Abstract
Methane emissions are mitigated by anaerobic methane-oxidizing archaea, including Methanoperedens. Some Methanoperedens host huge extrachromosomal genetic elements (ECEs) called Borgs that may modulate their activity, yet the broader diversity of Methanoperedens ECEs is understudied. Here we report small enigmatic linear ECEs, circular viruses and unclassified ECEs that are predicted to replicate within Methanoperedens. Linear ECEs have inverted terminal repeats, tandem repeats and coding patterns that are strongly reminiscent of Borgs, but they are only 52–145 kb in length. As they share proteins with Borgs and Methanoperedens, we refer to them as mini-Borgs. Mini-Borgs are genetically diverse and can be assigned to at least five family-level groups. We identify eight families of Methanoperedens viruses, some of which encode multi-haem cytochromes, and circular ECEs encoding transposon-associated TnpB genes with proximal population-heterogeneous CRISPR arrays. These ECEs exchange genetic information with each other and with Methanoperedens, probably impacting their archaeal host activity and evolution.





Similar content being viewed by others
Data availability
Metagenomic sequencing reads related to Methanoperedens ECEs are available under NCBI BioProject PRJNA999944. The genomes described in this study can be accessed at https://ggkbase.berkeley.edu/project_groups/methanoperedens_ece. Please note that it is necessary to sign up as a user (simply provide an email address) to download the data. We also submit genomes to figshare (https://doi.org/10.6084/m9.figshare.24481222)90. Reference protein sequences used in this study can be found in NCBI (https://www.ncbi.nlm.nih.gov/), KEGG (https://www.genome.jp/kegg/) and UniProt databases (https://www.uniprot.org/). Protein structures (for example, 6R10) can be accessed at RCSB PDB (https://www.rcsb.org/). Source data are provided with this paper.
References
Thauer, R. K., Kaster, A. K., Seedorf, H., Buckel, W. & Hedderich, R. Methanogenic archaea: ecologically relevant differences in energy conservation. Nat. Rev. Microbiol. 6, 579–591 (2008).
Evans, P. N. et al. An evolving view of methane metabolism in the Archaea. Nat. Rev. Microbiol. 17, 219–232 (2019).
Boetius, A. et al. A marine microbial consortium apparently mediating anaerobic oxidation of methane. Nature 407, 623–626 (2000).
Orphan, V. J., House, C. H., Hinrichs, K. U., McKeegan, K. D. & DeLong, E. F. Methane-consuming archaea revealed by directly coupled isotopic and phylogenetic analysis. Science 293, 484–487 (2001).
Niemann, H. et al. Novel microbial communities of the Haakon Mosby mud volcano and their role as a methane sink. Nature 443, 854–858 (2006).
Haroon, M. F. et al. Anaerobic oxidation of methane coupled to nitrate reduction in a novel archaeal lineage. Nature 500, 567–570 (2013).
Raghoebarsing, A. A. et al. A microbial consortium couples anaerobic methane oxidation to denitrification. Nature 440, 918–921 (2006).
Cai, C. et al. A methanotrophic archaeon couples anaerobic oxidation of methane to Fe(III) reduction. ISME J. 12, 1929–1939 (2018).
Leu, A. O. et al. Anaerobic methane oxidation coupled to manganese reduction by members of the Methanoperedenaceae. Isme J. 14, 1030–1041 (2020).
Leu, A. O. et al. Lateral gene transfer drives metabolic flexibility in the anaerobic methane-oxidizing archaeal family Methanoperedenaceae. Mbio 11, e01325-20 (2020).
Shi, L. D. et al. Coupled anaerobic methane oxidation and reductive arsenic mobilization in wetland soils. Nat. Geosci. 13, 799–805 (2020).
Al-Shayeb, B. et al. Borgs are giant genetic elements with potential to expand metabolic capacity. Nature 610, 731–736 (2022).
Schoelmerich, M. C. et al. A widespread group of large plasmids in methanotrophic Methanoperedens archaea. Nat. Commun. 13, 7085 (2022).
Schoelmerich, M. C. et al. Borg extrachromosomal elements of methane-oxidizing archaea have conserved and expressed genetic repertoires. Preprint at bioRxiv, https://doi.org/10.1101/2023.08.01.549754 (2023).
Chen, L. X. et al. Large freshwater phages with the potential to augment aerobic methane oxidation. Nat. Microbiol. 5, 1504–1515 (2020).
McIlroy, S. J. et al. Anaerobic methanotroph ‘Candidatus Methanoperedens nitroreducens’ has a pleomorphic life cycle. Nat. Microbiol. 8, 321–331 (2023).
Schoelmerich, M. C., Sachdeva, R., West-Roberts, J., Waldburger, L. & Banfield, J. F. Tandem repeats in giant archaeal Borg elements undergo rapid evolution and create new intrinsically disordered regions in proteins. PLoS Biol. 21, e3001980 (2023).
Brown, C. T., Olm, M. R., Thomas, B. C. & Banfield, J. F. Measurement of bacterial replication rates in microbial communities. Nat. Biotechnol. 34, 1256–1263 (2016).
Peeva, V. et al. Linear mitochondrial DNA is rapidly degraded by components of the replication machinery. Nat. Commun. 9, 1727 (2018).
Zhou, L. & Sazanov, L. A. Structure and conformational plasticity of the intact Thermus thermophilus V/A-type ATPase. Science 365, eaaw9144 (2019).
Santos-Pérez, I. et al. Structural basis for assembly of vertical single β-barrel viruses. Nat. Commun. 10, 1184 (2019).
Krupovic, M., Makarova, K. S. & Koonin, E. V. Cellular homologs of the double jelly-roll major capsid proteins clarify the origins of an ancient virus kingdom. Proc. Natl Acad. Sci. USA 119, e2120620119 (2022).
Camargo, A. P. et al. Identification of mobile genetic elements with geNomad. Nat. Biotechnol. https://doi.org/10.1038/s41587-023-01953-y (2023).
Hernsdorf, A. W. et al. Potential for microbial H2 and metal transformations associated with novel bacteria and archaea in deep terrestrial subsurface sediments. ISME J. https://doi.org/10.1038/ismej.2017.39 (2017).
Camargo, A. P. et al. IMG/VR v4: an expanded database of uncultivated virus genomes within a framework of extensive functional, taxonomic, and ecological metadata. Nucleic Acids Res. https://doi.org/10.1093/nar/gkac1037 (2022).
Laso-Perez, R. et al. Evolutionary diversification of methanotrophic ANME-1 archaea and their expansive virome. Nat. Microbiol. https://doi.org/10.1038/s41564-022-01297-4 (2023).
Shiimori, M., Garrett, S. C., Graveley, B. R. & Terns, M. P. Cas4 nucleases define the PAM, length, and orientation of DNA fragments integrated at CRISPR Loci. Mol. Cell 70, 814–824 (2018).
Zhang, Z. F., Pan, S. F., Liu, T., Li, Y. J. & Peng, N. Cas4 nucleases can effect specific integration of CRISPR spacers. J. Bacteriol. 201, e00747-18 (2019).
Hooton, S. P. T. & Connerton, I. F. Campylobacter jejuni acquire new host-derived CRISPR spacers when in association with bacteriophages harboring a CRISPR-like Cas4 protein. Front. Microbiol. 5, 744 (2015).
Ino, K. et al. Ecological and genomic profiling of anaerobic methane-oxidizing archaea in a deep granitic environment. ISME J. https://doi.org/10.1038/ismej.2017.140 (2017).
Pinilla-Redondo, R. et al. Type IV CRISPR Cas systems are highly diverse and involved in competition between plasmids. Nucleic Acids Res. 48, 2000–2012 (2020).
Zhou, Y. et al. Structure of a type IV CRISPR–Cas ribonucleoprotein complex. iScience 24, 102201 (2021).
Taylor, H. N. et al. Positioning diverse type IV structures and functions within class 1 CRISPR–Cas systems. Front. Microbiol. 12, 671522 (2021).
Kletzin, A. et al. Cytochromes c in Archaea: distribution, maturation, cell architecture, and the special case of Ignicoccus hospitalis. Front. Microbiol. 6, 439 (2015).
Edwards, M. J., Richardson, D. J., Paquete, C. M. & Clarke, T. A. Role of multiheme cytochromes involved in extracellular anaerobic respiration in bacteria. Protein Sci. 29, 830–842 (2020).
Baquero, D. P. et al. Extracellular cytochrome nanowires appear to be ubiquitous in prokaryotes. Cell https://doi.org/10.1016/j.cell.2023.05.012 (2023).
Jumper, J. et al. Highly accurate protein structure prediction with AlphaFold. Nature 596, 583–589 (2021).
Tyson, G. W. & Banfield, J. F. Rapidly evolving CRISPRs implicated in acquired resistance of microorganisms to viruses. Environ. Microbiol. 10, 200–207 (2008).
Chadwick, G. L. et al. Comparative genomics reveals electron transfer and syntrophic mechanisms differentiating methanotrophic and methanogenic archaea. PLoS Biol. 20, e3001508 (2022).
Costi, M. P. et al. Thymidylate synthase structure, function and implication in drug discovery. Curr. Med. Chem. 12, 2241–2258 (2005).
Manandhar, M., Boulware, K. S. & Wood, R. D. The ERCC1 and ERCC4 (XPF) genes and gene products. Gene 569, 153–161 (2015).
Johnson, A., Yao, N. Y., Bowman, G. D., Kuriyan, J. & O’Donnell, M. The replication factor C clamp loader requires arginine finger sensors to drive DNA binding and proliferating cell nuclear antigen loading. J. Biol. Chem. 281, 35531–35543 (2006).
Seybert, A., Scott, D. J., Scaife, S., Singleton, M. R. & Wigley, D. B. Biochemical characterisation of the clamp/clamp loader proteins from the euryarchaeon Archaeoglobus fulgidus. Nucleic Acids Res. 30, 4329–4338 (2002).
Kelman, Z. & Hurwitz, J. A unique organization of the protein subunits of the DNA polymerase clamp loader in the archaeon Methanobacterium thermoautotrophicum Delta H. J. Biol. Chem. 275, 7327–7336 (2000).
Mardanov, A. V., Kadnikov, V. V., Beletsky, A. V. & Ravin, N. V. Sulfur and methane-oxidizing microbial community in a terrestrial mud volcano revealed by metagenomics. Microorganisms 8, 1333 (2020).
Lindell, D., Jaffe, J. D., Johnson, Z. I., Church, G. M. & Chisholm, S. W. Photosynthesis genes in marine viruses yield proteins during host infection. Nature 438, 86–89 (2005).
Al-Shayeb, B. et al. Clades of huge phages from across Earth’s ecosystems. Nature 578, 425–431 (2020).
Shmakov, S. et al. Diversity and evolution of class 2 CRISPR–Cas systems. Nat. Rev. Microbiol. 15, 169–182 (2017).
Sasnauskas, G. et al. TnpB structure reveals minimal functional core of Cas12 nuclease family. Nature 616, 384–389 (2023).
Altae-Tran, H. et al. The widespread IS200/IS605 transposon family encodes diverse programmable RNA-guided endonucleases. Science 374, 57–65 (2021).
Karvelis, T. et al. Transposon-associated TnpB is a programmable RNA-guided DNA endonuclease. Nature 599, 692–696 (2021).
Chen, L. X., Anantharaman, K., Shaiber, A., Eren, A. M. & Banfield, J. F. Accurate and complete genomes from metagenomes. Genome Res. 30, 315–333 (2020).
Valentin-Alvarado, L. E. et al. Asgard archaea modulate potential methanogenesis substrates in wetland soil. Preprint at bioRxiv https://doi.org/10.1101/2023.11.21.568159 (2023).
Hyatt, D. et al. Prodigal: prokaryotic gene recognition and translation initiation site identification. BMC Bioinformatics 11, 119 (2010).
Meheust, R., Burstein, D., Castelle, C. J. & Banfield, J. F. The distinction of CPR bacteria from other bacteria based on protein family content. Nat. Commun. 10, 4173 (2019).
Oksanen, J. et al. vegan: Community Ecology Package. R package version 2.6-4 https://CRAN.R-project.org/package=vegan (R Foundation for Statistical Computing, 2022).
R Core Team R: A Language and Environment for Statistical Computing (R Foundation for Statistical Computing, 2022); https://www.R-project.org/
Gu, Z. Complex heatmap visualization. Imeta 1, e43 (2022).
Nicolas, A. M. et al. A subset of viruses thrives following microbial resuscitation during rewetting of a seasonally dry California grassland soil. Nat. Commun. 14, 5835 (2023).
Olm, M. R., Brown, C. T., Brooks, B. & Banfield, J. F. dRep: a tool for fast and accurate genomic comparisons that enables improved genome recovery from metagenomes through de-replication. ISME J. 11, 2864–2868 (2017).
Harrell, J. F. Hmisc: Harrell Miscellaneous. R package version 5.0-1 R Foundation for Statistical Computing https://CRAN.R-project.org/package=Hmisc (2023).
Russel, J., Pinilla-Redondo, R., Mayo-Munoz, D., Shah, S. A. & Sorensen, S. J. CRISPRCasTyper: automated identification, annotation, and classification of CRISPR–Cas loci. CRISPR J. 3, 462–469 (2020).
Boratyn, G. M. et al. BLAST: a more efficient report with usability improvements. Nucleic Acids Res. 41, W29–W33 (2013).
Ahlgren, N. A., Ren, J., Lu, Y. Y., Fuhrman, J. A. & Sun, F. Z. Alignment-free d2* oligonucleotide frequency dissimilarity measure improves prediction of hosts from metagenomically-derived viral sequences. Nucleic Acids Res. 45, 39–53 (2017).
Galiez, C., Siebert, M., Enault, F., Vincent, J. & Soding, J. WIsH: who is the host? Predicting prokaryotic hosts from metagenomic phage contigs. Bioinformatics 33, 3113–3114 (2017).
Lu, C. et al. Prokaryotic virus host predictor: a Gaussian model for host prediction of prokaryotic viruses in metagenomics. BMC Biol. 19, 5 (2021).
Coutinho, F. H. et al. RaFAH: host prediction for viruses of Bacteria and Archaea based on protein content. Patterns 2, 100274 (2021).
Roux, S. et al. iPHoP: an integrated machine learning framework to maximize host prediction for metagenome-derived viruses of archaea and bacteria. PLoS Biol. 21, e3002083 (2023).
Jang, H. B. et al. Taxonomic assignment of uncultivated prokaryotic virus genomes is enabled by gene-sharing networks. Nat. Biotechnol. 37, 632–639 (2019).
Medvedeva, S. et al. Three families of Asgard archaeal viruses identified in metagenome-assembled genomes. Nat. Microbiol. 7, 962–973 (2022).
Shannon, P. et al. Cytoscape: a software environment for integrated models of biomolecular interaction networks. Genome Res. 13, 2498–2504 (2003).
Nishimura, Y. et al. ViPTree: the viral proteomic tree server. Bioinformatics 33, 2379–2380 (2017).
Gilchrist, C. L. M. & Chooi, Y. H. clinker & clustermap.js: automatic generation of gene cluster comparison figures. Bioinformatics 37, 2473–2475 (2021).
Seemann, T. Prokka: rapid prokaryotic genome annotation. Bioinformatics 30, 2068–2069 (2014).
Kanehisa, M., Furumichi, M., Sato, Y., Kawashima, M. & Ishiguro-Watanabe, M. KEGG for taxonomy-based analysis of pathways and genomes. Nucleic Acids Res. https://doi.org/10.1093/nar/gkac963 (2022).
Bateman, A. et al. UniProt: the Universal Protein Knowledgebase in 2023. Nucleic Acids Res. https://doi.org/10.1093/nar/gkac1052 (2022).
Edgar, R. C. Search and clustering orders of magnitude faster than BLAST. Bioinformatics 26, 2460–2461 (2010).
Benson, D. A. et al. GenBank. Nucleic Acids Res. 41, D36–D42 (2012).
Blum, M. et al. The InterPro protein families and domains database: 20 years on. Nucleic Acids Res. 49, D344–D354 (2021).
Jones, P. et al. InterProScan 5: genome-scale protein function classification. Bioinformatics 30, 1236–1240 (2014).
Yu, N. Y. et al. PSORTb 3.0: improved protein subcellular localization prediction with refined localization subcategories and predictive capabilities for all prokaryotes. Bioinformatics 26, 1608–1615 (2010).
Katoh, K., Misawa, K., Kuma, K. & Miyata, T. MAFFT: a novel method for rapid multiple sequence alignment based on fast Fourier transform. Nucleic Acids Res. 30, 3059–3066 (2002).
Capella-Gutierrez, S., Silla-Martinez, J. M. & Gabaldon, T. trimAl: a tool for automated alignment trimming in large-scale phylogenetic analyses. Bioinformatics 25, 1972–1973 (2009).
Nguyen, L. T., Schmidt, H. A., von Haeseler, A. & Minh, B. Q. IQ-TREE: a fast and effective stochastic algorithm for estimating maximum-likelihood phylogenies. Mol. Biol. Evol. 32, 268–274 (2015).
Letunic, I. & Bork, P. Interactive Tree Of Life (iTOL) v5: an online tool for phylogenetic tree display and annotation. Nucleic Acids Res. 49, W293–W296 (2021).
Mirdita, M. et al. ColabFold: making protein folding accessible to all. Nat. Methods 19, 679–682 (2022).
van Kempen, M. et al. Fast and accurate protein structure search with Foldseek. Nat. Biotechnol. https://doi.org/10.1038/s41587-023-01773-0 (2023).
DeLano, W. L. The PyMOL molecular graphics system. Schrödinger http://www.pymol.org (2002).
Meng, E. C. et al. UCSF ChimeraX: tools for structure building and analysis. Protein Sci. 32, e4792 (2023).
Shi, L.-D. Materials involved in the Methanoperedens ECEs paper. figshare https://doi.org/10.6084/m9.figshare.24481222 (2024).
Acknowledgements
We thank J. A. Doudna and B. Adler for helpful discussion of the TnpB-associated CRISPR complex. We also thank Adam Abate’s lab (University of California, San Francisco) for their efforts to obtain cell concentrates from soil samples, and the Oxford Nanopore research team (New York) for the efforts to perform Hi-C sequencing. Funding for this research was provided by the Bill and Melinda Gates Foundation (grant number INV-037174 to J.F.B.). The findings and conclusions are those of the authors and do not necessarily reflect positions or policies of the Bill and Melinda Gates Foundation. Funding was also provided via the Innovative Genomics Institute Climate fund philanthropic donation to J.F.B., a DFG postdoctoral fellowship to M.C.S. (project number 447383558 to M.C.S.) and The Ministry of Economy, Trade and Industry of Japan as ‘The project for validating near-field system assessment methodology in geological disposal system’ (2022 FY, grant number JPJ007597).
Author information
Authors and Affiliations
Contributions
The study was designed by L.-D.S. and J.F.B. Binning and genome curation and analysis were performed by L.-D.S. and J.F.B. Proteome and phylogenetic analyses were carried out by L.-D.S. Correlation analyses were performed by L.-D.S. J.W.-R. provided the Corona Mine sequences and computational support. M.C.S. contributed to proteome analyses, and L.-D.S. and P.I.P. carried out the protein structural analyses. L.C. contributed to TnpB and CRISPR array analyses. Y.A. provided the Horonobe metagenomic sequences, and S.L. and R.S. contributed to the data handling and bioinformatic analyses. L.-D.S. and J.F.B. wrote the paper with input from all the authors.
Corresponding author
Ethics declarations
Competing interests
J.F.B. is a co-founder of Metagenomi. The other authors declare no competing interests.
Peer review
Peer review information
Nature Microbiology thanks Marc Strous and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Genomic architectures and predicted replichores of representative complete mini-Borgs.
Yellow blocks indicate predicted genes while red and blue lines indicate predicted replication origin and terminus, respectively. Gray dots and green lines show GC skew and cumulative GC skew across genomes.
Extended Data Fig. 2 Coverage profile of representative complete and near-complete mini-Borgs.
Yellow blocks indicate predicted genes. Red arrows in the Venus_106kb_34_23_complete denote inverted terminal repeats (ITR). Mapped reads are depicted below the genomes.
Extended Data Fig. 3 Examples of tandem repeats in the Jupiter_54kb_36_306_complete genome.
Yellow blocks indicate predicted genes; red and blue blocks denote, respectively, left inverted terminal repeat and perfect tandem repeats distributed in various regions.
Extended Data Fig. 4 Structural comparison of ATPase subunit K.
a. Representative proteins from mini-Borgs, Methanoperedens, and a reference Thermus thermophilus (PDB: 6R10) were superimposed and highlighted in red, blue, and cyan, respectively. b, c. Two copies of the T. thermophilus ATPase subunit K ring were substituted by that from mini-Borg and Methanoperedens, as shown in the surface form. The surface was colored based on calculated electrostatic potential, where kb, T, e in the unit indicate the Boltzmann’s constant, temperature in Kelvin, and the charge of an electron.
Extended Data Fig. 5 Phylogeny of Borgs and mini-Borgs inferred from a concatenation of two marker proteins.
Blastp of markers against the NCBI database recruited no significant hits, therefore no other sequences are included in the tree. Support values were calculated based on 1000 replicates.
Extended Data Fig. 6 Genomic comparison of Methanoperedens-associated ECEs to other archaeal viruses and plasmids.
References were downloaded from the NCBI genome database accessed on October 30, 2023. Colored blocks indicate dissimilarity between pairs of genomes, in which the redder color is, the more similar genomes are. The original vector figure can be found in figshare under the DOI of 10.6084/m9.figshare.24481222.
Extended Data Fig. 7 An example of recombination in the Jupiter_54kb_36_306_complete genome.
The obvious dip in the coverage profile (blue area in the upper panel) suggests a region for which the more divergent reads were not recruited. Sequence blocks surrounded by colored rectangles indicate alleles that are linked in a variety of configurations, consistent with recombination involving mini-Borg variants.
Extended Data Fig. 8 Modeled structure and genomic context of mini-Borg capsid-like proteins.
a. The reference is the icosahedral asymmetric unit of the major capsid from Haloarcula californiae icosahedral virus (PDB: 6H9C). Detailed superimposition is shown for the predicted structure of a representative mini-Borg capsid-like protein from Jupiter_54kb_36_306_complete and the cryo-EM structure of a capsid VP7 protein. b. The genes encoding capsid-like proteins are highlighted and identified in all 12 complete or near-complete mini-Borgs. Coordinates listed next to the names indicate the positions within aligned regions. Typically, these regions are located in the middle of corresponding genomes (note the genome length in mini-Borg names). Genes belonging to the same group are connected if pairwise amino acid identity is > 30%, as shown by shading.
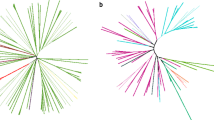
Extended Data Fig. 9 Phylogeny of TnpB transposase shared by Methanoperedens and associated ECEs.
Homologous references were recruited from the NCBI nr database. Arc colors indicate different taxonomic clades. Proteins found in Methanoperedens and associated ECEs are marked using colored dots. The tree was mid-point rerooted and support values were calculated based on 1000 replicates (only show those at the boundary of different taxonomic branches).
Supplementary information
Supplementary Information
Supplementary Figs. 1–23.
Supplementary Tables
Supplementary Tables 1–6.
Source data
Source Data Fig. 1
Plot source data.
Source Data Fig. 2
Plot source data.
Source Data Fig. 3
Original protein sequences.
Source Data Fig. 4
Original protein sequences.
Source Data Fig. 5
Plot source data.
Source Data Extended Data Fig. 4
Original protein sequences.
Source Data Extended Data Fig. 5
Original protein sequences.
Source Data Extended Data Fig. 6
Plot source data.
Source Data Extended Data Fig. 8
Original protein sequences.
Source Data Extended Data Fig. 9
Original protein sequences.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Shi, LD., West-Roberts, J., Schoelmerich, M.C. et al. Methanotrophic Methanoperedens archaea host diverse and interacting extrachromosomal elements. Nat Microbiol 9, 2422–2433 (2024). https://doi.org/10.1038/s41564-024-01740-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41564-024-01740-8
- Springer Nature Limited