Abstract
Campylobacter jejuni is the leading cause of bacterial diarrhoeal disease in many areas of the world. The high incidence of sporadic cases of disease in humans is largely due to its prevalence as a zoonotic agent in animals, both in agriculture and in the wild. Compared with many other enteric bacterial pathogens, C. jejuni has strict growth and nutritional requirements and lacks many virulence and colonization determinants that are typically used by bacterial pathogens to infect hosts. Instead, C. jejuni has a different collection of factors and pathways not typically associated together in enteric pathogens to establish commensalism in many animal hosts and to promote diarrhoeal disease in the human population. In this Review, we discuss the cellular architecture and structure of C. jejuni, intraspecies genotypic variation, the multiple roles of the flagellum, specific nutritional and environmental growth requirements and how these factors contribute to in vivo growth in human and avian hosts, persistent colonization and pathogenesis of diarrhoeal disease.
Similar content being viewed by others
Introduction
Campylobacter jejuni is a commensal bacterium that resides in the intestinal tracts of many wild and agriculture-associated animals. Poultry flocks, especially chickens, are commonly colonized with C. jejuni. As such, handling and consumption of poultry meats contaminated with the bacterium are leading risk factors for sporadic cases of diarrhoeal disease caused by C. jejuni in humans1,2. Outbreak cases of disease are often attributed to consumption of contaminated water or raw milk3,4. The rates of diarrhoeal disease caused by C. jejuni are high in both developed and developing countries5. Although C. jejuni is already the leading cause of bacterial diarrhoeal disease in many European countries, the Centers for Disease Control and Prevention reported that C. jejuni surpassed Salmonella species as the leading cause of food-borne bacterial diarrhoeal disease in the United States in 2017 (ref.6). The increased antibiotic resistance of C. jejuni and the difficulty in reducing rates of diarrhoeal disease in recent years have augmented the emphasis on identifying new targets for antimicrobials and vaccine development and on developing innovative strategies to reduce the presence of C. jejuni in agriculture, in meats that are consumed by humans and in environmental reservoirs to lower the risk of exposure to C. jejuni.
C. jejuni predominantly colonizes the mucous layers on the epithelium in the lower intestinal tract and associated crypts of hosts. These lower intestinal tract niches are ideal for colonization as they naturally harbour a plentiful supply of nutrients and carbon sources that support C. jejuni metabolism and robust growth. Furthermore, the intestinal microbiota in these locations has been proposed to contribute metabolites that influence the expression of C. jejuni colonization factors and promote growth7 (and reviewed in ref.8). However, subsequent steps in host interactions are likely to drive different outcomes of infection, with commensalism developing in many avian species and animals and diarrhoeal disease occurring in susceptible humans (Box 1). C. jejuni adherence to and invasion of the avian intestinal epithelium, as analysed in chickens, seem to be infrequent events that do not produce robust inflammation or pathological signs of disease except in some chicken breeds that seem to have defects in proper immune or inflammatory responses9,10. However, infections that progress to diarrhoeal disease in humans are characterized by invasion of the colonic epithelium and inflammation, which is likely to contribute to the pathology and symptoms of diarrhoeal disease11 (Box 2).
Although most associations of C. jejuni with animal hosts in nature result in commensalism, a more obvious pathogenic outcome due to C. jejuni infection occurs in ovine species. Whereas Campylobacter fetus is the predominant cause of ovine abortions in many parts of the world, a C. jejuni clone (clone SA) is the leading cause of ovine abortions in the United States12. Surprisingly, there are few differences in genomic content between C. jejuni isolates that promote commensalism in poultry and diarrhoeal disease in humans and the C. jejuni clone SA. A recent study has uncovered that amino acid changes in a specific region of the major outer membrane protein of C. jejuni clone SA are necessary and sufficient to transform an avian commensal C. jejuni strain into a virulent abortifacient strain in a guinea pig infection model13.
Overall, much knowledge regarding essential determinants and molecular mechanisms to explain how C. jejuni infects avian and human hosts to promote commensalism or diarrhoeal disease, respectively, is to still be discovered. The biology of C. jejuni is likely malleable and complex to enable persistence in nature as a member of the commensal flora of many animals, yet it promotes infection in humans to cause diarrhoeal disease. For instance, the avian and human host generally differ not only in diets that ultimately provide nutrition for C. jejuni but also in innate factors, such as body temperature (42°C in chickens versus 37°C in humans), that require C. jejuni to quickly adapt to changing conditions upon transmission from one host to another. Moreover, whereas the development of in vivo model systems to replicate diarrhoeal disease following infection with C. jejuni is improving, current models that involve the manipulation of the microbiota or the use of immune-deficient animals suffer owing to difficulty in animal rearing for infection, the general lack in replicating pathological features of diarrhoeal disease observed in humans and requiring large inocula for infection relative to the low inocula required for infection of humans. Furthermore, C. jejuni lacks or does not extensively rely on many virulence factors typically used by or found in other enteric bacterial pathogens (Box 3). For example, C. jejuni lacks a dedicated injectisome type III secretion system (T3SS) to transport effector proteins into eukaryotic cells to manipulate host cell biology. Instead, the bacterium has expanded the use of the flagellar T3SS to secrete proteins that modulate host interactions for colonization and infection. Additionally, C. jejuni produces lipooligosaccharide (LOS), a truncated form of lipopolysaccharide (LPS) that is produced by many enteric bacterial pathogens.
Recent technological advances, including metabolite profiling, second-generation sequencing upon in vitro and in vivo growth and genomic content comparisons between diverse isolates, have identified new and essential metabolic pathways for growth, mechanisms of generating genotypic and phenotypic diversity and antibiotic resistance and essential genes for infection. Additionally, many exciting insights into cellular architecture and surface structures have uncovered possible roles that these factors have in immune evasion, phage infection, motility, signal transduction and division.
In this Review, we explore the collective aspects of C. jejuni biology that contribute to its divergence from many enteric bacteria and how it efficiently creates niches to coexist as a commensal in many animal hosts while also being a prominent agent of bacterial diarrhoeal disease in humans. We explore how cell shape and surface structures contribute to cellular architecture and composition, mechanisms to generate genotypic and phenotypic diversity, how the flagellum affects motility and C. jejuni cellular biology and insights into metabolic requirements for growth. Although these collective determinants and activities of C. jejuni differentiate the bacterium from other enteric pathogens, the exact requirements for many of these factors for C. jejuni in host interactions remain elusive.
Cellular architecture
Cell shape
The helical shape of the C. jejuni cell body and its amphitrichous flagella are postulated to contribute to the natural high velocity of motility observed in viscous environments14,15,16,17. This motility facilitates colonization, spread and retention within mucous layers on intestinal epithelia for commensalism in animals and pathogenesis of diarrhoeal disease in humans. Correct processing of peptidoglycan subunits contributes to the helical cell shape, as mutations in peptidoglycan peptidases that cleave different sites along the peptide chains that crosslink peptidoglycan subunits result in straight cell bodies18,19,20 (Fig. 1). These peptidoglycan-processing mutants exhibit altered inflammatory responses and a decrease in pathology in mouse models of diarrhoeal disease, probably owing to reduced migration to the mucous layer of caecal crypts18,19,20. The straight body phenotype also reduces commensal colonization capacity in chicks18,19. Currently, it is unclear what other peptidoglycan processing mechanisms and modifications are necessary for the helical shape of C. jejuni.
The Campylobacter jejuni cell body with the characteristic helical shape and amphitrichous flagella is shown in the centre. Boxes outline areas enlarged to depict molecular substructures. Processes involved in generating cellular architecture and substructures include peptidoglycan processing, lipooligosaccharide (LOS) sialylation, general N-linked protein glycosylation and free oligosaccharide production, capsular polysaccharide (CPS) biogenesis and O-linked protein glycosylation of flagellin. Pgp1 and Pgp2 peptidoglycan peptidases cleave at different sites in the peptide chains (indicated by dashed arrows) that link peptidoglycan subunits. Peptidoglycan processing contributes to the generation of a proper peptidoglycan network that produces the helical cell shape of C. jejuni. Δpgp1 and Δpgp2 mutants exhibit altered ratios of different peptidoglycan subunits (the increase or decrease of different types of peptidoglycan subunits is indicated by arrows), which results in a straight cell body shape. The modifications of LOS outer core structures on the surface of C. jejuni mimic human ganglioside structures. These different ganglioside structures are composed of different linkages of N-acetylgalactosamine (GalNAc), N-acetylneuraminic acid (Neu5Ac), glucose (Glu) and galactose (Gal) and mimic a range of gangliosides (GM1, GM2, GM3, GD3, GT1a, GDa1 and GD1c). The general N-linked protein glycosylation system produces both free oligosaccharides in the periplasm and an identical oligosaccharide linked to asparagine residues on many inner membrane, periplasmic and outer membrane proteins. This oligosaccharide is composed of a chain of GalNAc residues with bacillosamine (Bac) and a phosphoethanolamine group. The extracellular CPS of many C. jejuni strains is attached to the outer membrane by a lipid anchor and is composed of linked subunits that consist of a backbone of two to three saccharides with different modifications. The most commonly recognized modifications are O-methyl phosphoramidate (MeOPN) and O-methyl (O-Me) groups, which can vary in amount and positioning on a CPS subunit. O-linked protein glycosylation of a flagellin subunit usually results in the modification of ~19 specific serine and threonine residues with pseudaminic acid (Pse) and related derivatives. Some strains also produce legionaminic acid (Leg) and related derivatives that are added to the flagellin subunit. d-iGlu, d-isomer of glucose; Kdo, ketodeoxyoctonic acid; meso-DAP, meso-diaminopimelic acid; NAG, N-acetylglucosamine; NAM, N-acetylmuramic acid.
Capsular polysaccharide
C. jejuni strains produce an extracellular capsular polysaccharide (CPS) that defines around 47 different serotypes21. CPS, which many other enteric bacteria also produce, is required by C. jejuni for wild-type levels of commensal colonization of chicks and affects many aspects of pathogenesis of diarrhoeal disease, including adherence to and invasion of intestinal epithelial cells, serum resistance, activation of Toll-like receptor (TLR) and cytokine stimulation (reviewed in ref.22). Generally, C. jejuni CPS is composed of a repeating backbone of two or more sugars that differ among serotypes, but the backbone does not seem to vary structurally within a strain (Fig. 1). Instead, intrastrain variation within a C. jejuni population occurs via modifications of the backbone with O-methyl phosphoramidate (MeOPN) and O-methyl (O-Me) groups23,24,25,26. More than 70% of C. jejuni strains produce MeOPN-modified CPS23,26,27,28,29. Variation in MeOPN and O-Me levels and placement is due to reversible phase variation of the respective transferases that modify specific sites on saccharides of the CPS backbone26,27,28,29,30.
Recent analysis of the C. jejuni CPS structure has revealed intriguing findings for how the CPS may vary to alter interactions that affect bacteriophage infectivity, chick colonization, invasion capacity for human epithelial cells and serum resistance. The CPS backbone, but not MeOPN modification, is required for wild-type levels of commensal colonization of chicks26,31. However, certain MeOPN arrangements on CPS form a receptor for bacteriophages, whereas O-Me at specific sites inhibits bacteriophage infection27,28,30. Thus, in vivo phase variation of MeOPN transferases to eliminate MeOPN modification enables C. jejuni to survive phage infection in chickens without altering commensal colonization. MeOPN modifications also influence virulence, as loss of MeOPN on CPS increases invasion potential but reduces serum resistance29,31,32. Thus, phase variation of MeOPN transferases may occur during human infection to switch from MeOPN-containing CPS to resist humoral immunity to MeOPN-deficient CPS at later times during infection to promote intestinal epithelial invasion.
Lipooligosaccharide
The outer membrane of C. jejuni contains LOS, which lacks O antigen that contains the repeating oligosaccharide polymer of the outer core of LPS of many bacteria. Instead, the outer cores of C. jejuni LOS are composed of a short oligosaccharide that is modified in some C. jejuni strains with sialic acid to create structures that resemble gangliosides found on human neurons and tissues, such as GM1, GM2, GM3, GD1a, GD3, GT1a and GD1c23,33 (Fig. 1). Producing human ganglioside-like epitopes may be a strategy to attenuate recognition by the host, and the initiation of an immune response is detrimental to the bacterium during infection. Addition of these ganglioside mimics reduces immunogenicity of the LOS while also increasing serum resistance34,35. Autoimmunity due to the production of antibodies against C. jejuni LOS that cross-react with gangliosides on human neurons can lead to Guillain–Barre syndrome and Miller–Fisher syndrome36,37,38,39.
Different genetic mechanisms contribute to interstrain and intrastrain diversity of LOS structures. C. jejuni strains can be grouped into 22 different classes based on LOS loci, with gene content influencing whether strains produce certain saccharides and transferases to modify LOS40. Further evolution of the range of ganglioside mimics produced has been promoted by natural frameshift or missense mutations that alter the activity of biosynthetic enzymes or transferases and by phase variation that reversibly turns on and off expression of key transferases (thoroughly discussed in ref.41). Altering ganglioside mimics through these mechanisms changes serum resistance, immunogenicity and invasion capacity of C. jejuni35. For example, phase variation of cgtA from the phase-on to phase-off state causes C. jejuni to switch from producing GM2-like LOS epitopes to GM3-like LOS epitopes, with a corresponding increase in serum resistance and invasion35.
Protein glycosylation
Although once considered largely a eukaryotic-specific activity, some bacteria have the ability to glycosylate specific proteins. C. jejuni contains two systems to specifically modify different sets of proteins: an O-linked protein glycosylation system that specifically modifies flagellins forming the extracellular flagellar filament, and an N-linked general protein glycosylation system that modifies many extra-cytoplasmic proteins.
O-linked glycosylation of C. jejuni flagellins results in up to 19 serine and threonine residues on a flagellin subunit being modified with pseudaminic acid (Pse) and derivatives containing acetyl and acetamindino groups (PseAcOAc or PseAm, respectively42,43,44) (Fig. 1). Some strains contain additional genes to synthesize related legionaminic acid (Leg) derivatives for flagellin glycosylation45,46,47. Flagellin glycosylation is essential for flagellins to polymerize into the extracellular filament for motility, although how the modifications contribute to filament polymerization is unknown42,44. As C. jejuni flagellin lacks a subdomain for TLR5 recognition that is involved in flagellin interactions for filament formation in other bacteria48, the O-linked glycans may directly contribute to subunit interactions or may indirectly contribute by altering flagellin conformation for subunit interactions.
Most, if not all, of the available ~19 serine and threonine residues in a flagellin subunit are glycosylated, which contributes to 10% of the protein mass. Although ~19 residues are glycosylated in a flagellin, glycosylation of only three to five residues is essential for filament formation, motility or autoagglutination facilitated by interactions between filaments on different C. jejuni cells, which may influence adhesion and microcolony formation on human intestinal cells49. For most strains, Pse is the predominant modification on serine or threonine residues of flagellin subunits. However, some residues also accommodate Pse derivatives, such as PseAm or PseAcOAc. By contrast, other residues seem to accept only a specific glycan (for example, PseAm) and are only modified with Pse or another moiety in mutants unable to produce the glycan usually attached to the amino acid residue42,44,45,46,47. This observation suggests a strategy to maintain maximal glycosylation independent of glycan availability. Heterogeneity of the modification of flagellin subunits is important as heterogeneity increases filament integrity, filament function and autoagglutination, which affects microcolony or biofilm formation to augment host interactions and persistence44,45,50,51.
Whereas many bacterial N-linked or O-linked glycosylation systems are dedicated to glycosylating one or just a few relatively abundant proteins, the C. jejuni N-linked protein glycosylation system modifies dozens of relatively average or low abundant periplasmic and outer membrane proteins with a specific glycan52,53,54 (Fig. 1). These proteins are modified at a specific asparagine residue within a consensus motif with a heptasaccharide containing five N-acetylgalactosamine residues, glucose, bacillosamine (Bac) and an attached phosphoethanolamine54,55. Mutation of this system affects adherence, invasion, chick colonization, mouse colonization, and the immunogenicity and lectin-binding ability of many proteins52,53,56,57,58. How glycosylation by this system may affect the biological activity of a protein is unknown. Owing to the conservation of this system in other Campylobacter strains such as Campylobacter coli, this system is expected to be broadly important across Campylobacter species. However, it is unclear if modification of only one or a small subset of C. jejuni proteins is essential for promoting beneficial activities or whether expansive modification of target proteins is required. So far, only the VirB10 protein involved in DNA uptake and transformation is known to require glycosylation to function59. N-linked glycosylation of surface proteins may prevent or lessen degradation by extracellular host proteases and facilitate interactions with the C-type lectin receptor MGL on immature dendritic cells and certain macrophages to limit the production of interleukin-6 (IL-6) during infection60,61. An alternative possibility has been put forward by the observation that only 10% of the glycan produced by this system is actually attached to a protein, with the remaining 90% found as free glycan in the periplasm62. This free glycan has been postulated to function as an osmoprotectant during osmotic stress. Therefore, it is possible that the C. jejuni N-linked protein glycosylation system produces glycans that affect C. jejuni biology in both the protein-linked and free states.
Genotypic and phenotypic variation
C. jejuni has an inherent ability to generate genotypic and phenotypic diversity owing to an augmented number of phase-variable loci. Furthermore, advantageous traits can quickly spread between strains, and the formation of new combinations of genetic alleles can be accelerated as the bacterium is naturally competent for DNA uptake and transformation. Recent genome-wide association studies have compared genetic variations enriched in human pathogenic isolates relative to their prevalence in poultry isolates in the same C. jejuni clonal complex63,64. These studies provide evidence for the generation of mutations that contribute to genetic and phenotypic variation as C. jejuni progresses from poultry farms to infection of humans. However, it is unclear when the mutations were acquired and whether the mutations benefit one or more steps in the progression of C. jejuni from growth in an avian host through meat processing and packaging to infection of humans.
Natural transformation
C. jejuni strains are genetically heterogenous with little clonality in population structure, which contributes to intraspecies variation. Nevertheless, C. jejuni can be highly structured into clusters of related isolates. This genetic heterogeneity creates a large gene pool for genomic recombination, adaptation and evolution of the species during colonization or transmission between hosts. Most intraspecies genetic diversity in C. jejuni is thought to be generated via horizontal gene transfer and recombination as the bacterium is naturally competent for DNA uptake and transformation65,66. Genomic rearrangements that contribute to genetic diversity have also been shown to occur during in vivo and in vitro growth of C. jejuni even in the absence of selection, although the mechanisms and stimuli that may promote these changes are unknown65,67.
C. jejuni uses an uptake system to transport DNA across membrane barriers into the cytoplasm, and the RecA recombinase for integration of homologous DNA into the chromosome68,69,70,71,72,73. A transposon mutagenesis screen identified components of the Cts type II secretion system (T2SS) as essential factors in C. jejuni for DNA uptake and transformation, which are conserved across strains69. Additionally, approximately 10% of C. jejuni strains contain the pVir plasmid, and VirB10 encoded on this plasmid contributes to natural transformation59,71. An essential DNA-binding receptor has remained elusive.
C. jejuni is highly discriminatory for the type of exogenous DNA for natural transformation, with selectivity for DNA from C. jejuni and other Campylobacter species over other bacteria74. Other naturally transformable bacteria, such as Neisseria species and Haemophilus species, recognize closely related DNA via a DNA uptake sequence (DUS) that occurs frequently in the genome75,76,77. However, C. jejuni is not naturally transformable by PCR-generated DNA or cloned C. jejuni DNA purified from Escherichia coli, suggesting that this bacterium does not use a DUS78. Recently, the Campylobacter transformation system methyltransferase (CtsM), which is highly conserved among Campylobacter species, was identified to methylate RAATTY sites on C. jejuni DNA to create suitable DNA substrates for transformation78. It is unclear whether this methylated DNA sequence is required for binding to a DNA receptor or a subsequent step in uptake and whether there is an advantage in using methylation versus a DUS for DNA recognition for C. jejuni in nature.
Phase variation
C. jejuni is exceptionally proficient at generating rapid intrastrain heterogeneity by phase variation. Phase variation enables the stochastic, reversible switching of phenotypes with high frequency owing to indels in homopolymeric nucleotide tracts or heteropolymeric nucleotide tracts in genes or promoter regions to promote efficient adaptation and survival of a subpopulation of the species. A given C. jejuni strain may contain around 30 contingency loci with nucleotide tracts that may reversibly phase vary during normal growth or specific conditions (Table 1). However, only around four to six of these contingency loci seem to be conserved across C. jejuni isolates, indicating that the number of contingency genes is variable across strains, but an individual contingency gene may differ between C. jejuni strains in possessing the nucleotide tracts for phase variation79.
Most contingency loci are involved in modification of surface structures (for example, CPS, LOS or flagellins). As discussed above, many transferases for LOS sialylation and CPS production phase vary to generate diverse structures within a C. jejuni population to evade immune responses or predation by bacteriophages. Another prominent phase-variable surface structure is the C. jejuni flagellum, with both the synthesis of the flagellum and the process of motility being phase variable80,81,82. In vivo and in vitro phase variation of flagellar biogenesis are due to polyA or polyT tracts within the coding sequence of flgS and flgR, which encode a two-component signal transduction system (TCS) for the expression of flagellar rod and hook genes essential for flagellar formation80,81. Additionally, the coding sequences for both genes can be restored by imperfect repair owing to insertion of a nucleotide elsewhere in the gene or indels of larger nucleotide sequences, demonstrating the broad intragenomic recombination capacity of C. jejuni to restore the highly beneficial motility phenotype80,81.
Flagellar motility phase varies owing to motA, which, along with motB, encodes the stators of the flagellar motor that generate power for flagellar rotation82. Phase variation of motA is due to loss of a nucleotide in a polyG tract or insertion of a 49-base repeat that disrupts the coding sequence, and both alterations can be restored completely. Loss of flagellar biogenesis or flagellar motility by phase variation hinders host colonization80,81. Thus, the advantages of in vivo phase variation of flagellar motility to create non-motile variants are unclear. However, an immune response to flagella during human infection may drive a subpopulation temporarily to a phase-off, non-motile state to survive a detrimental host response.
The genetic mechanisms of generating diversity by phase variation are unclear. C. jejuni has a functional RecA recombinase, a base-excision repair system and a nucleotide-excision repair system but lacks a mismatch repair (MMR) system containing MutS1, MutH and MutL proteins to repair DNA polymerase errors68. Instead, C. jejuni produces MutS2, the biological function of which is unknown. Recently, a natural mutator strain of C. jejuni has been identified that lacks a functional MutY protein, an adenine glycosylase involved in correcting adenine-7,8-dihydro-8-oxoguanine mismatches that lead to G:T and C:A transversions83. Mutations in mutY increase spontaneous mutation and emergence of antibiotic resistance rates 100-fold. A factor that seems to actually promote spontaneous mutation and fluoroquinolone resistance in C. jejuni is Mfd, a protein that in other bacteria removes stalled RNA polymerase at DNA lesions to facilitate DNA repair84. How Mfd facilitates spontaneous mutation and whether it promotes only specific mutations at certain loci for antibiotic resistance or is broadly mutagenic at other loci is unknown. Although the phase variation rates of C. jejuni are generally high compared with other bacteria, C. jejuni does not seem to be a natural hypermutator based on spontaneous point mutation rates79. It has been suggested that C. jejuni lacks a repair system for indels associated with polynucleotide tracts that contribute to high phase variation rates79.
The roles of the flagellum
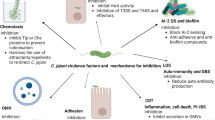
Besides being involved in chemotactic motility, the C. jejuni flagella also have roles in other important biological activities, including secretion of virulence and colonization factors, cellular division and affecting regulatory systems (Fig. 2). Thus, the C. jejuni flagellum must be inherently equipped for this multitasking.
The flagellum is composed of an MS ring (dark blue) and C ring (orange) that surround the flagellar type III secretion system (fT3SS) core in the inner membrane, a rod and hook structure (dark blue) that transverses the periplasm and outer membrane and an extracellular flagellar filament (red). Three disk structures, the basal disk (dark green), medial disk (bright green) and proximal disk (bright red), surround the flagellar rod in the periplasm. An isosurface rendering of a longitudinal slice of a tomogram of the C. jejuni flagellar motor obtained by electron cryotomography that reveals the flagellar motor structure is shown (upper left panel). The basal disk is composed of FlgP, the medial disk is composed of paralyzed flagellum protein A (PflA) and the proximal disk is composed of PflB and the MotAB stators. These disk structures incorporate an increased number of stator complexes into the motor and position them at a wider distance from the motor axis to contribute to an increased amount of torque relative to many other bacterial flagellar motors. The flagellar motor switch proteins FliF and FliG multimerize around the fT3SS core proteins in the inner membrane to form the MS ring and rotor of the C ring, respectively, during the initial stage of flagellar biogenesis. The FlgS sensor kinase detects the formation of the MS ring and rotor as a regulatory checkpoint during flagellar biogenesis, probably through direct interaction of multimers of FliF and/or FliG (upper right). After detection, autophosphorylation of FlgS initiates signal transduction to result in phosphorylation of the FlgR response regulator and expression of σ54-dependent flagellar rod and hook genes and eventually leads to expression of σ28-dependent flagellins and fed gene expression. Flagellar proteins are secreted in a specific order by the fT3SS to build the flagellum. Some Cia and Fed proteins are also secreted by the flagellum to influence host interactions. Polar flagellar biogenesis is regulated in a GTP-dependent manner by the flagellar motor switch protein FlhF GTPase and the FlhG ATPase. FlhG seems to control the active and inactive states of FlhF by promoting FlhF GTPase activity, which may influence both flagellar placement and the number of flagella at poles. FlhG also influences a process that prevents the polymerization of the cell division protein FtsZ into the septal Z ring at a pole so that the Z ring forms at the cellular midpoint for symmetrical division. FlhF, fT3SS proteins, FliF, and FliM and FliN C ring proteins influence the ability of FlhG alone or together with other unknown proteins to inhibit Z ring formation at poles. Tomogram of the C. jejuni flagellar motor courtesy of M. Beeby, Imperial College London, UK.
Motility and signal transduction
As with other motile bacteria, the C. jejuni flagellum is integrated with a chemotaxis system for chemoattraction and chemorepulsion to migrate to native niches within mucous layers of the lower intestinal tract in avian hosts and humans ideal for growth and host association9,58,85,86. C. jejuni displays chemotactic responses to aspartate, asparagine, tricarboxylic acid (TCA) cycle intermediates, formate, lactate and chicken mucus, with some strains having additional chemotactic responses towards galactose and fucose87,88,89,90,91.
As C. jejuni primarily exists in association with the host intestinal tract, the C. jejuni flagellum has evolved as an ideal motor to power propulsion through viscous milieus, such as intestinal mucus, with a velocity around ~40 μm per second in viscosities that normally impede other motile bacteria14,15,16,17. As such, the C. jejuni flagellum is equipped with additional components to generate power for creating a higher level of torque for flagellar rotation and motility relative to other bacteria. Electron cryotomography to resolve flagellar motor structures in situ revealed three large multimeric disk structures (annotated as basal, medial and proximal disks) that surround the flagellar rod and ring structures between the outer and inner membranes92 (Fig. 2). These disk structures are composed of FlgP, paralyzed flagellum protein A (PflA) and PflB and form scaffolds to incorporate MotAB stator complexes into the proximal disk to power rotation via proton transport and impart greater torque on the flagellar rotor93. Whereas E. coli flagellar motors contain at most 11 stators to generate power94, the C. jejuni flagellar motor incorporates 17 stators and orients the stators via the disk scaffolds at a greater radial distance from the central motor axis and rotor93. This numerical and spatial stator arrangement creates a more powerful motor that generates a higher level of torque for greater propulsion of C. jejuni through a range of viscosities93.
The cytoplasmic and inner membrane substructures of the flagellum influence other processes in C. jejuni. The core of the flagellar type III secretion system (fT3SS) is located in the inner membrane and is surrounded by the MS ring (a multimer of flagellar motor switch protein FliF) and the cytoplasmic C ring (multimers of the FliG rotor and flagellar switch proteins) (Fig. 2). These components are required for secretion of most proteins that form the flagellar rod, hook and filament. Formation of the MS ring and rotor around the fT3SS core by FliF and FliG in C. jejuni creates a regulatory checkpoint detected by the FlgSR TCS to activate σ54-dependent expression of flagellar rod and hook genes86,95,96,97,98,99. Mutation of any fT3SS protein, FliF or FliG abolishes FlgSR-dependent and σ54-dependent flagellar gene expression96,97,99. Physical detection of MS ring and rotor formation around the fT3SS core by two-component system histidine kinase (FlgS) ensures that a competent secretory system has formed before resources are expended to produce substrates for the fT3SS to build a flagellum96. Thus, the C. jejuni flagellum influences signal transduction for its own biogenesis.
The C. jejuni amphitrichous flagella also promote a darting motility characterized by rapid back-and-forth movements along a short linear path, which may increase contact with human epithelial cells to augment adherence and invasion15,16,17. C. jejuni spatially and numerically regulates flagellar biogenesis to create and maintain the amphitrichous flagellation pattern. The C. jejuni flagellar biosynthesis protein FlhF GTPase is required for flagellar biogenesis, and mutants with altered GTPase activity produce heterogenous flagellation phenotypes, including normal amphitrichous flagella, lateral flagella, polar hyperflagellation or aflagellation97,100,101 (Fig. 2). The placement of the fT3SS–MS ring–C ring complex on the cell body most likely determines the ultimate position of the nascent flagellum. Although it is unknown how FlhF functions in flagellation, FlhF may regulate polar positioning or organization of fT3SS, MS ring and C ring proteins in a GTP-dependent manner for polar flagellation100,101.
The FlhG ATPase seems to mediate numerical control of flagellation through FlhF101,102. FlhG stimulates the in vitro GTPase activity of FlhF, which likely converts it from an active GTP-bound state that facilitates a step in polar flagellar biogenesis to an inactive GDP-bound state101 (Fig. 2). It has been hypothesized that accurate control of FlhF activity via FlhG ensures exactly one flagellum is formed at each C. jejuni pole for amphitrichous flagellation, which is ideal for motility.
Flagellum-dependent regulation of cell division
C. jejuni flagellar components also influence spatial control of cell division through FlhG102. C. jejuni lacks a canonical bacterial Min system that spatially regulates septal Z ring formation so that it forms at the midcell for symmetrical division rather than at a pole. C. jejuni FlhG shares homology with the MinD ATPase of the Min system but does not encode orthologues of other Min proteins, such as MinC, which inhibits cell division protein FtsZ polymerization into the Z ring. C. jejuni ΔflhG produces a high level of non-viable minicells, which are products of asymmetrical division occurring at poles that often consequently lack chromosomal DNA102. C. jejuni mutants lacking the FliF MS ring protein, C ring switch proteins, fT3SS proteins or FlhF produce high levels of minicells102. Thus, polar flagellar formation, which may begin with FlhF producing the initial fT3SS-MS ring-C ring structure of a single flagellum at a new, unflagellated pole immediately after symmetrical division, appears to influence FlhG (either alone or with other unknown proteins) to inhibit septal Z ring formation at a pole so that a Z ring forms at the cellular midpoint for symmetrical division101,102 (Fig. 2). These findings suggest a possible additional explanation for an often-pondered question — why does C. jejuni produce a fairly rare amphitrichous flagellation pattern? The amphitrichous flagellation pattern appears ideal for the darting motility and efficient migration through viscous milieus encountered naturally but also has a role beyond motility in linking polar flagellation to a process that prevents division at a pole for accurate symmetrical division and efficient generation of viable progeny.
Flagellum-dependent protein secretion
Most fT3SSs specifically recognize only flagellar proteins as substrates for secretion to construct flagella. However, some bacterial fT3SSs, including those of Yersinia enterocolitica and C. jejuni, secrete proteins not involved in flagellar motility103,104,105,106,107,108,109,110. The Cia proteins, Fed proteins and flagellin C (FlaC) are secreted by the C. jejuni fT3SS (refs103,104,105,106,107,108,109,111,112). The Cia proteins were first discovered as bile-inducible, fT3SS-secreted proteins not involved in motility but that instead influence C. jejuni interactions with human intestinal cells103,111,112,113. Some Cia proteins have been reported to localize inside eukaryotic cells to influence C. jejuni interactions, although it is unclear the essentiality of secreted Cia proteins for adherence to and invasion of eukaryotic cells across different strains108,114,115,116,117. The Fed proteins are co-expressed with many flagellar proteins, and some of these proteins are secreted by the fT3SS (refs106,107,108,116). These proteins largely influence the commensal colonization capacity of C. jejuni for chicks, although individual functions of the Feds are unknown107,108. FlaC is a flagellin-like protein that is secreted but does not influence motility105. Instead, FlaC influences invasion of human intestinal cells and recently has been shown to modulate immune responses by promoting cross tolerance to some TLRs to reduce cytokine production105,109. This expanded secretory activity of the C. jejuni flagellum and its influence on signal transduction and cell division broaden the multitasking function of this organelle beyond its role as a powerful motor for propulsion.
Metabolism
C. jejuni has evolved metabolic pathways that are well adapted to the conditions (such as temperature and oxygen tension) and nutrients that are present in host intestinal niches. Reflective of its commensal lifestyle in avian hosts, C. jejuni grows well at 42ºC, which is the body temperature of chickens and fowl, with no obvious thermal stress normally observed for many bacterial enteric pathogens. C. jejuni is also an obligate microaerophile that requires oxygen for growth but at lower levels than in the atmosphere (~3–10% O2). However, many enteric bacterial pathogens are facultative anaerobes. The oxygen requirement for C. jejuni is due to an oxygen-dependent ribonucleotide reductase to generate nucleotides for DNA synthesis118. Oxygen is also a preferred electron acceptor for one of two respiratory oxidases for microaerobic growth119 (Fig. 3). However, the essential pyruvate:acceptor oxidoreductase (POR) and the TCA cycle 2-oxoglutarate:acceptor oxidoreductase (OOR) are rapidly inactivated by oxygen, and other important catabolism enzymes, such as SdaA, which converts serine into pyruvate, are also oxygen sensitive120. C. jejuni produces two hemerythrins, which are primarily found in anaerobes and microaerophiles and protect these enzymes during oxygen exposure120. Residing within the mucous layer, close to the epithelium of the lower intestinal tract of avian and human hosts, is expected to meet the low, but essential, oxygen needs of C. jejuni, as the lumen contains higher oxygen tension that may impair in vivo growth9,121.
Campylobacter jejuni predominantly relies on amino acids and keto acids as carbon sources to fuel the tricarboxylic acid (TCA) cycle and many anabolic pathways. The major metabolic pathways shown are for the amino acids serine, glutamine, asparagine, glutamate, aspartate and proline, which are those preferentially used by C. jejuni for metabolism. Some amino acids, such as glutamine and asparagine, may be converted by γ-glutamyltranspeptidase (GGT) or l-asparaginase (AnsA), respectively, to other amino acids prior to transport. Major transporters for amino acids and other metabolites are shown in the periplasm at the top. Serine is a major amino acid utilized for the generation of pyruvate, which can feed into the acetogenesis pathway or be converted to oxaloacetate for the TCA cycle or as a substrate for phosphoenolpyruvate (PEP). PEP is a substrate for gluconeogenesis required for capsular polysaccharide (CPS) and lipooligosaccharide (LOS) biogenesis and protein glycosylation. Lactate and fucose can be converted into pyruvate; however, not all C. jejuni strains have the capacity for fucose transport and catabolism. Respiratory oxidases (cbb3-type cytochrome c oxidase (Cco) and cyanide-insensitive cytochrome bd-like quinol oxidase (Cio)) that use oxygen as the major terminal electron acceptor and systems that use specific metabolites as electron donors (formate dehydrogenase (Fdh), hydrogenase (Hyd), sulfite:cytochrome c oxidoreductase (SOR) and gluconate dehydrogenase (GADH)) are shown at the bottom. Blue boxes indicate enzymes or transport systems that are not present in all strains. Yellow boxes indicate enzymes (l-serine dehydratase (SdaA), pyruvate:acceptor oxidoreductase (POR) and 2-oxoglutarate:acceptor oxidoreductase (OOR)) that are especially sensitive to oxygen exposure. Ac-CoA, acetyl-CoA; Frd,[definition]; FucP, l-fucose permease; LctP, l-lactate permease; PutP, proline:sodium symporter PutP; SdaC, l-serine transporter.
In vitro and in vivo analysis of C. jejuni catabolism and anabolism have revealed important insights into nutrients and pathways that are available and essential for growth. C. jejuni catabolism involves a fairly narrow substrate range of carbon and energy sources but an expansive anabolism122,123,124,125. These various catabolic and anabolic pathways generally have limited redundancy.
Carbohydrate catabolism is extremely limited in C. jejuni, with only a subset of strains that catabolize fucose89,126,127. Fucose catabolism is not essential for colonization of avian species but may provide a competitive advantage in other hosts126,127. Instead, amino acids and peptides provide carbon for energy generation and anabolism. Serine, aspartate, asparagine, glutamine, glutamate and proline are amino acids preferentially used for catabolism to support growth, with C. jejuni using specific transporters for the acquisition of these amino acids (reviewed in detail in ref.8) (Fig. 3). These amino acids are ideal for in vivo growth as they are the most abundant amino acids in the avian and human lower intestines128,129. Serine catabolism alone results in anabolism of at least ten other amino acids, and serine itself can be converted into pyruvate, which feeds into acetogenesis via the conversion of acetyl-CoA (Ac-CoA) and the TCA cycle via conversion of oxaloacetate122,124,130,131. The conversion of oxaloacetate into phosphoenolpyruvate is the entry point into gluconeogenesis, which is essential for the production of saccharides for LOS, CPS and protein glycosylation systems122,132. Also, glutamate and aspartate can be converted directly into 2-oxoglutarate and fumarate to feed the TCA cycle123. C. jejuni has many other biosynthetic pathways and transporters for other amino acids that are not catabolized but are directly shuttled into protein translation133.
Interactions with the gut microbiota and its metabolites are expected to affect C. jejuni colonization in both animal and human hosts. C. jejuni produces a number of respiratory systems that use a range of metabolites that are supplied by the host-associated microbiota as electron donors and acceptors, such as formate, hydrogen, sulfite, lactate, gluconate, mesaconate and crotonate, which expand its ability to thrive during in vivo intestinal growth (reviewed in ref.8). Furthermore, a recent study found that C. jejuni seems to sense in vivo short-chain fatty acids (SCFAs) and organic acids that are produced by the microbiota as cues to identify ideal niches in avian and human hosts that contain nutrients to support growth7. Transcription of many C. jejuni amino acid uptake and catabolism systems required for optimal host colonization is positively influenced by butyrate and acetate, which are major SCFAs produced by the microbiota in the niches of the lower intestinal tract for C. jejuni. However, lactate, an organic acid most abundant in the upper regions of the intestines, where C. jejuni less efficiently colonizes, represses the expression of these genes7. Thus, microbiota-derived metabolites may enable C. jejuni to spatially discriminate between different areas of the intestinal tracts in hosts in addition to supplying nutrients and metabolites for optimal growth and persistence.
Another recent report has suggested that microbiota activity in converting bile salts to secondary bile salts can influence the pathogenesis of C. jejuni134. Secreted bile salts in the intestines are converted to secondary bile salts by the microbiota, which were found to dampen the ability of C. jejuni to cause colitis in IL-10-deficient mice. An interesting finding from this work was that the generation of secondary bile salts did not decrease the viability or load of C. jejuni in the intestinal lumen but seemed to reduce the ability of C. jejuni to invade the intestinal mucosa of the mice. These recent studies suggest that much more knowledge is to be gained regarding the impact of the microbiota and its metabolism on the ability of C. jejuni to colonize natural hosts for commensalism or infect susceptible hosts to result in diarrhoeal disease.
Changes in the physiology of C. jejuni that are characterized by a metabolic downshift during which the bacterium does not replicate have been observed with intracellular C. jejuni in human intestinal cells135,136,137. During the adaptation to a low oxygen environment within cells, the levels of proteins that are involved in catabolic and anabolic pathways have been reported to be reduced, while fumarate respiration is increased135. Full recovery of C. jejuni after invasion requires either prolonged exposure to very low oxygen levels before growth in microaerobic conditions or exogenous electron donors to improve respiration135,136. These findings indicate the complex, yet efficient, means that C. jejuni possesses for growth in various hosts and intestinal environments, in addition to steps in pathogenesis of disease.
Conclusions
C. jejuni has differentiated itself from many enteric bacterial pathogens in terms of the types of relationships it forms with animal and human hosts that enable it to persist in nature and cause disease. C. jejuni has also distinguished itself by the collection of factors and mechanisms that are necessary for infection of and growth within a host and seems to lack some virulence factors commonly used by enteric pathogens for infection and disease (for example, multiple toxins, canonical injectisome T3SSs and effector proteins). The unique combination of determinants C. jejuni requires for infection include multiple systems that mediate LOS modification and two different protein glycosylation systems, a multifunctional flagellum, a variable CPS, an amino acid-based metabolism with multiple respiratory systems and a transformation system with altered target DNA recognition to promote genetic recombination and generate diversity. Many of these systems are complex and require numerous proteins, with many different metabolic and biochemical activities, and are unlikely to have been acquired through a single evolutionary step. It is unclear how C. jejuni evolved such complex systems, and owing to their conservation and retention in the species, these systems seem to be beneficial. Compared with typical enteric bacterial pathogens, the C. jejuni genome is two to three times smaller, but it has reserved genomic space to encode these large, presumably energy expensive, complex systems and encodes the composition of factors to efficiently promote infection in both avian and mammalian hosts. Furthermore, as a species, C. jejuni displays amazing intrastrain diversity, making it difficult to develop antibiotics, therapeutics or vaccines to combat its infectivity, virulence or presence in agriculture or food production.
An interesting and paradoxical observation about the biology of C. jejuni is that it can be such a major cause of diarrhoeal disease worldwide and is rampant in agriculture and many wild animals with such apparently restrictive growth requirements. For an organism that is a microaerophile requiring low oxygen concentrations in the environment and whose growth is hindered at temperatures below ~33ºC, it would seem difficult for the bacterium to survive ex vivo within environmental faecal droppings, contaminated aqueous sources, and frozen and refrigerated meats or raw milk, and to maintain viability and infectivity for transmission to subsequent hosts to achieve high persistence in the world. Undoubtedly, insightful and interesting ex vivo survival mechanisms related to transmission are yet to be discovered and revealed for this bacterium.
Many research groups have made great strides in unravelling the biological mechanisms of C. jejuni, providing some insights into why these factors might be important for infection and in vivo growth, as well as for the progression to commensalism or disease. For many of these factors, a mechanistic understanding of how they function in vivo is generally lacking. The C. jejuni field is in need of better animal model systems that efficiently reproduce pathological symptoms of diarrhoeal disease that are observed in humans infected with the bacterium. The development of mouse model systems, among others, has been improving by identifying microbiota components and knockout strains that better replicate diarrhoeal disease observed in humans upon infection with C. jejuni, but these models are not yet ideal. Furthermore, a natural commensalism model involving the use of chickens is currently available for C. jejuni. While chickens are easily colonized by C. jejuni, fewer reagents and molecular tools are available for the avian system than for other animal models. If both of these model systems continue to develop, the future will be a fascinating time to study the role and biological context of this unusual set of factors C. jejuni has acquired to infect and maintain residence in animals and humans.
References
Hale, C. R. et al. Estimates of enteric illness attributable to contact with animals and their environments in the United States. Clin. Infect. Dis. 54 (Suppl. 5), S472–S479 (2012).
Friedman, C. R. et al. Risk factors for sporadic Campylobacter infection in the United States: a case-control study in FoodNet sites. Clin. Infect. Dis. 38 (Suppl. 3), S285–S296 (2004).
Centers for Disease Control & Prevention. Surveillance for waterborne disease outbreaks associated with drinking water and other nonrecreational water - United States, 2009–2010. MMWR Morb. Mortal. Wkly Rep. 62, 714–720 (2013).
Costard, S., Espejo, L., Groenendaal, H. & Zagmutt, F. J. Outbreak-related disease burden associated with consumption of unpasteurized cow’s milk and cheese, United States, 2009–2014. Emerg. Infect. Dis. 23, 957–964 (2017).
Kaakoush, N. O., Castano-Rodriguez, N., Mitchell, H. M. & Man, S. M. Global epidemiology of Campylobacter infection. Clin. Microbiol. Rev. 28, 687–720 (2015).
Marder, E. P. et al. Incidence and trends of infections with pathogens transmitted commonly through food and the effect of increasing use of culture-independent diagnostic tests on surveillance - foodborne diseases active surveillance network, 10 U. S. Sites, 2013–2016. MMWR Morb. Mortal. Wkly Rep. 66, 397–403 (2017).
Luethy, P. M. et al. Microbiota-derived short-chain fatty acids modulate expression of Campylobacter jejuni determinants required for commensalism and virulence. mBio 8, e00407–00417 (2017).
Hofreuter, D. Defining the metabolic requirements for the growth and colonization capacity of Campylobacter jejuni. Front. Cell. Infect. Microbiol. 4, 137 (2014).
Beery, J. T., Hugdahl, M. B. & Doyle, M. P. Colonization of gastrointestinal tracts of chicks by Campylobacter jejuni. Appl. Environ. Microbiol. 54, 2365–2370 (1988).
Humphrey, S. et al. Campylobacter jejuni is not merely a commensal in commercial broiler chickens and affects bird welfare. mBio 5, e01364–01314 (2014).
Blaser, M. J. Epidemiologic and clinical features of Campylobacter jejuni infections. J. Infect. Dis. 176 (Suppl. 2), S103–S105 (1997).
Sahin, O., Yaeger, M., Wu, Z. & Zhang, Q. Campylobacter-associated diseases in animals. Annu. Rev. Anim. Biosci. 5, 21–42 (2017).
Wu, Z. et al. Point mutations in the major outer membrane protein drive hypervirulence of a rapidly expanding clone of Campylobacter jejuni. Proc. Natl Acad. Sci. USA 113, 10690–10695 (2016). This study reveals the very limited and specific genetic differences between an ovine C. jejuni abortifacient strain and a human C. jejuni diarrhoeal isolate.
Ferrero, R. L. & Lee, A. Motility of Campylobacter jejuni in a viscous environment: comparison with conventional rod-shaped bacteria. J. Gen. Microbiol. 134, 53–59 (1988).
Shigematsu, M., Umeda, A., Fujimoto, S. & Amako, K. Spirochaete-like swimming mode of Campylobacter jejuni in a viscous environment. J. Med. Microbiol. 47, 521–526 (1998).
Szymanski, C. M., King, M., Haardt, M. & Armstrong, G. D. Campylobacter jejuni motility and invasion of Caco-2 cells. Infect. Immun. 63, 4295–4300 (1995).
Karim, Q. N., Logan, R. P., Puels, J., Karnholz, A. & Worku, M. L. Measurement of motility of Helicobacter pylori. Campylobacter jejuni, and Escherichia coli by real time computer tracking using the Hobson BacTracker. J. Clin. Pathol. 51, 623–628 (1998).
Frirdich, E. et al. Peptidoglycan-modifying enzyme Pgp1 is required for helical cell shape and pathogenicity traits in Campylobacter jejuni. PLoS Pathog. 8, e1002602 (2012). This study identifies a type of peptidoglycan processing that is required for the characteristic helical cell shape of C. jejuni.
Frirdich, E. et al. Peptidoglycan LD-carboxypeptidase Pgp2 influences Campylobacter jejuni helical cell shape and pathogenic properties and provides the substrate for the DL-carboxypeptidase Pgp1. J. Biol. Chem. 289, 8007–8018 (2014).
Stahl, M. et al. The helical shape of Campylobacter jejuni promotes in vivo pathogenesis by aiding transit through intestinal mucus and colonization of crypts. Infect. Immun. 84, 3399–3407 (2016).
Karlyshev, A. V., Linton, D., Gregson, N. A., Lastovica, A. J. & Wren, B. W. Genetic and biochemical evidence of a Campylobacter jejuni capsular polysaccharide that accounts for Penner serotype specificity. Mol. Microbiol. 35, 529–541 (2000). This work demonstrates that the CPS determines the Penner serotype, which was previously thought to be due to LOS antigens.
Guerry, P. et al. Campylobacter polysaccharide capsules: virulence and vaccines. Front. Cell. Infect. Microbiol. 2, 7 (2012).
Szymanski, C. M. et al. Detection of conserved N-linked glycans and phase-variable lipooligosaccharides and capsules from campylobacter cells by mass spectrometry and high resolution magic angle spinning NMR spectroscopy. J. Biol. Chem. 278, 24509–24520 (2003).
St Michael, F. et al. The structures of the lipooligosaccharide and capsule polysaccharide of Campylobacter jejuni genome sequenced strain NCTC 11168. Eur. J. Biochem. 269, 5119–5136 (2002).
Karlyshev, A. V. et al. Analysis of Campylobacter jejuni capsular loci reveals multiple mechanisms for the generation of structural diversity and the ability to form complex heptoses. Mol. Microbiol. 55, 90–103 (2005). This report provides a summary of different mechanisms that contribute to CPS diversity in C. jejuni.
McNally, D. J. et al. Commonality and biosynthesis of the O-methyl phosphoramidate capsule modification in Campylobacter jejuni. J. Biol. Chem. 282, 28566–28576 (2007).
Holst Sorensen, M. C. et al. Phase variable expression of capsular polysaccharide modifications allows Campylobacter jejuni to avoid bacteriophage infection in chickens. Front. Cell. Infect. Microbiol. 2, 11 (2012). This study reports CPS changes that can occur in vitro and in vivo to protect C. jejuni from bacteriophage infection, while commensal colonization of chickens remains unaffected.
Sorensen, M. C. et al. Bacteriophage F336 recognizes the capsular phosphoramidate modification of Campylobacter jejuni NCTC11168. J. Bacteriol. 193, 6742–6749 (2011).
Pequegnat, B. et al. Phase-variable changes in the position of O-methyl phosphoramidate modifications on the polysaccharide capsule of Campylobacter jejuni modulate serum resistance. J. Bacteriol. 199, e00027–e00017 (2017).
Aidley, J., Sorensen, M. C. H., Bayliss, C. D. & Brondsted, L. Phage exposure causes dynamic shifts in the expression states of specific phase-variable genes of Campylobacter jejuni. Microbiology 163, 911–919 (2017).
van Alphen, L. B. et al. Biological roles of the O-methyl phosphoramidate capsule modification in Campylobacter jejuni. PLoS ONE 9, e87051 (2014).
Maue, A. C. et al. The polysaccharide capsule of Campylobacter jejuni modulates the host immune response. Infect. Immun. 81, 665–672 (2013).
Moran, A. P. Structure and conserved characteristics of Campylobacter jejuni lipopolysaccharides. J. Infect. Dis. 176 (Suppl. 2), S115–S121 (1997).
Guerry, P., Ewing, C. P., Hickey, T. E., Prendergast, M. M. & Moran, A. P. Sialylation of lipooligosaccharide cores affects immunogenicity and serum resistance of Campylobacter jejuni. Infect. Immun. 68, 6656–6662 (2000).
Guerry, P. et al. Phase variation of Campylobacter jejuni 81–176 lipooligosaccharide affects ganglioside mimicry and invasiveness in vitro. Infect. Immun. 70, 787–793 (2002).
Ang, C. W. et al. Structure of Campylobacter jejuni lipopolysaccharides determines antiganglioside specificity and clinical features of Guillain-Barre and Miller Fisher patients. Infect. Immun. 70, 1202–1208 (2002).
Yuki, N. Molecular mimicry between gangliosides and lipopolysaccharides of Campylobacter jejuni isolated from patients with Guillain-Barre syndrome and Miller Fisher syndrome. J. Infect. Dis. 176 (Suppl. 2), S150–153 (1997).
Yuki, N. & Miyatake, T. Guillain-Barre syndrome and Fisher’s syndrome following Campylobacter jejuni infection. Ann. NY Acad. Sci. 845, 330–340 (1998).
Yuki, N. et al. Carbohydrate mimicry between human ganglioside GM1 and Campylobacter jejuni lipooligosaccharide causes Guillain-Barre syndrome. Proc. Natl Acad. Sci. USA 101, 11404–11409 (2004).
Richards, V. P., Lefebure, T., Pavinski Bitar, P. D. & Stanhope, M. J. Comparative characterization of the virulence gene clusters (lipooligosaccharide [LOS] and capsular polysaccharide [CPS]) for Campylobacter coli. Campylobacter jejuni subsp. jejuni and related Campylobacter species. Infect. Genet. Evol. 14, 200–213 (2013).
Gilbert, M. et al. The genetic basis for the variation in the lipo-oligosaccharide of the mucosal pathogen, Campylobacter jejuni. J. Biol. Chem. 277, 327–337 (2002). The study presents various genetic mechanisms and loci compositions and provides insights into how variation of LOS structures is achieved across C. jejuni strains.
Thibault, P. et al. Identification of the carbohydrate moieties and glycosylation motifs in Campylobacter jejuni flagellin. J. Biol. Chem. 276, 34862–34870 (2001).
McNally, D. J. et al. Functional characterization of the flagellar glycosylation locus in Campylobacter jejuni 81–176 using a focused metabolomics approach. J. Biol. Chem. 281, 18489–18498 (2006).
Goon, S., Kelly, J. F., Logan, S. M., Ewing, C. P. & Guerry, P. Pseudaminic acid, the major modification on Campylobacter flagellin, is synthesized via the Cj1293 gene. Mol. Microbiol. 50, 659–671 (2003).
Howard, S. L. et al. Campylobacter jejuni glycosylation island important in cell charge, legionaminic acid biosynthesis, and colonization of chickens. Infect. Immun. 77, 2544–2556 (2009).
Ulasi, G. N., Creese, A. J., Hui, S. X., Penn, C. W. & Cooper, H. J. Comprehensive mapping of O-glycosylation in flagellin from Campylobacter jejuni 11168: a multienzyme differential ion mobility mass spectrometry approach. Proteomics 15, 2733–2745 (2015).
Zebian, N. et al. Comprehensive analysis of flagellin glycosylation in Campylobacter jejuni NCTC 11168 reveals incorporation of legionaminic acid and its importance for host colonization. Glycobiology 26, 386–397 (2016).
Andersen-Nissen, E. et al. Evasion of toll-like receptor 5 by flagellated bacteria. Proc. Natl Acad. Sci. USA 102, 9247–9252 (2005).
Ewing, C. P., Andreishcheva, E. & Guerry, P. Functional characterization of flagellin glycosylation in Campylobacter jejuni 81–176. J. Bacteriol. 191, 7086–7093 (2009). This study reports that only a small number of specific residues in a flagellin subunit must be glycosylated for full flagellin function in motility, flagellar formation and autoaggregation.
Guerry, P. et al. Changes in flagellin glycosylation affect Campylobacter autoagglutination and virulence. Mol. Microbiol. 60, 299–311 (2006).
van Alphen, L. B. et al. A functional Campylobacter jejuni maf4 gene results in novel glycoforms on flagellin and altered autoagglutination behaviour. Microbiology 154, 3385–3397 (2008).
Szymanski, C. M., Yao, R., Ewing, C. P., Trust, T. J. & Guerry, P. Evidence for a system of general protein glycosylation in Campylobacter jejuni. Mol. Microbiol. 32, 1022–1030 (1999). This study presents the initial evidence that C. jejuni has a general protein glycosylation system.
Linton, D., Allan, E., Karlyshev, A. V., Cronshaw, A. D. & Wren, B. W. Identification of N-acetylgalactosamine-containing glycoproteins PEB3 and CgpA in Campylobacter jejuni. Mol. Microbiol. 43, 497–508 (2002).
Young, N. M. et al. Structure of the N-linked glycan present on multiple glycoproteins in the Gram-negative bacterium, Campylobacter jejuni. J. Biol. Chem. 277, 42530–42539 (2002).
Scott, N. E. et al. Modification of the Campylobacter jejuni N-linked glycan by EptC protein-mediated addition of phosphoethanolamine. J. Biol. Chem. 287, 29384–29396 (2012).
Szymanski, C. M., Burr, D. H. & Guerry, P. Campylobacter protein glycosylation affects host cell interactions. Infect. Immun. 70, 2242–2244 (2002).
Karlyshev, A. V. et al. The Campylobacter jejuni general glycosylation system is important for attachment to human epithelial cells and in the colonization of chicks. Microbiology 150, 1957–1964 (2004).
Hendrixson, D. R. & DiRita, V. J. Identification of Campylobacter jejuni genes involved in commensal colonization of the chick gastrointestinal tract. Mol. Microbiol. 52, 471–484 (2004). This is the first study to use transposon mutagenesis to identify C. jejuni genes required for commensalism in chickens.
Larsen, J. C., Szymanski, C. & Guerry, P. N-Linked protein glycosylation is required for full competence in Campylobacter jejuni 81–176. J. Bacteriol. 186, 6508–6514 (2004).
van Sorge, N. M. et al. N-Glycosylated proteins and distinct lipooligosaccharide glycoforms of Campylobacter jejuni target the human C-type lectin receptor MGL. Cell. Microbiol. 11, 1768–1781 (2009).
Alemka, A., Nothaft, H., Zheng, J. & Szymanski, C. M. N-Glycosylation of Campylobacter jejuni surface proteins promotes bacterial fitness. Infect. Immun. 81, 1674–1682 (2013).
Nothaft, H., Liu, X., McNally, D. J., Li, J. & Szymanski, C. M. Study of free oligosaccharides derived from the bacterial N-glycosylation pathway. Proc. Natl Acad. Sci. USA 106, 15019–15024 (2009).
Yahara, K. et al. Genome-wide association of functional traits linked with Campylobacter jejuni survival from farm to fork. Environ. Microbiol. 19, 361–380 (2017).
Buchanan, C. J. et al. A genome-wide association study to identify diagnostic markers for human pathogenic Campylobacter jejuni strains. Front. Microbiol. 8, 1224 (2017).
de Boer, P. et al. Generation of Campylobacter jejuni genetic diversity in vivo. Mol. Microbiol. 44, 351–359 (2002).
Wang, Y. & Taylor, D. E. Natural transformation in Campylobacter species. J. Bacteriol. 172, 949–955 (1990).
Ridley, A. M., Toszeghy, M. J., Cawthraw, S. A., Wassenaar, T. M. & Newell, D. G. Genetic instability is associated with changes in the colonization potential of Campylobacter jejuni in the avian intestine. J. Appl. Microbiol. 105, 95–104 (2008).
Gaasbeek, E. J. et al. Functional characterization of excision repair and RecA-dependent recombinational DNA repair in Campylobacter jejuni. J. Bacteriol. 191, 3785–3793 (2009).
Wiesner, R. S., Hendrixson, D. R. & DiRita, V. J. Natural transformation of Campylobacter jejuni requires components of a type II secretion system. J. Bacteriol. 185, 5408–5418 (2003).
Beauchamp, J. M., Erfurt, R. S. & DiRita, V. J. Characterization and localization of the Campylobacter jejuni transformation system proteins CtsE, CtsP, and CtsX. J. Bacteriol. 197, 636–645 (2015).
Bacon, D. J. et al. Involvement of a plasmid in virulence of Campylobacter jejuni 81–176. Infect. Immun. 68, 4384–4390 (2000).
Jeon, B., Muraoka, W., Sahin, O. & Zhang, Q. Role of Cj1211 in natural transformation and transfer of antibiotic resistance determinants in Campylobacter jejuni. Antimicrob. Agents Chemother. 52, 2699–2708 (2008).
Jeon, B. & Zhang, Q. Cj0011c, a periplasmic single- and double-stranded DNA-binding protein, contributes to natural transformation in Campylobacter jejuni. J. Bacteriol. 189, 7399–7407 (2007).
Wassenaar, T. M., Fry, B. N. & van der Zeijst, B. A. Genetic manipulation of Campylobacter: evaluation of natural transformation and electro-transformation. Gene 132, 131–135 (1993).
Elkins, C., Thomas, C. E., Seifert, H. S. & Sparling, P. F. Species-specific uptake of DNA by gonococci is mediated by a 10-base-pair sequence. J. Bacteriol. 173, 3911–3913 (1991).
Mell, J. C., Hall, I. M. & Redfield, R. J. Defining the DNA uptake specificity of naturally competent Haemophilus influenzae cells. Nucleic Acids Res. 40, 8536–8549 (2012).
Goodman, S. D. & Scocca, J. J. Identification and arrangement of the DNA sequence recognized in specific transformation of Neisseria gonorrhoeae. Proc. Natl Acad. Sci. USA 85, 6982–6986 (1988).
Beauchamp, J. M., Leveque, R. M., Dawid, S. & DiRita, V. J. Methylation-dependent DNA discrimination in natural transformation of Campylobacter jejuni. Proc. Natl Acad. Sci. USA 114, E8053–E8061 (2017). This study shows that a methylase and specific modifications of a DNA sequence are important for DNA recognition by C. jejuni for natural transformation.
Bayliss, C. D. et al. Phase variable genes of Campylobacter jejuni exhibit high mutation rates and specific mutational patterns but mutability is not the major determinant of population structure during host colonization. Nucleic Acids Res. 40, 5876–5889 (2012).
Hendrixson, D. R. A phase-variable mechanism controlling the Campylobacter jejuni FlgR response regulator influences commensalism. Mol. Microbiol. 61, 1646–1659 (2006).
Hendrixson, D. R. Restoration of flagellar biosynthesis by varied mutational events in Campylobacter jejuni. Mol. Microbiol. 70, 519–536 (2008).
Mohawk, K. L., Poly, F., Sahl, J. W., Rasko, D. A. & Guerry, P. High frequency, spontaneous motA mutations in Campylobacter jejuni strain 81–176. PLoS ONE 9, e88043 (2014).
Dai, L., Sahin, O., Tang, Y. & Zhang, Q. A mutator phenotype promoting the emergence of spontaneous oxidative stress resistant mutants in Campylobacter jejuni. Appl. Environ. Microbiol. 83, e01685–17 (2017).
Han, J., Sahin, O., Barton, Y. W. & Zhang, Q. Key role of Mfd in the development of fluoroquinolone resistance in Campylobacter jejuni. PLoS Pathog. 4, e1000083 (2008). This report identifies Mfd as a factor that promotes spontaneous mutagenesis, rather than correcting DNA replication errors, that contributes to fluoroquinolone resistance in C. jejuni.
Black, R. E., Levine, M. M., Clements, M. L., Hughes, T. P. & Blaser, M. J. Experimental Campylobacter jejuni infection in humans. J. Infect. Dis. 157, 472–479 (1988).
Wosten, M. M. S. M., Wagenaar, J. A. & van Putten, J. P. M. The FlgS/FlgR two-component signal transduction system regulates the fla regulon in Campylobacter jejuni. J. Biol. Chem. 279, 16214–16222 (2004).
Hartley-Tassell, L. E. et al. Identification and characterization of the aspartate chemosensory receptor of Campylobacter jejuni. Mol. Microbiol. 75, 710–730 (2010).
Rahman, H. et al. Characterisation of a multi-ligand binding chemoreceptor CcmL (Tlp3) of Campylobacter jejuni. PLoS Pathog. 10, e1003822 (2014).
Dwivedi, R. et al. L-Fucose influences chemotaxis and biofilm formation in Campylobacter jejuni. Mol. Microbiol. 101, 575–589 (2016).
Vegge, C. S., Brondsted, L., Li, Y. P., Bang, D. D. & Ingmer, H. Energy taxis drives Campylobacter jejuni toward the most favorable conditions for growth. Appl. Environ. Microbiol. 75, 5308–5314 (2009).
Day, C. J. et al. A direct-sensing galactose chemoreceptor recently evolved in invasive strains of Campylobacter jejuni. Nat. Commun. 7, 13206 (2016).
Chen, S. et al. Structural diversity of bacterial flagellar motors. EMBO J. 30, 2972–2981 (2011).
Beeby, M. et al. Diverse high-torque bacterial flagellar motors assemble wider stator rings using a conserved protein scaffold. Proc. Natl Acad. Sci. USA 113, E1917–E1926 (2016). This work identifies a structural scaffold for incorporation of stator complexes into the C. jejuni flagellar motor that likely contributes to increased torque relative to other motile bacteria to power flagellar rotation and motility.
Reid, S. W. et al. The maximum number of torque-generating units in the flagellar motor of Escherichia coli is at least 11. Proc. Natl Acad. Sci. USA 103, 8066–8071 (2006).
Boll, J. M. & Hendrixson, D. R. A specificity determinant for phosphorylation in a response regulator prevents in vivo cross-talk and modification by acetyl phosphate. Proc. Natl Acad. Sci. USA 108, 20160–20165 (2011).
Boll, J. M. & Hendrixson, D. R. A regulatory checkpoint during flagellar biogenesis in Campylobacter jejuni initiates signal transduction to activate transcription of flagellar genes. mBio 4, e00432–e00413 (2013).
Hendrixson, D. R. & DiRita, V. J. Transcription of σ54-dependent but not σ28-dependent flagellar genes in Campylobacter jejuni is associated with formation of the flagellar secretory apparatus. Mol. Microbiol. 50, 687–702 (2003).
Joslin, S. N. & Hendrixson, D. R. Analysis of the Campylobacter jejuni FlgR response regulator suggests integration of diverse mechanisms to activate an NtrC-like protein. J. Bacteriol. 190, 2422–2433 (2008).
Joslin, S. N. & Hendrixson, D. R. Activation of the Campylobacter jejuni FlgSR two-component system is linked to the flagellar export apparatus. J. Bacteriol. 191, 2656–2667 (2009).
Balaban, M., Joslin, S. N. & Hendrixson, D. R. FlhF and its GTPase activity are required for distinct processes in flagellar gene regulation and biosynthesis in Campylobacter jejuni. J. Bacteriol. 191, 6602–6611 (2009).
Gulbronson, C. J. et al. FlhG employs diverse intrinsic domains and influences FlhF GTPase activity to numerically regulate polar flagellar biogenesis in Campylobacter jejuni. Mol. Microbiol. 99, 291–306 (2016).
Balaban, M. & Hendrixson, D. R. Polar flagellar biosynthesis and a regulator of flagellar number influence spatial parameters of cell division in Campylobacter jejuni. PLoS Pathog. 7, e1002420 (2011). This work demonstrates that FlhG and amphitrichous flagellation are involved in the spatial regulation of symmetrical division, which is the canonical function of the Min system that is lacking in C. jejuni.
Konkel, M. E., Kim, B. J., Rivera-Amill, V. & Garvis, S. G. Bacterial secreted proteins are required for the internaliztion of Campylobacter jejuni into cultured mammalian cells. Mol. Microbiol. 32, 691–701 (1999).
Konkel, M. E. et al. Secretion of virulence proteins from Campylobacter jejuni is dependent on a functional flagellar export apparatus. J. Bacteriol. 186, 3296–3303 (2004).
Song, Y. C. et al. FlaC, a protein of Campylobacter jejuni TGH9011 (ATCC43431) secreted through the flagellar apparatus, binds epithelial cells and influences cell invasion. Mol. Microbiol. 53, 541–553 (2004).
Poly, F. et al. Heterogeneity of a Campylobacter jejuni protein that is secreted through the flagellar filament. Infect. Immun. 75, 3859–3867 (2007).
Barrero-Tobon, A. M. & Hendrixson, D. R. Identification and analysis of flagellar coexpressed determinants (Feds) of Campylobacter jejuni involved in colonization. Mol. Microbiol. 84, 352–369 (2012).
Barrero-Tobon, A. M. & Hendrixson, D. R. Flagellar biosynthesis exerts temporal regulation of secretion of specific Campylobacter jejuni colonization and virulence determinants. Mol. Microbiol. 93, 957–974 (2014).
Faber, E. et al. Novel immunomodulatory flagellin-like protein FlaC in Campylobacter jejuni and other Campylobacterales. mSphere 1, 00028–00015 (2016).
Young, G. M., Schmiel, D. H. & Miller, V. L. A new pathway for the secretion of virulence factors by bacteria: the flagellar export apparatus functions as a protein-secretion system. Proc. Natl Acad. Sci. USA 96, 6456–6461 (1999).
Buelow, D. R., Christensen, J. E., Neal-McKinney, J. M. & Konkel, M. E. Campylobacter jejuni survival within human epithelial cells is enhanced by the secreted protein CiaI. Mol. Microbiol. 80, 1296–1312 (2011).
Christensen, J. E., Pacheco, S. A. & Konkel, M. E. Identification of a Campylobacter jejuni-secreted protein required for maximal invasion of host cells. Mol. Microbiol. 73, 650–662 (2009). This work identifies many potential proteins that are secreted by the flagella of C. jejuni that may have a role in host interactions.
Rivera-Amill, V., Kim, B. J., Seshu, J. & Konkel, M. E. Secretion of the virulence-associated Campylobacter invasion antigens from Campylobacter jejuni requires a stimulatory signal. J. Infect. Dis. 183, 1607–1616 (2001).
Neal-McKinney, J. M. & Konkel, M. E. The Campylobacter jejuni CiaC virulence protein is secreted from the flagellum and delivered to the cytosol of host cells. Front. Cell. Infect. Microbiol. 2, 31 (2012).
Samuelson, D. R. et al. The Campylobacter jejuni CiaD effector protein activates MAP kinase signaling pathways and is required for the development of disease. Cell Commun. Signal 11, 79 (2013).
Goon, S. et al. A σ28-regulated nonflagella gene contributes to virulence of Campylobacter jejuni 81–176. Infect. Immun. 74, 769–772 (2006).
Novik, V., Hofreuter, D. & Galan, J. E. Identification of Campylobacter jejuni genes involved in its interaction with epithelial cells. Infect. Immun. 78, 3540–3553 (2010).
Sellars, M. J., Hall, S. J. & Kelly, D. J. Growth of Campylobacter jejuni supported by respiration of fumarate, nitrate, nitrite, trimethylamine-N-oxide, or dimethyl sulfoxide requires oxygen. J. Bacteriol. 184, 4187–4196 (2002).
Jackson, R. J. et al. Oxygen reactivity of both respiratory oxidases in Campylobacter jejuni: the cydAB genes encode a cyanide-resistant, low-affinity oxidase that is not of the cytochrome bd type. J. Bacteriol. 189, 1604–1615 (2007).
Kendall, J. J., Barrero-Tobon, A. M., Hendrixson, D. R. & Kelly, D. J. Hemerythrins in the microaerophilic bacterium Campylobacter jejuni help protect key iron-sulphur cluster enzymes from oxidative damage. Environ. Microbiol. 16, 1105–1121 (2014).
Lee, A., O’Rourke, J. L., Barrington, P. J. & Trust, T. J. Mucus colonization as a determinant of pathogenicity in intestinal infection by Campylobacter jejuni: a mouse cecal model. Infect. Immun. 51, 536–546 (1986).
Gao, B. et al. Metabolic and fitness determinants for in vitro growth and intestinal colonization of the bacterial pathogen Campylobacter jejuni. PLoS Biol. 15, e2001390, (2017). This work reports a thorough isotopologue profiling analysis and an insertion sequencing (INSeq) genetic screen to identify metabolic pathways important for in vitro and in vivo growth of C. jejuni.
Guccione, E. et al. Amino acid-dependent growth of Campylobacter jejuni: key roles for aspartase (AspA) under microaerobic and oxygen-limited conditions and identification of AspB (Cj0762), essential for growth on glutamate. Mol. Microbiol. 69, 77–93 (2008).
Velayudhan, J., Jones, M. A., Barrow, P. A. & Kelly, D. J. L-Serine catabolism via an oxygen-labile L-serine dehydratase is essential for colonization of the avian gut by Campylobacter jejuni. Infect. Immun. 72, 260–268 (2004).
Leon-Kempis Mdel, R., Guccione, E., Mulholland, F., Williamson, M. P. & Kelly, D. J. The Campylobacter jejuni PEB1a adhesin is an aspartate/glutamate-binding protein of an ABC transporter essential for microaerobic growth on dicarboxylic amino acids. Mol. Microbiol. 60, 1262–1275 (2006).
Stahl, M. et al. L-Fucose utilization provides Campylobacter jejuni with a competitive advantage. Proc. Natl Acad. Sci. USA 108, 7194–7199 (2011).
Muraoka, W. T. & Zhang, Q. Phenotypic and genotypic evidence for L-fucose utilization by Campylobacter jejuni. J. Bacteriol. 193, 1065–1075 (2011).
Ahlman, B., Leijonmarck, C. E., Lind, C., Vinnars, E. & Wernerman, J. Free amino acids in biopsy specimens from the human colonic mucosa. J. Surg. Res. 55, 647–653 (1993).
Parsons, C. M., Potter, L. M. & Brown, R. D. Jr Effects of dietary carbohydrate and of intestinal microflora on excretion of endogenous amino acids by poultry. Poult Sci. 62, 483–489 (1983).
Wright, J. A. et al. Metabolite and transcriptome analysis of Campylobacter jejuni in vitro growth reveals a stationary-phase physiological switch. Microbiology 155, 80–94 (2009).
Mendz, G. L., Ball, G. E. & Meek, D. J. Pyruvate metabolism in Campylobacter spp. Biochim. Biophys. Acta 1334, 291–302 (1997).
Velayudhan, J. & Kelly, D. J. Analysis of gluconeogenic and anaplerotic enzymes in Campylobacter jejuni: an essential role for phosphoenolpyruvate carboxykinase. Microbiology 148, 685–694 (2002).
Ribardo, D. A. & Hendrixson, D. R. Analysis of the LIV system of Campylobacter jejuni reveals alternative roles for LivJ and LivK in commensalism beyond branched-chain amino acid transport. J. Bacteriol. 193, 6233–6243 (2011).
Sun, X. et al. Microbiota-derived metabolic factors reduce Campylobacteriosis in mice. Gastroenterology 154, 1751–1763.e2 (2018).
Watson, R. O. & Galan, J. E. Campylobacter jejuni survives within epithelial cells by avoiding delivery to lysosomes. PLoS Pathog. 4, e14 (2008). This work provides insights into the intracellular fate and residence of C. jejuni upon infection of human cells.
Pryjma, M., Apel, D., Huynh, S., Parker, C. T. & Gaynor, E. C. FdhTU-modulated formate dehydrogenase expression and electron donor availability enhance recovery of Campylobacter jejuni following host cell infection. J. Bacteriol. 194, 3803–3813 (2012).
Liu, X., Gao, B., Novik, V. & Galan, J. E. Quantitative proteomics of intracellular Campylobacter jejuni reveals metabolic reprogramming. PLoS Pathog. 8, e1002562 (2012).
Masanta, W. O. et al. Modification of intestinal microbiota and its consequences for innate immune response in the pathogenesis of campylobacteriosis. Clin. Dev. Immunol. 2013, 526860 (2013).
Bax, M. et al. Campylobacter jejuni lipooligosaccharides modulate dendritic cell-mediated T cell polarization in a sialic acid linkage-dependent manner. Infect. Immun. 79, 2681–2689 (2011).
Fox, J. G. et al. Gastroenteritis in NF-kappaB-deficient mice is produced with wild-type Camplyobacter jejuni but not with C. jejuni lacking cytolethal distending toxin despite persistent colonization with both strains. Infect. Immun. 72, 1116–1125 (2004).
Mansfield, L. S. et al. C57BL/6 and congenic interleukin-10-deficient mice can serve as models of Campylobacter jejuni colonization and enteritis. Infect. Immun. 75, 1099–1115 (2007).
Stahl, M. et al. A novel mouse model of Campylobacter jejuni gastroenteritis reveals key pro-inflammatory and tissue protective roles for Toll-like receptor signaling during infection. PLoS Pathog. 10, e1004264 (2014).
Watson, R. O., Novik, V., Hofreuter, D., Lara-Tejero, M. & Galan, J. E. A. MyD88-deficient mouse model reveals a role for Nramp1 in Campylobacter jejuni infection. Infect. Immun. 75, 1994–2003 (2007).
Han, Z. et al. Influence of the gut microbiota composition on Campylobacter jejuni colonization in chickens. Infect. Immun. 85, e00380–17 (2017).
Han, Z. et al. Differences in host breed and diet influence colonization by Campylobacter jejuni and induction of local immune responses in chicken. Gut Pathog. 8, 56 (2016).
Backert, S., Boehm, M., Wessler, S. & Tegtmeyer, N. Transmigration route of Campylobacter jejuni across polarized intestinal epithelial cells: paracellular, transcellular or both? Cell Commun. Signal 11, 72 (2013).
Rubinchik, S., Seddon, A. & Karlyshev, A. V. Molecular mechanisms and biological role of Campylobacter jejuni attachment to host cells. Eur. J. Microbiol. Immunol. 2, 32–40 (2012).
Hu, L. & Kopecko, D. J. Campylobacter jejuni 81–176 associates with microtubules and dynein during invasion of human intestinal cells. Infect. Immun. 67, 4171–4182 (1999).
Hu, L., Tall, B. D., Curtis, S. K. & Kopecko, D. J. Enhanced microscopic definition of Campylobacter jejuni 81–176 adherence to, invasion of, translocation across, and exocytosis from polarized human intestinal Caco-2 cells. Infect. Immun. 76, 5294–5304 (2008).
Oelschlaeger, T. A., Guerry, P. & Kopecko, D. J. Unusual microtubule-dependent endocytosis mechanisms triggered by Campylobacter jejuni and Citrobacter freundii. Proc. Natl Acad. Sci. USA 90, 6884–6888 (1993). This is one of the first analyses of how C. jejuni invades human intestinal epithelial cells.
Whitehouse, C. A. et al. Campylobacter jejuni cytolethal distending toxin causes a G2-phase cell cycle block. Infect. Immun. 66, 1934–1940 (1998).
Lara-Tejero, M. & Galan, J. E. A bacterial toxin that controls cell cycle progression as a deoxyribonuclease I-like protein. Science 290, 354–357 (2000).
Abuoun, M. et al. Cytolethal distending toxin (CDT)-negative Campylobacter jejuni strains and anti-CDT neutralizing antibodies are induced during human infection but not during colonization in chickens. Infect. Immun. 73, 3053–3062 (2005).
Mortensen, N. P. et al. The role of Campylobacter jejuni cytolethal distending toxin in gastroenteritis: toxin detection, antibody production, and clinical outcome. APMIS 119, 626–634 (2011).
Purdy, D. et al. Characterisation of cytolethal distending toxin (CDT) mutants of Campylobacter jejuni. J. Med. Microbiol. 49, 473–479 (2000).
Hickey, T. E. et al. Campylobacter jejuni cytolethal distending toxin mediates release of interleukin-8 from intestinal epithelial cells. Infect. Immun. 68, 6535–6541 (2000).
Zheng, J., Meng, J., Zhao, S., Singh, R. & Song, W. Campylobacter-induced interleukin-8 secretion in polarized human intestinal epithelial cells requires Campylobacter-secreted cytolethal distending toxin- and Toll-like receptor-mediated activation of NF-kappaB. Infect. Immun. 76, 4498–4508 (2008).
Joshua, G. W., Guthrie-Irons, C., Karlyshev, A. V. & Wren, B. W. Biofilm formation in Campylobacter jejuni. Microbiology 152, 387–396 (2006).
Kalmokoff, M. et al. Proteomic analysis of Campylobacter jejuni 11168 biofilms reveals a role for the motility complex in biofilm formation. J. Bacteriol. 188, 4312–4320 (2006).
Turonova, H. et al. Biofilm spatial organization by the emerging pathogen Campylobacter jejuni: comparison between NCTC 11168 and 81–176 strains under microaerobic and oxygen-enriched conditions. Front. Microbiol. 6, 709 (2015).
Ica, T. et al. Characterization of mono- and mixed-culture Campylobacter jejuni biofilms. Appl. Environ. Microbiol. 78, 1033–1038 (2012).
Reuter, M., Mallett, A., Pearson, B. M. & van Vliet, A. H. Biofilm formation by Campylobacter jejuni is increased under aerobic conditions. Appl. Environ. Microbiol. 76, 2122–2128 (2010).
Reeser, R. J., Medler, R. T., Billington, S. J., Jost, B. H. & Joens, L. A. Characterization of Campylobacter jejuni biofilms under defined growth conditions. Appl. Environ. Microbiol. 73, 1908–1913 (2007).
Teh, A. H., Lee, S. M. & Dykes, G. A. Does Campylobacter jejuni form biofilms in food-related environments? Appl. Environ. Microbiol. 80, 5154–5160 (2014).
Hanning, I., Jarquin, R. & Slavik, M. Campylobacter jejuni as a secondary colonizer of poultry biofilms. J. Appl. Microbiol. 105, 1199–1208 (2008).
Hilbert, F., Scherwitzel, M., Paulsen, P. & Szostak, M. P. Survival of Campylobacter jejuni under conditions of atmospheric oxygen tension with the support of Pseudomonas spp. Appl. Environ. Microbiol. 76, 5911–5917 (2010).
Kim, J. S. et al. Passage of Campylobacter jejuni through the chicken reservoir or mice promotes phase variation in contingency genes Cj0045 and Cj0170 that strongly associates with colonization and disease in a mouse model. Microbiology 158, 1304–1316 (2012).
Hitchen, P. et al. Modification of the Campylobacter jejuni flagellin glycan by the product of the Cj1295 homopolymeric-tract-containing gene. Microbiology 156, 1953–1962 (2010).
Miller, W. G. et al. Diversity within the Campylobacter jejuni type I restriction-modification loci. Microbiology 151, 337–351 (2005).
Karlyshev, A. V., Linton, D., Gregson, N. A. & Wren, B. W. A novel paralogous gene family involved in phase-variable flagella-mediated motility in Campylobacter jejuni. Microbiology 148, 473–480 (2002).
Acknowledgements
The authors apologize to colleagues whose relevant studies and works they could not specifically cite in this Review owing to space limitations. Work in the author’s laboratory was supported by National Institutes of Health (NIH) grant R01AI1065539.
Author contributions
P.M.B. and D.R.H. researched data for the article, made substantial contributions to discussions of the content, wrote the article and reviewed and/or edited the manuscript before submission.
Reviewer information
Nature Reviews Microbiology thanks Victoria Korolik, Mirko Rossi and the other, anonymous reviewer(s) for their contribution to the peer review of this work.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Glossary
- Amphitrichous flagella
-
A flagellation pattern in bacteria resulting in a single flagellum produced at each cellular pole.
- Peptidoglycan
-
A polymer of repeating N-acetylglucosamine and N-acetyl muramic acid subunits crosslinked by peptides that forms the major structural component of the cell wall of bacteria.
- Capsular polysaccharide
-
An extracellular polysaccharide that varies in composition between bacteria and may contribute to different activities, such as biofilm formation, immune evasion and protection from environmental stresses.
- Serotypes
-
Subsets of strains within a species that each produce different antigenic forms of a structural component that is usually on the surface of the bacterial cell.
- Phase variation
-
A reversible switch between phenotypes mediated by random DNA replication errors in homopolymeric or heteropolymeric nucleotide tracts within the coding sequence or promoter region of a gene.
- Gangliosides
-
Types of glycolipids that usually contain a sialic acid compound and are found in many mammalian tissues, especially neurons.
- O-linked protein glycosylation
-
The attachment of a glycan to the hydroxyl oxygen of an amino acid in a protein (specifically a serine or threonine residue for Campylobacter jejuni proteins).
- N-linked protein glycosylation
-
The attachment of a glycan to the amide nitrogen of an asparagine residue in a protein.
- Indels
-
An insertion or deletion of a base in a gene.
- Homopolymeric nucleotide tracts
-
A short stretch of identical nucleotides in a nucleic acid sequence.
- Heteropolymeric nucleotide tracts
-
A repeated DNA sequence composed of different nucleotides in a nucleic acid sequence.
- Min system
-
A spatial regulatory system that prevents the formation of the septal Z ring at the poles of bacteria so that it correctly forms at the midpoint for symmetrical division.
- Septal Z ring
-
A structure, usually composed of monomers of cell division protein FtsZ along with other proteins, that eventually divides the bacterial cell into two daughter cells during division.
Rights and permissions
About this article
Cite this article
Burnham, P.M., Hendrixson, D.R. Campylobacter jejuni: collective components promoting a successful enteric lifestyle. Nat Rev Microbiol 16, 551–565 (2018). https://doi.org/10.1038/s41579-018-0037-9
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41579-018-0037-9
- Springer Nature Limited
This article is cited by
-
Compendium of 5810 genomes of sheep and goat gut microbiomes provides new insights into the glycan and mucin utilization
Microbiome (2024)
-
Interplay of two small RNAs fine-tunes hierarchical flagella gene expression in Campylobacter jejuni
Nature Communications (2024)
-
Characterisation of N-linked protein glycosylation in the bacterial pathogen Campylobacter hepaticus
Scientific Reports (2023)
-
SVR-flaA typing of erythromycin- and ciprofloxacin-resistant Campylobacter jejuni strains isolated from poultry slaughterhouses in southern Brazil
Brazilian Journal of Microbiology (2023)
-
Development and validation of a CRISPR interference system for gene regulation in Campylobacter jejuni
BMC Microbiology (2022)