Abstract
Chromosomal instability (CIN) is a hallmark of cancer and a driver of metastatic dissemination, therapeutic resistance, and immune evasion. CIN is present in 60–80% of human cancers and poses a formidable therapeutic challenge as evidenced by the lack of clinically approved drugs that directly target CIN. This limitation in part reflects a lack of well-defined druggable targets as well as a dearth of tractable biomarkers enabling direct assessment and quantification of CIN in patients with cancer. Over the past decade, however, our understanding of the cellular mechanisms and consequences of CIN has greatly expanded, revealing novel therapeutic strategies for the treatment of chromosomally unstable tumours as well as new methods of assessing the dynamic nature of chromosome segregation errors that define CIN. In this Review, we describe advances that have shaped our understanding of CIN from a translational perspective, highlighting both challenges and opportunities in the development of therapeutic interventions for patients with chromosomally unstable cancers.
Key points
-
Chromosomal instability (CIN) has long been recognized as a hallmark of aggressive human malignancies and research over the past two decades has identified several novel therapeutic approaches for patients with chromosomally unstable cancers.
-
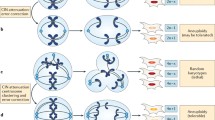
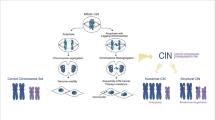
CIN drives cancer progression by generating genomic alterations such as chromosomal gains, losses and complex rearrangements. CIN can also drive the development of epigenetic abnormalities and chronic inflammation that facilitate both metastatic dissemination and immune evasion.
-
The development of synthetic lethal approaches in the context of CIN has led to the development of several novel treatment approaches, including KIF18A inhibitors, p53-reactivating agents and PLK4 inhibitors, all of which are currently being tested in clinical trials.
-
A mechanistic understanding of the cancer cell-intrinsic and immune-related vulnerabilities imposed by CIN will create opportunities for the development of a new class of therapies for patients with otherwise difficult-to-treat cancers.





Similar content being viewed by others
References
Orr, B., Godek, K. M. & Compton, D. Aneuploidy. Curr. Biol. 25, R538–542 (2015).
Lejeune, J. & Turpin, R. Chromosomal aberrations in man. Am. J. Hum. Genet. 13, 175–184 (1961).
Arber, D. A. et al. International consensus classification of myeloid neoplasms and acute leukemias: integrating morphologic, clinical, and genomic data. Blood 140, 1200–1228 (2022).
Torres, E. M. et al. Effects of aneuploidy on cellular physiology and cell division in haploid yeast. Science 317, 916–924 (2007).
Torres, E. M., Springer, M. & Amon, A. No current evidence for widespread dosage compensation in S. cerevisiae. eLife 5, e10996 (2016).
Santaguida, S., Vasile, E., White, E. & Amon, A. Aneuploidy-induced cellular stresses limit autophagic degradation. Genes Dev. 29, 2010–2021 (2015).
Lukow, D. A. et al. Chromosomal instability accelerates the evolution of resistance to anti-cancer therapies. Dev. Cell 56, 2427–2439.e4 (2021).
Ippolito, M. R. et al. Gene copy-number changes and chromosomal instability induced by aneuploidy confer resistance to chemotherapy. Dev. Cell 56, 2440–2454.e6 (2021).
Schvartzman, J. M., Duijf, P. H., Sotillo, R., Coker, C. & Benezra, R. Mad2 is a critical mediator of the chromosome instability observed upon Rb and p53 pathway inhibition. Cancer Cell 19, 701–714 (2011).
Nguyen, B. et al. Genomic characterization of metastatic patterns from prospective clinical sequencing of 25,000 patients. Cell 185, 563–575.e11 (2022).
Agustinus, A. S. et al. Epigenetic dysregulation from chromosomal transit in micronuclei. Nature 619, 176–183 (2023).
Papathanasiou, S. et al. Heritable transcriptional defects from aberrations of nuclear architecture. Nature 619, 184–192 (2023).
Beach, R. R. et al. Aneuploidy causes non-genetic individuality. Cell 169, 229–242.e21 (2017).
Martínez-Ruiz, C. et al. Genomic–transcriptomic evolution in lung cancer and metastasis. Nature 616, 543–552 (2023).
Bakhoum, S. F. et al. Chromosomal instability drives metastasis through a cytosolic DNA response. Nature 553, 467–472 (2018).
McPherson, A. et al. Divergent modes of clonal spread and intraperitoneal mixing in high-grade serous ovarian cancer. Nat. Genet 48, 758–767 (2016).
Roylance, R. et al. Relationship of extreme chromosomal instability with long-term survival in a retrospective analysis of primary breast cancer. Cancer Epidemiol. Biomark. Prev. 20, 2183–2194 (2011).
Stopsack, K. H. et al. Aneuploidy drives lethal progression in prostate cancer. Proc. Natl Acad. Sci. USA 116, 11390–11395 (2019).
Chibon, F., Lesluyes, T., Valentin, T. & Le Guellec, S. CINSARC signature as a prognostic marker for clinical outcome in sarcomas and beyond. Genes Chromosomes Cancer 58, 124–129 (2019).
Frankell, A. M. et al. The evolution of lung cancer and impact of subclonal selection in TRACERx. Nature 616, 525–533 (2023).
Sourty, B., La Basset, Ë., Garcion, E. & Rousseau, A. DNAR-01. Chromothripsis, one major genetic instability factor in glioblastoma, is rare in IDH-mutant gliomas. Neuro-Oncology 24, vii90 (2022).
Luebeck, J. et al. Extrachromosomal DNA in the cancerous transformation of Barrett’s oesophagus. Nature 616, 798–805 (2023).
Bakhoum, S. F. & Compton, D. A. Kinetochores and disease: keeping microtubule dynamics in check! Curr. Opin. Cell Biol. 24, 64–70 (2012).
de Oliveira Lisboa, M., Brofman, P. R. S., Schmid-Braz, A. T., Rangel-Pozzo, A. & Mai, S. Chromosomal instability in acute myeloid leukemia. Cancers 13, 2655 (2021).
Lee, A. J. et al. Chromosomal instability confers intrinsic multidrug resistance. Cancer Res 71, 1858–1870 (2011).
van Dijk, E. et al. Chromosomal copy number heterogeneity predicts survival rates across cancers. Nat. Commun. 12, 3188 (2021).
Al Bakir, M. et al. The evolution of non-small cell lung cancer metastases in TRACERx. Nature 616, 534–542 (2023).
Li, M. et al. The ATM–p53 pathway suppresses aneuploidy-induced tumorigenesis. Proc. Natl Acad. Sci. USA 107, 14188–14193 (2010).
Laughney, A. M., Elizalde, S., Genovese, G. & Bakhoum, S. F. Dynamics of tumor heterogeneity derived from clonal karyotypic evolution. Cell Rep. 12, 809–820 (2015).
Al-Rawi, D. H. & Bakhoum, S. F. Chromosomal instability as a source of genomic plasticity. Curr. Opin. Genet. Dev. 74, 101913 (2022).
Bakhoum, S. F. & Cantley, L. C. The multifaceted role of chromosomal instability in cancer and its microenvironment. Cell 174, 1347–1360 (2018).
Thompson, S. L., Bakhoum, S. F. & Compton, D. A. Mechanisms of chromosomal instability. Curr. Biol. 20, R285–295 (2010).
Bronder, D. & Bakhoum, S. F. A. CINful way to overcome addiction: how chromosomal instability enables cancer to overcome its oncogene addiction. EMBO Mol. Med. 12, e12017 (2020).
Sotillo, R., Schvartzman, J. M., Socci, N. D. & Benezra, R. Mad2-induced chromosome instability leads to lung tumour relapse after oncogene withdrawal. Nature 464, 436–440 (2010).
Harrold, E. et al. Molecular and clinical determinants of acquired resistance and treatment duration for targeted therapies in colorectal cancer. Clin. Cancer Res. 30, 2672–2683 (2024).
Liu, Y. L. et al. Subsequent therapies and survival after immunotherapy in recurrent ovarian cancer. Gynecol. Oncol. 155, 51–57 (2019).
Kaya, A. et al. Molecular signatures of aneuploidy-driven adaptive evolution. Nat. Commun. 11, 588 (2020).
Santaguida, S. & Amon, A. Short- and long-term effects of chromosome mis-segregation and aneuploidy. Nat. Rev. Mol. Cell Biol. 16, 473–485 (2015).
Schukken, K. M. & Foijer, F. CIN and aneuploidy: different concepts, different consequences. BioEssays 40, https://doi.org/10.1002/bies.201700147 (2018).
Baker, D. J. et al. BubR1 insufficiency causes early onset of aging-associated phenotypes and infertility in mice. Nat. Genet. 36, 744–749 (2004).
Macedo, J. C. et al. FoxM1 repression during human aging leads to mitotic decline and aneuploidy-driven full senescence. Nat. Commun. 9, 2834 (2018).
Andriani, G. A. et al. Whole chromosome instability induces senescence and promotes SASP. Sci. Rep. 6, 35218 (2016).
Santaguida, S. et al. Chromosome mis-segregation generates cell-cycle-arrested cells with complex karyotypes that are eliminated by the immune system. Dev. Cell 41, 638–651.e5 (2017).
Wang, R. W., Viganò, S., Ben-David, U., Amon, A. & Santaguida, S. Aneuploid senescent cells activate NF-κB to promote their immune clearance by NK cells. EMBO Rep. 22, e52032 (2021).
Hsu, S.-K. et al. Unfolded protein response (UPR) in survival, dormancy, immunosuppression, metastasis, and treatments of cancer cells. Int. J. Mol. Sci. 20, 2518 (2019).
Scharping, N. E. et al. The tumor microenvironment represses T cell mitochondrial biogenesis to drive intratumoral T cell metabolic insufficiency and dysfunction. Immunity 45, 374–388 (2016).
Reveil, P. O. Recherches de Physiologie Vegetale. De l’Action des Poisons sur les Plantes (E Martinet, 1865).
Gusev, Y., Kagansky, V. & Dooley, W. C. Long-term dynamics of chromosomal instability in cancer: a transition probability model. Math. Comput. Model. 33, 1253–1273 (2001).
Brennan, C. M. et al. Protein aggregation mediates stoichiometry of protein complexes in aneuploid cells. Genes Dev. 33, 1031–1047 (2019).
Oromendia, A. B., Dodgson, S. E. & Amon, A. Aneuploidy causes proteotoxic stress in yeast. Genes Dev. 26, 2696–2708 (2012).
Pfau, S. J. & Amon, A. Chromosomal instability and aneuploidy in cancer: from yeast to man. EMBO Rep. 13, 515–527 (2012).
Stingele, S. et al. Global analysis of genome, transcriptome and proteome reveals the response to aneuploidy in human cells. Mol. Syst. Biol. 8, 608 (2012).
Ohashi, A. et al. Aneuploidy generates proteotoxic stress and DNA damage concurrently with p53-mediated post-mitotic apoptosis in SAC-impaired cells. Nat. Commun. 6, 7668 (2015).
Deshaies, R. J. Proteotoxic crisis, the ubiquitin-proteasome system, and cancer therapy. BMC Biol. 12, 94 (2014).
Donnelly, N., Passerini, V., Dürrbaum, M., Stingele, S. & Storchová, Z. HSF1 deficiency and impaired HSP90‐dependent protein folding are hallmarks of aneuploid human cells. EMBO J. 33, 2374–2387 (2014).
Morimoto, R. I. Proteotoxic stress and inducible chaperone networks in neurodegenerative disease and aging. Genes Dev. 22, 1427–1438 (2008).
Li, J. et al. Non-cell-autonomous cancer progression from chromosomal instability. Nature 620, 1080–1088 (2023).
Li, J. et al. Metastasis and immune evasion from extracellular cGAMP hydrolysis. Cancer Discov. 11, 1212–1227 (2021).
Advani, V. M. & Ivanov, P. Translational control under stress: reshaping the translatome. BioEssays 41, e1900009 (2019).
Pan, T. Adaptive translation as a mechanism of stress response and adaptation. Annu. Rev. Genet. 47, 121–137 (2013).
Lee, Y. S., Shibata, Y., Malhotra, A. & Dutta, A. A novel class of small RNAs: tRNA-derived RNA fragments (tRFs). Genes Dev. 23, 2639–2649 (2009).
Sun, C. et al. Roles of tRNA-derived fragments in human cancers. Cancer Lett. 414, 16–25 (2018).
Calvo, V. et al. Discovery of 2-amino-3-amido-5-aryl-pyridines as highly potent, orally bioavailable, and efficacious PERK kinase inhibitors. Bioorg. Med. Chem. Lett. 43, 128058 (2021).
Stokes, M. E. et al. PERK inhibition by HC-5404 sensitizes renal cell carcinoma tumor models to antiangiogenic tyrosine kinase inhibitors. Clin. Cancer Res. 29, 4870–4882 (2023).
Ghosh, R. et al. Allosteric inhibition of the IRE1α RNase preserves cell viability and function during endoplasmic reticulum stress. Cell 158, 534–548 (2014).
Tufanli, O. et al. Targeting IRE1 with small molecules counteracts progression of atherosclerosis. Proc. Natl Acad. Sci. USA 114, E1395–E1404 (2017).
Gabrail, N. Y. et al. A phase 1/2 trial of ORIN1001, a first-in-class IRE1 inhibitor, in patients with advanced solid tumors. J. Clin. Oncol. 39, 3080 (2021).
Rimawi, M. F. et al. Early efficacy evaluation of ORIN1001, a first in class IRE1 alpha inhibitor, in advanced solid tumors. J. Clin. Oncol. 41, https://doi.org/10.1200/JCO.2023.41.16_suppl.1092 (2023).
Wilhelm, T. et al. Mild replication stress causes chromosome mis-segregation via premature centriole disengagement. Nat. Commun. 10, 3585 (2019).
Zeman, M. K. & Cimprich, K. A. Causes and consequences of replication stress. Nat. Cell Biol. 16, 2–9 (2014).
Gemble, S. et al. Genetic instability from a single S phase after whole-genome duplication. Nature 604, 146–151 (2022).
Peng, C. et al. The error-prone DNA polymerase κ promotes temozolomide resistance in glioblastoma through Rad17-dependent activation of ATR-Chk1 signaling. Cancer Res. 76, 2340–2353 (2016).
Patterson, K. et al. Altered RECQL5 expression in urothelial bladder carcinoma increases cellular proliferation and makes RECQL5 helicase activity a novel target for chemotherapy. Oncotarget 7, 76140–76150 (2016).
Urban, V., Dobrovolna, J. & Janscak, P. Distinct functions of human RecQ helicases during DNA replication. Biophys. Chem. 225, 20–26 (2017).
Viziteu, E. et al. RECQ1 helicase is involved in replication stress survival and drug resistance in multiple myeloma. Leukemia 31, 2104–2113 (2017).
Santana-Codina, N. et al. Oncogenic KRAS supports pancreatic cancer through regulation of nucleotide synthesis. Nat. Commun. 9, 4945 (2018).
Li, Q. et al. Rac1 activates non-oxidative pentose phosphate pathway to induce chemoresistance of breast cancer. Nat. Commun. 11, 1456 (2020).
Ge, X. Q., Jackson, D. A. & Blow, J. J. Dormant origins licensed by excess Mcm2–7 are required for human cells to survive replicative stress. Genes Dev. 21, 3331–3341 (2007).
Lee, C. H. et al. Telaglenastat plus everolimus in advanced renal cell carcinoma: a randomized, double-blinded, placebo-controlled, phase II ENTRATA trial. Clin. Cancer Res. 28, 3248–3255 (2022).
Meric-Bernstam, F. et al. Telaglenastat plus cabozantinib or everolimus for advanced or metastatic renal cell carcinoma: an open-label phase I trial. Clin. Cancer Res. 28, 1540–1548 (2022).
Turajlic, S. et al. Tracking cancer evolution reveals constrained routes to metastases: TRACERx Renal. Cell 173, 581–594.e12 (2018).
Addie, R. D. et al. Metabolic reprogramming related to whole-chromosome instability in models for Hürthle cell carcinoma. Sci. Rep. 10, 9578 (2020).
Warburg, O. On the origin of cancer cells. Science 123, 309–314 (1956).
Warburg, O. On respiratory impairment in cancer cells. Science 124, 269–270 (1956).
Dai, C., Sun, F., Zhu, C. & Hu, X. Tumor environmental factors glucose deprivation and lactic acidosis induce mitotic chromosomal instability – an implication in aneuploid human tumors. PLoS ONE 8, e63054 (2013).
Shaukat, Z. et al. Chromosomal instability causes sensitivity to metabolic stress. Oncogene 34, 4044–4055 (2015).
Bahreyni, A. et al. Role of adenosine signaling in the pathogenesis of breast cancer. J. Cell Physiol. 233, 1836–1843 (2018).
Barfeld, S. J. et al. Myc-dependent purine biosynthesis affects nucleolar stress and therapy response in prostate cancer. Oncotarget 6, 12587–12602 (2015).
Parker, W. B. Enzymology of purine and pyrimidine antimetabolites used in the treatment of cancer. Chem. Rev. 109, 2880–2893 (2009).
Newman, D. L. & Gregory, S. L. Operation between aneuploidy and metabolic changes in driving tumorigenesis. Int. J. Mol. Sci. 20, 4611 (2019).
Brault, V. et al. Opposite phenotypes of muscle strength and locomotor function in mouse models of partial trisomy and monosomy 21 for the proximal Hspa13-App region. PLoS Genet. 11, e1005062 (2015).
Clemente-Ruiz, M. et al. Gene dosage imbalance contributes to chromosomal instability-induced tumorigenesis. Dev. Cell 36, 290–302 (2016).
Prasad, S., Gupta, S. C. & Tyagi, A. K. Reactive oxygen species (ROS) and cancer: role of antioxidative nutraceuticals. Cancer Lett. 387, 95–105 (2017).
Ragu, S. et al. Oxygen metabolism and reactive oxygen species cause chromosomal rearrangements and cell death. Proc. Natl Acad. Sci. USA 104, 9747–9752 (2007).
Brunet, A. et al. Akt promotes cell survival by phosphorylating and inhibiting a forkhead transcription factor. Cell 96, 857–868 (1999).
Limaye, V. et al. Sphingosine kinase-1 enhances endothelial cell survival through a PECAM-1-dependent activation of PI-3K/Akt and regulation of Bcl-2 family members. Blood 105, 3169–3177 (2005).
Zhou, J., Chen, Y., Lang, J.-Y., Lu, J.-J. & Ding, J. Salvicine inactivates β1 integrin and inhibits adhesion of MDA-MB-435 cells to fibronectin via reactive oxygen species signaling. Mol. Cancer Res. 6, 194–204 (2008).
Liu, D. et al. Autophagy regulates the survival of cells with chromosomal instability. Oncotarget 7, 63913–63923 (2016).
Wu, H., Wang, M. C. & Bohmann, D. JNK protects Drosophila from oxidative stress by trancriptionally activating autophagy. Mech. Dev. 126, 624–637 (2009).
Cohen-Sharir, Y. et al. Aneuploidy renders cancer cells vulnerable to mitotic checkpoint inhibition. Nature 590, 486–491 (2021).
Payton, M. et al. Small-molecule inhibition of kinesin KIF18A reveals a mitotic vulnerability enriched in chromosomally unstable cancers. Nat. Cancer 5, 66–84 (2023).
Quinton, R. J. et al. Whole-genome doubling confers unique genetic vulnerabilities on tumour cells. Nature 590, 492–497 (2021).
Zhang, R. et al. Discovery of potent, orally active KIF18A inhibitors targeting CIN-high cancer cells. J. Clin. Oncol. 40, e15046 (2022).
Torres, E. M., Williams, B. R. & Amon, A. Aneuploidy: cells losing their balance. Genetics 179, 737–746 (2008).
Bielski, C. M. & Taylor, B. S. Homing in on genomic instability as a therapeutic target in cancer. Nat. Commun. 12, 3663 (2021).
Morden, C. R. et al. Chromosome instability is prevalent and dynamic in high-grade serous ovarian cancer patient samples. Gynecol. Oncol. 161, 769–778 (2021).
Colombo, N. et al. Tolerability of maintenance olaparib in newly diagnosed patients with advanced ovarian cancer and a BRCA mutation in the randomized phase III SOLO1 trial. Gynecol. Oncol. 163, 41–49 (2021).
Gonzalez-Martin, A. et al. Niraparib in patients with newly diagnosed advanced ovarian cancer. N. Engl. J. Med. 381, 2391–2402 (2019).
Wethington, S. L. et al. Combination ATR (ceralasertib) and PARP (olaparib) inhibitor (CAPRI) trial in acquired PARP inhibitor-resistant homologous recombination-deficient ovarian cancer. Clin. Cancer Res. 29, 2800–2807 (2023).
Kulkarni, S. et al. Evolving DNA repair synthetic lethality targets in cancer. Biosci. Rep. 42, BSR20221713 (2022).
Weaver, B. A. How Taxol/paclitaxel kills cancer cells. Mol. Biol. Cell 25, 2677–2681 (2014).
Marquis, C. et al. Chromosomally unstable tumor cells specifically require KIF18A for proliferation. Nat. Commun. 12, 1213 (2021).
Sousa-Pimenta, M. et al. Chemotherapeutic properties and side-effects associated with the clinical practice of terpene alkaloids: paclitaxel, docetaxel, and cabazitaxel. Front. Pharmacol. 14, 1157306 (2023).
Komlodi-Pasztor, E., Sackett, D. L. & Fojo, A. T. Inhibitors targeting mitosis: tales of how great drugs against a promising target were brought down by a flawed rationale. Clin. Cancer Res. 18, 51–63 (2012).
Van den Bossche, J. et al. Spotlight on volasertib: preclinical and clinical evaluation of a promising Plk1 inhibitor. Med. Res. Rev. 36, 749–786 (2016).
Benten, D. et al. Aurora kinase inhibitor PHA-739358 suppresses growth of hepatocellular carcinoma in vitro and in a xenograft mouse model. Neoplasia 11, 934–944 (2009).
Serrano-Del Valle, A. et al. Future prospects for mitosis-targeted antitumor therapies. Biochem. Pharmacol. 190, 114655 (2021).
Belmontes, B. et al. Abstract 516: dscovery and preclinical characterization of AMG 650, a first-in-class inhibitor of kinesin KIF18A motor protein with potent activity against chromosomally unstable cancers. Cancer Res. 83, 516 (2023).
Haykal, M. M., Rodrigues-Ferreira, S. & Nahmias, C. Aneuploidy triggers vulnerability to WEE1 inhibition via severe chromosome pulverization. Preprint at bioRxiv https://doi.org/10.1101/2023.09.19.558475 (2023).
Lheureux, S. et al. Adavosertib plus gemcitabine for platinum-resistant or platinum-refractory recurrent ovarian cancer: a double-blind, randomised, placebo-controlled, phase 2 trial. Lancet 397, 281–292 (2021).
Schutte, T. et al. Clinical development of WEE1 inhibitors in gynecological cancers: a systematic review. Cancer Treat. Rev. 115, 102531 (2023).
Liu, J. F. et al. Correlation of cyclin E1 expression and clinical outcomes in a phase 1b dose-escalation study of azenosertib (ZN-c3), a WEE1 inhibitor, in combination with chemotherapy (CT) in patients (pts) with platinum-resistant or refractory (R/R) epithelial ovarian, peritoneal, or fallopian tube cancer (EOC). J. Clin. Oncol. 41, https://doi.org/10.1200/JCO.2023.41.16_suppl.5513 (2023).
Gelderblom, H. et al. Debio 0123-101: A phase 1 trial of Debio 0123 in combination with carboplatin in advanced solid tumors — safety, pharmacokinetic, and preliminary antitumor activity data. J. Clin. Oncol. 41, https://doi.org/10.1200/JCO.2023.41.16_suppl.3012 (2023).
Guiley, K. Z. & Shokat, K. M. A small molecule reacts with the p53 somatic mutant Y220C to rescue wild-type thermal stability. Cancer Discov. 13, 56–69 (2023).
Dumbrava, E. E. et al. First-in-human study of PC14586, a small molecule structural corrector of Y220C mutant p53, in patients with advanced solid tumors harboring a TP53 Y220C mutation. J. Clin. Oncol. 40, 3003 (2022).
Murphy, T. et al. Preclinical characterization and clinical trial of CFI-400945, a polo-like kinase 4 inhibitor, in patients with relapsed/refractory acute myeloid leukemia and higher-risk myelodysplastic neoplasms. Leukemia 38, 502–512 (2024).
Meitinger, F. et al. TRIM37 controls cancer-specific vulnerability to PLK4 inhibition. Nature 585, 440–446 (2020).
Yeow, Z. Y. et al. Targeting TRIM37-driven centrosome dysfunction in 17q23-amplified breast cancer. Nature 585, 447–452 (2020).
Ippolito, M. R. et al. Increased RNA and protein degradation is required for counteracting transcriptional burden and proteotoxic stress in human aneuploid cells. Preprint at bioRxiv https://doi.org/10.1101/2023.01.27.525826 (2023).
Zerbib, J. et al. Human aneuploid cells depend on the RAF/MEK/ERK pathway for overcoming increased DNA damage. Preprint at bioRxiv https://doi.org/10.1101/2023.01.27.525822 (2023).
Flynn, P. J., Koch, P. D. & Mitchison, T. J. Chromatin bridges, not micronuclei, activate cGAS after drug-induced mitotic errors in human cells. Proc. Natl Acad. Sci. USA 118, e2103585118 (2021).
Bakhoum, S. F. Targeting the undruggable. Science 380, 47 (2023).
Tang, S., Stokasimov, E., Cui, Y. & Pellman, D. Breakage of cytoplasmic chromosomes by pathological DNA base excision repair. Nature 606, 930–936 (2022).
Hatch, E. M., Fischer, A. H., Deerinck, T. J. & Hetzer, M. W. Catastrophic nuclear envelope collapse in cancer cell micronuclei. Cell 154, 47–60 (2013).
Dou, Z. et al. Cytoplasmic chromatin triggers inflammation in senescence and cancer. Nature 550, 402–406 (2017).
Gluck, S. et al. Innate immune sensing of cytosolic chromatin fragments through cGAS promotes senescence. Nat. Cell. Biol. 19, 1061–1070 (2017).
Sun, L., Wu, J., Du, F., Chen, X. & Chen, Z. J. Cyclic GMP-AMP synthase is a cytosolic DNA sensor that activates the type I interferon pathway. Science 339, 786–791 (2013).
Wu, J. et al. Cyclic GMP-AMP is an endogenous second messenger in innate immune signaling by cytosolic DNA. Science 339, 826–830 (2013).
Liu, S. et al. Phosphorylation of innate immune adaptor proteins MAVS, STING, and TRIF induces IRF3 activation. Science 347, aaa2630 (2015).
Fitzgerald, K. A. et al. IKKε and TBK1 are essential components of the IRF3 signaling pathway. Nat. Immunol. 4, 491–496 (2003).
Sharma, S. et al. Triggering the interferon antiviral response through an IKK-related pathway. Science 300, 1148–1151 (2003).
Carozza, J. A. et al. Extracellular cGAMP is a cancer cell-produced immunotransmitter involved in radiation-induced anti-cancer immunity. Nat. Cancer 1, 184–196 (2020).
Li, X., Wang, F., Xu, X., Zhang, J. & Xu, G. The dual role of STAT1 in ovarian cancer: insight into molecular mechanisms and application potentials. Front. Cell Dev. Biol. 9, 636595 (2021).
Dunn, G. P. et al. A critical function for type I interferons in cancer immunoediting. Nat. Immunol. 6, 722–729 (2005).
Jin, J. et al. Noncanonical NF-κB pathway controls the production of type I interferons in antiviral innate immunity. Immunity 40, 342–354 (2014).
Xu, Y. et al. SN52, a novel nuclear factor-κB inhibitor, blocks nuclear import of RelB:p52 dimer and sensitizes prostate cancer cells to ionizing radiation. Mol. Cancer Ther. 7, 2367–2376 (2008).
Yamada, T. et al. Constitutive aryl hydrocarbon receptor signaling constrains type I interferon-mediated antiviral innate defense. Nat. Immunol. 17, 687–694 (2016).
Toufektchan, E. et al. Intratumoral TREX1 induction promotes immune evasion by limiting type I interferon. Cancer Immunol. Res. 12, 673–686 (2024).
Yap, T. A. et al. 27O First-in-class first-in-human phase I trial of RBN-2397 in patients with advanced solid tumors validates PARP7 as a novel anticancer therapeutic target. ESMO Open 8, 100993 (2023).
Gozgit, J. M. et al. PARP7 negatively regulates the type I interferon response in cancer cells and its inhibition triggers antitumor immunity. Cancer Cell 39, 1214–1226.e10 (2021).
Vanpouille-Box, C. et al. DNA exonuclease Trex1 regulates radiotherapy-induced tumour immunogenicity. Nat. Commun. 8, 15618 (2017).
Francica, B. et al. Abstract 2075: systemic small molecule TREX1 inhibitors to selectively activate STING in the TME of metastatic disease. Cancer Res. 82, 2075–2075 (2022).
Tani, T. et al. TREX1 inactivation unleashes cancer cell STING-interferon signaling and promotes anti-tumor immunity. Cancer Discov. 14, 752–765 (2024).
Zimmerli, D. et al. MYC promotes immune-suppression in triple-negative breast cancer via inhibition of interferon signaling. Nat. Commun. 13, 6579 (2022).
Muthalagu, N. et al. Repression of the type I interferon pathway underlies MYC- and KRAS-dependent evasion of NK and B cells in pancreatic ductal adenocarcinoma. Cancer Discov. 10, 872–887 (2020).
Li, Q. et al. Therapeutic development by targeting the cGAS-STING pathway in autoimmune disease and cancer. Front Pharm. 12, 779425 (2021).
Wang, Y. et al. Aneuploidy landscape in precursors of ovarian cancer. Clin. Cancer Res. 30, 600–615 (2023).
Carozza, J. A. et al. ENPP1’s regulation of extracellular cGAMP is a ubiquitous mechanism of attenuating STING signaling. Proc. Natl Acad. Sci. USA 119, e2119189119 (2022).
Cogan, D. & Bakhoum, S. F. Re-awakening innate immune signaling in cancer: the development of highly potent ENPP1 inhibitors. Cell Chem. Biol. 27, 1327–1328 (2020).
Solomon, P. E. et al. Discovery of VH domains that allosterically inhibit ENPP1. Nat. Chem. Biol. 20, 30–41 (2024).
Csiki, I. et al. 195TiP A first-in-human, phase I a/b dose escalation and expansion study to evaluate RBS2418 as monotherapy and in combination with pembrolizumab in subjects with advanced unresectable, recurrent or metastatic tumors. Immuno-Oncol. Technol. 16, 100307 (2022).
Thompson, E. A. & Powell, J. D. Inhibition of the adenosine pathway to potentiate cancer immunotherapy: potential for combinatorial approaches. Annu. Rev. Med. 72, 331–348 (2021).
Decout, A., Katz, J. D., Venkatraman, S. & Ablasser, A. The cGAS-STING pathway as a therapeutic target in inflammatory diseases. Nat. Rev. Immunol. 21, 548–569 (2021).
Li, J. & Bakhoum, S. F. The pleiotropic roles of cGAS-STING signaling in the tumor microenvironment. J. Mol. Cell. Biol. 14, mjac019 (2022).
Hong, C. et al. cGAS-STING drives the IL-6-dependent survival of chromosomally instable cancers. Nature 607, 366–373 (2022).
Laks, E. et al. Clonal decomposition and DNA replication states defined by scaled single-cell genome sequencing. Cell 179, 1207–1221.e22 (2019).
Funnell, T. et al. Single-cell genomic variation induced by mutational processes in cancer. Nature 612, 106–115 (2022).
Zhou, H. et al. Plasma cell-free DNA chromosomal instability analysis by low-pass whole-genome sequencing to monitor breast cancer relapse. Breast Cancer Res. Treat. 178, 63–73 (2019).
Christodoulou, E. et al. Combined low-pass whole genome and targeted sequencing in liquid biopsies for pediatric solid tumors. NPJ Precis. Oncol. 7, 21 (2023).
Bethune, G. et al. Impact of the 2013 American Society of Clinical Oncology/College of American Pathologists guideline recommendations for human epidermal growth factor receptor 2 (HER2) testing of invasive breast carcinoma: a focus on tumours assessed as ‘equivocal’ for HER2 gene amplification by fluorescence in-situ hybridization. Histopathology. 67, 880–887 (2015).
Akkari, Y. M. N. et al. Guiding the global evolution of cytogenetic testing for hematologic malignancies. Blood 139, 2273–2284 (2022).
Bakhoum, S. F., Danilova, O. V., Kaur, P., Levy, N. B. & Compton, D. A. Chromosomal instability substantiates poor prognosis in patients with diffuse large B-cell lymphoma. Clin. Cancer Res. 17, 7704–7711 (2011).
Xu, Z. et al. Deep learning predicts chromosomal instability from histopathology images. IScience 24, 102394 (2021).
Negoto, T. et al. Profiling chromosomal-level variations in gastric malignancies. Cancer Sci. 113, 3864–3876 (2022).
Matsuura, T. et al. Histological diagnosis of polyploidy discriminates an aggressive subset of hepatocellular carcinomas with poor prognosis. Br. J. Cancer 129, 1251–1260 (2023).
Paniagua, I. & Jacobs, J. J. L. Quantification of chromosomal aberrations in mammalian cells. Bio-Protocol 13, e4739 (2023).
Jones, G. D. et al. A genomic-pathologic annotated risk model to predict recurrence in early-stage lung adenocarcinoma. JAMA Surg. 156, e205601 (2021).
Tsang, E. S. et al. Homologous recombination deficiency signatures in gastrointestinal and thoracic cancers correlate with platinum therapy duration. NPJ Precis. Oncol. 7, 31 (2023).
Timms, K. M. et al. Association of BRCA1/2 defects with genomic scores predictive of DNA damage repair deficiency among breast cancer subtypes. Breast Cancer Res. 16, 475 (2014).
Vazquez-Garcia, I. et al. Ovarian cancer mutational processes drive site-specific immune evasion. Nature 612, 778–786 (2022).
Watkins, T. B. K. et al. Pervasive chromosomal instability and karyotype order in tumour evolution. Nature 587, 126–132 (2020).
Frezza, A. M. et al. CINSARC in high-risk soft tissue sarcoma patients treated with neoadjuvant chemotherapy: results from the ISG-STS 1001 study. Cancer Med. 12, 1350–1357 (2023).
Lynch, A. R., Arp, N. L., Zhou, A. S., Weaver, B. A. & Burkard, M. E. Quantifying chromosomal instability from intratumoral karyotype diversity using agent-based modeling and Bayesian inference. eLife 11, e69799 (2022).
Agustinus, A. S. et al. Epigenetic dysregulation from chromosomal transit in micronuclei. Nature 619, 176–183 (2023).
Coy, S. et al. 2D and 3D multiplexed subcellular profiling of nuclear instability in human cancer. Preprint at bioRxiv https://doi.org/10.1101/2023.11.07.566063 (2023).
Schonhoft, J. D. et al. Morphology-predicted large-scale transition number in circulating tumor cells identifies a chromosomal instability biomarker associated with poor outcome in castration-resistant prostate cancer. Cancer Res. 80, 4892–4903 (2020).
Freitas, M. O., Gartner, J., Rangel-Pozzo, A. & Mai, S. Genomic instability in circulating tumor cells. Cancers 12, 3001 (2020).
Chen, Z. et al. Chromosomal instability of circulating tumor DNA reflect therapeutic responses in advanced gastric cancer. Cell Death Dis. 10, 697 (2019).
Tamayo, N. A. et al. Targeting the mitotic kinesin KIF18A in chromosomally unstable cancers: hit optimization toward an in vivo chemical probe. J. Med. Chem. 65, 4972–4990 (2022).
Di Bona, M. & Bakhoum, S. F. Micronuclei and cancer. Cancer Discov. 14, 214–226 (2024).
Crasta, K. et al. DNA breaks and chromosome pulverization from errors in mitosis. Nature 482, 53–58 (2012).
Liu, S. et al. Nuclear envelope assembly defects link mitotic errors to chromothripsis. Nature 561, 551–555 (2018).
Maciejowski, J., Li, Y., Bosco, N., Campbell, P. J. & de Lange, T. Chromothripsis and kataegis induced by telomere crisis. Cell 163, 1641–1654 (2015).
Ly, P. et al. Chromosome segregation errors generate a diverse spectrum of simple and complex genomic rearrangements. Nat. Genet. 51, 705–715 (2019).
Shoshani, O. et al. Chromothripsis drives the evolution of gene amplification in cancer. Nature 591, 137–141 (2021).
Umbreit, N. T. et al. Mechanisms generating cancer genome complexity from a single cell division error. Science 368, eaba0712 (2020).
Acknowledgements
We thank all members of the Bakhoum laboratory for discussion and comments. We thank the following sources of funding: for S.F.B., National Institutes of Health/National Cancer Institute (grant nos. P50CA247749, DP5OD026395, R01CA256188, R01CA280572, P30CA008748) Department of Defense Era of Hope Award, Burroughs Wellcome Fund, the Pershing Square Sohn Foundation for Cancer Research, the Josie Robertson Foundation, the Mark Foundation for Cancer Research, and the Starr Cancer Consortium; for D.H.A.-R., Mary Kay Ash Foundation, OCRA Mentored Research Award, 1L32MD017781-01, Conquer Cancer Foundation, and Young Investigator Award.
Author information
Authors and Affiliations
Contributions
All authors researched data for this article and made a substantial contribution to discussions of content. D.H.A.-R., E.L., J.L. and S.F.B. wrote the manuscript. All authors reviewed and/or edited the manuscript prior to submission.
Corresponding author
Ethics declarations
Competing interests
S.F.B. has acted as a consultant and/or adviser of Meliora Therapeutics and Volastra Therapeutics, is a co-inventor on patents related to some of the research described in this manuscript on agents targeting chromosomal instability and the cGAS–STING pathway in patients with advanced-stage cancer (patent numbers WO2019014246A1, US20210130903A1 and EP4208572A1), and is the scientific co-founder of, holds equity in, and receives compensation from Volastra Therapeutics. The other authors declare no competing interests.
Peer review
Peer review information
Nature Reviews Clinical Oncology thanks T. Hirota and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Al-Rawi, D.H., Lettera, E., Li, J. et al. Targeting chromosomal instability in patients with cancer. Nat Rev Clin Oncol 21, 645–659 (2024). https://doi.org/10.1038/s41571-024-00923-w
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41571-024-00923-w
- Springer Nature Limited