg corrected FDR of 0.02 (Supplementary Tables 3, 5 and 6).
RNA analyses
Native RNA gel electrophoresis
We loaded purified GC and input homogenate RNA with GelRed, and ran them on a native 1% agarose gel in TBE (89 mM Tris-HCl, pH 7.8, 89 mM borate, 2 mM EDTA) alongside single-strand RNA ladder (NEB).
RNA capillary electrophoresis
We performed analysis of RNA on an Agilent 2100 Bioanalyzer following the manufacturer’s protocol. For mRNA-specific profiles, we used oligo (dT)25 beads to purify polyA-containing transcripts before analysis.
RT–PCR
We used equal masses of RNA from purified GC and input to make GC and input cDNA libraries, following manufacturer’s instructions (SuperScript III, ThermoFisher). We performed PCR on GC and input cDNA sample templates, amplifying Actb and Gfap for 32 cycles with primer pairs according to PrimerBank51. We imaged amplicon bands using standard agarose electrophoresis.
qPCR analysis
cDNA synthesis was performed on RNA purified from sorted GC and soma using SuperScript IV (ThermoFisher). qPCR analysis was performed on a BioRad CFX96 using the TaqMan Gene Expression Assay (Applied Biosystems, ThermoFisher) according to the manufacturer’s protocol. The following TaqMan probes were used: Rplp0 (#4453320; Mm00725448_s1), Rpl18a (#4448892; Mm04205642_gH), Rpsa (#4448892; Mm00726662_s1), Rps24 (#4448892; Mm01623058_s1), Rack1 (Gnb2l1) (#4448892; Mm01291968_g1), Eef1b2 (#4448892; Mm00516995_m1), Eef1g (#4448892; Mm02342826_g1), Cdkn1b (#4453320; Mm00438168_m1), Hspa5 (#4453320; Mm00517691_m1), Ppib (#4453320; Mm00478295_m1), Tubb2b (#4448892; Mm00849948_g1), Actb (#4453320; Mm02619580_g1), Gap43 (#4448892; Mm00500404_m1). Relative abundance in each sample was normalized by the mean expression of all transcripts tested, and enrichment is reported as the mean of GC/soma ratios across three biological replicates from independent litters. Sample order was randomized between replicate experiments. Agreement between RNA-seq and qPCR enrichment was assayed by calculating R2, the square of Pearson’s r, from log2-transformed normalized expression ratios.
Neuron culture
We cultured neurons from newborn mouse pup cortices that had been electroporated in utero at embryonic day 15 to label layer II/III inter-hemispheric projection neurons. We cultured neurons isolated from electroporated areas of cortex on poly-d-lysine-coated glass coverslips for 2–3 days, as previously described39.
Single molecule in situ hybridization
Single molecule in situ hybridization was performed using the RNAscope 2.5 HD RED kit according to the manufacturer’s instructions (Advanced Cell Diagnostics). In brief, primary cortical neurons cultured on glass coverslips for 2–3 days (described above) were fixed in 4% paraformaldehyde for 30 min at room temperature, followed by a series of ethanol dehydration and rehydration steps, pretreated first with hydrogen peroxide for 10 min, then with protease III (1:50) for 10 min. After the pretreatment steps, coverslips were incubated with individual probes for 2 h and the standard RNAscope protocol was followed. Incubation time of amplification step 5 and colour reaction were optimized for each probe per the recommendations of the manufacturer. After rinses, coverslips were immunostained for GFP using the standard procedure (described below). RNAscope probes targeting RPLP0 (315411, Entrez Gene: NM_007475.5), RACK1 (443621, Entrez Gene: NM_008143.3), PPIB (313911, Entrez Gene: NM_011149.2) and negative control DapB (#310043) were used. Images were acquired on a Zeiss LSM 880 confocal microscope using a 63× objective. Outlines were created using Trace Contour filter on Adobe Photoshop CC 2017.
Western blot
We performed western blots using standard Tris-glycine SDS–PAGE protocols. We determined total protein using the fluorometric Qubit protein assay (Thermo Fisher), and loaded equal amounts of total protein from input and GC fractions. We electroblotted resolved proteins onto PVDF membranes using semi-dry transfer. We followed standard western blot protocols, and incubated blots with primary antibodies diluted in 3% BSA in TBS with 0.2% Triton X-100, or in ‘Can Get Signal’ buffer (Toyobo). We used the following antibodies for immunoblotting: mouse-anti-β-actin, A5441, Sigma (1:2,000); mouse-anti-GAP43, MAB347, Chemicon (1:2,000); mouse-anti-GM130, 610823, BD Biosciences (1:3,000); mouse-anti-LAMP1, 1D4B, Developmental Studies Hybridoma Bank* (1:500); rabbit-anti-LARP1, #PA5-62398, ThermoFisher (1:1,000); mouse-anti-MAP2, #M1406, Sigma (1:1,000); rabbit-anti-mTOR, #2983, Cell Signaling Technology (1:1,000); rabbit-anti-mTOR, #A300-504A, Bethyl Labs (1:500); rabbit-anti-raptor, #42-4000, ThermoFisher (1:1,000); rabbit-anti-RICTOR, #2140, Cell Signaling Technology (1:1,000); rabbit-anti-TSC1, PA5-20131, ThermoFisher (1:1,000); mouse-anti-tubulin, MMS-435P, Covance (1:2,000).
Isotype-specific secondary antibodies used for ECL imaging were HRP-conjugated and cross-adsorbed (Life Technologies; Abcam). Immunoreactive bands were visualized through detection of chemiluminescence by SuperSignal West Pico PLUS (ThermoFisher) using a CCD camera imager (FluoroChemM, Protein Simple). To measure GC fraction enrichment, we quantified band intensities by densitometry, after applying despeckle filters using ImageJ software (NIH). Within each biological replicate, we normalized enrichment ratios (GC/input) for each protein of interest to the enrichment of GAP43 (the cardinal GC marker used to assess the degree of enrichment in each GC sample) and calculated mean and standard error using these normalized values. We used densitometry values from n ≥ 3 independent fractionation experiments from distinct litters (n = 7 for mTOR; n = 5 for LARP1, raptor and RICTOR; n = 3 for LAMP1 and TSC1; n = 8 for TUBB3; n = 6 for ACTB and GM130) to calculate means and standard deviations of GC/input ratios. We normalized expressed ratios to GAP43 ratios, and we calculated statistical significance using one-way ANOVA and post hoc analysis using Student’s t-tests.
Immunolabelling and imaging
For imaging, cultured neurons were fixed after 3 days in vitro in 4% paraformaldehyde. Brains were prepared by intracardial perfusion with 4% paraformaldehyde, and cut on a vibrating microtome (Leica) into 80 μm coronal sections. Immunolabelling of both coverslips and brain sections was performed in 3% bovine serum albumin and 0.2% Triton X-100. We used the following antibodies for immunolabelling: chicken-anti-GFP, A10262, Invitrogen (1:500); mouse-anti-LAMP1, 1D4B, Developmental Studies Hybridoma Bank (1:100); rabbit-anti-LARP1, PA5-62398, ThermoFisher (1:200); rabbit-anti-mTOR, 2983, Cell Signaling Technology (1:400); rabbit-anti-mTOR, A300-503A, Bethyl Labs (1:400); rabbit-anti-RAPTOR, 42-4000, ThermoFisher (1:200); mouse-anti-raptor, ab169506, Abcam (1:200); rabbit-anti-RFP, 600-401-379, Rockland (1:500); rabbit-anti-RICTOR, 2140, Cell Signaling Technology (1:200); rabbit-anti-TSC1, PA5-20131, ThermoFisher (1:500).
Isotype-specific secondary antibodies used for fluorescence imaging were Alexa Fluor-conjugated and cross-absorbed (Life Technologies). We acquired images on an epifluorescence microscope (Nikon 90i) with automated stage controller. Whole brain section images were generated using EDF z-stack projections and mosaic image stitching through the NIS Elements software (Nikon).
In vivo PI3K-DN and mTOR cKO analysis
To analyse electroporation position, migration and callosal axon extension in electroporated brains (Fig. 6 and Extended Data Fig. 10), four different brains from three independent litters were used for each experimental group. Animals for control and test electroporations were assigned at random, within the appropriate cohorts of experimental strains. Brain sections were immunostained against GFP, and imaged using an epifluorescence microscope (Nikon 90i). Images were used for quantifications as detailed below. Where counting was manual, obviously discernible phenotypes prevented effective blinding, thus a strict standardized process for measurement was used across samples. For quantification of migration, four different brains per condition were used, with minimum four sections per brain, with two non-overlapping rectangular areas (medial and lateral to midline, covering the entire cortical column) per brain section (average approximately 155 GFP+ cells per section, median 156 GFP+ cells per section). Each area was divided into 10 horizontal parallel bins, and the number of GFP+ cells was manually quantified. The percentage of GFP+ cells occupying the top three bins versus the percentage occupying the lower seven bins was plotted (Fig. 6a and Extended Data Fig. 10c). For quantification of alignment of electroporation areas and the extent of trans-hemispheric axon growth, four different brains per condition were used, with one section per brain (corresponding to sensorimotor area) used. Fiji was used for generation of bins in selected regions of interest and downstream automated intensity measurements52. For electroporation area measurements, a rectangular box covering the entire neocortical grey matter of the electroporated hemisphere (white matter cropped out before the measurement) was selected, binned into 200 parallel bins for intensity measurements, the background subtracted, and each bin normalized to the total intensity of the section. For callosal axon growth, rectangular boxes covering the entire callosum, centred to the midline, were selected, GFP+ cell bodies were manually removed, and the field divided into 400 bins for intensity measurements. After background intensity removal, the intensity of each bin was normalized to the average intensity of the first five bins of the same section.
Reporting summary
Further information on experimental design is available in the Nature Research Reporting Summary linked to this article.
Data availability
The datasets generated and analysed during the current study are available in the Harvard Dataverse repository https://doi.org/10.7910/DVN/ISOEB6.
References
Lowery, L. A. & Van Vactor, D. The trip of the tip: understanding the growth cone machinery. Nat. Rev. Mol. Cell Biol. 10, 332–343 (2009).
Saxton, R. A. & Sabatini, D. M. mTOR signaling in growth, metabolism, and disease. Cell 168, 960–976 (2017).
Meyuhas, O., Avni, D. & Shama, S. Translational control of ribosomal protein mRNAs in eukaryotes. 30, 363–388 (1996).
Fonseca, B. D. et al. La-related protein 1 (LARP1) represses terminal oligopyrimidine (TOP) mRNA translation downstream of mTOR complex 1 (mTORC1). J. Biol. Chem. 290, 15996–16020 (2015).
Greig, L. C., Woodworth, M. B., Galazo, M. J., Padmanabhan, H. & Macklis, J. D. Molecular logic of neocortical projection neuron specification, development and diversity. Nat. Rev. Neurosci. 14, 755–769 (2013).
Pfenninger, K. H., Ellis, L., Johnson, M. P., Friedman, L. B. & Somlo, S. Nerve growth cones isolated from fetal rat brain: subcellular fractionation and characterization. Cell 35, 573–584 (1983).
Lohse, K. et al. Axonal origin and purity of growth cones isolated from fetal rat brain. Brain Res. Dev. Brain Res. 96, 83–96 (1996).
Fame, R. M., MacDonald, J. L. & Macklis, J. D. Development, specification, and diversity of callosal projection neurons. Trends Neurosci. 34, 41–50 (2011).
Greig, L. C., Woodworth, M. B., Greppi, C. & Macklis, J. D. Ctip1 controls acquisition of sensory area identity and establishment of sensory input fields in the developing neocortex. Neuron 90, 261–277 (2016).
Llorca, O. et al. Eukaryotic type II chaperonin CCT interacts with actin through specific subunits. Nature 402, 693–696 (1999).
Moccia, R. et al. An unbiased cDNA library prepared from isolated Aplysia sensory neuron processes is enriched for cytoskeletal and translational mRNAs. J. Neurosci. 23, 9409–9417 (2003).
Leung, K.-M. et al. Asymmetrical beta-actin mRNA translation in growth cones mediates attractive turning to netrin-1. Nat. Neurosci. 9, 1247–1256 (2006).
Crino, P. B. & Eberwine, J. Molecular characterization of the dendritic growth cone: regulated mRNA transport and local protein synthesis. Neuron 17, 1173–1187 (1996).
Taylor, A. M. et al. Axonal mRNA in uninjured and regenerating cortical mammalian axons. J. Neurosci. 29, 4697–4707 (2009).
Zivraj, K. H. et al. Subcellular profiling reveals distinct and developmentally regulated repertoire of growth cone mRNAs. J. Neurosci. 30, 15464–15478 (2010).
Jung, H., Yoon, B. C. & Holt, C. E. Axonal mRNA localization and local protein synthesis in nervous system assembly, maintenance and repair. Nat. Rev. Neurosci. 13, 308–324 (2012).
Catapano, L. A., Arnold, M. W., Perez, F. A. & Macklis, J. D. Specific neurotrophic factors support the survival of cortical projection neurons at distinct stages of development. J. Neurosci. 21, 8863–8872 (2001).
Arlotta, P. et al. Neuronal subtype-specific genes that control corticospinal motor neuron development in vivo. Neuron 45, 207–221 (2005).
Özdinler, P. H. & Macklis, J. D. IGF-I specifically enhances axon outgrowth of corticospinal motor neurons. Nat. Neurosci. 9, 1371–1381 (2006).
Meyuhas, O. & Kahan, T. The race to decipher the top secrets of TOP mRNAs. Biochim. Biophys. Acta 1849, 801–811 (2015).
Geiger, T., Wehner, A., Schaab, C., Cox, J. & Mann, M. Comparative proteomic analysis of eleven common cell lines reveals ubiquitous but varying expression of most proteins. Mol. Cell. Proteomics 11, M111.014050 (2012).
Liebermeister, W. et al. Visual account of protein investment in cellular functions. Proc. Natl Acad. Sci. USA 111, 8488–8493 (2014).
Tang, H. et al. Amino acid-induced translation of TOP mRNAs is fully dependent on phosphatidylinositol 3-kinase-mediated signaling, is partially inhibited by rapamycin, and is independent of S6K1 and rpS6 phosphorylation. Mol. Cell. Biol. 21, 8671–8683 (2001).
Thoreen, C. C. et al. A unifying model for mTORC1-mediated regulation of mRNA translation. Nature 485, 109–113 (2012).
Laplante, M. & Sabatini, D. M. mTOR signaling in growth control and disease. Cell 149, 274–293 (2012).
Kye, M. J. et al. SMN regulates axonal local translation via miR-183/mTOR pathway. Hum. Mol. Genet. 23, 6318–6331 (2014).
Tcherkezian, J. et al. Proteomic analysis of cap-dependent translation identifies LARP1 as a key regulator of 5'TOP mRNA translation. Genes Dev. 28, 357–371 (2014).
Dhand, R. et al. PI 3-kinase: structural and functional analysis of intersubunit interactions. EMBO J. 13, 511–521 (1994).
Itoh, Y. et al. PDK1-Akt pathway regulates radial neuronal migration and microtubules in the developing mouse neocortex. Proc. Natl Acad. Sci. USA 113, E2955–E2964 (2016).
Lu, Y., Belin, S. & He, Z. Signaling regulations of neuronal regenerative ability. Curr. Opin. Neurobiol. 27, 135–142 (2014).
Okabe, M., Ikawa, M., Kominami, K., Nakanishi, T. & Nishimune, Y. ‘Green mice’ as a source of ubiquitous green cells. FEBS Lett. 407, 313–319 (1997).
Madisen, L. et al. A robust and high-throughput Cre reporting and characterization system for the whole mouse brain. Nat. Neurosci. 13, 133–140 (2010).
Gallardo, T., Shirley, L., John, G. B. & Castrillon, D. H. Generation of a germ cell-specific mouse transgenic Cre line, Vasa-Cre. Genesis 45, 413–417 (2007).
Risson, V. et al. Muscle inactivation of mTOR causes metabolic and dystrophin defects leading to severe myopathy. J. Cell Biol. 187, 859–874 (2009).
Rhee, J. M. et al. In vivo imaging and differential localization of lipid-modified GFP-variant fusions in embryonic stem cells and mice. Genesis 44, 202–218 (2006).
Matsuda, T. & Cepko, C. L. Electroporation and RNA interference in the rodent retina in vivo and in vitro. Proc. Natl Acad. Sci. USA 101, 16–22 (2004).
Saito, T. & Nakatsuji, N. Efficient gene transfer into the embryonic mouse brain using in vivo electroporation. Dev. Biol. 240, 237–246 (2001).
Biesemann, C. et al. Proteomic screening of glutamatergic mouse brain synaptosomes isolated by fluorescence activated sorting. EMBO J. 33, 157–170 (2014).
Catapano, L. A., Arlotta, P., Cage, T. A. & Macklis, J. D. Stage-specific and opposing roles of BDNF, NT-3 and bFGF in differentiation of purified callosal projection neurons toward cellular repair of complex circuitry. Eur. J. Neurosci. 19, 2421–2434 (2004).
Molyneaux, B. J. et al. Novel subtype-specific genes identify distinct subpopulations of callosal projection neurons. J. Neurosci. 29, 12343–12354 (2009).
Galazo, M. J., Emsley, J. G. & Macklis, J. D. Corticothalamic projection neuron development beyond subtype specification: Fog2 and intersectional controls regulate intraclass neuronal diversity. Neuron 91, 90–106 (2016).
Dobin, A. et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics 29, 15–21 (2013).
Patro, R., Mount, S. M. & Kingsford, C. Sailfish enables alignment-free isoform quantification from RNA-seq reads using lightweight algorithms. Nat. Biotechnol. 32, 462–464 (2014).
Okonechnikov, K., Conesa, A. & García-Alcalde, F. Qualimap 2: advanced multi-sample quality control for high-throughput sequencing data. Bioinformatics 32, 292–294 (2016).
Cox, J. et al. Accurate proteome-wide label-free quantification by delayed normalization and maximal peptide ratio extraction, termed MaxLFQ. Mol. Cell. Proteomics 13, 2513–2526 (2014).
Tyanova, S. et al. The Perseus computational platform for comprehensive analysis of (prote)omics data. Nat. Methods 13, 731–740 (2016).
Berriz, G. F. & Roth, F. P. The Synergizer service for translating gene, protein and other biological identifiers. Bioinformatics 24, 2272–2273 (2008).
Binder, J. X. et al. COMPARTMENTS: unification and visualization of protein subcellular localization evidence. Database (Oxford) 2014, bau012 (2014).
Szklarczyk, D. et al. STRING v10: protein–protein interaction networks, integrated over the tree of life. Nucleic Acids Res. 43, D447–D452 (2015).
Cox, J. & Mann, M. 1D and 2D annotation enrichment: a statistical method integrating quantitative proteomics with complementary high-throughput data. BMC Bioinformatics 13 (Suppl. 16), S12 (2012).
Spandidos, A., Wang, X., Wang, H. & Seed, B. PrimerBank: a resource of human and mouse PCR primer pairs for gene expression detection and quantification. Nucleic Acids Res. 38, D792–D799 (2010).
Schindelin, J. et al. Fiji: an open-source platform for biological-image analysis. Nat. Methods 9, 676–682 (2012).
Acknowledgements
We thank A.-K. Hadjantonakis and S. Srinivas for reagents; B. Noble and I. Florea for technical support; J. LaVecchio, G. Buruzula and S. Ionescu of HSCRB-HSCI Flow Cytometry Core; W. Lane, J. Neveu and B. Budnik of the Harvard FAS CSB Mass Spectrometry and Proteomics Resource Laboratory; the Harvard Center for Biological Imaging for infrastructure and support; L. Liang for statistical advice; N. Brose, C. Biesemann, E. Herzog, L. Reijmers and J. Rinn for discussions. This work was supported by grants to J.D.M. from the Paul G. Allen Frontiers Group, Brain Research Foundation Scientific Innovations Award program; NIH Pioneer Award DP1 NS106665, Emily and Robert Pearlstein Fund, and the Max and Anne Wien Professorship; with additional infrastructure support from NIH grants NS045523, NS075672, NS049553, and NS041590. A.P. was partially supported by a European Molecular Biology Organization Long Term Fellowship and a Human Frontier Science Program Long Term Fellowship. A.J.M. was partially supported by NIH Training Grant T32 AG000222. Work by R.K. and the Harvard Chan Bioinformatics Core was supported by funds from the Harvard NeuroDiscovery Center and Harvard Stem Cell Institute. J.D.M. is an Allen Distinguished Investigator of the Paul G. Allen Frontiers Group.
Reviewer information
Nature thanks D. Sabatini, J. Henley and the anonymous reviewer(s) for their contribution to the peer review of this work.
Author information
Authors and Affiliations
Contributions
A.P. and J.D.M. conceived of the project. A.P., A.J.M. and J.D.M. designed experiments. A.P., A.J.M., A.O., P.D. and J.H. performed experiments. A.P., A.J.M., A.O., J.H., R.K. and J.D.M. analysed data. A.P., A.J.M., P.D. and J.D.M. interpreted results. A.P. and J.D.M. wrote the manuscript. All authors contributed to discussions and manuscript editing.
Corresponding authors
Ethics declarations
Competing interests
A US patent application (15/144,660) is pending on growth cone sorting technology and implications. The application lists J.D.M., A.P. and A.J.M. as inventors. All other authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Isolated GC verification.
a, Enrichment analysis of GC fraction versus starting homogenate (input). Left, western blot detects protein enrichment of GAP43 (GC marker) and depletion of MAP2 (somato-dendritic protein marker) in GC fraction. Middle, native gel electrophoresis shows de-enriched presence of large (28S) and small (18S) ribosomal subunit rRNA in GC fraction. Right, RT–PCR detects mRNA for Actb (ubiquitous) but not Gfap (progenitor and glial marker) from the GC fraction. n = 6 biological replicates for protein and rRNA; n = 3 for mRNA. b, GC protection assay schema: bulk GC fraction isolated after subcellular fractionation is a suspension of GC particles enclosing GC-specific molecules (blue) within a medium that contains dilute soluble cytosolic molecules (red). Treatment with RNase or protease leads to hydrolysis of RNA and protein in the suspension medium not protected within GC particles, leaving only the GC-protected molecules (blue) after treatment. Addition of detergent before treatment results in hydrolysis of both cytosolic as well as GC-encapsulated molecules due to ruptures in the encapsulating GC plasma membrane, providing a control for the efficiency of enzymes. The difference in RNA or protein signal between hydrolysis control and GC-protected samples corresponds to the GC-encapsulated signal. c, d, GC protection assays with non-membrane-permeable degrading enzymes (protease in c and RNase in d) to test GC integrity and GC-specific membrane encapsulation of RNA and proteins in isolated GCs. Treatment with enzyme plus detergent, but neither alone, completely abolishes RNA and protein signal from GC fractions. Signals persisting in treatments with enzyme alone (lanes 3) correspond to RNA and protein encapsulated (protected) by the GC membrane, and correspond to the specific molecular content of isolated GCs. Treatment with protease alone has no effect on the signal from GAP43, confirming GC-specificity. Conversely, the signal from GM130, a Golgi matrix protein known to be excluded from GCs, is abolished with protease treatment alone, indicating there is no non-specific encapsulation in GCs. Reduced presence of both Actb and rRNA in samples treated with enzyme alone is consistent with their ubiquitous presence in both GCs and elsewhere in the homogenate. n = 5 independent biological replicates. e, Bioanalyzer profiles show GC-protected RNA compared to detergent-treated control, with characteristic peaks corresponding to 28S and 18S rRNA, and a spectrum of low intensity signal characteristic of mRNA. Experiments performed in n = 5 biological replicates with consistent results.
Extended Data Fig. 2 Quality control filtering of mass spectrometry measurements from sorted somata and GCs.
a, b, Multi-scatter plot of mass spectrometry signal intensity (log2-transformed label-free quantification (LFQ) of proteins) for each detected soma (a) and GC (b) protein in pairwise comparisons across six biological replicates. Quality control minimum stringency criteria were set based on average Pierson’s correlation coefficients across biological replicates. Soma samples displayed higher complexity than GC samples, which was reflected in the minimum acceptable correlation coefficients of 0.5 for somata (a) and 0.8 for GCs (b). All six soma replicates, and four out of six GC replicates met quality control criteria. Outlier GC samples (GC 3 and GC 6) are shaded grey (b).
Extended Data Fig. 3 Sorted GC–soma protein mass spectrometry intensities.
Scatter plot of paired protein intensities from trans-hemispheric sorted GC and sorted parent somata. Units represent log2-transformed peak-normalized intensities as measured by MaxLFQ45. Gene groups are coloured as indicated in the key. The GC marker GAP43 is indicated by a circled asterisk.
Extended Data Fig. 4 Sorted GC–soma proteome mapping.
Volcano plot of GC–soma proteome mapping, with log2-transformed λP values for each gene product plotted against significance (−log-transformed P values determined by two-tailed t-test). Significance thresholds were set to a 0.05 permutation-based FDR to indicate soma- and GC-specific mapping. n = 6 biological replicates, 3–6 litters each. Coloured gene groups are indicated. The proteome of sensorimotor cortex inter-hemispheric projection neurons distributes between cellular compartments with varying polarization that clusters with gene group, including GC-rich clusters (for example, proteasome), soma-rich clusters (for example, histones), and groups with moderate levels present in both GCs and somata, with moderate enrichments for one or the other compartment (for example, ribosomes and actin/tubulin).
Extended Data Fig. 5 Sorted GC mRNA-to-protein distribution.
Scatter plot pairing mRNA and corresponding protein relative abundance in trans-hemispheric-sorted GCs. Units represent log2-transformed peak-normalized mean intensities as measured by Sailfish TPM (mRNA) and MaxLFQ (protein) from 6 biological replicates of 3–6 litters each. Gene groups are indicated. The GC marker GAP43 is indicated by an asterisk. Trans-hemispheric GCs contain mRNA of select high-abundance GC proteins, as well as of most ribosomal protein mRNAs, whereas ribosomal proteins remain at GCs at moderate levels.
Extended Data Fig. 6 Subcellular RNA–proteome mapping.
2D RNA–proteome mapping of statistically significant enrichments (two-tailed t-test P value, significance threshold of 0.02 Benjamini–Hochberg corrected FDR, from 6 biological replicates of 3–6 litters each) in Gene Ontology (GO) terms and gene groups defined in Supplementary Table 7. Labels are displayed for non-redundant GO cell compartment terms and gene groups. Histograms along the x axis show RNA distributions of log2-transformed mean λR values. Histograms along the y axis on the right side show protein distributions of log2-transformed mean λP values of genes within each group. Four clusters emerge: the soma cluster (blue) contains groups of genes with both mRNA and protein enriched in the soma; the anterograde cluster (red) contains groups of genes with mRNA mapping to soma and protein to GC; the mito cluster (grey) contains mitochondrial genes with intermediate distributions; and the TOP cluster (green) exclusively contains TOP transcripts (including ribosomal protein genes), with mRNAs mapping to the GC and proteins to both GC and soma.
Extended Data Fig. 7 Subcellular transcriptome distribution follows mTOR dependence.
a, Sample transcripts from each class represented in Fig. 4 were verified by qPCR (y axis), and correlated with RNA-seq mapping values (x axis). Measurements from the two approaches displayed strong concurrence, with correlation coefficient R2 = 0.736. Centre points denote the mean, and error bars denote s.e.m. of three biological replicates from independent litters. Colour legend and schematics of mRNA classes based on known mTOR dependence: mRNAs containing a TOP motif are mTOR-dependent (green); schema indicates direct binding of the TOP motif by LARP1, which interacts with mTOR. mRNAs containing internal ribosome entry sites or lacking poly(A) tails are mTOR-independent (blue). Canonical mRNAs that undergo cap-dependent translation (grey) display moderate responses to mTOR. c, Expanded dataset presented in Fig. 4c. Single-molecule RNA chromogenic in situ hybridization (ISH, red) of two TOP transcripts: Rack1 (non-ribosomal TOP) and Rplp0 (ribosomal TOP), compared to a control transcript Ppib (soma-mapped canonical) in callosal projection neurons. Neurons were labelled with mem-GFP (green) via in utero electroporation at embryonic day (E) 15, cultured at P0, fixed and hybridized at in vitro day (DIV) 2–3. Nuclei were stained with DAPI (blue). Soma and GC close-ups shown in insets as overlays of transcript (red) with mem-GFP (green) in top rows, or with traced GC outlines in bottom rows. 92, 103 and 84 GCs imaged for Rack1, Rplp0 and Ppib probes, respectively, from n = 4 biological replicates from independent in utero electroporations. Five example GCs are shown per sample to capture the representative range. Scale bars, 10 μm.
Extended Data Fig. 8 Dense foci of mTOR in GCs.
a, Data relate to Fig. 5. Biochemical analysis of GC enrichment of mTOR pathway proteins and controls, shown in triplicate western blots of homogenate (input) and GC fraction pairs, derived from six independent preps. GAP43 is positive control for enrichment; GM130 is a negative control. Quantification as in Fig. 5. b, Close-ups of GCs from callosal projection neurons immunelabelled for endogenous mTOR pathway proteins (red in overlays, heat-mapped in underlying panels). Five example GCs are shown per sample to capture the representative range. Neurons were labelled via in utero electroporation at E15 with membrane-GFP (green in overlays, outlined in underlying panels), cultured at P0, fixed and stained at DIV2–DIV3. mTOR, LARP1, TSC1 and raptor (mTORC1 marker) appear in dense local foci within GCs. RICTOR (mTORC2 marker) and LAMP1 (lysosome marker) appear in fine granules distinct from GC foci. Bar (bottom right) indicates heat-map colour range, as well as 10-μm scale. 83 GCs imaged for mTOR, 47 for LARP1, 42 for LAMP1, 49 for TSC1, 26 for raptor, and 30 for RICTOR, from a minimum of n = 3 biological replicates from independent in utero electroporations.
Extended Data Fig. 9 GC-specific mTOR localization.
a, High-magnification views of representative GCs from callosal projection neurons immunolabelled for mTOR, equivalent to Fig. 5c and Extended Data Fig. 8b mTOR panels, but with a distinct mTOR antibody to independently confirm dense focal mTOR in GCs. Top, overlay images; bottom, heat maps of the same GCs. b, Example of mTOR labelling (red in two left panels; heat map in right panel) in 3-day-cultured neurons. A GFP-labelled axon from an electroporated inter-hemispheric neuron displays dense focal mTOR in its GC (arrow) compared to adjacent cell bodies (asterisks; DNA in blue indicates nuclei). Two other unlabelled GCs in the field can be recognized by virtue of their dense focal mTOR labelling alone (arrowheads). c, Example of dendritic GCs (arrows) lacking mTOR, juxtaposed to an unlabelled GC (arrowhead) with prominent mTOR focus. Bars in a–c (bottom right of each) indicate mTOR intensity heat-map colour range, as well as 10-μm scale. GCs imaged from n = 2 biological replicates from independent in utero electroporations.
Extended Data Fig. 10 mTOR signalling is required for the extension of trans-hemispheric axons.
a, Data relate to Fig. 6. Electroporation of callosal projection neurons with GFP and genetic payloads at E15, fixation and analysis at P3. Control electroporations (left column, grey in quantifications) show soma migration into upper layers (middle row insets, examples from four brains), and callosal projections well into the contralateral cortex (bottom insets, examples from four brains). Electroporation with PI3K-DN (middle column, green in quantifications) hinders migration of somata, with failure of callosal axon growth. Electroporation of Cre in mice with homozygous floxed-mTOR alleles for conditional mTOR gene knockout (mTOR-KO, right column, blue in quantifications) results specifically in failure of callosal axon growth. Scale bars, 100 μm. b, Quantification of the location of the electroporation field shows comparable mediolateral electroporation positions across all samples. Plotted are histograms of binned GFP intensities along the tangential axis of the ipsilateral cortex ending at the midline, as schematized above. c, Quantification of extent of migration, with percentage of somata in layers II/III (dark colours in graph) versus somata still en route in deeper layers (light colours in graph), as schematized above. Inhibition of PI3K signalling (green) interferes with migration, whereas acute mTOR deletion (blue) does not significantly affect migration. d, Quantification of callosal axon extension, showing that PI3K inhibition (green) as well as knockout of mTOR (blue) disrupt the formation of axon projection across the corpus callosum. Plotted are binned GFP intensity histograms within the corpus callosum from ipsilateral, through the midline (indicated by dotted line), to the contralateral side, as schematized above. Data are mean ± s.e.m. from n = 4 mice from different litters for each condition.
Supplementary information
Supplementary Table 1: Trans-hemispheric GC sub-proteome, {P}GC
List of proteins that are detected by mass-spec from sorted trans-hemispheric GCs. Columns correspond to gene names, protein names, theoretical protein molecular weight, and ENSEMBL gene ID.
Supplementary Table 2: GC-soma proteome mapping
List of protein GC-soma ratios λP. Columns correspond to gene names, λP values (log2), specific enrichment based on significance threshold set to 0.05 permutation-based false discovery rate (FDR), two-tailed t-test P values (-log), protein names, and gene group. n=6 biological replicates from 3-6 litters each.
Supplementary Table 3: Protein GC-soma annotation enrichment
List of significantly enriched Gene Ontology (GO) terms based on λP values. Columns correspond to term type (GO-Biological Function, GO-Cellular Component, and GO-Molecular Function), term name, number of genes, rank distribution score51, two-tailed t-test P value, significance based on Benjamini-Hochberg corrected FDR, and mean λP value.
Supplementary Table 4: GC-soma RNA mapping
List of mRNA GC-soma ratios λR. Columns correspond to gene names, λR values (log2), specific enrichment based on significance threshold set to 0.05 permutation-based FDR, two-tailed t-test P values (-log), gene names, and gene group. n=6 biological replicates from 3-6 litters each.
Supplementary Table 5: mRNA GC-soma annotation enrichment
List of significantly enriched Gene Ontology (GO) terms based on λR values. Columns correspond to term type (GO-Biological Function, GO-Cellular Component, and GO-Molecular Function), term name, number of genes, rank distribution score51, two-tailed t-test P value, significance threshold of 0.02 Benjamini-Hochberg corrected FDR, and mean λR value.
Supplementary Table 6: 2D RNA-proteome GC-soma annotation enrichment
List of significantly enriched Gene Ontology (GO) and gene group terms based on paired λR and λP values. Columns correspond to term type (GO-Biological Function, GO-Cellular Component, GO-Molecular Function, and gene group), term name, number of genes, two-tailed t-test P value, significance threshold of 0.02 Benjamini-Hochberg corrected FDR, mean λR, and mean λP values.
Supplementary Table 7: Gene groups
List of gene names and ENSEMBL gene IDs that constitute the annotated gene groups in Fig. 2, Fig. 4, Extended Data Fig.3-5, and Supplementary Table 6.
Rights and permissions
About this article
Cite this article
Poulopoulos, A., Murphy, A.J., Ozkan, A. et al. Subcellular transcriptomes and proteomes of developing axon projections in the cerebral cortex. Nature 565, 356–360 (2019). https://doi.org/10.1038/s41586-018-0847-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-018-0847-y
- Springer Nature Limited
This article is cited by
-
CPEB2-activated axonal translation of VGLUT2 mRNA promotes glutamatergic transmission and presynaptic plasticity
Journal of Biomedical Science (2024)
-
The role of TSC1 and TSC2 proteins in neuronal axons
Molecular Psychiatry (2024)
-
Mechanistic insights into the basis of widespread RNA localization
Nature Cell Biology (2024)
-
Human neuronal maturation comes of age: cellular mechanisms and species differences
Nature Reviews Neuroscience (2024)
-
Molecular programs guiding arealization of descending cortical pathways
Nature (2024)